Cardiomyocyte Transplantation after Myocardial Infarction Alters the Immune Response in the Heart
Abstract
1. Introduction
2. Materials and Methods
2.1. Mice
2.2. Neonatal Cardiomyocyte Isolation
2.3. Surgical Procedure
2.4. Isolation of Mononuclear Cells for Flow Cytometry
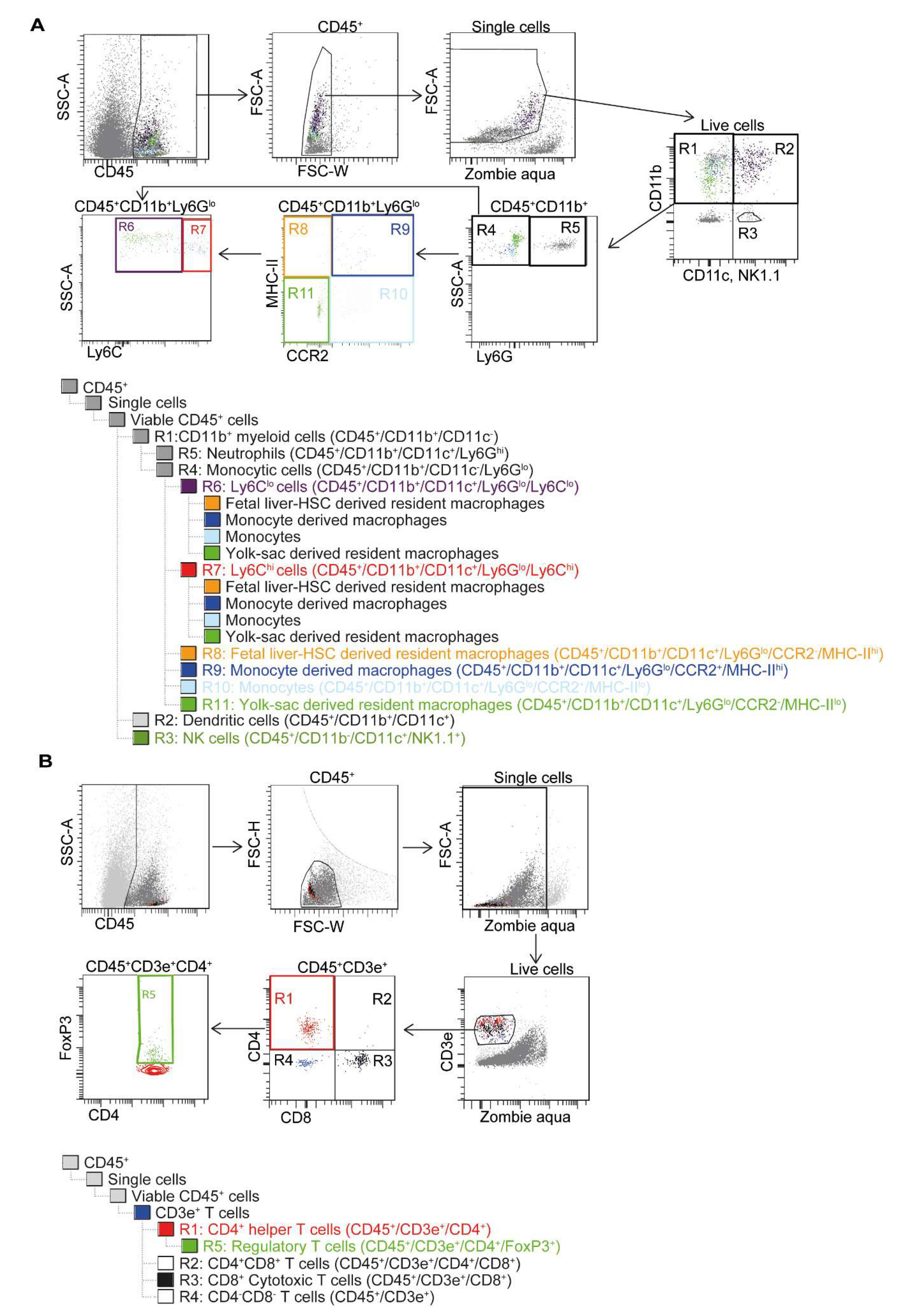
2.5. Flow Cytometry with Gating Strategy
2.6. Magnetic Resonance Imaging (MRI)
2.7. Conductance Catheter
2.8. Histological Analysis
2.9. Immunofluorescence Staining
2.10. Microarray
2.11. Microarray Analysis
2.12. Statistics
3. Results
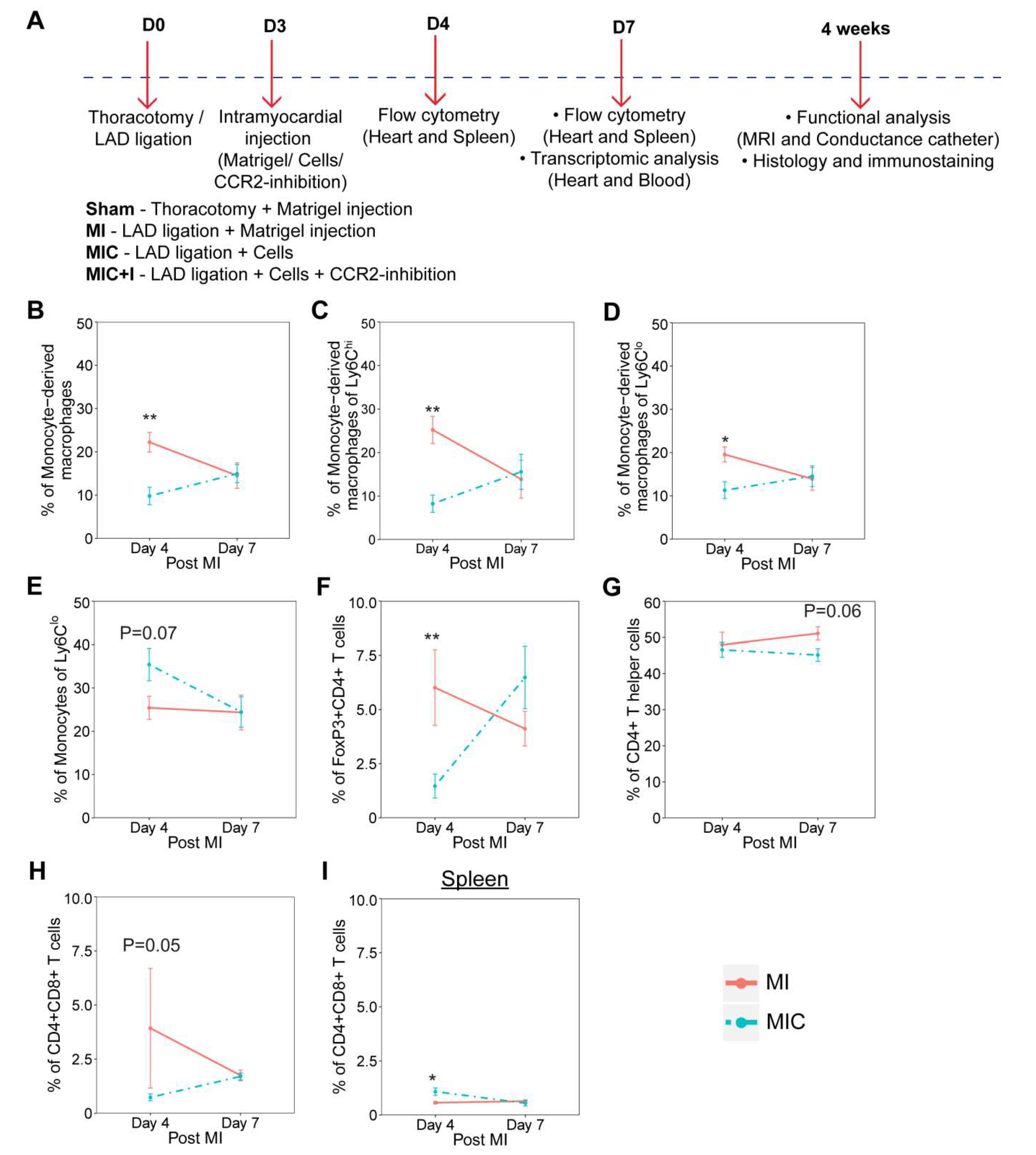
3.1. Cardiomyocyte Transplantation Alters the Dynamics of the Immune Response in the Heart after MI in C57BL/6J Mice
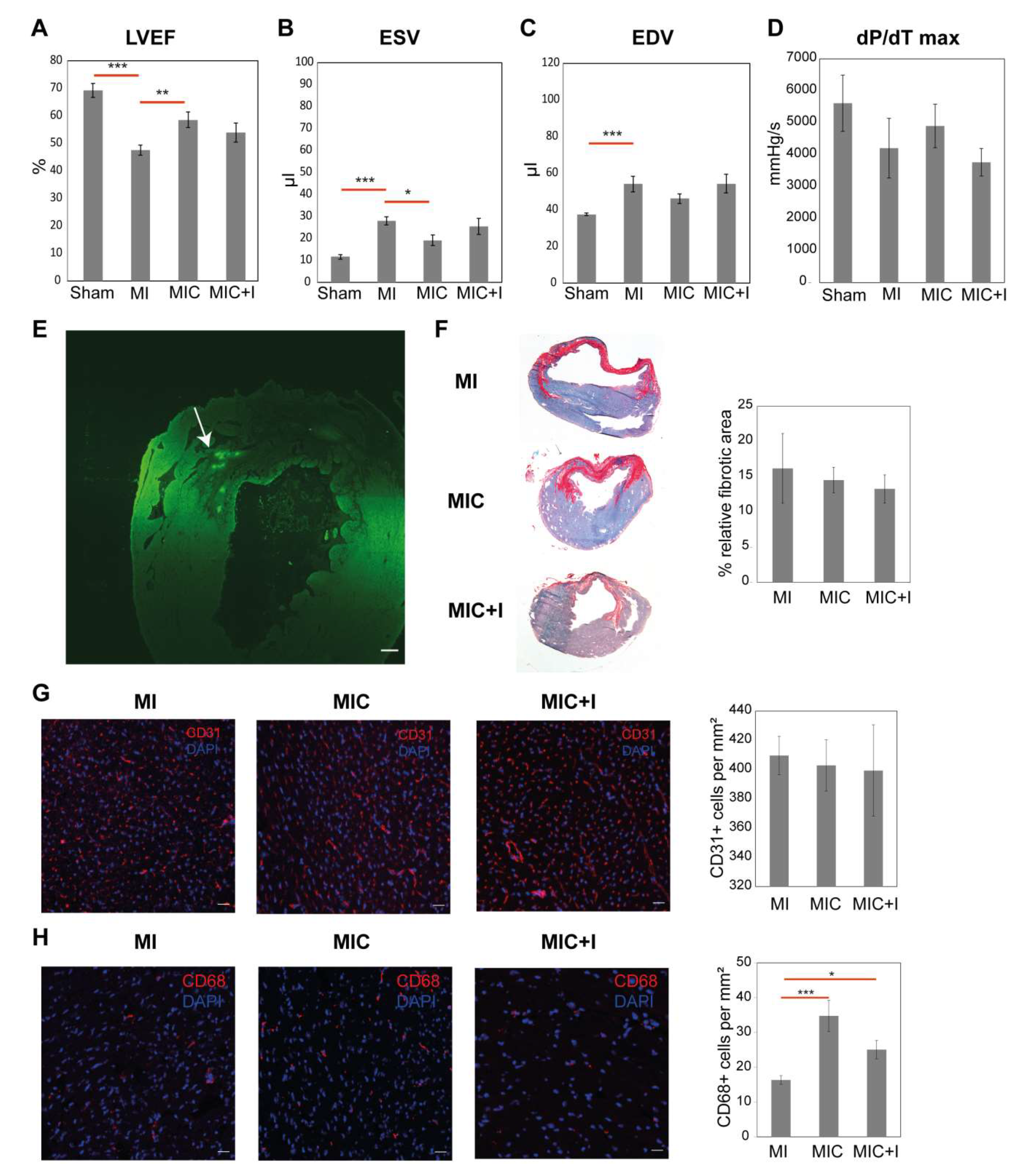
3.2. Intramyocardial Syngeneic Cardiomyocyte Transplantation Improves Cardiac Pump Function
3.3. CCR2 Inhibition Diminishes the Positive Effect of Cardiomyocyte Transplantation Alone
3.4. CD68+ Macrophage Numbers in the Remote Area of the Heart Varies between the Groups
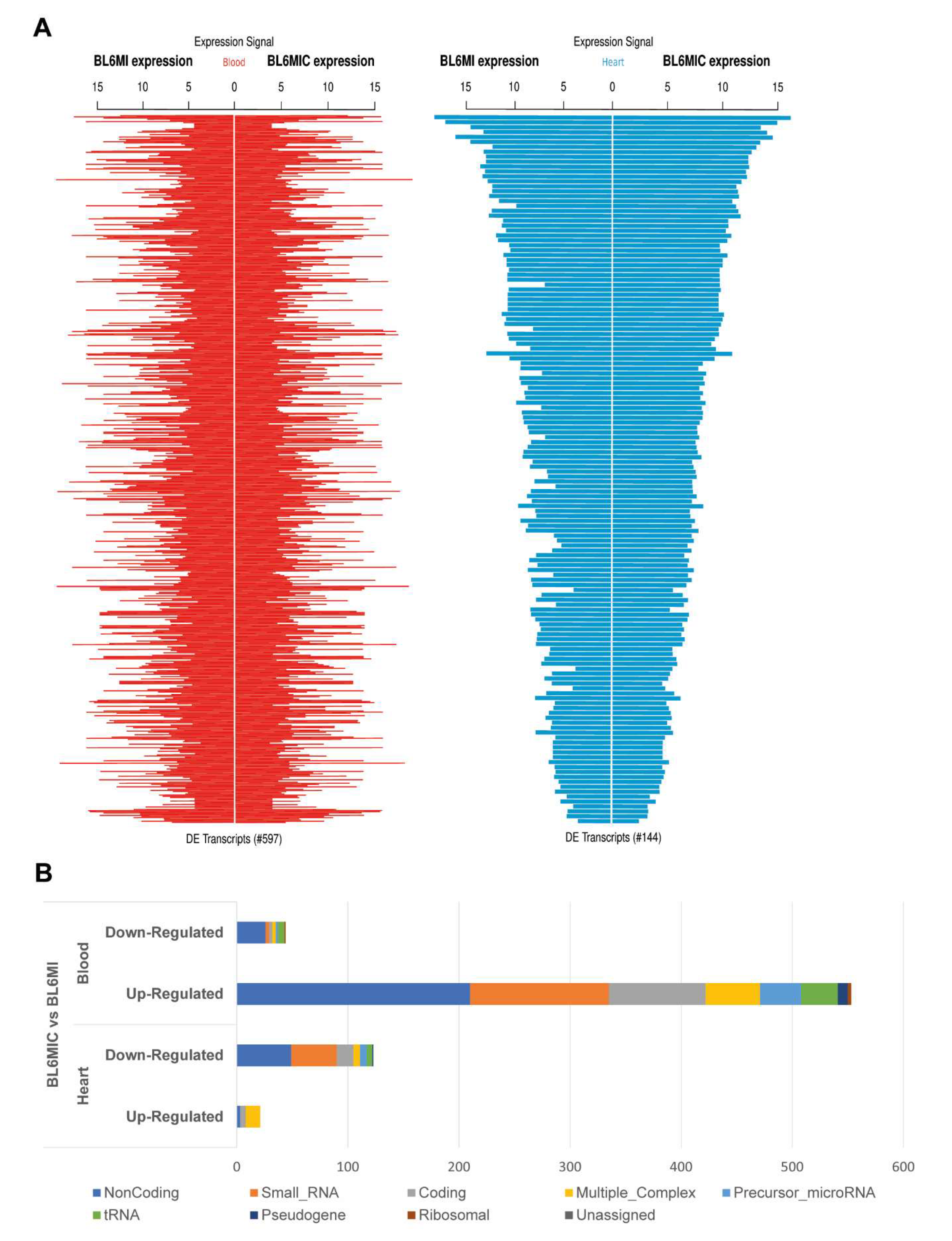
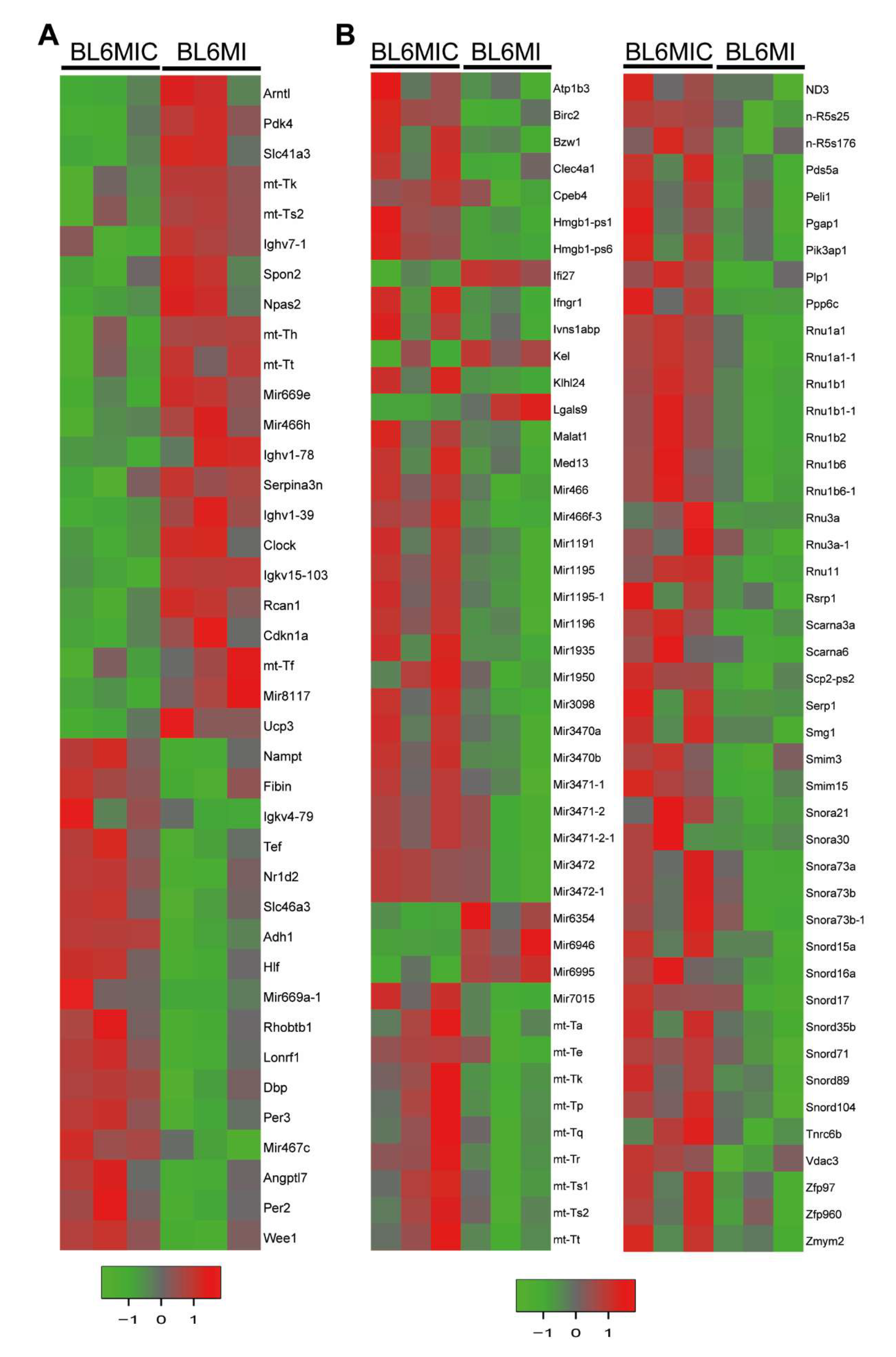
3.5. Transcriptomic Profiling of the Heart and Blood Revealed Striking Differences in Gene Expression Due to Cardiomyocyte Transplantation
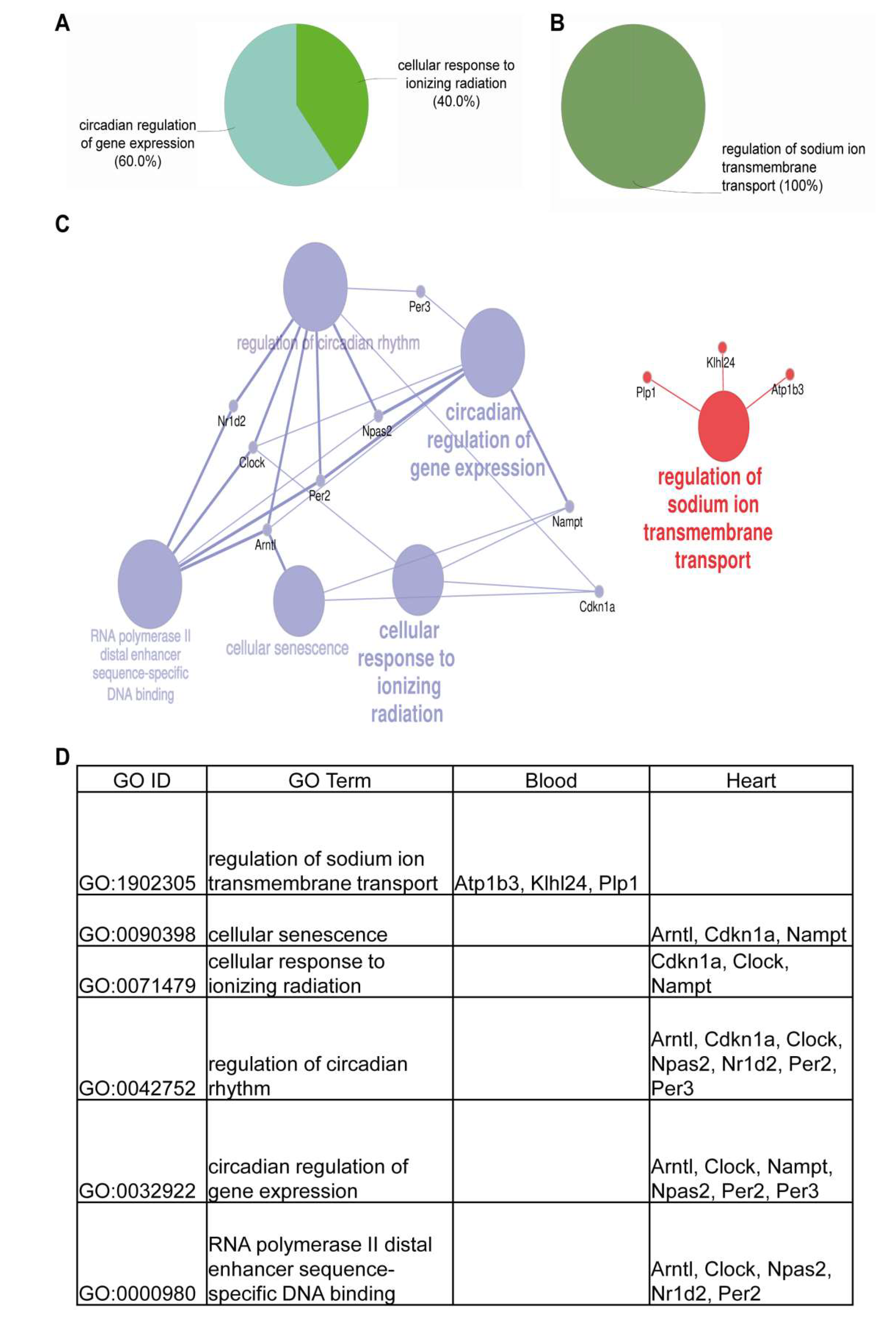
3.6. Cardiomyocyte Transplantation in C57BL/6J Mice Primarily Influences the Circadian Regulation and Mitochondrial Metabolism
4. Discussion
Author Contributions
Funding
Conflicts of Interest
References
- Benjamin, E.J.; Blaha, M.J.; Chiuve, S.E.; Cushman, M.; Das, S.R.; Deo, R.; de Ferranti, S.D.; Floyd, J.; Fornage, M.; Gillespie, C.; et al. Heart Disease and Stroke Statistics-2017 Update: A Report From the American Heart Association. Circulation 2017, 135, e146–e603. [Google Scholar] [CrossRef] [PubMed]
