Uridine Prevents Negative Effects of OXPHOS Xenobiotics on Dopaminergic Neuronal Differentiation
Abstract
1. Introduction
2. Materials and Methods
2.1. Cells, Mice and Materials
2.2. Cell Culture and Differentiation
2.3. Pharmacological Treatments
2.4. Cell Proliferation
2.5. Overexpressing OXPHOS-Related Mutant Proteins
2.6. Chromosomes and Mitochondrial DNA Analysis
2.7. DNA Methylation Analysis
2.8. RT-qPCR Analysis
2.9. Western Blotting
2.10. Flow Cytometry
2.11. Immunocytochemistry
2.12. ELISA
2.13. Analysis of Oxidative Phosphorylation Function
2.14. Mouse Experiments
2.15. Statistical Analysis
3. Results
3.1. OXPHOS Function and Neuronal Differentiation
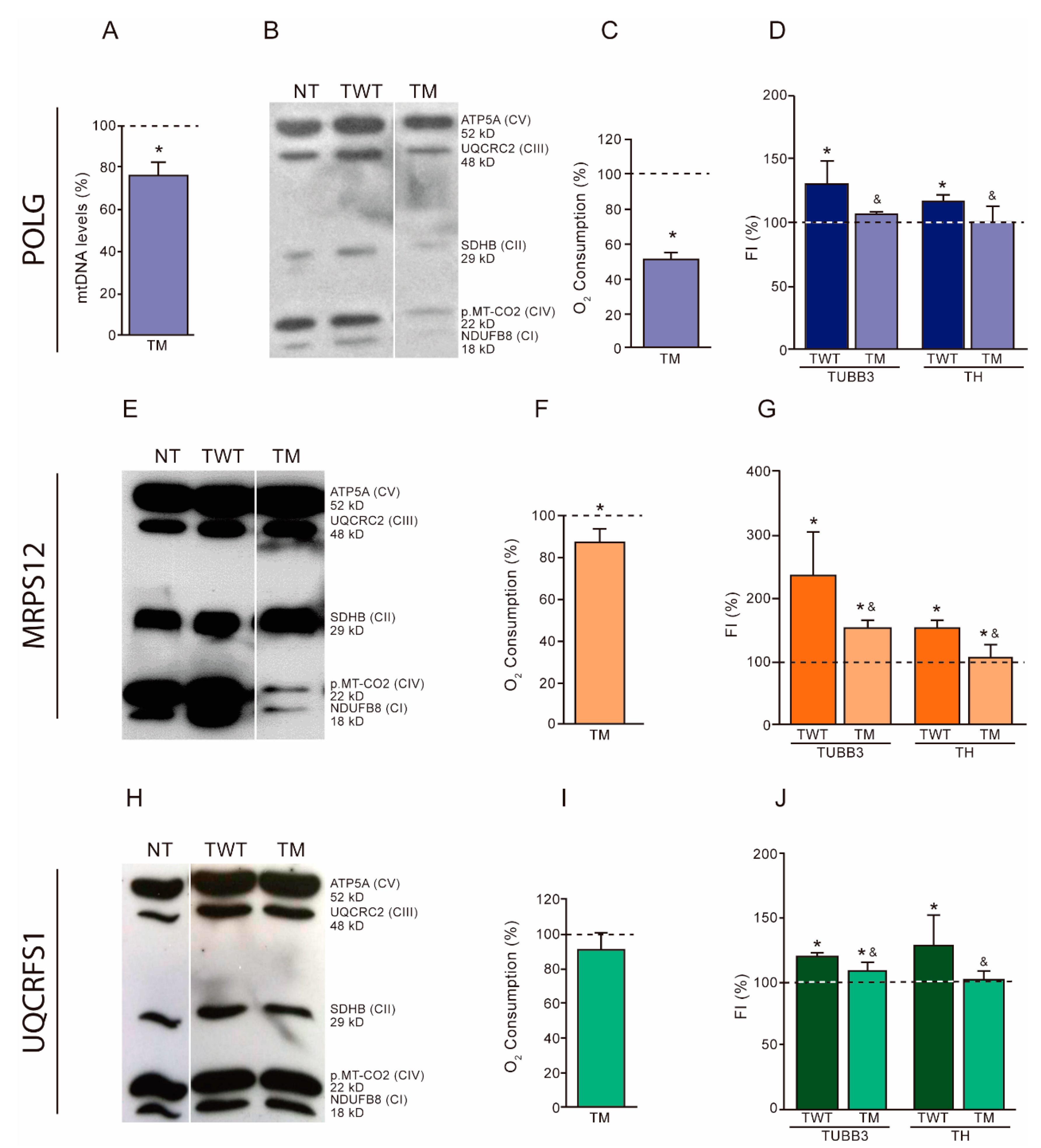
3.2. OXPHOS Dysfunction and Dopaminergic Neuronal Differentiation
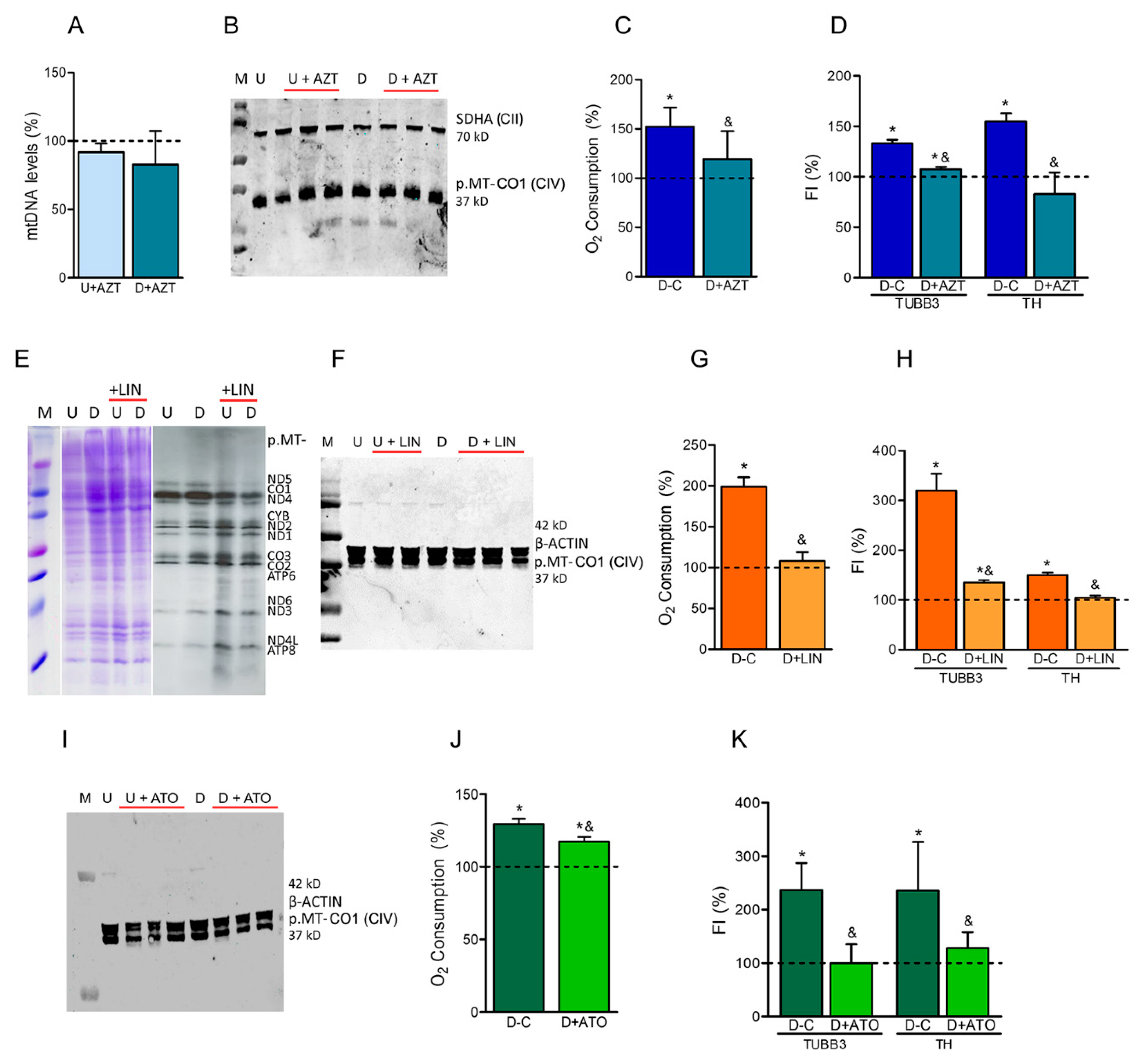
3.3. OXPHOS Xenobiotics and Dopaminergic Neuronal Differentiation
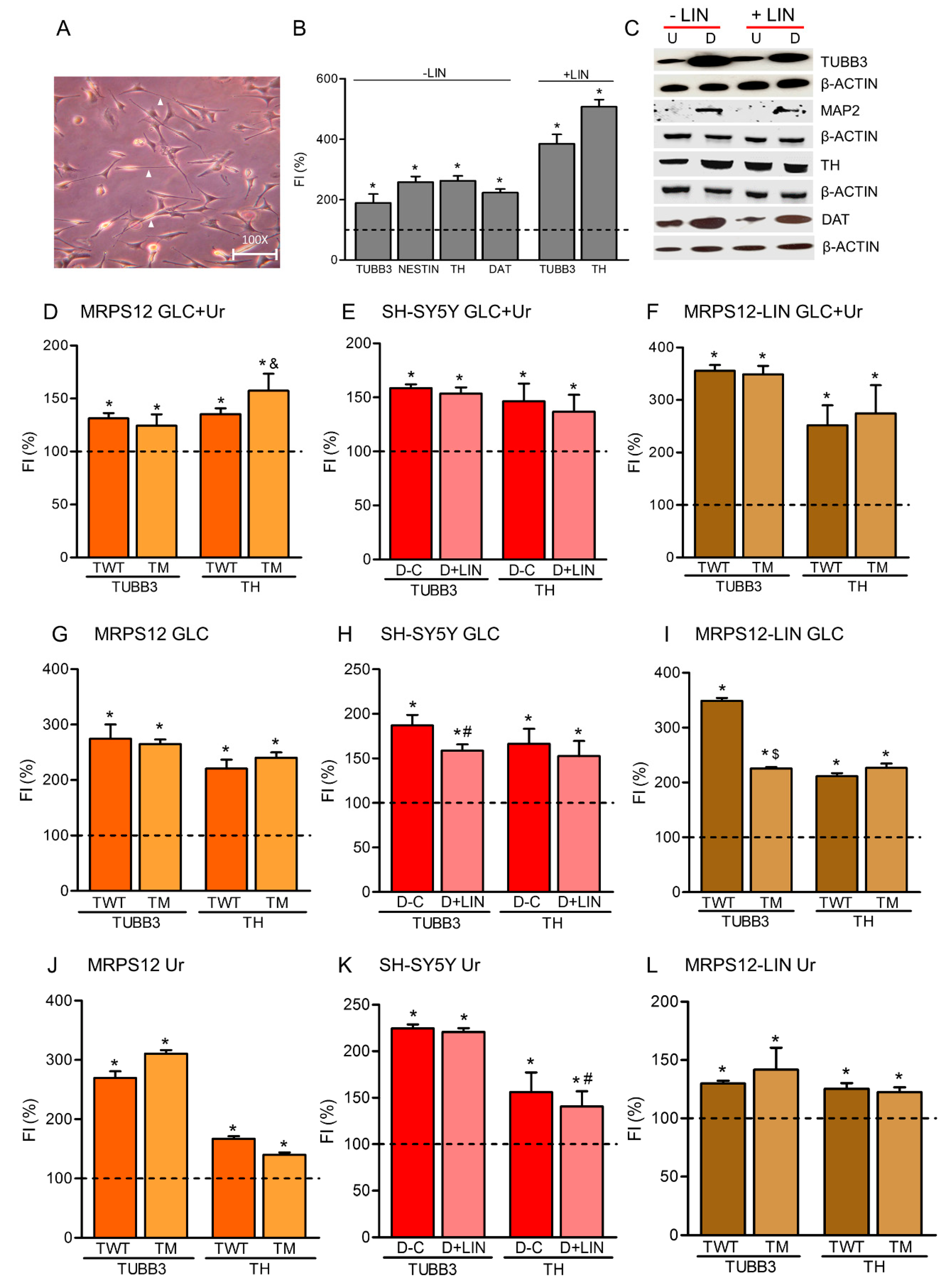
3.4. High Glucose or Uridine Compensate OXPHOS-Dysfunction Effects on Dopaminergic Neuronal Differentiation
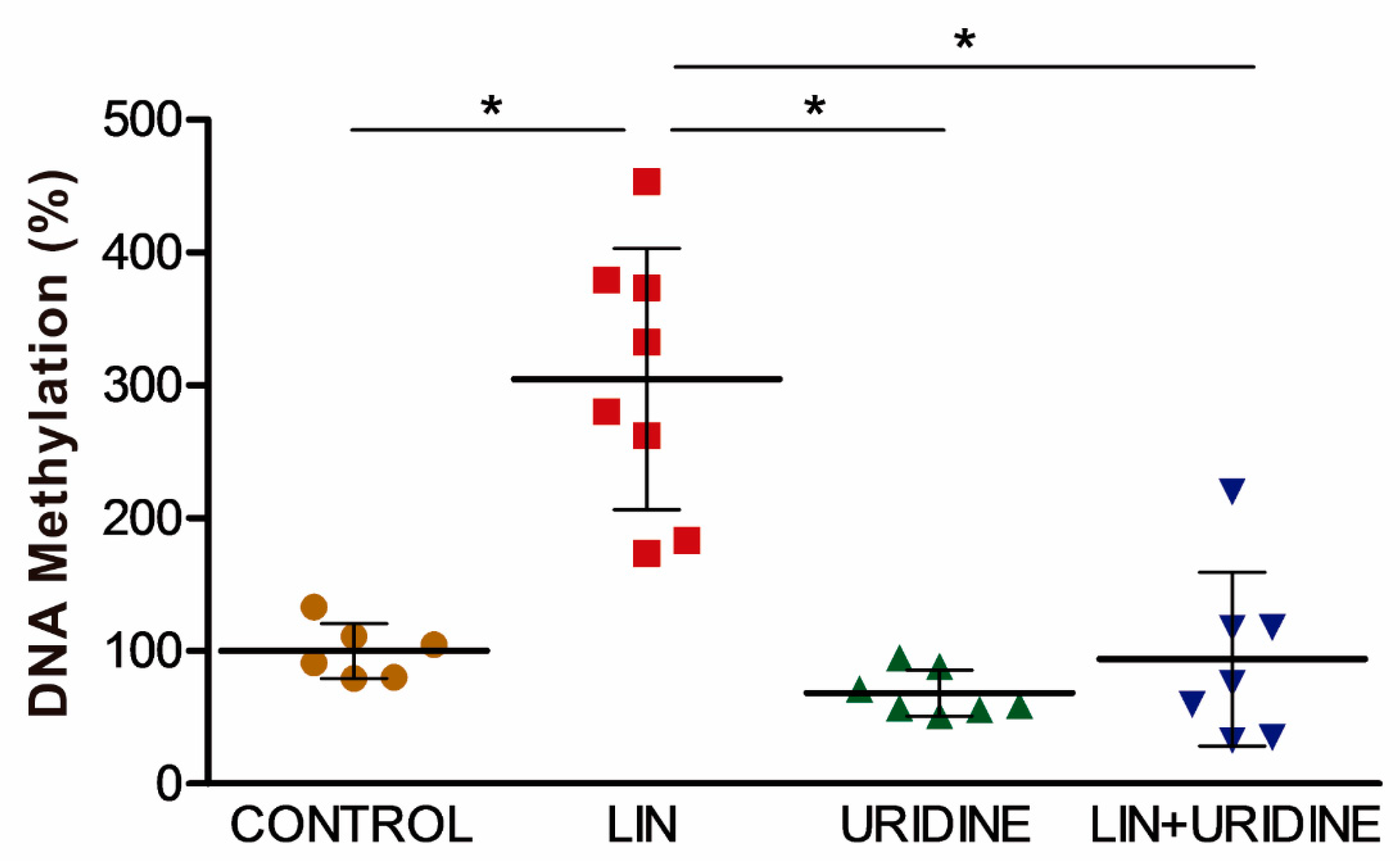
3.5. Effect of Prenatal Exposure to Linezolid on Mouse Brain
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Pacelli, C.; Giguere, N.; Bourque, M.J.; Levesque, M.; Slack, R.S.; Trudeau, L.E. Elevated mitochondrial bioenergetics and axonal arborization size are key contributors to the vulnerability of dopamine neurons. Curr. Biol. 2015, 25, 2349–2360. [Google Scholar] [CrossRef] [PubMed]
- Hall, C.N.; Klein-Flugge, M.C.; Howarth, C.; Attwell, D. Oxidative phosphorylation, not glycolysis, powers presynaptic and postsynaptic mechanisms underlying brain information processing. J. Neurosci. 2012, 32, 8940–8951. [Google Scholar] [CrossRef] [PubMed]
- Obeso, J.A.; Stamelou, M.; Goetz, C.G.; Poewe, W.; Lang, A.E.; Weintraub, D.; Burn, D.; Halliday, G.M.; Bezard, E.; Przedborski, S.; et al. Past, present, and future of Parkinson’s disease: A special essay on the 200th Anniversary of the Shaking Palsy. Mov. Disord. 2017, 32, 1264–1310. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Gallardo, E.; Iceta, R.; Iglesias, E.; Montoya, J.; Ruiz-Pesini, E. OXPHOS toxicogenomics and Parkinson’s disease. Mutat. Res. 2011, 728, 98–106. [Google Scholar] [CrossRef]
