Identification of Novel Molecular Markers of Human Th17 Cells
Abstract
1. Introduction
2. Materials and Methods
2.1. Naive CD4+ TCell Isolation and Differentiation
2.2. RNA Sequencing (RNA-seq) and Analysis of Differentially Expressed Genes (DEGs)
2.3. Gene Ontology
2.4. Quantitative PCR
2.5. Western Blotting
2.6. Analysis of Interleukin Production
2.7. Chromatin Immunoprecipitation Assay
2.8. Statistics
3. Results
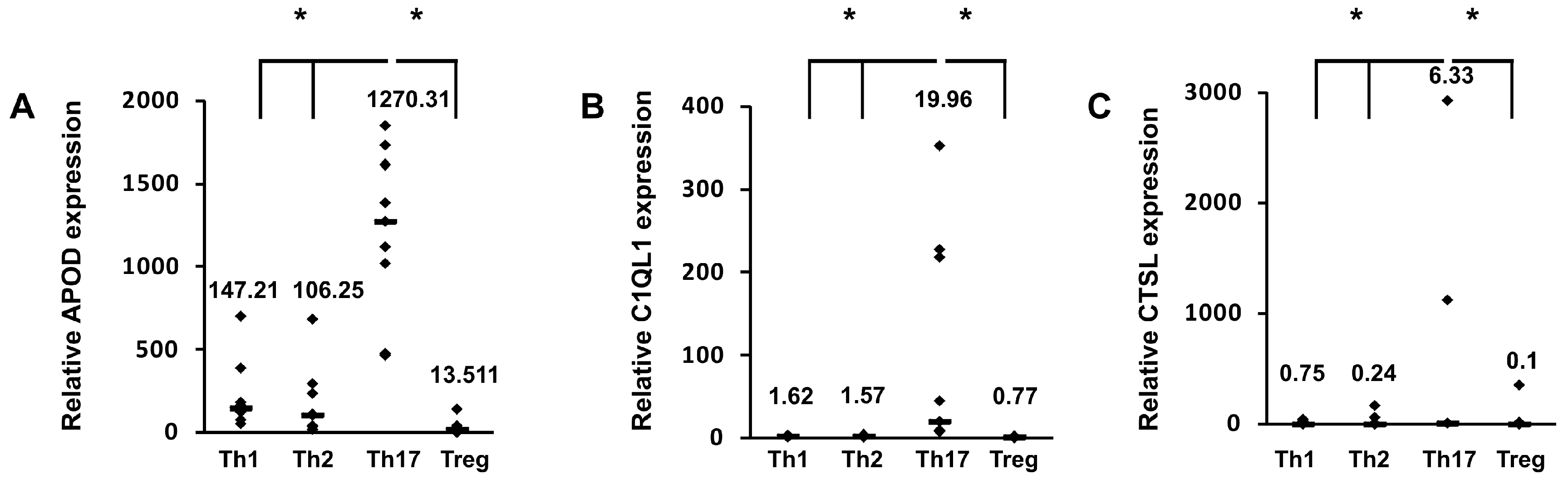
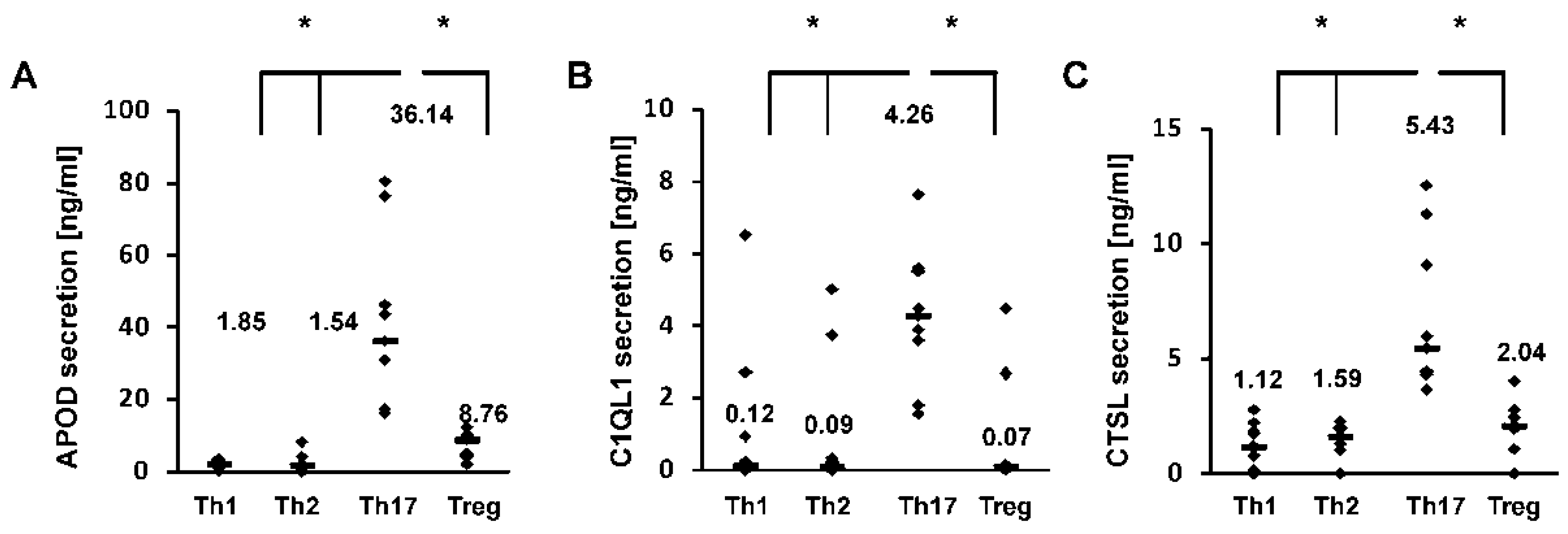
3.1. Identification of Novel Th17-Specific Genes
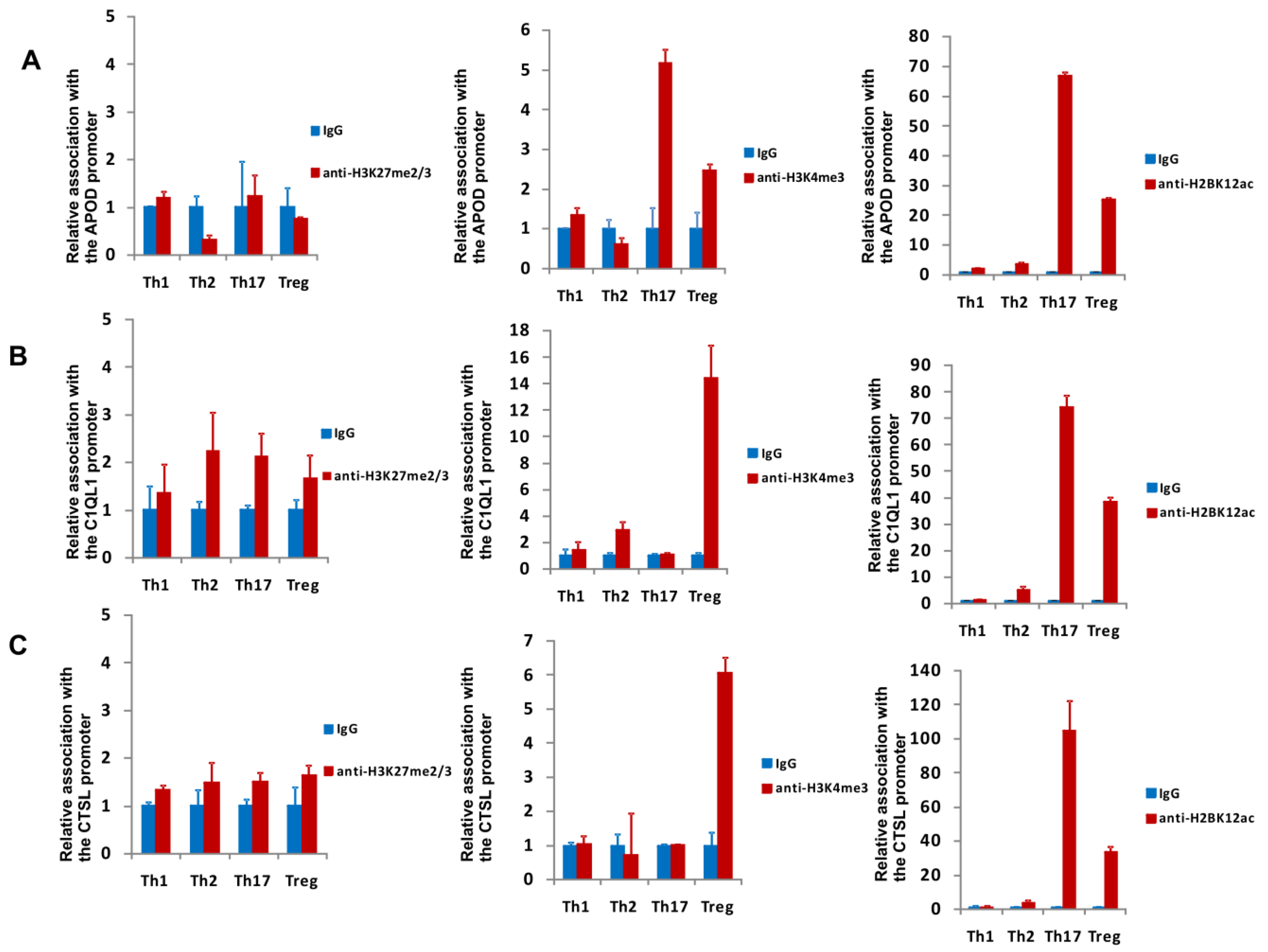
3.2. Analysis of Epigenetic Marks in the Loci of the Identified Genes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- E Harrington, L.; Hatton, R.D.; Mangan, P.R.; Turner, H.; Murphy, T.L.; Murphy, K.M.; Weaver, C.T. Interleukin 17–producing CD4+ effector T cells develop via a lineage distinct from the T helper type 1 and 2 lineages. Nat. Immunol. 2005, 6, 1123–1132. [Google Scholar] [CrossRef] [PubMed]
- Park, H.; Li, Z.; O Yang, X.; Chang, S.H.; Nurieva, R.; Wang, Y.-H.; Wang, Y.; Hood, L.; Zhu, Z.; Tian, Q.; et al. A distinct lineage of CD4 T cells regulates tissue inflammation by producing interleukin 17. Nat. Immunol. 2005, 6, 1133–1141. [Google Scholar] [CrossRef] [PubMed]
- Wei, L.; Laurence, A.; Elias, K.; O’Shea, J.J. IL-21 is produced by Th17 cells and drives IL-17 production in a STAT3-dependent manner. J. Biol. Chem. 2007, 282, 34605–34610. [Google Scholar] [CrossRef] [PubMed]
- Liang, S.C.; Tan, X.-Y.; Luxenberg, D.P.; Karim, R.; Dunussi-Joannopoulos, K.; Collins, M.; Fouser, L.A. Interleukin (IL)-22 and IL-17 are coexpressed by Th17 cells and cooperatively enhance expression of antimicrobial peptides. J. Exp. Med. 2006, 203, 2271–2279. [Google Scholar] [CrossRef] [PubMed]
- Medvedev, A.; Chistokhina, A.; Hirose, T.; Jetten, A.M. Genomic Structure and Chromosomal Mapping of the Nuclear Orphan Receptor RORγ (RORC) Gene. Genomics. 1997, 46, 93–102. [Google Scholar] [CrossRef] [PubMed]
- Crome, S.; Wang, A.Y.; Kang, C.Y.; Levings, M.K. The role of retinoic acid-related orphan receptor variant 2 and IL‐17 in the development and function of human CD4+T cells. Eur. J. Immunol. 2009, 39, 1480–1493. [Google Scholar] [CrossRef]
- Cosmi, L.; De Palma, R.; Santarlasci, V.; Maggi, L.; Capone, M.; Frosali, F.; Rodolico, G.; Querci, V.; Abbate, G.; Angeli, R.; et al. Human interleukin 17–producing cells originate from a CD161+CD4+ T cell precursor. J. Exp. Med. 2008, 205, 1903–1916. [Google Scholar] [CrossRef] [PubMed]
- Acosta-Rodriguez, E.; Rivino, L.; Geginat, J.; Jarrossay, D.; Gattorno, M.; Lanzavecchia, A.; Sallusto, F.; Napolitani, G. Surface phenotype and antigenic specificity of human interleukin 17–producing T helper memory cells. Nat. Immunol. 2007, 8, 639–646. [Google Scholar] [CrossRef]
