Oncogenic Role of ZFAS1 lncRNA in Head and Neck Squamous Cell Carcinomas
Abstract
1. Introduction
2. Materials and Methods
2.1. HNSCC Cell Culture and Quantification of ZFAS1 Expression
2.2. TCGA Data
2.3. Data Analysis
2.4. Gene Analysis
2.5. Targets Analysis
2.6. Statistical Analysis
2.7. Availability of Data and Materials
3. Results
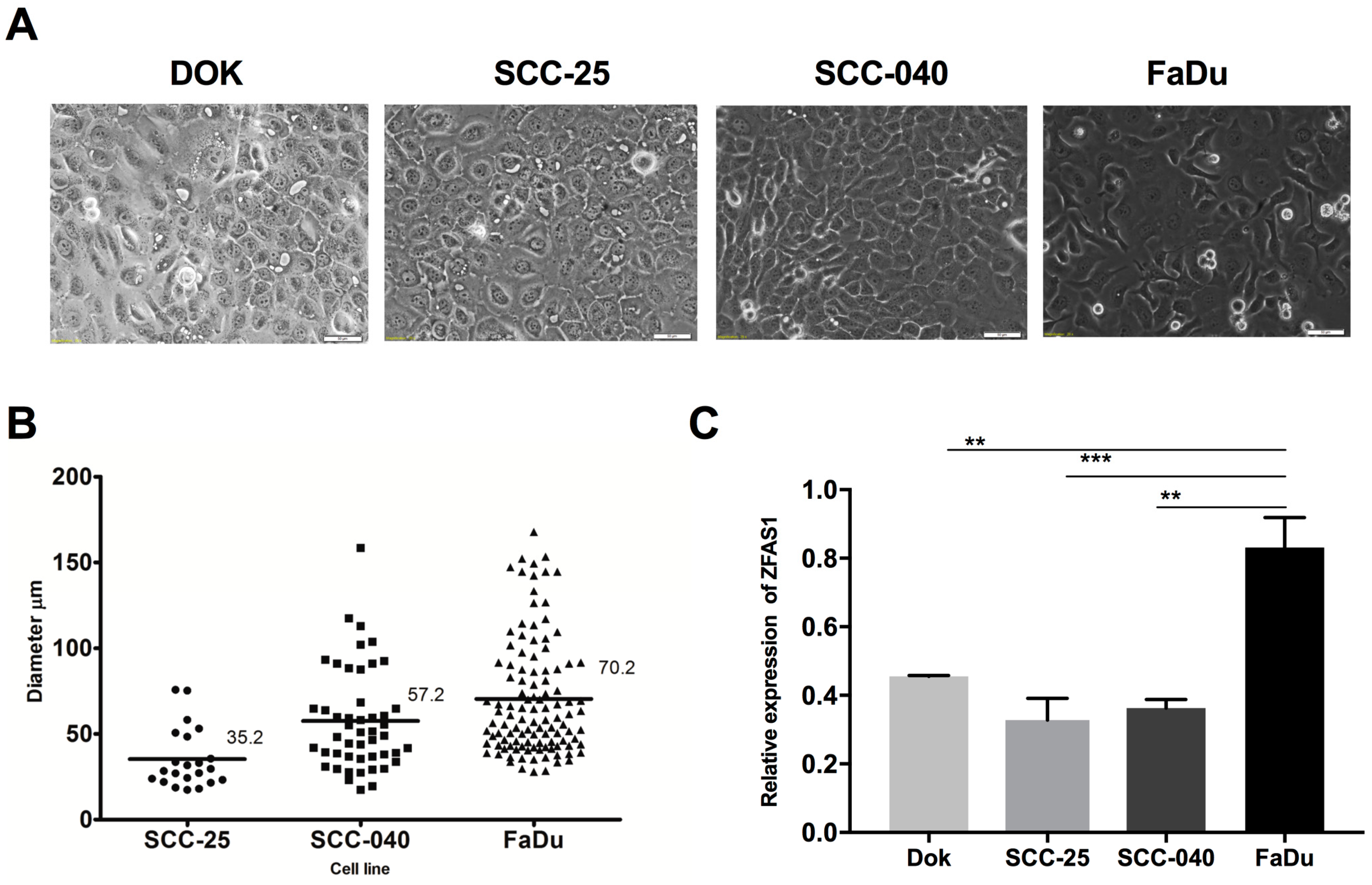
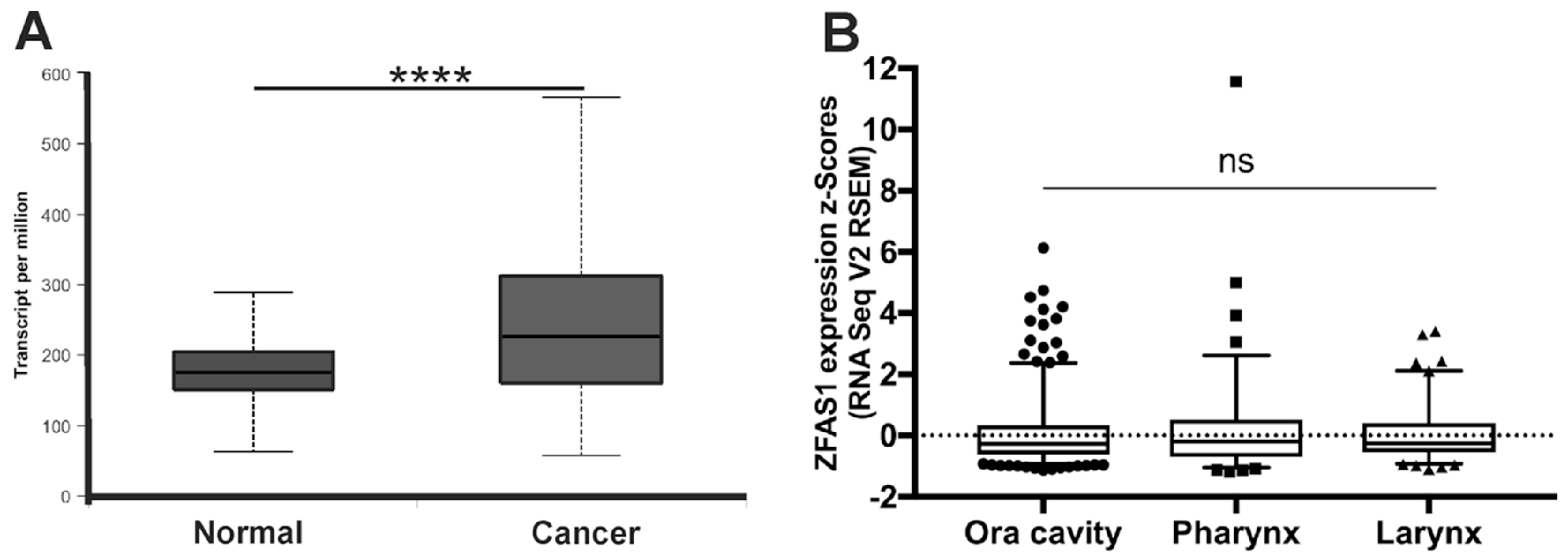
3.1. ZFAS1 is Up-Regulated in HNSCC Cell Lines and Cancer Samples of HNSCC Patients
3.2. ZFAS1 Levels Differ Depending on Clinicopathological Parameters
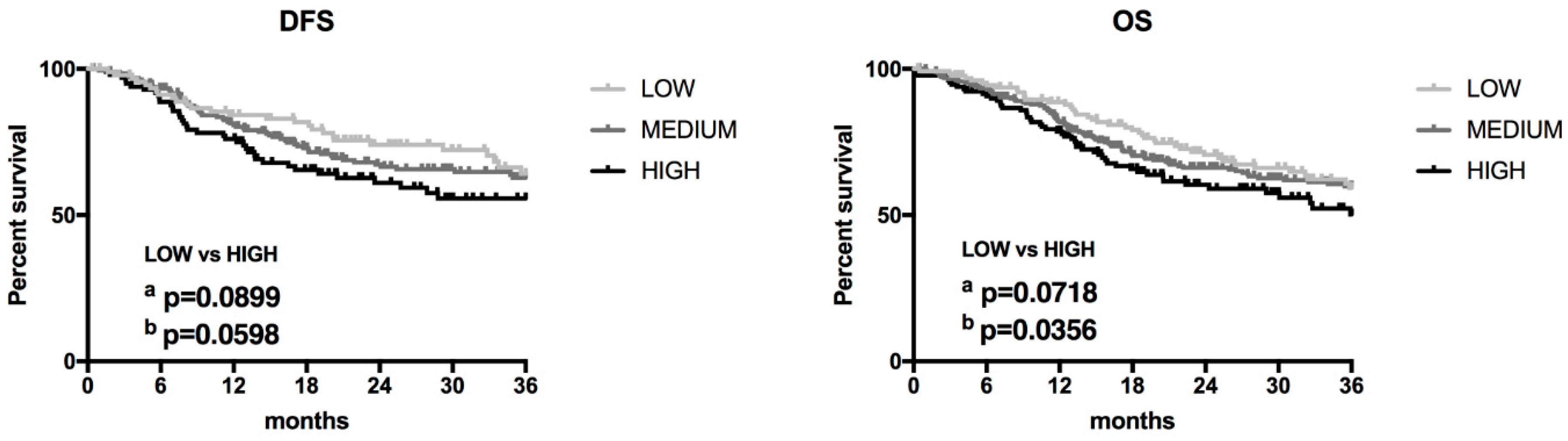
3.3. Association of ZFAS1 Expression and DFS and OS in the Studied Patients
3.4. ZFAS1 is Involved in Important Cellular Processes
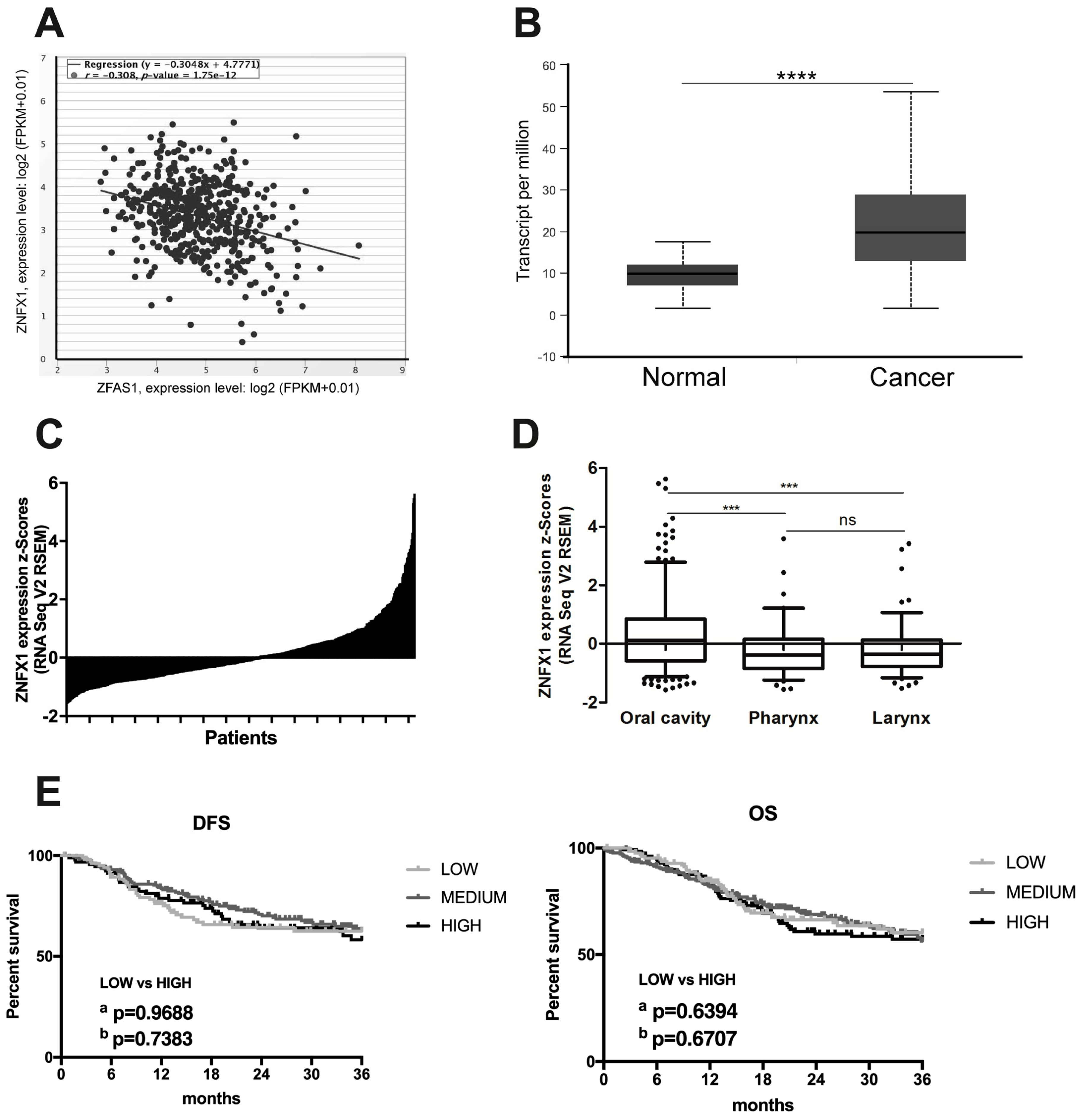
3.5. lncRNA ZFAS1 is Negatively Correlated with ZNFX1 mRNA in HNSCC
3.6. Role of ZFAS1 in the EMT Process, Cancer-Initiating Cells Maintenance, and Metastasis Process in HNSCC
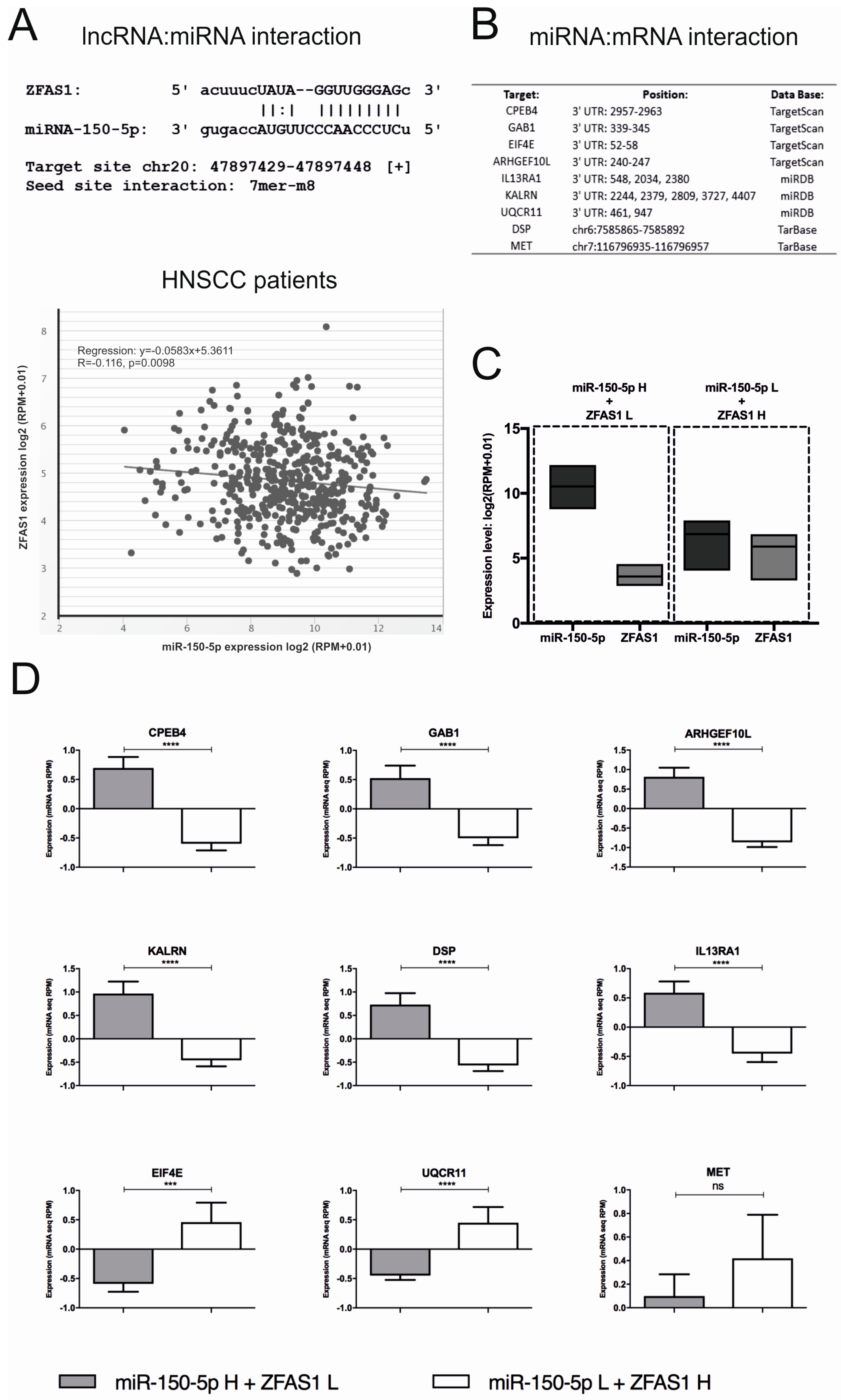
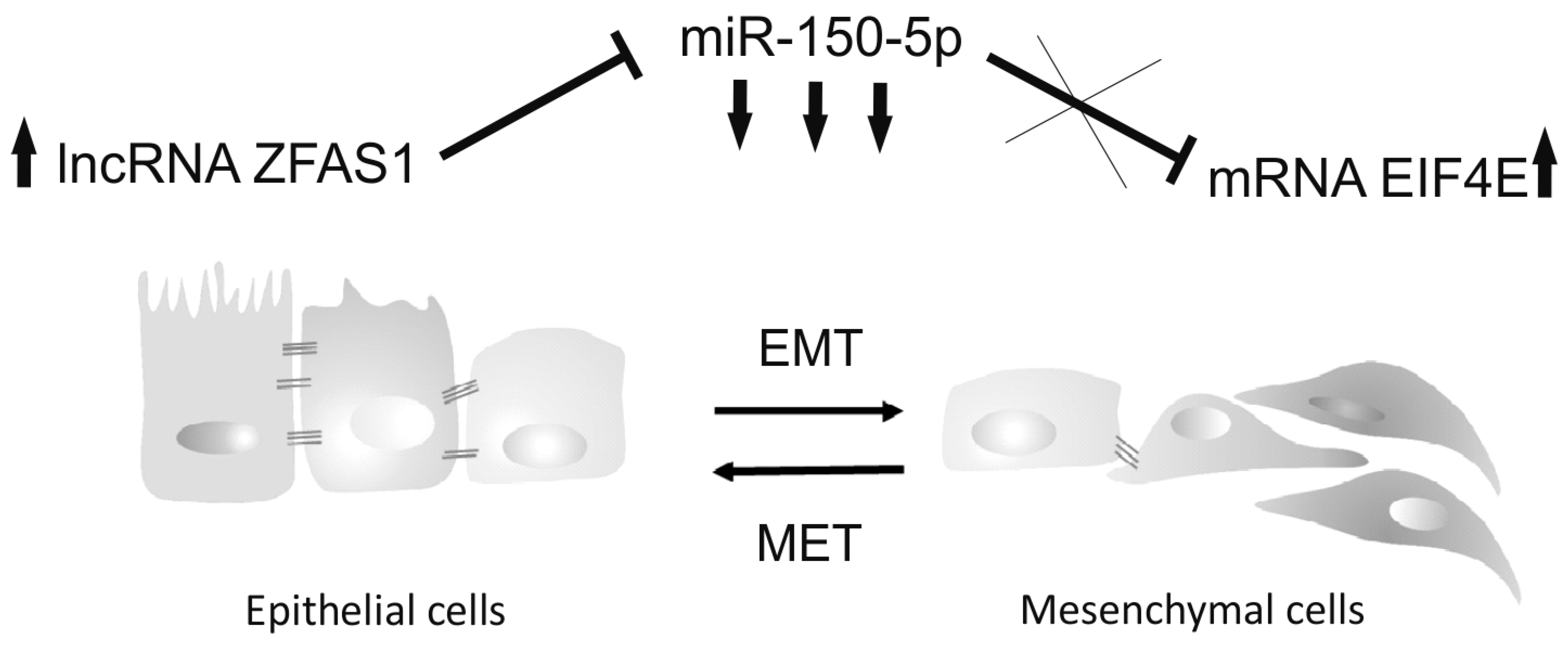
3.7. ZFAS1, As A Molecular Sponge, Regulates miR-150-5p and Influences the Cell Phenotype
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| HNSCC | head and neck squamous cell carcinoma |
| lncRNA | long non-coding RNA |
| qRT-PCR | quantitative reverse transcriptase PCR |
| TCGA | The Cancer Genome Atlas |
| EMT | epithelial-to-mesenchymal transition |
| HPV | human papillomavirus |
| PCR | polymerase chain reaction |
| B2M | beta-2 microglobulin |
| DFS | disease free survival |
| OS | overall survival |
| HR | Hazard Ratio |
| CI | Confidence Interval |
| SEM | standard error of the mean |
References
- Cohen, N.; Fedewa, S.; Chen, A.Y. Epidemiology and Demographics of the Head and Neck Cancer Population. Oral Maxillofac. Surg. Clin. N. Am. 2018, 30, 381–395. [Google Scholar] [CrossRef] [PubMed]
- Jou, A.; Hess, J. Epidemiology and Molecular Biology of Head and Neck Cancer. Oncol. Res. Treat. 2017, 40, 328–332. [Google Scholar] [CrossRef] [PubMed]
- Kolenda, T.; Przybyła, W.; Kapałczyńska, M.; Teresiak, A.; Zajaczkowska, M.; Blizniak, R.; Lamperska, K.M. Tumor microenvironment—Unknown niche with powerful therapeutic potentials. Rep. Pract. Oncol. Radiother. 2018, 23, 143–153. [Google Scholar] [CrossRef] [PubMed]
- Lasinska, I.; Kolenda, T.; Teresiak, A.; Lamperska, K.M.; Galus, L.; Mackiewicz, J. Immunotherapy in Patients with Recurrent and Metastatic Squamous Cell Carcinoma of the Head and Neck. Anticancer Agents Med. Chem. 2018, in press. [Google Scholar] [CrossRef]
- Zhao, G.; Fu, Y.; Su, Z.; Wu, R. How long-non coding RNAs and MicroRNAs mediate the endogenous RNA network of Head and Neck Squamous Cell Carcinoma: A comprehensive analysis. Cell Physiol. Biochem. 2018, 50, 332–341. [Google Scholar] [CrossRef]
- Lamperska, K.; Kozlowski, P.; Kolenda, T.; Teresiak, A.; Blizniak, R.; Przybyła, W.; Masternak, M.M.; Golusinski, P.; Golusinski, W. Unpredictable changes of selected miRNA in expression profile of HNSCC. Cancer Biomark. 2016, 16, 55–64. [Google Scholar] [CrossRef] [PubMed]