- Bender, A.; Krishnan, K.J.; Morris, C.M.; Taylor, G.A.; Reeve, A.K.; Perry, R.H.; Jaros, E.; Hersheson, J.S.; Betts, J.; Klopstock, T.; et al. High levels of mitochondrial DNA deletions in substantia nigra neurons in aging and Parkinson disease. Nat. Genet. 2006, 38, 515–517. [Google Scholar] [CrossRef]
- Coxhead, J.; Kurzawa-Akanbi, M.; Hussain, R.; Pyle, A.; Chinnery, P.; Hudson, G. Somatic mtDNA variation is an important component of Parkinson’s disease. Neurobiol. Aging 2016, 38, 217.e211–217.e216. [Google Scholar] [CrossRef]
- Grunewald, A.; Rygiel, K.A.; Hepplewhite, P.D.; Morris, C.M.; Picard, M.; Turnbull, D.M. Mitochondrial DNA depletion in respiratory chain-deficient Parkinson disease neurons. Ann. Neurol. 2016, 79, 366–378. [Google Scholar] [CrossRef]
- O’Brien, L.C.; Keeney, P.M.; Bennett, J.P., Jr. Differentiation of human neural stem cells into motor neurons stimulates mitochondrial biogenesis and decreases glycolytic flux. Stem Cells Dev. 2015, 24, 1984–1994. [Google Scholar] [CrossRef]
- Gothie, J.D.; Sebillot, A.; Luongo, C.; Legendre, M.; Nguyen Van, C.; Le Blay, K.; Perret-Jeanneret, M.; Remaud, S.; Demeneix, B.A. Adult neural stem cell fate is determined by thyroid hormone activation of mitochondrial metabolism. Mol. Metab. 2017, 6, 1551–1561. [Google Scholar] [CrossRef]
- Yokota, M.; Hatakeyama, H.; Ono, Y.; Kanazawa, M.; Goto, Y.I. Mitochondrial respiratory dysfunction disturbs neuronal and cardiac lineage commitment of human iPSCs. Cell Death Dis. 2017, 8, e2551. [Google Scholar] [CrossRef]
- Fang, D.; Qing, Y.; Yan, S.; Chen, D.; Yan, S.S. Development and dynamic regulation of mitochondrial network in human midbrain dopaminergic neurons differentiated from iPSCs. Stem Cell Rep. 2016, 7, 678–692. [Google Scholar] [CrossRef] [PubMed]
- Pashkovskaia, N.; Gey, U.; Rodel, G. Mitochondrial ROS direct the differentiation of murine pluripotent P19 cells. Stem Cell Res. 2018, 30, 180–191. [Google Scholar] [CrossRef] [PubMed]
