Controlling the Switch from Neurogenesis to Pluripotency during Marmoset Monkey Somatic Cell Reprogramming with Self-Replicating mRNAs and Small Molecules
Abstract
1. Introduction
2. Materials and Methods
2.1. Isolation of Marmoset Fetal Fibroblasts (cjFFs)
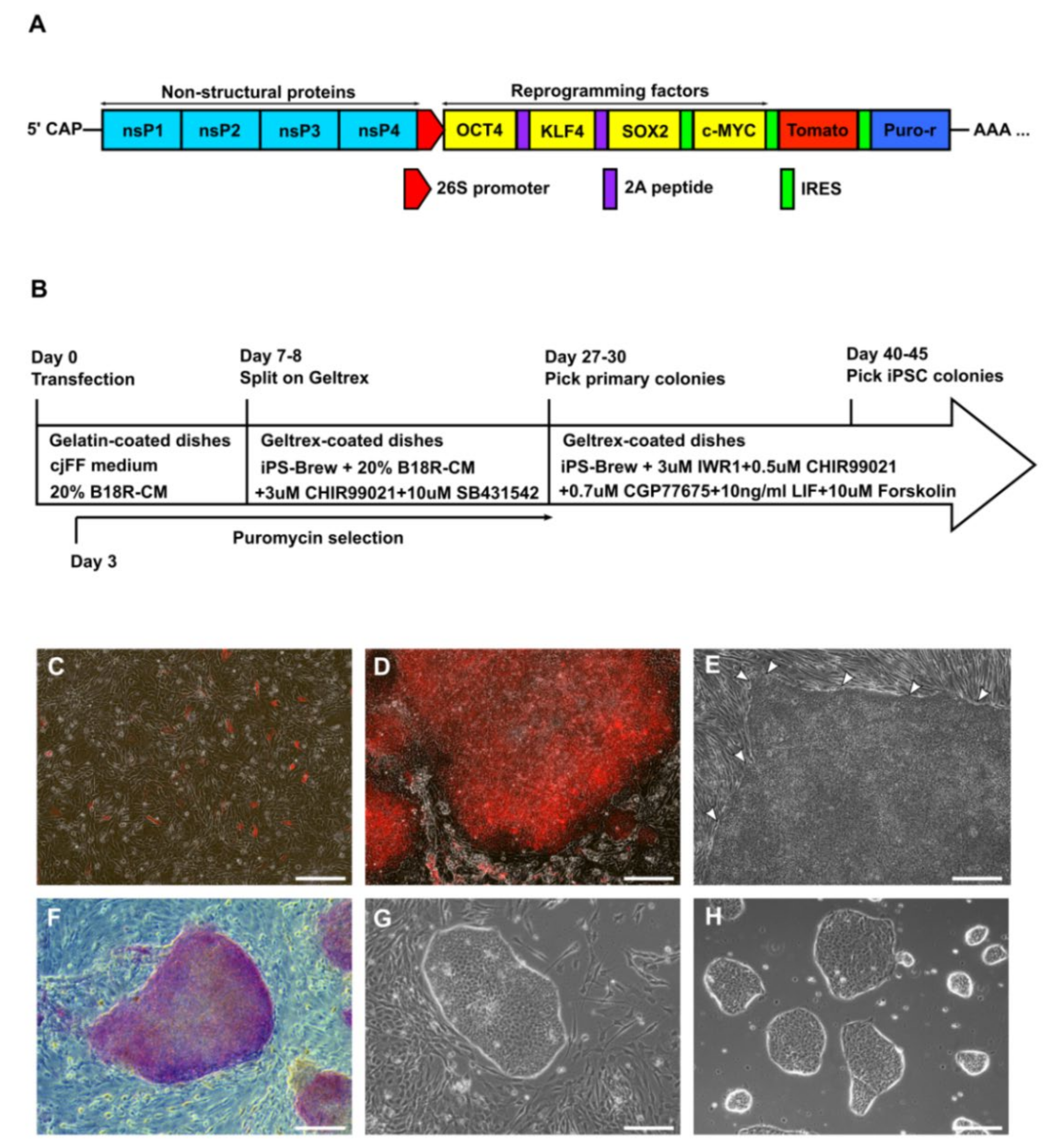
2.2. Synthesis of Self-Replicating mRNAs (VEE-OKS-iM-iTomato)
2.3. Production of B18R-Conditioned Medium (B18R-CM)
2.4. Electroporation of cjFFs with VEE-OKS-iM-iTomato and Generation of Intermediate Primary Colonies
2.5. Generation and Culture of Marmoset iPSCs
2.6. Directing Intermediate Primary Colonies into the Neural Lineages
2.7. Total RNA Isolation, PCR, RT-PCR, and Real-Time Relative Quantitation
2.8. Immunofluorescence and Alkaline Phosphatase Activity
2.9. In Vitro Differentiation of Marmoset iPSCs
2.10. Teratoma Assay
2.11. Karyotyping
3. Results
3.1. Derivation of Primary Colonies and iPSCs
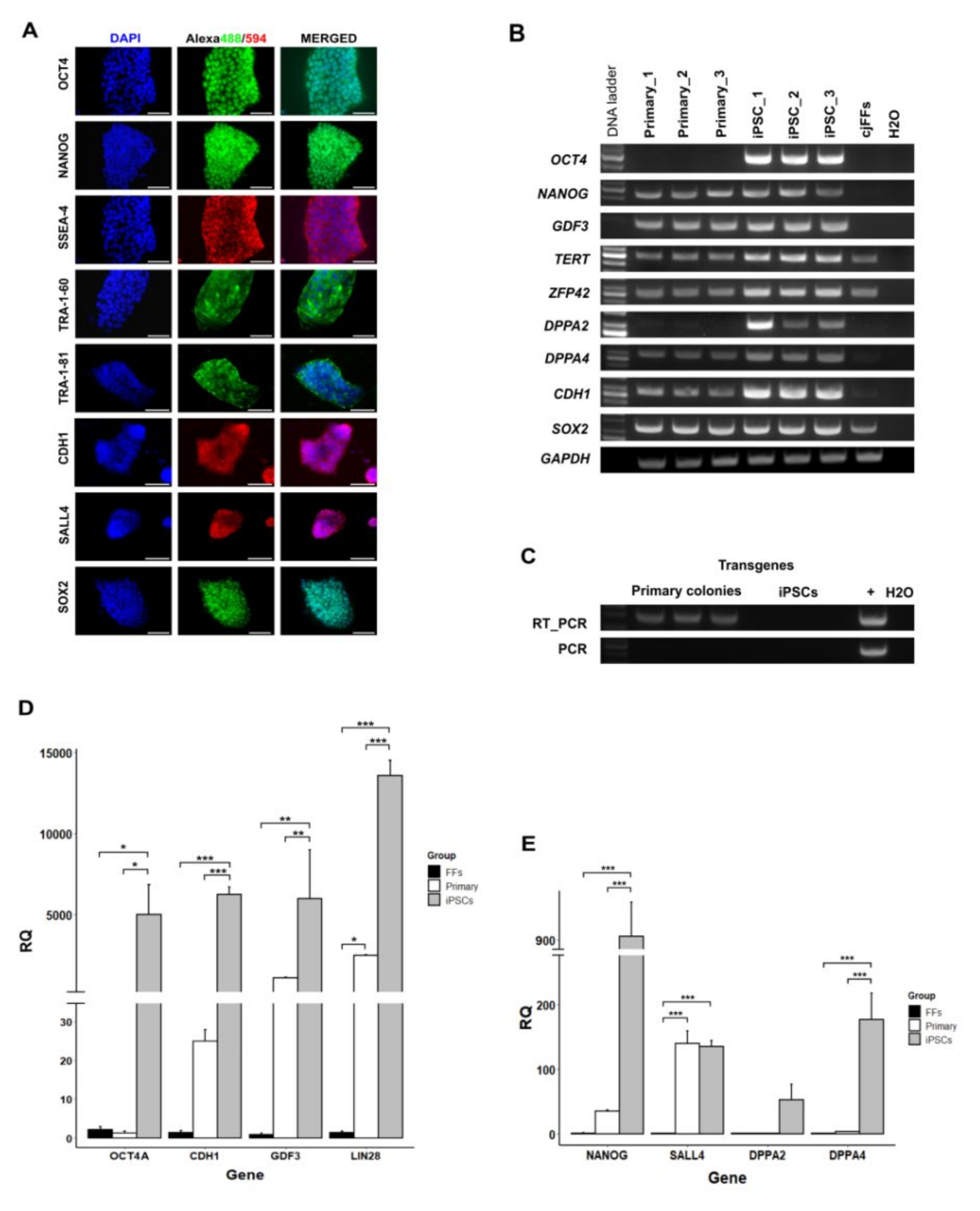
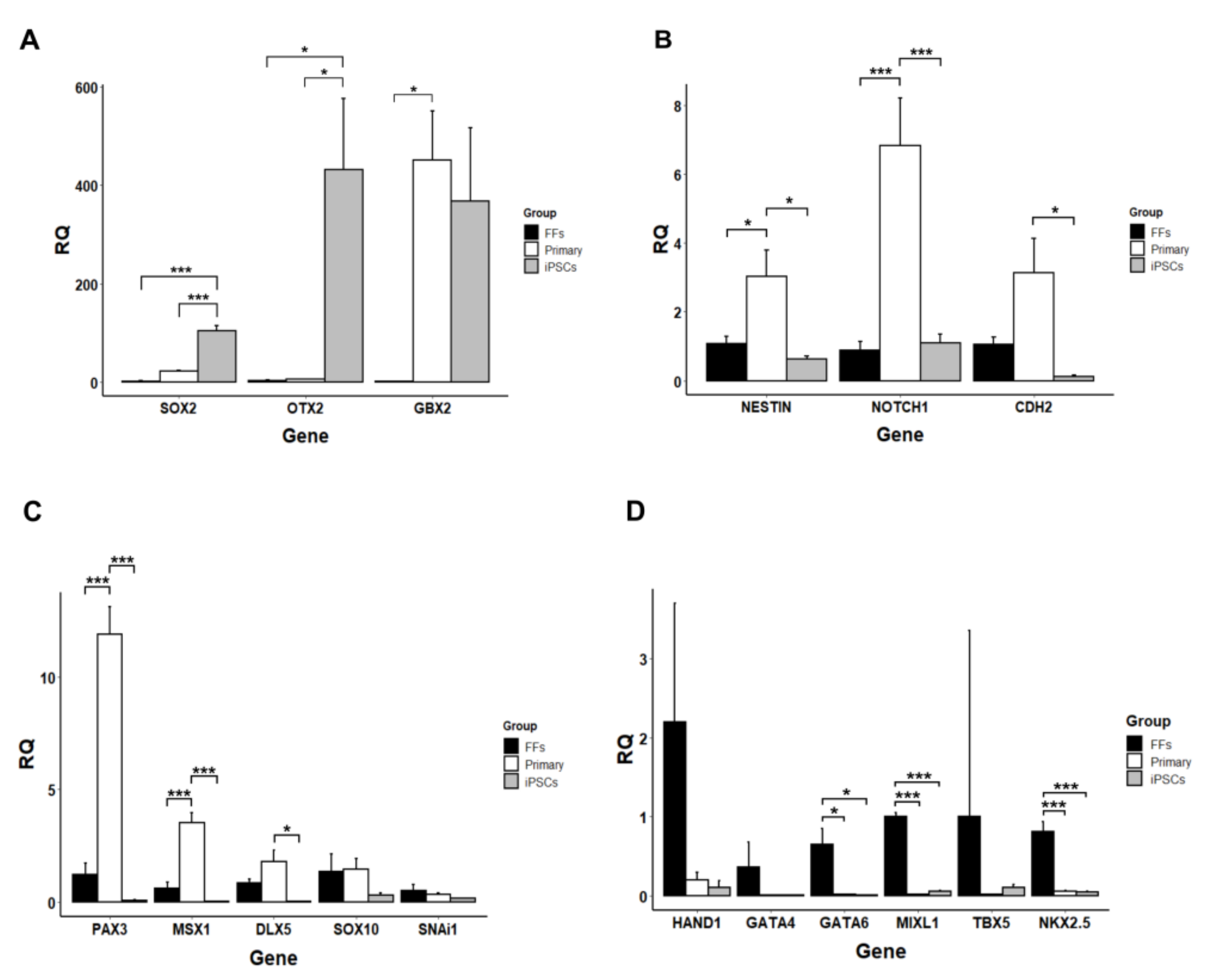
3.2. Gene Expression Analysis
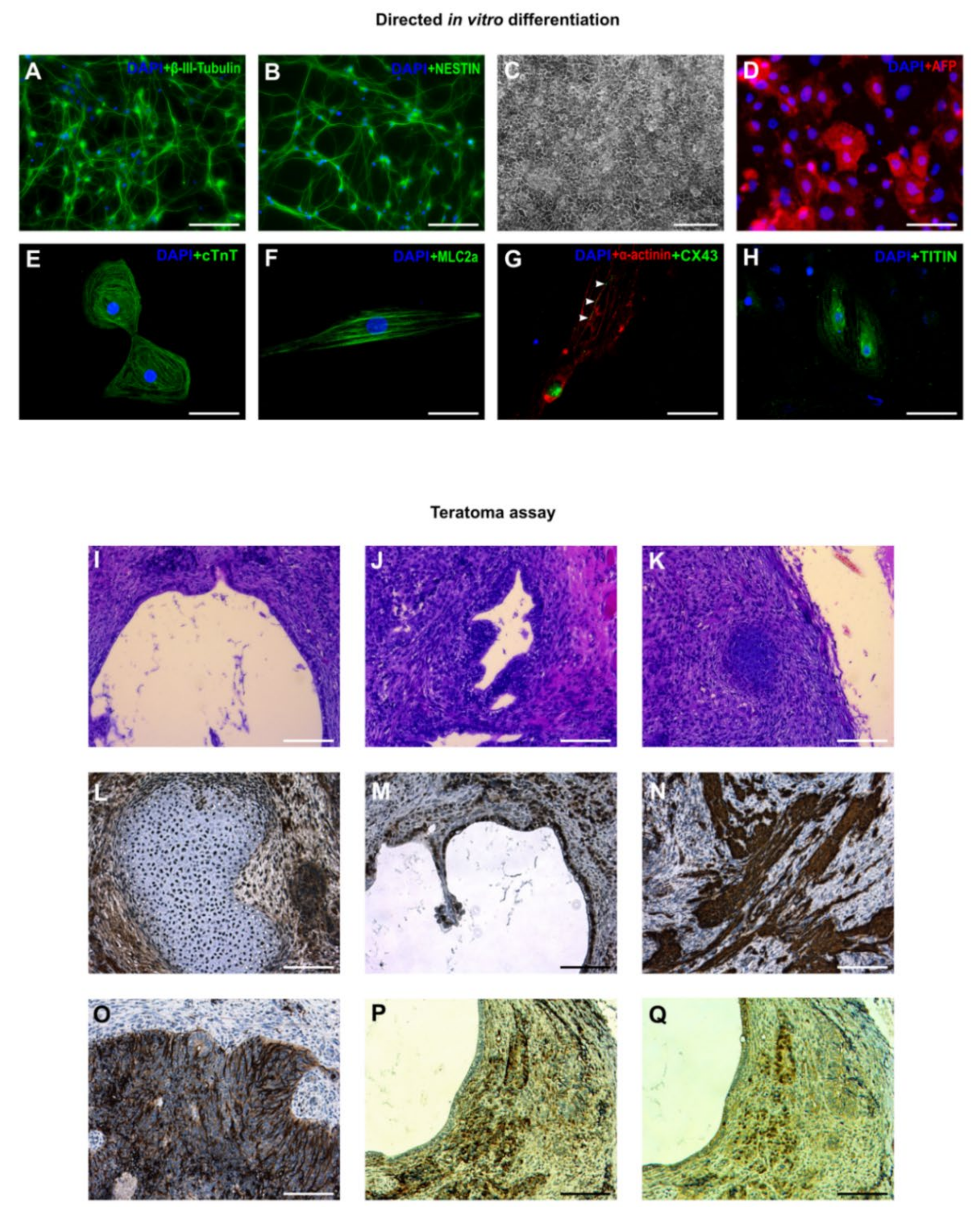
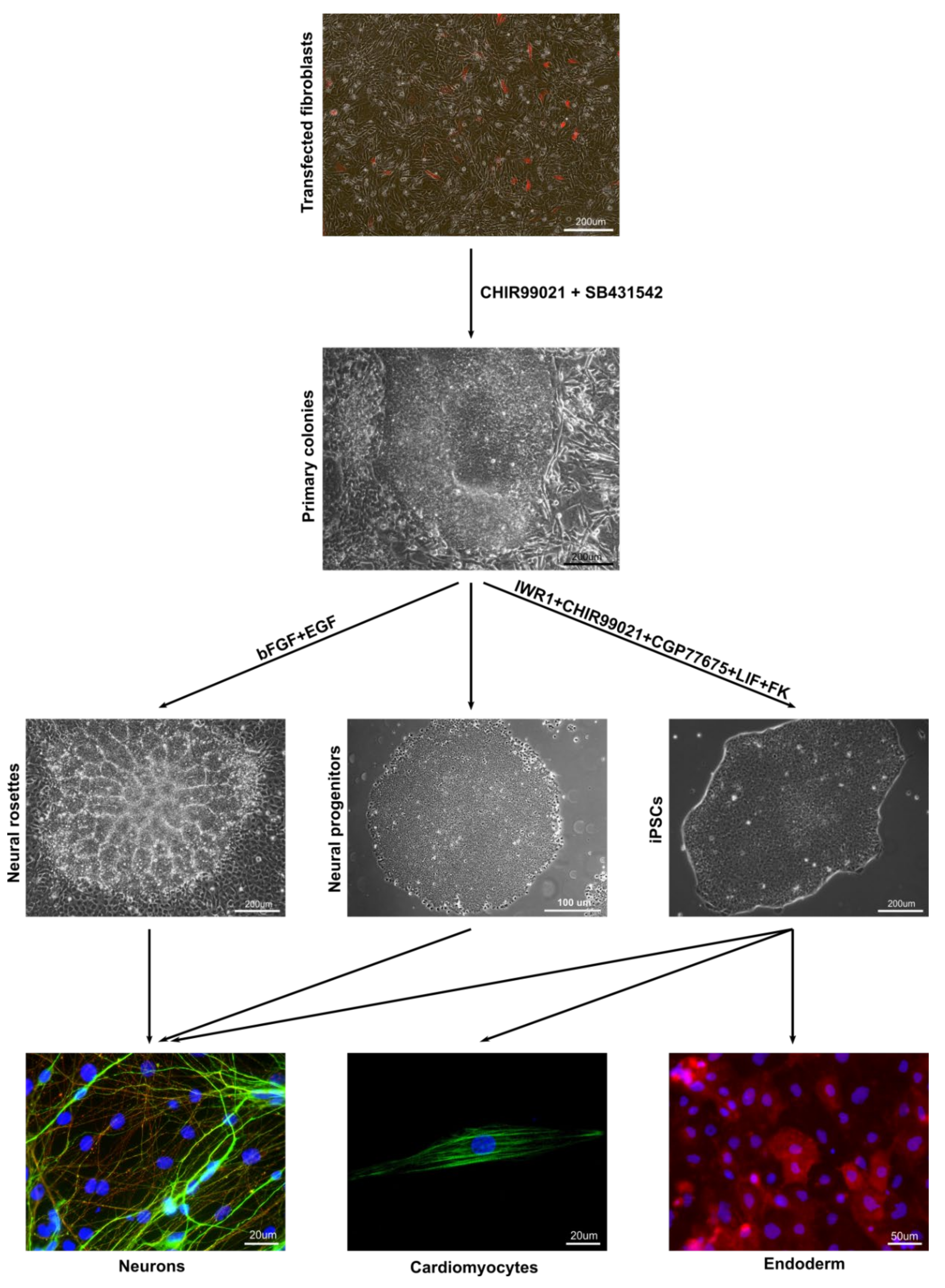
3.3. In Vitro and In Vivo Differentiation of Marmoset iPSCs
3.4. Karyotyping
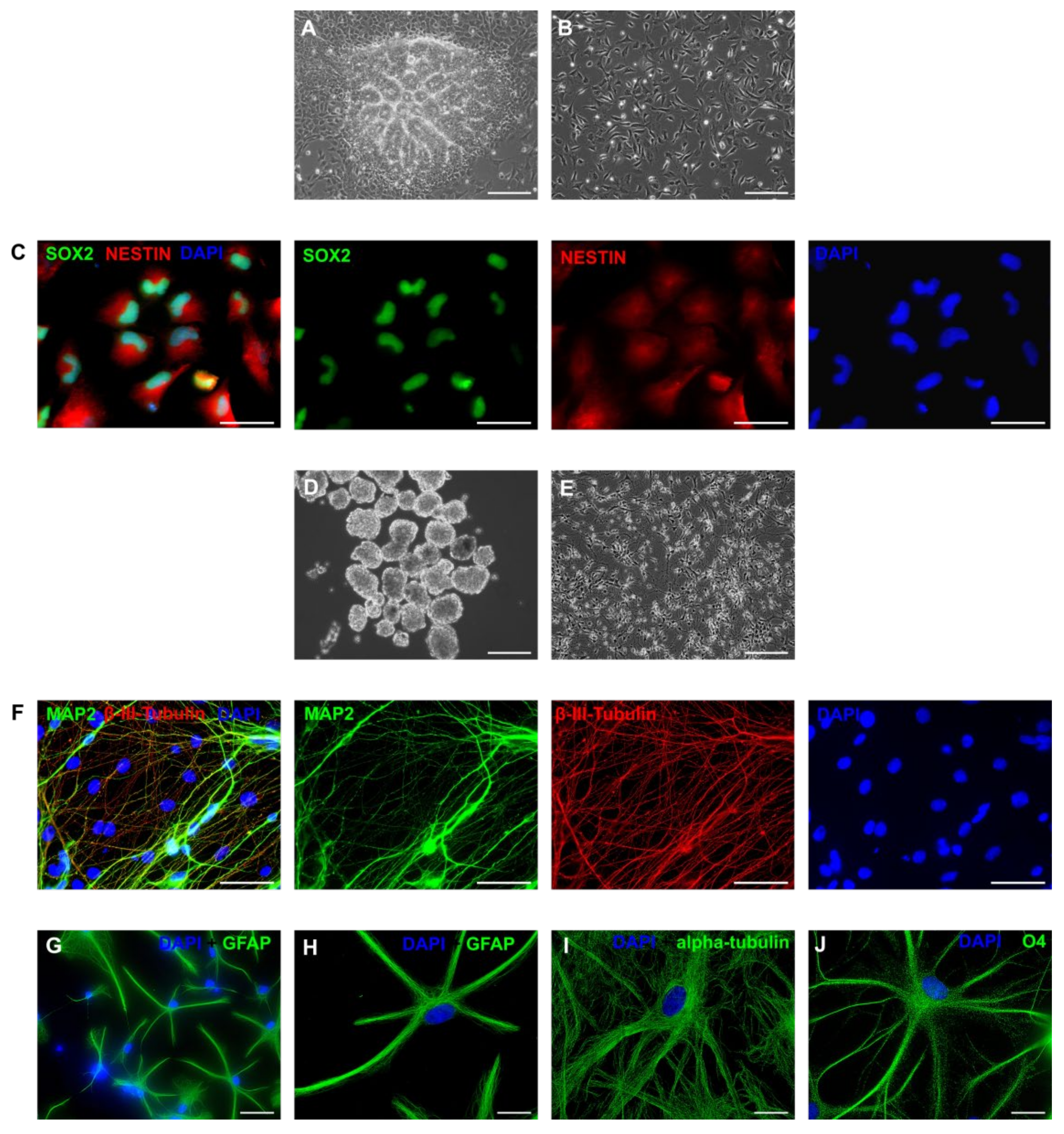
3.5. Neurogenic Potential of the Intermediate Primary Colonies
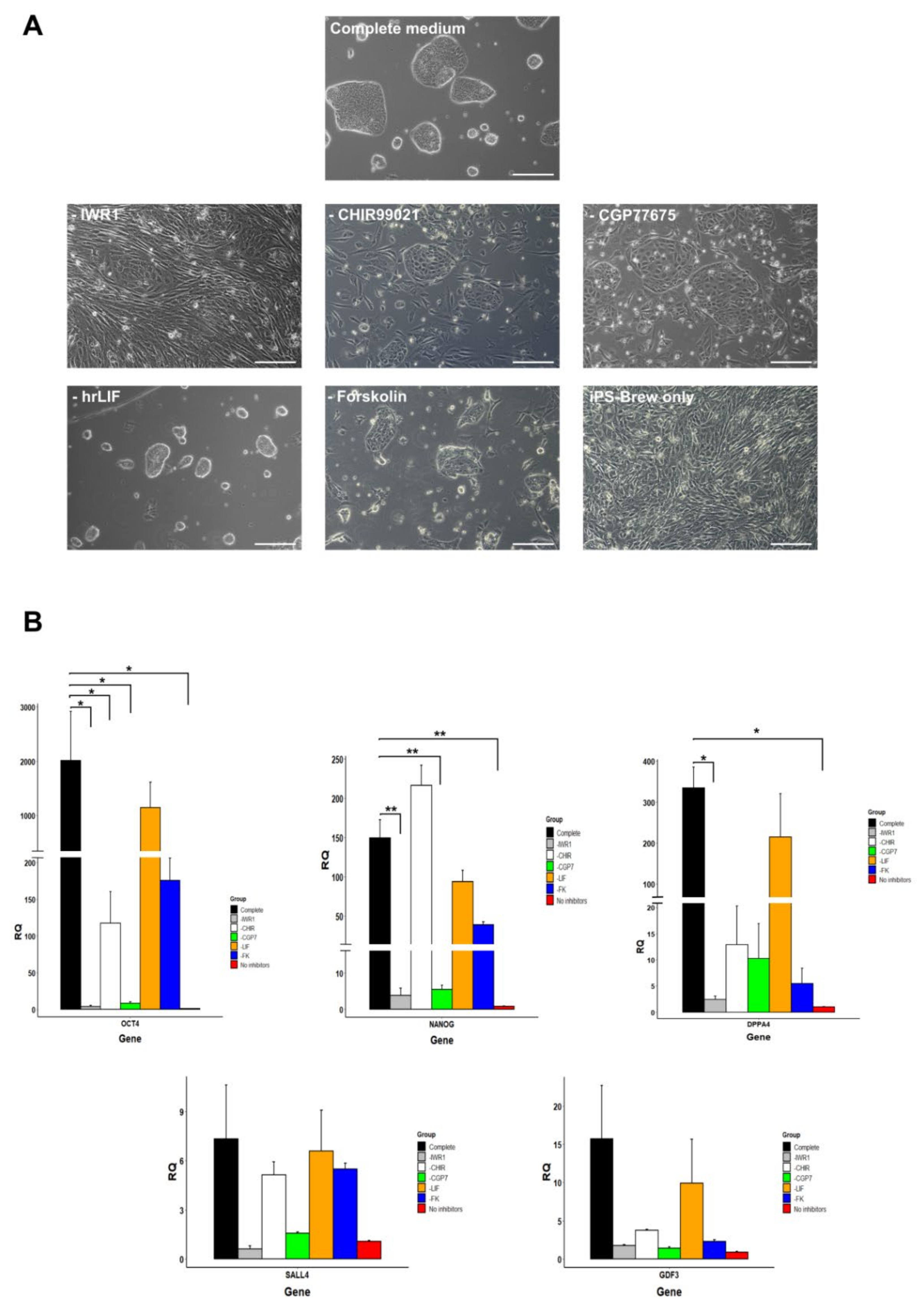
3.6. Small Molecule Inhibitors Are Necessary for Maintaining Pluripotency of Marmoset iPSCs
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Fleifel, D.; Rahmoon, M.A.; AlOkda, A.; Nasr, M.; Elserafy, M.; El-Khamisy, S.F. Recent advances in stem cells therapy: A focus on cancer, Parkinson’s and Alzheimer’s. J. Genet. Eng. Biotechnol. 2018, 16, 427–432. [Google Scholar] [CrossRef]
- Shroff, G. A review on stem cell therapy for multiple sclerosis: Special focus on human embryonic stem cells. Stem Cells Cloning Adv. Appl. 2018, 11, 1–11. [Google Scholar] [CrossRef]
- Solis, M.A.; Moreno Velásquez, I.; Correa, R.; Huang, L.L.H. Stem cells as a potential therapy for diabetes mellitus: A call-to-action in Latin America. Diabetol. Metab. Syndr. 2019, 11, 20. [Google Scholar] [CrossRef] [PubMed]
- Song, C.G.; Zhang, Y.Z.; Wu, H.N.; Cao, X.L.; Guo, C.J.; Li, Y.Q.; Zheng, M.H.; Han, H. Stem cells: A promising candidate to treat neurological disorders. Neural Regen. Res. 2018, 13, 1294–1304. [Google Scholar] [CrossRef] [PubMed]
