Abstract
Rheumatoid arthritis (RA) is a chronic systemic autoimmune disease that affects synovial joints, leading to inflammation, joint destruction, loss of function, and disability. Although recent pharmaceutical advances have improved the treatment of RA, patients often inquire about dietary interventions to improve RA symptoms, as they perceive pain and/or swelling after the consumption or avoidance of certain foods. There is evidence that some foods have pro- or anti-inflammatory effects mediated by diet-related metabolites. In addition, recent literature has shown a link between diet-related metabolites and microbiome changes, since the gut microbiome is involved in the metabolism of some dietary ingredients. But diet and the gut microbiome are not the only factors linked to circulating pro- and anti-inflammatory metabolites. Other factors including smoking, associated comorbidities, and therapeutic drugs might also modify the circulating metabolomic profile and play a role in RA pathogenesis. This article summarizes what is known about circulating pro- and anti-inflammatory metabolites in RA. It also emphasizes factors that might be involved in their circulating concentrations and diet-related metabolites with a beneficial effect in RA.
1. Introduction
Rheumatoid arthritis (RA) is an autoimmune inflammatory arthritis that affects approximately 1% of the world’s population. It is a potentially debilitating disease that affects women two to three times more frequently than men [1]. It is characterized by pain and swelling in joints and produces irreversible joint damage that negatively affects patients’ quality of life in the absence of treatment. In our clinical practice, patients often mention changes in their symptoms after the consumption or avoidance of certain foods and inquire about the adequate type of diet for this disease. However, there is very little knowledge on how diet or specific ingredients affect pain and inflammation in RA. Recently, a lot of research on diet, gut microbiome, and gut-microbe-derived metabolites has focused on explaining how this diet–microbiome-metabolomic axis can explain different symptoms and overall health status.
Several studies have employed different analytical methods (mass spectrometry, MS, nuclear magnetic resonance, NMR) to characterize the metabolomic profile in the blood (serum or plasma), urine, or synovial fluid in patients with rheumatoid arthritis. Due to the heterogeneity of the methods that were used, the results of most of the studies are not comparable; however, there are metabolites with similar changes across multiple studies (Figure 1 and Table 1). The objective of this work is to review the existing evidence for the relationship between diet, metabolites, and inflammation in RA.
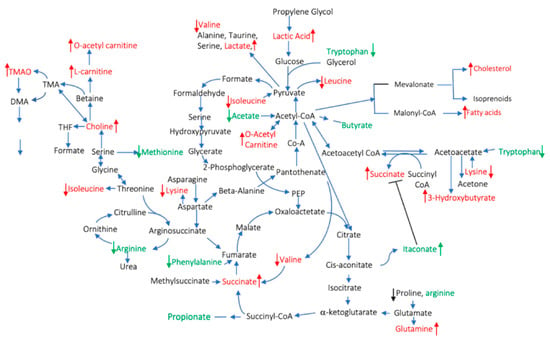
Figure 1.
Pro- and anti-inflammatory circulating metabolites described in rheumatoid arthritis (RA) patients. The red color indicates pro- and the green indicates anti-inflammatory metabolites. The arrows indicate increases/decreased concentrations of the metabolites compared to healthy controls.

Table 1.
Metabolic profile changes in plasma or serum of patients diagnosed with rheumatoid arthritis (RA). DMARD: disease-modifying antirheumatic drugs; GC: glucocorticoids; pSS primary Sjogren syndrome.
2. Factors That Influence Circulating Metabolites and Their Potential Role in Rheumatoid Arthritis
Metabolites reflect an organism’s state, which results from the interaction of internal and external factors, such as genetic and environmental/lifestyle factors, respectively. In disease states, the circulating metabolites are also affected by the pathological processes, and there are already well-studied metabolites that are considered to be disease reporters, like the increase of blood glucose levels in diabetes mellitus. In systemic diseases such as RA, the abnormal circulating metabolomic profile might reflect genetic predisposition, local inflammation, comorbidities, and several environmental factors including diet, smoking, or microbiome (Figure 2).
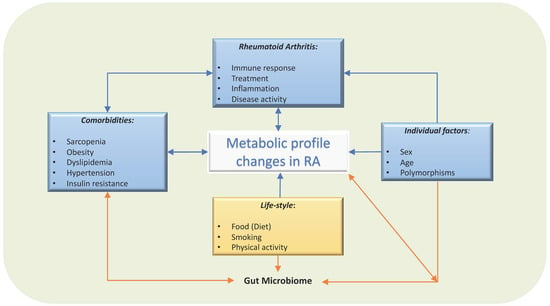
Figure 2.
Factors involved in circulating metabolic profile in patients with RA. Several factors influence the circulating metabolites levels. Not only dietary factors or local synovial metabolites, but also comorbidities, treatment and individual factors, such as sex, age and genetics, will modify their metabolism, gut microbiome, and therefore, the circulating metabolic profile.
2.1. Diet
Amongst environmental factors, diet is one that directly affects circulating metabolites. For example, essential amino acids (histidine, isoleucine, leucine, lysine, methionine, phenylalanine, threonine, tryptophan, and valine) and essential fatty acids (alpha-linolenic acid and linoleic acid) come from the diet. Of interest, some of these essential nutrients were found to be low in RA patients, including linoleic acid, and several amino acids (Table 1), suggesting a link between diet and inflammation in RA.
Epidemiological studies have shown a relationship between diet and RA; thus, some of the metabolomic changes observed in several fluids (serum/plasma or urine) in early arthritis could be related to differences in dietary patterns between RA patients and the healthy population. A study that included a large number of patients (15770 adult males and females) found that patients with arthritis (including RA and osteoarthritis) had lower quality diets compared to people without arthritis, based on HEI-2015, a healthy eating index created by the USDA and based on the Dietary Guidelines for Americans [9]. Patients with arthritis consumed less fruit, vegetables, greens and beans, whole grains, seafood, and plant protein, but more added sugars, saturated fats, and empty calories compared to those without arthritis [10]. The association of poor dietary quality with RA was also observed in other studies, in which RA patients had an inadequate intake of fruit, vegetables, dairy, fatty acids, and whole grains [11,12,13]. A study in a Chinese population found that RA patients were consuming different amounts of chicken, fish, mushrooms, beans, citrus, dairy products, and organ meats than healthy controls [14]. Another study that included a white population found that both women and men on a nonvegetarian diet were at higher risk of developing RA [15]. Hu et al. analyzed the cohort of women included in the Nurses’ Health Study and Nurses’s Health Study II that were followed from 1984 to the present-day, and found that good dietary quality, moderate alcohol consumption, and low intake of red meat were associated with a lower rate of RA incidence [13].
In the past, due to advances in the field of metabolomics, efforts have been made to predict food intake by measuring blood/urine/fecal metabolites. Two main techniques are being used: MS coupled with liquid- or gas-phase chromatography and proton (1H) NMR [16]. There are numerous metabolomics studies that have identified candidate biomarkers for different dietary patterns, as well as for different types of foods, ranging from meat to fruits and vegetables. Table 2 shows a summary of foods and the metabolites that have been found to be markers of their intake using metabolomics. Several studies have also found metabolites related to dietary patterns, like Mediterranean, high fat, or Western diets [17,18,19,20,21].

Table 2.
Food intake and candidate biomarkers identified by mass spectrometry (MS) and/or nuclear magnetic resonance (NMR).
The identification of food biomarkers is an ongoing process; a consensus has not been reached as to which metabolites would be the most adequate biomarkers for different types of foods. Moreover, some metabolites are markers of categories of food, not being able to discriminate between the exact types of foods being analyzed (1-methylhistidine and 3-methylhistidine are found in meat and are not useful in discriminating between types of meat). It is possible that for some foods, a combination of metabolites would be more suited as a marker than a single metabolite. Unfortunately, as of now, the metabolomic studies in RA (Table 1) have not collected food intake data nor used the same metabolomic platforms, making it difficult to associate specific food intake with metabolic changes in RA patients. However, some metabolites from Table 1 and Table 2 suggest an interaction between circulating metabolites and diet. For instance, some of these studies [5,59] showed higher levels of carnitine and taurine in RA patients, which are potential biomarkers of meat intake.
2.2. Drugs
Researchers have used a metabolomics approach to evaluate the changes in circulating metabolites from drugs used in RA treatment (Table 3). The study of these changes might help to understand RA pathogenesis, since the therapeutic effects of these drugs could potentially be driven by metabolic changes either by normalizing their abnormal values or by increasing anti-inflammatory metabolites. For instance, using a targeted metabolomic approach, Fu et al. compared the effect of oral glucocorticoids (GC) on serum polar lipids and observed an increase in lysophosphatidylcholines (LPC) and lysophosphatidylethanolamines (LPE) in females but not in male patients with RA [7]. GC inhibits phospholipase A, a key enzyme that hydrolyzes membrane phospholipids which is increased in inflammatory tissues. The effect of GC on phospholipase A will likely modify the phospholipid profile. Of interest, polyunsaturated acyl LPC and LPE presented an anti-inflammatory effect on animal models [60]. The effect of low dose GC (<10 mg/day) on arginine metabolism and cardiovascular risk in RA patients was also studied [61]. This study from Australia that included 36 RA patients, 18 of which were on GC (GC users) and 18 that were not receiving GC (non-GC-users), found that asymmetric dimethyl arginine (ADMA) and symmetric dimethyl arginine (SDMA) levels were lower in patients on chronic GC compared to non-GC users, suggesting that long-term treatment with GC had an improved endothelial function and a cardiovascular protective effect. by modulating arginine metabolism [61].

Table 3.
Metabolic profile modifications by drugs used in the treatment of RA.
Wang et al. [59] studied the change of the plasma metabolic profile in 29 RA patients after the initiation of treatment with methotrexate (14 patients) or a combination of methotrexate with a Chinese medicinal herb (15 patients). They found decreased levels of several amino acids (tryptophan, threonine, histidine, methionine, and glycine) as well as other metabolites (carnitine, hypoxanthine, cytosine, uracil, and uric acid), while taurine, aspartate, alanine, lactic acid, adenosine, and guanine were significantly increased in RA patients compared to controls. Interestingly, the treatment with methotrexate brought the levels of all these metabolites back to normal levels, suggesting a causative role of these amino acids in RA pathogenesis. The combination of MTX with tripterygium glycosides tablets was more effective in obtaining these results compared to monotherapy with MTX. Although more data is needed to link amino acid changes to abnormal immune response in RA, data in immune cells suggest a direct link between amino acid metabolism and T cell and macrophage responses by promoting and modulating inflammation, which could potentially be involved in RA pathogenesis [62,63,64,65]. In RA, tryptophan is the substrate of indoleamine-2,3-dioxygenase IDO2, which was demonstrated to be required for the activation of CD4+ Th cells, the production of pathogenic autoantibodies, and the subsequent development of arthritis in a KRN mouse model of arthritis [66,67,68]. This offers a possible explanation for the decrease in tryptophan levels that is then reversed by the addition of methotrexate. On the other hand, levels of S-adenosy-L-homocysteine, 5-formyltetrahydrofolate, and 5-methyltetrahydrofolate were similar between controls and RA patients before treatment, and decreased after 3 months of methotrexate, pointing to these methotrexate-associated metabolites as adherence biomarkers [59].
TNF is a potent pro-inflammatory cytokine that plays key role in cell metabolism, including glucose and lipid metabolism [72]; thus, changes in metabolic profile are expected after the administration of a TNF inhibitor. The first study that evaluated the changes in the metabolic profile of 16 RA and psoriatic arthritis (PsA) patients after TNFi treatment (etanercept and infliximab) used urine samples. The study described increases in hippuric acid, citrate, and lactic acid after infliximab treatment, while increases in choline, phenylacetic acid, urea, creatine, and methylamine were seen after etanercept treatment [71]. Another group evaluated the serum metabolomic profile in 20 patients with RA before and after treatment with TNFi (etanercept or adalimumab). Of the 20 patients, 55% of patients had a moderate EULAR response, while only 20% reached a good response. At 3 months posttreatment, 3-hydroxyisobutyrate, lysine, acetoacetate, acetylphosphocholine, creatine sn-glycero-3-phosphocholine, histidine, and phenylalanine levels decreased, while leucine, acetate, betaine, and formate levels increased, but they did not reach those of the healthy control [69]. The changes of the serum metabolic profile in response to treatment with a TNFi, etanercept, in 27 patients with active RA were also evaluated by Priori et al. These patients were receiving concomitant therapy with GC and disease-modifying antirheumatic drugs. After 3 months of treatment, isoleucine, leucine, valine, alanine, glutamine, tyrosine, and glucose levels were found to be increased in good responders as defined by EULAR-ESR criteria, whereas 3-hydroxybutyrate levels were reduced [70]. The decrease of 3-hydroxybutirate, acetoacetate, and acetylphosphocholine levels suggests a modulation of lipid metabolism after TNF inhibition, especially in responders. In addition, the increase of glucose and other amino acids suggests a decrease of glucose and amino acid metabolism by the inflamed tissues.
2.3. Comorbidities
RA patients present several comorbidities including obesity, metabolic syndrome, and sarcopenia, probably triggered by a disbalance of proinflammatory cytokines including TNF and IL-6 among other causes [73,74,75,76,77], that will modify the circulating metabolites [78]. Several studies have investigated circulating metabolic changes related to the metabolic syndrome and obesity [79,80]. Of interest, a lot of circulating metabolites that are different in RA patients compared to controls could be related to associated metabolic syndrome, since choline metabolism (especially TMAO and carnitine), aminoacids (alanine, glutamine, glutamate, arginine, aspartate, asparagine, histidine, methionine, cysteine, lysine, branched-chain amino acids (BCAA), phenylaniline, tyrosine, and tryptophan) and phospholipids (phosphatydilcholines) also change in those with metabolic syndrome [80]. Several works on muscle mass have also suggested that some circulating metabolites can be biomarkers of muscle mass and sarcopenia [81]. Even though both fat tissue and muscle, as well as associated immune cells in these inflamed tissues, can be sources of metabolites, it is unknown how much they can contribute to the pool of circulating metabolites. For example, studies measuring the metabolomics profile in visceral adipose tissue and serum from obese patients found low correlations between serum and adipose tissue metabolites [82]. On the other hand, we can speculate that there might be a competition between inflamed tissues (adipose tissue vs. synovial tissue) for the uptake of circulating anti-inflammatory metabolites.
2.4. Sex and Age
Several epidemiological studies have shown differences in metabolite concentrations according to sex and gender. A cross-sectional study in urine samples showed that some metabolites from the tricarboxylic acid cycle (TCA) cycle such as citrate and fumarate were elevated in women, while carnitine, acetylcarnitine, acetone, and creatinine were higher in men [83]. In addition, Fan et al. found that 2-hydroxyglutaric acid, α-ketoglutarate, and 2-oxyglutaric acid were higher in women. However, UDP-glucoronic acid was higher in men, suggesting that this could be linked to sex hormones [84]. Another study showed differences between sex and metabolic profile in serum, suggesting that glycine, serine, and sphingomyelines are upregulated in women, and ornithine, arginine, acyl carnitines, and amino acids derived from glutamine pathway are elevated in males [85]. Finally, a longitudinal cohort of adults showed a positive correlation of levels of glutamine, tyrosine, long chain fatty acids, acyl-carnitines, and sphingolipids, and a negative correlation of histidine, tryptophan, threonine, serine, and leucine levels with age [86].
2.5. Smoking and Exercise
Smoking is a known risk factor for RA and is associated with an increased risk of more severe arthritis, and less likelihood of achieving remission. Smoking also decreases the effectiveness of some disease-modifying antirheumatic drugs (DMARDs) [87,88]. The exact reason of these associations is not well understood, although the effect of smoking on immune cells, and cytokine production, and the increase of oxidative stress that it causes, have been described [89,90,91], and these likely affect the immune response in RA. Metabolomics has also identified blood biomarkers associated with chronic tobacco smoking. One study performed on a large number of healthy participants (892) from around the world found an association between smoking and three well-established nicotine metabolites (cotinine, hydroxycotinine, and cotinine N-oxide), and an additional 12 xenobiotic metabolites involved in benzoatic (e.g., 3-ethylphenylsulphate) or xanthine metabolism (e.g., 1-methylurate), three amino acids (o-cresol sulphate, serotonin, indolepropionate), two lipids (scyllo-inositol, pregnenolone sulphate), four vitamins or cofactors, and one carbohydrate (oxalate) [92]. Several of these metabolites, especially nicotine-derived metabolites, have been described to modulate the immune response [90], and other metabolic changes could be involved in smoking-induced methylation changes in immune cells [93]. Another study looked at the immediate effects of smoking on the metabolic profile. Thirty-one metabolites were shown to be acutely affected by cigarette smoking, including menthol-glucuronide, the reduction of glutamate, oleamide, and 13 glycerophospholipids. Moreover, detailed analysis revealed changes in 12 cancer-related metabolites, notably related with cAMP inhibition [94]. Since a known mechanism of methotrexate in treatment of RA is to induce an increase of cellular cAMP [95], the inhibition of this metabolite by smoking could explain the decrease in the effectiveness of this drug in RA.
