Transcriptional Analysis of Masson Pine (Pinus massoniana) under High CO2 Stress
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Material and Experimental Conditions
2.2. Total RNA Isolation, Complementary DNA Library Preparation and Sequencing
2.3. Transcriptome Assembly and Functional Annotation
2.4. Differential Expression Analysis
2.5. Quantitative Real-Time PCR Validation
3. Results
3.1. Transcriptome Sequencing and De Novo Assembly
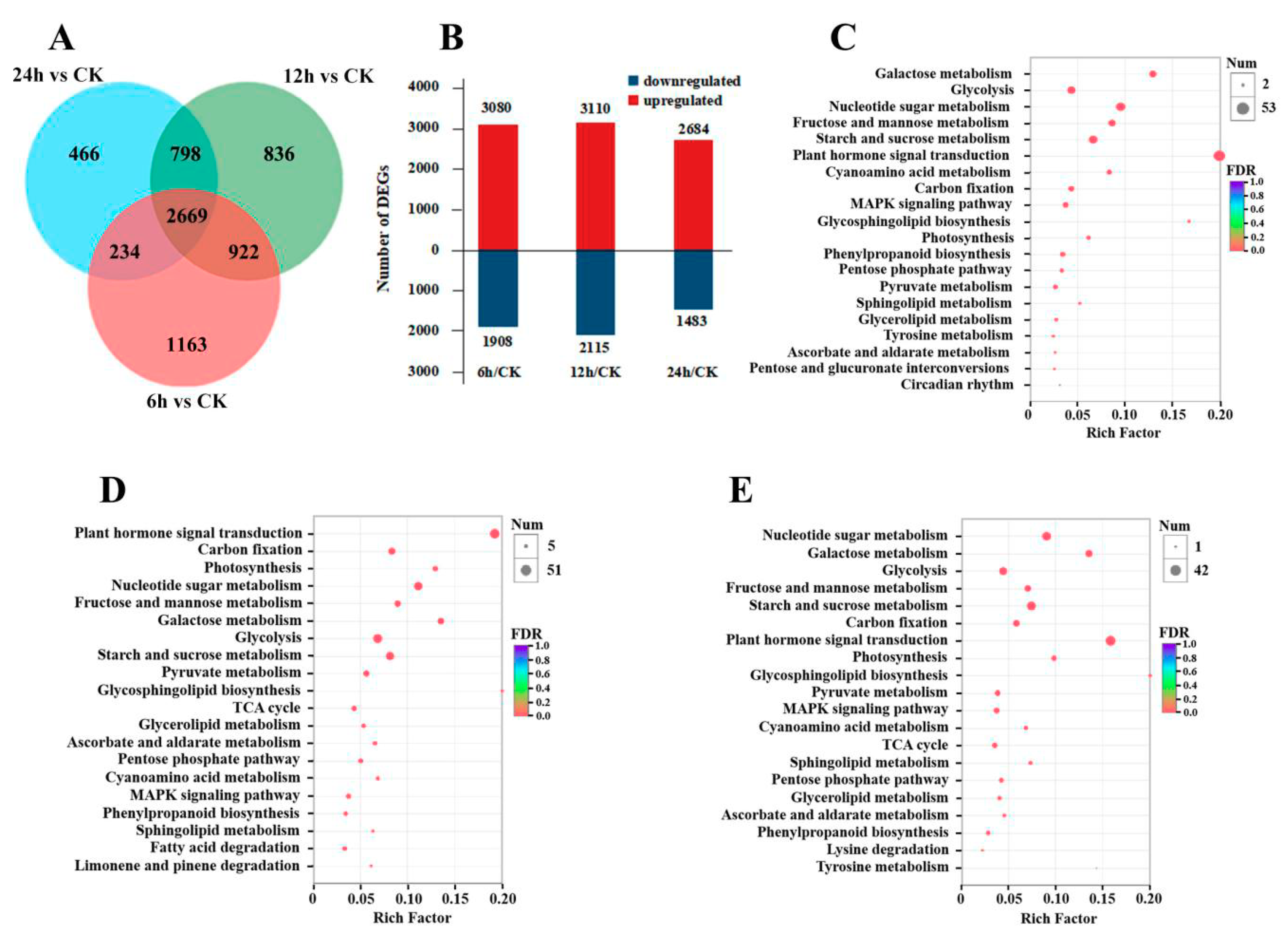
3.2. Gene Expression and KEGG Enrichment Analysis under CO2 Stress
3.3. Energy Metabolism under Elevated CO2 Stress
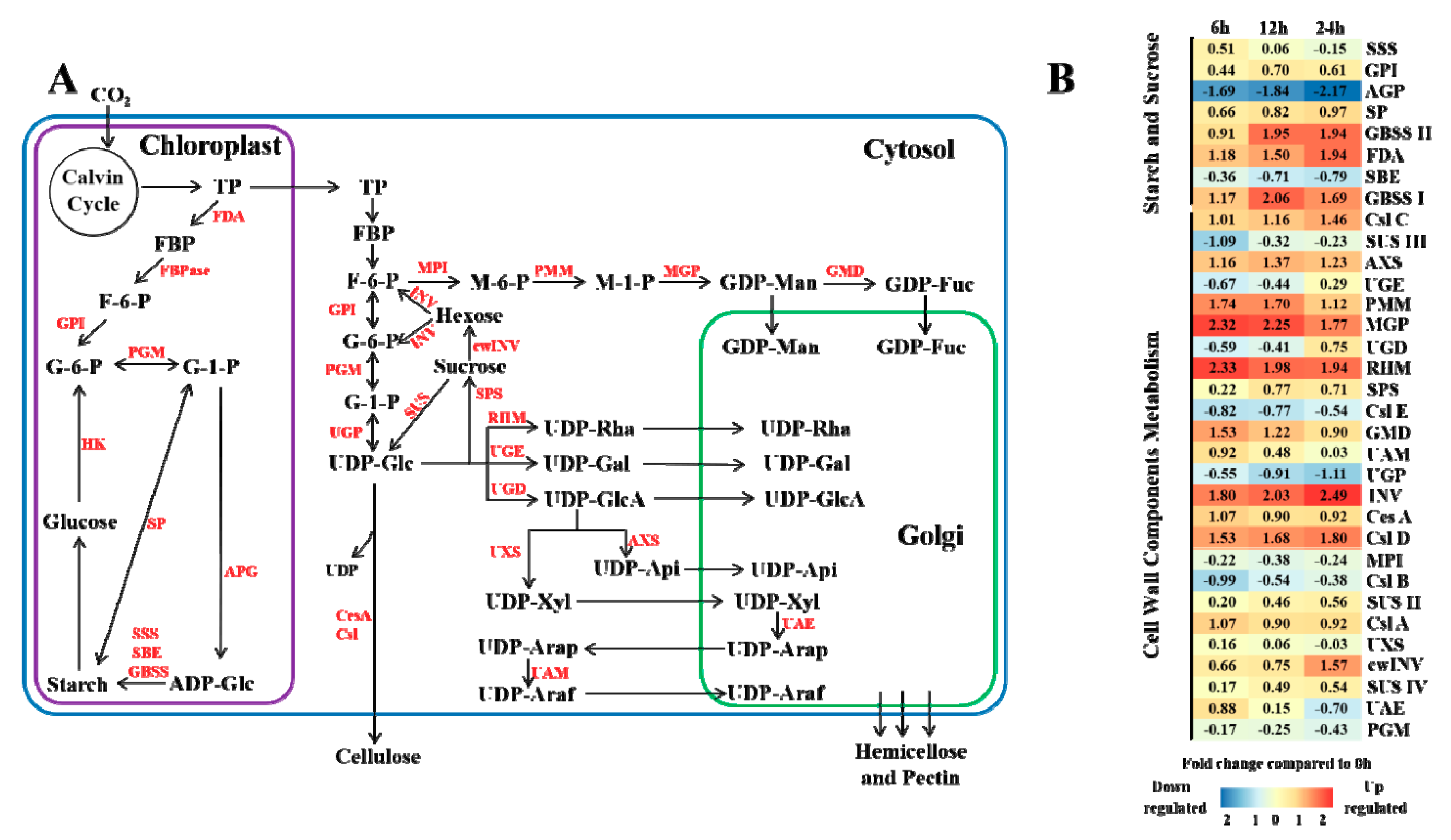
3.4. Biosynthesis of Sucrose, Starch and Cell Wall Components under Elevated CO2 Stress
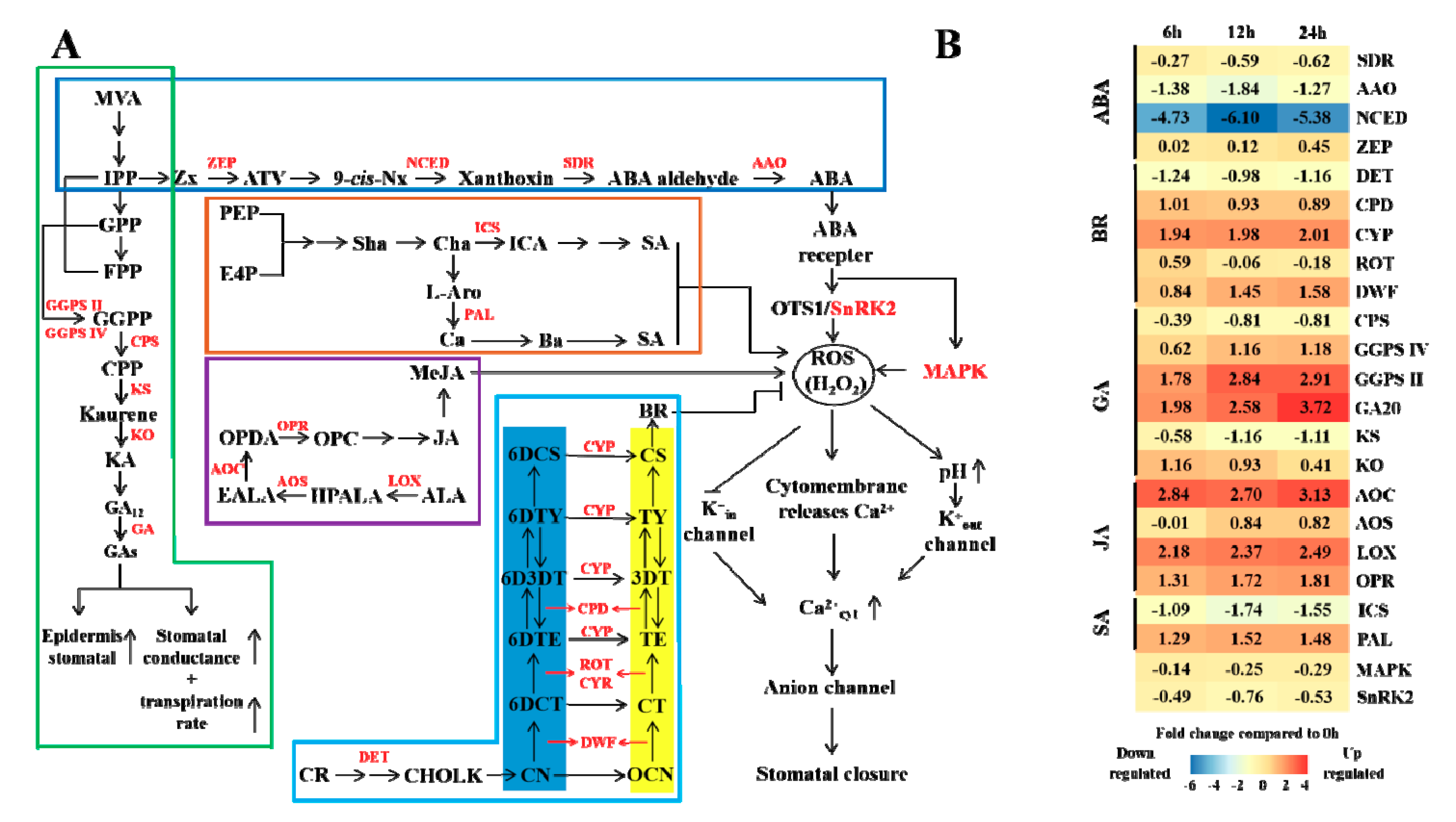
3.5. Hormone Regulation under Elevated CO2 Stress
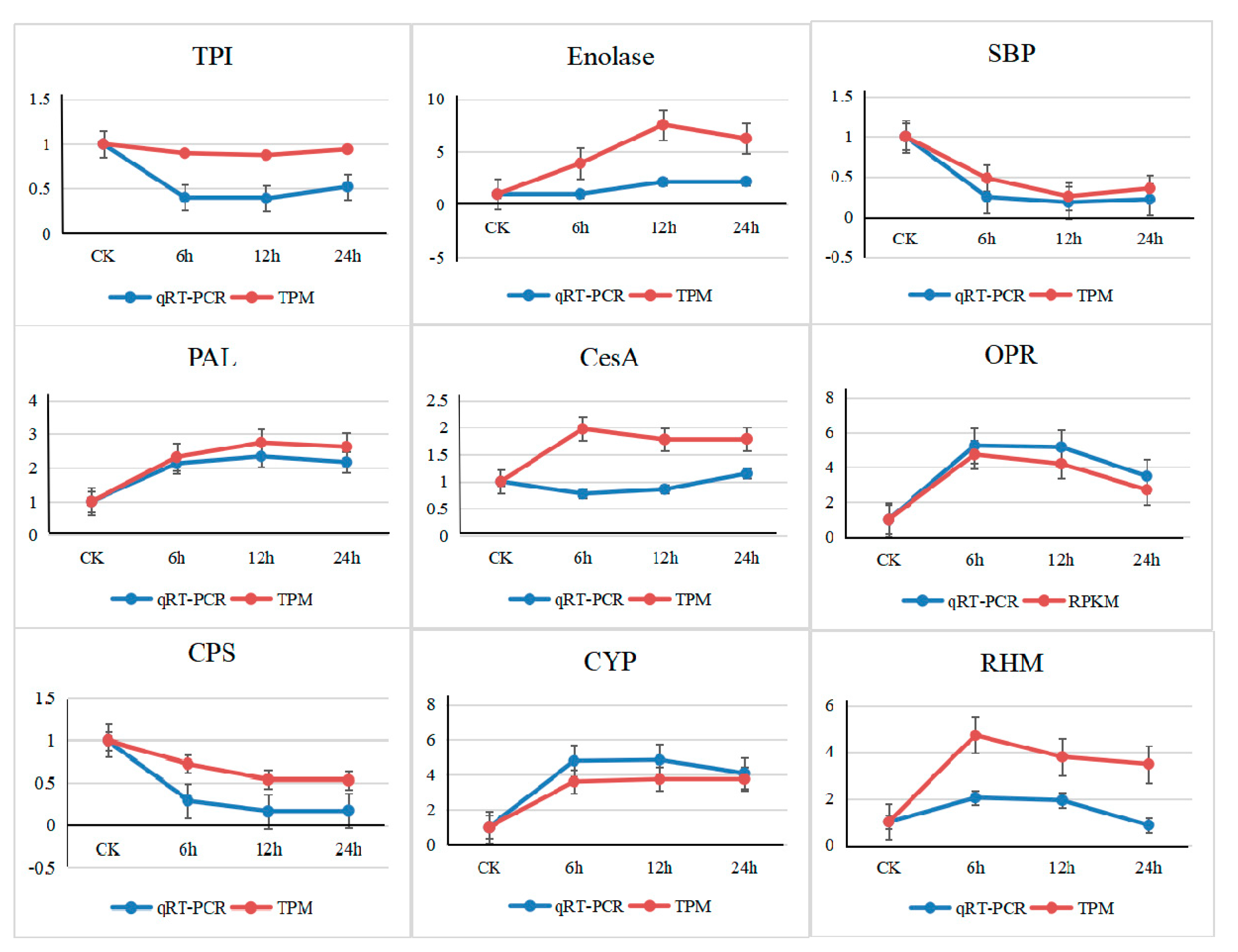
3.6. Validation by Quantitative Real-Time PCR
4. Discussion
4.1. Effects of Elevated CO2 on Masson Pine Energy Metabolism
4.2. Effects of Elevated CO2 on Masson Pine Carbohydrate and Cell Wall Component Synthesis
4.3. Effects of Elevated CO2 on Masson Pine Cell Wall Component Synthesis
4.4. Effects of Elevated CO2 on Masson Pine Hormones and Stomatal Regulation
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
| Primer | Sequence (5′→3′) | |
|---|---|---|
| RHM | Forward | TACGAATAGTCTCTGGCTTGTGAG |
| Reverse | TCTGGTTGTGTCCTTGACCTAATA | |
| CYP | Forward | TCTATGGTGATCACTGGAGAAAGA |
| Reverse | GATGAGAGAATGGTTGAGAATGTG | |
| CPS | Forward | TACTCGGTGTTATAAGTGCAGCTC |
| Reverse | CATGTAGCCCTTGACACAAAATAG | |
| OPR | Forward | TACGATACGGGAACAACTACTGAA |