- Okita, K.; Ichisaka, T.; Yamanaka, S. Generation of germline-competent induced pluripotent stem cells. Nature 2007, 448, 313–317. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Tanabe, K.; Ohnuki, M.; Narita, M.; Ichisaka, T.; Tomoda, K.; Yamanaka, S. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 2007, 131, 861–872. [Google Scholar] [CrossRef] [PubMed]
- Martin, U. Therapeutic Application of Pluripotent Stem Cells: Challenges and Risks. Front. Med. 2017, 4, 229. [Google Scholar] [CrossRef]
- Bauer, G.; Elsallab, M.; Abou-El-Enein, M. Concise Review: A Comprehensive Analysis of Reported Adverse Events in Patients Receiving Unproven Stem Cell-Based Interventions. Stem Cells Transl. Med. 2018, 7, 676–685. [Google Scholar] [CrossRef]
- Itakura, G.; Kawabata, S.; Ando, M.; Nishiyama, Y.; Sugai, K.; Ozaki, M.; Iida, T.; Ookubo, T.; Kojima, K.; Kashiwagi, R.; et al. Fail-Safe System against Potential Tumorigenicity after Transplantation of iPSC Derivatives. Stem Cell Rep. 2017, 8, 673–684. [Google Scholar] [CrossRef]
- Nakamura, M.; Okano, H. Cell transplantation therapies for spinal cord injury focusing on induced pluripotent stem cells. Cell Res. 2013, 23, 70–80. [Google Scholar] [CrossRef] [PubMed]
- Ross, C.N.; Davis, K.; Dobek, G.; Tardif, S.D. Aging Phenotypes of Common Marmosets (Callithrix jacchus). J. Aging Res. 2012, 2012, 567143. [Google Scholar] [CrossRef] [PubMed]
- Tardif, S.D.; Mansfield, K.G.; Ratnam, R.; Ross, C.N.; Ziegler, T.E. The marmoset as a model of aging and age-related diseases. Ilar J. 2011, 52, 54–65. [Google Scholar] [CrossRef]
- Yun, J.W.; Ahn, J.B.; Kang, B.C. Modeling Parkinson’s disease in the common marmoset (Callithrix jacchus): Overview of models, methods, and animal care. Lab. Anim. Res. 2015, 31, 155–165. [Google Scholar] [CrossRef]
- Senos, R.; Benedicto, H.G.; del Rio do Valle, C.M.; del Rio do Valle, R.; Nayudu, P.L.; Kfoury Junior, J.R.; Bombonato, P.P. Gross morphometry of the heart of the Common marmoset. Folia Morphol. 2014, 73, 37–41. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, E.; Suemizu, H.; Shimada, A.; Hanazawa, K.; Oiwa, R.; Kamioka, M.; Tomioka, I.; Sotomaru, Y.; Hirakawa, R.; Eto, T.; et al. Generation of transgenic non-human primates with germline transmission. Nature 2009, 459, 523–527. [Google Scholar] [CrossRef]