- Kolenda, T.; Guglas, K.; Ryś, M.; Bogaczyńska, M.; Łasińska, I.; Mackiewicz, J. Biological role of long non-coding RNA in head and neck cancers. Rep. Pract. Oncol. Radiother. 2017, 22, 378–388. [Google Scholar] [CrossRef] [PubMed]
- Guglas, K.; Bogaczyńska, M.; Kolenda, T.; Ryś, M.; Teresiak, A.; Bliźniak, R.; Łasińska, I.; Mackiewicz, J.; Lamperska, K. lncRNA in HNSCC: Challenges and potential. Contemp. Oncol. (Pozn). 2017, 21, 259–266. [Google Scholar] [CrossRef]
- Fatima, F.; Nawaz, M. Vesiculated Long Non-Coding RNAs: Offshore Packages Deciphering Trans-Regulation between Cells, Cancer Progression and Resistance to Therapies. Non-Coding RNA 2017, 3, 10. [Google Scholar] [CrossRef]
- Askarian-Amiri, M.E.; Crawford, J.; French, J.D.; Smart, C.E.; Smith, M.A.; Clark, M.B.; Ru, K.; Mercer, T.R.; Thompson, E.R.; Lakhani, S.R.; et al. Snord-host rna ZFAS1 is a regulator of mammary development and a potential marker for breast cancer. RNA 2011, 17, 878–891. [Google Scholar] [CrossRef] [PubMed]
- Gao, K.; Ji, Z.; She, K.; Yang, Q.; Shao, L. Long non-coding RNA ZFAS1 is an unfavourable prognostic factor and promotes glioma cell progression by activation of the Notch signaling pathway. Biomed. Pharmacother. 2017, 87, 555–560. [Google Scholar] [CrossRef] [PubMed]
- Lv, Q.L.; Chen, S.H.; Zhang, X.; Sun, B.; Hu, L.; Qu, Q.; Huang, Y.T.; Wang, G.H.; Liu, Y.L.; Zhang, Y.Y.; et al. Upregulation of long noncoding RNA zinc finger antisense 1 enhances epithelial–mesenchymal transition in vitro and predicts poor prognosis in glioma. Tumour Biol. 2017, 39. [Google Scholar] [CrossRef]
- Lee, J.; Park, H.Y.; Kim, W.W.; Lee, S.J.; Jeong, J.H.; Kang, S.H.; Jung, J.H.; Chae, Y.S. Biological function of long noncoding RNA snaR in Her2-positive breast cancer cells. Tumour Biol. 2017, 39. [Google Scholar] [CrossRef] [PubMed]
- Fan, S.; Fan, C.; Liu, N.; Huang, K.; Fang, X.; Wang, K. Down regulation of the long non-coding RNA ZFAS1 is associated with cell proliferation, migration and invasion in breast cancer. Mol. Med. Rep. 2018, 17, 6405–6412. [Google Scholar]
- Zhou, H.; Wang, F.; Chen, H.; Tan, Q.; Qiu, S.; Chen, S.; Jing, W.; Yu, M.; Liang, C.; Ye, S.; et al. Increased expression of long-noncoding rna ZFAS1 is associated with epithelial–mesenchymal transition of gastric cancer. Aging (Albany NY). 2016, 8, 2023–2038. [Google Scholar]
- Nie, F.; Yu, X.; Huang, M.; Wang, Y.; Xie, M.; Ma, H.; Wang, Z.; De, W.; Sun, M. Long noncoding rna ZFAS1 promotes gastric cancer cells proliferation by epigenetically repressing klf2 and nkd2 expression. Oncotarget 2017, 8, 38227–38238. [Google Scholar] [CrossRef] [PubMed]
- Pan, L.; Liang, W.; Fu, M.; Huang, Z.H.; Li, X.; Zhang, W.; Zhang, P.; Qian, H.; Jiang, P.C.; Xu, W.R.; et al. Exosomes-mediated transfer of long noncoding rna ZFAS1 promotes gastric cancer progression. J. Cancer Res. Clin. Oncol. 2017, 143, 991–1004. [Google Scholar] [CrossRef]