- Steinhoff, G.; Nesteruk, J.; Wolfien, M.; Große, J.; Ruch, U.; Vasudevan, P.; Müller, P. Stem cells and heart disease—Brake or accelerator? Adv. Drug Deliv. Rev. 2017, 120, 2–24. [Google Scholar] [CrossRef] [PubMed]
- A futile cycle in cell therapy. Nat. Biotechnol. 2017, 35, 291. [CrossRef] [PubMed]
- Chong, J.J.H.; Yang, X.; Don, C.W.; Minami, E.; Liu, Y.-W.; Weyers, J.J.; Mahoney, W.M.; Van Biber, B.; Cook, S.M.; Palpant, N.J.; et al. Human embryonic-stem-cell-derived cardiomyocytes regenerate non-human primate hearts. Nature 2014, 510, 273–277. [Google Scholar] [CrossRef] [PubMed]
- Aurora, A.B.; Olson, E.N. Immune Modulation of Stem Cells and Regeneration. Cell Stem Cell 2014, 15, 14–25. [Google Scholar] [CrossRef] [PubMed]
- Lavine, K.J.; Epelman, S.; Uchida, K.; Weber, K.J.; Nichols, C.G.; Schilling, J.D.; Ornitz, D.M.; Randolph, G.J.; Mann, D.L. Distinct macrophage lineages contribute to disparate patterns of cardiac recovery and remodeling in the neonatal and adult heart. Proc. Natl. Acad. Sci. USA 2014, 111, 16029–16034. [Google Scholar] [CrossRef]
- Ben-Mordechai, T.; Holbova, R.; Landa-Rouben, N.; Harel-Adar, T.; Feinberg, M.S.; Elrahman, I.A.; Blum, G.; Epstein, F.H.; Silman, Z.; Cohen, S.; et al. Macrophage subpopulations are essential for infarct repair with and without stem cell therapy. J. Am. Coll. Cardiol. 2013, 62, 1890–1901. [Google Scholar] [CrossRef]
- Matysiak, M.; Orlowski, W.; Fortak-Michalska, M.; Jurewicz, A.; Selmaj, K. Immunoregulatory function of bone marrow mesenchymal stem cells in EAE depends on their differentiation state and secretion of PGE2. J. Neuroimmunol. 2011, 233, 106–111. [Google Scholar] [CrossRef]
- Corcione, A.; Benvenuto, F.; Ferretti, E.; Giunti, D.; Cappiello, V.; Cazzanti, F.; Risso, M.; Gualandi, F.; Mancardi, G.L.; Pistoia, V.; et al. Human mesenchymal stem cells modulate B-cell functions. Blood 2006, 107, 367–372. [Google Scholar] [CrossRef]