Exercise is another factor that might change the metabolomic profile. However, these changes depend of the quantity and type of exercise. For example, in people who exercise more than 2 h per day, some metabolites, including medium and long fatty acids, ketones, sulfated bile acids, palmitate, linoleate, stearate, and palmitoleate, increased two-fold in their plasma concentration. Decreases of pyruvate and lactate, among others intermediates of TCA, have been reported after a short running period [96]. The reader can find an extensive review of these changes in a recently published review [96] about metabolic changes after exercising.
2.6. Genetics: Polymorphisms and Metabolism
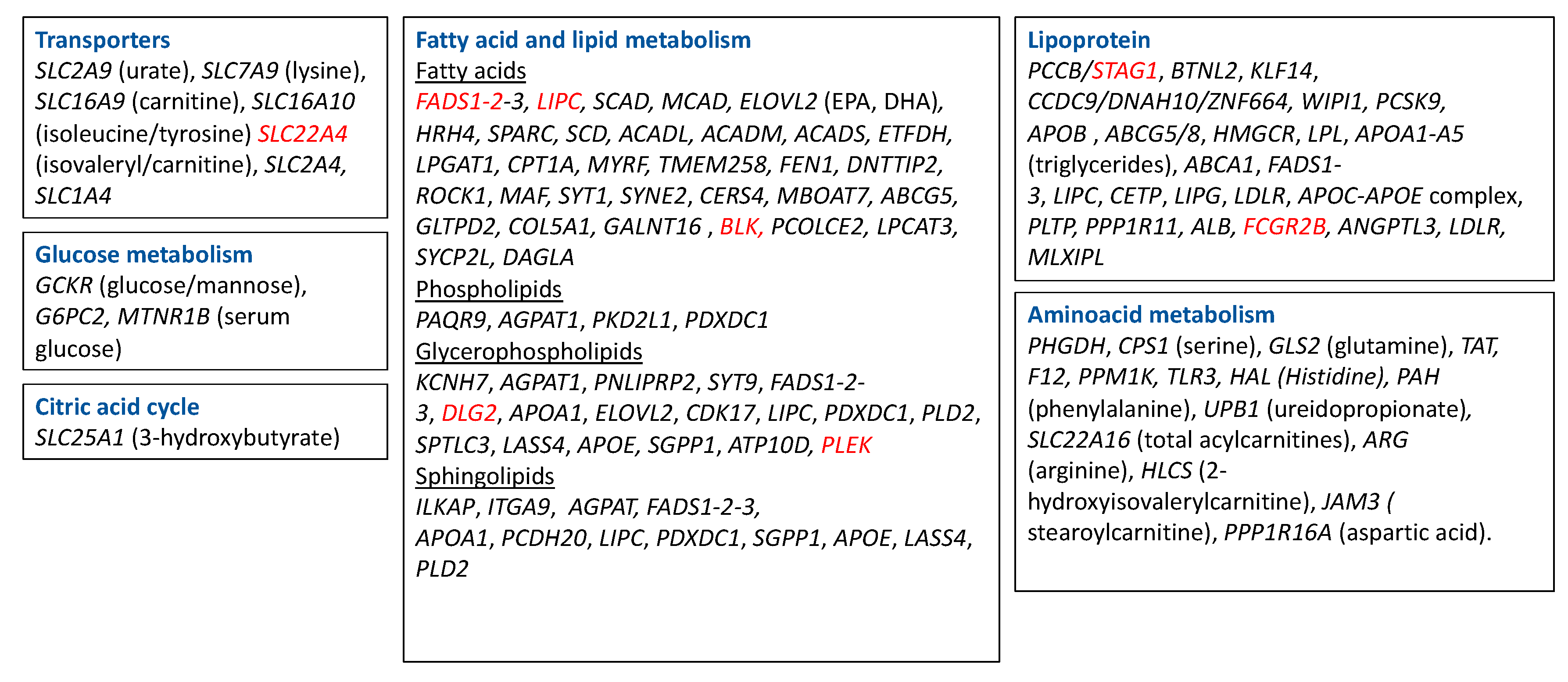
Genome-wide association studies (GWAS) uncovered multiple loci that are associated with the level of metabolites, which involve a large number of metabolic pathways, indicating widespread genetic influences on the human metabolome (Figure 3). The loci that have been described involve amino acids, intermediates of lipid metabolism, including sterols, carnitines, and intermediates of inositol and fatty acid metabolism, intermediates of purine and pyrimidine metabolism, glucose homeostasis, and vitamin and cofactor levels [97,98,99,100]. Polymorphisms in these metabolite-associated genes were also described in RA GWAS. In Figure 3, we put together a summary of metabolism-related genes described in genome-wide association studies (GWAS; https://www.ebi.ac.uk/gwas/). Polymorphisms in the genes underlined in red were found to be associated with RA. These genes are mostly related to lipid metabolism. Interestingly, lipid metabolites are considered pro-inflammatory metabolites (see Section 3), and higher levels of lipids were described in serum of RA patients compared to control subjects (Table 1). DLG2 (Discs Large MAGUK Scaffold Protein 2), which was found to be associated with glycerophospholipid metabolism [101], was also found to be related to response to TNF inhibitors in RA patients [102]. FADS1 and 2 (Fatty Acid Desaturase) and BLK (BLK Proto-Oncogene, Src Family Tyrosine Kinase), involved in fatty acid metabolism, and STAG1 (Stromal Antigen 1) and FCGR2B (Fc Fragment Of IgG Receptor IIb), involved in lipoprotein metabolism, were found to be associated with susceptibility to developing RA in several studies [103,104,105,106,107]. SLC22A4, a transporter related to isovaleryl/carnitine, was found to be associated with RA in a Japanese population [108], but not in a Canadian one [109]. Finally, Geiger et al. described 2 SNPs (single nucleotide polymorphism), rs9309413 and rs4775041, found on PLEK (Pleckstrin) and LIPC (Hepatic Triacylglycerol Lipase) genes, associated with sphingomyelin associated and phosphatidylethanolamine (PE) [110], that were associated with RA in a previous study [111]. Little is known about the role of these genes in inflammation and autoimmunity, so more studies are needed to determine whether some of these pathways are critical for the pathogenesis of RA.
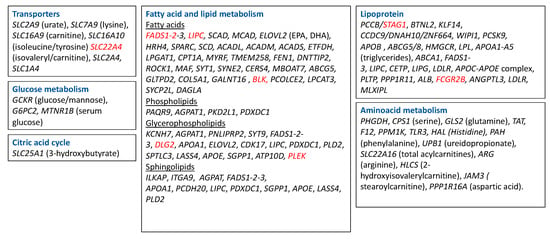
Figure 3.
Metabolism-related genes described in GWAS. Highlighted in red are the genes that were found to be associated with RA. SLC2A9—Solute Carrier Family 2 Member 9; SLC7A9—Solute Carrier Family 7 Member 9; SLC16A9—Solute Carrier Family 16 Member 9; SLC16A10—Solute Carrier Family 16 Member 10; SLC22A4—Solute Carrier Family 22 Member 4; SLC22A4—Solute Carrier Family 2 Member 4; SCL1A4—Solute Carrier Family 1 Member 4; SLC25A1—Solute Carrier Family 25 Member 1; FADS—fatty acid desaturase; LIPC—Hepatic Triacylglycerol Lipase; SCAD—Short-chain acyl-CoA dehydrogenase; MCAD—Medium-chain acyl-CoA dehydrogenase; ELOVL2—Fatty Acid Elongase 2; HRH 4—Histamine Receptor 4; SPARC—Secreted Protein Acidic And Cysteine Rich; SPTLC3—Serine Palmitoyltransferase Long Chain Base Subunit 3; LASS4—Ceramide Synthase 4, SGPP1—Sphingosine-1-phosphate, ATP10D—ATPase Phospholipid Transporting 10D (Putative), SCD—Stearoyl-CoA Desaturase; ACADL—Acyl-CoA Dehydrogenase Long Chain; ACADM—Acyl-CoA Dehydrogenase Medium Chain; ACADS—Acyl-CoA Dehydrogenase Short Chain; ETFDH—Electron Transfer Flavoprotein Dehydrogenase; LPGAT1—Lysophosphatidylglycerol Acyltransferase 1; CPT1—Carnitine Palmitoyltransferase 1; PHGDH—Phosphoglycerate Dehydrogenase; CPS1—Carbamoyl-Phosphate Synthase 1; GLS2—glutaminase; EPA—eicosapentaenoic acid; DHA—docosahexaenoic acid; GCKR—Glucokinase Regulator; APOA—Apolipoprotein; MYRF—Myelin Regulatory Factor; TMEM258—transmembrane protein 258; FEN1—flap structure-specific endonuclease 1; DNTTIP2—deoxynucleotidyl transferase terminal interacting protein 2; ROCK1—Rho Associated Coiled-Coil Containing Protein Kinase 1; MAF—MAF BZIP Transcription Facto; SYT1—Synaptotagmin 1; SYNE2—Spectrin Repeat Containing Nuclear Envelope Protein 2; CERS4—Ceramide Synthase 4; MBOAT7—Membrane Bound O-Acyltransferase Domain Containing 7; ABCG5—ATP Binding Cassette Subfamily G Member 5; GLTPD2—Glycolipid Transfer Protein Domain Containing 2; COL5A1—Collagen Type V Alpha 1 Chain; GALNT16—Polypeptide N-Acetylgalactosaminyltransferase 16; BLK—BLK Proto-Oncogene, Src Family Tyrosine Kinase; PCOLCE2—Procollagen C-Endopeptidase Enhancer 2; LPCAT3—Lysophosphatidylcholine Acyltransferase 3; SYCP2L—Synaptonemal Complex Protein 2 Like; DAGLA—Diacylglycerol Lipase Alpha; PAQR9—Progestin And AdipoQ Receptor Family Member 9; AGPAT1—1-Acylglycerol-3-Phosphate O-Acyltransferase 1; PKD2L1—Polycystin 2 Like 1, Transient Receptor Potential Cation Channel, PDXDC1—yridoxal Dependent Decarboxylase Domain Containing 1; KCNH7—Potassium Voltage-Gated Channel Subfamily H Member 7; PNLIPRP2—Pancreatic Lipase Related Protein 2 (Gene/Pseudogene); SYT9—Synaptotagmin 9; DLG2—Discs Large MAGUK Scaffold Protein 2; CDK17—Cyclin Dependent Kinase 17; PDXDC1—Pyridoxal Dependent Decarboxylase Domain Containing 1; PLD2—Phospholipase D2; APOE—Apolipoprotein E; ILKAP—ILK Associated Serine/Threonine Phosphatase; ITGA9—Integrin Subunit Alpha 9; PCDH20—Protocadherin 20; GCKR—Glucokinase Regulator; G6PC2—Glucose-6-Phosphatase Catalytic Subunit 2; MTNR1B—Melatonin Receptor 1B; PCCB—Propionyl-CoA Carboxylase Subunit Beta; STAG1—Stromal Antigen 1; BTNL2—Butyrophilin Like 2, KLF14—Kruppel Like Factor 14, CCDC9—Coiled-Coil Domain Containing 9; DNAH10—Dynein Axonemal Heavy Chain 10; ZNF664—Zinc Finger Protein 664; WIPI1-WD Repeat Domain, Phosphoinositide Interacting 1; PCSK9 -, Proprotein Convertase Subtilisin/Kexin Type 9; APOB—Apolipoprotein B; HMGCR—3-Hydroxy-3-Methylglutaryl-CoA Reductase; LPL—Lipoprotein Lipase; ABCA1—ATP Binding Cassette Subfamily A Member 1; CETP—Cholesteryl Ester Transfer Protein; LIPG—Lipase G, Endothelial Type; LDLR—Low Density Lipoprotein Receptor; PLTP—Phospholipid Transfer Protein; PPP1R11—Protein Phosphatase 1 Regulatory Inhibitor Subunit 11; ALB—Albumin; FCGR2B—Fc Fragment Of IgG Receptor IIb; ANGPTL3—Angiopoietin Like 3; MLXIPL—MLX Interacting Protein Like; GLS2—Glutaminase 2, TAT—Tyrosine Aminotransferase; F12—; Coagulation Factor XII; PPM1K—Protein Phosphatase, Mg2+/Mn2+ Dependent 1K, TLR3—Toll Like Receptor 3; HAL—Histidine Ammonia-Lyase; PAH—Phenylalanine Hydroxylase; UPB1—Beta-Ureidopropionase 1; SLC22A16—Solute Carrier Family 22 Member 16; ARG—Arginase; HLCS—Holocarboxylase Synthetase; JAM3—Junctional Adhesion Molecule 3; PPP1R16A—Protein Phosphatase 1 Regulatory Subunit 16A.
2.7. Gut Microbiome/Absorption
The gut microbiome represents the collection of microbes that inhabit the intestines. Its composition is shaped by several factors, like genetics, age, delivery pattern, diet, antibiotic use, and other treatments [112,113,114,115]. It can also be modulated by prebiotics [116,117], probiotics [118], and fecal microbiota transplantation. Bacteria in the gut are important not only in the absorption of certain vitamins and in the synthesis of bile acids, but they also have the potential to modify circulating pro- or anti-inflammatory mediators, since they are involved in the metabolism of some dietary components [119]. For example, trimethylamine-N-oxide, a pro-inflammatory metabolite that derives from choline and carnitine present in red meat, eggs, and dairy products, is produced by Prevotella copri among other bacteria [120,121]. An increased abundance of Prevotella copri was found in new-onset untreated RA patients, suggesting P. copri may be pathogenic in this disease [122]. In contrast, bacteria that have an almost exclusive saccharolytic metabolism, such as lactobacilli and bifidobacterial, are considered potentially beneficial [123], since they produce a variety of tryptophan catabolites (indole, tryptamine, indoleethanol (IE), indolepropionic acid (IPA), indolelactic acid (ILA), indoleacetic acid (IAA), skatole, indolealdehyde (IAld), and indoleacrylic acid (IA)) which are critical for intestinal homeostasis by decreasing intestinal permeability [124]. In addition, some of these catabolites enter the bloodstream and may have anti-inflammatory and anti-oxidative effects [124]. The microbial degradation of whole-grain complex carbohydrates increases short-chain fatty acids (SCFA; butyrate, acetate and propionate), which were also shown to be beneficial to the intestinal immune response [125]. Microbial bile acid metabolites have recently been linked to colonic homeostasis [126].
The modulation of the microbiome through diet interventions is a potential strategy in the treatment of diseases, since microbiome alterations are related to disease, i.e., inflammatory bowel disease, obesity, cardiovascular diseases, autoimmune diseases, and others. It seems that the microbiome response to diet is variable and is highly influenced by the subject’s baseline microbiome. Several studies found differences in the baseline microbiome of responders versus nonresponders to different diet interventions. Additionally, individuals with differing bacterial gene richness appear to have differing baseline gut microbiota communities that respond distinctively to a given dietary intervention which will influence the diversity of the gut microbe-derived specialized metabolites and circulating metabolites [127,128,129,130,131,132,133,134,135,136].