- Pesini, A.; Iglesias, E.; Bayona-Bafaluy, M.P.; Garrido-Perez, N.; Meade, P.; Gaudo, P.; Jimenez-Salvador, I.; Andres-Benito, P.; Montoya, J.; Ferrer, I.; et al. Brain pyrimidine nucleotide synthesis and Alzheimer disease. Aging 2019, 11, 8433–8462. [Google Scholar] [CrossRef] [PubMed]
- Iglesias, E.; Pesini, A.; Garrido-Perez, N.; Meade, P.; Bayona-Bafaluy, M.P.; Montoya, J.; Ruiz-Pesini, E. Prenatal exposure to oxidative phosphorylation xenobiotics and late-onset Parkinson disease. Ageing Res. Rev. 2018, 45, 24–32. [Google Scholar] [CrossRef] [PubMed]
- Renault, D.; Yousef, H.; Mohamed, A.A. The multilevel antibiotic-induced perturbations to biological systems: Early-life exposure induces long-lasting damages to muscle structure and mitochondrial metabolism in flies. Environ. Pollut. 2018, 241, 821–833. [Google Scholar] [CrossRef] [PubMed]
- Llobet, L.; Toivonen, J.M.; Montoya, J.; Ruiz-Pesini, E.; Lopez-Gallardo, E. Xenobiotics that affect oxidative phosphorylation alter differentiation of human adipose-derived stem cells at concentrations that are found in human blood. Dis. Model. Mech. 2015, 8, 1441–1455. [Google Scholar] [CrossRef] [PubMed]
- Llobet, L.; Bayona-Bafaluy, M.P.; Pacheu-Grau, D.; Torres-Perez, E.; Arbones-Mainar, J.M.; Navarro, M.A.; Gomez-Diaz, C.; Montoya, J.; Lopez-Gallardo, E.; Ruiz-Pesini, E. Pharmacologic concentrations of linezolid modify oxidative phosphorylation function and adipocyte secretome. Redox Biol. 2017, 13, 244–254. [Google Scholar] [CrossRef]
- Le Grand, J.N.; Gonzalez-Cano, L.; Pavlou, M.A.; Schwamborn, J.C. Neural stem cells in Parkinson’s disease: A role for neurogenesis defects in onset and progression. Cell. Mol. Life Sci. 2015, 72, 773–797. [Google Scholar] [CrossRef]
- King, M.P.; Attardi, G. Human cells lacking mtDNA: Repopulation with exogenous mitochondria by complementation. Science 1989, 246, 500–503. [Google Scholar] [CrossRef]
- King, M.P.; Attardi, G. Isolation of human cell lines lacking mitochondrial DNA. Methods Enzymol. 1996, 264, 304–313. [Google Scholar] [CrossRef]