- Sotiropoulou, P.A.; Perez, S.A.; Gritzapis, A.D.; Baxevanis, C.N.; Papamichail, M. Interactions between human mesenchymal stem cells and natural killer cells. Stem Cells 2006, 24, 74–85. [Google Scholar] [CrossRef]
- Karakikes, I.; Ameen, M.; Termglinchan, V.; Wu, J.C. Human Induced Pluripotent Stem Cell-Derived Cardiomyocytes: Insights Into Molecular, Cellular, and Functional Phenotypes. Circ. Res. 2015, 117, 80–88. [Google Scholar] [CrossRef] [PubMed]
- Kishino, Y.; Fujita, J.; Tohyama, S.; Okada, M.; Tanosaki, S.; Someya, S.; Fukuda, K. Toward the realization of cardiac regenerative medicine using pluripotent stem cells. Inflamm. Regen. 2020, 40, 1. [Google Scholar] [CrossRef] [PubMed]
- Vasudevan, P.; Gaebel, R.; Döring, P.; Müller, P.; Lemcke, H.; Stenzel, J.; Lindner, T.; Kurth, J.; Steinhoff, G.; Vollmar, B.; et al. 18F-FDG PET-Based Imaging of Myocardial Inflammation Predicts a Functional Outcome Following Transplantation of mESC-Derived Cardiac Induced Cells in a Mouse Model of Myocardial Infarction. Cells 2019, 8, 1613. [Google Scholar] [CrossRef] [PubMed]
- Heiberg, E.; Sjögren, J.; Ugander, M.; Carlsson, M.; Engblom, H.; Arheden, H. Design and validation of Segment—Freely available software for cardiovascular image analysis. BMC Med. Imaging 2010, 10, 1. [Google Scholar] [CrossRef] [PubMed]
- Koczan, D.; Fitzner, B.; Zettl, U.K.; Hecker, M. Microarray data of transcriptome shifts in blood cell subsets during S1P receptor modulator therapy. Sci. Data 2018, 5, 180145. [Google Scholar] [CrossRef]
- Bindea, G.; Mlecnik, B.; Hackl, H.; Charoentong, P.; Tosolini, M.; Kirilovsky, A.; Fridman, W.-H.; Pagès, F.; Trajanoski, Z.; Galon, J. ClueGO: A Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 2009, 25, 1091–1093. [Google Scholar] [CrossRef]