2.8. Metabolite Released from or Uptaken by Inflamed Tissues
Another potential factor that determines the concentrations of the circulating metabolites is represented by the release of metabolites from the inflamed joint or their uptake by the synovium. Little is known about metabolic or lipidomic profiling of synovial tissue [137,138]. In addition, no study has, to date, evaluated the relation between circulating metabolites in serum or plasma and synovial metabolites, although there might be a correlation. For instance, the synovial tissue of RA patients presents an enhanced level of lactate compared to noninflamed synovial tissue [138], which suggests an increase in the anaerobic cellular metabolism of resident cells [139,140]. Lactate has also been one of the metabolites described to be upregulated in patients with RA [5]. Of note, inflammatory pathways increase the expression of nutrient transporters [141,142,143,144]; therefore, this highly metabolic tissue will consume high amounts of metabolites, either to feed the increased metabolism of activated cells (pro-inflammatory metabolites) or to resolve inflammation (anti-inflammatory metabolites); this could be reflected by a decrease of circulating metabolites described in RA (Table 1 and Figure 1): such as glucose and amino acids (alanine, serine, methionine, threonine, leucine, valine, isoleucine, aspartate, phenylalanine, tyrosine, and proline) [6,145].
Fibroblast-like synoviocites (FLS), key cells in the pathogenesis and progression of RA, have an activated metabolism and can potentially release metabolites into the bloodstream [146]. Ahn et al. [147] characterized the intracellular metabolic profile of RA and osteoarthritis (OA) by an untargeted metabolomic approach using GC/TOF-MS. The results revealed that a high number of metabolites were increased in RA compared to OA FLS; these metabolites were amines (inosine, urate, 5′-deoxy-5′-methylthioadenosine, guanine, benzamide), fatty acids (behenic acid, palmitoleic acid, arachidic acid, oleic acid, myristic acid, stearic acid, palmitic acid, octadecanol, linoleic acid, lauric acid), phosphates (glucose-6-phosphate, phosphogluconic acid, adenosine-5-monophosphate, phosphate, fructose-6-phosphate), organic acids (aspartate, adipate, 2-ketoisocaproate 3-phenyllactate, 2-hydroxyvaleric acid, phenylacetate, glycolate, oxalate, benzoate), amino acids (asparagine, glutamine), sugars and sugar alcohols (lactose fucose, mannose) and salicylaldehyde. Other metabolites, mostly amino acids (isoleucine, leucine, histidine, valine, ornithine, lysine, methionine sulfoxide, tryptophan, N-methylalanine, tyrosine, phenylalanine, citrulline, oxoproline, threonine, serine) were decreased in RA compared to OA FLS. At the same time, the glycolysis and pentose phosphate pathways were more activated in RA than OA FLS. RA FLS are aggressive cells, similar to cancer cells, and require high amounts of energy to fulfill their pathogenetic functions in RA, which include proliferation, migration, and invasion [148,149].
Macrophages and T cells are the other dominant type of synovial cells in the inflamed joint, and are important in the progression of the disease, with their abundance being correlated with disease activity but also response to treatment [150]. Similar to the FLS, activated macrophages and T cells also rely on glycolysis and have alterations of the TCA cycle [150], which is consistent with the high levels of lactic acid, citrate, and succinate found in the synovial fluid of RA patients [151]. Although metabolic profiling of RA synovial macrophages and T cells hasn’t yet been undertaken, they are probably a source of circulating metabolites, while metabolites also exert their effect on synovial cells [152,153,154,155].
3. Evidence for a Pro-/Anti-Inflammatory Role of Metabolites in RA
RA is a chronic autoimmune disease, with a systemic immune response to autoantigens that may exist years before the onset of clinical symptoms, and a local immune activation of the synovial tissue which becomes inflamed, hyperplastic, and invasive of local cartilage and bone [156]. Whether or not pro- or anti-inflammatory metabolites play a role in RA pathogenesis is still unknown. However, the explosive growth of the field of tissue immunometabolism and its description of multiple critical metabolic pathways in the activation and differentiation of immune cells such as T and B lymphocytes, macrophages, dendritic cells, and fibroblasts, among others (see reviews in [157,158,159,160,161]), suggests that most of the metabolites involved in the immune response can also be important in RA. Here, we describe pro- and anti-inflammatory metabolites associated with RA pathogenesis (Figure 4).
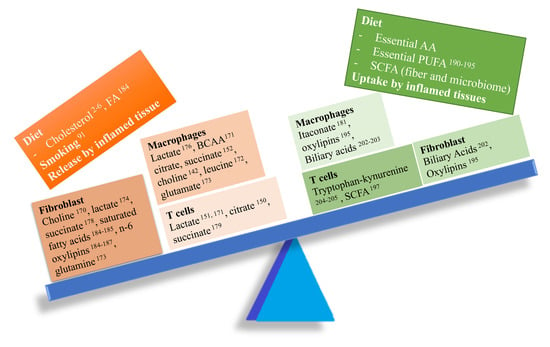
Figure 4.
Imbalance between pro- and anti-inflammatory metabolites in RA. Several pro-inflammatory metabolites (left side of the balance) might play a key role in RA pathogenesis modulating the function of several cell types involved in synovial inflammation.
3.1. Pro-Inflammatory Metabolites
Choline and Trimethylamine-N-oxide (TMAO). Metabolites related to the choline pathway were identified in several studies in synovial tissue, synovial fluid, and blood (serum/plasma) samples in both animal models and human studies. Diet is the main source of choline [162], whose metabolites (trimethylamine-N-oxide, TMAO) have already been related with cardiovascular inflammation [121,163]. Choline and other dietary trimethylamine (TMA) containing species like carnitine are metabolized to TMA by the gut microbiota. TMA is subsequently oxidized by at least one member of the flavin-containing monooxygenases, FMO3, forming trimethylamine-N oxide (TMAO), which is then released into circulation [164]. TMAO is a candidate biomarker for meat and fish intake, as can be seen in Table 2.
Despite being so well studied in relation to cardiovascular inflammation, we were not able to find studies evaluating the role of TMAO in RA. Our group found that serum TMAO was associated with measures of joint (tender joint count, swollen joint count, DAS28-CRP) and skin inflammation (body surface area affected by psoriasis) in a small cohort of patients with psoriasis and PsA [165]. The increased TMAO in patients with psoriasis and PsA, two diseases associated with metabolic syndrome [166], could be due either to an increased activity of FMO3, which has been described to be upregulated in obesity [167], but also to changes in the microbiome composition, which is an intermediate component of TMAO synthesis. TMAO, as well as choline, was found to be increased in serum samples in the murine K/BxN model of arthritis compared to control mice [168]. Choline is also a nutrient uptaken by the cells and metabolized via the Kennedy pathway, during which several phospholipids that function as signaling molecules are produced, such as glycerol-phosphocholine (GPC), phosphocholine, phosphatidylcholine (PC), lyso-PC, diacylglycerol, and lysophosphatidic acid [169]. Importantly, choline metabolism has been related to the RA FLS phenotype [170] and IL-1β secretion in macrophages [142].
BCAA (Branched-chain amino acids). Decreased levels of valine, leucine, and isoleucine were found in RA patients. Decreased levels of BCAA could be explained by low dietary consumption or by a higher intake of these amino acids by inflamed tissue. These are essential amino acids, so their source is the diet, and lately, they have been related to inflammation by inducing oxidative stress (via NADPH and Akt-mTOR signaling) and promoting the secretion of proinflammatory cytokines (IL-6, TNF) as well as the migration of peripheral blood mononuclear cells [171]. Branched chain aminotransferases1 (BCAT1), an enzyme that initiates BCAA metabolism, is the predominant isoform in human primary macrophages. Its action on leucine produces acetyl-CoA and glutamate, which enter the TCA cycle. Treatment of LPS and TNF stimulated human macrophages with ERG240, a leucine analogue that blocks BCAT1 activity, decreased oxygen consumption and glycolysis. Moreover, oral administration of ERG240 reduced the severity of collagen-induced arthritis in mice [172].
Glutamine is an amino acid used as a source to fuel metabolism. Glutaminase 1, the enzyme responsible of glutaminolysis, is upregulated in RA synovial fibroblasts [173], and inhibition of this enzyme decreased the aggressive phenotype of the FLS and improved the severity of arthritis in the SKG murine model of arthritis.
Glycolytic intermediates: The RA joint is characterized by a shift of the aerobic oxidative phosphorylation to a glycolytic state, in which less ATP is produced but at a faster rate, to be able to ensure the necessary energetic requirements for the highly active cells. Metabolites related to the glycolytic pathway have been detected in several studies on animal models, as well as human metabolomics studies.
Lactate is the end product of glycolysis, a metabolic pathway that is upregulated in activated FLS and macrophages. High concentrations of lactic acid are found in both blood and synovial fluid from inflamed joints in RA patients. Several studies have shown that lactate promotes the aggressive phenotype of FLS [174], the pro-inflammatory properties of macrophages [152,153,154], stimulates IL-17 secretion by CD4+ T cells and, at the same time, decrease CD4+T migration, which is related to the maintenance of a chronic inflammatory infiltrate [155,175]. Moreover, recently, a new lactate induced histone modification was described, lactylation, which correlates with the levels of lactate and is different from acetylation [176]. These findings require further study to evaluate their role in disease states, since altered epigenetic marks have been recently described in RA FLS [177].
Succinate is elevated in the synovial fluid of patients with RA [151]. The TCA metabolite promotes inflammation by stimulating IL-1β secretion in murine macrophages through HIF-1α [152]. Moreover, succinate activates NLRP3 inflammasome inducing IL-1β secretion by synovial fibroblasts in a rat model of RA [178]. It seems that succinate also plays a role in innate and adaptive immune responses. Researchers found that the genetic deficiency of Sucnr1, a succinate receptor expressed by immune cells, decreases trafficking of dendritic cells and reduces expansion of Th17 cells in the lymph nodes, reducing the symptoms of arthritis in the mouse antigen-induced arthritis model [179].
Itaconate, a macrophage activation marker, is thought to play an anti-inflammatory role, since it inhibits the succinate dehydrogenase-mediated oxidation of succinate, and through this, exerts anti-inflammatory effects in activated macrophages, as shown in an in vivo model of ischemia-reperfusion injury [180]. However, in an animal model of RA, higher levels of itaconate were found to be associated with high disease activity [181].
Cholesterol comes from the diet and its levels are increased in RA patients [2,6]; this was found to be predictive of RA in women, but not men [182]. Lipid metabolism is altered in RA, but cholesterol metabolism in RA has not been specifically studied. Interestingly, cholesterol was recently found to be high in OA chondrocytes, due to an increased uptake, upregulation of cholesterol hydroxylases, and increased production of oxysterol metabolites [183].
Free fatty acids (FFA) can be either taken from the diet (essential FA, alpha-linolenic acid -an omega-3 FA- and linoleic acid -an omega-6 FA-) or synthesized in the organism. it was suggested that they were proinflammatory, since they contribute to low-level inflammation in obese patients. FFA levels were higher in the serum of RA patients than in control subjects, and correlated with disease activity [184]. Frommer et al. showed that FFA contribute to the pathogenesis and damage in RA, OA, and PsA, since stimulation of FLS with oleic, palmitic, and linoleic acid induced the secretion of proinflammatory cytokine IL-6, the chemokines IL-8 and MCP-1, as well as the matrix metalloproteinases pro-MMP1 and MMP3 [185]. Arachidonic acid (ARA) is the precursor of classically described prostaglandins (PGE2), which are known to be involved in inflammation in general, but also in arthritis [186,187].
3.2. Anti-Inflammatory Metabolites
Polyunsaturated Fatty Acids (PUFA) Related Metabolites. Docosahexaenoic acid, DHA, and eicosapentaenoic acid, EPA have anti-inflammatory properties, mainly because they compete with ARA for the action of the enzymes (cyclooxygenase-COX, lypooxigenase-LOX, cytochrome P450) which results in a decreased production of ARA derived proinflammatory oxylipins and an increased production of DHA and EPA derived anti-inflammatory oxylipins (Figure 5) [188,189]. Several studies have described improved outcomes in RA patients after dietary intervention [188,190,191]. Decreased levels of EPA and DHA were described in Spanish RA patients, and were associated with higher disease duration, positivity for rheumatoid factor, erosive disease and with a worse response to TNF inhibitors [192]. Gene variants of the enzymes involved in the PUFA metabolism can determine the metabolic and clinical response to dietary intake of PUFA. For example, 5-lipoxygenase (ALOX5) gene variants were found to influence response to fish oil supplementation, changing the oxylipin profile and, consequently, having a different effect on cardiovascular risk [193]. Another study checked the association between genetic variants of ALOX5, ALOX12, ALOX12B, and ALOX15, and type-2 diabetes mellitus (T2D), and found that ALOX12 and ALOX12B genetic variants increased susceptibility to T2D development, possibly though alterations in PUFA/ARA metabolism.
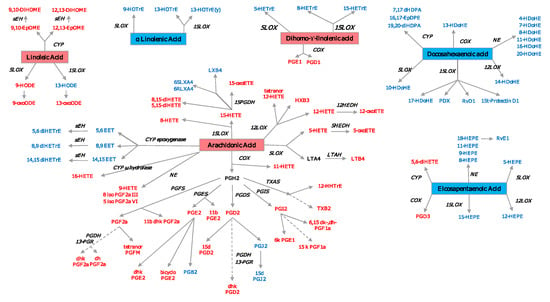
Figure 5.
Oxylipin derived from PUFA. Pro-inflammatory oxylipins are marked in red, while anti-inflammatory ones are marked in blue. The precursor n3-PUFAs are marked in a red square, while the n6-PUFAs are marked in a blue square. COX—cyclooxygenase; LOX—lypooxigenase; CYP—cytochrome P450; NE—nonenzymatic; PGFS—prostaglandin F synthase; PGES—prostaglandin E synthase; PGDS—prostaglandin D synthase; PGIS—prostaglandin I synthase; TXAS—thromboxane A2 synthase; LTAH—leukotriene A4 hydrolase; MDB—membrane dipeptidase; HEDH—hydroxyeicosanoid dehydrogenase; PGDH—hydroxyprostaglandin dehydrogenase; 13-PGR—15-ketoprostaglandin∆13 reductase; sEH—soluble epoxide hydrolase. A list with all the oxylipins can be found in Supplementary Table S1.
Oxylipin Related Pathways. Prostaglandins, thromboxanes and leukotrienes are the classically described oxylipins involved in the pathogenesis of RA. The newer methods of LC/MS and NMR make it possible to identify several other oxylipins, e.g., 8-HETE, 12-HETE, and 12-HEPE are products of the 12-lypoxygenase pathway. Liagre et al. demonstrated the presence of 12-LOX in type B synoviocytes and found that IL-1β and TNF stimulation increased 12-HETE production, while IL-6 and IL-4 did not have the same effect [194]. This pathway was also studied by Kronke et al., who showed that the deletion of 12/15-LOX in two models of arthritis (the K/BxN serum-transfer and a TNF transgenic mouse model) led to uncontrolled inflammation and tissue damage [195]. LTB4 and 5-HETE are products of ARA via 5-LOX pathway; 5- and 15-LOX are expressed in both OA and RA synovium in the lining and sublining macrophages, neutrophils, and mast cells, and have been shown to be involved in RA pathogenesis, promoting inflammation [196].
Short Chain Fatty Acids (SCFA) are byproducts of the metabolism of dietary fiber by the gut microbiome. They modulate immune and inflammatory responses via the activation of free fatty acid (FFA) receptors type 2 and 3 (FFA2 and FFA3 receptors) and G protein-coupled receptor 109A (GPR109A), and via inhibition of histone deacetylases (HDACs). A metabolomic study performed on a CIA rat model found decreased levels of acetate, propionate, butyrate, and valerate in fecal samples of arthritic rats compared to controls [197]. The administration of butyrate inhibited collagen-induced arthritis via Treg/IL10/Th17 axis [198].
Bile acids (BA) seem to have anti-inflammatory properties. Primary BAs are synthesized in the liver and are liberated in the gastrointestinal tract to help with lipid digestion. Gut bacteria metabolize primary bile acids and can deconjugate them, synthesizing secondary BAs. BAs have been detected in the systemic circulation, where their concentrations vary with diet, and have been related to insulin resistance [199]. High concentrations of BA can actually kill intestinal bacteria to prevent colonization, but they also regulate the mucosal immune functions through several receptors. A recent study showed the role of BA in the maintenance of the homeostasis of the mucosal immune function in the gut, through the vitamin D receptor [126]. An in vitro study found that taurolithocholic acid suppressed the expression of genes involved in mediating pro-inflammatory effects, phagocytosis, interactions with pathogens and autophagy, as well as the recruitment of immune cells, such as NK cells, neutrophils and T cells [200]. BA exert their actions through both specific and nonspecific receptors. The activation of TGR5 receptor by endogenous BA suppressed the production of LPS induced inflammatory cytokines in macrophages, while no effect was seen in macrophages that lacked this receptor [201,202]. Of interest, a study described an anti-inflammatory role of taurochenodeoxycholic acid in RA FLS [203]. Most studies focused on the effects of the BA on the gut mucosal immunity, and hence, future studies are needed to elucidate the roles of circulating BAs in disease states.