- Thorenoor, N.; Faltejskova-Vychytilova, P.; Hombach, S.; Mlcochova, J.; Kretz, M.; Svoboda, M.; Slaby, O. Long non-coding rna ZFAS1 interacts with cdk1 and is involved in p53-dependent cell cycle control and apoptosis in colorectal cancer. Oncotarget 2016, 7, 622–637. [Google Scholar] [CrossRef]
- Wang, W.; Xing, C. Upregulation of long noncoding rna ZFAS1 predicts poor prognosis and prompts invasion and metastasis in colorectal cancer. Pathol. Res. Pract. 2016, 212, 690–695. [Google Scholar] [CrossRef]
- Xie, S.; Quanxing, G.; Wang, X.; Sun, X.; Kang, Y. Long non-coding RNA ZFAS1 sponges miR-484 to promote cell proliferation and invasion in colorectal cancer. Cell Cycle 2018, 17, 154–161. [Google Scholar] [CrossRef]
- Fang, C.; Zan, J.; Yue, B.; Liu, C.; He, C.; Yan, D. Long non-coding ribonucleic acid zinc finger antisense 1 promotes the progression of colonic cancer by modulating zeb1 expression. J. Gastroenterol. Hepatol. 2017, 32, 1204–1211. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Xie, J.; Shen, C.; Cheng, D.; Shi, Y.; Wu, Z.; Deng, X.; Chen, H.; Shen, B.; Peng, C.; et al. Amplification of long noncoding rna ZFAS1 promotes metastasis in Hepatocellular carcinoma. Cancer Res. 2015, 75, 3181–3191. [Google Scholar] [CrossRef]
- Xia, B.; Hou, Y.; Chen, H.; Yang, S.; Liu, T.; Lin, M.; Lou, G. Long non-coding rna ZFAS1 interacts with mir-150-5p to regulate sp1 expression and ovarian cancer cell malignancy. Oncotarget 2017, 8, 19534–19546. [Google Scholar] [PubMed]
- Liu, R.; Zeng, Y.; Zhou, C.F.; Wang, Y.; Li, X.; Liu, Z.Q.; Chen, X.P.; Zhang, W.; Zhou, H.H. Long noncoding rna expression signature to predict platinum-based chemotherapeutic sensitivity of ovarian cancer patients. Sci. Rep. 2017, 7, 18. [Google Scholar] [CrossRef]
- Wei, Y.H.; Fu, Y.; Luo, H.J.; Li, R.; Li, H.Y.; Zhang, Z.; Zhu, Y.H.; Gao, Y.; Liu, X.L. Higher expression of ZFAS1 is associated with poor prognosis in malignant melanoma and promotes cell proliferation and invasion. Int.J. Clin. Exp. Pathol. 2017, 10, 4640–4646. [Google Scholar]
- Tian, F.M.; Meng, F.Q.; Wang, X.B. Overexpression of long-noncoding rna ZFAS1 decreases survival in human NSCLC patients. Eur. Rev. Med. Pharmacol. Sci. 2016, 20, 5126–5131. [Google Scholar] [PubMed]
- Liu, G.; Wang, L.; Han, H.; Li, Y.; Lu, S.; Li, T.; Cheng, C. Lncrna ZFAS1 promotes growth and metastasis by regulating bmi1 and zeb2 in osteosarcoma. Am. J. Cancer Res. 2017, 7, 1450–1462. [Google Scholar]
- Shi, H.; Liu, Z.; Pei, D.; Jiang, Y.; Zhu, H.; Chen, B. Development and validation of nomogram based on lncrna ZFAS1 for predicting survival in lymph node-negative esophageal squamous cell carcinoma patients. Oncotarget 2017, 8, 59048–59057. [Google Scholar] [CrossRef]
- Guo, H.; Wu, L.; Zhao, P.; Feng, A. Overexpression of long non-coding rna zinc finger antisense 1 in acute myeloid leukemia cell lines influences cell growth and apoptosis. Exp. Ther. Med. 2017, 14, 647–651. [Google Scholar] [CrossRef] [PubMed]