- Babicki, S.; Arndt, D.; Marcu, A.; Liang, Y.; Grant, J.R.; Maciejewski, A.; Wishart, D.S. Heatmapper: Web-enabled heat mapping for all. Nucleic Acids Res. 2016, 44, W147–W153. [Google Scholar] [CrossRef]
- Parri, M.; Pietrovito, L.; Grandi, A.; Campagnoli, S.; De Camilli, E.; Bianchini, F.; Marchiò, S.; Bussolino, F.; Jin, B.; Sarmientos, P.; et al. Angiopoietin-like 7, a novel pro-angiogenetic factor over-expressed in cancer. Angiogenesis 2014, 17, 881–896. [Google Scholar] [CrossRef]
- Dick, S.A.; Macklin, J.A.; Nejat, S.; Momen, A.; Clemente-Casares, X.; Althagafi, M.G.; Chen, J.; Kantores, C.; Hosseinzadeh, S.; Aronoff, L.; et al. Self-renewing resident cardiac macrophages limit adverse remodeling following myocardial infarction. Nat. Immunol. 2019, 20, 29–39. [Google Scholar] [CrossRef] [PubMed]
- Epelman, S.; Lavine, K.J.; Beaudin, A.E.; Sojka, D.K.; Carrero, J.A.; Calderon, B.; Brija, T.; Gautier, E.L.; Ivanov, S.; Satpathy, A.T.; et al. Embryonic and adult-derived resident cardiac macrophages are maintained through distinct mechanisms at steady state and during inflammation. Immunity 2014, 40, 91–104. [Google Scholar] [CrossRef] [PubMed]
- Farbehi, N.; Patrick, R.; Dorison, A.; Xaymardan, M.; Janbandhu, V.; Wystub-Lis, K.; Ho, J.W.K.; Nordon, R.E.; Harvey, R.P. Single-cell expression profiling reveals dynamic flux of cardiac stromal, vascular and immune cells in health and injury. Elife 2019, 8, 1–39. [Google Scholar] [CrossRef] [PubMed]
- Kolossov, E.; Bostani, T.; Roell, W.; Breitbach, M.; Pillekamp, F.; Nygren, J.M.; Sasse, P.; Rubenchik, O.; Fries, J.W.U.; Wenzel, D.; et al. Engraftment of engineered ES cell-derived cardiomyocytes but not BM cells restores contractile function to the infarcted myocardium. J. Exp. Med. 2006, 203, 2315–2327. [Google Scholar] [CrossRef] [PubMed]
