Transcriptomic Revelation of Phenolic Compounds Involved in Aluminum Toxicity Responses in Roots of Cunninghamia lanceolata (Lamb.) Hook
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials and Growth Conditions
2.2. RNA Extraction, mRNA-seq Library Construction and Sequencing
2.3. De Novo Assembly and Functional Annotation
2.4. Differentially Expressed Gene (DEG) Analysis
2.5. Transcription Factor Analysis
2.6. Quantitative Real-Time PCR (qRT-PCR)
3. Results
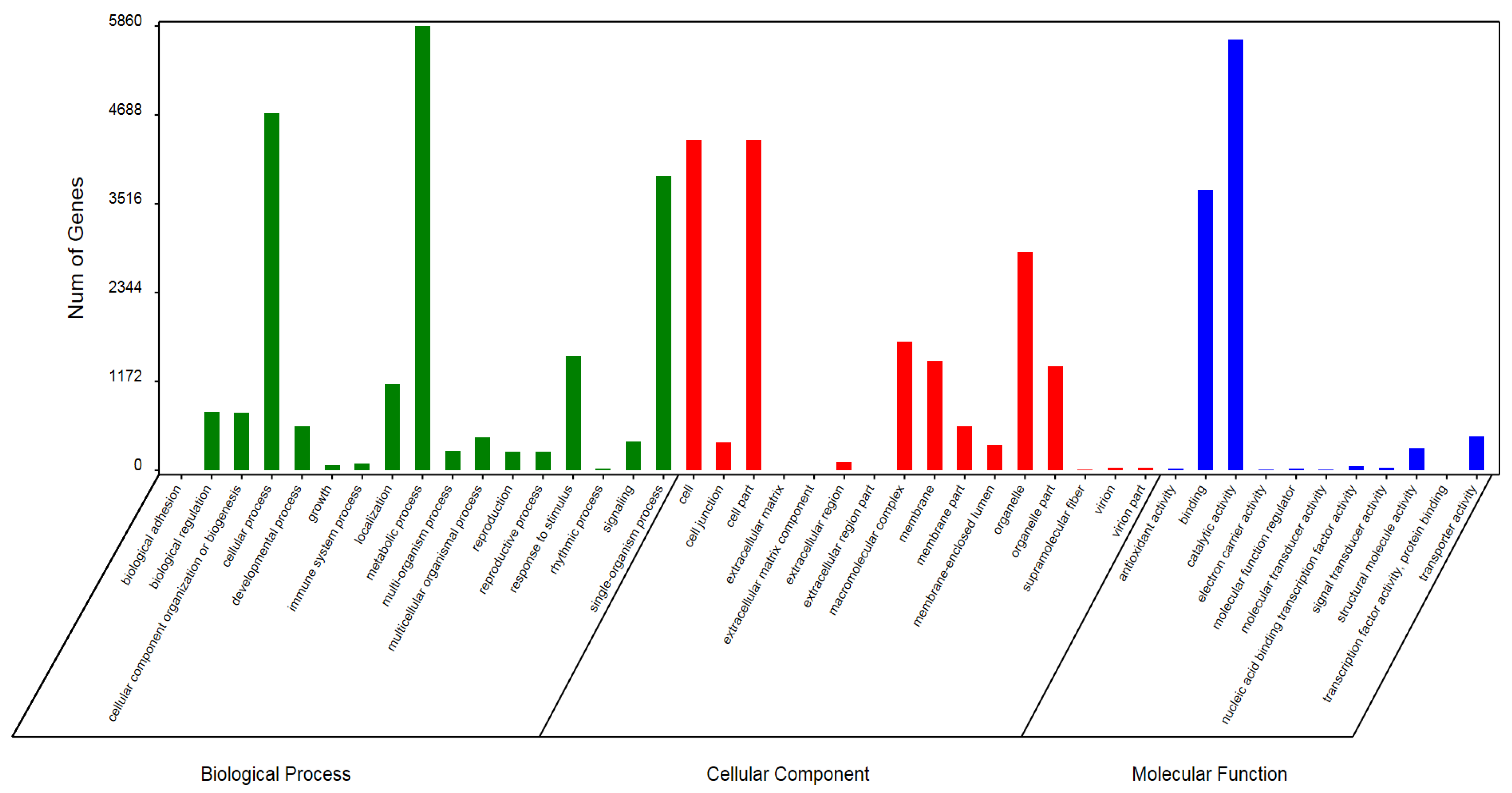
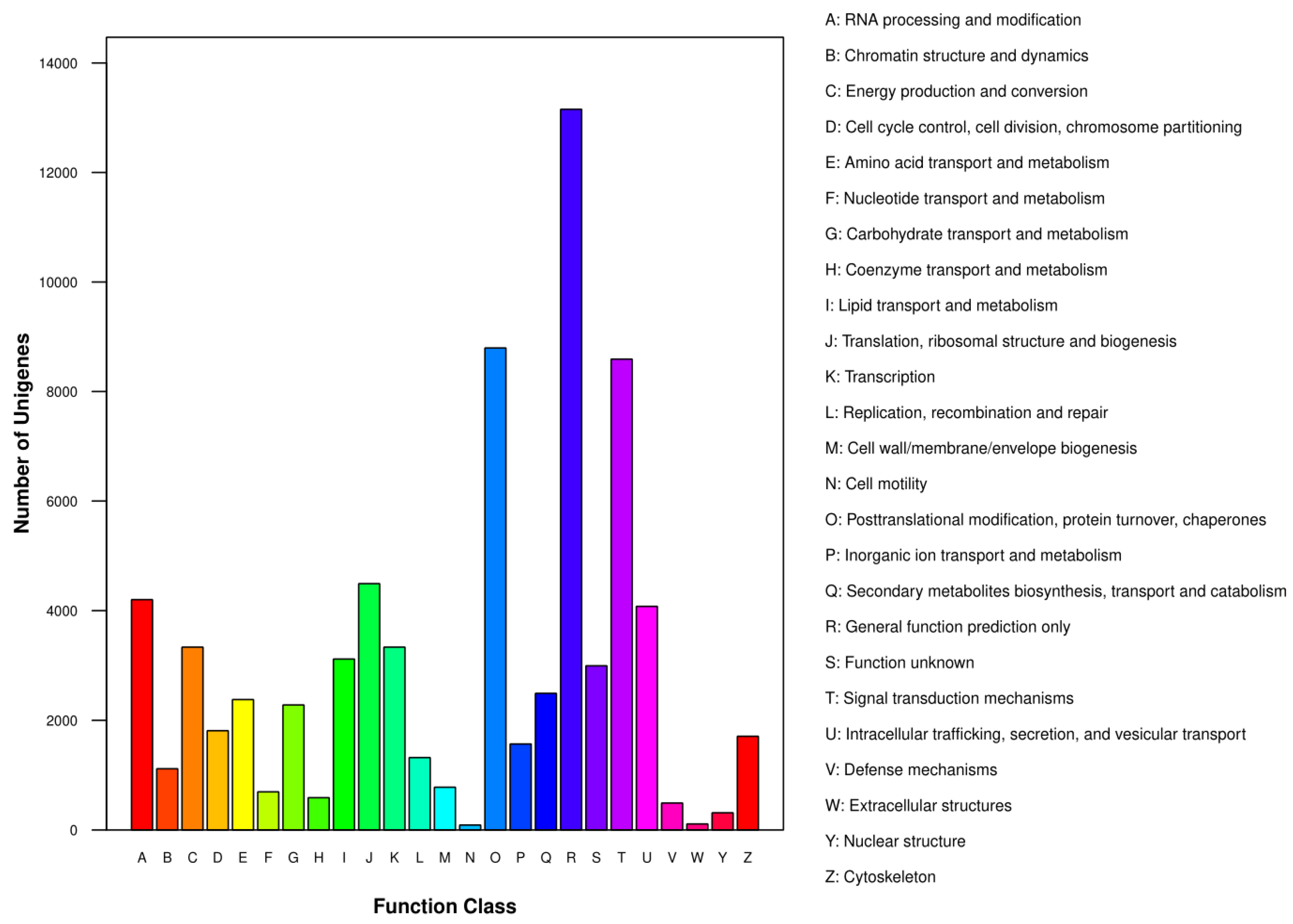
3.1. RNA Sequencing, Assembly and Functional Annotation
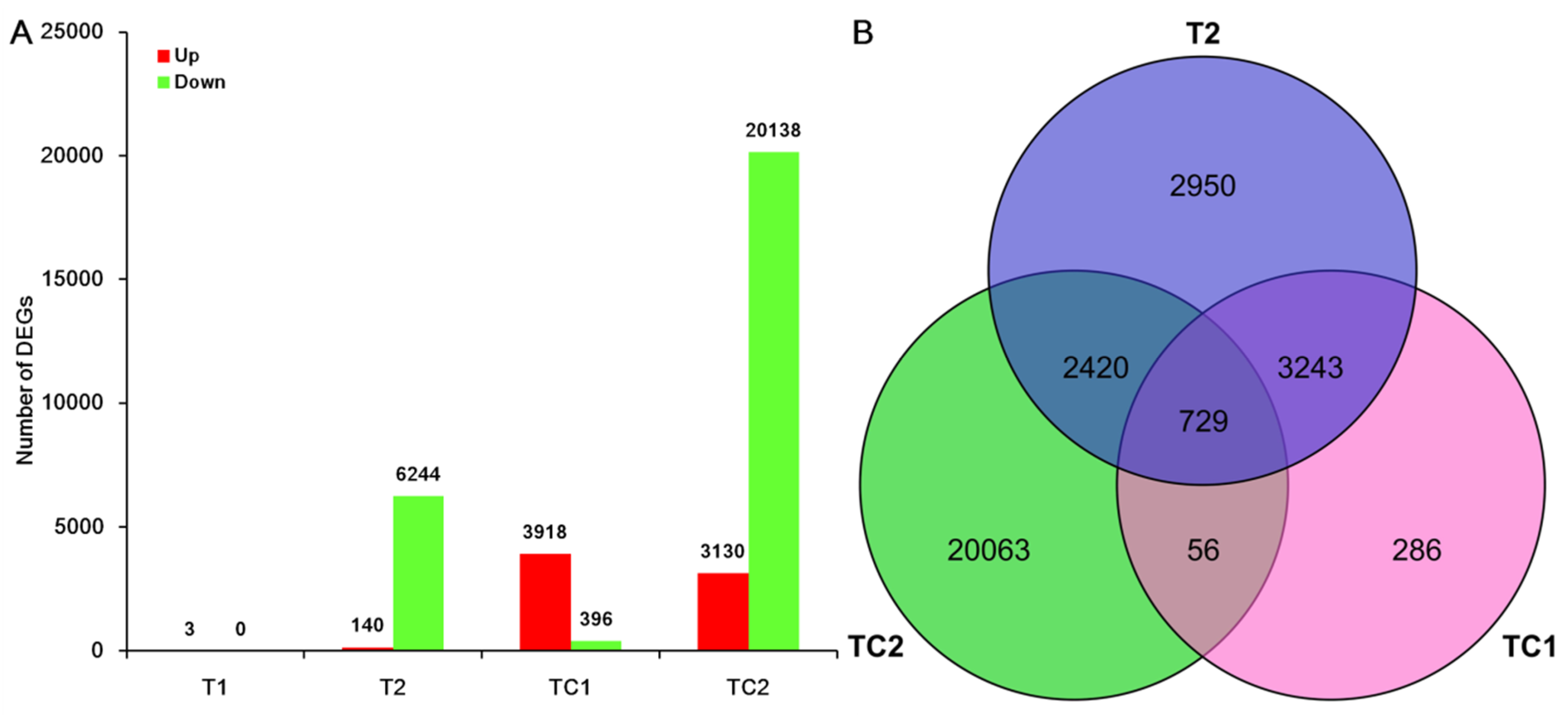
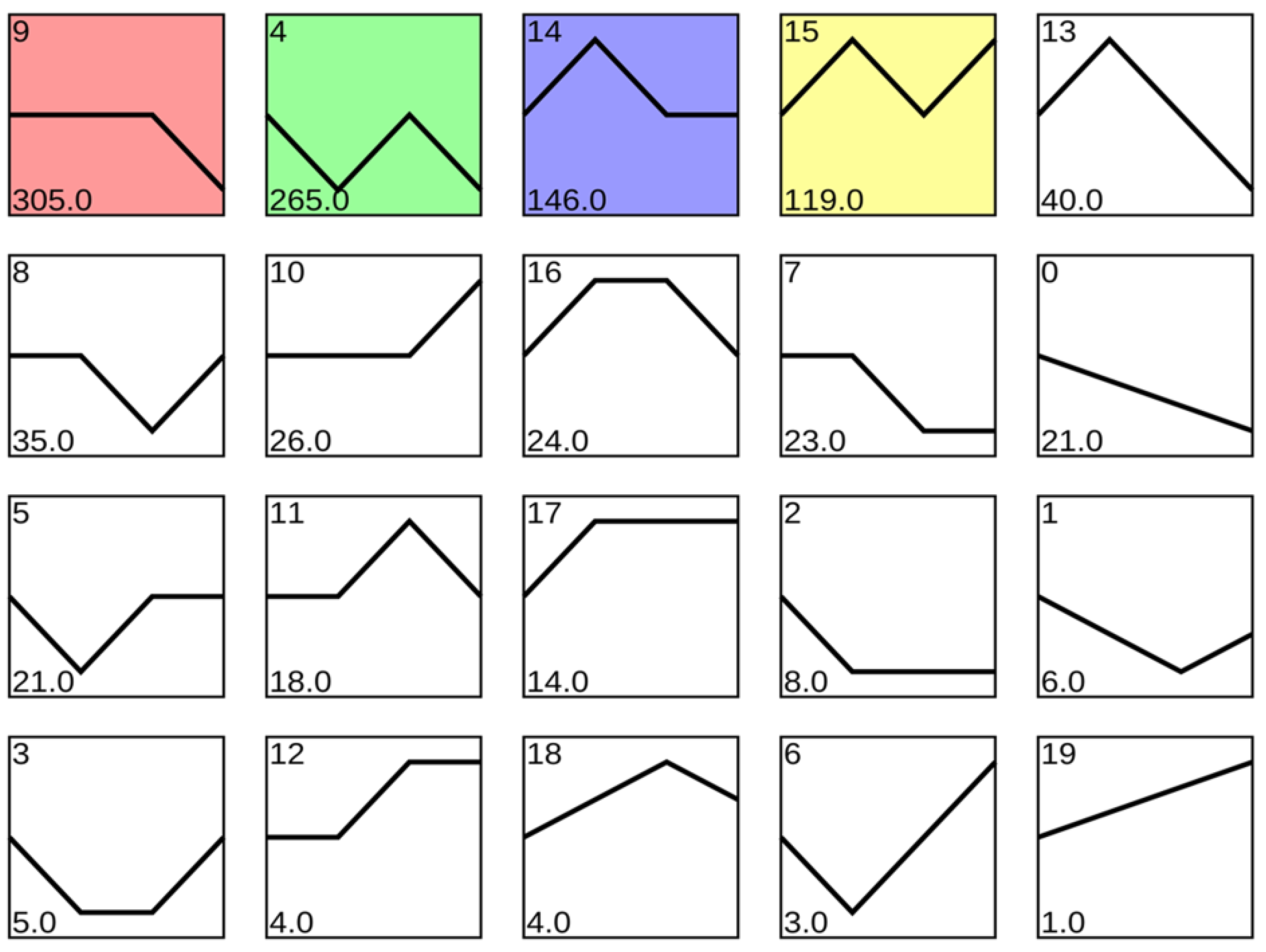
3.2. Analysis of Differential Expressed Genes (DEGs)
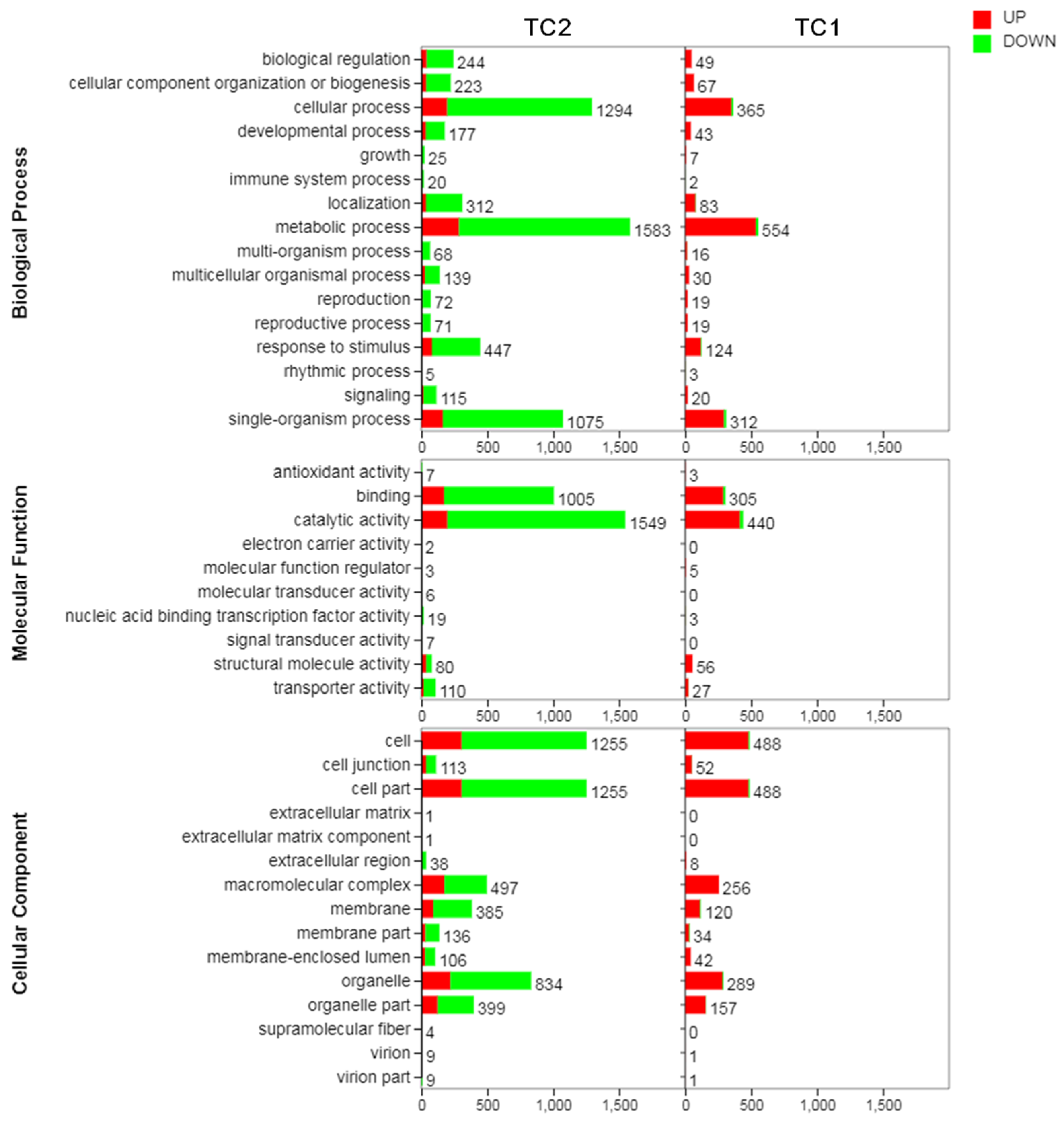
3.3. GO Analysis of DEGs
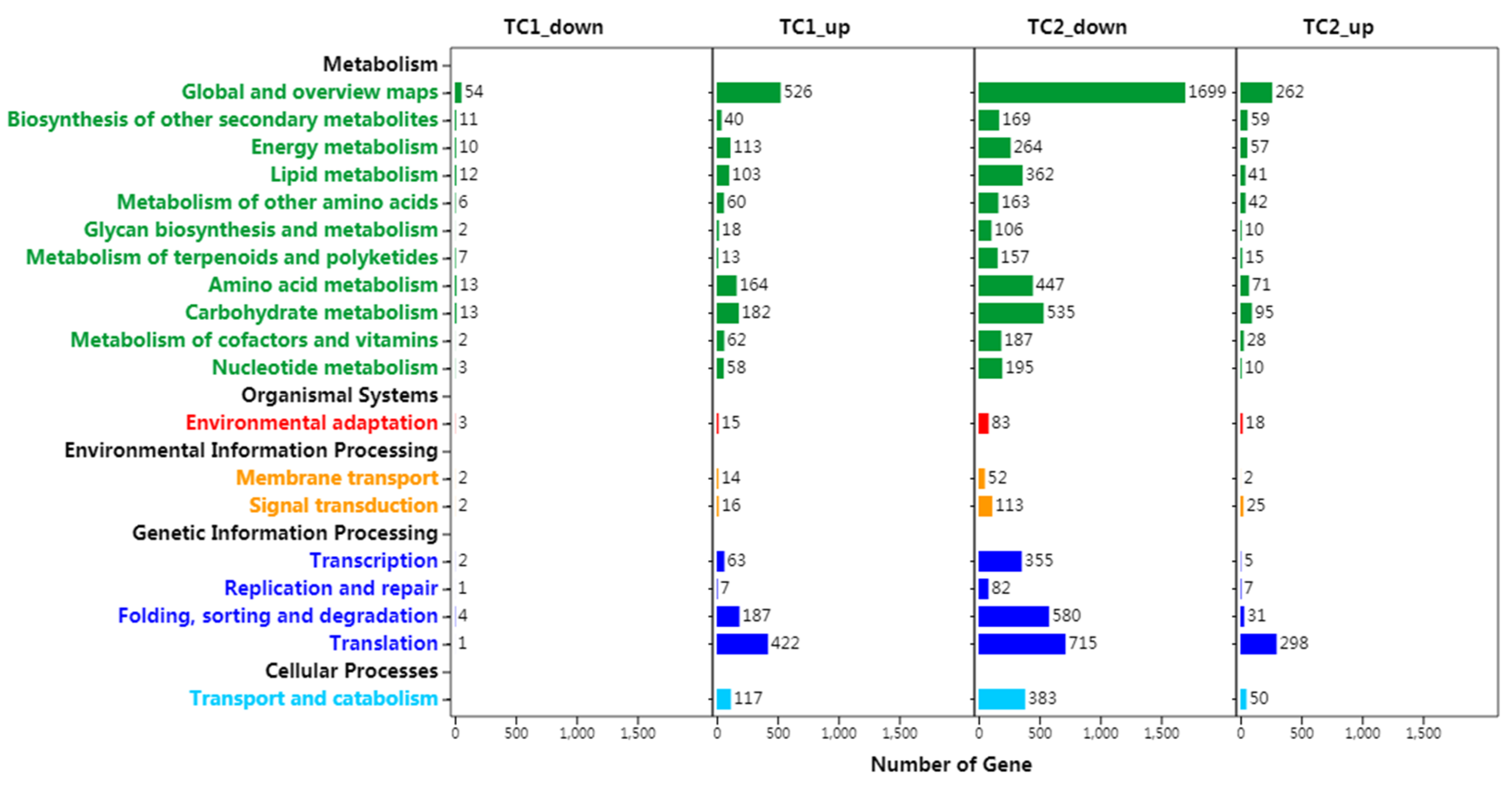
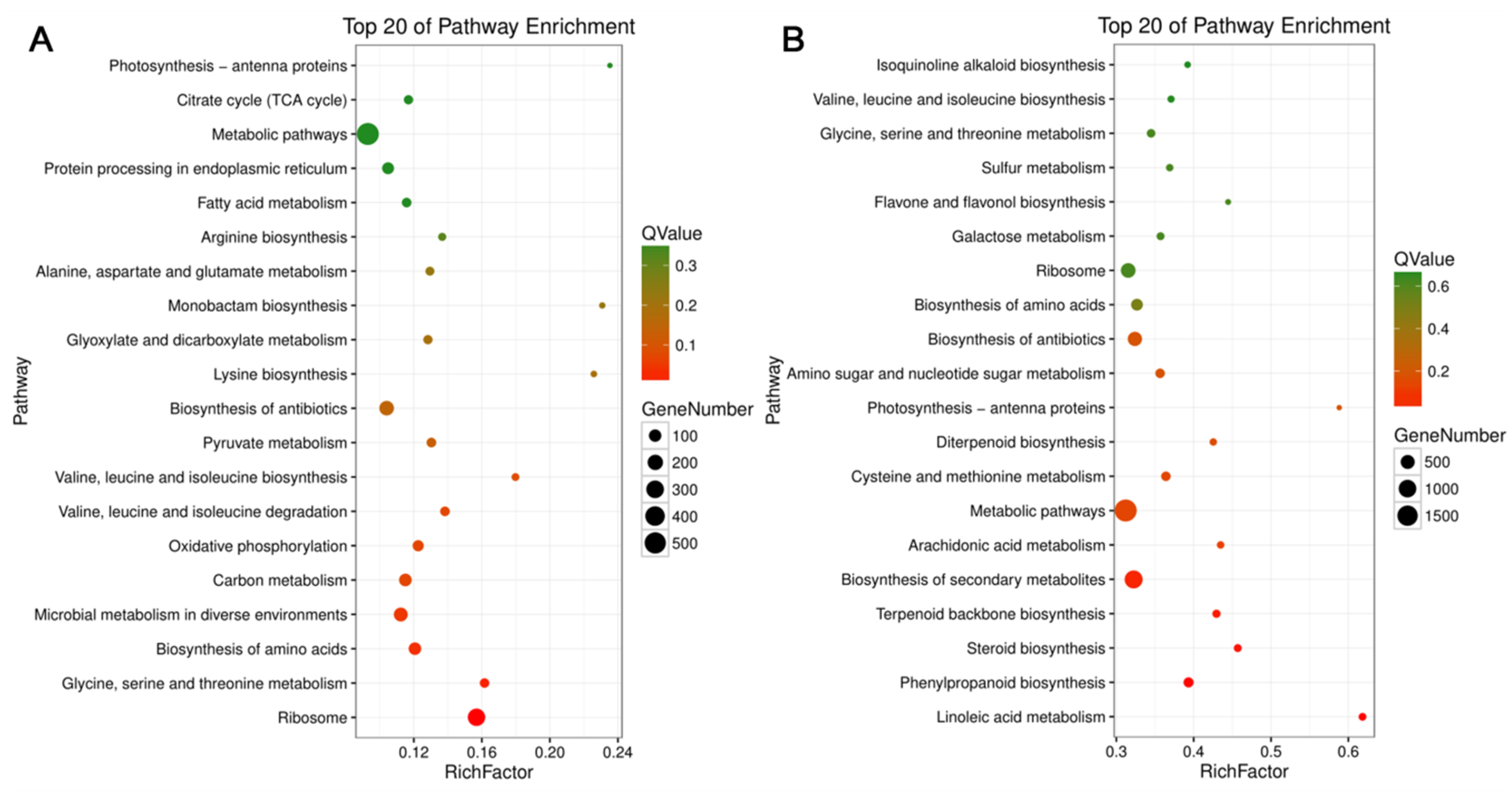
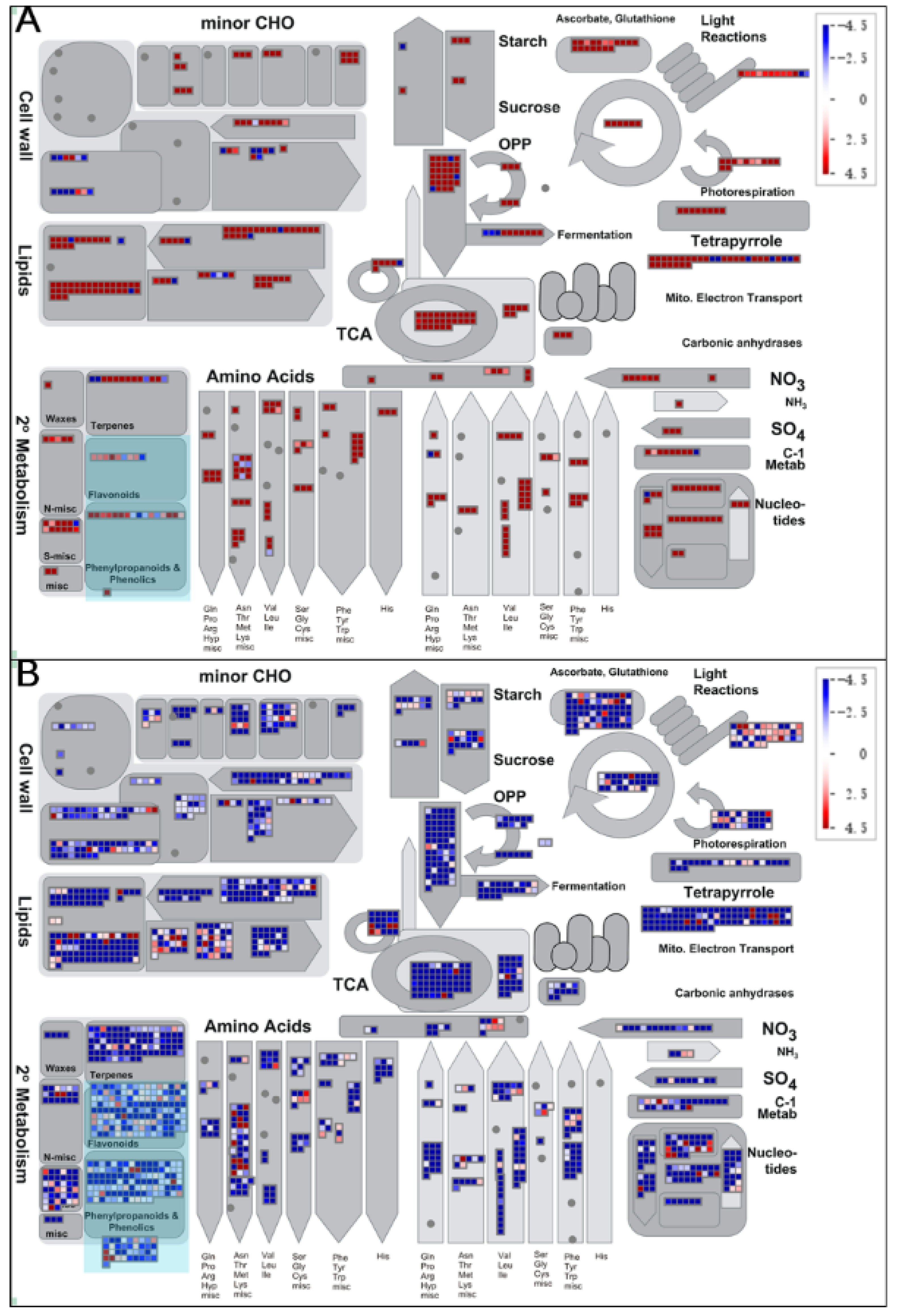
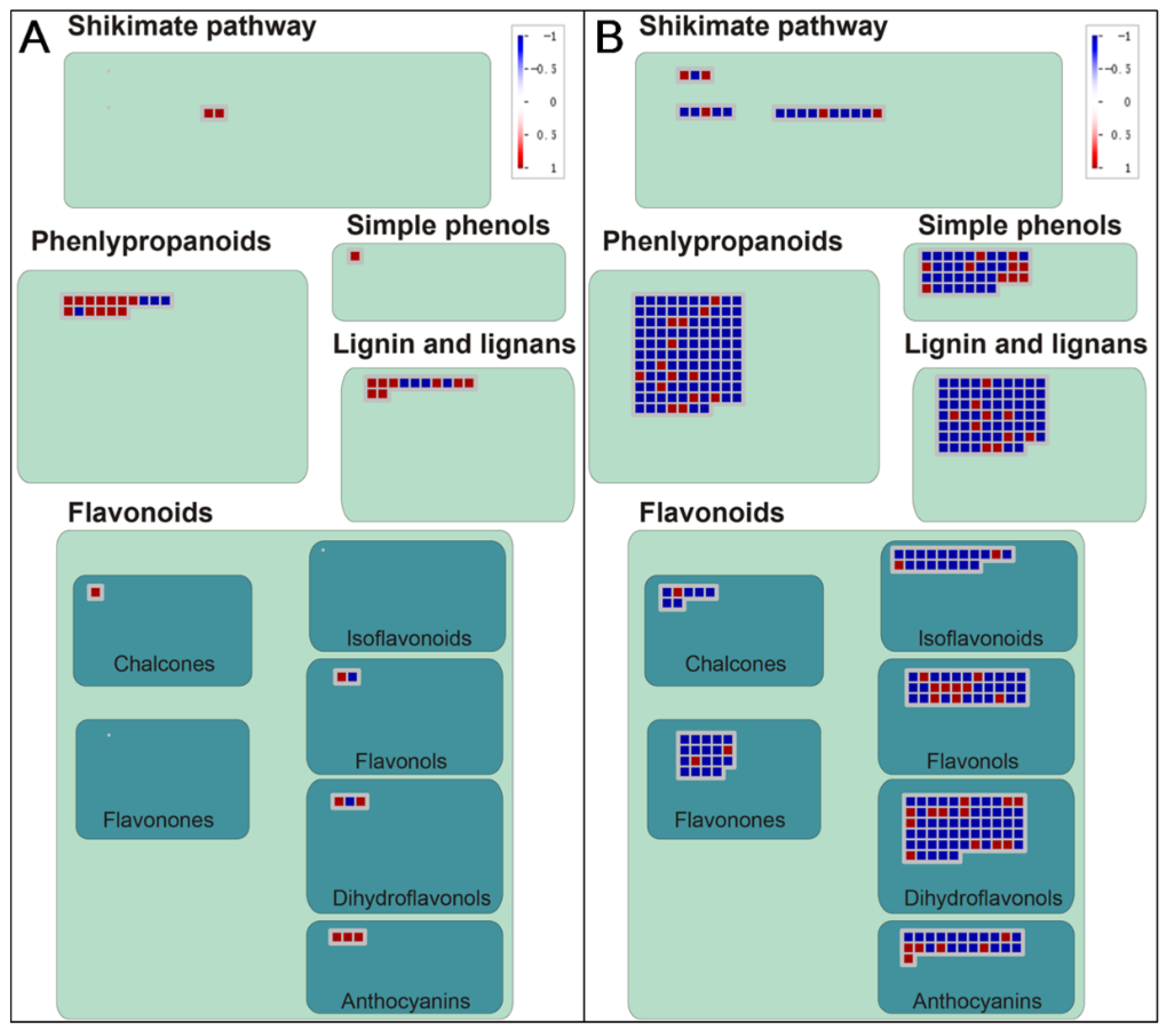
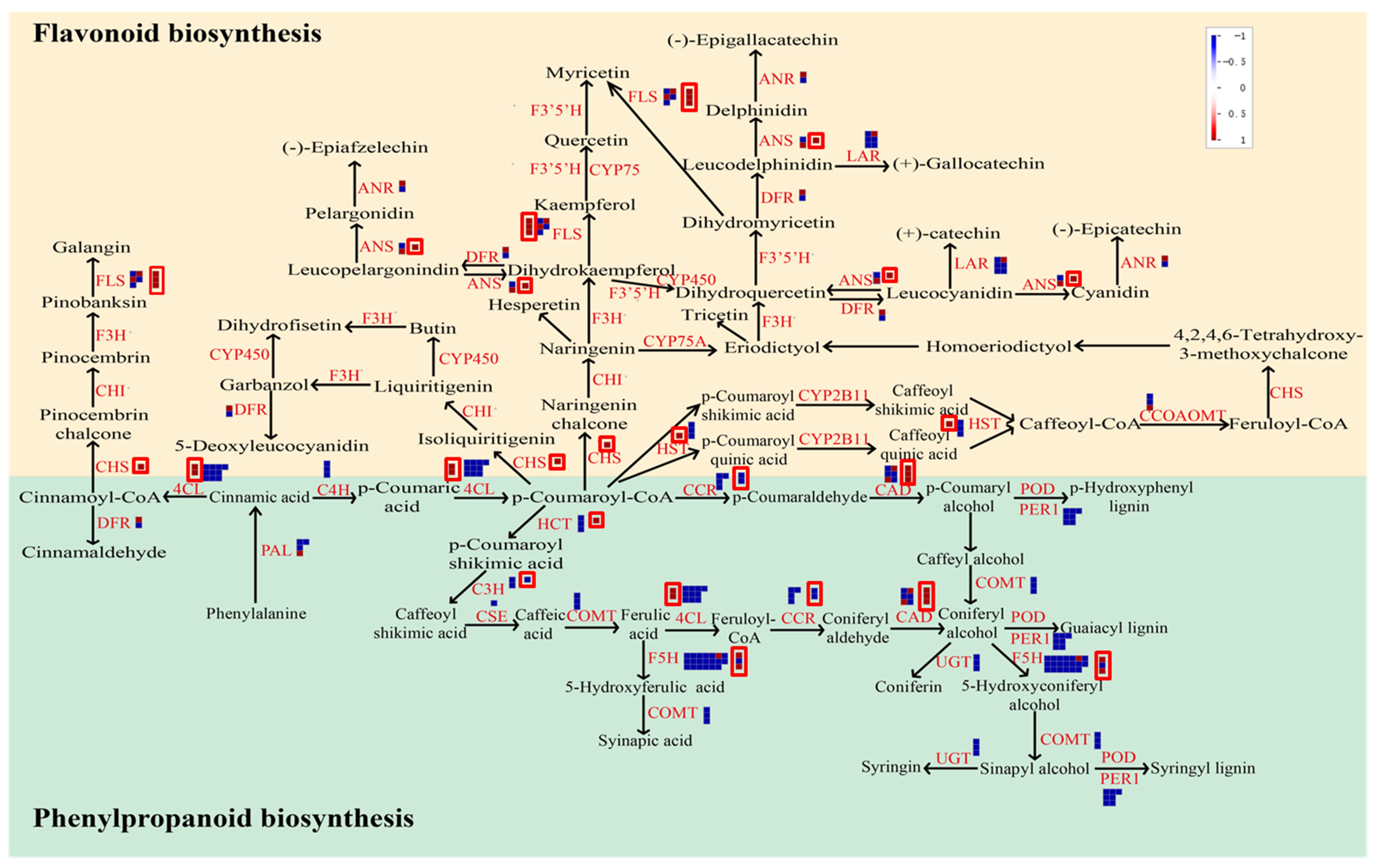
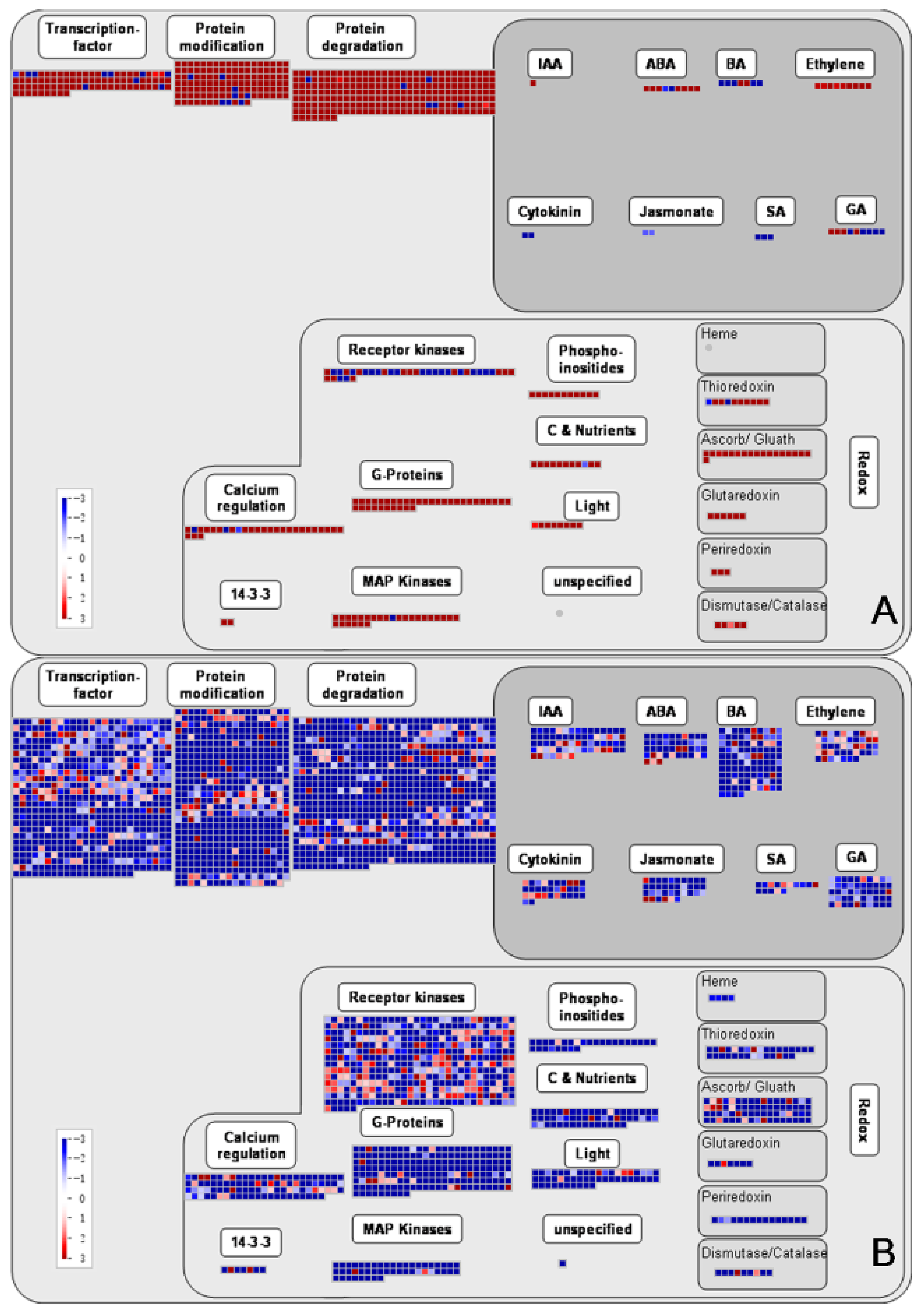
3.4. KEGG Pathway Analysis of DEGs
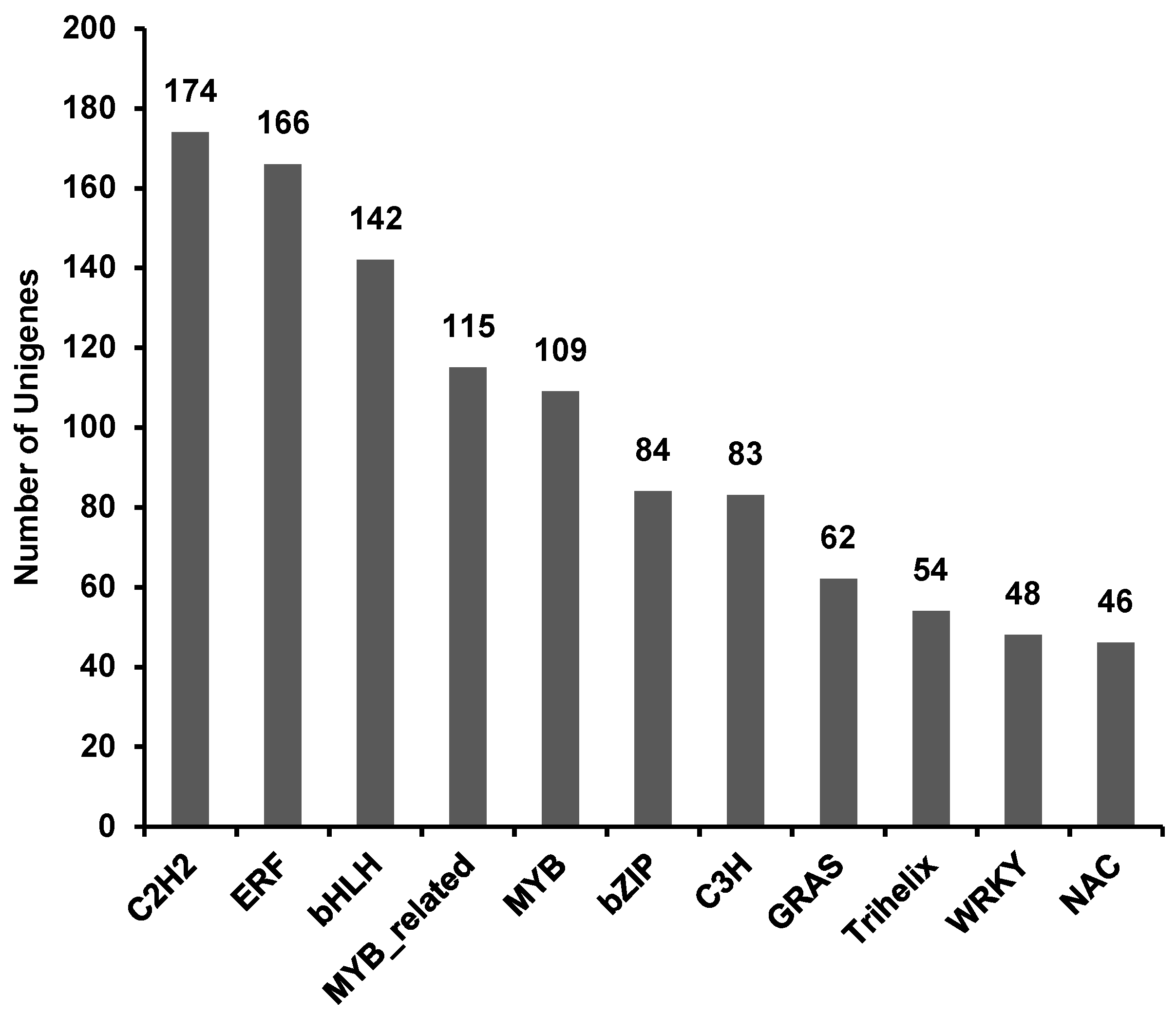
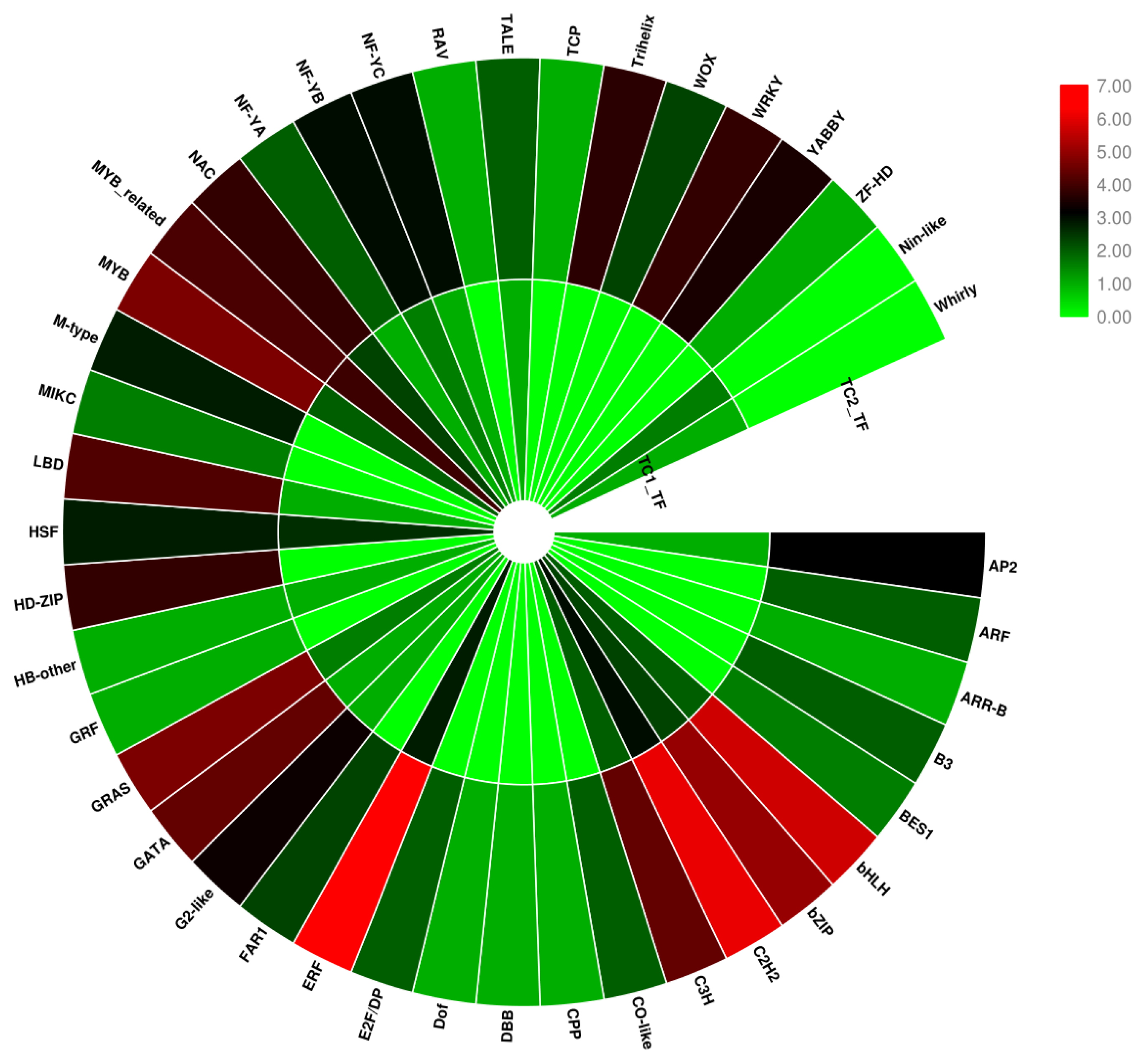
3.5. Transcription Factor Analysis
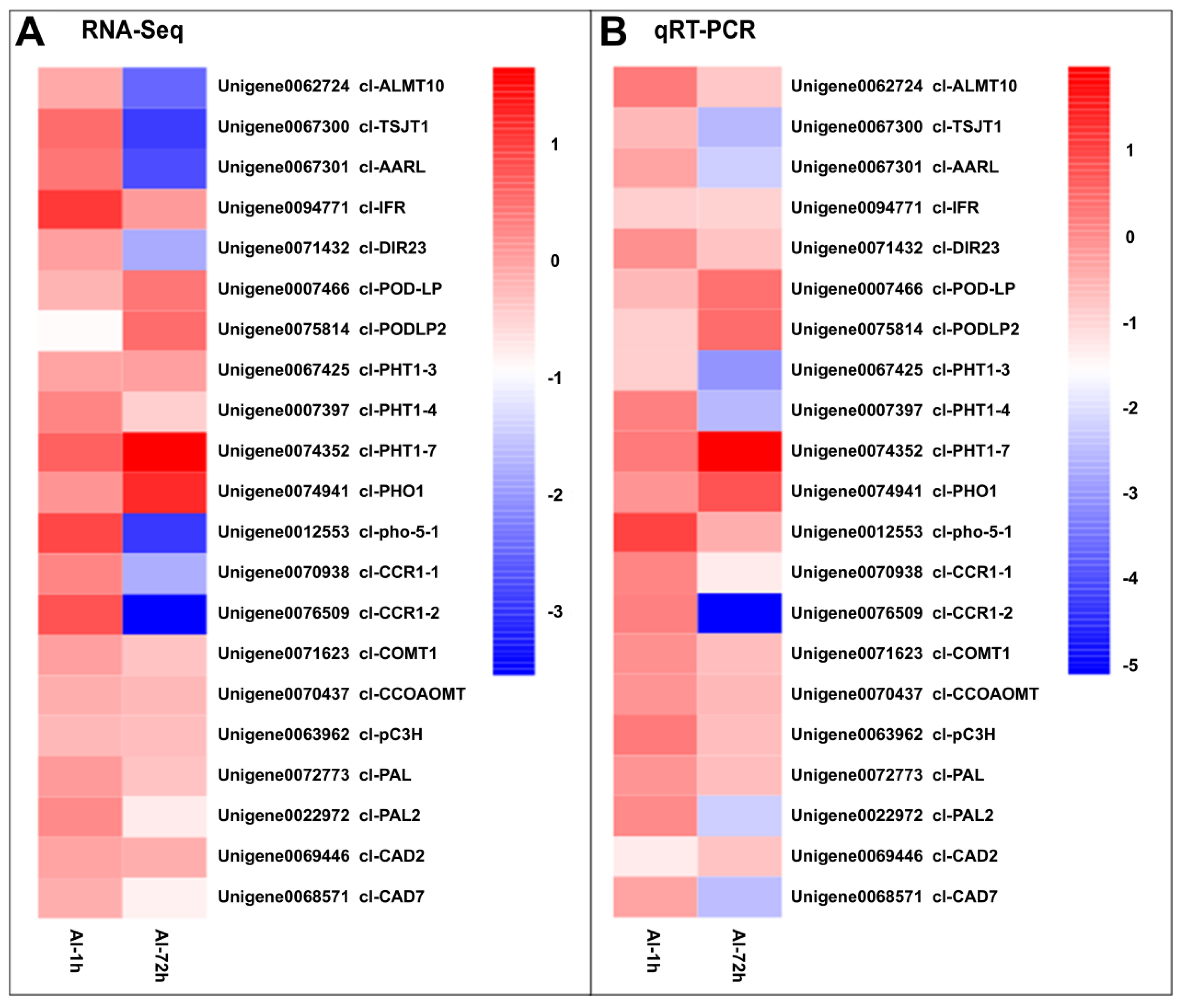
3.6. Validation of RNA-Seq Results by qRT-PCR
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Dong, T.; Duan, B.; Zhang, S.; Korpelainen, H.; Niinemets, Ü.; Li, C. Growth, biomass allocation and photosynthetic responses are related to intensity of root severance and soil moisture conditions in the plantation tree Cunninghamia lanceolata. Tree Physiol. 