| Reverse | TCGAGCTCTAAAAACTGAGGAGAT | |
| PAL | Forward | GAAGCCTGAGTTTACAGATCCATT |
| Reverse | CGTAAACCACTTCAATCACTTCAC | |
| CesA | Forward | GGAAGGCTGTACTTTATCCTTCAA |
| Reverse | ATGCAAGACCAGATACAAGAGACA | |
| SBP | Forward | TACCAGCCCAATAACAATAACCAC |
| Reverse | CTCTCATCCACGAAGCTAATAACC | |
| TPI | Forward | CCCTCTGCCACTTTCTTTATGTC |
| Reverse | TCTAAGACTTCCTCACTTCTCCG | |
| Enolase | Forward | AAGAGCTGCAAGGTAAAGTCTGTT |
| Reverse | TCTGATTCACCTTCAGCAGTAAAG | |
| qRT-PCR-RHM | Forward | CCACATCCTCACAGTAGAGATAGC |
| Reverse | CCGGTGATTACTACCAGAGGTAAC | |
| qRT-PCR-CYP | Forward | AGGCCCTTCCTCAGAGGTTATCT |
| Reverse | CCGGAGTTGGTACTAGTCTTGGTA | |
| qRT-PCR-CPS | Forward | GTCTAGAGCGGTTCACTCAGAT |
| Reverse | CCTCTCTCCAACTATCACTGTGTC | |
| qRT-PCR-OPR | Forward | GGTACCGTTCTTACTGGTTTGAGG |
| Reverse | GATCCTGTAGTTGGCTACACAGAC | |
| qRT-PCR-PAL | Forward | CCCTCAGGTGGAGATTATCAG |
| Reverse | CCTCCATGTAGAGCTTTGTCTC | |
| qRT-PCR-CesA | Forward | CCTGTACGGAGTAAGTTTGGTG |
| Reverse | ACCAGTGGAGGTAGATATGCTG | |
| qRT-PCR-SBP | Forward | CTACAGAGATAGGAGAGGGGAAAC |
| Reverse | CTCCTGTGTATCGGAGTGTGTACT | |
| qRT-PCR-TPI | Forward | CCTCCCACTTCTACTAGGGTTT |
| Reverse | ACCAGCCAGGAGTAGTTAAGAGTG | |
| qRT-PCR-Enolase | Forward | GGCCAGACAGATTATAGACAGC |
| Reverse | CTCTCATCTCTAGGGCCTCATA |
| Abbr. | Full Name | Abbr. | Full Name |
|---|---|---|---|
| PS | Photosystem | LEC | Light Harvesting Complex |
| PC | Plastocyanin | RuBP | Ribulose-1,5-Bisphosphate |
| 3-PG | 3-Phosphoglycerate | 1,3-DPG | 1,3-Diphosphoglycerate |
| PGAL | 3-Phosphoglyceraldehyde | DHAP | Dihydroxy-Acetone Phosphate |
| FBP | Fructose-1,6-Diphosphate | F-6-P | Fructose-6-Phosphate |