- Presgraves, S.P.; Ahmed, T.; Borwege, S.; Joyce, J.N. Terminally differentiated SH-SY5Y cells provide a model system for studying neuroprotective effects of dopamine agonists. Neurotox. Res. 2004, 5, 579–598. [Google Scholar] [CrossRef] [PubMed]
- Swistowski, A.; Peng, J.; Han, Y.; Swistowska, A.M.; Rao, M.S.; Zeng, X. Xeno-free defined conditions for culture of human embryonic stem cells, neural stem cells and dopaminergic neurons derived from them. PLoS ONE 2009, 4, e6233. [Google Scholar] [CrossRef] [PubMed]
- Gardner, K.; Hall, P.A.; Chinnery, P.F.; Payne, B.A. HIV treatment and associated mitochondrial pathology: Review of 25 years of in vitro, animal, and human studies. Toxicol. Pathol. 2014, 42, 811–822. [Google Scholar] [CrossRef] [PubMed]
- Demir, M.; Laywell, E.D. Neurotoxic effects of AZT on developing and adult neurogenesis. Front. Neurosci. 2015, 9, 93. [Google Scholar] [CrossRef]
- Pacheu-Grau, D.; Gomez-Duran, A.; Iglesias, E.; Lopez-Gallardo, E.; Montoya, J.; Ruiz-Pesini, E. Mitochondrial antibiograms in personalized medicine. Hum. Mol. Genet. 2013, 22, 1132–1139. [Google Scholar] [CrossRef]
- Dryden, M.S. Linezolid pharmacokinetics and pharmacodynamics in clinical treatment. J Antimicrob. Chemother. 2011, 66 (Suppl. S4), iv7–iv15. [Google Scholar] [CrossRef]
- Gomez-Duran, A.; Pacheu-Grau, D.; Lopez-Perez, M.J.; Montoya, J.; Ruiz-Pesini, E. Mitochondrial pharma-Q-genomics: Targeting the OXPHOS cytochrome b. Drug Discov. Today 2011, 16, 176–180. [Google Scholar] [CrossRef]
- Song, Z.; Laleve, A.; Vallieres, C.; McGeehan, J.E.; Lloyd, R.E.; Meunier, B. Human mitochondrial cytochrome b variants studied in yeast: Not all are silent polymorphisms. Hum. Mutat. 2016, 37, 933–941. [Google Scholar] [CrossRef]
- Lv, Z.; Yan, X.; Lu, L.; Su, C.; He, Y. Atovaquone enhances doxorubicin’s efficacy via inhibiting mitochondrial respiration and STAT3 in aggressive thyroid cancer. J. Bioenerg. Biomembr. 2018, 50, 263–270. [Google Scholar] [CrossRef]
- Calderon, M.M.; Penzak, S.R.; Pau, A.K.; Kumar, P.; McManus, M.; Alfaro, R.M.; Kovacs, J.A. Efavirenz but not atazanavir/ritonavir significantly reduces atovaquone concentrations in HIV-infected subjects. Clin. Infect. Dis. 2016, 62, 1036–1042. [Google Scholar] [CrossRef]