2016, 36, 807–817. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Han, X.; Sang, J.; He, X.; Liu, M.; Qiao, G.; Zhuo, R.; Hu, J. Transcriptome analysis of immature xylem in the Chinese fir at different developmental phases. PeerJ 2016, 4, e2097. [Google Scholar] [CrossRef] [PubMed]
- Von Uexkull, H.R.; Mutert, E.; Uexkull, H.R. Global extent, development and economic impact of acid soils. Plant Soil 1995, 171, 1–15. [Google Scholar] [CrossRef]
- Bojórquez-Quintal, E.; Escalante-Magaña, C.; Echevarría-Machado, I.; Martínez-Estévez, M. Aluminum, a Friend or Foe of Higher Plants in Acid Soils. Front. Plant Sci. 2017, 8, 1767. [Google Scholar] [CrossRef] [PubMed]
- Zahra, T.; Amin, B.; Shekofeh, E. The effects of aluminum and phosphorous on Brassica napus. Am. Eurasian J. Agric. Environ. Sci. 2009, 6, 137–142. [Google Scholar]
- Kochian, L.V.; Piñeros, M.A.; Liu, J.; Magalhaes, J.V. Plant Adaptation to Acid Soils: The Molecular Basis for Crop Aluminum Resistance. Annu. Rev. Plant Boil. 2015, 66, 571–598. [Google Scholar] [CrossRef]
- Cárcamo, M.P.; Reyes-Díaz, M.; Rengel, Z.; Alberdi, M.; Omena-Garcia, R.P.; Nunes-Nesi, A.; Inostroza-Blancheteau, C. Aluminum stress differentially affects physiological performance and metabolic compounds in cultivars of highbush blueberry. Sci. Rep. 2019, 9, 11275. [Google Scholar]
- Yang, L.T.; Qi, Y.P.; Jiang, H.X.; Chen, L.S. Roles of organic acid anion secretion in aluminium tolerance of higher plants. BioMed Res. Int. 2013, 2013, 173682. [Google Scholar] [CrossRef]