| E-4-P | Erythrose 4-Phosphate | SBP | Sedoheptulose-1,7-Diphosphate |
| S-7-P | Sedoheptulose-7-Phosphate | R-5-P | Ribose-5-Phosphate |
| Ru-5-P | Ribulose-1,5-Phosphate | Xu-5-P | Xylulose-5-Phosphate |
| 6-PG | 6-Phosphogluconate | 6-PG-L | Gluconolactone-6-Phosphate |
| G-6-P | Glucose-6-Phosphate | 2-PG | 2-Phosphoglycerate |
| PEP | Phosphoenolpyruvic acid | EP | Enolpyruvic Acid |
| Pyr | Pyruvic Acid | AcCoA | Acetyl CoA |
| ICL | Isocitric Acid | KGA | α-Ketoglutaric Acid |
| SUA | Succinate Acid | H2MA | Malic Acid |
| OAA | Oxaloacetic Acid | RCA | Rubisco Activase |
| PGK | Phosphoglycerate Kinase | GAPD | Glyceraldehyde-3-Phosphate Dehydrogenase |
| TPI | Triose-Phosphate Isomerase | TRA | Transaldolase |
| FBPase | Fructose-1,6-Bisphosphatase | TKL | Transketolase |
| RPI | Ribulose Phosphate Isomerase | SBPase | Sedoheptulose-1,7-Bisphosphatase |
| RPK | Phosphoribulokinase | RPE | Ribulose Phosphate Epimerase |
| 6-PGDH | 6-Phosphogluconate Dehydrogenase | G-6-PD | Glucose-6-Phosphate-Dehydrogenase |
| G-6-Pase | Glucose-6-Phosphatase | HK | Hexokinase |
| PK | Pyruvate Kinase | PGAM | Phosphoglycerate Mutase |
| CS | Citrate Synthase | ICD | Isocitrate Dehydrogenase |
| OGDC | α-Ketoglutarate Dehydrogenase | SDH | Succinate Dehydrogenase |
| MDH | Malate Dehydrogenase | TP | Triose Phosphate |
| ADP-Glc | Adenosine Diphosphate Glucose | UDP-Glc | Uridine Diphosphate Glucose |
| M-6-P | Mannose-6-Phosphate | M-1-P | Mannose-1-Phosphate |
| GDP-Man | Guanosine Diphosphate Mannose | GDP-Fuc | Guanosine Diphosphate Fucose |
| UDP-Rha | Uridine Diphosphate Rhamnose | UDP-Gal | Uridine Diphosphate Galacturonate |