Tryptophan metabolism. Tryptophan is an essential amino acid that must be provided in the diet. It has been described that microbes-derived tryptophan metabolites can exert systemic and anti-inflammatory effects [124]. Moreover, tryptophan and its catabolic metabolites generated through the kynurenine pathway are involved in inflammation. Kynurenine has known anti-inflammatory effects that are toxic to T cells and induce cell death by apoptosis. Kynurenine is formed from tryptophan by the activity of indoleamine 2, 3-dioxygenase (IDO). The activation of IDO is actively involved in the resolution of arthritis in mice associated with an increase in kynurenine metabolites [204]. Kynurenine itself has been identified as a ligand for the aryl hydrocarbon receptor, which is important in the maturation of immune cells, and its addition promotes the differentiation of regulatory T cells and suppresses the differentiation of pathogenic Th17 cells [205].
4. Studies of Beneficial Effect of Diet in RA
In spite of the growing evidence of the relationship between diet and RA symptoms, research in the field is still limited to mostly observational studies; however, there are quite a few interventional studies in which diet has been evaluated as a strategy to improve RA symptoms. A detailed review of the studies can be found here [206,207,208]. Most interventions combine a diet with high intake of vegetables, fruit, and antioxidants with periods of fasting. The outcomes used in the majority of the studies include tender and swollen joint, disability index score, general health assessment scores, and a few inflammatory markers that can be quantified in blood, i.e., C reactive protein, erythrocyte sedimentation rate and proinflammatory cytokine (IL-6, TNF alpha).
These dietary intervention studies don’t shed any light on the specific metabolites that are responsible for the effect, or their mechanism of action. One study evaluated the anti-inflammatory effects of a low ARA diet and fish oil in patients with RA. Besides the usual outcomes, this study also quantified fatty acids and eicosanoids, using radioimmunoassay and gas chromatography coupled with MS [209]. They found that the diet improved RA clinical signs. In terms of fatty acid changes, they observed an enrichment of eicosapentaenoic acid in erythrocyte lipids and lower formation of urinary leukotriene B(4), 11-dehydro-thromboxane B(2) and prostaglandin metabolites in patients receiving the fish oil diet, especially when fish oil was given for a longer period of time (up to 8 months).
The concentrations of plasma phospholipid related fatty acids were evaluated after a vegan and lacto-vegetarian diet intervention in RA patients. It was found that 20:3n-6 and 20:4n-6 were significantly reduced after 3.5 months of vegan diet, but the concentration increased to baseline values with a lactovegetarian diet. Also, 20:5n-3 was significantly reduced after both vegan and lactovegetarian diet periods. However, no significant difference in fatty acid concentrations was detected between diet responders and diet nonresponders after both diet periods, which suggests that the changes in the fatty acid concentrations were not in response to diet [210].
Another study in an RA Swedish population found that EPA and DHA were both increased in erythrocytes after a blue mussel diet compared to a control diet, resulting in a decrease of n-6 PUFA ARA (20:4 n-6) and dihomo-gamma-linolenic acid (DGLA; 20:3 n−6), as well as a small decrease in saturated fatty acids and the monounsaturated fatty acid palmitoleic acid (16:1 n-7). Baseline EPA and DHA levels in this population were higher than in healthy men and women, but this group had already reported that the RA population from the studied area had a higher fish and shellfish intake compared to the general population of Sweden [211]. In contrast to these findings, erythrocyte levels of α-linolenic acid (ALA; 18:3n3), EPA 20:5n3, and the omega-3 index (EPA plus DHA) were found to be significantly lower in RA patients compared to healthy controls in a Korean population, although diet was not accounted for in this study. In addition, EPA and ALA were negatively associated with the risk of RA in Korean women [212].
Additionally, the study of other autoimmune diseases does not help in establishing a link between diet and metabolites with their pro or anti-inflammatory effect. In Crohn’s disease, for instance, animal studies suggest the potential beneficial effect of short chain fatty acids, tryptophan, arginine, and glutamine due to their roles in the modulation of the immune system, but no clinical studies have been performed to date [213]. A cross-sectional metabolomics study also found decreased levels of essential PUFA in patients with lupus, but diet was not taken into account in this study [214]. Further studies are needed before we can make conclusions about the role of diet in the levels of circulating and local metabolites and their relationship with clinical outcomes in RA and other autoimmune diseases.
5. Conclusions
Metabolomics studies have clearly shown that there is an alteration of the metabolic profile in patients with RA which may be related to the pathogenesis of the disease, but also to exposure to external factors, since the levels of the metabolites are influenced by several factors, including genetic ones, diet, sex, drugs, comorbidities, and microbiome (Figure 1). The possibility of altering some of these factors represents an attractive approach for future therapeutically interventions. Diet is a modifiable factor, and studies have shown that it can be effective in improving RA symptoms. Understanding the complex relation between diet, metabolites, microbiome, and disease status is still an ongoing process, but existing studies are promising. The field of RA needs more studies, including mendelian randomization studies and randomized clinical trials, combining the use of metabolomics, transcriptomics, and the microbiome to help understand how these elements interact with each other, identify patients who would benefit from a dietary intervention, and design the correct intervention.
Supplementary Materials
The following are available online at https://www.mdpi.com/2073-4409/9/4/827/s1. Table S1. Oxylipin abbreviations and names.
Author Contributions
All authors have reviewed the literature and contributed to writing the manuscript. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the National Institutes of Health (R01AR073324 to M.G., T32AR064194 to J.D.M.-S. and Krupp Endowed Foundation to R.C. and M.G.
Conflicts of Interest
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
References
- Hochberg, M.C.; Spector, T.D. Epidemiology of rheumatoid arthritis: Update. Epidemiol. Rev. 1990, 12, 247–252. [Google Scholar] [CrossRef] [PubMed]
- Lauridsen, M.B.; Bliddal, H.; Christensen, R.; Danneskiold-Samsøe, B.; Bennett, R.; Keun, H.; Lindon, J.C.; Nicholson, J.K.; Dorff, M.H.; Jaroszewski, J.W.; et al. 1H NMR spectroscopy-based interventional metabolic phenotyping: A cohort study of rheumatoid arthritis patients. J. Proteome Res. 2010, 9, 4545–4553. [Google Scholar] [CrossRef] [PubMed]
- Surowiec, I.; Gjesdal, C.G.; Jonsson, G.; Norheim, K.B.; Lundstedt, T.; Trygg, J.; Omdal, R. Metabolomics study of fatigue in patients with rheumatoid arthritis naïve to biological treatment. Rheumatol. Int. 2016, 36, 703–711. [Google Scholar] [CrossRef]
- Yousri, N.A.; Bayoumy, K.; Elhaq, W.G.; Mohney, R.P.; Emadi, S.A.; Hammoudeh, M.; Halabi, H.; Masri, B.; Badsha, H.; Uthman, I.; et al. Large Scale Metabolic Profiling identifies Novel Steroids linked to Rheumatoid Arthritis. Sci. Rep. 2017, 7, 9137. [Google Scholar] [CrossRef]
- Young, S.P.; Kapoor, S.R.; Viant, M.R.; Byrne, J.J.; Filer, A.; Buckley, C.D.; Kitas, G.D.; Raza, K. The impact of inflammation on metabolomic profiles in patients with arthritis. Arthritis Rheum. 2013, 65, 2015–2023. [Google Scholar] [CrossRef]
- Zhou, J.; Chen, J.; Hu, C.; Xie, Z.; Li, H.; Wei, S.; Wang, D.; Wen, C.; Xu, G. Exploration of the serum metabolite signature in patients with rheumatoid arthritis using gas chromatography-mass spectrometry. J. Pharm. Biomed. Anal. 2016, 127, 60–67. [Google Scholar] [CrossRef]
- Fu, J.; Cuppen, B.V.; Welsing, P.M.; van Wietmarschen, H.; Harms, A.C.; Berger, R.; Koval, S.; Fritsch-Stork, R.D.; Bijlsma, J.W.; Hankemeier, T.; et al. Differences between serum polar lipid profiles of male and female rheumatoid arthritis patients in response to glucocorticoid treatment. Inflammopharmacology 2016, 24, 397–402. [Google Scholar] [CrossRef]
- Li, J.; Che, N.; Xu, L.; Zhang, Q.; Wang, Q.; Tan, W.; Zhang, M. LC-MS-based serum metabolomics reveals a distinctive signature in patients with rheumatoid arthritis. Clin. Rheumatol. 2018, 37, 1493–1502. [Google Scholar] [CrossRef]
- Krebs-Smith, S.M.; Pannucci, T.E.; Subar, A.F.; Kirkpatrick, S.I.; Lerman, J.L.; Tooze, J.A.; Wilson, M.M.; Reedy, J. Update of the Healthy Eating Index: HEI-2015. J. Acad. Nutr. Diet 2018, 118, 1591–1602. [Google Scholar] [CrossRef]
- Comee, L.; Taylor, C.A.; Nahikian-Nelms, M.; Ganesan, L.P.; Krok-Schoen, J.L. Dietary patterns and nutrient intake of individuals with rheumatoid arthritis and osteoarthritis in the United States. Nutrition 2019, 67-68, 110533. [Google Scholar] [CrossRef]
- Berube, L.T.; Kiely, M.; Yazici, Y.; Woolf, K. Diet quality of individuals with rheumatoid arthritis using the Healthy Eating Index (HEI)-2010. Nutr. Health 2017, 23, 17–24. [Google Scholar] [CrossRef] [PubMed]
- Barebring, L.; Winkvist, A.; Gjertsson, I.; Lindqvist, H.M. Poor Dietary Quality Is Associated with Increased Inflammation in Swedish Patients with Rheumatoid Arthritis. Nutrients 2018, 10, 535. [Google Scholar] [CrossRef]
- Hu, Y.; Sparks, J.A.; Malspeis, S.; Costenbader, K.H.; Hu, F.B.; Karlson, E.W.; Lu, B. Long-term dietary quality and risk of developing rheumatoid arthritis in women. Ann. Rheum. Dis. 2017, 76, 1357–1364. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Wang, Y.; Feng, M.; Zhang, X.; Jin, Y.B.; Li, X.; Su, L.C.; Liu, S.; Wang, A.X.; Chen, X.M.; et al. Dietary intake and risk of rheumatoid arthritis—A cross section multicenter study. Clin. Rheumatol. 2016, 35, 2901–2908. [Google Scholar] [CrossRef]
- Fraser, G.E. Associations between diet and cancer, ischemic heart disease, and all-cause mortality in non-Hispanic white California Seventh-day Adventists. Am. J. Clin. Nutr. 1999, 70, 532S–538S. [Google Scholar] [CrossRef]
- Wishart, D. Metabolomics: Applications to food science and nutrition research. Trends Food Sci. Tech. 2008, 19, 482–493. [Google Scholar] [CrossRef]
- Lindqvist, H.M.; Rådjursöga, M.; Malmodin, D.; Winkvist, A.; Ellegård, L. Serum metabolite profiles of habitual diet: Evaluation by 1H-nuclear magnetic resonance analysis. Am. J. Clin. Nutr. 2019, 110, 53–62. [Google Scholar] [CrossRef]
- Rådjursöga, M.; Lindqvist, H.M.; Pedersen, A.; Karlsson, B.G.; Malmodin, D.; Ellegård, L.; Winkvist, A. Nutritional Metabolomics: Postprandial Response of Meals Relating to Vegan, Lacto-Ovo Vegetarian, and Omnivore Diets. Nutrients 2018, 10, 63. [Google Scholar] [CrossRef]
- Stella, C.; Beckwith-Hall, B.; Cloarec, O.; Holmes, E.; Lindon, J.C.; Powell, J.; van der Ouderaa, F.; Bingham, S.; Cross, A.J.; Nicholson, J.K. Susceptibility of human metabolic phenotypes to dietary modulation. J. Proteome Res. 2006, 5, 2780–2788. [Google Scholar] [CrossRef]
- Macias, S.; Kirma, J.; Yilmaz, A.; Moore, S.E.; McKinley, M.C.; McKeown, P.P.; Woodside, J.V.; Graham, S.F.; Green, B.D. Application of. Metabolites 2019, 9. [Google Scholar]
- Wellington, N.; Shanmuganathan, M.; de Souza, R.J.; Zulyniak, M.A.; Azab, S.; Bloomfield, J.; Mell, A.; Ly, R.; Desai, D.; Anand, S.S.; et al. Metabolic Trajectories Following Contrasting Prudent and Western Diets from Food Provisions: Identifying Robust Biomarkers of Short-Term Changes in Habitual Diet. Nutrients 2019, 11, 407. [Google Scholar] [CrossRef] [PubMed]
- Myint, T.; Fraser, G.E.; Lindsted, K.D.; Knutsen, S.F.; Hubbard, R.W.; Bennett, H.W. Urinary 1-methylhistidine is a marker of meat consumption in Black and in White California Seventh-day Adventists. Am. J. Epidemiol. 2000, 152, 752–755. [Google Scholar] [CrossRef]
- Bertram, H.C.; Hoppe, C.; Petersen, B.O.; Duus, J.; Mølgaard, C.; Michaelsen, K.F. An NMR-based metabonomic investigation on effects of milk and meat protein diets given to 8-year-old boys. Br. J. Nutr. 2007, 97, 758–763. [Google Scholar] [CrossRef] [PubMed]
- Yin, X.; Gibbons, H.; Rundle, M.; Frost, G.; McNulty, B.A.; Nugent, A.P.; Walton, J.; Flynn, A.; Gibney, M.J.; Brennan, L. Estimation of Chicken Intake by Adults Using Metabolomics-Derived Markers. J. Nutr. 2017. [Google Scholar] [CrossRef] [PubMed]