- Baytak, E.; Gong, Q.; Akman, B.; Yuan, H.; Chan, W.C.; Kucuk, C. Whole transcriptome analysis reveals dysregulated oncogenic lncRNAs in natural killer/t-cell lymphoma and establishes mir155hg as a target of prdm1. Tumour Biol. 2017, 39. [Google Scholar] [CrossRef]
- Wang, T.; Ma, S.; Qi, X.; Tang, X.; Cui, D.; Wang, Z.; Chi, J.; Li, P.; Zhai, B. Long noncoding rna znfx1-as1 suppresses growth of Hepatocellular carcinoma cells by regulating the methylation of mir-9. OncoTargets Ther. 2016, 9, 5005–5014. [Google Scholar]
- Lamperska, K.; Kolenda, T.; Teresiak, A.; Kowalik, A.; Kruszyna-Mochalska, M.; Jackowiak, W.; Blizniak, R.; Przybyła, W.; Kapałczyńska, M.; Kozłowski, P. Different levels of let-7d expression modulate response of FaDu cells to irradiation and chemotherapeutics. PLoS ONE 2017, 12, e0180265. [Google Scholar] [CrossRef]
- Feng, L.L.; Shen, F.R.; Zhou, J.H.; Chen, Y.G. Expression of the lncRNA ZFAS1 in cervical cancer and its correlation with prognosis and chemosensitivity. Gene 2019, 696, 105–112. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Aksoy, B.A.; Dogrusoz, U.; Dresdner, G.; Gross, B.; Sumer, S.O.; Sun, Y.; Jacobsen, A.; Sinha, R.; Larsson, E.; et al. Integrative Analysis of Complex Cancer Genomics and Clinical Profiles Using the cBioPortal. Sci. Signal. 2013, 6, pl1. [Google Scholar] [CrossRef]
- Chandrashekar, D.S.; Bashel, B.; Balasubramanya, S.A.H.; Creighton, C.J.; Ponce-Rodriguez, I.; Chakravarthi, B.V.S.K.; Varambally, S. UALCAN: A Portal for Facilitating Tumor Subgroup Gene Expression and Survival Analyses. Neoplasia 2017, 19, 649–658. [Google Scholar] [CrossRef]
- Li, J.H.; Liu, S.; Zhou, H.; Qu, L.H.; Yang, J.H. starBase v2.0: Decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networksfrom large-scale CLIP-Seq data. Nucleic Acids Res. 2014, 42, D92–D97. [Google Scholar] [CrossRef]
- Mi, H.; Muruganujan, A.; Casagrande, J.T.; Thomas, P.D. Large-scale gene function analysis with the PANTHER classification system. Nat. Protoc. 2013, 8, 1551–1566. [Google Scholar] [CrossRef] [PubMed]
- Xiao, M.; Liu, L.; Zhang, S.; Yang, X.; Wang, Y. Cancer stem cell biomarkers for head and neck squamous cell carcinoma: A bioinformatic analysis. Oncol. Rep. 2018, 40, 3843–3851. [Google Scholar] [CrossRef] [PubMed]
- Natarajan, J.; Chandrashekar, C.; Radhakrishnan, R. Critical biomarkers of epithelial-mesenchymal transition in the head and neck cancers. J. Cancer Res. Ther. 2014, 10, 512–518. [Google Scholar] [PubMed]
- Paczkowska, J.; Szyfter, K.; Giefing, M.; Wierzbicka, M. Genetic signature and profiling of head and neck cancer: Where do we stand? Curr. Opin. Otolaryngol. Head Neck Surg. 2017, 25, 154–158. [Google Scholar] [CrossRef]
- Rodrigo, J.P.; Martínez, P.; Allonca, E.; Alonso-Durán, L.; Suárez, C.; Astudillo, A.; García-Pedrero, J.M. Immunohistochemical markers of distant metastasis in laryngeal and hypopharyngeal squamous cell carcinomas. Clin. Exp. Metastasis 2014, 31, 317–325. [Google Scholar] [CrossRef] [PubMed]