| UDP-GlcA | Uridine Diphosphate Glucuronate | UDP-Xyl | Uridine Diphosphate Xylose |
| UDP-Api | Uridine Diphosphate Apiose | UDP-Arap | Uridine Diphosphate Arabinose, Pyranose form |
| UDP-Araf | Uridine Diphosphate, Furanose form | FDA | Fructose-Bisphosphate Aldolase |
| GPI | Phosphoglucose Isomerase | PGM | Phosphoglucomutase |
| SP | Starch Phosphorylase | AGP | Adenosine Diphosphoglucose Pyrophosphorylase |
| SSS | Soluble Starch Synthase | GBSS | Granule-bound Starch Synthase |
| SBE | Starch Branching Enzyme | UGP | UDP-Glucose Pyrophosphorylase |
| CesA | Cellulose Synthase Catalytic Subunit | Csl | Cellulose Synthase |
| MPI | Phosphomannose Isomerase | PMM | Phosphomannomutase |
| MGP | Mannose-1-Phosphate Guanylyltransferase | GMD | GDP-Mannose 4,6-Dehydratase |
| INV | Invertase | cwINV | Cell Wall Invertase |
| SPS | Sucrose-Phosphate Synthase | SUS | Sucrose Synthase |
| RHM | Rhamnose Biosynthetic Enzyme | UGE | UDP-Glucose 4-Epimerase |
| UGD | UDP-Glc Dehydrogenase | UXS/AXS | UDP-Glucuronate Decarboxylases |
| UAE | UDP-Arabinose 4-Epimerase | UAM | UDP-Arabinose Mutase |
| ABA | Abscisic Acid | GA | Gibberellin |
| SA | Salicylic Acid | JA | Jasmonate |
| BR | Brassinolide | MVA | Mevalonic Acid |
| IPP | Isopentenyl Pyrophosphate | Zx | Zeaxanthin |
| ATV | All-trans Violaxanthin | 9-cis-Nx | 9-cis-Neoxanthin |
| GPP | Geranyl- Pyrophosphate | FPP | Farnesyl Pyrophosphate |
| GGPP | Geranylgeranyl Pyrophosphate | CPP | Cuban Pyrophosphate |
| KA | Kaurenoic Acid | Sha | Shikimic Acid |
| Cha | Chorismic acid | ICA | Isochorismic Acid |