- Harris, K.M.; Ramachandran, G.; Basu, S.; Rollins, S.; Mann, D.; Cross, A.S. The il-23/th17 axis is involved in the adaptive immune response to bacillus anthracis in humans. Eur J. Immunol. 2014, 44, 752–762. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.; Ibrahim, A.S.; Xu, X.; Farber, J.M.; Avanesian, V.; Baquir, B.; Fu, Y.; French, S.W.; Edwards, J.E.; Spellberg, B. Th1-Th17 Cells Mediate Protective Adaptive Immunity against Staphylococcus aureus and Candida albicans Infection in Mice. PLOS Pathog. 2009, 5, e1000703. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.; Na, L.; Fidel, P.L.; Schwarzenberger, P. Requirement of Interleukin?17A for Systemic Anti– Candida albicans Host Defense in Mice. J. Infect. Dis. 2004, 190, 624–631. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.O.; Panopoulos, A.D.; Nurieva, R.; Chang, S.H.; Wang, D.; Watowich, S.S.; Dong, C. STAT3 Regulates Cytokine-mediated Generation of Inflammatory Helper T Cells. J. Biol. Chem. 2007, 282, 9358–9363. [Google Scholar] [CrossRef] [PubMed]
- Buckley, R.H. Primary immunodeficiency diseases: Dissectors of the immune system. Immunol. Rev. 2002, 185, 206–219. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.S.; Chew, G.Y.; Simpson, N.; Priyadarshi, A.; Wong, M.; Grimbacher, B.; Fulcher, D.A.; Tangye, S.G.; Cook, M.C. Deficiency of Th17 cells in hyper IgE syndrome due to mutations in STAT3. J. Exp. Med. 2008, 205, 1551–1557. [Google Scholar] [CrossRef] [PubMed]
- Milner, J.D.; Sandler, N.G.; Douek, D.C. Th17 cells, Job’s syndrome and HIV: Opportunities for bacterial and fungal infections. Curr. Opin. HIV AIDS 2010, 5, 179–183. [Google Scholar] [CrossRef]
- Eyerich, K.; Foerster, S.; Rombold, S.; Seidl, H.-P.; Behrendt, H.; Hofmann, H.; Ring, J.; Traidl-Hoffmann, C. Patients with Chronic Mucocutaneous Candidiasis Exhibit Reduced Production of Th17-Associated Cytokines IL-17 and IL-22. J. Invest. Dermatol. 2008, 128, 2640–2645. [Google Scholar] [CrossRef] [PubMed]
- Hirota, K.; Hashimoto, M.; Yoshitomi, H.; Tanaka, S.; Nomura, T.; Yamaguchi, T.; Iwakura, Y.; Sakaguchi, N.; Sakaguchi, S. T cell self-reactivity forms a cytokine milieu for spontaneous development of IL-17+ Th cells that cause autoimmune arthritis. J. Exp. Med. 2007, 204, 41–47. [Google Scholar] [CrossRef] [PubMed]
- Kébir, H.; Kreymborg, K.; Ifergan, I.; Dodelet-Devillers, A.; Cayrol, R.; Bernard, M.; Giuliani, F.; Arbour, N.; Becher, B.; Prat, A. Human TH17 lymphocytes promote blood-brain barrier disruption and central nervous system inflammation. Nat. Med. 2007, 13, 1173–1175. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Chen, X.; Liu, Z.; Yue, Q.; Liu, H. Expression of Th17 cytokines in skin lesions of patients with psoriasis. J. Huazhong Univ. Sci. Technol. Med. Sci. 2007, 27, 330–332. [Google Scholar] [CrossRef]
- Sheibanie, A.F.; Yen, J.-H.; Khayrullina, T.; Emig, F.; Zhang, M.; Tuma, R.; Ganea, I. The proinflammatory effect of prostaglandin E2 in experimental inflammatory bowel disease is mediated through the IL-23-->IL-17 axis. J. Immunol. 2007, 178, 8138–8147. [Google Scholar] [CrossRef]
- Zheng, L.; Ye, P.; Liu, C. The role of the il-23/il-17 axis in the pathogenesis of graves’ disease. Endocr J. 2013, 60, 591–597. [Google Scholar] [CrossRef] [PubMed]
- Jandus, C.; Bioley, G.; Rivals, J.-P.; Dudler, J.; Speiser, D.E.; Romero, P. Increased numbers of circulating polyfunctional Th17 memory cells in patients with seronegative spondylarthritides. Arthritis Rheum. 2008, 58, 2307–2317. [Google Scholar] [CrossRef] [PubMed]