- Fan, Z.; Li, M.; Chen, X.; Wang, J.; Liang, X.; Wang, H.; Wang, Z.; Cheng, B.; Xia, J. Prognostic Value of Cancer Stem Cell Markers in Head and Neck Squamous Cell Carcinoma: A Meta-analysis. Sci. Rep. 2017, 7, 43008. [Google Scholar] [CrossRef]
- Dahiya, K.; Dhankhar, R. Updated overview of current biomarkers in head and neck carcinoma. World J. Methodol. 2016, 6, 77–86. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, V.; Bell, G.W.; Nam, JW.; Bartel, D.P. Predicting effective microRNA target sites in mammalian mRNAs. Life 2015, 4, e05005. [Google Scholar] [CrossRef]
- Wong, N.; Wang, X. miRDB: An online resource for microRNA target prediction and functional annotations. Nucleic Acids Res. 2014, 43, D146–D152. [Google Scholar] [CrossRef]
- Vlachos, I.S.; Paraskevopoulou, M.D.; Karagkouni, D.; Georgakilas, G.; Vergoulis, T.; Kanellos, I.; Anastasopoulos, I.-L.; Maniou, S.; Karathanou, K.; Fevgas, D.K.A.; et al. DIANA-TarBase v7.0: Indexing more than half a million experimentally supported miRNA:mRNA interactions. Nucleic Acids Res. 2014, 43, D153–D159. [Google Scholar] [CrossRef]
- Dong, D.; Mu, Z.; Zhao, C.; Sun, M. ZFAS1: A novel tumor-related long non-coding RNA. Cancer Cell Int. 2018, 18, 125. [Google Scholar] [CrossRef]
- Wirth, M.; Jira, D.; Ott, A.; Piontek, G.; Pickhard, A. High NOTCH1 mRNA Expression Is Associated with Better Survival in HNSCC. Int. J. Mol. Sci. 2018, 19, 830. [Google Scholar] [CrossRef] [PubMed]
- Fan, S.; Ramalingam, S.S.; Kauh, J.; Xu, Z.; Khuri, F.R.; Sun, S. Phosphorylated eukaryotic translation initiation factor 4 (eIF4E) is elevated in human cancer tissues. Cancer Biol. Ther. 2009, 8, 1463–1469. [Google Scholar] [CrossRef]
- Siddiqui, N.; Sonenberg, N. Signalling to eIF4E in cancer. Biochem. Soc. Trans. 2015, 43, 763–772. [Google Scholar] [CrossRef] [PubMed]
- Pettersson, F.; Del Rincon, S.V.; Emond, A.; Huor, B.; Ngan, E.; Ng, J.; Dobocan, M.C.; Siegel, P.M.; Miller, W.H., Jr. Genetic and pharmacologic inhibition of eIF4E reduces breast cancer cell migration, invasion, and metastasis. Cancer Res. 2015, 75, 1102–1112. [Google Scholar] [CrossRef]
- Robichaud, N.; del Rincon, S.V.; Huor, B.; Alain, T.; Petruccelli, L.A.; Hearnden, J.; Goncalves, C.; Grotegut, S.; Spruck, C.H.; Furic, L.; et al. Phosphorylation of eIF4E promotes EMT and metastasis via translational control of SNAIL and MMP-3. Oncogene 2014, 34, 2032–2042. [Google Scholar] [CrossRef] [PubMed]
- Singh, J.; Jayaraj, R.; Baxi, S.; Mileva, M.; Skinner, J.; Dhand, N.K.; Thomas, M. Immunohistochemical expression levels of p53 and eIF4E markers in histologically negative surgical margins, and their association with the clinical outcome of patients with head and neck squamous cell carcinoma. Mol. Clin. Oncol. 2015, 4, 166–172. [Google Scholar] [CrossRef] [PubMed]
- DeFatta, R.J.; Nathan, C.O.; De Benedetti, A. Antisense RNA to eIF4E suppresses oncogenic properties of a head and neck squamous cell carcinoma cell line. Laryngoscope 2000, 110, 928–933. [Google Scholar] [CrossRef] [PubMed]
| Parameter | Group | Mean ± SEM | P-val |