- Sato, K.; Oiwa, R.; Kumita, W.; Henry, R.; Sakuma, T.; Ito, R.; Nozu, R.; Inoue, T.; Katano, I.; Sato, K.; et al. Generation of a Nonhuman Primate Model of Severe Combined Immunodeficiency Using Highly Efficient Genome Editing. Cell Stem Cell 2016, 19, 127–138. [Google Scholar] [CrossRef]
- Shimozawa, A.; Ono, M.; Takahara, D.; Tarutani, A.; Imura, S.; Masuda-Suzukake, M.; Higuchi, M.; Yanai, K.; Hisanaga, S.I.; Hasegawa, M. Propagation of pathological α-synuclein in marmoset brain. Acta Neuropathol. Commun. 2017, 5, 12. [Google Scholar] [CrossRef]
- Tomioka, I.; Ishibashi, H.; Minakawa, E.N.; Motohashi, H.H.; Takayama, O.; Saito, Y.; Popiel, H.A.; Puentes, S.; Owari, K.; Nakatani, T.; et al. Transgenic Monkey Model of the Polyglutamine Diseases Recapitulating Progressive Neurological Symptoms. eNeuro 2017, 4. [Google Scholar] [CrossRef]
- Tomioka, I.; Maeda, T.; Shimada, H.; Kawai, K.; Okada, Y.; Igarashi, H.; Oiwa, R.; Iwasaki, T.; Aoki, M.; Kimura, T.; et al. Generating induced pluripotent stem cells from common marmoset (Callithrix jacchus) fetal liver cells using defined factors, including Lin28. Genes Cells Devoted Mol. Cell. Mech. 2010, 15, 959–969. [Google Scholar] [CrossRef]
- Wu, Y.; Zhang, Y.; Mishra, A.; Tardif, S.D.; Hornsby, P.J. Generation of induced pluripotent stem cells from newborn marmoset skin fibroblasts. Stem Cell Res. 2010, 4, 180–188. [Google Scholar] [CrossRef]
- Debowski, K.; Warthemann, R.; Lentes, J.; Salinas-Riester, G.; Dressel, R.; Langenstroth, D.; Gromoll, J.; Sasaki, E.; Behr, R. Non-viral generation of marmoset monkey iPS cells by a six-factor-in-one-vector approach. PLoS ONE 2015, 10, e0118424. [Google Scholar] [CrossRef]
- Vermilyea, S.C.; Guthrie, S.; Meyer, M.; Smuga-Otto, K.; Braun, K.; Howden, S.; Thomson, J.A.; Zhang, S.C.; Emborg, M.E.; Golos, T.G. Induced Pluripotent Stem Cell-Derived Dopaminergic Neurons from Adult Common Marmoset Fibroblasts. Stem Cells Dev. 2017, 26, 1225–1235. [Google Scholar] [CrossRef] [PubMed]
- Nakajima, M.; Yoshimatsu, S.; Sato, T.; Nakamura, M.; Okahara, J.; Sasaki, E.; Shiozawa, S.; Okano, H. Establishment of induced pluripotent stem cells from common marmoset fibroblasts by RNA-based reprogramming. Biochem. Biophys. Res. Commun. 2019, 515, 593–599. [Google Scholar] [CrossRef]
