Characterization of RNP Networks of PUM1 and PUM2 Post-Transcriptional Regulators in TCam-2 Cells, a Human Male Germ Cell Model
Abstract
1. Introduction
2. Materials and Methods
2.1. RNA Immunoprecipitation and Sequencing
2.2. Western Blot Analysis
2.3. Bioinformatic Analysis of PUM1- and PUM2-Bound mRNAs
2.4. siRNA Silencing of PUM Proteins
2.5. Bioinformatic Identification of mRNAs Under Regulation by PUM Proteins
2.6. RT-qPCR Analysis of mRNA Expression after PUM1 and PUM2 Knockdown
2.7. Mass Spectrometry Analysis after Anti-PUM1 and Anti-PUM2 Immunoprecipitation
2.8. Bioinformatic Construction of the PUM RNA Regulon
2.9. Quantification and Statistical Analysis
3. Results
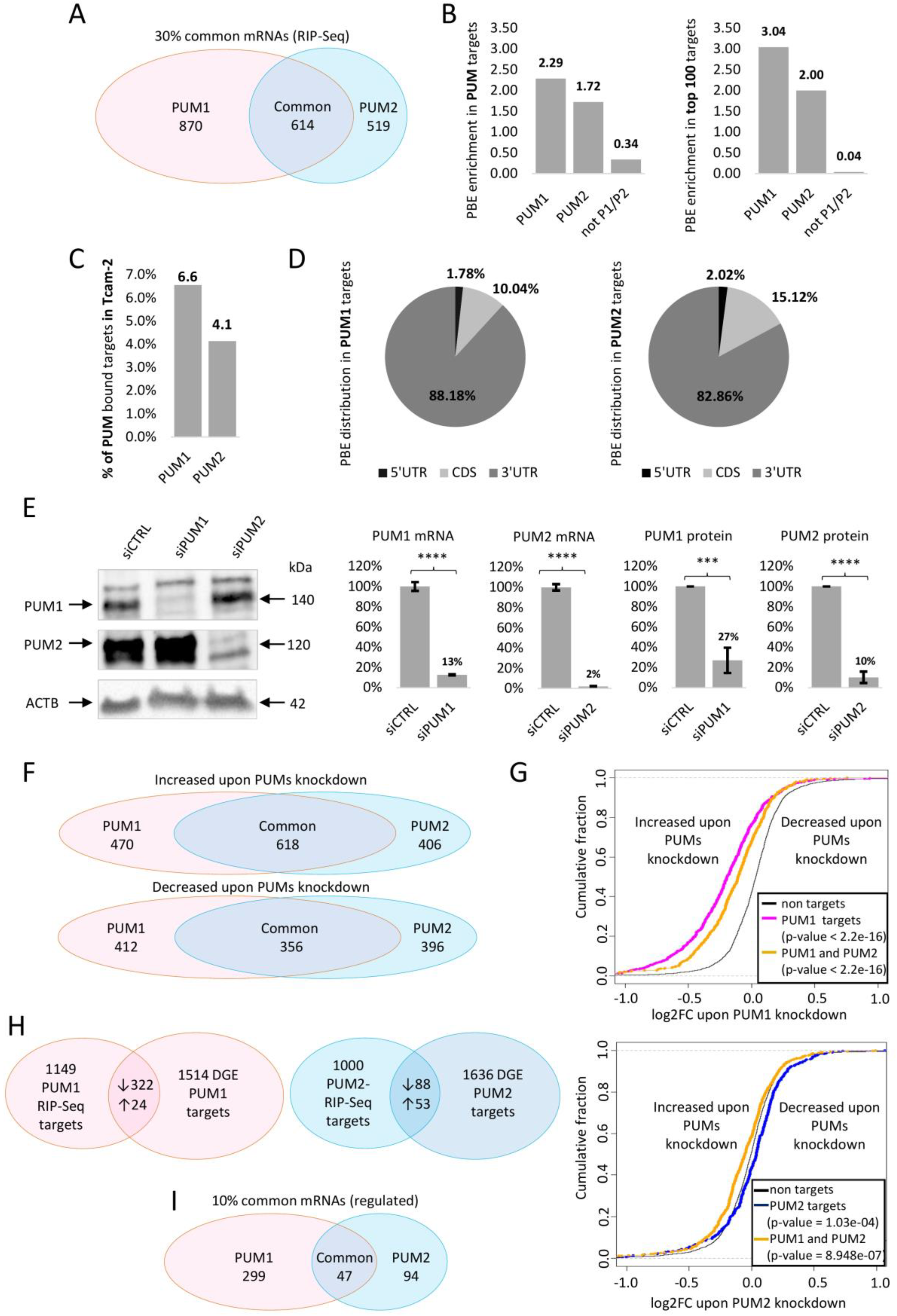
3.1. Identification of PUM1 and PUM2 mRNA Targets by RIP-Seq
3.2. Differential Gene Expression Analysis upon PUM1 or PUM2 siRNA Knockdown
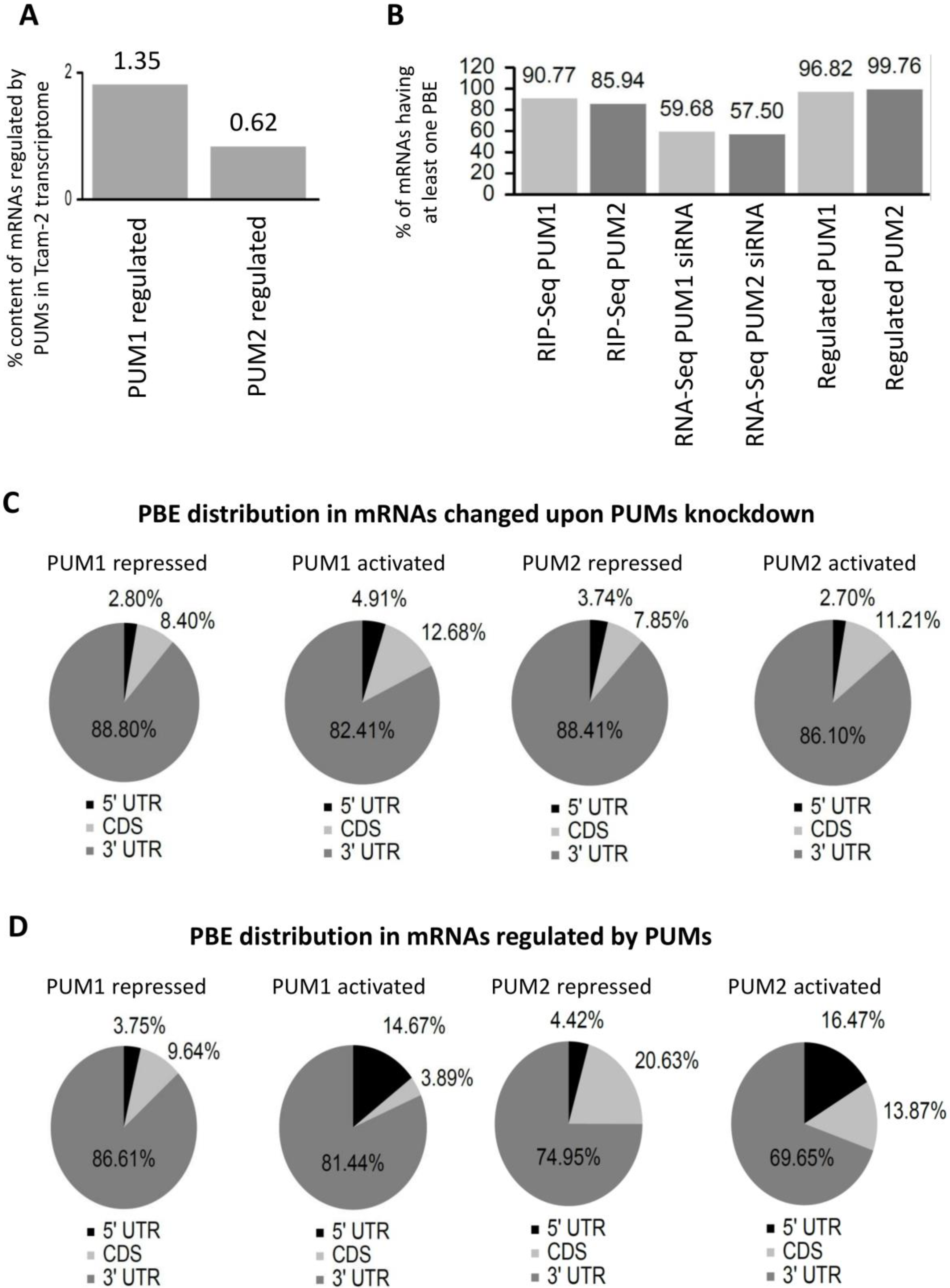
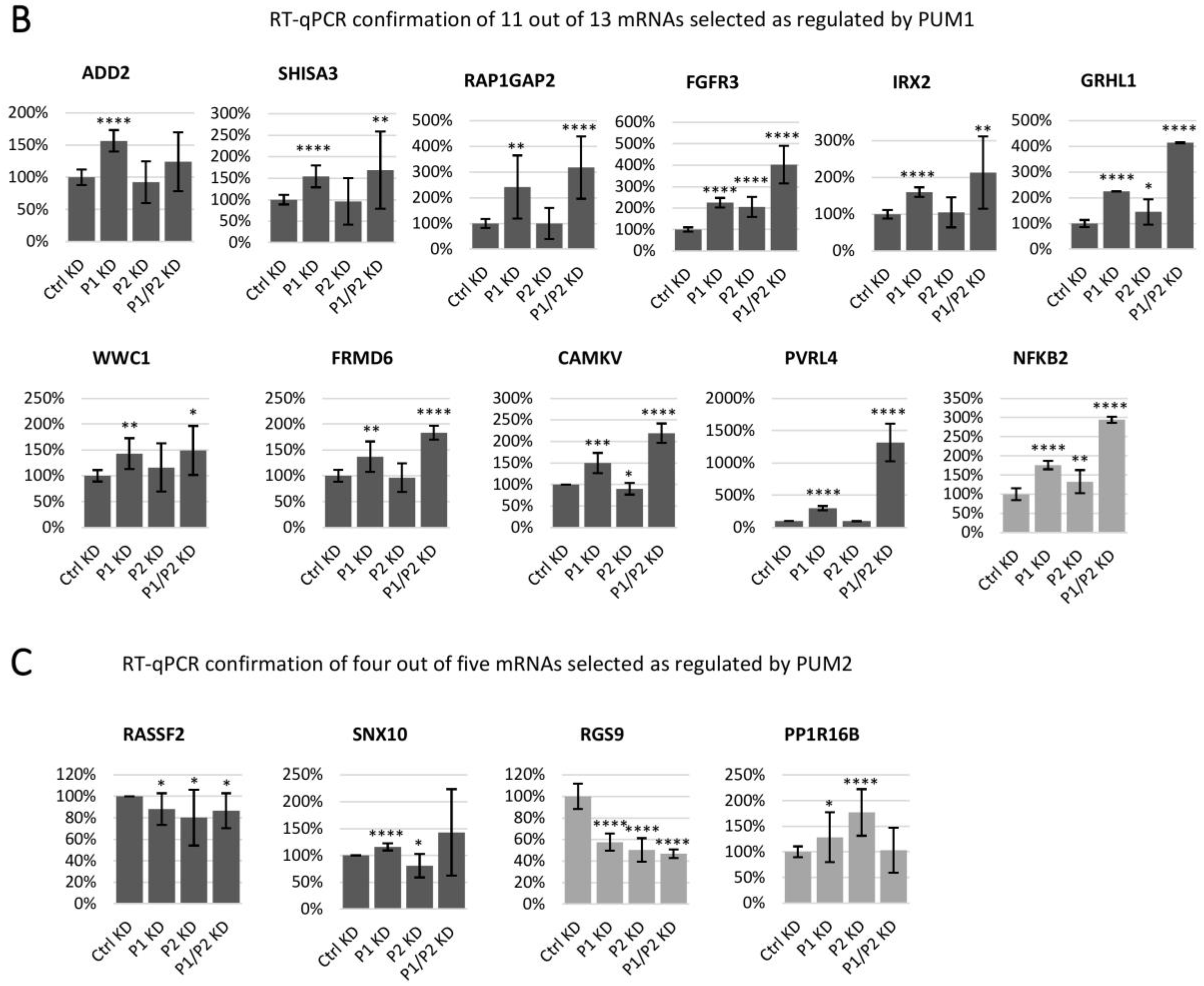
3.3. Selection of PUM1- and PUM2-Regulated mRNA Targets Based on RIP-Seq and PUMs Knockdown Analysis
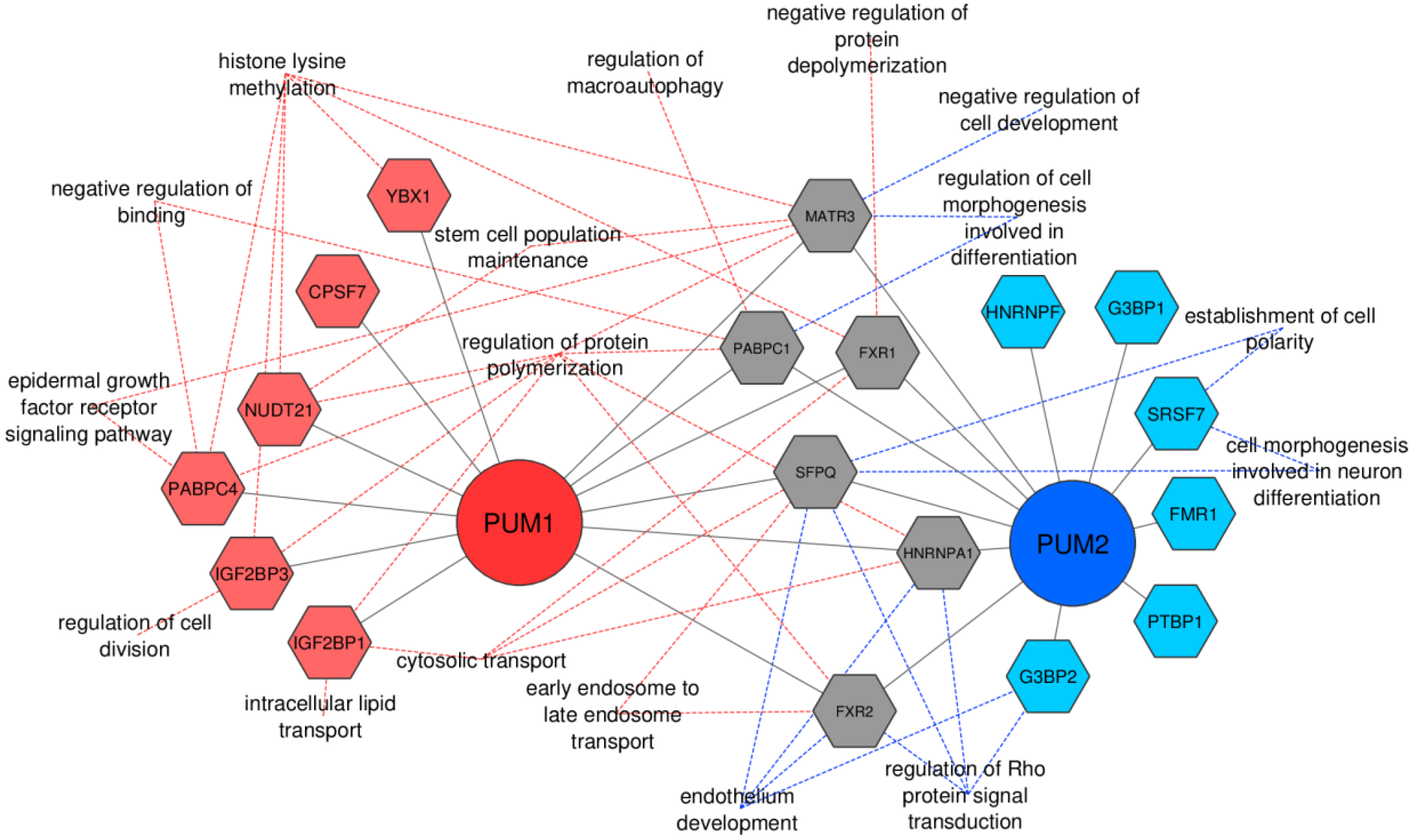
3.4. Global Profiling Reveals Many Putative Protein Cofactors Interacting with PUM1 and PUM2 to be RBPs
3.5. PUM1 and PUM2 Form Separate Regulons in TCam-2 Cells
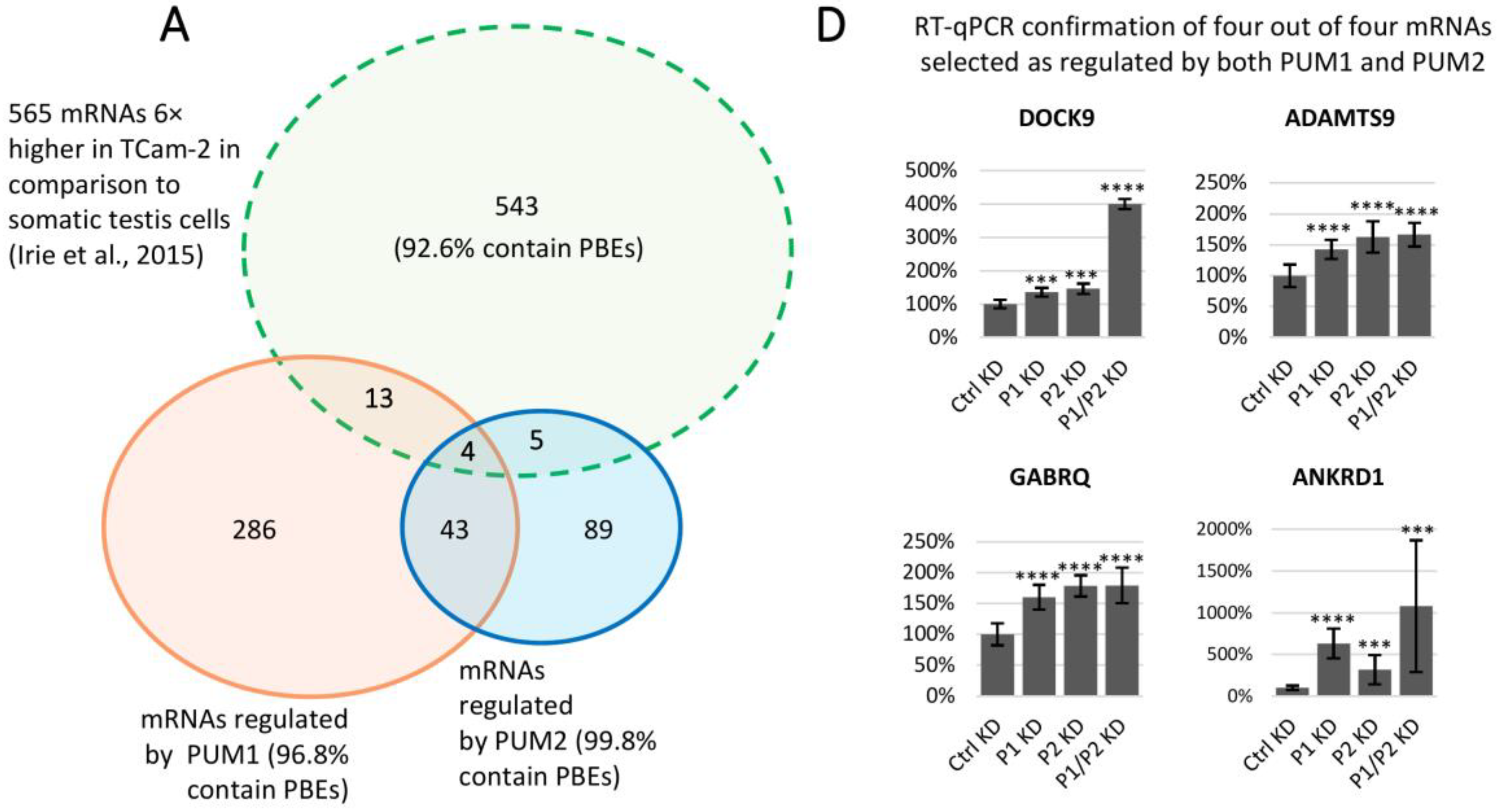
3.6. Several mRNAs Highly Expressed in TCam-2 Cells Compared to Somatic Gonadal Tissue are Regulated by PUM Proteins
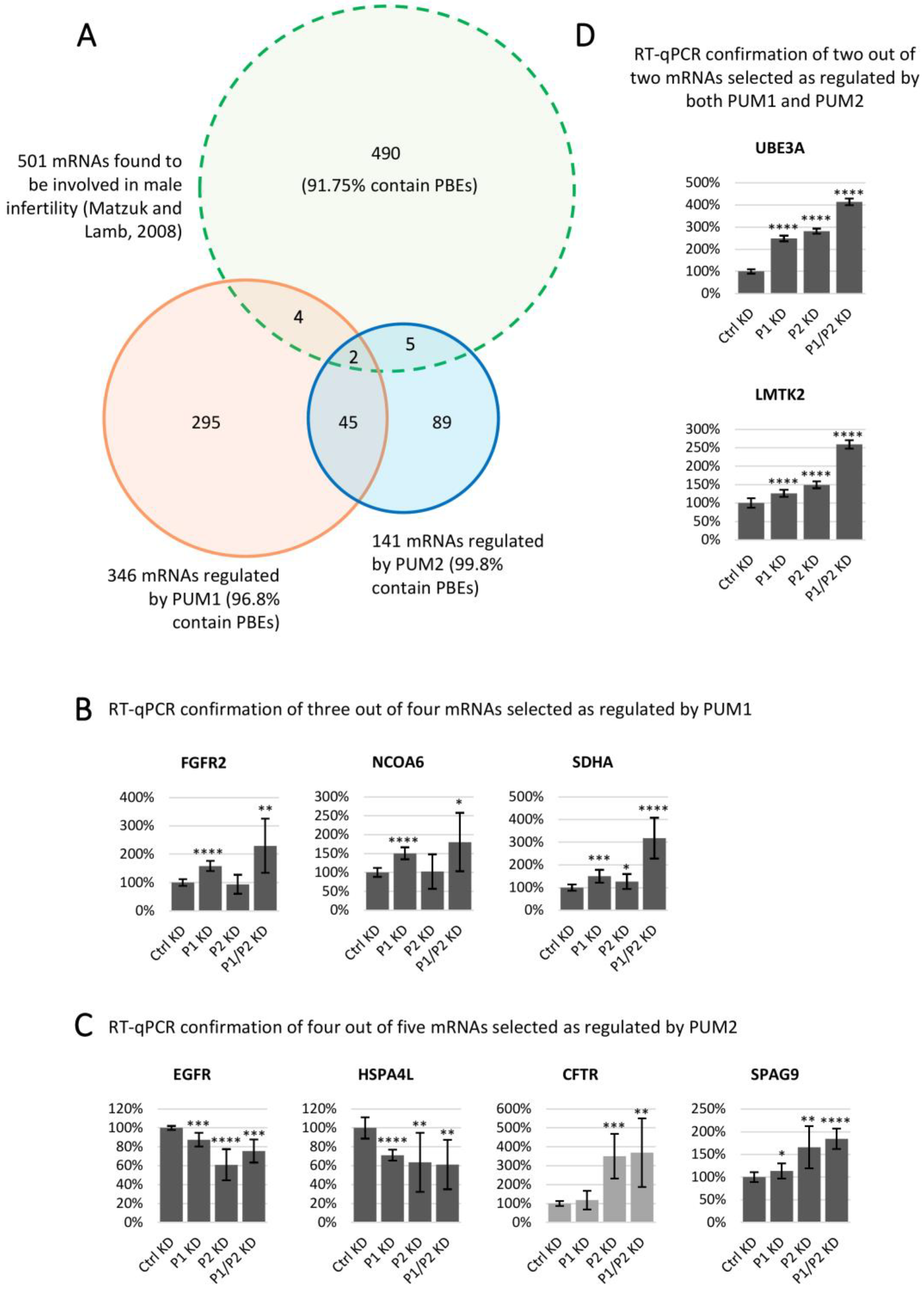
3.7. Genes Associated with Male Infertility in Humans and/or Mice are Regulated by PUM Proteins
4. Discussion
5. Conclusions
Supplementary Materials
Data Availability
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Gerstberger, S.; Hafner, M.; Tuschl, T. A census of human RNA-binding proteins. Nat. Rev. Genet. 2014, 15, 829–845. [Google Scholar] [CrossRef]
- Mukherjee, N.; Wessels, H.H.; Lebedeva, S.; Sajek, M.; Ghanbari, M.; Garzia, A.; Munteanu, A.; Yusuf, D.; Farazi, T.; Hoell, J.I.; et al. Deciphering human ribonucleoprotein regulatory networks. Nucleic Acids Res. 2019, 47, 570–581. [Google Scholar] [CrossRef] [PubMed]
