Adaptation of Proteasomes and Lysosomes to Cellular Environments
Abstract
1. Background
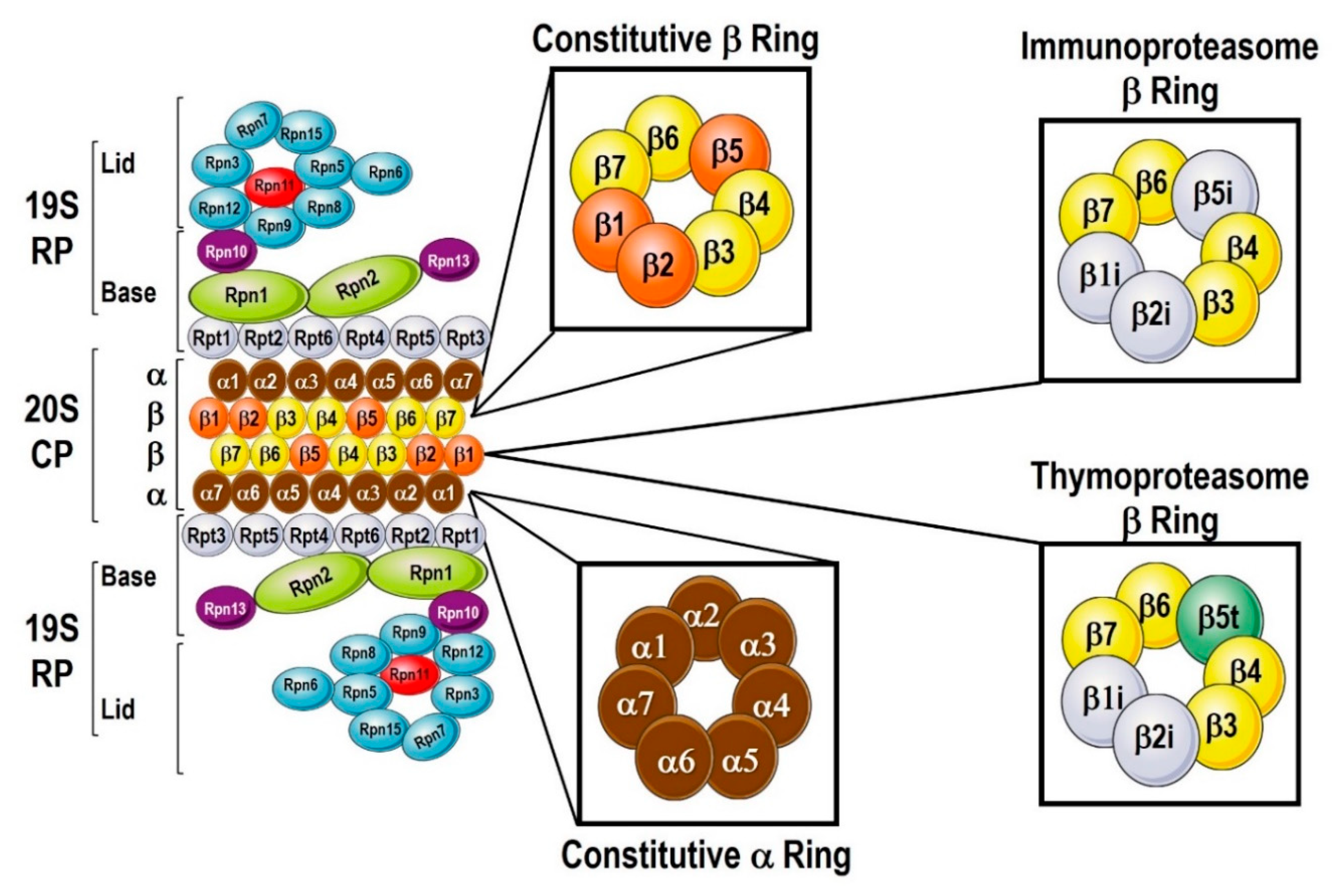
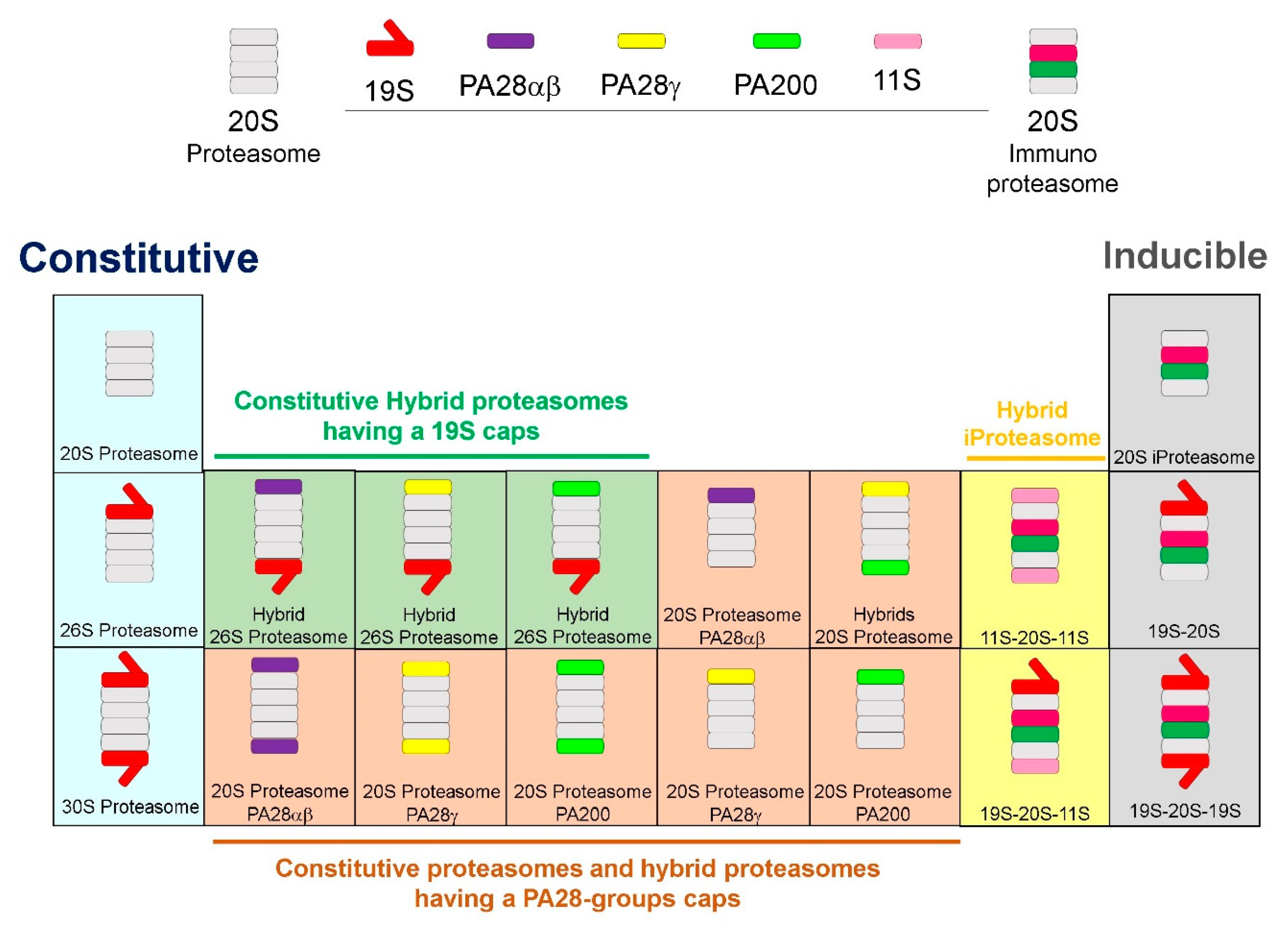
2. Proteasome Plasticity
3. Recognition of Proteins by the Proteasome
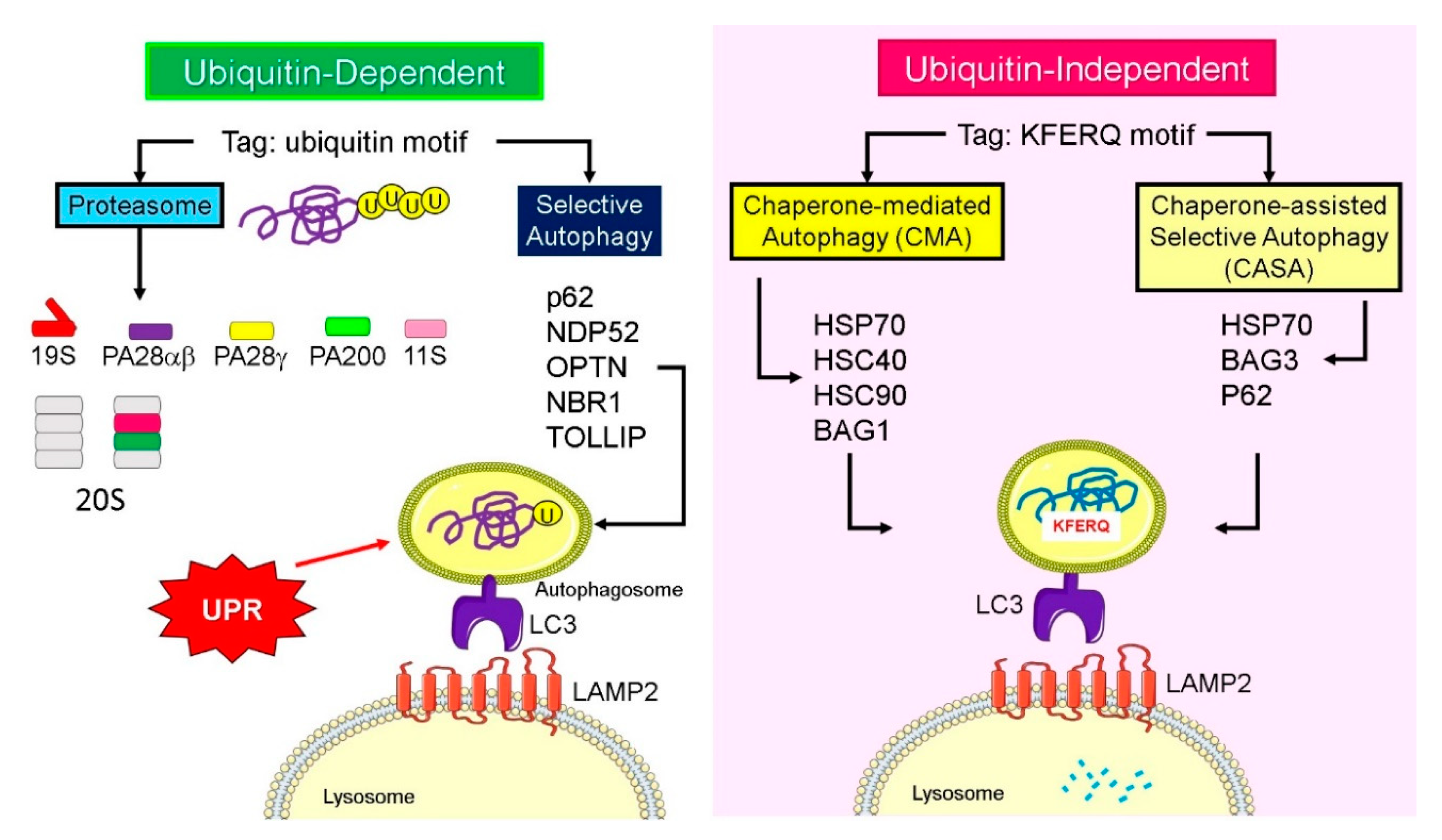
4. Lysosomal Degradation and Recognition of Ubiquitylated Proteins
5. Ubiquitin-Like Proteins and Adaptors for Selective Autophagy
6. Signals That Determine Proteasomal and Lysosomal Degradation
7. Concluding Remarks
Funding
Conflicts of Interest
References
- Dong, Z.; Cui, H. The Autophagy-Lysosomal Pathways and Their Emerging Roles in Modulating Proteostasis in Tumors. Cells 2018, 8, 4. [Google Scholar] [CrossRef] [PubMed]
- Clague, M.J.; Urbé, S. Ubiquitin: Same Molecule, Different Degradation Pathways. Cell 2010, 143, 682–685. [Google Scholar] [CrossRef] [PubMed]
- Kim, E.; Park, S.; Lee, J.H.; Mun, J.Y.; Choi, W.H.; Yun, Y.; Lee, J.; Kim, J.H.; Kang, M.-J.; Lee, M.J. Dual Function of USP14 Deubiquitinase in Cellular Proteasomal Activity and Autophagic Flux. Cell Rep. 2018, 24, 732–743. [Google Scholar] [CrossRef] [PubMed]
- Gan, J.; Leestemaker, Y.; Sapmaz, A.; Ovaa, H. Highlighting the Proteasome: Using Fluorescence to Visualize Proteasome Activity and Distribution. Front. Mol. Biosci. 2019, 6, 14. [Google Scholar] [CrossRef]
- Morozov, A.V.; Karpov, V.L. Proteasomes and Several Aspects of Their Heterogeneity Relevant to Cancer. Front. Oncol. 2019, 9, 761. [Google Scholar] [CrossRef]
- Tanaka, K. The proteasome: Overview of structure and functions. Proc. Jpn. Acad. Ser. B 2009, 85, 12–36. [Google Scholar] [CrossRef]
- Livneh, I.; Cohen-Kaplan, V.; Cohen-Rosenzweig, C.; Avni, N.; Ciechanover, A. The life cycle of the 26S proteasome: From birth, through regulation and function, and onto its death. Cell Res. 2016, 26, 869–885. [Google Scholar] [CrossRef]
- Saeki, Y. Ubiquitin recognition by the proteasome. J. Biochem. 2017, 161, 113–124. [Google Scholar] [CrossRef]
- Hoeller, D.; Dikic, I. How the proteasome is degraded. Proc. Natl. Acad. Sci. USA 2016, 113, 13266–13268. [Google Scholar] [CrossRef]
- Koulich, E.; Li, X.; DeMartino, G.N. Relative Structural and Functional Roles of Multiple Deubiquitylating Proteins Associated with Mammalian 26S Proteasome. Mol. Boil. Cell 2008, 19, 1072–1082. [Google Scholar] [CrossRef]
- Finley, D. Recognition and processing of ubiquitin-protein conjugates by the proteasome. Annu. Rev. Biochem. 2009, 78, 477–513. [Google Scholar] [CrossRef] [PubMed]
- Krzywda, S.; Brzozowski, A.M.; Higashitsuji, H.; Fujita, J.; Welchman, R.; Dawson, S.; Mayer, R.J.; Wilkinson, A.J. The Crystal Structure of Gankyrin, an Oncoprotein Found in Complexes with Cyclin-dependent Kinase 4, a 19 S Proteasomal ATPase Regulator, and the Tumor Suppressors Rb and p53. J. Boil. Chem. 2003, 279, 1541–1545. [Google Scholar] [CrossRef] [PubMed]