| L-Aro | L-Arogenate | Ca | Cinnamic Acid |
| Ba | Benzoic Acid | ALA | Linolenic Acid |
| HPALA | 13-Hydrogen Peroxide Linolenic Acid | EALA | 12,13-Epoxylinolenic Acid |
| OPDA | 12-Oxophytodienoic Acid | CR | Campesterol |
| CHOLK | Cholesten-3-Ketone | CN | Campestanol |
| OCN | 6-Oxocampestanol | CT | Cathasterone |
| 6DCT | 6-Deoxycathasterone | TE | Teasterone |
| 6DTE | 6-Deoxyteasterone | 3DT | 3-Dehydrocathasterone |
| 6D3DT | 6-Deoxy-3-Dehydrocathasterone | TY | Typhasterol |
| 6DTY | 6-Deoxytyphasterol | CS | Castasterone |
| 6DCS | 6-Deoxycastasterone | OTS1 | Open Stomata 1 |
| SnRK2 | Sucrose Non-fermenting 1-related Kinase | MAPK | Mitogen Activated Protein Kinase |
| ZEP | Zeaxanthin Epoxidase | NCED | 9-cis-Epoxycarotenoid Dioxygenase |
| SDR | Short-chain Dehydrogenase/reductase | AAO | ABA Aldehyde Oxidase |
| ICS | Isochorismate Synthase | PAL | Phenylalanine Ammonialyase |
| GGPS | Geranylgeranylpyrophosphate | CPS | Copalyl Pyrophosphate Synthase |
| KS | Kaurene Synthase | KO | Kaurene Oxidase |
| GA | Gibberellin Oxidase | LOX | Lipoxygenase |
| AOS | Allene Oxide Synthase | AOC | Allene Oxide Cyclase |
| OPR | 12-Oxophytodienoate Reductase | DWF | Trans-Cinnamate 4-Hydroxylase |
| ROT | C-23 Hydroxylase | CPD | Coumarate-3-Hydroxylase |
References
- Liu, Y.H.; Wang, H.J.; Zhang, X.Q. The background of climate change, diplomatic and political games. In Climate Change and Forest Carbon in China; Li, T.Y., Lin, H., Eds.; China Meteorological Press: Beijing, China, 2014; pp. 9–14. [Google Scholar]
- GCP. Carbon Budget. Available online: http://www.globalcarbonproject.org/carbonbudget (accessed on 6 July 2019).