- Watanabe, T.; Yamazaki, S.; Yoneda, N.; Shinohara, H.; Tomioka, I.; Higuchi, Y.; Yagoto, M.; Ema, M.; Suemizu, H.; Kawai, K.; et al. Highly efficient induction of primate iPS cells by combining RNA transfection and chemical compounds. Genes Cells Devoted Mol. Cell. Mech. 2019, 24, 473–484. [Google Scholar] [CrossRef]
- Kinney, R.M.; Johnson, B.J.; Welch, J.B.; Tsuchiya, K.R.; Trent, D.W. The full-length nucleotide sequences of the virulent Trinidad donkey strain of Venezuelan equine encephalitis virus and its attenuated vaccine derivative, strain TC-83. Virology 1989, 170, 19–30. [Google Scholar] [CrossRef]
- Petrakova, O.; Volkova, E.; Gorchakov, R.; Paessler, S.; Kinney, R.M.; Frolov, I. Noncytopathic replication of Venezuelan equine encephalitis virus and eastern equine encephalitis virus replicons in Mammalian cells. J. Virol. 2005, 79, 7597–7608. [Google Scholar] [CrossRef]
- Yoshioka, N.; Gros, E.; Li, H.R.; Kumar, S.; Deacon, D.C.; Maron, C.; Muotri, A.R.; Chi, N.C.; Fu, X.D.; Yu, B.D.; et al. Efficient generation of human iPSCs by a synthetic self-replicative RNA. Cell Stem Cell 2013, 13, 246–254. [Google Scholar] [CrossRef]
- Tiburcy, M.; Hudson, J.E.; Balfanz, P.; Schlick, S.; Meyer, T.; Chang Liao, M.L.; Levent, E.; Raad, F.; Zeidler, S.; Wingender, E.; et al. Defined Engineered Human Myocardium With Advanced Maturation for Applications in Heart Failure Modeling and Repair. Circulation 2017, 135, 1832–1847. [Google Scholar] [CrossRef]
- Petkov, S.; Kahland, T.; Shomroni, O.; Lingner, T.; Salinas, G.; Fuchs, S.; Debowski, K.; Behr, R. Immortalization of common marmoset monkey fibroblasts by piggyBac transposition of hTERT. PLoS ONE 2018, 13, e0204580. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Hu, K.; Smuga-Otto, K.; Tian, S.; Stewart, R.; Slukvin, I.I.; Thomson, J.A. Human induced pluripotent stem cells free of vector and transgene sequences. Science 2009, 324, 797–801. [Google Scholar] [CrossRef] [PubMed]
- Okita, K.; Matsumura, Y.; Sato, Y.; Okada, A.; Morizane, A.; Okamoto, S.; Hong, H.; Nakagawa, M.; Tanabe, K.; Tezuka, K.; et al. A more efficient method to generate integration-free human iPS cells. Nat. Methods 2011, 8, 409–412. [Google Scholar] [CrossRef] [PubMed]
- Schlaeger, T.M.; Daheron, L.; Brickler, T.R.; Entwisle, S.; Chan, K.; Cianci, A.; DeVine, A.; Ettenger, A.; Fitzgerald, K.; Godfrey, M.; et al. A comparison of non-integrating reprogramming methods. Nat. Biotechnol. 2015, 33, 58–63. [Google Scholar] [CrossRef]