- Kaneko, T.; Hamazaki, J.; Iemura, S.-I.; Sasaki, K.; Furuyama, K.; Natsume, T.; Tanaka, K.; Murata, S. Assembly Pathway of the Mammalian Proteasome Base Subcomplex Is Mediated by Multiple Specific Chaperones. Cell 2009, 137, 914–925. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Wu, J.; Dong, Y.; Chen, S.; Sun, S.; Ma, Y.-B.; Ouyang, Q.; Finley, D.; Kirschner, M.W.; Mao, Y.J. Conformational Landscape of the p28-Bound Human Proteasome Regulatory Particle. Mol. Cell 2017, 67, 322–333.e6. [Google Scholar] [CrossRef] [PubMed]
- Livnat-Levanon, N.; Kevei, E.; Kleifeld, O.; Krutauz, D.; Segref, A.; Rinaldi, T.; Erpapazoglou, Z.; Cohen, M.M.; Reis, N.; Hoppe, T.; et al. Reversible 26S Proteasome Disassembly upon Mitochondrial Stress. Cell Rep. 2014, 7, 1371–1380. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Chemmama, I.E.; Yu, C.; Huszagh, A.; Xu, Y.; Viner, R.; Block, S.A.; Cimermancic, P.; Rychnovsky, S.D.; Ye, Y.; et al. The proteasome-interacting Ecm29 protein disassembles the 26S proteasome in response to oxidative stress. J. Boil. Chem. 2017, 292, 16310–16320. [Google Scholar] [CrossRef]
- Pickering, A.M.; Davies, K.J. Differential roles of proteasome and immunoproteasome regulators Pa28alphabeta, Pa28gamma and Pa200 in the degradation of oxidized proteins. Arch. Biochem. Biophys. 2012, 523, 181–190. [Google Scholar] [CrossRef] [PubMed]
- Yewdell, J.W. Immunoproteasomes: Regulating the regulator. Proc. Natl. Acad. Sci. USA 2005, 102, 9089–9090. [Google Scholar] [CrossRef]
- Ferrington, D.A.; Gregerson, D.S. Immunoproteasomes: Structure, function, and antigen presentation. Prog. Mol. Biol. Transl. Sci. 2012, 109, 75–112. [Google Scholar]
- Murata, S.; Takahama, Y.; Tanaka, K. Thymoproteasome: Probable role in generating positively selecting peptides. Curr. Opin. Immunol. 2008, 20, 192–196. [Google Scholar] [CrossRef]
- Wendler, P.; Enenkel, C. Nuclear Transport of Yeast Proteasomes. Front. Mol. Biosci. 2019, 6, 34. [Google Scholar] [CrossRef] [PubMed]
- Palmer, A.; Mason, G.G.; Paramio, J.M.; Knecht, E.; Rivett, A.J. Changes in proteasome localization during the cell cycle. Eur. J. Cell Boil. 1994, 64, 163–175. [Google Scholar]
- Albert, S.; Schaffer, M.; Beck, F.; Mosalaganti, S.; Asano, S.; Thomas, H.F.; Plitzko, J.M.; Beck, M.; Baumeister, W.; Engel, B.D. Proteasomes tether to two distinct sites at the nuclear pore complex. Proc. Natl. Acad. Sci. USA 2017, 114, 13726–13731. [Google Scholar] [CrossRef] [PubMed]
- Koch, B.; Yu, H.-G. Regulation of inner nuclear membrane associated protein degradation. Nucleus 2019, 10, 169–180. [Google Scholar] [CrossRef] [PubMed]