- Emperador, S.; Pacheu-Grau, D.; Bayona-Bafaluy, M.P.; Garrido-Perez, N.; Martin-Navarro, A.; Lopez-Perez, M.J.; Montoya, J.; Ruiz-Pesini, E. An MRPS12 mutation modifies aminoglycoside sensitivity caused by 12S rRNA mutations. Front. Genet. 2014, 5, 469. [Google Scholar] [CrossRef] [PubMed]
- Gomez-Duran, A.; Pacheu-Grau, D.; Martinez-Romero, I.; Lopez-Gallardo, E.; Lopez-Perez, M.J.; Montoya, J.; Ruiz-Pesini, E. Oxidative phosphorylation differences between mitochondrial DNA haplogroups modify the risk of Leber’s hereditary optic neuropathy. Biochim. Biophys. Acta 2012, 1822, 1216–1222. [Google Scholar] [CrossRef] [PubMed]
- Kost, G.C.; Selvaraj, S.; Lee, Y.B.; Kim, D.J.; Ahn, C.H.; Singh, B.B. Clavulanic acid increases dopamine release in neuronal cells through a mechanism involving enhanced vesicle trafficking. Neurosci. Lett. 2011, 504, 170–175. [Google Scholar] [CrossRef] [PubMed]
- Gomez-Duran, A.; Pacheu-Grau, D.; Lopez-Gallardo, E.; Diez-Sanchez, C.; Montoya, J.; Lopez-Perez, M.J.; Ruiz-Pesini, E. Unmasking the causes of multifactorial disorders: OXPHOS differences between mitochondrial haplogroups. Hum. Mol. Genet. 2010, 19, 3343–3353. [Google Scholar] [CrossRef]
- Emperador, S.; Vidal, M.; Hernandez-Ainsa, C.; Ruiz-Ruiz, C.; Woods, D.; Morales-Becerra, A.; Arruga, J.; Artuch, R.; Lopez-Gallardo, E.; Bayona-Bafaluy, M.P.; et al. The decrease in mitochondrial DNA mutation load parallels visual recovery in a Leber hereditary optic neuropathy patient. Front. Neurosci. 2018, 12, 61. [Google Scholar] [CrossRef]
- Nair, A.B.; Jacob, S. A simple practice guide for dose conversion between animals and human. J. Basic Clin. Pharm. 2016, 7, 27–31. [Google Scholar] [CrossRef]
- Robinson, R.; Chapman, K.; Hudson, S.; Sparrow, S.; Spencer-Briggs, D.; Danks, A.; Hill, R.; Everett, D.; Mulier, B.; Old, S.; et al. Guidance on Dose Level Selection for Regulatory General Toxicology Studies for Pharmaceuticals. 2009. Available online: http://www.lasa.co.uk/PDF/LASA-NC3RsDoseLevelSelection.pdf (accessed on 20 January 2019).