- Tidball, A.M.; Neely, M.D.; Chamberlin, R.; Aboud, A.A.; Kumar, K.K.; Han, B.; Bryan, M.R.; Aschner, M.; Ess, K.C.; Bowman, A.B. Genomic Instability Associated with p53 Knockdown in the Generation of Huntington’s Disease Human Induced Pluripotent Stem Cells. PLoS ONE 2016, 11, e0150372. [Google Scholar] [CrossRef]
- Li, W.; Zhou, H.; Abujarour, R.; Zhu, S.; Young Joo, J.; Lin, T.; Hao, E.; Schöler, H.R.; Hayek, A.; Ding, S. Generation of human-induced pluripotent stem cells in the absence of exogenous Sox2. Stem Cells (Dayt. Ohio) 2009, 27, 2992–3000. [Google Scholar] [CrossRef]
- Lin, T.; Ambasudhan, R.; Yuan, X.; Li, W.; Hilcove, S.; Abujarour, R.; Lin, X.; Hahm, H.S.; Hao, E.; Hayek, A.; et al. A chemical platform for improved induction of human iPSCs. Nat. Methods 2009, 6, 805–808. [Google Scholar] [CrossRef] [PubMed]
- Zhu, S.; Li, W.; Zhou, H.; Wei, W.; Ambasudhan, R.; Lin, T.; Kim, J.; Zhang, K.; Ding, S. Reprogramming of human primary somatic cells by OCT4 and chemical compounds. Cell Stem Cell 2010, 7, 651–655. [Google Scholar] [CrossRef]
- Lu, J.; Liu, H.; Huang, C.T.; Chen, H.; Du, Z.; Liu, Y.; Sherafat, M.A.; Zhang, S.C. Generation of integration-free and region-specific neural progenitors from primate fibroblasts. Cell Rep. 2013, 3, 1580–1591. [Google Scholar] [CrossRef]
- Li, W.; Sun, W.; Zhang, Y.; Wei, W.; Ambasudhan, R.; Xia, P.; Talantova, M.; Lin, T.; Kim, J.; Wang, X.; et al. Rapid induction and long-term self-renewal of primitive neural precursors from human embryonic stem cells by small molecule inhibitors. Proc. Natl. Acad. Sci. USA 2011, 108, 8299–8304. [Google Scholar] [CrossRef]
- Denham, M.; Hasegawa, K.; Menheniott, T.; Rollo, B.; Zhang, D.; Hough, S.; Alshawaf, A.; Febbraro, F.; Ighaniyan, S.; Leung, J.; et al. Multipotent caudal neural progenitors derived from human pluripotent stem cells that give rise to lineages of the central and peripheral nervous system. Stem Cells 2015, 33, 1759–1770. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.B.; Sebastiano, V.; Wu, G.; Araúzo-Bravo, M.J.; Sasse, P.; Gentile, L.; Ko, K.; Ruau, D.; Ehrich, M.; van den Boom, D.; et al. Oct4-induced pluripotency in adult neural stem cells. Cell 2009, 136, 411–419. [Google Scholar] [CrossRef]