- Niepel, M.; Molloy, K.R.; Williams, R.; Farr, J.C.; Meinema, A.C.; Vecchietti, N.; Cristea, I.M.; Chait, B.T.; Rout, M.P.; Strambio-De-Castillia, C. The nuclear basket proteins Mlp1p and Mlp2p are part of a dynamic interactome including Esc1p and the proteasome. Mol. Boil. Cell 2013, 24, 3920–3938. [Google Scholar] [CrossRef]
- Husnjak, K.; Dikic, I. Ubiquitin-Binding Proteins: Decoders of Ubiquitin-Mediated Cellular Functions. Annu. Rev. Biochem. 2012, 81, 291–322. [Google Scholar] [CrossRef]
- Marblestone, J.G.; Butt, S.; McKelvey, D.M.; Sterner, D.E.; Mattern, M.R.; Nicholson, B.; Eddins, M.J. Comprehensive Ubiquitin E2 Profiling of Ten Ubiquitin E3 Ligases. Cell Biophys. 2013, 67, 161–167. [Google Scholar] [CrossRef]
- Collins, G.A.; Goldberg, A.L. The Logic of the 26S Proteasome. Cell 2017, 169, 792–806. [Google Scholar] [CrossRef]
- Shoji, S.; Hanada, K.; Ohsawa, N.; Shirouzu, M. Central catalytic domain of BRAP (RNF52) recognizes the types of ubiquitin chains and utilizes oligo-ubiquitin for ubiquitylation. Biochem. J. 2017, 474, 3207–3226. [Google Scholar] [CrossRef]
- Hurley, J.H.; Lee, S.; Prag, G. Ubiquitin-binding domains. Biochem. J. 2006, 399, 361–372. [Google Scholar] [CrossRef]
- Grice, G.L.; Nathan, J.A. The recognition of ubiquitinated proteins by the proteasome. Cell. Mol. Life Sci. 2016, 73, 3497–3506. [Google Scholar] [CrossRef] [PubMed]
- Asher, G.; Lotem, J.; Sachs, L.; Kahana, C.; Shaul, Y. Mdm-2 and ubiquitin-independent p53 proteasomal degradation regulated by NQO1. Proc. Natl. Acad. Sci. USA 2002, 99, 13125–13130. [Google Scholar] [CrossRef] [PubMed]
- Wiggins, C.M.; Tsvetkov, P.; Johnson, M.; Joyce, C.L.; Lamb, C.A.; Bryant, N.J.; Komander, D.; Shaul, Y.; Cook, S.J. BIM(EL), an intrinsically disordered protein, is degraded by 20S proteasomes in the absence of poly-ubiquitylation. J. Cell Sci. 2011, 124, 969–977. [Google Scholar] [CrossRef] [PubMed]
- Craxton, A.; Butterworth, M.; Harper, N.; Fairall, L.; Schwabe, J.; Ciechanover, A.; Cohen, G.M. NOXA, a sensor of proteasome integrity, is degraded by 26S proteasomes by an ubiquitin-independent pathway that is blocked by MCL-1. Cell Death Differ. 2012, 19, 1424–1434. [Google Scholar] [CrossRef]
- Prakash, S.; Tian, L.; Ratliff, K.S.; E Lehotzky, R.; Matouschek, A. An unstructured initiation site is required for efficient proteasome-mediated degradation. Nat. Struct. Mol. Boil. 2004, 11, 830–837. [Google Scholar] [CrossRef]
- Krappmann, D.; Wulczyn, F.G.; Scheidereit, C. Different mechanisms control signal-induced degradation and basal turnover of the NF-kappaB inhibitor IkappaB alpha in vivo. EMBO J. 1996, 15, 6716–6726. [Google Scholar] [CrossRef]
- Saftig, P.; Klumperman, J. Lysosome biogenesis and lysosomal membrane proteins: Trafficking meets function. Nat. Rev. Mol. Cell Boil. 2009, 10, 623–635. [Google Scholar] [CrossRef]
- Braulke, T.; Bonifacino, J.S. Sorting of lysosomal proteins. Biochim. Biophys. Acta (BBA)-Bioenerg. 2009, 1793, 605–614. [Google Scholar] [CrossRef]