- Annunziato, F.; Cosmi, L.; Santarlasci, V.; Maggi, L.; Liotta, F.; Mazzinghi, B.; Parente, E.; Filiì, L.; Ferri, S.; Frosali, F.; et al. Phenotypic and functional features of human Th17 cells. J. Exp. Med. 2007, 204, 1849–1861. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Churchill, M.J.; Nagar, K.K.; Tailor, Y.H.; Chu, T.; Rush, B.S.; Jiang, Z.; Wang, E.B.; Renz, B.W.; Wang, H.; et al. IL-17 producing mast cells promote the expansion of myeloid-derived suppressor cells in a mouse allergy model of colorectal cancer. Oncotarget 2015, 6, 32966–32979. [Google Scholar] [CrossRef] [PubMed]
- Cosmi, L.; Cimaz, R.; Maggi, L.; Santarlasci, V.; Capone, M.; Borriello, F.; Frosali, F.; Querci, V.; Simonini, G.; Barra, G.; et al. Evidence of the transient nature of the Th17 phenotype of CD4+CD161+ T cells in the synovial fluid of patients with juvenile idiopathic arthritis. Arthritis Rheum. 2011, 63, 2504–2515. [Google Scholar] [CrossRef]
- Mazzoni, A.; Santarlasci, V.; Maggi, L.; Capone, M.; Rossi, M.C.; Querci, V.; De Palma, R.; Chang, H.-D.; Thiel, A.; Cimaz, R.; et al. Demethylation of the RORC2 and IL17A in Human CD4+ T Lymphocytes Defines Th17 Origin of Nonclassic Th1 Cells. J. Immunol. 2015, 194, 3116–3126. [Google Scholar] [CrossRef]
- Maggi, L.; Mazzoni, A.; Cimaz, R.; Liotta, F.; Annunziato, F.; Cosmi, L. Th17 and Th1 Lymphocytes in Oligoarticular Juvenile Idiopathic Arthritis. Front. Immunol. 2019, 10, 450. [Google Scholar] [CrossRef] [PubMed]
- Wilson, N.J.; Boniface, K.; Chan, J.R.; McKenzie, B.S.; Blumenschein, W.M.; Mattson, J.D.; Basham, B.; Smith, K.; Chen, T.; Morel, F.; et al. Development, cytokine profile and function of human interleukin 17–producing helper T cells. Nat. Immunol. 2007, 8, 950–957. [Google Scholar] [CrossRef]
- Patro, R.; Duggal, G.; Love, M.I.; A Irizarry, R.; Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 2017, 14, 417–419. [Google Scholar] [CrossRef]
- Anders, S.; Huber, W. Differential expression analysis for sequence count data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef]
- Mi, H.; Huang, X.; Muruganujan, A.; Tang, H.; Mills, C.; Kang, D.; Thomas, P. PANTHER version 11: Expanded annotation data from Gene Ontology and Reactome pathways, and data analysis tool enhancements. Nucleic Acids Res. 2016, 45, D183–D189. [Google Scholar] [CrossRef] [PubMed]
- Ratajewski, M.; Walczak-Drzewiecka, A.; Salkowska, A.; Dastych, J. Upstream stimulating factors regulate the expression of rorgammat in human lymphocytes. J. Immunol. 2012, 189, 3034–3042. [Google Scholar] [CrossRef] [PubMed]
- Hoene, V.; Fischer, M.; Ivanova, A.; Wallach, T.; Berthold, F.; Dame, C. GATA factors in human neuroblastoma: Distinctive expression patterns in clinical subtypes. Br. J. Cancer 2009, 101, 1481–1489. [Google Scholar] [CrossRef] [PubMed]
- Vandesompele, J.; De Preter, K.; Pattyn, F.; Poppe, B.; Van Roy, N.; De Paepe, A.; Speleman, F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002, 3, 0034. [Google Scholar] [CrossRef] [PubMed]