- Laflamme, M.A.; Chen, K.Y.; Naumova, A.V.; Muskheli, V.; Fugate, J.A.; Dupras, S.K.; Reinecke, H.; Xu, C.; Hassanipour, M.; Police, S.; et al. Cardiomyocytes derived from human embryonic stem cells in pro-survival factors enhance function of infarcted rat hearts. Nat. Biotechnol. 2007, 25, 1015–1024. [Google Scholar] [CrossRef] [PubMed]
- Kehat, I.; Khimovich, L.; Caspi, O.; Gepstein, A.; Shofti, R.; Arbel, G.; Huber, I.; Satin, J.; Itskovitz-Eldor, J.; Gepstein, L. Electromechanical integration of cardiomyocytes derived from human embryonic stem cells. Nat. Biotechnol. 2004, 22, 1282–1289. [Google Scholar] [CrossRef] [PubMed]
- Halbach, M.; Pfannkuche, K.; Pillekamp, F.; Ziomka, A.; Hannes, T.; Reppel, M.; Hescheler, J.; Müller-Ehmsen, J. Electrophysiological maturation and integration of murine fetal cardiomyocytes after transplantation. Circ. Res. 2007, 101, 484–492. [Google Scholar] [CrossRef] [PubMed]
- Funakoshi, S.; Miki, K.; Takaki, T.; Okubo, C.; Hatani, T.; Chonabayashi, K.; Nishikawa, M.; Takei, I.; Oishi, A.; Narita, M.; et al. Enhanced engraftment, proliferation, and therapeutic potential in heart using optimized human iPSC-derived cardiomyocytes. Sci. Rep. 2016, 6, 19111. [Google Scholar] [CrossRef]
- Masumoto, H.; Matsuo, T.; Yamamizu, K.; Uosaki, H.; Narazaki, G.; Katayama, S.; Marui, A.; Shimizu, T.; Ikeda, T.; Okano, T.; et al. Pluripotent stem cell-engineered cell sheets reassembled with defined cardiovascular populations ameliorate reduction in infarct heart function through cardiomyocyte-mediated neovascularization. Stem Cells 2012, 30, 1196–1205. [Google Scholar] [CrossRef]
- Caspi, O.; Huber, I.; Kehat, I.; Habib, M.; Arbel, G.; Gepstein, A.; Yankelson, L.; Aronson, D.; Beyar, R.; Gepstein, L. Transplantation of human embryonic stem cell-derived cardiomyocytes improves myocardial performance in infarcted rat hearts. J. Am. Coll. Cardiol. 2007, 50, 1884–1893. [Google Scholar] [CrossRef]