- Kinraide, T.B. Identity of the rhizotoxic aluminium species. Plant Soil 1991, 134, 167–178. [Google Scholar] [CrossRef]
- Kopittke, P.M.; Moore, K.L.; Lombi, E.; Gianoncelli, A.; Ferguson, B.J.; Blamey, F.P.C.; Menzies, N.W.; Nicholson, T.M.; McKenna, B.A.; Wang, P.; et al. Identification of the primary lesion of toxic aluminum in plant roots. Plant Physiol. 2015, 167, 1402–1411. [Google Scholar] [CrossRef]
- Zhao, C.R.; Ikka, T.; Sawaki, Y.; Kobayashi, Y.; Suzuki, Y.; Hibino, T.; Sato, S.; Sakurai, N.; Shibata, D.; Koyama, H. Comparative transcriptomic characterization of aluminum, sodium chloride, cadmium and copper rhizotoxicities in Arabidopsis thaliana. BMC Plant Biol. 2009, 9, 32. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Wang, T.; Zhao, M.; Tian, Q.; Zhang, W.H. Identification of aluminum-responsive microRNAs in Medicago truncatula by genome-wide high-throughput sequencing. Planta 2012, 235, 375–386. [Google Scholar] [CrossRef] [PubMed]
- Rengel, Z. Uptake of aluminium by plant cells. New Phytol. 2010, 134, 389–406. [Google Scholar] [CrossRef]
- Schaedle, M.; Thornton, F.C.; Raynal, D.J.; Tepper, H.B. Response of tree seedlings to aluminum. Tree Physiol. 1989, 5, 337–356. [Google Scholar] [CrossRef] [PubMed]
- Silva, I.R.; Novais, R.F.; Jham, G.N.; Barros, N.F.; Gebrim, F.O.; Nunes, F.N.; Neves, J.C.L.; Leite, F.P. Responses of eucalypt species to aluminum: The possible involvement of low molecular weight organic acids in the Al tolerance mechanism. Tree Physiol. 2004, 24, 1267–1277. [Google Scholar] [CrossRef] [PubMed]