- Joshi, S.; Pantalena, L.-C.; Liu, X.K.; Gaffen, S.L.; Liu, H.; Rohowsky-Kochan, C.; Ichiyama, K.; Yoshimura, A.; Steinman, L.; Christakos, S.; et al. 1,25-Dihydroxyvitamin D3 Ameliorates Th17 Autoimmunity via Transcriptional Modulation of Interleukin-17A. Mol. Cell. Biol. 2011, 31, 3653–3669. [Google Scholar] [CrossRef] [PubMed]
- Schleussner, N.; Merkel, O.; Costanza, M.; Liang, H.C.; Hummel, F.; Romagnani, C.; Durek, P.; Anagnostopoulos, I.; Hummel, M.; Johrens, K.; et al. The ap-1-batf and -batf3 module is essential for growth, survival and th17/ilc3 skewing of anaplastic large cell lymphoma. Leukemia 2018, 32, 1994–2007. [Google Scholar] [CrossRef]
- Shi, L.Z.; Wang, R.; Huang, G.; Vogel, P.; Neale, G.; Green, D.R.; Chi, H. HIF1α–dependent glycolytic pathway orchestrates a metabolic checkpoint for the differentiation of TH17 and Treg cells. J. Exp. Med. 2011, 208, 1367–1376. [Google Scholar] [CrossRef]
- Tuomela, S.; Salo, V.; Tripathi, S.K.; Chen, Z.; Laurila, K.; Gupta, B.; Aijo, T.; Oikari, L.; Stockinger, B.; Lähdesmäki, H.; et al. Identification of early gene expression changes during human Th17 cell differentiation. Blood 2012, 119, e151–e160. [Google Scholar] [CrossRef] [PubMed]
- Szabo, S.J.; Kim, S.T.; Costa, G.L.; Zhang, X.; Fathman, C.G.; Glimcher, L.H. A Novel Transcription Factor, T-bet, Directs Th1 Lineage Commitment. Cell 2000, 100, 655–669. [Google Scholar] [CrossRef]
- Zheng, W.-P.; Flavell, R.A. The Transcription Factor GATA-3 Is Necessary and Sufficient for Th2 Cytokine Gene Expression in CD4 T Cells. Cell 1997, 89, 587–596. [Google Scholar] [CrossRef]
- Chen, W.; Jin, W.; Hardegen, N.; Lei, K.J.; Li, L.; Marinos, N.; McGrady, G.; Wahl, S.M. Conversion of peripheral cd4+cd25- naive t cells to cd4+cd25+ regulatory t cells by tgf-beta induction of transcription factor foxp3. J. Exp. Med. 2003, 198, 1875–1886. [Google Scholar] [CrossRef] [PubMed]
- Cherwinski, H.M.; Schumacher, J.H.; Brown, K.D.; Mosmann, T.R. Two types of mouse helper T cell clone. III. Further differences in lymphokine synthesis between Th1 and Th2 clones revealed by RNA hybridization, functionally monospecific bioassays, and monoclonal antibodies. J. Exp. Med. 1987, 166, 1229–1244. [Google Scholar] [CrossRef] [PubMed]
- Mosmann, T.R.; Coffman, R.L. Th1 and th2 cells: Different patterns of lymphokine secretion lead to different functional properties. Annu Rev. Immunol 1989, 7, 145–173. [Google Scholar] [CrossRef] [PubMed]
- E Harrington, L.; Mangan, P.R.; Weaver, C.T. Expanding the effector CD4 T-cell repertoire: The Th17 lineage. Curr. Opin. Immunol. 2006, 18, 349–356. [Google Scholar] [CrossRef] [PubMed]
- Weaver, C.T.; Hatton, R.D.; Mangan, P.R.; Harrington, L.E. IL-17 Family Cytokines and the Expanding Diversity of Effector T Cell Lineages. Annu. Rev. Immunol. 2007, 25, 821–852. [Google Scholar] [CrossRef]
- Read, S.; Malmström, V.; Powrie, F. Cytotoxic T Lymphocyte–Associated Antigen 4 Plays an Essential Role in the Function of Cd25+Cd4+ Regulatory Cells That Control Intestinal Inflammation. J. Exp. Med. 2000, 192, 295–302. [Google Scholar] [CrossRef]
- Barski, A.; Cuddapah, S.; Cui, K.; Roh, T.-Y.; Schones, D.E.; Wang, Z.; Wei, G.; Chepelev, I.; Zhao, K. High-Resolution Profiling of Histone Methylations in the Human Genome. Cell 2007, 129, 823–837. [Google Scholar] [CrossRef]
