Overexpression of Rice Rab7 Gene Improves Drought and Heat Tolerance and Increases Grain Yield in Rice (Oryza sativa L.)
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material and Growth Conditions
2.2. Plasmid Construction and Rice Transformation
2.3. Molecular Analysis of Transgenic Plants and Transgene Expression
2.4. Drought and Heat Stress Treatments
2.5. Determination of Growth Traits and Relative Water Content
2.6. Estimation of Oxidative Stress Biomarker Levels
2.7. Estimation of Gas-Exchange Characteristics
2.8. Determination of Soluble Proteins, Soluble Sugars, Proline and Chlorophyll Contents
2.9. Antioxidant Enzyme Assays
2.10. Stress-Related Genes Expression Analysis
2.11. Statistical Analysis
3. Results and Discussion
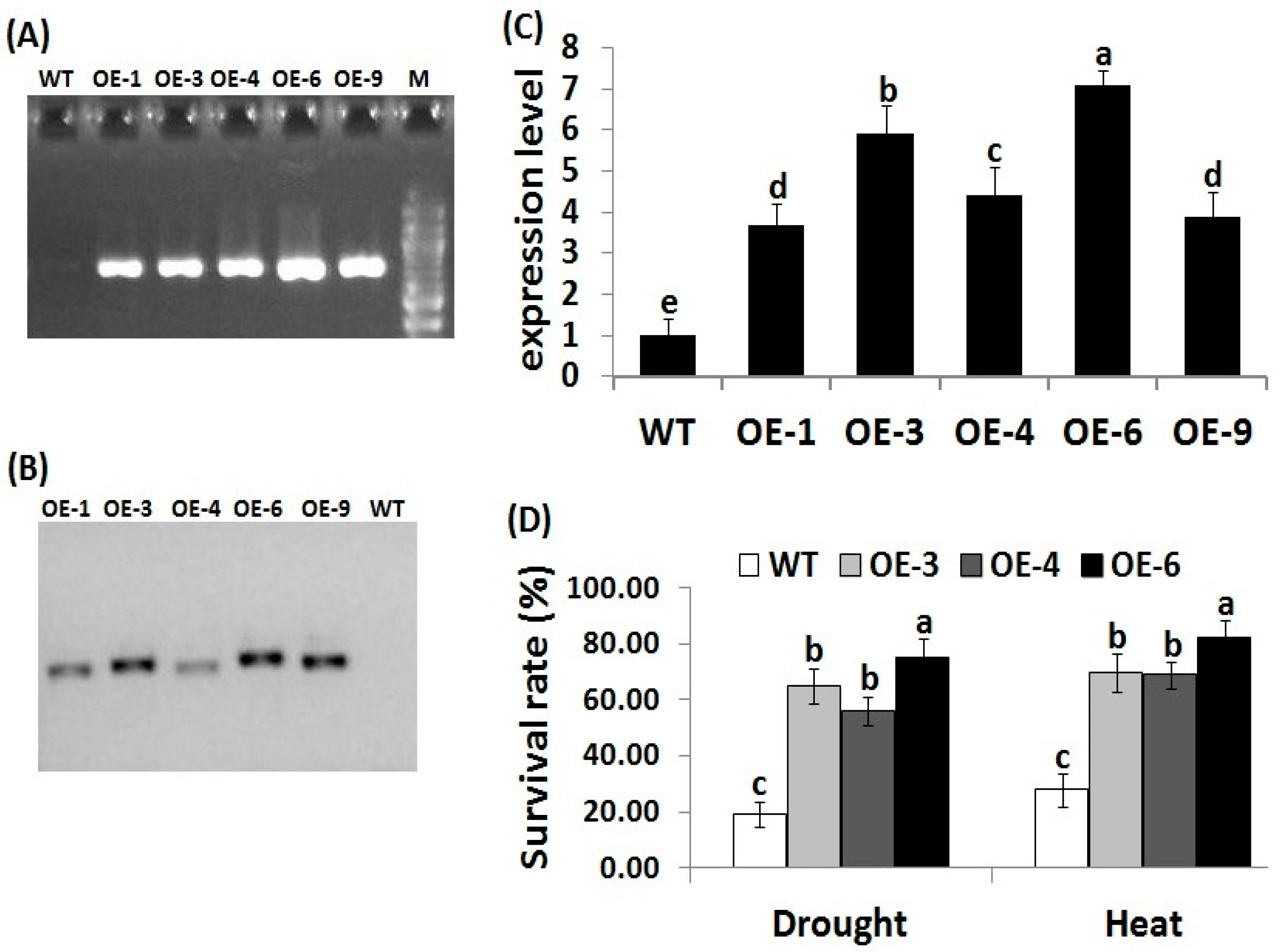
3.1. Transformation and Molecular Analysis of Transgenic Rice Lines
3.2. OsRab7 Overexpression in Rice Enhances Survival Rate, Plant Growth, and Relative Water Content under Drought and Heat Stress Conditions
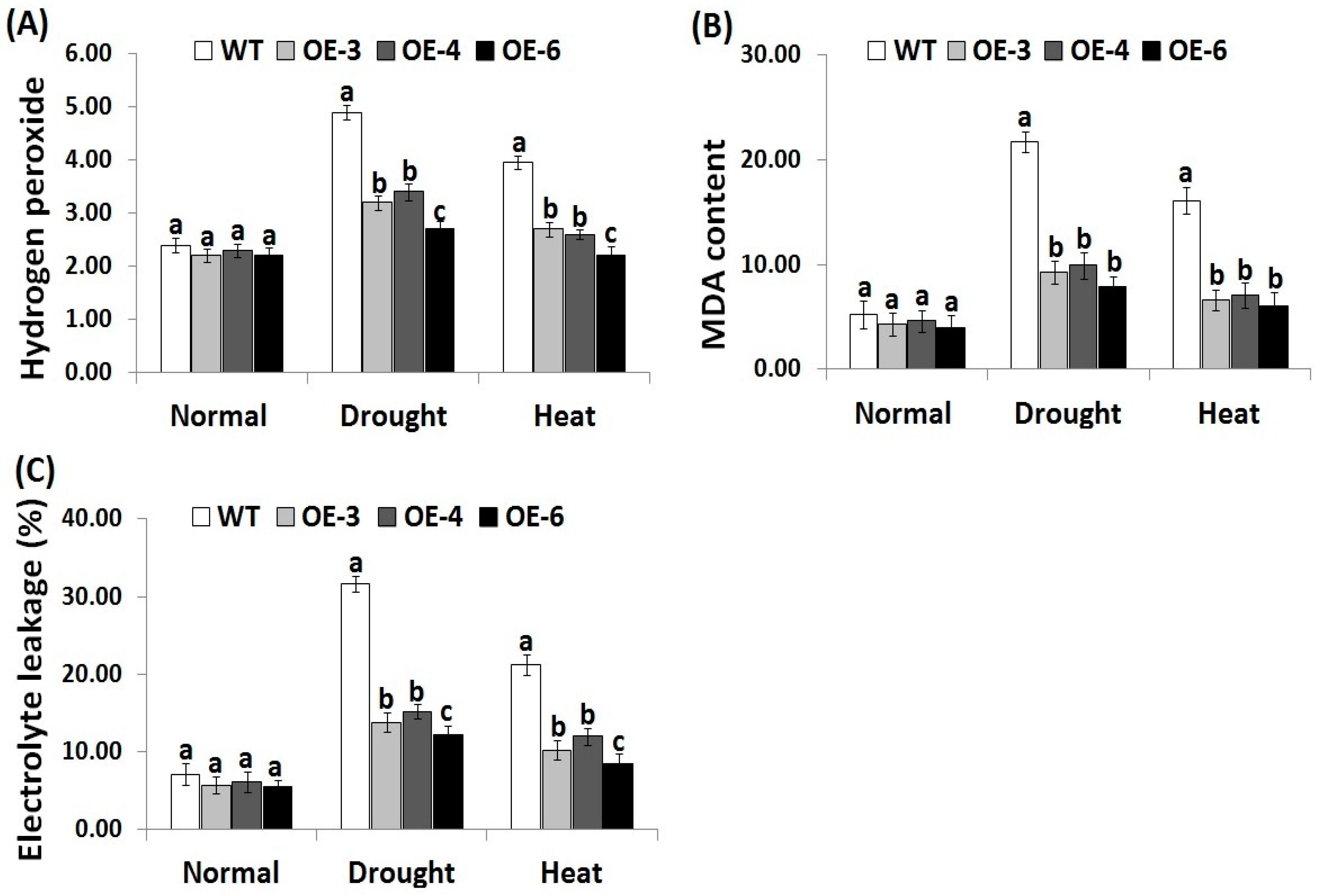
3.3. OsRab7 Overexpression in Rice Reduces Oxidative Stress Biomarkers under Drought and Heat Stress Conditions
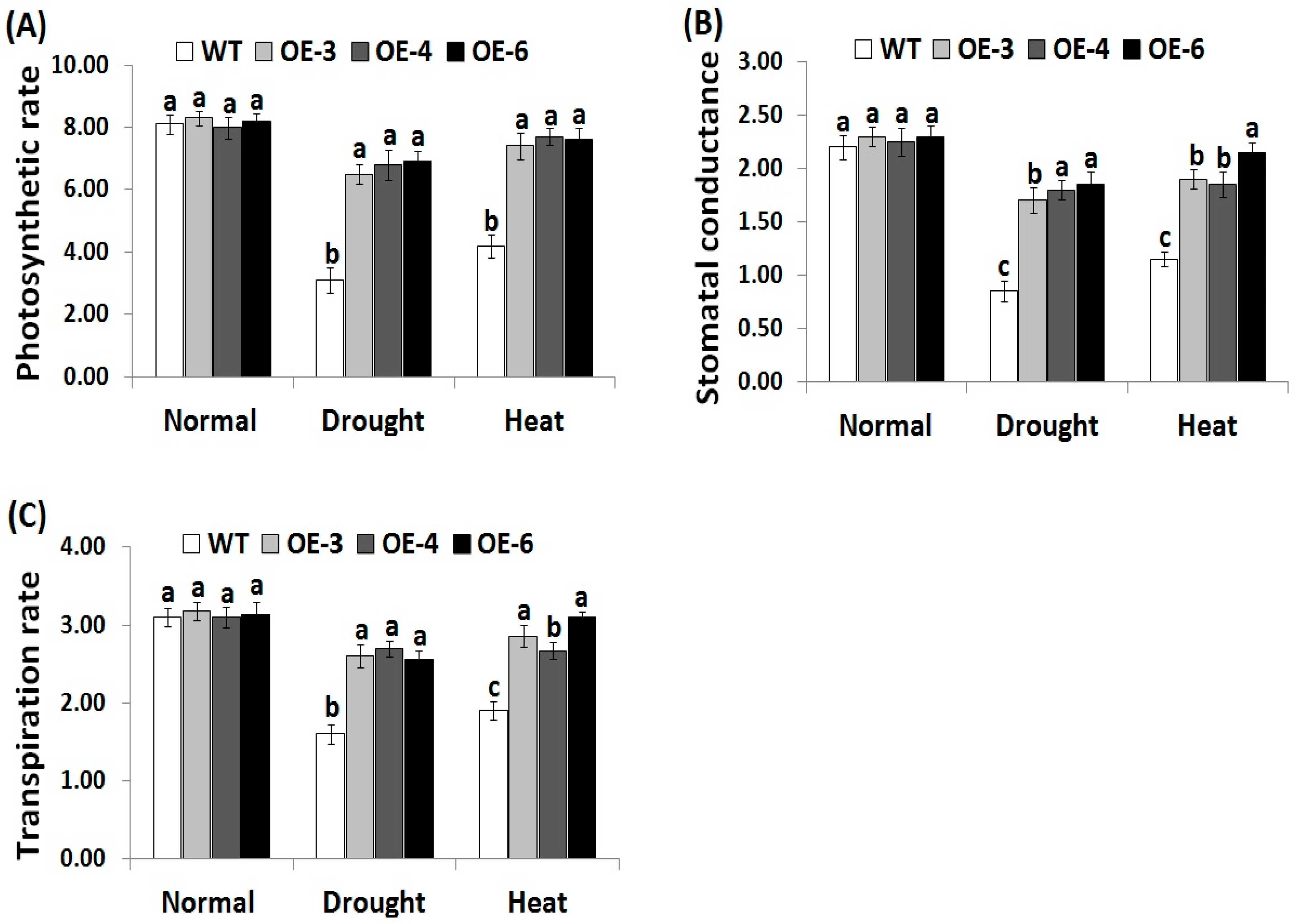
3.4. OsRab7 Overexpression in Rice Improves Gas-Exchange Characteristics Under Drought and Heat Stress Conditions
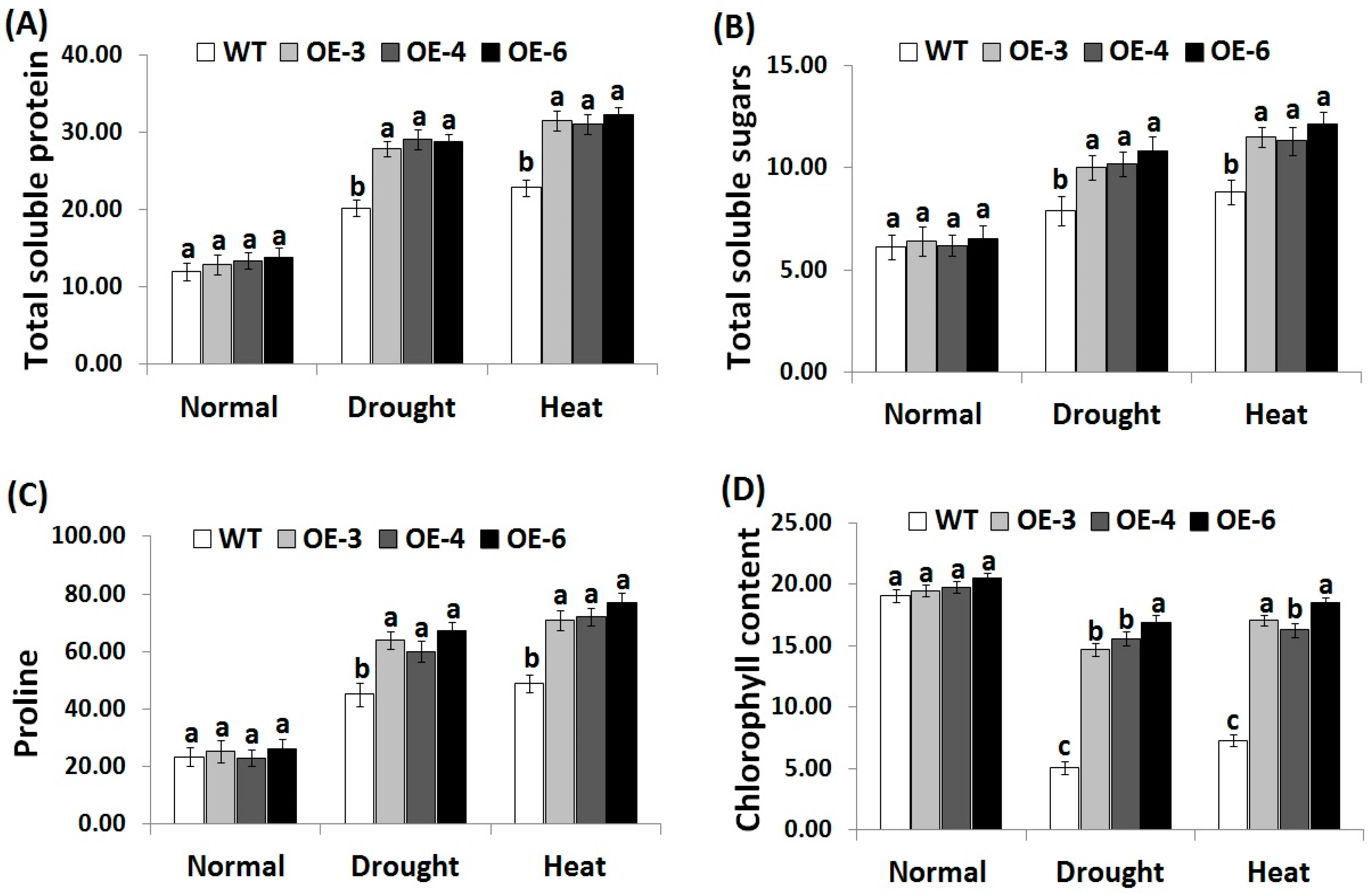
3.5. OsRab7 Overexpression in Rice Improves Osmolytes and Chlorophyll Content under Drought and Heat Stress Conditions
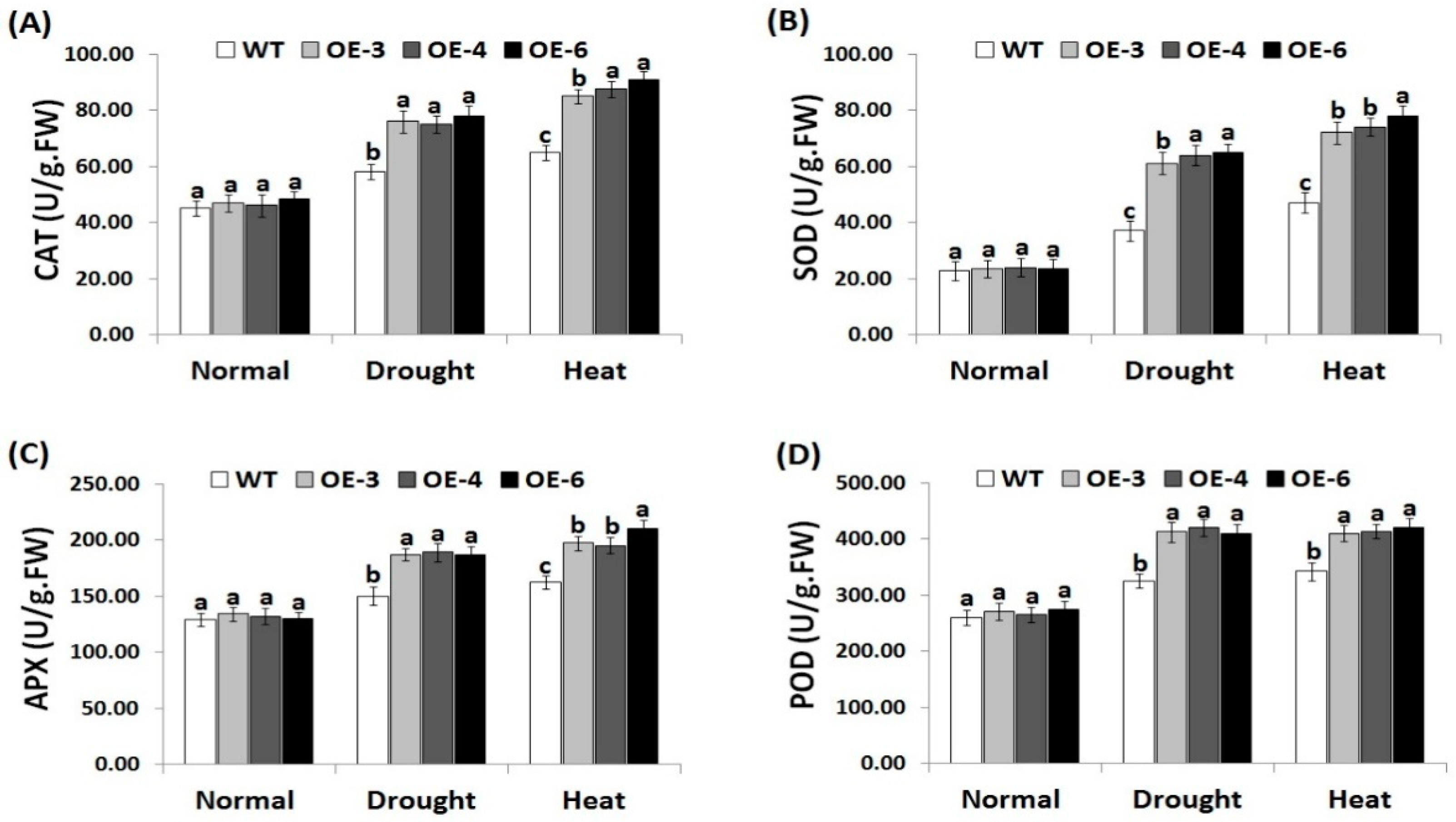
3.6. OsRab7 Overexpression in Rice Induces Antioxidant Enzyme Activities under Drought and Heat Stress Conditions
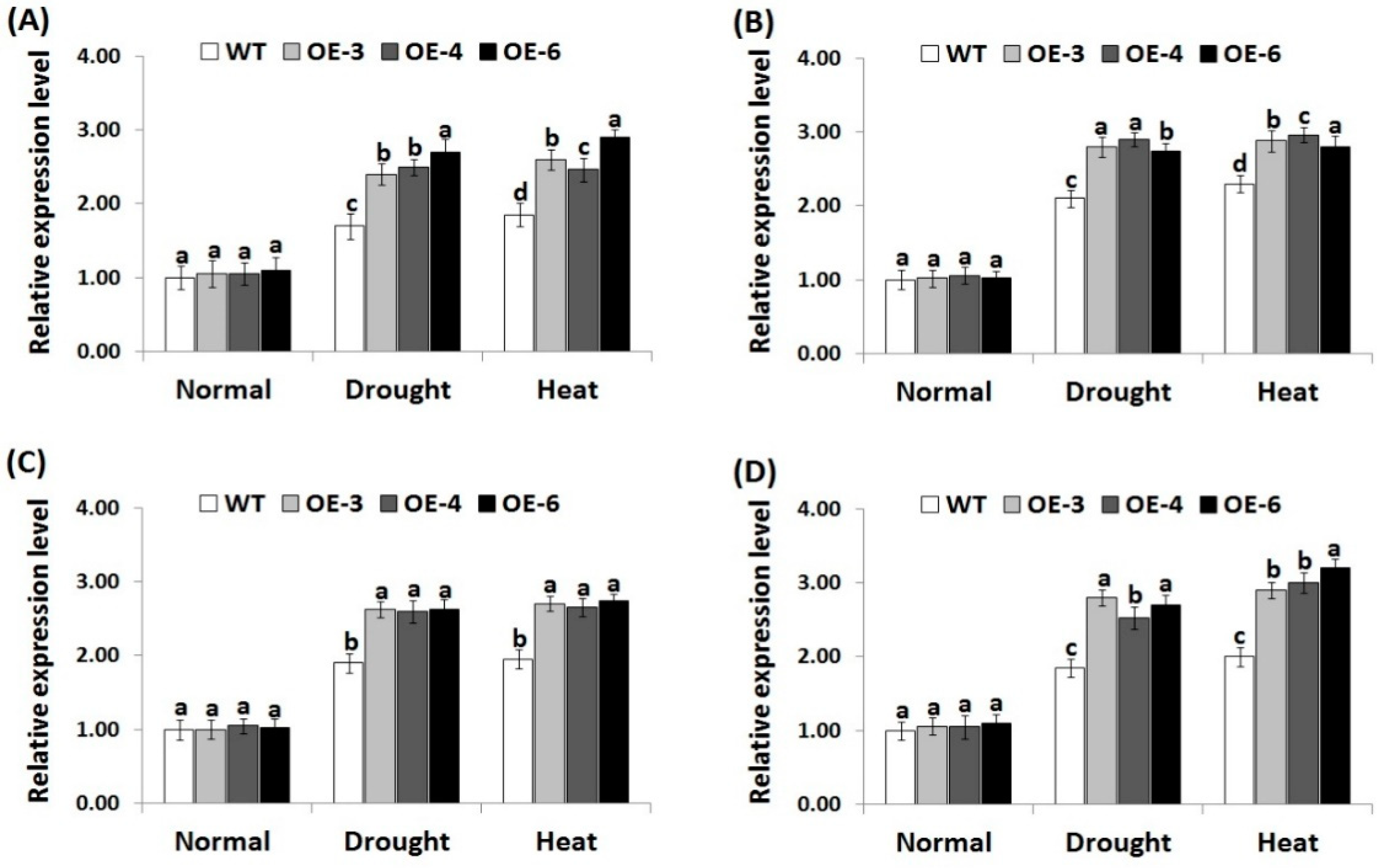
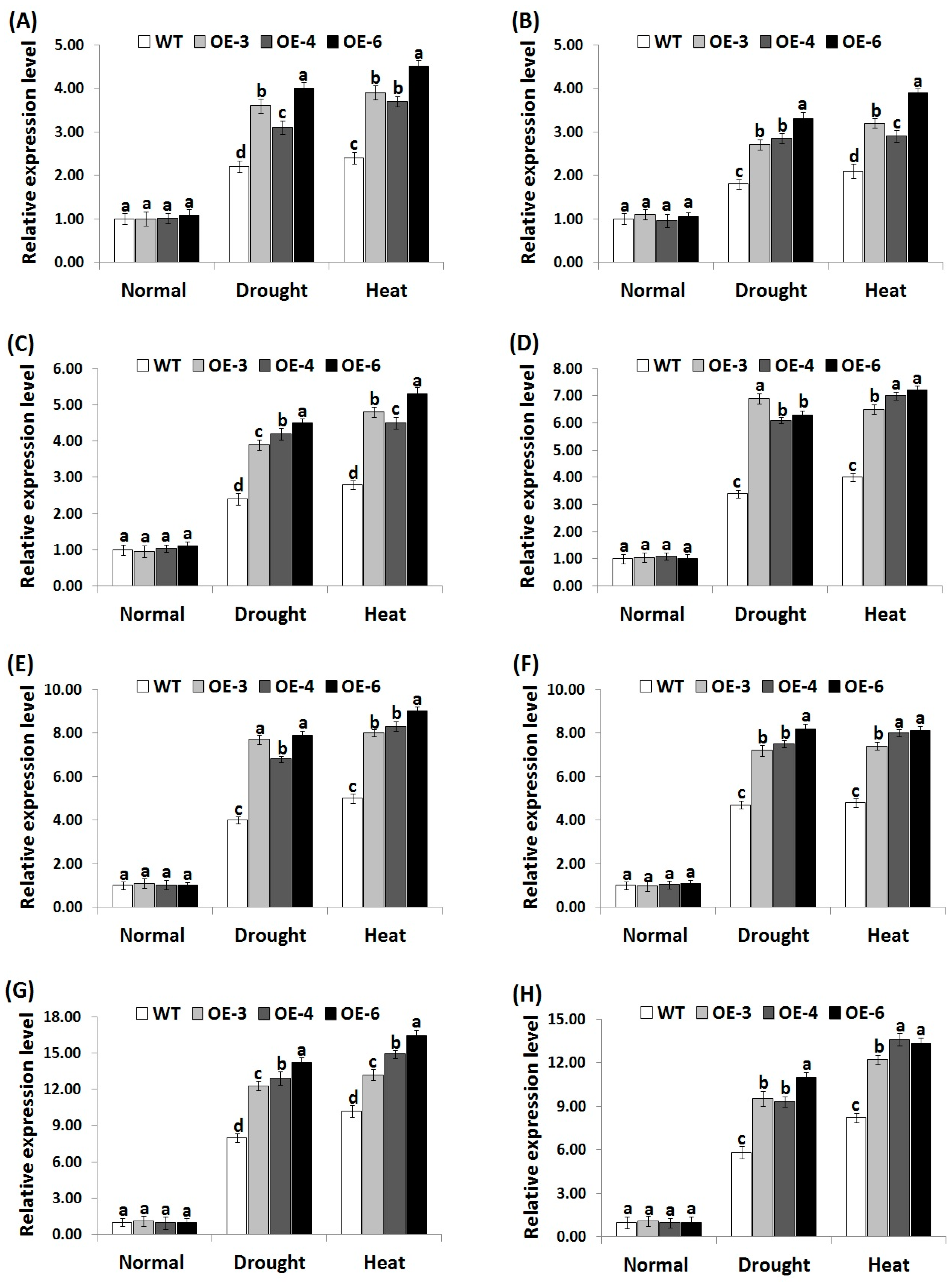
3.7. OsRab7 Overexpression in Rice Induces Abiotic Stress-Related Genes Expression under Drought and Heat Stress Conditions
3.8. OsRab7 Overexpression in Rice Increases Grain Yield under Drought and Heat Stress Conditions
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Zang, X.; Geng, X.; He, K.; Wang, F.; Tian, X.; Xin, M.; Yao, Y.; Hu, Z.; Ni, Z.; Sun, Q.; et al. Overexpression of the Wheat (Triticum aestivum L.) TaPEPKR2 gene enhances heat and dehydration tolerance in both wheat and Arabidopsis. Front. Plant Sci. 2018, 9, 1710. [Google Scholar] [CrossRef] [PubMed]
- Ahuja, I.; de Vos, R.C.; Bones, A.M.; Hall, R.D. Plant molecular stress responses face climate change. Trends Plant Sci. 2010, 15, 664–674. [Google Scholar] [CrossRef] [PubMed]