- Birket, M.J.; Orr, A.L.; Gerencser, A.A.; Madden, D.T.; Vitelli, C.; Swistowski, A.; Brand, M.D.; Zeng, X. A reduction in ATP demand and mitochondrial activity with neural differentiation of human embryonic stem cells. J. Cell Sci. 2011, 124, 348–358. [Google Scholar] [CrossRef]
- Schneider, L.; Giordano, S.; Zelickson, B.R.; Johnson, M.S.; Benavides, G.A.; Ouyang, X.; Fineberg, N.; Darley-Usmar, V.M.; Zhang, J. Differentiation of SH-SY5Y cells to a neuronal phenotype changes cellular bioenergetics and the response to oxidative stress. Free Radic. Biol. Med. 2011, 51, 2007–2017. [Google Scholar] [CrossRef]
- Xun, Z.; Lee, D.Y.; Lim, J.; Canaria, C.A.; Barnebey, A.; Yanonne, S.M.; McMurray, C.T. Retinoic acid-induced differentiation increases the rate of oxygen consumption and enhances the spare respiratory capacity of mitochondria in SH-SY5Y cells. Mech. Ageing Dev. 2012, 133, 176–185. [Google Scholar] [CrossRef]
- Ducray, A.D.; Felser, A.; Zielinski, J.; Bittner, A.; Burgi, J.V.; Nuoffer, J.M.; Frenz, M.; Mevissen, M. Effects of silica nanoparticle exposure on mitochondrial function during neuronal differentiation. J. Nanobiotechnol. 2017, 15, 49. [Google Scholar] [CrossRef]
- Miller, S.W.; Trimmer, P.A.; Parker, W.D., Jr.; Davis, R.E. Creation and characterization of mitochondrial DNA-depleted cell lines with “neuronal-like” properties. J. Neurochem. 1996, 67, 1897–1907. [Google Scholar] [CrossRef] [PubMed]
- Wong, A.; Cavelier, L.; Collins-Schramm, H.E.; Seldin, M.F.; McGrogan, M.; Savontaus, M.L.; Cortopassi, G.A. Differentiation-specific effects of LHON mutations introduced into neuronal NT2 cells. Hum. Mol. Genet. 2002, 11, 431–438. [Google Scholar] [CrossRef] [PubMed]
- Ahlqvist, K.J.; Hamalainen, R.H.; Yatsuga, S.; Uutela, M.; Terzioglu, M.; Gotz, A.; Forsstrom, S.; Salven, P.; Angers-Loustau, A.; Kopra, O.H.; et al. Somatic progenitor cell vulnerability to mitochondrial DNA mutagenesis underlies progeroid phenotypes in Polg mutator mice. Cell Metab. 2012, 15, 100–109. [Google Scholar] [CrossRef] [PubMed]
- Lorenz, C.; Lesimple, P.; Bukowiecki, R.; Zink, A.; Inak, G.; Mlody, B.; Singh, M.; Semtner, M.; Mah, N.; Aure, K.; et al. Human iPSC-derived neural progenitors are an effective drug discovery model for neurological mtDNA disorders. Cell Stem Cell 2017, 20, 659–674.e659. [Google Scholar] [CrossRef]
- Villanueva-Paz, M.; Povea-Cabello, S.; Villalon-Garcia, I.; Suarez-Rivero, J.M.; Alvarez-Cordoba, M.; de la Mata, M.; Talaveron-Rey, M.; Jackson, S.; Sanchez-Alcazar, J.A. Pathophysiological characterization of MERRF patient-specific induced neurons generated by direct reprogramming. Biochim. Biophys. Acta Mol. Cell Res. 2019, 1866, 861–881. [Google Scholar] [CrossRef]
- Connolly, N.M.C.; Theurey, P.; Adam-Vizi, V.; Bazan, N.G.; Bernardi, P.; Bolanos, J.P.; Culmsee, C.; Dawson, V.L.; Deshmukh, M.; Duchen, M.R.; et al. Guidelines on experimental methods to assess mitochondrial dysfunction in cellular models of neurodegenerative diseases. Cell Death Differ. 2018, 25, 542–572. [Google Scholar] [CrossRef]
- Umeda, S.; Muta, T.; Ohsato, T.; Takamatsu, C.; Hamasaki, N.; Kang, D. The D-loop structure of human mtDNA is destabilized directly by 1-methyl-4-phenylpyridinium ion (MPP+), a parkinsonism-causing toxin. Eur. J. Biochem. 2000, 267, 200–206. [Google Scholar] [CrossRef]
- Pan-Zhou, X.R.; Cui, L.; Zhou, X.J.; Sommadossi, J.P.; Darley-Usmar, V.M. Differential effects of antiretroviral nucleoside analogs on mitochondrial function in HepG2 cells. Antimicrob. Agents Chemother. 2000, 44, 496–503. [Google Scholar] [CrossRef]
- Liu, Y.; Nguyen, P.; Baris, T.Z.; Poirier, M.C. Molecular analysis of mitochondrial compromise in rodent cardiomyocytes exposed long term to nucleoside reverse transcriptase inhibitors (NRTIs). Cardiovasc. Toxicol. 2012, 12, 123–134. [Google Scholar] [CrossRef]