- Fredericks, A.M.; Cygan, K.J.; Brown, B.A.; Fairbrother, W.G. RNA-Binding Proteins: Splicing Factors and Disease. Biomolecules 2015, 5, 893–909. [Google Scholar] [CrossRef] [PubMed]
- Goldstrohm, A.C.; Hall, T.M.T.; McKenney, K.M. Post-transcriptional Regulatory Functions of Mammalian Pumilio Proteins. Trends. Genet. 2018. [Google Scholar] [CrossRef] [PubMed]
- Uyhazi, K.E. Pumilio Proteins Regulate Translation in Embryonic Stem Cells and are Essential for Early Embryonic Development. Ph.D. Thesis, Yale University, New Haven, CT, USA, 2012. [Google Scholar]
- Wang, X.; McLachlan, J.; Zamore, P.D.; Hall, T.M. Modular recognition of RNA by a human pumilio-homology domain. Cell 2002, 110, 501–512. [Google Scholar] [CrossRef]
- Gerber, A.P.; Herschlag, D.; Brown, P.O. Extensive association of functionally and cytotopically related mRNAs with Puf family RNA-binding proteins in yeast. PLoS Biol. 2004, 2, E79. [Google Scholar] [CrossRef] [PubMed]
- Keene, J.D. RNA regulons: Coordination of post-transcriptional events. Nat. Rev. Genet. 2007, 8, 533–543. [Google Scholar] [CrossRef]
- Spassov, D.S.; Jurecic, R. The PUF family of RNA-binding proteins: Does evolutionarily conserved structure equal conserved function? IUBMB Life 2003, 55, 359–366. [Google Scholar] [CrossRef]
- Galgano, A.; Forrer, M.; Jaskiewicz, L.; Kanitz, A.; Zavolan, M.; Gerber, A.P. Comparative analysis of mRNA targets for human PUF-family proteins suggests extensive interaction with the miRNA regulatory system. PLoS ONE 2008, 3, e3164. [Google Scholar] [CrossRef]
- Sternburg, E.L.; Estep, J.A.; Nguyen, D.K.; Li, Y.; Karginov, F.V. Antagonistic and cooperative AGO2-PUM interactions in regulating mRNAs. Sci. Rep. 2018, 8, 15316. [Google Scholar] [CrossRef]
- Sajek, M.; Janecki, D.M.; Smialek, M.J.; Ginter-Matuszewska, B.; Spik, A.; Oczkowski, S.; Ilaslan, E.; Kusz-Zamelczyk, K.; Kotecki, M.; Blazewicz, J.; et al. PUM1 and PUM2 exhibit different modes of regulation for SIAH1 that involve cooperativity with NANOS paralogues. Cell. Mol. Life Sci. 2019, 76, 147–161. [Google Scholar] [CrossRef]