- Wang, Z.; Schones, D.E.; Zhao, K. Characterization of human epigenomes. Curr. Opin. Genet. Dev. 2009, 19, 127–134. [Google Scholar] [CrossRef] [PubMed]
- McConathy, W.J.; Alaupovic, P. Studies on the isolation and partial characterization of apolipoprotein D and lipoprotein D of human plasma. Biochemistry. 1976, 15, 515–520. [Google Scholar] [CrossRef]
- Rassart, E.; Bedirian, A.; Carmo, S.D.; Guinard, O.; Sirois, J.; Terrisse, L.; Milne, R. Apolipoprotein D. Biochim. Biophys. Acta 2000, 1482, 185–198. [Google Scholar] [CrossRef]
- Rickhag, M.; Deierborg, T.; Patel, S.; Ruscher, K.; Wieloch, T. Apolipoprotein D is Elevated in Oligodendrocytes in the Peri-Infarct Region after Experimental Stroke: Influence of Enriched Environment. J. Cereb. Blood Flow Metab. 2008, 28, 551–562. [Google Scholar] [CrossRef] [PubMed]
- Eichinger, A.; Nasreen, A.; Kim, H.J.; Skerra, A. Structural Insight into the Dual Ligand Specificity and Mode of High Density Lipoprotein Association of Apolipoprotein D. J. Biol. Chem. 2007, 282, 31068–31075. [Google Scholar] [CrossRef] [PubMed]
- Perdomo, G.; Dong, H.H. Apolipoprotein D in Lipid Metabolism and Its Functional Implication in Atherosclerosis and Aging. Aging (Albany NY) 2009, 1, 17–27. [Google Scholar] [CrossRef] [PubMed]
- Cabral, J.H.M.; Atkins, G.L.; Sánchez, L.M.; López-Boado, Y.S.; Lopez-Otin, C.; Sawyer, L. Arachidonic acid binds to apolipoprotein D: Implications for the protein’s function. FEBS Lett. 1995, 366, 53–56. [Google Scholar] [CrossRef]
- Zeng, C.; Spielman, A.I.; Vowels, B.R.; Leyden, J.J.; Biemann, K.; Preti, G. A human axillary odorant is carried by apolipoprotein D. Proc. Natl. Acad. Sci. USA 1996, 93, 6626–6630. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, S.K.; Välikangas, T.; Shetty, A.; Khan, M.M.; Moulder, R.; Bhosale, S.D.; Komsi, E.; Salo, V.; De Albuquerque, R.S.; Rasool, O.; et al. Quantitative Proteomics Reveals the Dynamic Protein Landscape during Initiation of Human Th17 Cell Polarization. iScience 2019, 11, 334–355. [Google Scholar] [CrossRef]
- Cheng, X.; Yu, X.; Ding, Y.J.; Fu, Q.Q.; Xie, J.J.; Tang, T.T.; Yao, R.; Chen, Y.; Liao, Y.H. The th17/treg imbalance in patients with acute coronary syndrome. Clin. Immunol 2008, 127, 89–97. [Google Scholar] [CrossRef]
- Pejnovic, N.; Vratimos, A.; Lee, S.H.; Popadić, D.; Takeda, K.; Akira, S.; Chan, W.L. Increased atherosclerotic lesions and Th17 in interleukin-18 deficient apolipoprotein E-knockout mice fed high-fat diet. Mol. Immunol. 2009, 47, 37–45. [Google Scholar] [CrossRef]
- Shi, L.; Ji, Q.; Liu, L.; Shi, Y.; Lu, Z.; Ye, J.; Zeng, T.; Xue, Y.; Yang, Z.; Liu, Y.; et al. Il-22 produced by th22 cells aggravates atherosclerosis development in apoe(-/-) mice by enhancing dc-induced th17 cell proliferation. J. Cell Mol. Med. 2020, 24, 3064–3078. [Google Scholar] [CrossRef]
- Frostegård, J. Atherosclerosis in Patients With Autoimmune Disorders. Arter. Thromb. Vasc. Biol. 2005, 25, 1776–1785. [Google Scholar] [CrossRef]
- Matsuura, E.; Atzeni, F.; Sarzi-Puttini, P.; Turiel, M.; Lopez, L.R.; Nurmohamed, M.T. Is atherosclerosis an autoimmune disease? BMC Med. 2014, 12, 47. [Google Scholar] [CrossRef] [PubMed]