- Hung, K.M.; Chen, P.C.; Hsieh, H.C.; Calkins, M.J. Mitochondrial defects arise from nucleoside/nucleotide reverse transcriptase inhibitors in neurons: Potential contribution to HIV-associated neurocognitive disorders. Biochim. Biophys. Acta Mol. Basis Dis. 2017, 1863, 406–413. [Google Scholar] [CrossRef]
- Sanchez, A.B.; Varano, G.P.; de Rozieres, C.M.; Maung, R.; Catalan, I.C.; Dowling, C.C.; Sejbuk, N.E.; Hoefer, M.M.; Kaul, M. Antiretrovirals, Methamphetamine, and HIV-1 envelope protein gp120 compromise neuronal energy homeostasis in association with various degrees of synaptic and neuritic damage. Antimicrob. Agents Chemother. 2016, 60, 168–179. [Google Scholar] [CrossRef] [PubMed]
- Vayssiere, J.L.; Cordeau-Lossouarn, L.; Larcher, J.C.; Basseville, M.; Gros, F.; Croizat, B. Participation of the mitochondrial genome in the differentiation of neuroblastoma cells. In Vitro Cell Dev. Biol. 1992, 28A, 763–772. [Google Scholar] [CrossRef] [PubMed]
- Uittenbogaard, M.; Brantner, C.A.; Chiaramello, A. Epigenetic modifiers promote mitochondrial biogenesis and oxidative metabolism leading to enhanced differentiation of neuroprogenitor cells. Cell Death Dis. 2018, 9, 360. [Google Scholar] [CrossRef] [PubMed]
- Agostini, M.; Romeo, F.; Inoue, S.; Niklison-Chirou, M.V.; Elia, A.J.; Dinsdale, D.; Morone, N.; Knight, R.A.; Mak, T.W.; Melino, G. Metabolic reprogramming during neuronal differentiation. Cell Death Differ. 2016, 23, 1502–1514. [Google Scholar] [CrossRef] [PubMed]
- Nagiec, E.E.; Wu, L.; Swaney, S.M.; Chosay, J.G.; Ross, D.E.; Brieland, J.K.; Leach, K.L. Oxazolidinones inhibit cellular proliferation via inhibition of mitochondrial protein synthesis. Antimicrob. Agents Chemother. 2005, 49, 3896–3902. [Google Scholar] [CrossRef] [PubMed]
- Gregoire, M.; Morais, R.; Quilliam, M.A.; Gravel, D. On auxotrophy for pyrimidines of respiration-deficient chick embryo cells. Eur. J. Biochem. 1984, 142, 49–55. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Pooler, A.M.; Albrecht, M.A.; Wurtman, R.J. Dietary uridine-5′-monophosphate supplementation increases potassium-evoked dopamine release and promotes neurite outgrowth in aged rats. J. Mol. Neurosci. 2005, 27, 137–145. [Google Scholar] [CrossRef]
- Pesini, A.; Iglesias, E.; Garrido, N.; Bayona-Bafaluy, M.P.; Montoya, J.; Ruiz-Pesini, E. OXPHOS, pyrimidine nucleotides, and Alzheimer’s disease: A pharmacogenomics approach. J. Alzheimers Dis. 2014, 42, 87–96. [Google Scholar] [CrossRef]
- Morrison, B.E. Discovery of nigral dopaminergic neurogenesis in adult mice. Neural Regen. Res. 2016, 11, 878–881. [Google Scholar] [CrossRef]
- Rice, D.; Barone, S., Jr. Critical periods of vulnerability for the developing nervous system: Evidence from humans and animal models. Environ. Health Perspect. 2000, 108 (Suppl. S3), 511–533. [Google Scholar] [CrossRef]
- Bayer, S.A.; Altman, J.; Russo, R.J.; Zhang, X. Timetables of neurogenesis in the human brain based on experimentally determined patterns in the rat. Neurotoxicology 1993, 14, 83–144. [Google Scholar] [PubMed]
- Dutta, S.; Sengupta, P. Men and mice: Relating their ages. Life Sci. 2016, 152, 244–248. [Google Scholar] [CrossRef] [PubMed]
- Tartaglione, A.M.; Venerosi, A.; Calamandrei, G. Early-Life Toxic insults and onset of sporadic neurodegenerative diseases-an overview of experimental studies. Curr. Top. Behav. Neurosci. 2016, 29, 231–264. [Google Scholar] [CrossRef] [PubMed]
- Mandelbrot, L.; Tubiana, R.; Le Chenadec, J.; Dollfus, C.; Faye, A.; Pannier, E.; Matheron, S.; Khuong, M.A.; Garrait, V.; Reliquet, V.; et al. No perinatal HIV-1 transmission from women with effective antiretroviral therapy starting before conception. Clin. Infect. Dis. 2015, 61, 1715–1725. [Google Scholar] [CrossRef] [PubMed]
- Panel on Treatment of HIV-Infected Pregnant Women and Prevention of Perinatal Transmission. Recommendations for Use of Antiretroviral Drugs in Pregnant HIV-1-Infected Women for Maternal Health and Interventions to Reduce Perinatal HIV Transmission in the United States. Available online: http://aidsinfo.nih.gov/contentfiles/lvguidelines/PerinatalGL.pdf (accessed on 10 October 2016).