- Zhou, R.; Yu, X.; Ottosen, C.; Rosenqvist, E.; Zhao, L.; Wang, Y.; Yu, W.; Zhao, T.; Wu, Z. Drought stress had a predominant effect over heat stress on three tomato cultivars subjected to combined stress. BMC Plant Biol. 2017, 17, 24. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Wang, Z.; Ke, Q.; Ji, C.Y.; Jeong, J.C.; Lee, H.S.; Lim, Y.P.; Xu, B.; Deng, X.; Kwak, S.S. Overexpression of codA gene confers enhanced tolerance to abiotic stresses in alfalfa. Plant Physiol. Biochem. 2014, 85, 31–40. [Google Scholar] [CrossRef] [PubMed]
- El-Esawi, M.A.; Al-Ghamdi, A.A.; Ali, H.M.; Alayafi, A.A.; Witczak, J.; Ahmad, M. Analysis of genetic variation and enhancement of salt tolerance in French pea. Int. J. Mol. Sci. 2018, 19, 2433. [Google Scholar] [CrossRef] [PubMed]
- El-Esawi, M.A.; Alaraidh, I.A.; Alsahli, A.A.; Ali, H.M.; Alayafi, A.A.; Witczak, J.; Ahmad, M. Genetic variation and alleviation of salinity stress in barley. Molecules 2018, 23, 2488. [Google Scholar] [CrossRef] [PubMed]
- El-Esawi, M.A.; Alaraidh, I.A.; Alsahli, A.A.; Alamri, S.A.; Ali, H.M.; Alayafi, A.A. Bacillus firmus (SW5) augments salt tolerance in soybean (Glycine max L.) by modulating root system architecture, antioxidant defense systems and stress-responsive genes expression. Plant Physiol. Biochem. 2018, 132, 375–384. [Google Scholar] [CrossRef]
- El-Esawi, M.A.; Alaraidh, I.A.; Alsahli, A.A.; Alzahrani, S.M.; Ali, H.M.; Alayafi, A.A.; Ahmad, M. Serratia liquefaciens KM4 improves salt stress tolerance in maize by regulating redox potential, ion homeostasis, leaf gas exchange and stress-related gene expression. Int. J. Mol. Sci. 2018, 19, 3310. [Google Scholar] [CrossRef]
- El-Esawi, M.A.; Al-Ghamdi, A.A.; Ali, H.M.; Alayafi, A.A. Azospirillum lipoferum FK1 confers improved salt tolerance in chickpea (Cicer arietinum L.) by modulating osmolytes, antioxidant machinery and stress-related genes expression. Environ. Exp. Bot. 2019, 159, 55–65. [Google Scholar] [CrossRef]
- Nuoffer, C.; Balch, W.E. GTPases: Multifunctional molecular switches regulating/vesicular traffic. Annu. Rev. Biochem. 1994, 63, 949–990. [Google Scholar] [CrossRef]
- Pitakrattananukool, S.; Kawakatsu, T.; Anuntalabhochal, S.; Takaiwa, F. Overexpression of OsRab7B3, a small GTP binding protein gene, enhances leaf senescence in transgenic rice. Biosci. Biotechnol. Biochem. 2012, 76, 1296–1302. [Google Scholar] [CrossRef] [PubMed]
- Takai, Y.; Sasaki, T.; Matozaki, T. Small GTP-binding proteins. Physiol. Rev. 2001, 81, 153–208. [Google Scholar] [CrossRef] [PubMed]
- Sohn, E.J.; Kim, E.S.; Zhao, M.; Kim, S.J.; Kim, H.; Kim, Y.W.; Lee, Y.J.; Hillmer, S.; Sohn, U.; Jiang, L.; et al. Rha1, an Arabidopsis Rab5 homolog, plays a critical role in the vacuolar trafficking of soluble cargo proteins. Plant Cell 2003, 15, 1057–1070. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, P.; Reddy, M.K.; Sopory, S.K.; Agarwal, P.K. Plant rabs: Characterization, functional diversity and role in stress tolerance. Plant Mol. Biol. Rep. 2009, 27, 417–430. [Google Scholar] [CrossRef]