- Halbach, M.; Krausgrill, B.; Hannes, T.; Wiedey, M.; Peinkofer, G.; Baumgartner, S.; Sahito, R.G.A.; Pfannkuche, K.; Pillekamp, F.; Reppel, M.; et al. Time-course of the electrophysiological maturation and integration of transplanted cardiomyocytes. J. Mol. Cell. Cardiol. 2012, 53, 401–408. [Google Scholar] [CrossRef]
- Muller-Ehmsen, J. Rebuilding a Damaged Heart: Long-Term Survival of Transplanted Neonatal Rat Cardiomyocytes After Myocardial Infarction and Effect on Cardiac Function. Circulation 2002, 105, 1720–1726. [Google Scholar] [CrossRef]
- Shiba, Y.; Gomibuchi, T.; Seto, T.; Wada, Y.; Ichimura, H.; Tanaka, Y.; Ogasawara, T.; Okada, K.; Shiba, N.; Sakamoto, K.; et al. Allogeneic transplantation of iPS cell-derived cardiomyocytes regenerates primate hearts. Nature 2016, 538, 388–391. [Google Scholar] [CrossRef] [PubMed]
- Tachibana, A.; Santoso, M.R.; Mahmoudi, M.; Shukla, P.; Wang, L.; Bennett, M.; Goldstone, A.B.; Wang, M.; Fukushi, M.; Ebert, A.D.; et al. Paracrine Effects of the Pluripotent Stem Cell-Derived Cardiac Myocytes Salvage the Injured Myocardium. Circ. Res. 2017, 121, e22–e36. [Google Scholar] [CrossRef] [PubMed]
- Zhu, K.; Wu, Q.; Ni, C.; Zhang, P.; Zhong, Z.; Wu, Y.; Wang, Y.; Xu, Y.; Kong, M.; Cheng, H.; et al. Lack of Remuscularization Following Transplantation of Human Embryonic Stem Cell-Derived Cardiovascular Progenitor Cells in Infarcted Nonhuman Primates. Circ. Res. 2018, 122, 958–969. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.-W.; Chen, B.; Yang, X.; Fugate, J.A.; Kalucki, F.A.; Futakuchi-Tsuchida, A.; Couture, L.; Vogel, K.W.; Astley, C.A.; Baldessari, A.; et al. Human embryonic stem cell–derived cardiomyocytes restore function in infarcted hearts of non-human primates. Nat. Biotechnol. 2018, 36, 597–605. [Google Scholar] [CrossRef]