|---|---|---|---|
| Age | <61.5 | 0.05596 ± 0.07434 N = 280 | |
| >61.5 | 0.09350 ± 0.06536 N = 240 | 0.2296 | |
| Gender | Female | −0.1504 ± 0.05728 N = 137 | |
| Male | 0.1511 ± 0.06425 N = 384 | 0.0923 | |
| Alcohol | Positive | 0.09964 ± 0.06692 N = 348 | |
| Negative | −0.002307 ± 0.07054 N = 162 | 0.8429 | |
| Smoking | No/Ex | 0.002190 ± 0.05292 N = 334 | |
| Yes | 0.1925 ± 0.1063 N = 177 | 0.1634 | |
| Cancer Stage | I + II | −0.1802 ± 0.07576 N = 98 | |
| III + IV | 0.1760 ± 0.06747 N = 348 | 0.0091 | |
| T Stage | T1 + T2 | −0.002843 ± 0.09133 N = 185 | |
| T3 + T4 | 0.1698 ± 0.06775 N = 274 | 0.0169 | |
| N Stage | N0 + N1 | 0.01680 ± 0.06265 N = 327 | |
| N2 + N3 | 0.09821 ± 0.08352 N = 172 | 0.5544 | |
| Grade | G1 + G2 | 0.01891 ± 0.05334 N = 367 | |
| G3 + G4 | 0.2433 ± 0.1274 N = 132 | 0.0891 | |
| Perineural Invasion | Positive | 0.09063 ± 0.07960 N = 168 | |
| Negative | 0.1079 ± 0.09359 N = 195 | 0.8824 | |
| Lymph Node Neck Dissection | Positive | 0.1145 ± 0.05814 N = 421 | |
| Negative | −0.1111 ± 0.08960 N = 97 | 0.0667 | |
| Angiolymphatic Invasion | Positive | 0.1644 ± 0.1395 N = 124 | |
| Negative | 0.03061 ± 0.05976 N = 225 | 0.5053 | |
| HPV p16 status | Negative | −0.1243 ± 0.1195 N = 72 | |
| Positive | 0.3604 ± 0.3343 N = 39 | 0.1090 |
| Positively Correlated with ZFAS1 | |
|---|---|
| Process | Genes |
| Cell cycle (GO:0007049/P00013) | MAD2L2, C10orf2, MND1, ANAPC11, S100A13, NAP1L1, POLL, RPA3, EIF3F |
| Cell adhesion (GO:0007155) | ITGAE |
| Signal transduction (GO:0007165) | RWDD3, ARL3, CNPY2, RPS3, RAE1, C14orf153, ARL16, SSR2, IFI27L1 |
| Death (GO:0016265) | NME2P1, RPS3, ANP32B, NME2, NAP1L1, C14orf153, IFI27L1, NME1 |
| Response to stimulus (GO:0050896) | RWDD3, C6orf154, PARK7, CNPY2, RPS3, C12orf44, C9orf119, C14orf153, SSR2, IFI27L1, POLR2I |
| Apoptosis signaling pathway (P00006) | ATF4, DIABLO |
| FAS signaling pathway (P00020) | CYC1 |
| Integrin signaling pathway (P00034) | ITGAE |
| mRNA splicing (P00058) | SNRPB2, SNRPA |
| Negatively Correlated with ZFAS1 | |
| Process | Genes |
| Cell adhesion (GO:0007155) | CELSR2, ADAP2 |
| Signal transduction (GO:0007165) | BMP2K, MAPK3, ARHGAP32, TOM1L2, SASH1, RAB10, TOM1, CTNND1, SNRK, RHBDF2, PKP1, RASAL1, CASP10, PPP1R9B, ADAP2, DOCK9, GAB1, KALRN, PDPK1, MAST4, HTT |
| Cell differentiation (GO:0030154) | PPARD, CYFIP1, PPP1R9B, TMOD3 |
| Death (GO:0016265) | CASP10 |
| Response to stimulus (GO:0050896) | BMP2K, MAPK3, ARHGAP32, SASH1, RAB10, IL13RA1, CTNND1, SNRK, RHBDF2, PPARD, PKP1, CYFIP1, RASAL1, IL4R, CASP10, SLC30A4, PPP1R9B, MAPK3, GAB1, KALRN, PDPK1, MAST4 |
| Angiogenesis (P00005) | MAPK3, JAK1 |
| Apoptosis signaling pathway (P00006) | IGF2R, CASP10, MAPK3 |
| Cadherin signaling pathway (P00012) | CTNND1, CELSR2 |
| EGF receptor signaling pathway (P00018) | RASAL1, MAPK3, GAB1 |
| Endothelial signaling pathway (P00019) | FURIN, MAPK3 |
| FAS signaling pathway (P00020) | CASP10 |
| FGF signaling pathway (P00021) | RASAL1, MAPK3 |