- Pereira-Leal, J.B.; Seabra, M.C. Evolution of the Rab family of small GTP-binding Proteins. J. Mol. Biol. 2001, 313, 889–901. [Google Scholar] [CrossRef] [PubMed]
- Mazel, A.; Leshem, Y.; Tiwari, B.S.; Levine, A. Induction of salt and osmotic stress tolerance by overexpression of an intracellular vesicle trafficking protein AtRab7 (AtRabG3e). Plant Physiol. 2004, 134, 118–128. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Xia, M.; Chen, Q.; Wu, Z.; Wu, P. Identification of a new small GTP-binding protein gene OsRab5a, genomic organization, and expression pattern analysis during nitrate supply and early nutrient starvation in rice (Oryza sativa L.) root. Plant Sci. 2002, 163, 273–280. [Google Scholar] [CrossRef]
- Wang, Y.; Ren, Y.; Liu, X.; Jiang, L.; Chen, L.; Han, X.; Jin, M.; Liu, S.; Liu, F.; Lu, J.; et al. OsRab5a regulates endomembrane organization and storage protein trafficking in rice endosperm cells. Plant J. 2010, 64, 812–824. [Google Scholar] [CrossRef] [PubMed]
- Fukuda, M.; Satoh-Cruz, M.; Wen, L.; Crofts, A.J.; Sugino, A.; Washida, H.; Okita, T.W.; Ogawa, M.; Kawagoe, Y.; Maeshima, M.; et al. The small GTPase Rab5a is essential for intracellular transport of proglutelin from the Golgi apparatus to the protein storage vacuole and endosomal membrane organization in developing rice endosperm. Plant Physiol. 2011, 157, 632–644. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, P.K.; Agarwal, P.; Jain, P.; Jha, B.; Reddy, M.K.; Sopory, S.K. Constitutive overexpression of a stress-inducible small GTP-binding protein PgRab7 from Pennisetum glaucum enhances abiotic stress tolerance in transgenic tobacco. Plant Cell Rep. 2008, 27, 105–115. [Google Scholar] [CrossRef] [PubMed]
- Kwon, S.; Cho, H.J.; Bae, K.; Jung, J.H.; Jin, H.C.; Park, O.K. Role of an Arabidopsis Rab GTPase RabG3b in pathogen response and leaf senescence. J. Plant Biol. 2009, 52, 79–87. [Google Scholar] [CrossRef]
- Peng, X.; Ding, X.; Chang, T.; Wang, Z.; Liu, R.; Zeng, X.; Cai, Y.; Zhu, Y. Overexpression of a vesicle trafficking gene, OsRab7, enhances salt tolerance in rice. Sci. World J. 2014, 2014, 483526. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Shiroto, Y.; Kishitani, S.; Ito, Y.; Toriyama, K. Enhanced heat and drought tolerance in transgenic rice seedlings overexpressing OsWRKY11 under the control of HSP101 promoter. Plant Cell Rep. 2009, 28, 21–30. [Google Scholar] [CrossRef] [PubMed]