- Tomioka, R.; Takenaka, C. Difference in response to aluminum among Japanese coniferous species. Soil Sci. Plant Nutr. 2004, 50, 755–761. [Google Scholar] [CrossRef]
- Nowak, J.; Friend, A.L. Loblolly pine and slash pine responses to acute aluminum and acid exposures. Tree Physiol. 2006, 26, 1207–1215. [Google Scholar] [CrossRef]
- Nowak, J.; Friend, A.L. Aluminum fractions in root tips of slash pine and loblolly pine families differing in Al resistance. Tree Physiol. 2005, 25, 245–250. [Google Scholar] [CrossRef][Green Version]
- Kelly, J.M.; Schaedle, M.; Thornton, F.C.; Joslin, J.D. Sensitivity of tree seedlings to aluminum. II. Red oak, sugar maple, and European beech. J. Environ. Qual. 1990, 19, 172–179. [Google Scholar] [CrossRef]
- Thornton, F.C.; Schaedle, M.; Raynal, D.J. Effect of aluminum on the growth of sugar maple in solution culture. Can. J. For. Res. 1986, 16, 892–896. [Google Scholar] [CrossRef]
- Chen, Z.C.; Liao, H. Organic acid anions: An effective defensive weapon for plants against aluminum toxicity and phosphorus deficiency in acidic soils. J. Genet. Genom. 2016, 43, 631–638. [Google Scholar] [CrossRef]
- Osawa, H.; Endo, I.; Hara, Y.; Matsushima, Y.; Tange, T. Transient proliferation of proanthocyanidin-accumulating cells on the epidermal apex contributes to highly aluminum-resistant root elongation in camphor tree. Plant Physiol. 2011, 155, 433–446. [Google Scholar] [CrossRef] [PubMed]