- Bonam, S.R.; Wang, F.; Muller, S. Lysosomes as a therapeutic target. Nat. Rev. Drug Discov. 2019, 18, 923–948. [Google Scholar] [CrossRef]
- Griffiths, G.; Hoflack, B.; Simons, K.; Mellman, I.; Kornfeld, S. The mannose 6-phosphate receptor and the biogenesis of lysosomes. Cell 1988, 52, 329–341. [Google Scholar] [CrossRef]
- Jongsma, M.L.; Berlin, I.; Wijdeven, R.H.; Janssen, L.; Janssen, G.M.; Garstka, M.A.; Janssen, H.; Mensink, M.; Van Veelen, P.; Spaapen, R.M.; et al. An ER-Associated Pathway Defines Endosomal Architecture for Controlled Cargo Transport. Cell 2016, 166, 152–166. [Google Scholar] [CrossRef] [PubMed]
- Ballabio, A.; Bonifacino, J.S. Lysosomes as dynamic regulators of cell and organismal homeostasis. Nat. Rev. Mol. Cell Boil. 2019, 21, 101–118. [Google Scholar] [CrossRef] [PubMed]
- Bonifacino, J.S.; Neefjes, J. Moving and positioning the endolysosomal system. Curr. Opin. Cell Boil. 2017, 47, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Dubnikov, T.; Ben-Gedalya, T.; Cohen, E. Protein Quality Control in Health and Disease. Cold Spring Harb. Perspect. Boil. 2016, 9, a023523. [Google Scholar] [CrossRef] [PubMed]
- Lecker, S.H.; Goldberg, A.L.; Mitch, W.E. Protein Degradation by the Ubiquitin–Proteasome Pathway in Normal and Disease States. J. Am. Soc. Nephrol. 2006, 17, 1807–1819. [Google Scholar] [CrossRef] [PubMed]
- Qin, H.; Shao, Q.; Igdoura, S.A.; Alaoui-Jamali, M.A.; Laird, D. Lysosomal and Proteasomal Degradation Play Distinct Roles in the Life Cycle of Cx43 in Gap Junctional Intercellular Communication-deficient and -competent Breast Tumor Cells. J. Boil. Chem. 2003, 278, 30005–30014. [Google Scholar] [CrossRef]
- Mizushima, N.; Yoshimori, T.; Ohsumi, Y. The Role of Atg Proteins in Autophagosome Formation. Annu. Rev. Cell Dev. Boil. 2011, 27, 107–132. [Google Scholar] [CrossRef]
- Galluzzi, L.; Baehrecke, E.H.; Ballabio, A.; Boya, P.; Pedro, J.M.B.-S.; Cecconi, F.; Choi, A.M.; Chu, C.T.; Codogno, P.; Colombo, M.I.; et al. Molecular definitions of autophagy and related processes. EMBO J. 2017, 36, 1811–1836. [Google Scholar] [CrossRef]
- Stolz, A.; Ernst, A.; Dikic, I. Cargo recognition and trafficking in selective autophagy. Nat. Cell Biol. 2014, 16, 495–501. [Google Scholar] [CrossRef]
- Krick, R.; Bremer, S.; Welter, E.; Schlotterhose, P.; Muehe, Y.; Eskelinen, E.L.; Thumm, M. Cdc48/p97 and Shp1/p47 regulate autophagosome biogenesis in concert with ubiquitin-like Atg8. J. Cell Biol. 2010, 190, 965–973. [Google Scholar] [CrossRef]
- Hara, T.; Nakamura, K.; Matsui, M.; Yamamoto, A.; Nakahara, Y.; Suzuki-Migishima, R.; Yokoyama, M.; Mishima, K.; Saito, I.; Okano, H.; et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 2006, 441, 885–889. [Google Scholar] [CrossRef] [PubMed]