| Insulin/IGF pathway-mitogen activated protein kinase kinase/MAP kinase cascade (P00032) | IGF2R, MAPK3, PDPK1 |
| Integrin signalling pathway (P00034)/TGF-beta signaling pathway (P00052)/VEGF signaling pathway (P00056) | MAPK3 |
| Interleukin signaling pathway (P00036) | IL13RA1, IL4R, MAPK3, PDPK1 |
| JAK/STAT signaling pathway (P00038) | JAK1 |
| Oxidative stress response (P00046) | DUSP18 |
| PDGF signaling pathway (P00047) | RASAL1, JAK1, MAPK3, GAB1, PDPK1 |
| PI3 kinase pathway (P00048)/p53 pathway feedback loops 2 (P04398) | PDPK1 |
| Ras Pathway (P04393) | MAPK3, PDPK1 |
| Toll receptor signaling pathway (P00054) | MYD88, MAPK3 |
| Wnt signaling pathway (P00057) | CELSR2, PPARD |
| p38 MAPK pathway (P05918) | MAPK3, TAB2 |
| p53 pathway (P00059) | KAT2B, PDPK1 |
| Parameter | Group | Mean ± SEM | P-val |
|---|---|---|---|
| Age | <61.5 | 0.06453 ± 0.06357 N = 280 | |
| >61.5 | 0.1444 ± 0.07727 N = 240 | 0.7490 | |
| Gender | Female | 0.3651 ± 0.1037 N = 137 | |
| Male | 0.009726 ± 0.05510 N = 384 | 0.0004 | |
| Alcohol | Positive | 0.07041 ± 0.05853 N = 348 | |
| Negative | 0.1828 ± 0.09533 N = 162 | 0.4850 | |
| Smoking | No/Ex | 0.2099 ± 0.06822 N = 334 | |
| Yes | −0.1030 ± 0.06162 N = 177 | 0.0614 | |
| Cancer Stage | I + II | 0.5139 ± 0.1312 N = 98 | |
| III + IV | 0.03312 ± 0.05857 N = 348 | <0.0001 | |
| T Stage | T1 + T2 | 0.2607 ± 0.08801 N = 185 | |
| T3 + T4 | 0.04948 ± 0.06795 N = 274 | 0.0240 | |
| N Stage | N0 + N1 | 0.1589 ± 0.06431 N = 327 | |
| N2 + N3 | −0.03349 ± 0.07555 N = 172 | 0.0898 | |
| Grade | G1 + G2 | 0.1542 ± 0.05590 N = 367 | |
| G3 + G4 | 0.03624 ± 0.1150 N = 132 | 0.0158 | |
| Perineural Invasion | Positive | 0.3096 ± 0.09042 N = 168 | |
| Negative | −0.03063 ± 0.07312 N = 195 | 0.0022 | |
| Lymph Node Neck Dissection | Positive | 0.08454 ± 0.05560 N = 421 | |
| Negative | 0.1518 ± 0.1051 N = 97 | 0.2833 | |
| Angiolymphatic Invasion | Positive | −0.0378 ± 0.08767 N = 124 | |
| Negative | 0.2003 ± 0.07913 N = 225 | 0.0791 | |
| HPV p16 status | Negative | 0.2192 ± 0.1358 N = 72 | |
| Positive | −0.3231 ± 0.1469 N = 39 | 0.0086 |
| Gene | ZFAS1 Low | ZFAS1 High | P-val |
|---|---|---|---|
| Mean ± SEM | Mean ± SEM | ||
| POU5F1 | −0.1129 ± 0.07364 | 0.2302 ± 0.1369 | 0.1369 |
| CD44 | 0.3144 ± 0.1627 | −0.2309 ± 0.09515 | 0.007 |
| MET | 0.2658 ± 0.2658 | −0.2342 ± 0.08113 | 0.0002 |
| NOTCH1 | 0.3202 ± 0.119 | −0.1167 ± 0.1057 | <0.0001 |
| MME | 0.04295 ± 0.09562 | −0.05979 ± 0.03238 | 0.013 |
| BMI1 | 0.05728 ± 0.0754 | −0.1872 ± 0.09101 | 0.0119 |
| CDH11 | 0.2674 ± 0.0987 | −0.2047 ± 0.06785 | <0.0001 |
| CTNNB1 | −0.2204 ± 0.06578 | −0.5712 ± 0.08014 | <0.0001 |
| SMAD2 | −0.04951 ± 0.1148 | −0.5928 ± 0.1116 | 0.0005 |
| CXCR4 | 0.02447 ± 0.06266 | 0.05141 ± 0.1126 | 0.007 |
| MMP3 | 0.03104 ± 0.07116 | −0.1548 ± 0.06001 | 0.0179 |
| CXCR2 | 0.052 ± 0.07529 | −0.3646 ± 0.03099 | <0.0001 |
| SMAD3 | 0.05268 ± 0.08117 | −0.149 ± 0.1202 | 0.001 |
| MMP9 | 0.11 ± 0.0769 | 0.1212 ± 0.1297 | 0.0182 |
| MMP8 | 0.1216 ± 0.1174 | −0.07822 ± 0.0292 | 0.0278 |