- Weidmann, C.A.; Goldstrohm, A.C. Drosophila Pumilio protein contains multiple autonomous repression domains that regulate mRNAs independently of Nanos and brain tumor. Mol. Cell. Biol. 2012, 32, 527–540. [Google Scholar] [CrossRef] [PubMed]
- Jaruzelska, J.; Kotecki, M.; Kusz, K.; Spik, A.; Firpo, M.; Reijo Pera, R.A. Conservation of a Pumilio-Nanos complex from Drosophila germ plasm to human germ cells. Dev. Genes Evol. 2003, 213, 120–126. [Google Scholar] [CrossRef] [PubMed]
- Kusz-Zamelczyk, K.; Sajek, M.; Spik, A.; Glazar, R.; Jedrzejczak, P.; Latos-Bielenska, A.; Kotecki, M.; Pawelczyk, L.; Jaruzelska, J. Mutations of NANOS1, a human homologue of the Drosophila morphogen, are associated with a lack of germ cells in testes or severe oligo-astheno-teratozoospermia. J. Med. Genet. 2013, 50, 187–193. [Google Scholar] [CrossRef] [PubMed]
- Moore, F.L.; Jaruzelska, J.; Fox, M.S.; Urano, J.; Firpo, M.T.; Turek, P.J.; Dorfman, D.M.; Pera, R.A. Human Pumilio-2 is expressed in embryonic stem cells and germ cells and interacts with DAZ (Deleted in AZoospermia) and DAZ-like proteins. Proc. Natl. Acad. Sci. USA 2003, 100, 538–543. [Google Scholar] [CrossRef]
- Reijo, R.; Lee, T.Y.; Salo, P.; Alagappan, R.; Brown, L.G.; Rosenberg, M.; Rozen, S.; Jaffe, T.; Straus, D.; Hovatta, O.; et al. Diverse spermatogenic defects in humans caused by Y chromosome deletions encompassing a novel RNA-binding protein gene. Nat. Genet. 1995, 10, 383–393. [Google Scholar] [CrossRef]
- Santos, M.G.; Machado, A.Z.; Martins, C.N.; Domenice, S.; Costa, E.M.; Nishi, M.Y.; Ferraz-de-Souza, B.; Jorge, S.A.; Pereira, C.A.; Soardi, F.C.; et al. Homozygous inactivating mutation in NANOS3 in two sisters with primary ovarian insufficiency. Biomed. Res. Int. 2014, 2014, 787465. [Google Scholar] [CrossRef]
- Wu, X.; Wang, B.; Dong, Z.; Zhou, S.; Liu, Z.; Shi, G.; Cao, Y.; Xu, Y. A NANOS3 mutation linked to protein degradation causes premature ovarian insufficiency. Cell Death Dis. 2013, 4, e825. [Google Scholar] [CrossRef]