- Grisel, N.; Zoller, S.; Künzli-Gontarczyk, M.; Lampart, T.; Münsterkötter, M.; Brunner, I.; Bovet, L.; Métraux, J.-P.; Sperisen, C. Transcriptome responses to aluminum stress in roots of aspen (Populus tremula). BMC Plant Boil. 2010, 10, 185. [Google Scholar] [CrossRef] [PubMed]
- Naik, D.; Smith, E.; Cumming, J.R. Rhizosphere carbon deposition, oxidative stress and nutritional changes in two poplar species exposed to aluminum. Tree Physiol. 2009, 29, 423–436. [Google Scholar] [CrossRef] [PubMed]
- Tahara, K.; Hashida, K.; Otsuka, Y.; Ohara, S.; Kojima, K.; Shinohara, K. Identification of a hydrolyzable tannin, oenothein B, as an aluminum-detoxifying ligand in a highly aluminum-resistant tree, Eucalyptus camaldulensis. Plant Physiol. 2014, 164, 683–693. [Google Scholar] [CrossRef] [PubMed]
- Maejima, E.; Osaki, M.; Wagatsuma, T.; Watanabe, T. Contribution of constitutive characteristics of lipids and phenolics in roots of tree species in Myrtales to aluminum tolerance. Physiol. Plant. 2017, 160, 11–20. [Google Scholar] [CrossRef]
- Ofei-Manu, P.; Wagatsuma, T.; Ishikawa, S.; Tawaraya, K. The plasma membrane strength of the root-tip cells and root phenolic compounds are correiated with Al tolerance in several common woody plants. Soil Sci. Plant Nutr. 2001, 47, 359–375. [Google Scholar] [CrossRef]
- Moghadam, N.D.; Poozesh, V.; Rezaei, A. Biochemical Responses of Two Soybean (Glycine max) Varieties to Aluminum Stress in Nutrient Solution. J. Chem. Health Risks 2016, 6, 237–247. [Google Scholar]