- Juhasz, G.; Érdi, B.; Sass, M.; Neufeld, T.P. Atg7-dependent autophagy promotes neuronal health, stress tolerance, and longevity but is dispensable for metamorphosis in Drosophila. Genes Dev. 2007, 21, 3061–3066. [Google Scholar] [CrossRef] [PubMed]
- Bjørkøy, G.; Lamark, T.; Brech, A.; Outzen, H.; Perander, M.; Øvervatn, A.; Stenmark, H.; Johansen, T. p62/SQSTM1 forms protein aggregates degraded by autophagy and has a protective effect on huntingtin-induced cell death. J. Cell Biol. 2005, 171, 603–614. [Google Scholar] [CrossRef] [PubMed]
- Pankiv, S.; Clausen, T.H.; Lamark, T.; Brech, A.; Bruun, J.A.; Outzen, H.; Øvervatn, A.; Bjørkøy, G.; Johansen, T. p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitinated protein aggregates by autophagy. J. Biol. Chem. 2007, 282, 24131–24145. [Google Scholar] [CrossRef] [PubMed]
- Moscat, J.; Diaz-Meco, M.T.; Wooten, M.W. Signal integration and diversification through the p62 scaffold protein. Trends Biochem. Sci. 2007, 32, 95–100. [Google Scholar] [CrossRef]
- Sumimoto, H.; Kamakura, S.; Ito, T. Structure and Function of the PB1 Domain, a Protein Interaction Module Conserved in Animals, Fungi, Amoebas, and Plants. Sci. STKE 2007, 2007, re6. [Google Scholar] [CrossRef]
- Johansen, T.; Lamark, T. Selective autophagy mediated by autophagic adapter proteins. Autophagy 2011, 7, 279–296. [Google Scholar] [CrossRef]
- Komatsu, M.; Kurokawa, H.; Waguri, S.; Taguchi, K.; Kobayashi, A.; Ichimura, Y.; Sou, Y.-S.; Ueno, I.; Sakamoto, A.; Tong, K.I.; et al. The selective autophagy substrate p62 activates the stress responsive transcription factor Nrf2 through inactivation of Keap1. Nature 2010, 12, 213–223. [Google Scholar] [CrossRef]
- Rogov, V.; Dötsch, V.; Johansen, T.; Kirkin, V. Interactions between Autophagy Receptors and Ubiquitin-like Proteins Form the Molecular Basis for Selective Autophagy. Mol. Cell 2014, 53, 167–178. [Google Scholar] [CrossRef]
- Fujita, N.; Morita, E.; Itoh, T.; Tanaka, A.; Nakaoka, M.; Osada, Y.; Umemoto, T.; Saitoh, T.; Nakatogawa, H.; Kobayashi, S.; et al. Recruitment of the autophagic machinery to endosomes during infection is mediated by ubiquitin. J. Cell Boil. 2013, 203, 115–128. [Google Scholar] [CrossRef]
- Alemu, E.A.; Lamark, T.; Torgersen, K.M.; Birgisdottir, Å.B.; Larsen, K.B.; Jain, A.; Olsvik, H.; Øvervatn, A.; Kirkin, V.; Johansen, T. ATG8 Family Proteins Act as Scaffolds for Assembly of the ULK Complex. J. Boil. Chem. 2012, 287, 39275–39290. [Google Scholar] [CrossRef] [PubMed]
- Lamark, T.; Perander, M.; Outzen, H.; Kristiansen, K.; Øvervatn, A.; Michaelsen, E.; Bjørkøy, G.; Johansen, T. Interaction Codes within the Family of Mammalian Phox and Bem1p Domain-containing Proteins. J. Boil. Chem. 2003, 278, 34568–34581. [Google Scholar] [CrossRef] [PubMed]