- Yue, J.W. Dynamics, Potential and Mechanism of Carbon Sequestration in Major Forest Types in Gansu Province, China. Ph.D. Thesis, University of Chinese Academy of Sciences, Beijing, China, 2018. [Google Scholar]
- Urban, O. Physiological Impacts of Elevated CO2 Concentration Ranging from Molecular to Whole Plant Responses. Photosynthetica 2003, 41, 9–20. [Google Scholar] [CrossRef]
- Chen, G.Y.; Yong, Z.H. Photosynthetic Acclimation in Rice Leaves to Free-air CO2 Enrichment Related to Both Ribulose-1,5-bisphosphate Carboxylation Limitation and Ribulose-1,5-bisphosphate Regeneration Limitation. Plant Cell Physiol. 2005, 46, 1036–1045. [Google Scholar] [CrossRef] [PubMed]
- Ainsworth, E.A.; Rogers, A. The response of photosynthesis and stomatal conductance to rising [CO2]: Mechanisms and environmental interactions. Plant Cell Environ. 2007, 30, 258–270. [Google Scholar] [CrossRef] [PubMed]
- Ashraf, M.; Akram, N.A.; Arteca, R.N. The Physiological, Biochemical and Molecular Roles of Brassinosteroids and Salicylic Acid in Plant Processes and Salt Tolerance. Crit. Rev. Plant Sci. 2010, 29, 162–190. [Google Scholar] [CrossRef]
- Pan, C.Z.; Zhang, H.; Ma, Q.M.; Fan, F.J. Role of ethylene biosynthesis and signaling in elevated CO2-induced heat stress response in tomato. Planta 2019, 250, 563–572. [Google Scholar] [CrossRef] [PubMed]
- Romero, I.; Alegria, C.E.; Pradena, A.G. WRKY transcription factors in the response of table grapes (cv. Autumn Royal) to high CO2 levels and low temperature. Postharvest Biol. Technol. 2018, 150, 42–51. [Google Scholar]
- Zou, X.L.; Shen, Q.J.X.; Neuman, D. An ABA inducible WRKY gene integrates responses of creosote bush (Larrea tridentata) to elevated CO2 and abiotic stresses. Plant Sci. 2007, 172, 997–1004. [Google Scholar] [CrossRef]
- Ni, Z.X.; Ye, Y.J.; Bai, T.D.; Xu, M.; Xu, L.A. Complete Chloroplast Genome of Pinus massoniana (Pinaceae): Gene Rearrangements, Loss of ndh Genes, and Short Inverted Repeats Contraction, Expansion. Molecules 2017, 22, 1528. [Google Scholar] [CrossRef]
- Guo, Y. Changes in Total Stand Biomass of Pinus Massoniana Forests and Its Response to Age Class Structure in Hunan Province, China. Master’s, Thesis, Central South University of Forestry and Technology, Hunan, China, 2015. [Google Scholar]
- Justine, M.F.; Yang, W.Q.; Wu, F.Z.; Khan, M.N. Dynamics of biomass and carbon sequestration across a chronosequence of Masson pine plantations. J. Geophys. Res. Biogeosci. 2017, 122, 578–591. [Google Scholar] [CrossRef]
- Elias, M.; Potvin, C. Assessing inter-and intra-specific variation in trunk carbon concentration for 32 neotropical tree species. Can. J. For. Res. 2003, 33, 1039–1045. [Google Scholar] [CrossRef]
- Banerjee, S.; Siemianowski, O.; Liu, M.L.; Lind, K.R. Stress response to CO2 deprivation by Arabidopsis thaliana in plant cultures. PLoS ONE 2019, 14, e0212462. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Castellano, I.; Patti, F.P. Molecular response of Sargassum vulgare to acidification at volcanic CO2 vents: Insights from de novo transcriptomic analysis. Mol. Ecol. 2017, 26, 2276–2290. [Google Scholar] [CrossRef] [PubMed]
- De Souza, A.P.; Cocuron, J.C.; Garcia, A.C.; Alonso, A.P.; Buckeridge, M.S. Changes in Whole-Plant Metabolism during the Grain-Filling Stage in Sorghum Grown under Elevated CO2 and Drought. Plant Physiol. 2015, 169, 1755–1765. [Google Scholar] [CrossRef] [PubMed]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Haas, B.J.; Papanicolaou, A.; Yassour, M.; Grabherr, M.; Blood, P.D.; Bowden, J.; Couger, M.B.; Eccles, D.; Li, B.; Lieber, M.; et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 2013, 8, 1494–1512. [Google Scholar] [CrossRef] [PubMed]
- Smith-Unna, R.; Boursnell, C.; Patro, R.; Hibberd, J.M.; Kelly, S. TransRate: Reference-free quality assessment of de novo transcriptome assemblies. Genome Res. 2016, 26, 1134–1144. [Google Scholar] [CrossRef] [PubMed]
- Simão, F.A.; Waterhouse, R.M.; Ioannidis, P. BUSCO: Assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 2015, 31, 3210–3212. [Google Scholar] [CrossRef]
- Li, B.; Dewey, C.N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef]
- Wang, L.K.; Feng, Z.X.; Wang, X.; Wang, X.W.; Zhang, X.G. DEGseq: An R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics 2010, 26, 136–138. [Google Scholar] [CrossRef]
- Anders, S.; Huber, W. Differential expression analysis for sequence count data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef]
- Zhu, P.; Ma, Y.; Zhu, L.; Chen, Y.; Li, R.; Ji, K. Selection of Suitable Reference Genes in Pinus massoniana Lamb. Under Different Abiotic Stresses for qPCR Normalization. Forests 2019, 10, 632. [Google Scholar] [CrossRef]