- Zhang, X.; Long, Y.; Huang, J.; Xia, J. Molecular Mechanisms for Coping with Al Toxicity in Plants. Int. J. Mol. Sci. 2019, 20, 1551. [Google Scholar] [CrossRef]
- Ma, J.F.; Chen, Z.C.; Shen, R.F. Molecular mechanisms of Al tolerance in gramineous plants. Plant Soil 2014, 381, 1–12. [Google Scholar] [CrossRef]
- Hirano, Y.; Pannatier, E.G.; Zimmermann, S.; Brunner, I. Induction of callose in roots of Norway spruce seedlings after short-term exposure to aluminum. Tree Physiol. 2004, 24, 1279–1283. [Google Scholar] [CrossRef] [PubMed]
- Umemura, T.; Usami, Y.; Aizawa, S.-I.; Tsunoda, K.-I.; Satake, K.-I. Seasonal change in the level and the chemical forms of aluminum in soil solution under a Japanese cedar forest. Sci. Total Environ. 2003, 317, 149–157. [Google Scholar] [CrossRef]
- Halman, J.M.; Schaberg, P.G.; Hawley, G.J.; Pardo, L.H.; Fahey, T.J. Calcium and aluminum impacts on sugar maple physiology in a northern hardwood forest. Tree Physiol. 2013, 33, 1242–1251. [Google Scholar] [CrossRef] [PubMed]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644. [Google Scholar] [CrossRef] [PubMed]
- Mortazavi, A.; Williams, B.A.; McCue, K.; Schaeffer, L.; Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 2008, 5, 621–628. [Google Scholar] [CrossRef] [PubMed]
- Conesa, A.; Götz, S.; García-Gómez, J.M.; Terol, J.; Talón, M.; Robles, M. Blast2GO: A universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 2005, 21, 3674–3676. [Google Scholar] [CrossRef]