- Aghamolki, M.T.K.; Yusop, M.K.; Oad, F.C.; Zakikhani, H.; Jaafar, H.Z.; Kharidah, S.; Musa, M.H. Heat stress effects on yield parameters of selected rice cultivars at reproductive growth stages. J. Food Agric. Environ. 2014, 12, 741–746. [Google Scholar]
- Kilasi, N.L.; Singh, J.; Vallejos, C.E.; Ye, C.; Jagadish, S.V.K.; Kusolwa, P.; Rathinasabapathi, B. Heat stress tolerance in rice (Oryza sativa L.): Identification of quantitative trait loci and candidate genes for seedling growth under heat stress. Front. Plant Sci. 2018, 9, 1578. [Google Scholar] [CrossRef] [PubMed]
- Liu, A.L.; Zou, J.; Liu, C.F.; Zhou, X.Y.; Zhang, X.W.; Luo, G.Y.; Chen, X.B. Over-expression of OsHsfA7 enhanced salt and drought tolerance in transgenic rice. BMB Rep. 2013, 46, 31–36. [Google Scholar] [CrossRef] [PubMed]
- Xiao, B.; Huang, Y.; Tang, N.; Xiong, L. Over-expression of a LEA gene in rice improves drought resistance under the field conditions. Theor. Appl. Genet. 2007, 115, 35–46. [Google Scholar] [CrossRef] [PubMed]
- Shim, J.S.; Oh, N.; Chung, P.J.; Kim, Y.S.; Choi, Y.D.; Kim, J.-K. Overexpression of OsNAC14 improves drought tolerance in rice. Front. Plant Sci. 2018, 9, 310. [Google Scholar] [CrossRef]
- Huang, L.; Wang, Y.; Wang, W.; Zhao, X.; Qin, Q.; Sun, F.; Hu, F.; Zhao, Y.; Li, Z.; Fu, B.; et al. Characterization of transcription factor gene OsDRAP1 conferring drought tolerance in rice. Front. Plant Sci. 2018, 9, 94. [Google Scholar] [CrossRef]
- Zeng, D.E.; Hou, P.; Xiao, F.L.Y. Overexpression of Arabidopsis XERICO gene confers enhanced drought and salt stress tolerance in rice (Oryza Sativa L.). J. Plant Biochem. Biotechnol. 2013, 24, 56–64. [Google Scholar] [CrossRef]
- Xiong, H.; Yu, J.; Miao, J.; Li, J.; Zhang, H.; Wang, X.; Liu, P.; Zhao, Y.; Jiang, C.; Yin, Z. Natural variation in OsLG3 increases drought tolerance in rice by inducing ROS scavenging. Plant Physiol. 2018, 178, 451–467. [Google Scholar] [CrossRef] [PubMed]
- Xiang, J.; Chen, X.; Hu, W.; Xiang, Y.; Yan, M.; Wang, J. Overexpressing heat-shock protein OsHSP50.2 improves drought tolerance in rice. Plant Cell Rep. 2018, 37, 1585–1595. [Google Scholar] [CrossRef] [PubMed]
- Sato, H.; Todaka, D.; Kudo, M. The Arabidopsis transcriptional regulator DPB3-1 enhances heat stress tolerance without growth retardation in rice. Plant Biotechnol. J. 2016, 14, 1756–1767. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Zhang, C.; Wei, C.; Liu, X.; Wang, M.; Yu, F.; Xie, Q.; Tu, J. The RING finger ubiquitin E3 ligase OsHTAS enhances heat tolerance by promoting H2O2-induced stomatal closure in rice. Plant Physiol. 2016, 170, 429–443. [Google Scholar] [CrossRef] [PubMed]