- O’Flynn O’Brien, K.L.; Varghese, A.C.; Agarwal, A. The genetic causes of male factor infertility: A review. Fertil. Steril. 2010, 93, 1–12. [Google Scholar] [CrossRef]
- Ibtisham, F.; Wu, J.; Xiao, M.; An, L.; Banker, Z.; Nawab, A.; Zhao, Y.; Li, G. Progress and future prospect of in vitro spermatogenesis. Oncotarget 2017, 8, 66709–66727. [Google Scholar] [CrossRef]
- Rajpert-De Meyts, E.; McGlynn, K.A.; Okamoto, K.; Jewett, M.A.; Bokemeyer, C. Testicular germ cell tumours. Lancet 2016, 387, 1762–1774. [Google Scholar] [CrossRef]
- Van de Geijn, G.J.; Hersmus, R.; Looijenga, L.H. Recent developments in testicular germ cell tumor research. Birth Defects Res. C Embryo Today 2009, 87, 96–113. [Google Scholar] [CrossRef] [PubMed]
- De Jong, J.; Stoop, H.; Gillis, A.J.; Hersmus, R.; van Gurp, R.J.; van de Geijn, G.J.; van Drunen, E.; Beverloo, H.B.; Schneider, D.T.; Sherlock, J.K.; et al. Further characterization of the first seminoma cell line TCam-2. Genes Chromosomes Cancer 2008, 47, 185–196. [Google Scholar] [CrossRef] [PubMed]
- Prasad, A.; Porter, D.F.; Kroll-Conner, P.L.; Mohanty, I.; Ryan, A.R.; Crittenden, S.L.; Wickens, M.; Kimble, J. The PUF binding landscape in metazoan germ cells. RNA 2016, 22, 1026–1043. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Pachter, L.; Salzberg, S.L. TopHat: Discovering splice junctions with RNA-Seq. Bioinformatics 2009, 25, 1105–1111. [Google Scholar] [CrossRef]
- Maere, S.; Heymans, K.; Kuiper, M. BiNGO: A Cytoscape plugin to assess overrepresentation of gene ontology categories in biological networks. Bioinformatics 2005, 21, 3448–3449. [Google Scholar] [CrossRef]
- Karolchik, D.; Hinrichs, A.S.; Furey, T.S.; Roskin, K.M.; Sugnet, C.W.; Haussler, D.; Kent, W.J. The UCSC Table Browser data retrieval tool. Nucleic Acids Res. 2004, 32, D493–D496. [Google Scholar] [CrossRef]
- Barsnes, H.; Vaudel, M. SearchGUI: A Highly Adaptable Common Interface for Proteomics Search and de Novo Engines. J. Proteome Res. 2018, 17, 2552–2555. [Google Scholar] [CrossRef]