- Ye, J.; Fang, L.; Zheng, H.; Zhang, Y.; Chen, J.; Zhang, Z.; Wang, J.; Li, S.; Li, R.; Bolund, L.; et al. WEGO: A web tool for plotting GO annotations. Nucleic Acids Res. 2006, 34, W293–W297. [Google Scholar] [CrossRef]
- Xu, H.; Cao, D.; Feng, J.; Wu, H.; Lin, J.; Wang, Y. Transcriptional regulation of vascular cambium activity during the transition from juvenile to mature stages in Cunninghamia lanceolata. J. Plant Physiol. 2016, 200, 7–17. [Google Scholar] [CrossRef]
- Chakraborty, S.; Mishra, A.; Verma, E.; Tiwari, B.; Mishra, A.K.; Singh, S.S. Physiological mechanisms of aluminum (Al) toxicity tolerance in nitrogen-fixing aquatic macrophyte Azolla microphylla Kaulf: Phytoremediation, metabolic rearrangements, and antioxidative enzyme responses. Environ. Sci. Pollut. Res. 2019, 26, 9041–9054. [Google Scholar] [CrossRef]
- Kidd, P.; Poschenrieder, C.; Llugany, M.; Gunse, B.; Barceló, J. The role of root exudates in aluminium resistance and silicon-induced amelioration of aluminium toxicity in three varieties of maize (Zea mays L.). J. Exp. Bot. 2001, 52, 1339–1352. [Google Scholar]
- Kumari, M.; Taylor, G.J.; Deyholos, M.K. Transcriptomic responses to aluminum stress in roots of Arabidopsis thaliana. Mol. Genet. Genom. 2008, 279, 339. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhang, J.; Guo, J.; Zhou, F.; Singh, S.; Xu, X.; Xie, Q.; Yang, Z.; Huang, C.-F. F-box protein RAE1 regulates the stability of the aluminum-resistance transcription factor STOP1 in Arabidopsis. Proc. Natl. Acad. Sci. USA 2019, 116, 319–327. [Google Scholar] [CrossRef] [PubMed]