- Cheung, W.; Keski-Rahkonen, P.; Assi, N.; Ferrari, P.; Freisling, H.; Rinaldi, S.; Slimani, N.; Zamora-Ros, R.; Rundle, M.; Frost, G.; et al. A metabolomic study of biomarkers of meat and fish intake. Am. J. Clin. Nutr. 2017, 105, 600–608. [Google Scholar] [CrossRef] [PubMed]
- Ross, A.B.; Svelander, C.; Undeland, I.; Pinto, R.; Sandberg, A.S. Herring and Beef Meals Lead to Differences in Plasma 2-Aminoadipic Acid, β-Alanine, 4-Hydroxyproline, Cetoleic Acid, and Docosahexaenoic Acid Concentrations in Overweight Men. J. Nutr. 2015, 145, 2456–2463. [Google Scholar] [CrossRef]
- Lenz, E.M.; Bright, J.; Wilson, I.D.; Hughes, A.; Morrisson, J.; Lindberg, H.; Lockton, A. Metabonomics, dietary influences and cultural differences: A 1H NMR-based study of urine samples obtained from healthy British and Swedish subjects. J. Pharm. Biomed. Anal. 2004, 36, 841–849. [Google Scholar] [CrossRef]
- Lloyd, A.J.; Favé, G.; Beckmann, M.; Lin, W.; Tailliart, K.; Xie, L.; Mathers, J.C.; Draper, J. Use of mass spectrometry fingerprinting to identify urinary metabolites after consumption of specific foods. Am. J. Clin. Nutr. 2011, 94, 981–991. [Google Scholar] [CrossRef]
- Andersen, M.B.; Kristensen, M.; Manach, C.; Pujos-Guillot, E.; Poulsen, S.K.; Larsen, T.M.; Astrup, A.; Dragsted, L. Discovery and validation of urinary exposure markers for different plant foods by untargeted metabolomics. Anal. Bioanal. Chem. 2014, 406, 1829–1844. [Google Scholar] [CrossRef]
- Guertin, K.A.; Moore, S.C.; Sampson, J.N.; Huang, W.Y.; Xiao, Q.; Stolzenberg-Solomon, R.Z.; Sinha, R.; Cross, A.J. Metabolomics in nutritional epidemiology: Identifying metabolites associated with diet and quantifying their potential to uncover diet-disease relations in populations. Am. J. Clin. Nutr. 2014, 100, 208–217. [Google Scholar] [CrossRef]
- Hanhineva, K.; Lankinen, M.A.; Pedret, A.; Schwab, U.; Kolehmainen, M.; Paananen, J.; de Mello, V.; Sola, R.; Lehtonen, M.; Poutanen, K.; et al. Nontargeted metabolite profiling discriminates diet-specific biomarkers for consumption of whole grains, fatty fish, and bilberries in a randomized controlled trial. J. Nutr. 2015, 145, 7–17. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Yang, S.; Cai, S.; Dong, J.; Li, X.; Chen, Z. Identification of biochemical changes in lactovegetarian urine using 1H NMR spectroscopy and pattern recognition. Anal. Bioanal. Chem. 2010, 396, 1451–1463. [Google Scholar] [CrossRef] [PubMed]
- O’Sullivan, A.; Gibney, M.J.; Brennan, L. Dietary intake patterns are reflected in metabolomic profiles: Potential role in dietary assessment studies. Am. J. Clin. Nutr. 2011, 93, 314–321. [Google Scholar] [CrossRef] [PubMed]
- Posma, J.M.; Garcia-Perez, I.; Heaton, J.C.; Burdisso, P.; Mathers, J.C.; Draper, J.; Lewis, M.; Lindon, J.C.; Frost, G.; Holmes, E.; et al. Integrated Analytical and Statistical Two-Dimensional Spectroscopy Strategy for Metabolite Identification: Application to Dietary Biomarkers. Anal. Chem. 2017, 89, 3300–3309. [Google Scholar] [CrossRef]
- Mennen, L.I.; Sapinho, D.; Ito, H.; Bertrais, S.; Galan, P.; Hercberg, S.; Scalbert, A. Urinary flavonoids and phenolic acids as biomarkers of intake for polyphenol-rich foods. Br. J. Nutr. 2006, 96, 191–198. [Google Scholar] [CrossRef]
- Edmands, W.M.; Ferrari, P.; Rothwell, J.A.; Rinaldi, S.; Slimani, N.; Barupal, D.K.; Biessy, C.; Jenab, M.; Clavel-Chapelon, F.; Fagherazzi, G.; et al. Polyphenol metabolome in human urine and its association with intake of polyphenol-rich foods across European countries. Am. J. Clin. Nutr. 2015, 102, 905–913. [Google Scholar] [CrossRef]
- May, D.H.; Navarro, S.L.; Ruczinski, I.; Hogan, J.; Ogata, Y.; Schwarz, Y.; Levy, L.; Holzman, T.; McIntosh, M.W.; Lampe, J.W. Metabolomic profiling of urine: Response to a randomised, controlled feeding study of select fruits and vegetables, and application to an observational study. Br. J. Nutr. 2013, 110, 1760–1770. [Google Scholar] [CrossRef]
- Pujos-Guillot, E.; Hubert, J.; Martin, J.F.; Lyan, B.; Quintana, M.; Claude, S.; Chabanas, B.; Rothwell, J.A.; Bennetau-Pelissero, C.; Scalbert, A.; et al. Mass spectrometry-based metabolomics for the discovery of biomarkers of fruit and vegetable intake: Citrus fruit as a case study. J. Proteome Res. 2013, 12, 1645–1659. [Google Scholar] [CrossRef]
- Heinzmann, S.S.; Brown, I.J.; Chan, Q.; Bictash, M.; Dumas, M.E.; Kochhar, S.; Stamler, J.; Holmes, E.; Elliott, P.; Nicholson, J.K. Metabolic profiling strategy for discovery of nutritional biomarkers: Proline betaine as a marker of citrus consumption. Am. J. Clin. Nutr. 2010, 92, 436–443. [Google Scholar] [CrossRef]
- Lloyd, A.J.; Beckmann, M.; Favé, G.; Mathers, J.C.; Draper, J. Proline betaine and its biotransformation products in fasting urine samples are potential biomarkers of habitual citrus fruit consumption. Br. J. Nutr. 2011, 106, 812–824. [Google Scholar] [CrossRef]
- Rangel-Huerta, O.D.; Aguilera, C.M.; Perez-de-la-Cruz, A.; Vallejo, F.; Tomas-Barberan, F.; Gil, A.; Mesa, M.D. A serum metabolomics-driven approach predicts orange juice consumption and its impact on oxidative stress and inflammation in subjects from the BIONAOS study. Mol. Nutr. Food Res. 2017, 61. [Google Scholar] [CrossRef]
- Beckmann, M.; Lloyd, A.J.; Haldar, S.; Seal, C.; Brandt, K.; Draper, J. Hydroxylated phenylacetamides derived from bioactive benzoxazinoids are bioavailable in humans after habitual consumption of whole grain sourdough rye bread. Mol. Nutr. Food Res. 2013, 57, 1859–1873. [Google Scholar] [CrossRef] [PubMed]
- Hanhineva, K.; Keski-Rahkonen, P.; Lappi, J.; Katina, K.; Pekkinen, J.; Savolainen, O.; Timonen, O.; Paananen, J.; Mykkänen, H.; Poutanen, K. The postprandial plasma rye fingerprint includes benzoxazinoid-derived phenylacetamide sulfates. J. Nutr. 2014, 144, 1016–1022. [Google Scholar] [CrossRef] [PubMed]
- Trimigno, A.; Münger, L.; Picone, G.; Freiburghaus, C.; Pimentel, G.; Vionnet, N.; Pralong, F.; Capozzi, F.; Badertscher, R.; Vergères, G. GC-MS Based Metabolomics and NMR Spectroscopy Investigation of Food Intake Biomarkers for Milk and Cheese in Serum of Healthy Humans. Metabolites 2018, 8, 26. [Google Scholar] [CrossRef] [PubMed]
- Münger, L.H.; Trimigno, A.; Picone, G.; Freiburghaus, C.; Pimentel, G.; Burton, K.J.; Pralong, F.P.; Vionnet, N.; Capozzi, F.; Badertscher, R.; et al. Identification of Urinary Food Intake Biomarkers for Milk, Cheese, and Soy-Based Drink by Untargeted GC-MS and NMR in Healthy Humans. J. Proteome Res. 2017, 16, 3321–3335. [Google Scholar] [CrossRef]
- Pallister, T.; Haller, T.; Thorand, B.; Altmaier, E.; Cassidy, A.; Martin, T.; Jennings, A.; Mohney, R.P.; Gieger, C.; MacGregor, A.; et al. Metabolites of milk intake: A metabolomic approach in UK twins with findings replicated in two European cohorts. Eur. J. Nutr. 2017, 56, 2379–2391. [Google Scholar] [CrossRef]
- Stalmach, A.; Mullen, W.; Barron, D.; Uchida, K.; Yokota, T.; Cavin, C.; Steiling, H.; Williamson, G.; Crozier, A. Metabolite profiling of hydroxycinnamate derivatives in plasma and urine after the ingestion of coffee by humans: Identification of biomarkers of coffee consumption. Drug. Metab. Dispos. 2009, 37, 1749–1758. [Google Scholar] [CrossRef]
- Rothwell, J.A.; Fillâtre, Y.; Martin, J.F.; Lyan, B.; Pujos-Guillot, E.; Fezeu, L.; Hercberg, S.; Comte, B.; Galan, P.; Touvier, M.; et al. New biomarkers of coffee consumption identified by the non-targeted metabolomic profiling of cohort study subjects. PLoS ONE 2014, 9, e93474. [Google Scholar] [CrossRef]
- Cornelis, M.C.; Erlund, I.; Michelotti, G.A.; Herder, C.; Westerhuis, J.A.; Tuomilehto, J. Metabolomic response to coffee consumption: Application to a three-stage clinical trial. J. Intern. Med. 2018, 283, 544–557. [Google Scholar] [CrossRef]
- Daykin, C.A.; Van Duynhoven, J.P.; Groenewegen, A.; Dachtler, M.; Van Amelsvoort, J.M.; Mulder, T.P. Nuclear magnetic resonance spectroscopic based studies of the metabolism of black tea polyphenols in humans. J. Agric. Food Chem. 2005, 53, 1428–1434. [Google Scholar] [CrossRef] [PubMed]
- Mulder, T.P.; Rietveld, A.G.; van Amelsvoort, J.M. Consumption of both black tea and green tea results in an increase in the excretion of hippuric acid into urine. Am. J. Clin. Nutr. 2005, 81, 256S–260S. [Google Scholar] [CrossRef]
- Van der Hooft, J.J.; de Vos, R.C.; Mihaleva, V.; Bino, R.J.; Ridder, L.; de Roo, N.; Jacobs, D.M.; van Duynhoven, J.P.; Vervoort, J. Structural elucidation and quantification of phenolic conjugates present in human urine after tea intake. Anal. Chem. 2012, 84, 7263–7271. [Google Scholar] [CrossRef] [PubMed]
- Van Dorsten, F.A.; Daykin, C.A.; Mulder, T.P.; Van Duynhoven, J.P. Metabonomics approach to determine metabolic differences between green tea and black tea consumption. J. Agric. Food Chem. 2006, 54, 6929–6938. [Google Scholar] [CrossRef] [PubMed]
- Esteban-Fernández, A.; Ibañez, C.; Simó, C.; Bartolomé, B.; Moreno-Arribas, M.V. An Ultrahigh-Performance Liquid Chromatography-Time-of-Flight Mass Spectrometry Metabolomic Approach to Studying the Impact of Moderate Red-Wine Consumption on Urinary Metabolome. J. Proteome Res. 2018, 17, 1624–1635. [Google Scholar] [CrossRef] [PubMed]
- Urpi-Sarda, M.; Boto-Ordóñez, M.; Queipo-Ortuño, M.I.; Tulipani, S.; Corella, D.; Estruch, R.; Tinahones, F.J.; Andres-Lacueva, C. Phenolic and microbial-targeted metabolomics to discovering and evaluating wine intake biomarkers in human urine and plasma. Electrophoresis 2015, 36, 2259–2268. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Aloy, M.; Llorach, R.; Urpi-Sarda, M.; Tulipani, S.; Estruch, R.; Martínez-González, M.A.; Corella, D.; Fitó, M.; Ros, E.; Salas-Salvadó, J.; et al. Novel multimetabolite prediction of walnut consumption by a urinary biomarker model in a free-living population: The PREDIMED study. J. Proteome Res. 2014, 13, 3476–3483. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Aloy, M.; Llorach, R.; Urpi-Sarda, M.; Jáuregui, O.; Corella, D.; Ruiz-Canela, M.; Salas-Salvadó, J.; Fitó, M.; Ros, E.; Estruch, R.; et al. A metabolomics-driven approach to predict cocoa product consumption by designing a multimetabolite biomarker model in free-living subjects from the PREDIMED study. Mol. Nutr. Food Res. 2015, 59, 212–220. [Google Scholar] [CrossRef]
- Llorach-Asunción, R.; Jauregui, O.; Urpi-Sarda, M.; Andres-Lacueva, C. Methodological aspects for metabolome visualization and characterization: A metabolomic evaluation of the 24 h evolution of human urine after cocoa powder consumption. J. Pharm. Biomed. Anal. 2010, 51, 373–381. [Google Scholar] [CrossRef]
- Wang, M.; Huang, J.; Fan, H.; He, D.; Zhao, S.; Shu, Y.; Li, H.; Liu, L.; Lu, S.; Xiao, C.; et al. Treatment of Rheumatoid Arthritis Using Combination of Methotrexate and Tripterygium Glycosides Tablets-A Quantitative Plasma Pharmacochemical and Pseudotargeted Metabolomic Approach. Front. Pharmacol. 2018, 9, 1051. [Google Scholar] [CrossRef]
- Hung, N.D.; Kim, M.R.; Sok, D.E. 2-Polyunsaturated acyl lysophosphatidylethanolamine attenuates inflammatory response in zymosan A-induced peritonitis in mice. Lipids 2011, 46, 893–906. [Google Scholar] [CrossRef]
- Radhakutty, A.; Mangelsdorf, B.L.; Drake, S.M.; Rowland, A.; Smith, M.D.; Mangoni, A.A.; Thompson, C.H.; Burt, M.G. Opposing effects of rheumatoid arthritis and low dose prednisolone on arginine metabolomics. Atherosclerosis 2017, 266, 190–195. [Google Scholar] [CrossRef]
- Viola, A.; Munari, F.; Sanchez-Rodriguez, R.; Scolaro, T.; Castegna, A. The Metabolic Signature of Macrophage Responses. Front. Immunol. 2019, 10, 1462. [Google Scholar] [CrossRef]
- Nouwen, L.V.; Everts, B. Pathogens MenTORing Macrophages and Dendritic Cells: Manipulation of mTOR and Cellular Metabolism to Promote Immune Escape. Cells 2020, 9, 161. [Google Scholar] [CrossRef]
- Ren, W.; Liu, G.; Yin, J.; Tan, B.; Wu, G.; Bazer, F.W.; Peng, Y.; Yin, Y. Amino-acid transporters in T-cell activation and differentiation. Cell Death Dis. 2017, 8, e2655. [Google Scholar] [CrossRef]
- Nakaya, M.; Xiao, Y.; Zhou, X.; Chang, J.H.; Chang, M.; Cheng, X.; Blonska, M.; Lin, X.; Sun, S.C. Inflammatory T cell responses rely on amino acid transporter ASCT2 facilitation of glutamine uptake and mTORC1 kinase activation. Immunity 2014, 40, 692–705. [Google Scholar] [CrossRef]
- Merlo, L.M.F.; Pigott, E.; DuHadaway, J.B.; Grabler, S.; Metz, R.; Prendergast, G.C.; Mandik-Nayak, L. IDO2 is a critical mediator of autoantibody production and inflammatory pathogenesis in a mouse model of autoimmune arthritis. J. Immunol. 2014, 192, 2082–2090. [Google Scholar] [CrossRef]