- Elias, J.E.; Gygi, S.P. Target-decoy search strategy for mass spectrometry-based proteomics. Methods Mol. Biol. 2010, 604, 55–71. [Google Scholar] [CrossRef]
- Apweiler, R.; Bairoch, A.; Wu, C.H.; Barker, W.C.; Boeckmann, B.; Ferro, S.; Gasteiger, E.; Huang, H.; Lopez, R.; Magrane, M.; et al. UniProt: The Universal Protein knowledgebase. Nucleic Acids Res. 2004, 32, D115–D119. [Google Scholar] [CrossRef]
- Vaudel, M.; Burkhart, J.M.; Zahedi, R.P.; Oveland, E.; Berven, F.S.; Sickmann, A.; Martens, L.; Barsnes, H. PeptideShaker enables reanalysis of MS-derived proteomics data sets. Nat. Biotechnol. 2015, 33, 22–24. [Google Scholar] [CrossRef] [PubMed]
- Cook, K.B.; Kazan, H.; Zuberi, K.; Morris, Q.; Hughes, T.R. RBPDB: A database of RNA-binding specificities. Nucleic Acids Res. 2011, 39, D301–D308. [Google Scholar] [CrossRef] [PubMed]
- Ray, D.; Kazan, H.; Cook, K.B.; Weirauch, M.T.; Najafabadi, H.S.; Li, X.; Gueroussov, S.; Albu, M.; Zheng, H.; Yang, A.; et al. A compendium of RNA-binding motifs for decoding gene regulation. Nature 2013, 499, 172–177. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Xu, G.; Yang, Y.T.; Xu, Z.; Chen, X.; Shi, B.; Xie, D.; Lu, Z.J.; Wang, P. POSTAR2: Deciphering the post-transcriptional regulatory logics. Nucleic Acids Res. 2019, 47, D203–D211. [Google Scholar] [CrossRef]
- Grant, C.E.; Bailey, T.L.; Noble, W.S. FIMO: Scanning for occurrences of a given motif. Bioinformatics 2011, 27, 1017–1018. [Google Scholar] [CrossRef]
- Bindea, G.; Mlecnik, B.; Hackl, H.; Charoentong, P.; Tosolini, M.; Kirilovsky, A.; Fridman, W.H.; Pages, F.; Trajanoski, Z.; Galon, J. ClueGO: A Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 2009, 25, 1091–1093. [Google Scholar] [CrossRef]
- Irie, N.; Weinberger, L.; Tang, W.W.; Kobayashi, T.; Viukov, S.; Manor, Y.S.; Dietmann, S.; Hanna, J.H.; Surani, M.A. SOX17 is a critical specifier of human primordial germ cell fate. Cell 2015, 160, 253–268. [Google Scholar] [CrossRef]