- Young, M.E.; Razeghi, P.; Cedars, A.M.; Guthrie, P.H.; Taegtmeyer, H. Intrinsic diurnal variations in cardiac metabolism and contractile function. Circ. Res. 2001, 89, 1199–1208. [Google Scholar] [CrossRef]
- Durgan, D.J.; Trexler, N.A.; Egbejimi, O.; McElfresh, T.A.; Suk, H.Y.; Petterson, L.E.; Shaw, C.A.; Hardin, P.E.; Bray, M.S.; Chandler, M.P.; et al. The circadian clock within the cardiomyocyte is essential for responsiveness of the heart to fatty acids. J. Biol. Chem. 2006, 281, 24254–24269. [Google Scholar] [CrossRef]
- Bass, J.; Takahashi, J.S. Circadian integration of metabolism and energetics. Science 2010, 330, 1349–1354. [Google Scholar] [CrossRef]
- Haus, E.; Smolensky, M.H. Biologic rhythms in the immune system. Chronobiol. Int. 1999, 16, 581–622. [Google Scholar] [CrossRef]
- Nguyen, K.D.; Fentress, S.J.; Qiu, Y.; Yun, K.; Cox, J.S.; Chawla, A. Circadian gene Bmal1 regulates diurnal oscillations of Ly6C(hi) inflammatory monocytes. Science 2013, 341, 1483–1488. [Google Scholar] [CrossRef]
- du Pré, B.C.; Dierickx, P.; Crnko, S.; Doevendans, P.A.; Vos, M.A.; Geijsen, N.; Neutel, D.; van Veen, T.A.B.; van Laake, L.W. Neonatal rat cardiomyocytes as an in vitro model for circadian rhythms in the heart. J. Mol. Cell. Cardiol. 2017, 112, 58–63. [Google Scholar] [CrossRef]
- Dierickx, P.; Vermunt, M.W.; Muraro, M.J.; Creyghton, M.P.; Doevendans, P.A.; van Oudenaarden, A.; Geijsen, N.; Van Laake, L.W. Circadian networks in human embryonic stem cell-derived cardiomyocytes. EMBO Rep. 2017, 18, 1199–1212. [Google Scholar] [CrossRef] [PubMed]
- Vagnozzi, R.J.; Maillet, M.; Sargent, M.A.; Khalil, H.; Johansen, A.K.Z.; Schwanekamp, J.A.; York, A.J.; Huang, V.; Nahrendorf, M.; Sadayappan, S.; et al. An acute immune response underlies the benefit of cardiac stem cell therapy. Nature 2020, 577, 405–409. [Google Scholar] [CrossRef] [PubMed]
| Target | Clone | Source |