- Merlo, L.M.F.; Grabler, S.; DuHadaway, J.B.; Pigott, E.; Manley, K.; Prendergast, G.C.; Laury-Kleintop, L.D.; Mandik-Nayak, L. Therapeutic antibody targeting of indoleamine-2,3-dioxygenase (IDO2) inhibits autoimmune arthritis. Clin. Immunol. 2017, 179, 8–16. [Google Scholar] [CrossRef]
- Collison, J. Experimental arthritis: Do you want to treat arthritis? IDO2! Nat. Rev. Rheumatol. 2017, 13, 196–197. [Google Scholar]
- Zabek, A.; Swierkot, J.; Malak, A.; Zawadzka, I.; Deja, S.; Bogunia-Kubik, K.; Mlynarz, P. Application of (1)H NMR-based serum metabolomic studies for monitoring female patients with rheumatoid arthritis. J. Pharm. Biomed. Anal. 2016, 117, 544–550. [Google Scholar] [CrossRef]
- Kapoor, S.R.; Filer, A.; Fitzpatrick, M.A.; Fisher, B.A.; Taylor, P.C.; Buckley, C.D.; McInnes, I.B.; Raza, K.; Young, S.P. Metabolic profiling predicts response to anti-tumor necrosis factor alpha therapy in patients with rheumatoid arthritis. Arthritis Rheum. 2013, 65, 1448–1456. [Google Scholar] [CrossRef]
- Priori, R.; Casadei, L.; Valerio, M.; Scrivo, R.; Valesini, G.; Manetti, C. ¹H-NMR-Based Metabolomic Study for Identifying Serum Profiles Associated with the Response to Etanercept in Patients with Rheumatoid Arthritis. PLoS ONE 2015, 10, e0138537. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; Fan, J.; Su, Q.; Yang, Z. Cytokines and Abnormal Glucose and Lipid Metabolism. Front. Endocrinol. (Lausanne) 2019, 10, 703. [Google Scholar] [CrossRef] [PubMed]
- Kerekes, G.; Nurmohamed, M.T.; González-Gay, M.A.; Seres, I.; Paragh, G.; Kardos, Z.; Baráth, Z.; Tamási, L.; Soltész, P.; Szekanecz, Z. Rheumatoid arthritis and metabolic syndrome. Nat. Rev. Rheumatol. 2014, 10, 691–696. [Google Scholar] [CrossRef]
- Kuriya, B.; Schieir, O.; Valois, M.F.; Pope, J.E.; Boire, G.; Bessette, L.; Hazlewood, G.; Thorne, J.C.; Tin, D.; Hitchon, C.; et al. Prevalence and Characteristics of Metabolic Syndrome Differ in Men and Women with Early Rheumatoid Arthritis. ACR Open Rheumatol. 2019, 1, 535–541. [Google Scholar] [CrossRef]
- Akbal, N.; Aydin, K.; Tezcan, M.E. Metabolic syndrome is not uncommon in treatment-naïve rheumatoid arthritis patients. Neth. J. Med. 2019, 77, 204–209. [Google Scholar]
- Müller, R.; Kull, M.; Põlluste, K.; Valner, A.; Lember, M.; Kallikorm, R. Factors Associated With Low Lean Mass in Early Rheumatoid Arthritis: A Cross-Sectional Study. Medicina (Kaunas) 2019, 55, 730. [Google Scholar] [CrossRef]
- Barone, M.; Viggiani, M.T.; Anelli, M.G.; Fanizzi, R.; Lorusso, O.; Lopalco, G.; Cantarini, L.; Di Leo, A.; Lapadula, G.; Iannone, F. Sarcopenia in Patients with Rheumatic Diseases: Prevalence and Associated Risk Factors. J. Clin. Med. 2018, 7, 504. [Google Scholar] [CrossRef]
- Hallajzadeh, J.; Safiri, S.; Mansournia, M.A.; Khoramdad, M.; Izadi, N.; Almasi-Hashiani, A.; Pakzad, R.; Ayubi, E.; Sullman, M.J.; Karamzad, N. Metabolic syndrome and its components among rheumatoid arthritis patients: A comprehensive updated systematic review and meta-analysis. PLoS ONE 2017, 12, e0170361. [Google Scholar] [CrossRef]
- Rangel-Huerta, O.D.; Pastor-Villaescusa, B.; Gil, A. Are we close to defining a metabolomic signature of human obesity? A systematic review of metabolomics studies. Metabolomics 2019, 15, 93. [Google Scholar] [CrossRef]
- Lent-Schochet, D.; McLaughlin, M.; Ramakrishnan, N.; Jialal, I. Exploratory metabolomics of metabolic syndrome: A status report. World J. Diabetes 2019, 10, 23–36. [Google Scholar] [CrossRef]
- Zhao, Q.; Shen, H.; Su, K.J.; Tian, Q.; Zhao, L.J.; Qiu, C.; Garrett, T.J.; Liu, J.; Kakhniashvili, D.; Deng, H.W. A joint analysis of metabolomic profiles associated with muscle mass and strength in Caucasian women. Aging (Albany NY) 2018, 10, 2624–2635. [Google Scholar] [CrossRef]
- Petrus, P.; Lecoutre, S.; Dollet, L.; Wiel, C.; Sulen, A.; Gao, H.; Tavira, B.; Laurencikiene, J.; Rooyackers, O.; Checa, A.; et al. Glutamine Links Obesity to Inflammation in Human White Adipose Tissue. Cell Metab. 2020, 31, 375–390. [Google Scholar] [CrossRef]
- Slupsky, C.M.; Rankin, K.N.; Wagner, J.; Fu, H.; Chang, D.; Weljie, A.M.; Saude, E.J.; Lix, B.; Adamko, D.J.; Shah, S.; et al. Investigations of the effects of gender, diurnal variation, and age in human urinary metabolomic profiles. Anal. Chem. 2007, 79, 6995–7004. [Google Scholar] [CrossRef]
- Fan, S.; Yeon, A.; Shahid, M.; Anger, J.T.; Eilber, K.S.; Fiehn, O.; Kim, J. Sex-associated differences in baseline urinary metabolites of healthy adults. Sci. Rep. 2018, 8, 11883. [Google Scholar] [CrossRef]
- Mittelstrass, K.; Ried, J.S.; Yu, Z.; Krumsiek, J.; Gieger, C.; Prehn, C.; Roemisch-Margl, W.; Polonikov, A.; Peters, A.; Theis, F.J.; et al. Discovery of sexual dimorphisms in metabolic and genetic biomarkers. PLoS Genet. 2011, 7, e1002215. [Google Scholar] [CrossRef]
- Darst, B.F.; Koscik, R.L.; Hogan, K.J.; Johnson, S.C.; Engelman, C.D. Longitudinal plasma metabolomics of aging and sex. Aging (Albany NY) 2019, 11, 1262–1282. [Google Scholar] [CrossRef]
- Klareskog, L.; Stolt, P.; Lundberg, K.; Källberg, H.; Bengtsson, C.; Grunewald, J.; Rönnelid, J.; Harris, H.E.; Ulfgren, A.K.; Rantapää-Dahlqvist, S.; et al. A new model for an etiology of rheumatoid arthritis: Smoking may trigger HLA-DR (shared epitope)-restricted immune reactions to autoantigens modified by citrullination. Arthritis Rheum. 2006, 54, 38–46. [Google Scholar] [CrossRef]
- Karlson, E.W.; Chang, S.C.; Cui, J.; Chibnik, L.B.; Fraser, P.A.; De Vivo, I.; Costenbader, K.H. Gene-environment interaction between HLA-DRB1 shared epitope and heavy cigarette smoking in predicting incident rheumatoid arthritis. Ann. Rheum. Dis. 2010, 69, 54–60. [Google Scholar] [CrossRef]
- Qiu, F.; Liang, C.L.; Liu, H.; Zeng, Y.Q.; Hou, S.; Huang, S.; Lai, X.; Dai, Z. Impacts of cigarette smoking on immune responsiveness: Up and down or upside down? Oncotarget 2017, 8, 268–284. [Google Scholar] [CrossRef]
- Gomes, J.P.; Watad, A.; Shoenfeld, Y. Nicotine and autoimmunity: The lotus’ flower in tobacco. Pharmacol. Res. 2018, 128, 101–109. [Google Scholar] [CrossRef]
- Baka, Z.; Buzas, E.; Nagy, G. Rheumatoid arthritis and smoking: Putting the pieces together. Arthritis Res. Ther. 2009, 11, 238. [Google Scholar] [CrossRef]
- Gu, F.; Derkach, A.; Freedman, N.D.; Landi, M.T.; Albanes, D.; Weinstein, S.J.; Mondul, A.M.; Matthews, C.E.; Guertin, K.A.; Xiao, Q.; et al. Cigarette smoking behaviour and blood metabolomics. Int. J. Epidemiol. 2016, 45, 1421–1432. [Google Scholar] [CrossRef]
- Bauer, M.; Fink, B.; Thurmann, L.; Eszlinger, M.; Herberth, G.; Lehmann, I. Tobacco smoking differently influences cell types of the innate and adaptive immune system-indications from CpG site methylation. Clin. Epigenetics 2015, 7, 83. [Google Scholar] [CrossRef]
- Hsu, P.C.; Lan, R.S.; Brasky, T.M.; Marian, C.; Cheema, A.K.; Ressom, H.W.; Loffredo, C.A.; Pickworth, W.B.; Shields, P.G. Metabolomic profiles of current cigarette smokers. Mol. Carcinog. 2017, 56, 594–606. [Google Scholar] [CrossRef]
- Friedman, B.; Cronstein, B. Methotrexate mechanism in treatment of rheumatoid arthritis. Jt. Bone Spine 2019, 86, 301–307. [Google Scholar] [CrossRef]
- Sakaguchi, C.A.; Nieman, D.C.; Signini, E.F.; Abreu, R.M.; Catai, A.M. Metabolomics-Based Studies Assessing Exercise-Induced Alterations of the Human Metabolome: A Systematic Review. Metabolites 2019, 9, 164. [Google Scholar] [CrossRef]
- Kettunen, J.; Tukiainen, T.; Sarin, A.P.; Ortega-Alonso, A.; Tikkanen, E.; Lyytikäinen, L.P.; Kangas, A.J.; Soininen, P.; Würtz, P.; Silander, K.; et al. Genome-wide association study identifies multiple loci influencing human serum metabolite levels. Nat. Genet. 2012, 44, 269–276. [Google Scholar] [CrossRef]
- Shin, S.Y.; Fauman, E.B.; Petersen, A.K.; Krumsiek, J.; Santos, R.; Huang, J.; Arnold, M.; Erte, I.; Forgetta, V.; Yang, T.P.; et al. An atlas of genetic influences on human blood metabolites. Nat. Genet. 2014, 46, 543–550. [Google Scholar] [CrossRef]
- Kettunen, J.; Demirkan, A.; Würtz, P.; Draisma, H.H.; Haller, T.; Rawal, R.; Vaarhorst, A.; Kangas, A.J.; Lyytikäinen, L.P.; Pirinen, M.; et al. Genome-wide study for circulating metabolites identifies 62 loci and reveals novel systemic effects of LPA. Nat. Commun. 2016, 7, 11122. [Google Scholar] [CrossRef]
- Illig, T.; Gieger, C.; Zhai, G.; Römisch-Margl, W.; Wang-Sattler, R.; Prehn, C.; Altmaier, E.; Kastenmüller, G.; Kato, B.S.; Mewes, H.W.; et al. A genome-wide perspective of genetic variation in human metabolism. Nat. Genet. 2010, 42, 137–141. [Google Scholar] [CrossRef]
- Demirkan, A.; van Duijn, C.M.; Ugocsai, P.; Isaacs, A.; Pramstaller, P.P.; Liebisch, G.; Wilson, J.F.; Johansson, Å.; Rudan, I.; Aulchenko, Y.S.; et al. Genome-wide association study identifies novel loci associated with circulating phospho- and sphingolipid concentrations. PLoS Genet. 2012, 8, e1002490. [Google Scholar] [CrossRef] [PubMed]
- Massey, J.; Plant, D.; Hyrich, K.; Morgan, A.W.; Wilson, A.G.; Spiliopoulou, A.; Colombo, M.; McKeigue, P.; Isaacs, J.; Cordell, H.; et al. Genome-wide association study of response to tumour necrosis factor inhibitor therapy in rheumatoid arthritis. Pharm. J. 2018, 18, 657–664. [Google Scholar] [CrossRef] [PubMed]
- Hu, H.J.; Jin, E.H.; Yim, S.H.; Yang, S.Y.; Jung, S.H.; Shin, S.H.; Kim, W.U.; Shim, S.C.; Kim, T.G.; Chung, Y.J. Common variants at the promoter region of the APOM confer a risk of rheumatoid arthritis. Exp. Mol. Med. 2011, 43, 613–621. [Google Scholar] [CrossRef]
- Julià, A.; Ballina, J.; Cañete, J.D.; Balsa, A.; Tornero-Molina, J.; Naranjo, A.; Alperi-López, M.; Erra, A.; Pascual-Salcedo, D.; Barceló, P.; et al. Genome-wide association study of rheumatoid arthritis in the Spanish population: KLF12 as a risk locus for rheumatoid arthritis susceptibility. Arthritis Rheum. 2008, 58, 2275–2286. [Google Scholar] [CrossRef]
- Freudenberg, J.; Lee, H.S.; Han, B.G.; Shin, H.D.; Kang, Y.M.; Sung, Y.K.; Shim, S.C.; Choi, C.B.; Lee, A.T.; Gregersen, P.K.; et al. Genome-wide association study of rheumatoid arthritis in Koreans: Population-specific loci as well as overlap with European susceptibility loci. Arthritis Rheum. 2011, 63, 884–893. [Google Scholar] [CrossRef]
- Terao, C.; Yamada, R.; Ohmura, K.; Takahashi, M.; Kawaguchi, T.; Kochi, Y.; Okada, Y.; Nakamura, Y.; Yamamoto, K.; Melchers, I.; et al. The human AIRE gene at chromosome 21q22 is a genetic determinant for the predisposition to rheumatoid arthritis in Japanese population. Hum. Mol. Genet. 2011, 20, 2680–2685. [Google Scholar] [CrossRef]
- Myouzen, K.; Kochi, Y.; Okada, Y.; Terao, C.; Suzuki, A.; Ikari, K.; Tsunoda, T.; Takahashi, A.; Kubo, M.; Taniguchi, A.; et al. Functional variants in NFKBIE and RTKN2 involved in activation of the NF-κB pathway are associated with rheumatoid arthritis in Japanese. PLoS Genet. 2012, 8, e1002949. [Google Scholar] [CrossRef]
- Tokuhiro, S.; Yamada, R.; Chang, X.; Suzuki, A.; Kochi, Y.; Sawada, T.; Suzuki, M.; Nagasaki, M.; Ohtsuki, M.; Ono, M.; et al. An intronic SNP in a RUNX1 binding site of SLC22A4, encoding an organic cation transporter, is associated with rheumatoid arthritis. Nat. Genet. 2003, 35, 341–348. [Google Scholar] [CrossRef]
- Newman, B.; Wintle, R.F.; van Oene, M.; Yazdanpanah, M.; Owen, J.; Johnson, B.; Gu, X.; Amos, C.I.; Keystone, E.; Rubin, L.A.; et al. SLC22A4 polymorphisms implicated in rheumatoid arthritis and Crohn’s disease are not associated with rheumatoid arthritis in a Canadian Caucasian population. Arthritis Rheum. 2005, 52, 425–429. [Google Scholar] [CrossRef]
- Gieger, C.; Geistlinger, L.; Altmaier, E.; Hrabé de Angelis, M.; Kronenberg, F.; Meitinger, T.; Mewes, H.W.; Wichmann, H.E.; Weinberger, K.M.; Adamski, J.; et al. Genetics meets metabolomics: A genome-wide association study of metabolite profiles in human serum. PLoS Genet. 2008, 4, e1000282. [Google Scholar] [CrossRef]
- Consortium, W.T.C.C. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 2007, 447, 661–678. [Google Scholar]
- Goodrich, J.K.; Waters, J.L.; Poole, A.C.; Sutter, J.L.; Koren, O.; Blekhman, R.; Beaumont, M.; Van Treuren, W.; Knight, R.; Bell, J.T.; et al. Human genetics shape the gut microbiome. Cell 2014, 159, 789–799. [Google Scholar] [CrossRef]
- Goodrich, J.K.; Davenport, E.R.; Beaumont, M.; Jackson, M.A.; Knight, R.; Ober, C.; Spector, T.D.; Bell, J.T.; Clark, A.G.; Ley, R.E. Genetic Determinants of the Gut Microbiome in UK Twins. Cell Host Microbe 2016, 19, 731–743. [Google Scholar] [CrossRef]
- Xie, H.; Guo, R.; Zhong, H.; Feng, Q.; Lan, Z.; Qin, B.; Ward, K.J.; Jackson, M.A.; Xia, Y.; Chen, X.; et al. Shotgun Metagenomics of 250 Adult Twins Reveals Genetic and Environmental Impacts on the Gut Microbiome. Cell Syst. 2016, 3, 572–584.e573. [Google Scholar] [CrossRef]