- Ramirez, F.; Ryan, D.P.; Gruning, B.; Bhardwaj, V.; Kilpert, F.; Richter, A.S.; Heyne, S.; Dundar, F.; Manke, T. deepTools2: A next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 2016, 44, W160–W165. [Google Scholar] [CrossRef]
- Bohn, J.A.; Van Etten, J.L.; Schagat, T.L.; Bowman, B.M.; McEachin, R.C.; Freddolino, P.L.; Goldstrohm, A.C. Identification of diverse target RNAs that are functionally regulated by human Pumilio proteins. Nucleic Acids Res. 2018, 46, 362–386. [Google Scholar] [CrossRef]
- Morris, A.R.; Mukherjee, N.; Keene, J.D. Ribonomic analysis of human Pum1 reveals cis-trans conservation across species despite evolution of diverse mRNA target sets. Mol. Cell. Biol. 2008, 28, 4093–4103. [Google Scholar] [CrossRef]
- Jarmoskaite, I.; Denny, S.K.; Vaidyanathan, P.P.; Becker, W.R.; Andreasson, J.O.L.; Layton, C.J.; Kappel, K.; Shivashankar, V.; Sreenivasan, R.; Das, R.; et al. A Quantitative and Predictive Model for RNA Binding by Human Pumilio Proteins. Mol. Cell 2019, 74, 966–981.e18. [Google Scholar] [CrossRef]
- Hafner, M.; Landthaler, M.; Burger, L.; Khorshid, M.; Hausser, J.; Berninger, P.; Rothballer, A.; Ascano, M., Jr.; Jungkamp, A.C.; Munschauer, M.; et al. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 2010, 141, 129–141. [Google Scholar] [CrossRef] [PubMed]
- Matzuk, M.M.; Lamb, D.J. The biology of infertility: Research advances and clinical challenges. Nat. Med. 2008, 14, 1197–1213. [Google Scholar] [CrossRef] [PubMed]
- Kershner, A.M.; Kimble, J. Genome-wide analysis of mRNA targets for Caenorhabditis elegans FBF, a conserved stem cell regulator. Proc. Natl. Acad. Sci. USA 2010, 107, 3936–3941. [Google Scholar] [CrossRef] [PubMed]
- Hafner, M.; Landthaler, M.; Burger, L.; Khorshid, M.; Hausser, J.; Berninger, P.; Rothballer, A.; Ascano, M.; Jungkamp, A.C.; Munschauer, M.; et al. PAR-CliP--a method to identify transcriptome-wide the binding sites of RNA binding proteins. J. Vis. Exp. 2010. [Google Scholar] [CrossRef]