- Chen, R.; Zhao, X.; Shao, Z.; Wei, Z.; Wang, Y.; Zhu, L.; Zhao, J.; Sun, M.; He, R.; He, G. Rice UDP-glucose pyrophosphorylase1 is essential for pollen callose deposition and its cosuppression results in a new type of thermosensitive genic male sterility. Plant Cell 2007, 19, 847–861. [Google Scholar] [CrossRef] [PubMed]
- Liang, Z.; Ma, D.; Tang, L.; Hong, Y.; Luo, A.; Zhou, J.; Dai, X. Expression of the spinach betaine aldehyde dehydrogenase (BADH) gene in transgenic tobacco plants. Chin. J. Biotechnol. 1997, 13, 153. [Google Scholar] [PubMed]
- Yamasaki, S.; Dillenburg, L.C. Measurements of leaf relative water content in Araucaria angustifolia. Rev. Bras. Fisiol. Veg. 1999, 11, 69–75. [Google Scholar]
- Dionisio-Sese, M.L.; Tobita, S. Antioxidant responses of rice seedlings to salinity stress. Plant Sci. 1998, 135, 1–9. [Google Scholar] [CrossRef]
- Velikova, V.; Yordanov, I.; Edreva, A. Oxidative stress and some antioxidant systems in acid rain-treated bean plants: Protective role of exogenous polyamines. Plant Sci. 2000, 151, 59–66. [Google Scholar] [CrossRef]
- Rao, K.V.M.; Sresty, T.V.S. Antioxidative parameters in the seedlings of pigeon pea (Cajanus cajan L. Millspaugh) in response to Zn and Ni stresses. Plant Sci. 2000, 157, 113–128. [Google Scholar]
- Holá, D.; Benešová, M.; Honnerová, J.; Hnilička, F.; Rothová, O.; Kočová, M.; Hniličková, H. The evaluation of photosynthetic parameters in maize inbred lines subjected to water deficiency: Can these parameters be used for the prediction of performance of hybrid progeny? Photosynthetica 2010, 4, 545–558. [Google Scholar] [CrossRef]
- Bradford, M.M. A rapid and sensitive method for the quantification of microgram quantities of protein utilizing the principle of protein dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Dey, P.M. Oligosaccharides. In Methods in Plant Biochemistry, Carbohydrates; Dey, P.M., Ed.; Academic Press: London, UK, 1990; Volume 2, pp. 189–218. [Google Scholar]
- Bates, L.; Waldren, P.P.; Teare, J.D. Rapid determination of free proline of water stress studies. Plant Soil 1973, 39, 205–207. [Google Scholar] [CrossRef]
- Aebi, H. Catalase in vitro. Method Enzymol. 1984, 105, 121–126. [Google Scholar]
- Zhang, X.Z. The measurement and mechanism of lipid peroxidation and SOD, POD and CAT activities in biological system. In Research Methodology of Crop Physiology; Zhang, X.Z., Ed.; Agriculture Press: Beijing, China, 1992; pp. 208–211. [Google Scholar]
- Nakano, Y.; Asada, K. Hydrogen peroxide is scavenged by ascorbate-specific peroxidase in spinach chloroplasts. Plant Cell Physiol. 1981, 22, 867–880. [Google Scholar]
- Vighi, I.L.; Benitez, L.C.; Amaral, M.N.; Moraes, G.P.; Auler, P.A.; Rodrigues, G.S.; Deuner, S.; Maia, L.C.; Braga, E.J.B. Functional characterization of the antioxidant enzymes in rice plants exposed to salinity stress. Biol. Plant. 2017, 61, 1–11. [Google Scholar] [CrossRef]