| NUAK1 | 0.122 ± 0.07912 | −0.0688 ± 0.09496 | 0.0003 |
| LEF1 | 0.1263 ± 0.06421 | 0.1775 ± 0.1233 | 0.0453 |
| VIM | 0.1266 ± 0.0797 | 0.04328 ± 0.1193 | 0.0069 |
| NFKB1 | 0.1475 ± 0.08634 | −0.5829 ± 0.0967 | <0.0001 |
| CDH1 | 0.1571 ± 0.1007 | −0.4887 ± 0.07985 | <0.0001 |
| CCR7 | 0.1597 ± 0.08847 | −0.08974 ± 0.1232 | <0.0001 |
| DSP | 0.1695 ± 0.09073 | −0.5766 ± 0.05892 | <0.0001 |
| MMP2 | 0.1806 ± 0.0956 | −0.1522 ± 0.08578 | <0.0001 |
| RPS6KB1 | 0.2459 ± 0.0791 | −0.2194 ± 0.1326 | <0.0001 |
| COL1A1 | 0.2989 ± 0.1166 | −0.04641 ± 0.1686 | <0.0001 |
| ETS1 | 0.3457 ± 0.1039 | −0.4395 ± 0.06999 | <0.0001 |
| DNMT3B | 0.3723 ± 0.1346 | −0.06854 ± 0.06941 | 0.0397 |
| COL4A1 | 0.4047 ± 0.1015 | −0.3437 ± 0.06195 | <0.0001 |
| TJP1 | 0.4241 ± 0.08394 | −0.8134 ± 0.05945 | <0.0001 |
| CTNND1 | 0.4876 ± 0.08717 | −0.6306 ± 0.07966 | <0.0001 |
| CD274 | 0.4998 ± 0.2477 | −0.1627 ± 0.1442 | <0.0001 |
| PTK2 | 0.9937 ± 0.1134 | 0.7089 ± 0.1479 | 0.0146 |
| EGFR | 1.746 ± 0.434 | 0.09098 ± 0.1739 | <0.0001 |
| SLC3A2 | −0.2395 ± 0.08464 | 0.2994 ± 0.08896 | <0.0001 |
| EPCAM | −0.03831 ± 0.07123 | 0.4533 ± 0.1358 | 0.0023 |
| TAZ | −0.2428 ± 0.06541 | 0.9798 ± 0.135 | <0.0001 |
| JMJD6 | −0.2956 ± 0.05952 | 0.7102 ± 0.127 | <0.0001 |
| ABCG2 | −0.02039 ± 0.03598 | 0.04807 ± 0.1264 | <0.0001 |
| ABCG5 | −0.05742 ± 0.09303 | 0.4499 ± 0.1921 | <0.0001 |
| HSPA5 | −0.02533 ± 0.08621 | 0.3028 ± 0.1066 | 0.0248 |
| S100A4 | −0.1642 ± 0.07136 | 0.6315 ± 0.1802 | 0.0002 |
| EIF4E | −0.4339 ± 0.07892 | −0.03224 ± 0.1098 | 0.0053 |
| ANXA2 | −0.3988 ± 0.06529 | 0.4008 ± 0.119 | <0.0001 |
| ILK | −0.2853 ± 0.06627 | 0.3292 ± 0.1328 | 0.0007 |
| GSK3A | −0.2842 ± 0.09702 | 0.06703 ± 0.1189 | 0.0187 |
| TRIM28 | −0.2113 ± 0.09974 | 0.7619 ± 0.1348 | <0.0001 |
| COL2A1 | −0.1365 ± 0.08125 | 0.4195 ± 0.2156 | 0.002 |
| FN1 | −0.005116 ± 0.0545 | 0.04132 ± 0.1179 | <0.0001 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kolenda, T.; Guglas, K.; Kopczyńska, M.; Teresiak, A.; Bliźniak, R.; Mackiewicz, A.; Lamperska, K.; Mackiewicz, J. Oncogenic Role of ZFAS1 lncRNA in Head and Neck Squamous Cell Carcinomas. Cells 2019, 8, 366. https://doi.org/10.3390/cells8040366
Kolenda T, Guglas K, Kopczyńska M, Teresiak A, Bliźniak R, Mackiewicz A, Lamperska K, Mackiewicz J. Oncogenic Role of ZFAS1 lncRNA in Head and Neck Squamous Cell Carcinomas. Cells. 2019; 8(4):366. https://doi.org/10.3390/cells8040366
Chicago/Turabian StyleKolenda, Tomasz, Kacper Guglas, Magda Kopczyńska, Anna Teresiak, Renata Bliźniak, Andrzej Mackiewicz, Katarzyna Lamperska, and Jacek Mackiewicz. 2019. "Oncogenic Role of ZFAS1 lncRNA in Head and Neck Squamous Cell Carcinomas" Cells 8, no. 4: 366. https://doi.org/10.3390/cells8040366
APA StyleKolenda, T., Guglas, K., Kopczyńska, M., Teresiak, A., Bliźniak, R., Mackiewicz, A., Lamperska, K., & Mackiewicz, J. (2019). Oncogenic Role of ZFAS1 lncRNA in Head and Neck Squamous Cell Carcinomas. Cells, 8(4), 366. https://doi.org/10.3390/cells8040366