- Moon, J.H.; Heo, J.S.; Kwon, S.; Kim, J.; Hwang, J.; Kang, P.J.; Kim, A.; Kim, H.O.; Whang, K.Y.; Yoon, B.S.; et al. Two-step generation of induced pluripotent stem cells from mouse fibroblasts using Id3 and Oct4. J. Mol. Cell Biol. 2012, 4, 59–62. [Google Scholar] [CrossRef] [PubMed]
- Jho, E.H.; Zhang, T.; Domon, C.; Joo, C.K.; Freund, J.N.; Costantini, F. Wnt/beta-catenin/Tcf signaling induces the transcription of Axin2, a negative regulator of the signaling pathway. Mol. Cell. Biol. 2002, 22, 1172–1183. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Chadarevian, J.P.; Ruiz, B.; Ying, Q.L. Cytoplasmic and Nuclear TAZ Exert Distinct Functions in Regulating Primed Pluripotency. Stem Cell Rep. 2017, 9, 732–741. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.; Huebner, A.J.; Clement, K.; Walsh, R.M.; Savol, A.; Lin, K.; Gu, H.; Di Stefano, B.; Brumbaugh, J.; Kim, S.Y.; et al. Prolonged Mek1/2 suppression impairs the developmental potential of embryonic stem cells. Nature 2017, 548, 219–223. [Google Scholar] [CrossRef]
- Gao, X.; Nowak-Imialek, M.; Chen, X.; Chen, D.; Herrmann, D.; Ruan, D.; Chen, A.C.H.; Eckersley-Maslin, M.A.; Ahmad, S.; Lee, Y.L.; et al. Establishment of porcine and human expanded potential stem cells. Nat. Cell Biol. 2019, 21, 687–699. [Google Scholar] [CrossRef]
- Stauske, M.; Rodriguez Polo, I.; Haas, W.; Knorr, D.Y.; Borchert, T.; Streckfuss-Bömeke, K.; Dressel, R.; Bartels, I.; Tiburcy, M.; Zimmermann, W.H.; et al. Non-Human Primate iPSC Generation, Cultivation, and Cardiac Differentiation under Chemically Defined Conditions. Cells 2020, 9, 1349. [Google Scholar] [CrossRef]
- Acampora, D.; Di Giovannantonio, L.G.; Simeone, A. Otx2 is an intrinsic determinant of the embryonic stem cell state and is required for transition to a stable epiblast stem cell condition. Development 2013, 140, 43–55. [Google Scholar] [CrossRef]
- Tai, C.I.; Ying, Q.L. Gbx2, a LIF/Stat3 target, promotes reprogramming to and retention of the pluripotent ground state. J. Cell Sci. 2013, 126, 1093–1098. [Google Scholar] [CrossRef]
- Ma, H.; Zhai, J.; Wan, H.; Jiang, X.; Wang, X.; Wang, L.; Xiang, Y.; He, X.; Zhao, Z.A.; Zhao, B.; et al. In vitro culture of cynomolgus monkey embryos beyond early gastrulation. Science 2019, 366. [Google Scholar] [CrossRef]
- Niu, Y.; Sun, N.; Li, C.; Lei, Y.; Huang, Z.; Wu, J.; Si, C.; Dai, X.; Liu, C.; Wei, J.; et al. Dissecting primate early post-implantation development using long-term in vitro embryo culture. Science 2019, 366. [Google Scholar] [CrossRef]