- Cai, W.; Liu, W.; Wang, W.S.; Fu, Z.W.; Han, T.T.; Lu, Y.T. Overexpression of rat neurons nitric oxide synthase in rice enhances drought and salt tolerance. PLoS ONE 2015, 10, e0131599. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Lai, Y.; Wu, X.; Wu, G.; Guo, C. Overexpression of OsEm1 encoding a group I LEA protein confers enhanced drought tolerance in rice. Biochem. Biophys. Res. Commun. 2016, 478, 703–709. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Siddiqui, M.H.; Al-Khaishany, M.Y.; Al-Qutami, M.A.; Al-Whaibi, M.H.; Grover, A.; Ali, H.M.; Al-Wahibi, M.S. Morphological and physiological characterization of different genotypes of faba bean under heat stress. Saudi J. Biol. Sci. 2015, 22, 656–663. [Google Scholar] [CrossRef]
- Sekmen, A.H.; Ozgur, R.; Uzilday, B.; Turkan, I. Reactive oxygen species scavenging capacities of cotton (Gossypium hirsutum) cultivars under combined drought and heat induced oxidative stress. Environ. Exp. Bot. 2014, 99, 141–149. [Google Scholar] [CrossRef]
- Alexieva, V.; Sergiev, I.; Mapelli, S.; Karanov, E. The effect of drought and ultraviolet radiation on growth and stress markers in pea and wheat. Plant Cell Environ. 2001, 24, 1337–1344. [Google Scholar] [CrossRef]
- Bajji, M.; Kinet, J.; Lutts, S. The use of the electrolyte leakage method for assessing cell membrane stability as a water stress tolerance test in durum wheat. Plant Growth Regul. 2002, 36, 61–70. [Google Scholar] [CrossRef]
- Ganguli, L.J. Lipid peroxidation-DNA damage by malondialdehyde. Mutat. Res. Fundam. Mol. Mech. 1999, 424, 83–95. [Google Scholar]
- Ben, K.R.; Abdelly, C.; Savouré, A. Proline, a multifunctional amino-acid involved in plant adaptation to environmental constraints. Biol. Aujourdhui 2012, 206, 291. [Google Scholar]
- Chaleff, R.S. Further characterization of picloram tolerant mutance of Nicotinana tabacum. Theor. Appl. Genet. 1980, 58, 91–95. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Liu, P.; Zhu, J. Effect of magnesium (Mg) on contents of free proline, soluble sugar and protein in soybean leaves. J. Henan Agric. Sci. 2004, 6, 35–38. [Google Scholar]
- Nahm, M.Y.; Kim, S.W.; Yun, D.; Lee, S.Y.; Cho, M.J.; Bahk, J.D. Molecular and biochemical analyses of OsRab7, a rice Rab7 homolog. Plant Cell Physiol. 2003, 44, 1341–1349. [Google Scholar] [CrossRef]
- Saito, C.; Ueda, T.; Abe, H.; Wada, Y.; Kuroiwa, T.; Hisada, A.; Furuya, M.; Nakano, A. A complex and mobile structure forms a distinct subregion within the continuous vacuolar membrane in young cotyledons of Arabidopsis. Plant J. 2002, 29, 245–255. [Google Scholar] [CrossRef]