- Cottrell, K.A.; Chaudhari, H.G.; Cohen, B.A.; Djuranovic, S. PTRE-seq reveals mechanism and interactions of RNA binding proteins and miRNAs. Nat. Commun. 2018, 9, 301. [Google Scholar] [CrossRef]
- Hubstenberger, A.; Courel, M.; Benard, M.; Souquere, S.; Ernoult-Lange, M.; Chouaib, R.; Yi, Z.; Morlot, J.B.; Munier, A.; Fradet, M.; et al. P-Body Purification Reveals the Condensation of Repressed mRNA Regulons. Mol. Cell 2017, 68, 144–157.e5. [Google Scholar] [CrossRef]
- Hein, M.Y.; Hubner, N.C.; Poser, I.; Cox, J.; Nagaraj, N.; Toyoda, Y.; Gak, I.A.; Weisswange, I.; Mansfeld, J.; Buchholz, F.; et al. A human interactome in three quantitative dimensions organized by stoichiometries and abundances. Cell 2015, 163, 712–723. [Google Scholar] [CrossRef]
- Jain, S.; Wheeler, J.R.; Walters, R.W.; Agrawal, A.; Barsic, A.; Parker, R. ATPase-Modulated Stress Granules Contain a Diverse Proteome and Substructure. Cell 2016, 164, 487–498. [Google Scholar] [CrossRef]
- Vessey, J.P.; Vaccani, A.; Xie, Y.; Dahm, R.; Karra, D.; Kiebler, M.A.; Macchi, P. Dendritic localization of the translational repressor Pumilio 2 and its contribution to dendritic stress granules. J. Neurosci. 2006, 26, 6496–6508. [Google Scholar] [CrossRef]
- Zhang, M.; Chen, D.; Xia, J.; Han, W.; Cui, X.; Neuenkirchen, N.; Hermes, G.; Sestan, N.; Lin, H. Post-transcriptional regulation of mouse neurogenesis by Pumilio proteins. Genes Dev. 2017, 31, 1354–1369. [Google Scholar] [CrossRef] [PubMed]
- Kedde, M.; van Kouwenhove, M.; Zwart, W.; Oude Vrielink, J.A.; Elkon, R.; Agami, R. A Pumilio-induced RNA structure switch in p27-3′ UTR controls miR-221 and miR-222 accessibility. Nat. Cell Biol. 2010, 12, 1014–1020. [Google Scholar] [CrossRef] [PubMed]
- Smialek, M.J.; Kuczynska, B.; Ilaslan, E.; Janecki, D.M.; Sajek, M.P.; Kusz-Zamelczyk, K.; Jaruzelska, J. Kinesin KIF18A is a novel PUM regulated target promoting mitotic progression and survival of human male germ cell line. J. Cell Sci. 2020. [Google Scholar] [CrossRef] [PubMed]