- Bérubé, N.G.; Swanson, X.H.; Bertram, M.; Kittle, J.D.; Didenko, V.; Baskin, D.S.; Smith, J.R.; Pereira-Smith, O.M. Cloning and characterization of CRF, a novel C1q-related factor, expressed in areas of the brain involved in motor function. Mol. Brain Res. 1999, 63, 233–240. [Google Scholar] [CrossRef]
- Iijima, T.; Miura, E.; Watanabe, M.; Yuzaki, M. Distinct expression of C1q-like family mRNAs in mouse brain and biochemical characterization of their encoded proteins. Eur. J. Neurosci. 2010, 31, 1606–1615. [Google Scholar] [CrossRef] [PubMed]
- Bolliger, M.F.; Martinelli, D.C.; Südhof, T.C. The cell-adhesion G protein-coupled receptor BAI3 is a high-affinity receptor for C1q-like proteins. Proc. Natl. Acad. Sci. U S A 2011, 108, 2534–2539. [Google Scholar] [CrossRef]
- Yuzaki, M. The C1q complement family of synaptic organizers: Not just complementary. Curr. Opin. Neurobiol. 2017, 45, 9–15. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Tan, A.; Yang, R.; Xue, Y.; Zhang, M.; Chen, L.; Xiao, L.; Yang, X.; Yu, Y. C1ql1/ctrp14 and c1ql4/ctrp11 promote angiogenesis of endothelial cells through activation of erk1/2 signal pathway. Mol. Cell Biochem. 2017, 424, 57–67. [Google Scholar] [CrossRef] [PubMed]
- Walsh, D.A.; I Pearson, C. Angiogenesis in the pathogenesis of inflammatory joint and lung diseases. Arthritis Res. 2001, 3, 147–153. [Google Scholar] [CrossRef] [PubMed]
- Aguilar-Cazares, D.; Chavez-Dominguez, R.; Carlos-Reyes, A.; Lopez-Camarillo, C.; De La Cruz, O.N.H.; Lopez-Gonzalez, J.S. Contribution of Angiogenesis to Inflammation and Cancer. Front. Oncol. 2019, 9, 1399. [Google Scholar] [CrossRef]
- Liu, J.; Duan, Y.; Cheng, X.; Chen, X.; Xie, W.; Long, H.; Lin, Z.H.; Zhu, B. IL-17 is associated with poor prognosis and promotes angiogenesis via stimulating VEGF production of cancer cells in colorectal carcinoma. Biochem. Biophys. Res. Commun. 2011, 407, 348–354. [Google Scholar] [CrossRef]
- Du, J.-W.; Xu, K.-Y.; Fang, L.-Y.; Qi, X.-L. Interleukin-17, produced by lymphocytes, promotes tumor growth and angiogenesis in a mouse model of breast cancer. Mol. Med. Rep. 2012, 6, 1099–1102. [Google Scholar] [CrossRef]
- Singh, T.P.; Schön, M.P.; Wallbrecht, K.; Gruber-Wackernagel, A.; Wang, X.-J.; Wolf, P. Involvement of IL-9 in Th17-Associated Inflammation and Angiogenesis of Psoriasis. PLoS ONE 2013, 8, e51752. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.-J.; Chen, Z.; Chamberlain, N.D.; Volin, M.V.; Swedler, W.; Volkov, S.; Sweiss, N.; Shahrara, S. Angiogenesis in rheumatoid arthritis is fostered directly by toll-like receptor 5 ligation and indirectly through interleukin-17 induction. Arthritis Rheum. 2013, 65, 2024–2036. [Google Scholar] [CrossRef] [PubMed]
- Kollmann, T.R.; Levy, O.; Montgomery, R.R.; Goriely, S. Innate immune function by Toll-like receptors: Distinct responses in newborns and the elderly. Immunity. 2012, 37, 771–783. [Google Scholar] [CrossRef]
- Lissner, M.M.; Thomas, B.J.; Wee, K.; Tong, A.-J.; Kollmann, T.R.; Smale, S.T. Age-Related Gene Expression Differences in Monocytes from Human Neonates, Young Adults, and Older Adults. PLoS ONE 2015, 10, e0132061. [Google Scholar] [CrossRef] [PubMed]