- Fan, F.H.; Wang, Q.H.; Wen, X.P.; Ding, G.J. Transcriptome-wide identification and expression profiling of Pinus massoniana MYB transcription factors responding to phosphorus deficiency. J. For. Res. 2019, 20. [Google Scholar] [CrossRef]
- Du, M.F.; Ding, G.J.; Cai, Q. The Transcriptomic Responses of Pinus massoniana to Drought Stress. Forests 2018, 9, 326. [Google Scholar] [CrossRef]
- Sarka, E.; Dvoracek, V. Biosynthesis of waxy starch-a review. Plant Soil Environ. 2017, 63, 335–341. [Google Scholar]
- Verbancic, J.; Lunn, J.E.; Stitt, M.; Persson, S. Carbon Supply and the Regulation of Cell Wall Synthesis. Mol. Plant 2018, 11, 75–94. [Google Scholar] [CrossRef]
- Bashar, K.K.; Tareq, M.Z.; Amin, M.R.; Honi, U. Phytohormone-Mediated Stomatal Response, Escape and Quiescence Strategies in Plants under Flooding Stress. Agronomy 2019, 9, 43. [Google Scholar] [CrossRef]
- Huang, H.; Liu, B.; Liu, L.Y.; Song, S.S. Jasmonate action in plant growth and development. J. Exp. Bot. 2017, 68, 1349–1359. [Google Scholar] [CrossRef] [Green Version]
- Ha, Y.M.; Shang, Y.; Yang, D.; Nam, K.H. Brassinosteroid reduces ABA accumulation leading to the inhibition of ABA-induced stomatal closure. Biochem. Biophys. Res. Commun. 2018, 504, 143–148. [Google Scholar] [CrossRef]
- Miura, K.; Tada, Y. Regulation of water, salinity, and cold stress responses by salicylic acid. Front. Plant Sci. 2014, 5, 4. [Google Scholar] [CrossRef] [Green Version]
- Hernandez-Ruiz, J.; Arnao, M.B. Relationship of Melatonin and Salicylic Acid in Biotic/Abiotic Plant Stress Responses. Agronomy 2018, 8, 33. [Google Scholar] [CrossRef]
- Zhao, B.L.; Li, J. Regulation of Brassinosteroid Biosynthesis and Inactivation. J. Integra. Plant Biol. 2012, 54, 746–759. [Google Scholar] [CrossRef] [PubMed]
- Xie, H.; Fan, G.Z.; Jin, Y.H.; Dong, L.; Deng, F.H. Progress of Research on Photosynthetic Acclimation of Plant to Elevated Atmospheric CO2. Rev. China Agric. Sci. Technol. 2006, 8, 29–34. [Google Scholar]
- Drake, B.G.; Gonzalez-Meler, M.A.; Long, S.P. More Efficient Plants: A Consequence of Rising Atmospheric CO2. Ann. Rev. Plant Biol. 1997, 48, 609–639. [Google Scholar] [CrossRef] [PubMed]
- Sreeharsha, R.V.; Mudalkar, S.; Sengupta, D.; Unnikrishnan, D.K.; Reddy, A.R. Mitigation of drought-induced oxidative damage by enhanced carbon assimilation and an efficient antioxidative metabolism under high CO2 environment in pigeonpea (Cajanus cajan L.). Photosynth. Res. 2019, 139, 425–439. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.Y. Effect of Elevated CO2 on Photosynthetic Characteristics and Molecular Mechanism in Deyeuxia Angustifolia. Ph.D. Thesis, Northeast Agricultural University, Harbin, China, 2015. [Google Scholar]
- Xu, S.; Chen, W.; He, X.Y.; Huang, Y.Q.; Gao, J.Y.; Zhao, Y.; Li, B. Research advance in effect of elevated CO2 on eco-physiology of trees. Acta Ecol. Sin. 2015, 35, 2452–2460. [Google Scholar]
- Turnbull, M.H.; Tissue, D.T.; Griffin, K.L.; Rogers, G.N.D.; Whitehead, D. Photosynthetic acclimation to long-term exposure to elevated CO2 concentration in Pinus radiata D. Don is related to age of needles. Plant Cell Environ. 1998, 21, 1019–1028. [Google Scholar] [CrossRef]
- Fukayama, H.; Sugino, M.; Fukuda, T. Gene expression profiling of rice grown in free air CO2 enrichment (FACE) and elevated soil temperature. Field Crops Res. 2011, 121, 195–199. [Google Scholar] [CrossRef]
- Sicher, R. Combined effects of CO2 enrichment and elevated growth temperatures on metabolites in soybean leaflets: Evidence for dynamic changes of TCA cycle intermediates. Planta 2013, 238, 369–380. [Google Scholar] [CrossRef]
- Liu, L.X.; Shen, F.F.; Lu, H.C.; Han, O.D.; Liu, Y.G. Research Advance on Sucrose Phosphate Synthase in Sucrose Metabolism. Mol. Plant Breed. 2005, 3, 275–281. [Google Scholar]
- Van Le, Q.U.Y.; Samson, G.U.Y.; Desjardins, Y. Opposite effects of exogenous sucrose on growth, photosynthesis and carbon metabolism of in vitro plantlets of tomato (L. Esculentum Mill.) grown under two levels of irradiances and CO2 concentration. J. Plant Physiol. 2001, 158, 599–605. [Google Scholar] [CrossRef]
- Qian, W.; Yue, C.; Wang, Y.; Cao, H.; Li, N.; Wang, L.; Hao, X.; Wang, X.; Xiao, B.; Yang, Y. Identification of the invertase gene family (INVs) in tea plant and their expression analysis under abiotic stress. Plant Cell Rep. 2016, 35, 2269–2283. [Google Scholar] [CrossRef] [PubMed]
- Yue, C.; Cao, H.L.; Wang, L.; Zhou, Y.H.; Huang, Y.T.; Hao, X.Y.; Wang, Y.C.; Wang, B.; Yang, Y.J.; Wang, X.C. Effects of cold acclimation on sugar metabolism and sugar-related gene expression in tea plant during the winter season. Plant Mol. Biol. 2015, 88, 591–608. [Google Scholar] [CrossRef] [PubMed]
- Tetlow, I.J. Understanding storage starch biosynthesis in plants: A means to quality improvement. Can. J. Bot. 2006, 84, 1167–1185. [Google Scholar] [CrossRef]