- Uyhazi, K.E.; Yang, Y.; Liu, N.; Qi, H.; Huang, X.A.; Mak, W.; Weatherbee, S.D.; de Prisco, N.; Gennarino, V.A.; Song, X.; et al. Pumilio proteins utilize distinct regulatory mechanisms to achieve complementary functions required for pluripotency and embryogenesis. Proc. Natl. Acad. Sci. USA 2020. [Google Scholar] [CrossRef]
- Vizcaino, J.A.; Csordas, A.; Del-Toro, N.; Dianes, J.A.; Griss, J.; Lavidas, I.; Mayer, G.; Perez-Riverol, Y.; Reisinger, F.; Ternent, T.; et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 2016, 44, 11033. [Google Scholar] [CrossRef]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Smialek, M.J.; Ilaslan, E.; Sajek, M.P.; Swiercz, A.; Janecki, D.M.; Kusz-Zamelczyk, K.; Wozniak, T.; Kotecki, M.; Handschuh, L.; Figlerowicz, M.; et al. Characterization of RNP Networks of PUM1 and PUM2 Post-Transcriptional Regulators in TCam-2 Cells, a Human Male Germ Cell Model. Cells 2020, 9, 984. https://doi.org/10.3390/cells9040984
Smialek MJ, Ilaslan E, Sajek MP, Swiercz A, Janecki DM, Kusz-Zamelczyk K, Wozniak T, Kotecki M, Handschuh L, Figlerowicz M, et al. Characterization of RNP Networks of PUM1 and PUM2 Post-Transcriptional Regulators in TCam-2 Cells, a Human Male Germ Cell Model. Cells. 2020; 9(4):984. https://doi.org/10.3390/cells9040984
Chicago/Turabian StyleSmialek, Maciej J., Erkut Ilaslan, Marcin P. Sajek, Aleksandra Swiercz, Damian M. Janecki, Kamila Kusz-Zamelczyk, Tomasz Wozniak, Maciej Kotecki, Luiza Handschuh, Marek Figlerowicz, and et al. 2020. "Characterization of RNP Networks of PUM1 and PUM2 Post-Transcriptional Regulators in TCam-2 Cells, a Human Male Germ Cell Model" Cells 9, no. 4: 984. https://doi.org/10.3390/cells9040984
APA StyleSmialek, M. J., Ilaslan, E., Sajek, M. P., Swiercz, A., Janecki, D. M., Kusz-Zamelczyk, K., Wozniak, T., Kotecki, M., Handschuh, L., Figlerowicz, M., & Jaruzelska, J. (2020). Characterization of RNP Networks of PUM1 and PUM2 Post-Transcriptional Regulators in TCam-2 Cells, a Human Male Germ Cell Model. Cells, 9(4), 984. https://doi.org/10.3390/cells9040984