- Honey, K.; Nakagawa, T.; Peters, C.; Rudensky, A. Cathepsin L Regulates CD4+ T Cell Selection Independently of Its Effect on Invariant Chain. J. Exp. Med. 2002, 195, 1349–1358. [Google Scholar] [CrossRef] [PubMed]
- Honey, K.; Benlagha, K.; Beers, C.; Forbush, K.; Teyton, L.; Kleijmeer, M.J.; Rudensky, A.Y.; Bendelac, A. Thymocyte expression of cathepsin L is essential for NKT cell development. Nat. Immunol. 2002, 3, 1069–1074. [Google Scholar] [CrossRef]
- Hou, L.; Cooley, J.; Swanson, R.; Ong, P.C.; Pike, R.N.; Bogyo, M.; Olson, S.T.; Remold-O’Donnell, E. The protease cathepsin L regulates Th17 cell differentiation. J. Autoimmun. 2015, 65, 56–63. [Google Scholar] [CrossRef]
- Ruffell, B.; Affara, N.I.; Cottone, L.; Junankar, S.R.; Johansson, M.; DeNardo, D.G.; Korets, L.; Reinheckel, T.; Sloane, B.F.; Bogyo, M.; et al. Cathepsin C is a tissue-specific regulator of squamous carcinogenesis. Genes Dev. 2013, 27, 2086–2098. [Google Scholar] [CrossRef]
- Solau-Gervais, E.; Zerimech, F.; Lemaire, R.; Fontaine, C.; Huet, G.; Flipo, R.-M. Cysteine and serine proteases of synovial tissue in rheumatoid arthritis and osteoarthritis. Scand. J. Rheumatol. 2007, 36, 373–377. [Google Scholar] [CrossRef]
- Hsing, L.C.; Kirk, E.A.; McMillen, T.S.; Hsiao, S.-H.; Caldwell, M.; Houston, B.; Rudensky, A.Y.; Leboeuf, R.C. Roles for cathepsins S, L, and B in insulitis and diabetes in the NOD mouse. J. Autoimmun. 2010, 34, 96–104. [Google Scholar] [CrossRef] [PubMed]
- Vidak, E.; Javoršek, U.; Vizovišek, M.; Turk, B. Cysteine Cathepsins and their Extracellular Roles: Shaping the Microenvironment. Cells 2019, 8, 264. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Fabregas, J.; Prescott, A.; Van Kasteren, S.I.; Pedrioli, D.L.; McLean, W.H.I.; Moles, A.; Reinheckel, T.; Poli, V.; Watts, C. Lysosomal protease deficiency or substrate overload induces an oxidative-stress mediated STAT3-dependent pathway of lysosomal homeostasis. Nat. Commun. 2018, 9, 5343. [Google Scholar] [CrossRef] [PubMed]
- Jakoš, T.; Pišlar, A.; Jewett, A.; Kos, J. Cysteine Cathepsins in Tumor-Associated Immune Cells. Front. Immunol. 2019, 10, 2037. [Google Scholar] [CrossRef] [PubMed]
- Roh, T.Y.; Cuddapah, S.; Cui, K.; Zhao, K. The genomic landscape of histone modifications in human T cells. Proc. Natl. Acad. Sci. USA 2006, 103, 15782–15787. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Zang, C.; A Rosenfeld, J.; E Schones, D.; Barski, A.; Cuddapah, S.; Cui, K.; Roh, T.-Y.; Peng, W.; Zhang, M.Q.; et al. Combinatorial patterns of histone acetylations and methylations in the human genome. Nat. Genet. 2008, 40, 897–903. [Google Scholar] [CrossRef] [PubMed]
- Singh, P.; Cho, J.; Tsai, S.Y.; Rivas, G.E.; Larson, G.P.; Szabó, P. Coordinated allele-specific histone acetylation at the differentially methylated regions of imprinted genes. Nucleic Acids Res. 2010, 38, 7974–7990. [Google Scholar] [CrossRef]
- Akimzhanov, A.M.; O Yang, X.; Dong, C. Chromatin Remodeling of Interleukin-17 (IL-17)-IL-17F Cytokine Gene Locus during Inflammatory Helper T Cell Differentiation. J. Biol. Chem. 2007, 282, 5969–5972. [Google Scholar] [CrossRef]