- Visconti, A.; Le Roy, C.I.; Rosa, F.; Rossi, N.; Martin, T.C.; Mohney, R.P.; Li, W.; de Rinaldis, E.; Bell, J.T.; Venter, J.C.; et al. Interplay between the human gut microbiome and host metabolism. Nat. Commun. 2019, 10, 4505. [Google Scholar] [CrossRef]
- Enam, F.; Mansell, T.J. Prebiotics: Tools to manipulate the gut microbiome and metabolome. J. Ind. Microbiol. Biotechnol. 2019, 46, 1445–1459. [Google Scholar] [CrossRef]
- Halkjær, S.I.; Christensen, A.H.; Lo, B.Z.S.; Browne, P.D.; Günther, S.; Hansen, L.H.; Petersen, A.M. Faecal microbiota transplantation alters gut microbiota in patients with irritable bowel syndrome: Results from a randomised, double-blind placebo-controlled study. Gut 2018, 67, 2107–2115. [Google Scholar] [CrossRef] [PubMed]
- Hemarajata, P.; Versalovic, J. Effects of probiotics on gut microbiota: Mechanisms of intestinal immunomodulation and neuromodulation. Therap Adv. Gastroenterol. 2013, 6, 39–51. [Google Scholar] [CrossRef]
- Schroeder, B.O.; Backhed, F. Signals from the gut microbiota to distant organs in physiology and disease. Nat. Med. 2016, 22, 1079–1089. [Google Scholar] [CrossRef]
- Koeth, R.A.; Wang, Z.; Levison, B.S.; Buffa, J.A.; Org, E.; Sheehy, B.T.; Britt, E.B.; Fu, X.; Wu, Y.; Li, L.; et al. Intestinal microbiota metabolism of L-carnitine, a nutrient in red meat, promotes atherosclerosis. Nat. Med. 2013, 19, 576–585. [Google Scholar] [CrossRef]
- Wang, Z.; Klipfell, E.; Bennett, B.J.; Koeth, R.; Levison, B.S.; Dugar, B.; Feldstein, A.E.; Britt, E.B.; Fu, X.; Chung, Y.M.; et al. Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature 2011, 472, 57–63. [Google Scholar] [CrossRef]
- Scher, J.U.; Sczesnak, A.; Longman, R.S.; Segata, N.; Ubeda, C.; Bielski, C.; Rostron, T.; Cerundolo, V.; Pamer, E.G.; Abramson, S.B.; et al. Expansion of intestinal Prevotella copri correlates with enhanced susceptibility to arthritis. Elife 2013, 2, e01202. [Google Scholar] [CrossRef]
- Gaucher, F.; Bonnassie, S.; Rabah, H.; Marchand, P.; Blanc, P.; Jeantet, R.; Jan, G. Review: Adaptation of Beneficial Propionibacteria, Lactobacilli, and Bifidobacteria Improves Tolerance Toward Technological and Digestive Stresses. Front. Microbiol. 2019, 10, 841. [Google Scholar] [CrossRef]
- Roager, H.M.; Licht, T.R. Microbial tryptophan catabolites in health and disease. Nat. Commun. 2018, 9, 3294. [Google Scholar] [CrossRef]
- Bach Knudsen, K.E. Microbial degradation of whole-grain complex carbohydrates and impact on short-chain fatty acids and health. Adv. Nutr. 2015, 6, 206–213. [Google Scholar] [CrossRef]
- Song, X.; Sun, X.; Oh, S.F.; Wu, M.; Zhang, Y.; Zheng, W.; Geva-Zatorsky, N.; Jupp, R.; Mathis, D.; Benoist, C.; et al. Microbial bile acid metabolites modulate gut RORγ. Nature 2019. [Google Scholar] [CrossRef]
- Vanegas, S.M.; Meydani, M.; Barnett, J.B.; Goldin, B.; Kane, A.; Rasmussen, H.; Brown, C.; Vangay, P.; Knights, D.; Jonnalagadda, S.; et al. Substituting whole grains for refined grains in a 6-wk randomized trial has a modest effect on gut microbiota and immune and inflammatory markers of healthy adults. Am. J. Clin. Nutr. 2017, 105, 635–650. [Google Scholar] [CrossRef]
- Bushman, F.D.; Lewis, J.D.; Wu, G.D. Diet, gut enterotypes and health: Is there a link? Nestle Nutr. Inst. Workshop Ser. 2013, 77, 65–73. [Google Scholar]
- Shoaie, S.; Ghaffari, P.; Kovatcheva-Datchary, P.; Mardinoglu, A.; Sen, P.; Pujos-Guillot, E.; de Wouters, T.; Juste, C.; Rizkalla, S.; Chilloux, J.; et al. Quantifying Diet-Induced Metabolic Changes of the Human Gut Microbiome. Cell Metab. 2015, 22, 320–331. [Google Scholar] [CrossRef]
- Roager, H.M.; Vogt, J.K.; Kristensen, M.; Hansen, L.B.S.; Ibrügger, S.; Mærkedahl, R.B.; Bahl, M.I.; Lind, M.V.; Nielsen, R.L.; Frøkiær, H.; et al. Whole grain-rich diet reduces body weight and systemic low-grade inflammation without inducing major changes of the gut microbiome: A randomised cross-over trial. Gut 2019, 68, 83–93. [Google Scholar] [CrossRef]
- Fujisaka, S.; Avila-Pacheco, J.; Soto, M.; Kostic, A.; Dreyfuss, J.M.; Pan, H.; Ussar, S.; Altindis, E.; Li, N.; Bry, L.; et al. Diet, Genetics, and the Gut Microbiome Drive Dynamic Changes in Plasma Metabolites. Cell Rep. 2018, 22, 3072–3086. [Google Scholar] [CrossRef] [PubMed]
- Leeming, E.R.; Johnson, A.J.; Spector, T.D.; Le Roy, C.I. Effect of Diet on the Gut Microbiota: Rethinking Intervention Duration. Nutrients 2019, 11. [Google Scholar] [CrossRef]
- Miyamoto, J.; Igarashi, M.; Watanabe, K.; Karaki, S.I.; Mukouyama, H.; Kishino, S.; Li, X.; Ichimura, A.; Irie, J.; Sugimoto, Y.; et al. Gut microbiota confers host resistance to obesity by metabolizing dietary polyunsaturated fatty acids. Nat. Commun. 2019, 10, 4007. [Google Scholar] [CrossRef]
- Singh, R.K.; Chang, H.W.; Yan, D.; Lee, K.M.; Ucmak, D.; Wong, K.; Abrouk, M.; Farahnik, B.; Nakamura, M.; Zhu, T.H.; et al. Influence of diet on the gut microbiome and implications for human health. J. Transl. Med. 2017, 15, 73. [Google Scholar] [CrossRef]
- Shin, J.H.; Jung, S.; Kim, S.A.; Kang, M.S.; Kim, M.S.; Joung, H.; Hwang, G.S.; Shin, D.M. Differential Effects of Typical Korean Versus American-Style Diets on Gut Microbial Composition and Metabolic Profile in Healthy Overweight Koreans: A Randomized Crossover Trial. Nutrients 2019, 11. [Google Scholar] [CrossRef]
- Healey, G.R.; Murphy, R.; Brough, L.; Butts, C.A.; Coad, J. Interindividual variability in gut microbiota and host response to dietary interventions. Nutr. Rev. 2017, 75, 1059–1080. [Google Scholar] [CrossRef]
- Hitchon, C.A.; El-Gabalawy, H.S.; Bezabeh, T. Characterization of synovial tissue from arthritis patients: A proton magnetic resonance spectroscopic investigation. Rheumatol. Int. 2009, 29, 1205–1211. [Google Scholar] [CrossRef] [PubMed]
- Volchenkov, R.; Dung Cao, M.; Elgstoen, K.B.; Goll, G.L.; Eikvar, K.; Bjorneboe, O.; Bathen, T.F.; Holen, H.L.; Kvien, T.K.; Skalhegg, B.S. Metabolic profiling of synovial tissue shows altered glucose and choline metabolism in rheumatoid arthritis samples. Scand. J. Rheumatol. 2017, 46, 160–161. [Google Scholar] [CrossRef] [PubMed]
- Quinonez-Flores, C.M.; Gonzalez-Chavez, S.A.; Pacheco-Tena, C. Hypoxia and its implications in rheumatoid arthritis. J. Biomed. Sci. 2016, 23, 62. [Google Scholar] [CrossRef] [PubMed]
- Hua, S.; Dias, T.H. Hypoxia-Inducible Factor (HIF) as a Target for Novel Therapies in Rheumatoid Arthritis. Front. Pharmacol. 2016, 7, 184. [Google Scholar] [CrossRef]
- Snider, S.A.; Margison, K.D.; Ghorbani, P.; LeBlond, N.D.; O’Dwyer, C.; Nunes, J.R.C.; Nguyen, T.; Xu, H.; Bennett, S.A.L.; Fullerton, M.D. Choline transport links macrophage phospholipid metabolism and inflammation. J. Biol. Chem. 2018, 293, 11600–11611. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Lopez, E.; Zhong, Z.; Stubelius, A.; Sweeney, S.R.; Booshehri, L.M.; Antonucci, L.; Liu-Bryan, R.; Lodi, A.; Terkeltaub, R.; Lacal, J.C.; et al. Choline Uptake and Metabolism Modulate Macrophage IL-1beta and IL-18 Production. Cell Metab. 2019, 29, 1350–1362 e1357. [Google Scholar] [CrossRef] [PubMed]
- Park, J.; Lee, I.S.; Kim, K.H.; Kim, Y.; An, E.J.; Jang, H.J. GI inflammation Increases Sodium-Glucose Cotransporter Sglt1. Int. J. Mol. Sci. 2019, 20. [Google Scholar] [CrossRef]
- Almutairi, S.M.; Ali, A.K.; He, W.; Yang, D.S.; Ghorbani, P.; Wang, L.; Fullerton, M.D.; Lee, S.H. Interleukin-18 up-regulates amino acid transporters and facilitates amino acid-induced mTORC1 activation in natural killer cells. J. Biol. Chem. 2019, 294, 4644–4655. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.Y.; Zheng, K.D.; Lin, K.; Zheng, G.; Zou, H.; Wang, J.M.; Lin, Y.Y.; Chuka, C.M.; Ge, R.S.; Zhai, W.; et al. Energy Metabolism Disorder as a Contributing Factor of Rheumatoid Arthritis: A Comparative Proteomic and Metabolomic Study. PLoS ONE 2015, 10, e0132695. [Google Scholar]
- Bartok, B.; Firestein, G.S. Fibroblast-like synoviocytes: Key effector cells in rheumatoid arthritis. Immunol. Rev. 2010, 233, 233–255. [Google Scholar] [CrossRef]
- Ahn, J.K.; Kim, S.; Hwang, J.; Kim, J.; Kim, K.H.; Cha, H.S. GC/TOF-MS-based metabolomic profiling in cultured fibroblast-like synoviocytes from rheumatoid arthritis. Joint Bone Spine 2016, 83, 707–713. [Google Scholar] [CrossRef]
- Bustamante, M.F.; Garcia-Carbonell, R.; Whisenant, K.D.; Guma, M. Fibroblast-like synoviocyte metabolism in the pathogenesis of rheumatoid arthritis. Arthritis Res. Ther. 2017, 19, 110. [Google Scholar] [CrossRef]
- Bottini, N.; Firestein, G.S. Duality of fibroblast-like synoviocytes in RA: Passive responders and imprinted aggressors. Nat. Rev. Rheumatol. 2013, 9, 24–33. [Google Scholar] [CrossRef]
- Haringman, J.J.; Gerlag, D.M.; Zwinderman, A.H.; Smeets, T.J.; Kraan, M.C.; Baeten, D.; McInnes, I.B.; Bresnihan, B.; Tak, P.P. Synovial tissue macrophages: A sensitive biomarker for response to treatment in patients with rheumatoid arthritis. Ann. Rheum. Dis. 2005, 64, 834–838. [Google Scholar] [CrossRef]
- Kim, S.; Hwang, J.; Xuan, J.; Jung, Y.H.; Cha, H.S.; Kim, K.H. Global metabolite profiling of synovial fluid for the specific diagnosis of rheumatoid arthritis from other inflammatory arthritis. PloS ONE 2014, 9, e97501. [Google Scholar] [CrossRef]
- Tannahill, G.M.; Curtis, A.M.; Adamik, J.; Palsson-McDermott, E.M.; McGettrick, A.F.; Goel, G.; Frezza, C.; Bernard, N.J.; Kelly, B.; Foley, N.H.; et al. Succinate is an inflammatory signal that induces IL-1β through HIF-1α. Nature 2013, 496, 238–242. [Google Scholar] [CrossRef] [PubMed]
- Shime, H.; Yabu, M.; Akazawa, T.; Kodama, K.; Matsumoto, M.; Seya, T.; Inoue, N. Tumor-secreted lactic acid promotes IL-23/IL-17 proinflammatory pathway. J. Immunol. 2008, 180, 7175–7183. [Google Scholar] [CrossRef] [PubMed]
- Littlewood-Evans, A.; Sarret, S.; Apfel, V.; Loesle, P.; Dawson, J.; Zhang, J.; Muller, A.; Tigani, B.; Kneuer, R.; Patel, S.; et al. GPR91 senses extracellular succinate released from inflammatory macrophages and exacerbates rheumatoid arthritis. J. Exp. Med. 2016, 213, 1655–1662. [Google Scholar] [CrossRef]
- Haas, R.; Smith, J.; Rocher-Ros, V.; Nadkarni, S.; Montero-Melendez, T.; D’Acquisto, F.; Bland, E.J.; Bombardieri, M.; Pitzalis, C.; Perretti, M.; et al. Lactate Regulates Metabolic and Pro-inflammatory Circuits in Control of T Cell Migration and Effector Functions. PLoS Biol 2015, 13, e1002202. [Google Scholar] [CrossRef]
- Falconer, J.; Murphy, A.N.; Young, S.; Clark, A.R.; Tiziani, S.; Guma, M.; Buckley, C.D. Synovial cell metabolism and chronic inflammation in rheumatoid arthritis. Arthritis Rheumatol. 1002. [Google Scholar]
- Newton, R.; Priyadharshini, B.; Turka, L.A. Immunometabolism of regulatory T cells. Nat. Immunol. 2016, 17, 618–625. [Google Scholar] [CrossRef]
- Ryan, D.G.; O’Neill, L.A.J. Krebs Cycle Reborn in Macrophage Immunometabolism. Annu. Rev. Immunol. 2020. [Google Scholar] [CrossRef]
- Weyand, C.M.; Goronzy, J.J. Immunometabolism in the development of rheumatoid arthritis. Immunol. Rev. 2020, 294, 177–187. [Google Scholar] [CrossRef]
- Singer, B.D.; Chandel, N.S. Immunometabolism of pro-repair cells. J. Clin. Invest. 2019, 129, 2597–2607. [Google Scholar] [CrossRef] [PubMed]
- Poznanski, S.M.; Barra, N.G.; Ashkar, A.A.; Schertzer, J.D. Immunometabolism of T cells and NK cells: Metabolic control of effector and regulatory function. Inflamm. Res. 2018, 67, 813–828. [Google Scholar] [CrossRef] [PubMed]
- Zeisel, S.H.; Mar, M.H.; Howe, J.C.; Holden, J.M. Concentrations of choline-containing compounds and betaine in common foods. J. Nutr. 2003, 133, 1302–1307. [Google Scholar] [CrossRef] [PubMed]
- Guasch-Ferré, M.; Hu, F.B.; Ruiz-Canela, M.; Bulló, M.; Toledo, E.; Wang, D.D.; Corella, D.; Gómez-Gracia, E.; Fiol, M.; Estruch, R.; et al. Plasma Metabolites From Choline Pathway and Risk of Cardiovascular Disease in the PREDIMED (Prevention With Mediterranean Diet) Study. J. Am. Heart Assoc. 2017, 6. [Google Scholar] [CrossRef] [PubMed]
- Lang, D.H.; Yeung, C.K.; Peter, R.M.; Ibarra, C.; Gasser, R.; Itagaki, K.; Philpot, R.M.; Rettie, A.E. Isoform specificity of trimethylamine N-oxygenation by human flavin-containing monooxygenase (FMO) and P450 enzymes: Selective catalysis by FMO3. Biochem. Pharmacol. 1998, 56, 1005–1012. [Google Scholar] [CrossRef]
- Coras, R.; Kavanaugh, A.; Boyd, T.; Huynh, D.; Lagerborg, K.A.; Xu, Y.J.; Rosenthal, S.B.; Jain, M.; Guma, M. Choline metabolite, trimethylamine N-oxide (TMAO), is associated with inflammation in psoriatic arthritis. Clin. Exp. Rheumatol. 2019, 37, 481–484. [Google Scholar] [PubMed]
- Gelfand, J.M.; Yeung, H. Metabolic syndrome in patients with psoriatic disease. J. Rheumatol. Suppl. 2012, 89, 24–28. [Google Scholar] [CrossRef] [PubMed]