- Barret, B.; Tardieu, M.; Rustin, P.; Lacroix, C.; Chabrol, B.; Desguerre, I.; Dollfus, C.; Mayaux, M.J.; Blanche, S.; French Perinatal Cohort Study Group. Persistent mitochondrial dysfunction in HIV-1-exposed but uninfected infants: Clinical screening in a large prospective cohort. AIDS 2003, 17, 1769–1785. [Google Scholar] [CrossRef]
- Tan, K.R.; Fairley, J.K.; Wang, M.; Gutman, J.R. A survey on outcomes of accidental atovaquone-proguanil exposure in pregnancy. Malar. J. 2018, 17, 198. [Google Scholar] [CrossRef]
- Andrejko, K.L.; Mayer, R.C.; Kovacs, S.; Slutsker, E.; Bartlett, E.; Tan, K.R.; Gutman, J.R. The safety of atovaquone-proguanil for the prevention and treatment of malaria in pregnancy: A systematic review. Travel Med. Infect. Dis. 2019, 27, 20–26. [Google Scholar] [CrossRef]
- Sugarman, J.; Colvin, C.; Moran, A.C.; Oxlade, O. Tuberculosis in pregnancy: An estimate of the global burden of disease. Lancet Glob. Health 2014, 2, e710–e716. [Google Scholar] [CrossRef]
- Esmail, H.; Barry, C.E., 3rd; Young, D.B.; Wilkinson, R.J. The ongoing challenge of latent tuberculosis. Philos. Trans. R. Soc. Lond B Biol. Sci. 2014, 369, 20130437. [Google Scholar] [CrossRef]
- Vergani, L.; Prescott, A.R.; Holt, I.J. Rhabdomyosarcoma rho(0) cells: Isolation and characterization of a mitochondrial DNA depleted cell line with ‘muscle-like’ properties. Neuromuscul. Disord. 2000, 10, 454–459. [Google Scholar] [CrossRef]
- Herst, P.M.; Levine, D.M.; Berridge, M.V. Mitochondrial gene knockout HL60rho0 cells show preferential differentiation into monocytes/macrophages. Leuk. Res. 2005, 29, 1163–1170. [Google Scholar] [CrossRef] [PubMed]
- Pfenninger, K.H. Plasma membrane expansion: A neuron’s Herculean task. Nat. Rev. Neurosci. 2009, 10, 251–261. [Google Scholar] [CrossRef] [PubMed]
- Nyberg, G.B.; Balcarcel, R.R.; Follstad, B.D.; Stephanopoulos, G.; Wang, D.I. Metabolic effects on recombinant interferon-gamma glycosylation in continuous culture of Chinese hamster ovary cells. Biotechnol. Bioeng. 1999, 62, 336–347. [Google Scholar] [CrossRef]
- Klivenyi, P.; Gardian, G.; Calingasan, N.Y.; Yang, L.; von Borstel, R.; Saydoff, J.; Browne, S.E.; Beal, M.F. Neuroprotective effects of oral administration of triacetyluridine against MPTP neurotoxicity. Neuromolecul. Med. 2004, 6, 87–92. [Google Scholar] [CrossRef]
- Cansev, M.; Ulus, I.H.; Wang, L.; Maher, T.J.; Wurtman, R.J. Restorative effects of uridine plus docosahexaenoic acid in a rat model of Parkinson’s disease. Neurosci. Res. 2008, 62, 206–209. [Google Scholar] [CrossRef] [PubMed][Green Version]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Iglesias, E.; Bayona-Bafaluy, M.P.; Pesini, A.; Garrido-Pérez, N.; Meade, P.; Gaudó, P.; Jiménez-Salvador, I.; Montoya, J.; Ruiz-Pesini, E. Uridine Prevents Negative Effects of OXPHOS Xenobiotics on Dopaminergic Neuronal Differentiation. Cells 2019, 8, 1407. https://doi.org/10.3390/cells8111407
Iglesias E, Bayona-Bafaluy MP, Pesini A, Garrido-Pérez N, Meade P, Gaudó P, Jiménez-Salvador I, Montoya J, Ruiz-Pesini E. Uridine Prevents Negative Effects of OXPHOS Xenobiotics on Dopaminergic Neuronal Differentiation. Cells. 2019; 8(11):1407. https://doi.org/10.3390/cells8111407
Chicago/Turabian StyleIglesias, Eldris, M. Pilar Bayona-Bafaluy, Alba Pesini, Nuria Garrido-Pérez, Patricia Meade, Paula Gaudó, Irene Jiménez-Salvador, Julio Montoya, and Eduardo Ruiz-Pesini. 2019. "Uridine Prevents Negative Effects of OXPHOS Xenobiotics on Dopaminergic Neuronal Differentiation" Cells 8, no. 11: 1407. https://doi.org/10.3390/cells8111407
APA StyleIglesias, E., Bayona-Bafaluy, M. P., Pesini, A., Garrido-Pérez, N., Meade, P., Gaudó, P., Jiménez-Salvador, I., Montoya, J., & Ruiz-Pesini, E. (2019). Uridine Prevents Negative Effects of OXPHOS Xenobiotics on Dopaminergic Neuronal Differentiation. Cells, 8(11), 1407. https://doi.org/10.3390/cells8111407