- Zerial, M.; McBride, H. Rab proteins as membrane organizers. Nat. Rev. Mol. Cell Biol. 2001, 2, 107–117. [Google Scholar] [CrossRef]
- Limpens, E.; Lvanov, S.; van Esse, W.; Voets, G.; Fedorova, E.; Bisseling, T. Medicago N2-Fixing symbiosomes acquire the endocytic identity marker Rab7 but delay the acquisition of vacuolar identity. Plant Cell 2009, 21, 2811–2828. [Google Scholar] [CrossRef] [PubMed]
- Levine, A. Regulation of stress responses by intracellular vesicle trafficking? Plant Physiol. Biochem. 2002, 40, 531–535. [Google Scholar] [CrossRef]
- Zhu, J.K. Salt and drought stress signal transduction in plants. Annu. Rev. Plant Biol. 2002, 53, 247–273. [Google Scholar] [CrossRef] [PubMed]
- Oh, S.J.; Kim, Y.S.; Kwon, C.W.; Park, H.K.; Jeong, J.S.; Kim, J.K. Overexpression of the transcription factor AP37 in rice improves grain yield under drought conditions. Plant Physiol. 2009, 150, 1368–1379. [Google Scholar] [CrossRef] [PubMed]
- Hakata, M.; Kuroda, M.; Ohsumi, A.; Hirose, T.; Nakamura, H.; Muramatsu, M.; Ichikawa, H.; Yamakawa, H. Overexpression of a rice TIFY gene increases grain size through enhanced accumulation of carbohydrates in the stem. Biosci. Biotechnol. Biochem. 2012, 76, 2129–2134. [Google Scholar] [CrossRef] [PubMed]
- Jiang, D.; Chen, W.; Dong, J.; Li, J.; Yang, F.; Wu, Z.; Zhou, H. Overexpression of OsmiR164b-resistant OsNAC2 improves plant architecture and grain yield in rice. J. Exp. Bot. 2018, 69, 1533–1543. [Google Scholar] [CrossRef] [PubMed]
- Luo, B.; Chen, J.; Zhu, L.; Liu, S.; Li, B.; Lu, H.; Ye, G.; Xu, G.; Fan, X. Overexpression of a high-affinity nitrate transporter OsNRT2.1 increases yield and manganese accumulation in rice under alternating wet and dry condition. Front. Plant Sci. 2018, 9, 1192. [Google Scholar] [CrossRef] [PubMed]
| Treatment | Line | Plant Height (cm) | Root Fresh Weight (mg) | Shoot Fresh Weight (mg) | Relative Water Content (%) |
|---|---|---|---|---|---|
| Normal | WT | 27.3 ± 0.94 a | 8.4 ± 0.24 a | 23.3 ± 0.56 a | 93.0 ± 6.8 a |
| OE-3 | 27.1 ± 0.78 a | 8.6 ± 0.31 a | 22.8 ± 0.51 a | 92.1 ± 4.8 a | |
| OE-4 | 26.7 ± 0.88 a | 7.9 ± 0.42 a | 23.4 ± 0.44 a | 94.4 ± 5.5 a | |
| OE-6 | 27.4 ± 0.78 a | 8.2 ± 0.32 a | 23.9 ± 0.49 a | 95.2 ± 4.9 a | |
| Drought | WT | 19.4 ± 0.66 b | 4.9 ± 0.29 b | 16.4 ± 0.51 b | 40.2 ± 5.6 b |
| OE-3 | 25.5 ± 0.84 a | 7.9 ± 0.33 a | 21.9 ± 0.62 a | 68.6 ± 4.8 a | |
| OE-4 | 25.1 ± 0.82 a | 7.8 ± 0.41 a | 22.2 ± 0.53 a | 66.2 ± 6.4 a | |
| OE-6 | 26.4 ± 0.77 a | 8.1 ± 0.37 a | 23.3 ± 0.66 a | 73.5 ± 5.3 a | |
| Heat | WT | 21.8 ± 0.83 b | 5.6 ± 0.28 b | 19.4 ± 0.56 b | 56.5 ± 4.9 b |
| OE-3 | 27.9 ± 0.88 a | 8.3 ± 0.32 a | 23.5 ± 0.52 a | 81.1 ± 5.1 a | |