- Schugar, R.C.; Shih, D.M.; Warrier, M.; Helsley, R.N.; Burrows, A.; Ferguson, D.; Brown, A.L.; Gromovsky, A.D.; Heine, M.; Chatterjee, A.; et al. The TMAO-Producing Enzyme Flavin-Containing Monooxygenase 3 Regulates Obesity and the Beiging of White Adipose Tissue. Cell Rep. 2017, 20, 279. [Google Scholar] [CrossRef]
- Weljie, A.M.; Dowlatabadi, R.; Miller, B.J.; Vogel, H.J.; Jirik, F.R. An inflammatory arthritis-associated metabolite biomarker pattern revealed by 1H NMR spectroscopy. J. Proteome Res. 2007, 6, 3456–3464. [Google Scholar] [CrossRef]
- Wu, Y.; Lau, B.; Smith, S.; Troyan, K.; Barnett Foster, D.E. Enteropathogenic Escherichia coli infection triggers host phospholipid metabolism perturbations. Infect. Immun. 2004, 72, 6764–6772. [Google Scholar] [CrossRef]
- Guma, M.; Sanchez-Lopez, E.; Lodi, A.; Garcia-Carbonell, R.; Tiziani, S.; Karin, M.; Lacal, J.C.; Firestein, G.S. Choline kinase inhibition in rheumatoid arthritis. Ann. Rheum. Dis. 2015, 74, 1399–1407. [Google Scholar] [CrossRef]
- Zhenyukh, O.; Civantos, E.; Ruiz-Ortega, M.; Sánchez, M.S.; Vázquez, C.; Peiró, C.; Egido, J.; Mas, S. High concentration of branched-chain amino acids promotes oxidative stress, inflammation and migration of human peripheral blood mononuclear cells via mTORC1 activation. Free Radic. Biol. Med. 2017, 104, 165–177. [Google Scholar] [CrossRef]
- Papathanassiu, A.E.; Ko, J.H.; Imprialou, M.; Bagnati, M.; Srivastava, P.K.; Vu, H.A.; Cucchi, D.; McAdoo, S.P.; Ananieva, E.A.; Mauro, C.; et al. BCAT1 controls metabolic reprogramming in activated human macrophages and is associated with inflammatory diseases. Nat. Commun. 2017, 8, 16040. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, S.; Saegusa, J.; Sendo, S.; Okano, T.; Akashi, K.; Irino, Y.; Morinobu, A. Glutaminase 1 plays a key role in the cell growth of fibroblast-like synoviocytes in rheumatoid arthritis. Arthritis Res. Ther. 2017, 19, 76. [Google Scholar] [CrossRef] [PubMed]
- Biniecka, M.; Canavan, M.; McGarry, T.; Gao, W.; McCormick, J.; Cregan, S.; Gallagher, L.; Smith, T.; Phelan, J.J.; Ryan, J.; et al. Dysregulated bioenergetics: A key regulator of joint inflammation. Ann. Rheum. Dis. 2016, 75, 2192–2200. [Google Scholar] [CrossRef] [PubMed]
- Pucino, V.; Certo, M.; Bulusu, V.; Cucchi, D.; Goldmann, K.; Pontarini, E.; Haas, R.; Smith, J.; Headland, S.E.; Blighe, K.; et al. Lactate Buildup at the Site of Chronic Inflammation Promotes Disease by Inducing CD4. Cell Metab. 2019, 30, 1055–1074. [Google Scholar] [CrossRef]
- Zhang, D.; Tang, Z.; Huang, H.; Zhou, G.; Cui, C.; Weng, Y.; Liu, W.; Kim, S.; Lee, S.; Perez-Neut, M.; et al. Metabolic regulation of gene expression by histone lactylation. Nature 2019, 574, 575–580. [Google Scholar] [CrossRef] [PubMed]
- Ai, R.; Laragione, T.; Hammaker, D.; Boyle, D.L.; Wildberg, A.; Maeshima, K.; Palescandolo, E.; Krishna, V.; Pocalyko, D.; Whitaker, J.W.; et al. Comprehensive epigenetic landscape of rheumatoid arthritis fibroblast-like synoviocytes. Nat. Commun. 2018, 9, 1921. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zheng, J.Y.; Liu, J.Q.; Yang, J.; Liu, Y.; Wang, C.; Ma, X.N.; Liu, B.L.; Xin, G.Z.; Liu, L.F. Succinate/NLRP3 Inflammasome Induces Synovial Fibroblast Activation: Therapeutical Effects of Clematichinenoside AR on Arthritis. Front. Immunol. 2016, 7, 532. [Google Scholar] [CrossRef]
- Saraiva, A.L.; Veras, F.P.; Peres, R.S.; Talbot, J.; de Lima, K.A.; Luiz, J.P.; Carballido, J.M.; Cunha, T.M.; Cunha, F.Q.; Ryffel, B.; et al. Succinate receptor deficiency attenuates arthritis by reducing dendritic cell traffic and expansion of T. FASEB J. 2018. [Google Scholar] [CrossRef]
- Lampropoulou, V.; Sergushichev, A.; Bambouskova, M.; Nair, S.; Vincent, E.E.; Loginicheva, E.; Cervantes-Barragan, L.; Ma, X.; Huang, S.C.; Griss, T.; et al. Itaconate Links Inhibition of Succinate Dehydrogenase with Macrophage Metabolic Remodeling and Regulation of Inflammation. Cell Metab. 2016, 24, 158–166. [Google Scholar] [CrossRef]
- Michopoulos, F.; Karagianni, N.; Whalley, N.M.; Firth, M.A.; Nikolaou, C.; Wilson, I.D.; Critchlow, S.E.; Kollias, G.; Theodoridis, G.A. Targeted Metabolic Profiling of the Tg197 Mouse Model Reveals Itaconic Acid as a Marker of Rheumatoid Arthritis. J. Proteome Res. 2016, 15, 4579–4590. [Google Scholar] [CrossRef]
- Turesson, C.; Bergström, U.; Pikwer, M.; Nilsson, J.; Jacobsson, L.T. High serum cholesterol predicts rheumatoid arthritis in women, but not in men: A prospective study. Arthritis Res. Ther. 2015, 17, 284. [Google Scholar] [CrossRef]
- Choi, W.S.; Lee, G.; Song, W.H.; Koh, J.T.; Yang, J.; Kwak, J.S.; Kim, H.E.; Kim, S.K.; Son, Y.O.; Nam, H.; et al. The CH25H-CYP7B1-RORα axis of cholesterol metabolism regulates osteoarthritis. Nature 2019, 566, 254–258. [Google Scholar] [CrossRef] [PubMed]
- Tang, M.W.; Koopman, F.A.; Visscher, J.P.; de Hair, M.J.; Gerlag, D.M.; Tak, P.P. Hormone, metabolic peptide, and nutrient levels in the earliest phases of rheumatoid arthritis-contribution of free fatty acids to an increased cardiovascular risk during very early disease. Clin. Rheumatol. 2017, 36, 269–278. [Google Scholar] [CrossRef] [PubMed]
- Frommer, K.W.; Schäffler, A.; Rehart, S.; Lehr, A.; Müller-Ladner, U.; Neumann, E. Free fatty acids: Potential proinflammatory mediators in rheumatic diseases. Ann. Rheum. Dis. 2015, 74, 303–310. [Google Scholar] [CrossRef]
- McCoy, J.M.; Wicks, J.R.; Audoly, L.P. The role of prostaglandin E2 receptors in the pathogenesis of rheumatoid arthritis. J. Clin. Invest. 2002, 110, 651–658. [Google Scholar] [CrossRef] [PubMed]
- Peng, A.; Lu, X.; Huang, J.; He, M.; Xu, J.; Huang, H.; Chen, Q. Rheumatoid arthritis synovial fibroblasts promote TREM-1 expression in monocytes via COX-2/PGE. Arthritis Res. Ther. 2019, 21, 169. [Google Scholar] [CrossRef] [PubMed]
- Schmöcker, C.; Zhang, I.W.; Kiesler, S.; Kassner, U.; Ostermann, A.I.; Steinhagen-Thiessen, E.; Schebb, N.H.; Weylandt, K.H. Effect of Omega-3 Fatty Acid Supplementation on Oxylipins in a Routine Clinical Setting. Int. J. Mol. Sci. 2018, 19. [Google Scholar] [CrossRef] [PubMed]
- Calder, P.C. Omega-3 fatty acids and inflammatory processes. Nutrients 2010, 2, 355–374. [Google Scholar] [CrossRef]
- Senftleber, N.K.; Nielsen, S.M.; Andersen, J.R.; Bliddal, H.; Tarp, S.; Lauritzen, L.; Furst, D.E.; Suarez-Almazor, M.E.; Lyddiatt, A.; Christensen, R. Marine Oil Supplements for Arthritis Pain: A Systematic Review and Meta-Analysis of Randomized Trials. Nutrients 2017, 9, 42. [Google Scholar] [CrossRef]
- Veselinovic, M.; Vasiljevic, D.; Vucic, V.; Arsic, A.; Petrovic, S.; Tomic-Lucic, A.; Savic, M.; Zivanovic, S.; Stojic, V.; Jakovljevic, V. Clinical Benefits of n-3 PUFA and ɤ-Linolenic Acid in Patients with Rheumatoid Arthritis. Nutrients 2017, 9, 325. [Google Scholar] [CrossRef]
- Rodriguez-Carrio, J.; Alperi-Lopez, M.; Lopez, P.; Ballina-Garcia, F.J.; Suarez, A. Non-Esterified Fatty Acids Profiling in Rheumatoid Arthritis: Associations with Clinical Features and Th1 Response. PLoS ONE 2016, 11, e0159573. [Google Scholar] [CrossRef]
- Stephensen, C.B.; Armstrong, P.; Newman, J.W.; Pedersen, T.L.; Legault, J.; Schuster, G.U.; Kelley, D.; Vikman, S.; Hartiala, J.; Nassir, R.; et al. ALOX5 gene variants affect eicosanoid production and response to fish oil supplementation. J. Lipid Res. 2011, 52, 991–1003. [Google Scholar] [CrossRef] [PubMed]
- Liagre, B.; Vergne, P.; Rigaud, M.; Beneytout, J.L. Expression of arachidonate platelet-type 12-lipoxygenase in human rheumatoid arthritis type B synoviocytes. FEBS Lett. 1997, 414, 159–164. [Google Scholar] [CrossRef]
- Krönke, G.; Katzenbeisser, J.; Uderhardt, S.; Zaiss, M.M.; Scholtysek, C.; Schabbauer, G.; Zarbock, A.; Koenders, M.I.; Axmann, R.; Zwerina, J.; et al. 12/15-lipoxygenase counteracts inflammation and tissue damage in arthritis. J. Immunol. 2009, 183, 3383–3389. [Google Scholar] [CrossRef] [PubMed]
- Gheorghe, K.R.; Korotkova, M.; Catrina, A.I.; Backman, L.; af Klint, E.; Claesson, H.E.; Rådmark, O.; Jakobsson, P.J. Expression of 5-lipoxygenase and 15-lipoxygenase in rheumatoid arthritis synovium and effects of intraarticular glucocorticoids. Arthritis Res. Ther. 2009, 11, R83. [Google Scholar] [CrossRef] [PubMed]
- Shan, J.; Peng, L.; Qian, W.; Xie, T.; Kang, A.; Gao, B.; Di, L. Integrated Serum and Fecal Metabolomics Study of Collagen-Induced Arthritis Rats and the Therapeutic Effects of the Zushima Tablet. Front. Pharmacol. 2018, 9, 891. [Google Scholar] [CrossRef] [PubMed]
- Hui, W.; Yu, D.; Cao, Z.; Zhao, X. Butyrate inhibit collagen-induced arthritis via Treg/IL-10/Th17 axis. Int. Immunopharmacol. 2019, 68, 226–233. [Google Scholar] [CrossRef]
- Ginos, B.N.R.; Navarro, S.L.; Schwarz, Y.; Gu, H.; Wang, D.; Randolph, T.W.; Shojaie, A.; Hullar, M.A.J.; Lampe, P.D.; Kratz, M.; et al. Circulating bile acids in healthy adults respond differently to a dietary pattern characterized by whole grains, legumes and fruits and vegetables compared to a diet high in refined grains and added sugars: A randomized, controlled, crossover feeding study. Metabolism 2018, 83, 197–204. [Google Scholar]
- Wammers, M.; Schupp, A.K.; Bode, J.G.; Ehlting, C.; Wolf, S.; Deenen, R.; Köhrer, K.; Häussinger, D.; Graf, D. Reprogramming of pro-inflammatory human macrophages to an anti-inflammatory phenotype by bile acids. Sci. Rep. 2018, 8, 255. [Google Scholar] [CrossRef]
- Pols, T.W.; Nomura, M.; Harach, T.; Lo Sasso, G.; Oosterveer, M.H.; Thomas, C.; Rizzo, G.; Gioiello, A.; Adorini, L.; Pellicciari, R.; et al. TGR5 activation inhibits atherosclerosis by reducing macrophage inflammation and lipid loading. Cell Metab. 2011, 14, 747–757. [Google Scholar] [CrossRef]
- Yoneno, K.; Hisamatsu, T.; Shimamura, K.; Kamada, N.; Ichikawa, R.; Kitazume, M.T.; Mori, M.; Uo, M.; Namikawa, Y.; Matsuoka, K.; et al. TGR5 signalling inhibits the production of pro-inflammatory cytokines by in vitro differentiated inflammatory and intestinal macrophages in Crohn’s disease. Immunology 2013, 139, 19–29. [Google Scholar] [CrossRef]
- Li, L.; Liu, C.; Liu, M.; Shi, L.; Liu, Q.; Guan, H.; Li, P. Taurochenodeoxycholic acid induces apoptosis of fibroblast-like synoviocytes. Eur. J. Pharmacol. 2013, 706, 36–40. [Google Scholar] [CrossRef] [PubMed]
- Szanto, S.; Koreny, T.; Mikecz, K.; Glant, T.T.; Szekanecz, Z.; Varga, J. Inhibition of indoleamine 2,3-dioxygenase-mediated tryptophan catabolism accelerates collagen-induced arthritis in mice. Arthritis Res. Ther. 2007, 9, R50. [Google Scholar] [CrossRef] [PubMed]
- Kang, K.Y.; Lee, S.H.; Jung, S.M.; Park, S.H.; Jung, B.H.; Ju, J.H. Downregulation of Tryptophan-related Metabolomic Profile in Rheumatoid Arthritis Synovial Fluid. J. Rheumatol. 2015, 42, 2003–2011. [Google Scholar] [CrossRef] [PubMed]
- Khanna, S.; Jaiswal, K.S.; Gupta, B. Managing Rheumatoid Arthritis with Dietary Interventions. Front. Nutr. 2017, 4, 52. [Google Scholar] [CrossRef] [PubMed]
- Masuko, K. A Potential Benefit of “Balanced Diet” for Rheumatoid Arthritis. Front. Med. (Lausanne) 2018, 5, 141. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Micheletti, R. Role of diet in rheumatic disease. Rheum. Dis. Clin. North. Am. 2011, 37, 119–133. [Google Scholar] [CrossRef]
- Adam, O.; Beringer, C.; Kless, T.; Lemmen, C.; Adam, A.; Wiseman, M.; Adam, P.; Klimmek, R.; Forth, W. Anti-inflammatory effects of a low arachidonic acid diet and fish oil in patients with rheumatoid arthritis. Rheumatol. Int. 2003, 23, 27–36. [Google Scholar] [CrossRef]
- Haugen, M.A.; Kjeldsen-Kragh, J.; Bjerve, K.S.; Høstmark, A.T.; Førre, O. Changes in plasma phospholipid fatty acids and their relationship to disease activity in rheumatoid arthritis patients treated with a vegetarian diet. Br. J. Nutr. 1994, 72, 555–566. [Google Scholar] [CrossRef][Green Version]
- Lindqvist, H.M.; Gjertsson, I.; Andersson, S.; Calder, P.C.; Bärebring, L. Influence of blue mussel (Mytilus edulis) intake on fatty acid composition in erythrocytes and plasma phospholipids and serum metabolites in women with rheumatoid arthritis. Prostaglandins Leukot. Essent. Fatty Acids 2019, 150, 7–15. [Google Scholar] [CrossRef]
- Lee, A.L.; Park, Y. The association between n-3 polyunsaturated fatty acid levels in erythrocytes and the risk of rheumatoid arthritis in Korean women. Ann. Nutr. Metab. 2013, 63, 88–95. [Google Scholar] [CrossRef]
- Sugihara, K.; Morhardt, T.L.; Kamada, N. The Role of Dietary Nutrients in Inflammatory Bowel Disease. Front. Immunol. 2018, 9, 3183. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.; Xie, C.; Han, J.; Ye, Y.; Weiel, J.; Li, Q.; Blanco, I.; Ahn, C.; Olsen, N.; Putterman, C.; et al. Metabolic disturbances associated with systemic lupus erythematosus. PloS ONE 2012, 7, e37210. [Google Scholar]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