- Kirkin, V.; Lamark, T.; Sou, Y.-S.; Bjørkøy, G.; Nunn, J.L.; Bruun, J.A.; Shvets, E.; McEwan, D.G.; Clausen, T.H.; Wild, P.; et al. A Role for NBR1 in Autophagosomal Degradation of Ubiquitinated Substrates. Mol. Cell 2009, 33, 505–516. [Google Scholar] [CrossRef]
- Deosaran, E.; Larsen, K.B.; Hua, R.; Sargent, G.; Wang, Y.; Kim, S.; Lamark, T.; Jauregui, M.; Law, K.; Lippincott-Schwartz, J.; et al. NBR1 acts as an autophagy receptor for peroxisomes. J. Cell Sci. 2012, 126, 939–952. [Google Scholar] [CrossRef] [PubMed]
- Wild, P.; Farhan, H.; McEwan, D.G.; Wagner, S.; Rogov, V.V.; Brady, N.R.; Richter, B.; Korac, J.; Waidmann, O.; Choudhary, C.; et al. Phosphorylation of the Autophagy Receptor Optineurin Restricts Salmonella Growth. Science 2011, 333, 228–233. [Google Scholar] [CrossRef]
- Korac, J.; Schaeffer, V.; Kovacević, I.; Clement, A.M.; Jungblut, B.; Behl, C.; Terzic, J.; Dikic, I. Ubiquitin-independent function of optineurin in autophagic clearance of protein aggregates. J. Cell Sci. 2012, 126, 580–592. [Google Scholar] [CrossRef]
- Kachaner, D.; Génin, P.; Laplantine, E.; Weil, R. Toward an integrative view of Optineurin functions. Cell Cycle 2012, 11, 2808–2818. [Google Scholar] [CrossRef]
- Von Muhlinen, N.; Akutsu, M.; Ravenhill, B.J.; Foeglein, Á.; Bloor, S.; Rutherford, T.J.; Freund, S.M.; Komander, D.; Randow, F. LC3C, Bound Selectively by a Noncanonical LIR Motif in NDP52, Is Required for Antibacterial Autophagy. Mol. Cell 2012, 48, 329–342. [Google Scholar] [CrossRef]
- Newman, A.C.; Scholefield, C.L.; Kemp, A.J.; Newman, M.; McIver, E.G.; Kamal, A.; Wilkinson, S. TBK1 kinase addiction in lung cancer cells is mediated via autophagy of Tax1bp1/Ndp52 and non-canonical NF-κB signalling. PLoS ONE 2012, 7, e50672. [Google Scholar] [CrossRef]
- Thurston, T.L.M.; Ryzhakov, G.; Bloor, S.; Von Muhlinen, N.; Randow, F. The TBK1 adaptor and autophagy receptor NDP52 restricts the proliferation of ubiquitin-coated bacteria. Nat. Immunol. 2009, 10, 1215–1221. [Google Scholar] [CrossRef]
- Farré, J.-C.; Burkenroad, A.; Burnett, S.F.; Subramani, S. Phosphorylation of mitophagy and pexophagy receptors coordinates their interaction with Atg8 and Atg11. EMBO Rep. 2013, 14, 441–449. [Google Scholar] [CrossRef] [PubMed]
- Schweers, R.L.; Zhang, J.; Randall, M.S.; Loyd, M.R.; Li, W.; Dorsey, F.C.; Kundu, M.; Opferman, J.T.; Cleveland, J.L.; Miller, J.L.; et al. NIX is required for programmed mitochondrial clearance during reticulocyte maturation. Proc. Natl. Acad. Sci. USA 2007, 104, 19500–19505. [Google Scholar] [CrossRef] [PubMed]
- Lampert, M.A.; Orogo, A.M.; Najor, R.H.; Hammerling, B.C.; Leon, L.J.; Wang, B.; Kim, T.; Sussman, M.A.; Gustafsson, Å.B. BNIP3L/NIX and FUNDC1-mediated mitophagy is required for mitochondrial network remodeling during cardiac progenitor cell differentiation. Autophagy 2019, 15, 1182–1198. [Google Scholar] [CrossRef] [PubMed]