| OE-4 | 27.3 ± 0.69 a | 7.7 ± 0.42 a | 23.7 ± 0.51 a | 83.6 ± 6.3 a | |
| OE-6 | 26.5 ± 0.73 a | 8.5 ± 0.39 a | 24.5 ± 0.62 a | 88.2 ± 5.2 a |
| Treatment | Line | Panicle Length (cm) | Number of Spikelets per Panicle | Total no. of Spikelets per Hill | Number of Filled Grains Per Hill | Filling Rate (%) | Total Grain Weight (g) |
|---|---|---|---|---|---|---|---|
| Normal | WT | 17.8 ± 1.11 a | 89.8 ± 8.52 a | 1188.4 ± 132.3 a | 1042.6 ± 141.3 a | 87.3 ± 12.11 a | 18.2 ± 1.32 a |
| OE-3 | 18.2 ± 1.07 a | 91.3 ± 7.61 a | 1166.8 ± 143.4 a | 1044.9 ± 113.5 a | 89.2 ± 10.14 a | 17.1 ± 1.41 a | |
| OE-4 | 18.1 ± 1.18 a | 88.6 ± 9.44 a | 1180.6 ± 128.7 a | 1068.7 ± 131.7 a | 90.2 ± 12.73 a | 18.7 ± 1.36 a | |
| OE-6 | 17.8 ± 1.22 a | 92.4 ± 8.66 a | 1201.9 ± 144.5 a | 1076.9 ± 122.8 a | 89.7 ± 10.52 a | 13.9 ± 1.45 a | |
| Drought | WT | 13.1 ± 1.34 b | 79.7 ± 7.94 b | 1020.2 ± 125.7 b | 698.30 ± 133.5 b | 68.1 ± 13.46 b | 17.8 ± 1.41 b |
| OE-3 | 16.9 ± 1.52 a | 87.2 ± 8.14 a | 1178.8 ± 112.6 a | 993.7 ± 124.5 a | 84.5 ± 10.32 a | 17.3 ± 1.38 a | |
| OE-4 | 17.1 ± 1.44 a | 88.9 ± 7.91 a | 1163.9 ± 118.9 a | 982.4 ± 121.5 a | 84.7 ± 11.43 a | 18.1 ± 1.32 a | |
| OE-6 | 17.7 ± 1.56 a | 90.3 ± 8.11 a | 1188.3 ± 155.3 a | 1028.7 ± 118.6 a | 86.6 ± 13.37 a | 17.4 ± 1.31 a | |
| Heat | WT | 14.2 ± 1.39 b | 81.4 ± 7.92 b | 1076.9 ± 162.3 b | 734.51 ± 109.2 b | 68.9 ± 15.62 b | 14.2 ± 1.28 b |
| OE-3 | 17.8 ± 1.68 a | 90.6 ± 8.55 a | 1192.7 ± 133.5 a | 982.6 ± 111.5 a | 82.8 ± 16.74 a | 17.9 ± 1.31 a | |
| OE-4 | 17.5 ± 1.12 a | 88.5 ± 9.12 a | 1203.9 ± 126.7 a | 964.4 ± 108.4 a | 80.6 ± 14.77 a | 18.3 ± 1.28 a | |
| OE-6 | 18.3 ± 1.88 a | 93.1 ± 8.32 a | 1201.2 ± 152.5 a | 1048.9 ± 126.8 a | 87.4 ± 13.56 a | 18.1 ± 1.31 a |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
El-Esawi, M.A.; Alayafi, A.A. Overexpression of Rice Rab7 Gene Improves Drought and Heat Tolerance and Increases Grain Yield in Rice (Oryza sativa L.). Genes 2019, 10, 56. https://doi.org/10.3390/genes10010056
El-Esawi MA, Alayafi AA. Overexpression of Rice Rab7 Gene Improves Drought and Heat Tolerance and Increases Grain Yield in Rice (Oryza sativa L.). Genes. 2019; 10(1):56. https://doi.org/10.3390/genes10010056
Chicago/Turabian StyleEl-Esawi, Mohamed A., and Aisha A. Alayafi. 2019. "Overexpression of Rice Rab7 Gene Improves Drought and Heat Tolerance and Increases Grain Yield in Rice (Oryza sativa L.)" Genes 10, no. 1: 56. https://doi.org/10.3390/genes10010056
APA StyleEl-Esawi, M. A., & Alayafi, A. A. (2019). Overexpression of Rice Rab7 Gene Improves Drought and Heat Tolerance and Increases Grain Yield in Rice (Oryza sativa L.). Genes, 10(1), 56. https://doi.org/10.3390/genes10010056