- An, J.-P.; Yao, J.-F.; Ren, Y.-R.; You, C.-X.; Wang, X.-N.; Hao, Y.-J. Apple MdMYC2 reduces aluminum stress tolerance by directly regulating MdERF3 gene. Plant Soil 2017, 418, 255–266. [Google Scholar] [CrossRef]
- Yao, Y.; He, R.J.; Xie, Q.L.; Zhao, X.H.; Deng, X.M.; He, J.B.; Song, L.; He, J.; Marchant, A.; Chen, X.-Y.; et al. ETHYLENE RESPONSE FACTOR 74 (ERF74) plays an essential role in controlling a respiratory burst oxidase homolog D (RbohD)-dependent mechanism in response to different stresses in Arabidopsis. New Phytol. 2017, 213, 1667–1681. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ma, Z.; Lin, S. Transcriptomic Revelation of Phenolic Compounds Involved in Aluminum Toxicity Responses in Roots of Cunninghamia lanceolata (Lamb.) Hook. Genes 2019, 10, 835. https://doi.org/10.3390/genes10110835
Ma Z, Lin S. Transcriptomic Revelation of Phenolic Compounds Involved in Aluminum Toxicity Responses in Roots of Cunninghamia lanceolata (Lamb.) Hook. Genes. 2019; 10(11):835. https://doi.org/10.3390/genes10110835
Chicago/Turabian StyleMa, Zhihui, and Sizu Lin. 2019. "Transcriptomic Revelation of Phenolic Compounds Involved in Aluminum Toxicity Responses in Roots of Cunninghamia lanceolata (Lamb.) Hook" Genes 10, no. 11: 835. https://doi.org/10.3390/genes10110835
APA StyleMa, Z., & Lin, S. (2019). Transcriptomic Revelation of Phenolic Compounds Involved in Aluminum Toxicity Responses in Roots of Cunninghamia lanceolata (Lamb.) Hook. Genes, 10(11), 835. https://doi.org/10.3390/genes10110835