- Novak, I.; Kirkin, V.; McEwan, D.G.; Zhang, J.; Wild, P.; Rozenknop, A.; Rogov, V.; Löhr, F.; Popovic, D.; Occhipinti, A.; et al. Nix is a selective autophagy receptor for mitochondrial clearance. EMBO Rep. 2009, 11, 45–51. [Google Scholar] [CrossRef]
- Liu, L.; Feng, D.; Chen, G.; Chen, M.; Zheng, Q.; Song, P.; Ma, Q.; Zhu, C.; Wang, R.; Qi, W.; et al. Mitochondrial outer-membrane protein FUNDC1 mediates hypoxia-induced mitophagy in mammalian cells. Nature 2012, 14, 177–185. [Google Scholar] [CrossRef]
- Lu, J.-H.; He, L.; Behrends, C.; Araki, M.; Araki, K.; Wang, Q.J.; Catanzaro, J.M.; Friedman, S.L.; Zong, W.-X.; Fiel, M.I.; et al. NRBF2 regulates autophagy and prevents liver injury by modulating Atg14L-linked phosphatidylinositol-3 kinase III activity. Nat. Commun. 2014, 5, 3920. [Google Scholar] [CrossRef]
- B’chir, W.; Maurin, A.C.; Carraro, V.; Averous, J.; Jousse, C.; Muranishi, Y.; Parry, L.; Stepien, G.; Fafournoux, P.; Bruhat, A. The eIF2alpha/ATF4 pathway is essential for stress-induced autophagy gene expression. Nucleic Acids Res. 2013, 41, 7683–7699. [Google Scholar] [CrossRef]
- Bao, Y.; Pu, Y.; Yu, X.; Gregory, B.D.; Srivastava, R.; Howell, S.H.; Bassham, D.C. IRE1B degrades RNAs encoding proteins that interfere with the induction of autophagy by ER stress in Arabidopsis thaliana. Autophagy 2018, 14, 1562–1573. [Google Scholar] [CrossRef]
- Deegan, S.; Koryga, I.; Glynn, S.A.; Gupta, S.; Gorman, A.M.; Samali, A. A close connection between the PERK and IRE arms of the UPR and the transcriptional regulation of autophagy. Biochem. Biophys. Res. Commun. 2015, 456, 305–311. [Google Scholar] [CrossRef]
- Guilbert, S.M.; Lambert, H.; Rodrigue, M.-A.; Fuchs, M.; Landry, J.; Lavoie, J.N. HSPB8 and BAG3 cooperate to promote spatial sequestration of ubiquitinated proteins and coordinate the cellular adaptive response to proteasome insufficiency. FASEB J. 2018, 32, 3518–3535. [Google Scholar] [CrossRef]
- Gao, Y.-S.; Hubbert, C.C.; Lu, J.; Lee, Y.-S.; Lee, J.Y.; Yao, T.-P. Histone Deacetylase 6 Regulates Growth Factor-Induced Actin Remodeling and Endocytosis. Mol. Cell. Boil. 2007, 27, 8637–8647. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.Y.; Koga, H.; Kawaguchi, Y.; Tang, W.; Wong, E.; Gao, Y.-S.; Pandey, U.B.; Kaushik, S.; Tresse, E.; Lu, J.; et al. HDAC6 controls autophagosome maturation essential for ubiquitin-selective quality-control autophagy. EMBO J. 2010, 29, 969–980. [Google Scholar] [CrossRef] [PubMed]
- Ju, J.S.; Miller, S.E.; Hanson, P.I.; Weihl, C.C. Impaired protein aggregate handling and clearance underlie the pathogenesis of p97/VCP-associated disease. J. Biol. Chem. 2008, 283, 30289–30299. [Google Scholar] [CrossRef] [PubMed]