| Primers for RT-PCR | |||
|---|---|---|---|
| Gene | Accession Code | Primer Sequences | Amplicon (bp) |
| OCT4A | ENSCJAT00000038869.2 | Fw: GGC TTG GGG CGC CTT CCT TC | |
| Rv: CAG GGT GAT CCT CTT CTG CTT C | 503 | ||
| NANOG | XM_002752302.3 | Fw: GCC ACC TGA AGA TGT GTG AAG ATG AAT G | |
| Rv: GGG TAG GCA TAA TGT AAA CAG AAC ACG | 149 | ||
| ZFP42 | XM_003732392.3 | Fw: GAAACCACGTCTGTGCAGAGTGTG | |
| Rv: GCATGAGTTAGGATGTGGGCTTTCAG | 256 | ||
| GDF3 | XM_002752299.3 | Fw: GCTGGATGTAGCTAAGGATTGGAATG | |
| Rv: GAAGTTAATGAATAGCTGGTGACGGTG | 271 | ||
| DPPA2 | XM_008982501.2 | Fw: CACTTTGCGGAACTGGTGTCAAG | |
| Rv: CAGTCTTAGGCTGAACAGCTCTG | 311 | ||
| DPPA4 | XM_002758824 | Fw: GTCCATGGGAAAAGTCTCCCTGCAG | |
| Rv: CACCACGGAATCCGACTCTCCAG | 287 | ||
| SALL4 | XM_002747676 | Fw: CTACTGACAGCGTTCCCAAACACCAG | |
| Rv: CAGCATAGCAACAATCGTGATTGT | 252 | ||
| TERT | ENSCJAT00000074768.1 | Fw: CTGCTCCTGCGTTTGGTGGATG | |
| Rv: CGTCTGGAGGCTGTTCACCTG | 405 | ||
| SOX2 | ENSCJAT00000016325.3 | Fw: GCTCGCAGACCTACATGAACG | |
| Rv: GCGGTCCGGCCCTCACAT | 332 | ||
| NESTIN | XM_008984607.2 | Fw: GCGTTGGAACAGAGGTTGGAG | |
| Rv: GTCTCAAGGGTAGCAGGCAAG | 377 | ||
| NOTCH1 | XM_017963672.1 | Fw: CTTACAGATGCAGCCACAGAACCTG | |
| Rv: GCGGGCAATCTGAGACTGCATG | 491 | ||
| OTX2 | ENSCJAT00000036430.2 | Fw: GGGCTGAGTCTGACCACTTCG | |
| Rv: GAGATGGCTGGTGACTGCATTG | 657 | ||
| GAPDH | XM_017976537.1 | Fw: CACTGGCGTCTTCACCACCATG | |
| Rv: GACACGGAAAGCCATGCCAGTG | 305 | ||
| Primers for real-time qPCR | |||
| Gene | Accession code | Primer sequences | Amplicon (bp) |
| OCT4A | ENSCJAT00000038869.2 | Fw: GGCGCCTTCCTTCCCCATGG | |
| Rv: GGCGAGAAGGCAAAATCCGAAG | 54 | ||
| NANOG | XM_002752302.3 | Fw: GCCACCTGAAGATGTGTGAAGATGAATG | |
| Rv: GGATTCAGCCAGTGCTCAGAGTG | 71 | ||
| ZFP42 | XM_003732392.3 | Fw: GTGTCCCTTTGAAGGCTGTAGGAAG | |
| Rv: GCATGAGTTAGGATGTGGGCTTTCAG | 72 | ||
| GDF3 | XM_002752299.3 | Fw: GCTGGATGTAGCTAAGGATTGGAATG | |
| Rv:CAGTATCTCCAGGAATAGCCCGAAG | 67 | ||
| DPPA2 | XM_008982501.2 | Fw: CACTTTGCGGAACTGGTGTCAAG | |
| Rv: CAGGCATATCTTGCTGTTGTTCAGG | 113 | ||
| DPPA4 | XM_002758824 | Fw: GTCCATGGGAAAAGTCTCCCTGCAG | |
| Rv: GAACCCAGGCCTGACCAGCATG | 76 | ||
| SALL4 | XM_002747676 | Fw: CTACTGACAGCGTTCCCAAACACCAG | |
| Rv:GCACGTTCTCCTTTAGCTTAGCTG | 89 | ||
| SOX2 | ENSCJAT00000016325.3 | Fw: GCTCGCAGACCTACATGAACG | |