|---|---|---|
| CD45 | 30-F11 | Biolegend |
| CD11b | M1/70 | Biolegend |
| CD11c | N418 | Biolegend |
| NK1.1 | PK136 | Biolegend |
| Ly6G | 1A8 | Biolegend |
| Ly6C | Hk1.4 | Biolegend |
| CCR2 | 475301 | R and D |
| MHC II | AF6-120.1 | Biolegend |
| CD3e | 145-2C11 | Biolegend |
| CD8a | 53-6.7 | Biolegend |
| CD4 | RM4-5 | Biolegend |
| FoxP3 | MF-14 | Biolegend |
| Anti-CD31 | MEC 7.46 | Abcam |
| Anti-CD68 | FA-11 | Invitrogen |
| Anti-GFP | Rabbit polyclonal | Abcam |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vasudevan, P.; Wolfien, M.; Lemcke, H.; Lang, C.I.; Skorska, A.; Gaebel, R.; Koczan, D.; Lindner, T.; Engelmann, R.; Vollmar, B.; et al. Cardiomyocyte Transplantation after Myocardial Infarction Alters the Immune Response in the Heart. Cells 2020, 9, 1825. https://doi.org/10.3390/cells9081825
Vasudevan P, Wolfien M, Lemcke H, Lang CI, Skorska A, Gaebel R, Koczan D, Lindner T, Engelmann R, Vollmar B, et al. Cardiomyocyte Transplantation after Myocardial Infarction Alters the Immune Response in the Heart. Cells. 2020; 9(8):1825. https://doi.org/10.3390/cells9081825
Chicago/Turabian StyleVasudevan, Praveen, Markus Wolfien, Heiko Lemcke, Cajetan Immanuel Lang, Anna Skorska, Ralf Gaebel, Dirk Koczan, Tobias Lindner, Robby Engelmann, Brigitte Vollmar, and et al. 2020. "Cardiomyocyte Transplantation after Myocardial Infarction Alters the Immune Response in the Heart" Cells 9, no. 8: 1825. https://doi.org/10.3390/cells9081825
APA StyleVasudevan, P., Wolfien, M., Lemcke, H., Lang, C. I., Skorska, A., Gaebel, R., Koczan, D., Lindner, T., Engelmann, R., Vollmar, B., Krause, B. J., Wolkenhauer, O., Lang, H., Steinhoff, G., & David, R. (2020). Cardiomyocyte Transplantation after Myocardial Infarction Alters the Immune Response in the Heart. Cells, 9(8), 1825. https://doi.org/10.3390/cells9081825