- Kalveram, B.; Schmidtke, G.; Groettrup, M. The ubiquitin-like modifier FAT10 interacts with HDAC6 and localizes to aggresomes under proteasome inhibition. J. Cell Sci. 2008, 121, 4079–4088. [Google Scholar] [CrossRef] [PubMed]
- Meriin, A.B.; Narayanan, A.; Meng, L.; Alexandrov, I.; Varelas, X.; Cissé, I.I.; Sherman, M.Y. Hsp70–Bag3 complex is a hub for proteotoxicity-induced signaling that controls protein aggregation. Proc. Natl. Acad. Sci. USA 2018, 115, E7043–E7052. [Google Scholar] [CrossRef] [PubMed]
- Schneider, J.L.; Suh, Y.; Cuervo, A.M. Deficient Chaperone-Mediated Autophagy in Liver Leads to Metabolic Dysregulation. Cell Metab. 2014, 20, 417–432. [Google Scholar] [CrossRef]

| Ubiquitin Ligases | Number of Ligases | Deubiquitinases | Number of DUBs |
|---|---|---|---|
| Tag Proteins with Ubiquitin Residues | Remove a Ubiquitin Residue from Proteins. | ||
| Ubiquitin activating enzyme E1 Ligase | 2 | Cystein Proteases: i. Ubiquitin specific proteases (USPs) ii. Ubiquitin carboxy-terminal hydrolases (UCHs) iii. Ovarian-tumor proteases (OTUs) iv. Machado-Joseph disease protein domain proteases (MJDs) v. Monocyte chemotactic protein-induced proteins (MCPIPs) vi. Permuted papain fold peptidases of dsRNA viruses and eukaryotes (PPPDEs) | 62 USPs 4 UCHs 15 OTUs 4 MJDs 7 MCPIPs |
| Ubiquitin conjugating enzyme E2 Ligase | 40 | Zinc-dependent metalloproteinases: i. JAMMs/MPN+ proteases | 4 JAMMs |
| Ubiquitin ligating enzymeE3 Ligase | 600 |
| Adapter Protein | Interaction Domain | Selective Autophagy Involved in |
|---|---|---|
| p62/SQSTM1 | UBA | Aggrephagy, mitophagy, xenophagy, pexophagy and zymophagy |
| NBR1 | UBA | Aggrephagy, mitophagy, xenophagy |
| Optineurin | UBAN, UBZ | Aggrephagy, mitophagy, xenophagy |
| NDP52 | UBZ | Aggrephagy, mitophagy, xenophagy |
| BNIP3L/NIX | - | Mitophagy |
| FUNDC1 | - | Mitophagy |
| HDAC6 | - | Aggrephagy and mitophagy |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mebratu, Y.A.; Negasi, Z.H.; Dutta, S.; Rojas-Quintero, J.; Tesfaigzi, Y. Adaptation of Proteasomes and Lysosomes to Cellular Environments. Cells 2020, 9, 2221. https://doi.org/10.3390/cells9102221
Mebratu YA, Negasi ZH, Dutta S, Rojas-Quintero J, Tesfaigzi Y. Adaptation of Proteasomes and Lysosomes to Cellular Environments. Cells. 2020; 9(10):2221. https://doi.org/10.3390/cells9102221
Chicago/Turabian StyleMebratu, Yohannes Afework, Zerihun Hailemariam Negasi, Saugata Dutta, Joselyn Rojas-Quintero, and Yohannes Tesfaigzi. 2020. "Adaptation of Proteasomes and Lysosomes to Cellular Environments" Cells 9, no. 10: 2221. https://doi.org/10.3390/cells9102221
APA StyleMebratu, Y. A., Negasi, Z. H., Dutta, S., Rojas-Quintero, J., & Tesfaigzi, Y. (2020). Adaptation of Proteasomes and Lysosomes to Cellular Environments. Cells, 9(10), 2221. https://doi.org/10.3390/cells9102221