| Rv: GACTTGACCACCGAGCCCATG | 103 | ||
| NESTIN | XM_008984607.2 | Fw: GCGTTGGAACAGAGGTTGGAG | |
| Rv: GACATCTTGAGGTGCGCCAG | 163 | ||
| NOTCH1 | XM_017963672.1 | Fw: CATTCCAACGTCTCTGACTGGTCTG | |
| Rv: GCGGGCAATCTGAGACTGCATG | 77 | ||
| OTX2 | ENSCJAT00000036430.2 | Fw: CCAACCATTGCCAGCAGCAGTG | |
| Rv: GAAGAGGAGGTGGACAAGGGATCTG | 89 | ||
| GBX2 | XM_017973818 | Fw: GAGTAGCACCGCCTTCAGCATAG | |
| Rv: GGGTAGCCGGTGTAGACGAAATGG | 83 | ||
| PAX3 | XM_008999566 | Fw: GCTGTCGGAGACCTCTTACCAG | |
| Rv: GAACCGTGCTGCTGGGATCCG | 68 | ||
| MSX1 | XM_003732530 | Fw: GACGAACCGTAAGCCACGGACG | |
| Rv: GCTCGGCGATGGACAGGTACTG | 95 | ||
| DLX5 | XM_017975343 | Fw: GCTCTCCTACCTCGGCTTCCTATG | |
| Rv: GCCGTTCACGCCGTGATACTG | 77 | ||
| SOX10 | XM_017963624 | Fw: GACATCGGTGAGATCAGCCACGAG | |
| Rv: CGGCAGGTACTGGTCCAACTCAG | 80 | ||
| SNAi1 | XM_003732844 | Fw: CTTTGCCGACCGCTCCAACCTG | |
| Rv: GGAGCAGGGACATTCGGGAGAAG | 107 | ||
| 18S | XM_002746395 | Fw: ATTAAGGGTGTGGGCCGAAG | |
| Rv: GAGTTCTCCTGCCCTCTTGG | 81 | ||
| PCR primers for detection of the T7-VEE-OKS-iM-iTomato plasmid DNA | |||
| VEE_OKSiM_F1: CAC CAC TCT GGG CTC TCC CAT G | |||
| VEE_OKSiM_R1: GTC CAG GTC CAG GAG ATC GTT G 388 | |||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Petkov, S.; Dressel, R.; Rodriguez-Polo, I.; Behr, R. Controlling the Switch from Neurogenesis to Pluripotency during Marmoset Monkey Somatic Cell Reprogramming with Self-Replicating mRNAs and Small Molecules. Cells 2020, 9, 2422. https://doi.org/10.3390/cells9112422
Petkov S, Dressel R, Rodriguez-Polo I, Behr R. Controlling the Switch from Neurogenesis to Pluripotency during Marmoset Monkey Somatic Cell Reprogramming with Self-Replicating mRNAs and Small Molecules. Cells. 2020; 9(11):2422. https://doi.org/10.3390/cells9112422
Chicago/Turabian StylePetkov, Stoyan, Ralf Dressel, Ignacio Rodriguez-Polo, and Rüdiger Behr. 2020. "Controlling the Switch from Neurogenesis to Pluripotency during Marmoset Monkey Somatic Cell Reprogramming with Self-Replicating mRNAs and Small Molecules" Cells 9, no. 11: 2422. https://doi.org/10.3390/cells9112422
APA StylePetkov, S., Dressel, R., Rodriguez-Polo, I., & Behr, R. (2020). Controlling the Switch from Neurogenesis to Pluripotency during Marmoset Monkey Somatic Cell Reprogramming with Self-Replicating mRNAs and Small Molecules. Cells, 9(11), 2422. https://doi.org/10.3390/cells9112422