- Yang, X.O.; Pappu, B.P.; Nurieva, R.; Akimzhanov, A.; Kang, H.S.; Chung, Y.; Ma, L.; Shah, B.; Panopoulos, A.D.; Schluns, K.S.; et al. T Helper 17 Lineage Differentiation Is Programmed by Orphan Nuclear Receptors RORα and RORγ. Immunity. 2008, 28, 29–39. [Google Scholar] [CrossRef]
- Xu, L.; Kitani, A.; Fuss, I.; Strober, W. Cutting edge: Regulatory T cells induce CD4+CD25-Foxp3- T cells or are self-induced to become Th17 cells in the absence of exogenous TGF-beta. J Immunol 2007, 178, 6725–6729. [Google Scholar] [CrossRef]
- Osorio, F.; LeibundGut-Landmann, S.; Lochner, M.; Lahl, K.; Sparwasser, T.; Eberl, G.; E Sousa, C.R. DC activated via dectin-1 convert Treg into IL-17 producers. Eur. J. Immunol. 2008, 38, 3274–3281. [Google Scholar] [CrossRef]
- Kleinewietfeld, M.; Hafler, D.A. The plasticity of human Treg and Th17 cells and its role in autoimmunity. Semin. Immunol. 2013, 25, 305–312. [Google Scholar] [CrossRef]
- Lee, Y.K.; Mukasa, R.; Hatton, R.D.; Weaver, C.T. Developmental plasticity of Th17 and Treg cells. Curr. Opin. Immunol. 2009, 21, 274–280. [Google Scholar] [CrossRef] [PubMed]
- Karmaus, P.W.; Chen, X.; Lim, S.A.; Herrada, A.A.; Nguyen, T.-L.M.; Xu, B.; Dhungana, Y.; Rankin, S.; Chen, W.; Rosencrance, C.; et al. Metabolic heterogeneity underlies reciprocal fates of TH17 cell stemness and plasticity. Nature 2019, 565, 101–105. [Google Scholar] [CrossRef] [PubMed]
- Wei, G.; Wei, L.; Zhu, J.; Zang, C.; Hu-Li, J.; Yao, Z.; Cui, K.; Kanno, Y.; Roh, T.-Y.; Watford, W.T.; et al. Global Mapping of H3K4me3 and H3K27me3 Reveals Specificity and Plasticity in Lineage Fate Determination of Differentiating CD4+ T Cells. Immunity. 2009, 30, 155–167. [Google Scholar] [CrossRef] [PubMed]
- Dong, C. Genetic controls of Th17 cell differentiation and plasticity. Exp. Mol. Med. 2010, 43, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Paksa, A.; Rajagopal, J. The epigenetic basis of cellular plasticity. Curr. Opin. Cell Biol. 2018, 49, 116–122. [Google Scholar] [CrossRef] [PubMed]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sałkowska, A.; Karaś, K.; Karwaciak, I.; Walczak-Drzewiecka, A.; Krawczyk, M.; Sobalska-Kwapis, M.; Dastych, J.; Ratajewski, M. Identification of Novel Molecular Markers of Human Th17 Cells. Cells 2020, 9, 1611. https://doi.org/10.3390/cells9071611
Sałkowska A, Karaś K, Karwaciak I, Walczak-Drzewiecka A, Krawczyk M, Sobalska-Kwapis M, Dastych J, Ratajewski M. Identification of Novel Molecular Markers of Human Th17 Cells. Cells. 2020; 9(7):1611. https://doi.org/10.3390/cells9071611
Chicago/Turabian StyleSałkowska, Anna, Kaja Karaś, Iwona Karwaciak, Aurelia Walczak-Drzewiecka, Mariusz Krawczyk, Marta Sobalska-Kwapis, Jarosław Dastych, and Marcin Ratajewski. 2020. "Identification of Novel Molecular Markers of Human Th17 Cells" Cells 9, no. 7: 1611. https://doi.org/10.3390/cells9071611
APA StyleSałkowska, A., Karaś, K., Karwaciak, I., Walczak-Drzewiecka, A., Krawczyk, M., Sobalska-Kwapis, M., Dastych, J., & Ratajewski, M. (2020). Identification of Novel Molecular Markers of Human Th17 Cells. Cells, 9(7), 1611. https://doi.org/10.3390/cells9071611