- Cui, H. Effects of CO2 Concentration and Nitrogen Application on Grain Yield, Quality Formation and Their Physiology Mechanism in Winter Wheat. Master’s Thesis, Nanjing Agricultural University, Nanjing, China, 2011. [Google Scholar]
- Anderson, C.T.; Wallace, I.S. Illuminating the wall: Using click chemistry to image pectins in Arabidopsis cell walls. Plant Signal. Behav. 2012, 7, 661–663. [Google Scholar] [CrossRef] [PubMed]
- Diez, M.D.A.; Peiru, S.; Demonte, A.M.; Gramajo, H.; Iglesias, A.A. Characterization of Recombinant UDP-and ADP-Glucose Pyrophosphorylases and Glycogen Synthase to Elucidate Glucose-1-Phosphate Partitioning into Oligo- and Polysaccharides in Streptomyces coelicolor. J. Bacteriol. 2012, 194, 1485–1493. [Google Scholar] [CrossRef] [PubMed]
- Hwang, S.G.; Chen, H.C.; Huang, W.Y.; Chu, Y.C.; Shii, C.T.; Cheng, W.H. Ectopic expression of rice OsNCED3 in Arabidopsis increases ABA level and alters leaf morphology. Plant Sci. 2010, 178, 12–22. [Google Scholar] [CrossRef]
- De Souza, A.M.S.; Batista, V.G.L.; Pinheiro, M.P.N.; Suassuna, J.F.; de Lima, L.M.; Fernandes, P.D. Expression of NCED gene in colored cotton genotypes subjected to water stress. Rev. Bras. Eng. Agric. Ambient. 2016, 20, 692–696. [Google Scholar] [CrossRef]
- Haubrick, L.L.; Torsethaugen, G.; Assmann, S.M. Effect of brassinolide, alone and in concert with abscisic acid, on control of stomatal aperture and potassium currents of Vicia faba guard cell protoplasts. Physiol. Plant. 2006, 128, 134–143. [Google Scholar] [CrossRef]
- Hu, Y.R.; Yu, D.Q. BRASSINOSTEROID INSENSITIVE2 Interacts with ABSCISIC ACID INSENSITIVE5 to Mediate the Antagonism of Brassinosteroids to Abscisic Acid during Seed Germination in Arabidopsis. Plant Cell 2014, 26, 4394–4408. [Google Scholar] [CrossRef]
- Barton, C.V.M.; Duursma, R.A.; Medlyn, B.E.; Ellsworth, D.S. Effects of elevated atmospheric [CO2] on instantaneous transpiration efficiency at leaf and canopy scales in Eucalyptus saligna. Glob. Chang. Biol. 2012, 18, 585–595. [Google Scholar] [CrossRef]
- Gao, C.J.; Xia, X.J.; Shi, K.; Zhou, Y.H.; Yu, J.Q. Response of Stomata to Global Climate Changes and the Underlying Regulation Mechanism of Stress Responses. Plant Physiol. J. 2012, 48, 19–28. [Google Scholar]
- Islam, M.M.; Munemasa, S.; Hossain, M.A.; Nakamura, Y.; Mori, I.C.; Murata, Y. Roles of AtTPC1, Vacuolar Two Pore Channel 1, in Arabidopsis Stomatal Closure. Plant Cell Physiol. 2010, 51, 302–311. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.J.; Fan, L.; Loescher, W.; Duan, W.; Liu, G.J. Salicylic acid alleviates decreases in photosynthesis under heat stress and accelerates recovery in grapevine leaves. BMC Plant Biol. 2010, 10, 34. [Google Scholar] [CrossRef] [PubMed]

| CK | 6 h | 12 h | 24 h | |
|---|---|---|---|---|
| Raw reads | 49,314,299 | 47,459,322 | 45,980,036 | 60,876,932 |
| Raw bases | 7,446,459,199 | 7,166,357,723 | 6,942,985,436 | 9,192,416,732 |
| Clean reads | 48,795,571 | 46,976,134 | 45,496,760 | 60,205,674 |
| Clean bases | 7,292,383,178 | 7,017,302,463 | 6,802,876,221 | 9,000,454,730 |
| Error rate (%) | 0.02 | 0.02 | 0.03 | 0.03 |
| Mapped reads | 16,879,027 | 16,459,249 | 15,859,340 | 21,108,414 |
| Mapped ratio (%) | 0.69 | 0.70 | 0.70 | 0.70 |
| GC content (%) | 47.47 | 46.74 | 46.24 | 46.32 |
| Q20 (%) | 98.12 | 98.20 | 98.06 | 97.86 |
| Q30 (%) | 94.04 | 94.22 | 93.86 | 93.40 |
| Type | Transcript | Unigenes |
|---|---|---|
| <500 bp | 61,696 | 48,592 |
| 501~1000 bp | 37,586 | 22,267 |
| 1001~2000 bp | 25,994 | 12,887 |
| >2000 bp | 15,587 | 8,678 |
| Total | 140,863 | 92,424 |
| Min length (bp) | 201 | 201 |
| Max length (bp) | 15,491 | 15,491 |
| Mean length (bp) | 891 | 935 |
| N50 (bp) | 1463 | 1550 |
| TransRate score | 0.20045 | 0.30498 |
| BUSCO score | 77.7% | 74.2% |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, F.; Sun, X.; Zou, B.; Zhu, P.; Lin, N.; Lin, J.; Ji, K. Transcriptional Analysis of Masson Pine (Pinus massoniana) under High CO2 Stress. Genes 2019, 10, 804. https://doi.org/10.3390/genes10100804
Wu F, Sun X, Zou B, Zhu P, Lin N, Lin J, Ji K. Transcriptional Analysis of Masson Pine (Pinus massoniana) under High CO2 Stress. Genes. 2019; 10(10):804. https://doi.org/10.3390/genes10100804
Chicago/Turabian StyleWu, Fan, Xiaobo Sun, Bingzhang Zou, Peihuang Zhu, Nengqing Lin, Jingquan Lin, and Kongshu Ji. 2019. "Transcriptional Analysis of Masson Pine (Pinus massoniana) under High CO2 Stress" Genes 10, no. 10: 804. https://doi.org/10.3390/genes10100804
APA StyleWu, F., Sun, X., Zou, B., Zhu, P., Lin, N., Lin, J., & Ji, K. (2019). Transcriptional Analysis of Masson Pine (Pinus massoniana) under High CO2 Stress. Genes, 10(10), 804. https://doi.org/10.3390/genes10100804





