Spatiotemporal Regulation of Nuclear Transport Machinery and Microtubule Organization
Abstract
:1. Introduction
2. Research History of Ran-dependent Transport Machinery and Spindle Microtubule Assembly
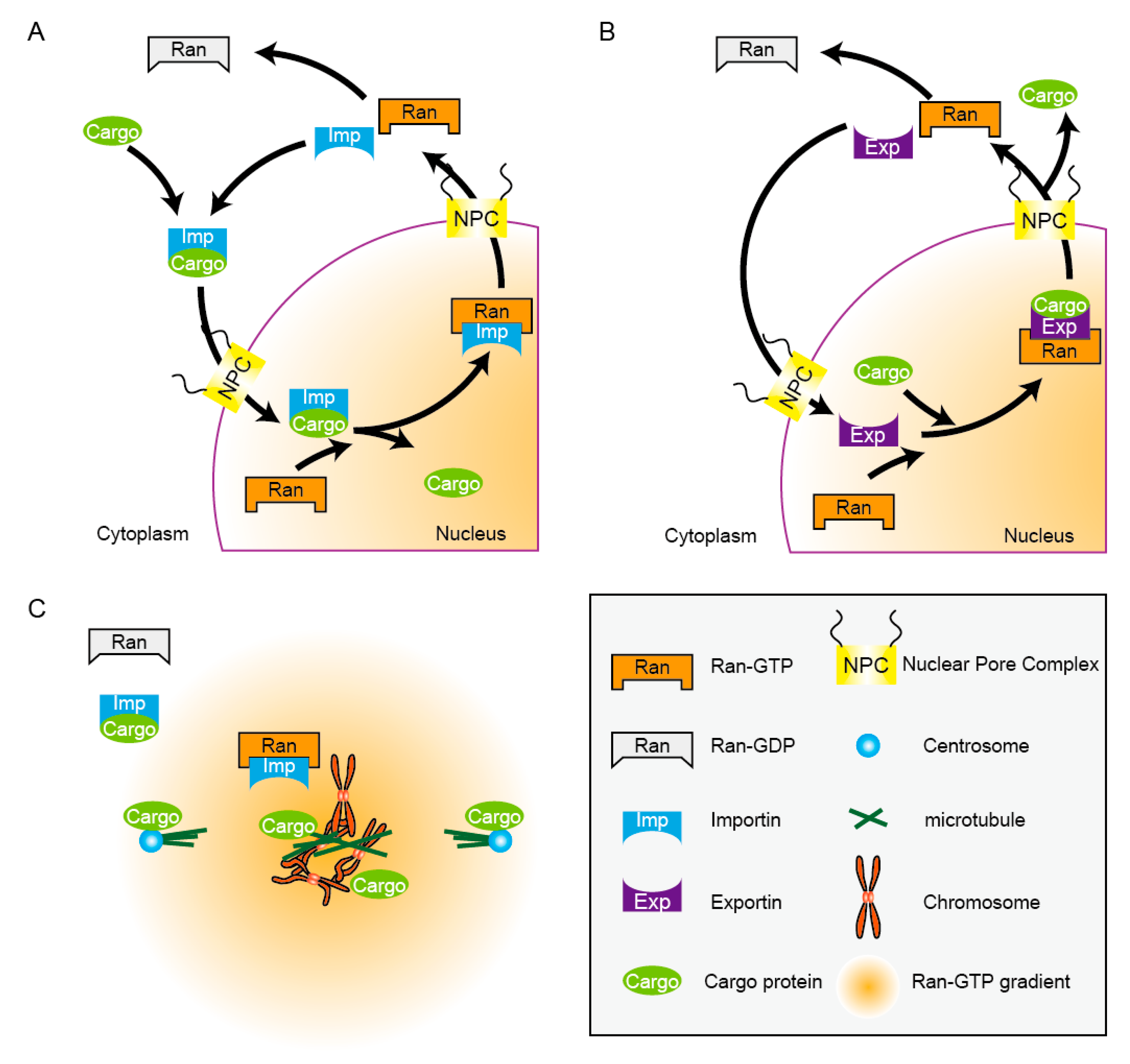
2.1. Ran-Dependent Nucleocytoplamic Transport Machinery
| Cargo | Class of Protein | Nuclear Export Receptor | Organism | Reference |
|---|---|---|---|---|
| Alp14 | MAP | Crm1/Exportin-1 | S. pombe | [31] |
| Klp5/Klp6 | kinesin | Crm1/Exportin-1 | S. pombe | [32] |
| Cyclin B1 | B-type cyclin | Crm1/Exportin-1 | H. sapiens | [33] |
| Survivin | CPC (Chromosome Passenger Complex) component | Crm1/Exportin-1 | M. musculus H. sapiens | [34,35] |
| TPX2 | MAP | Crm1/Exportin-1 | A. thaliana H. sapiens | [36,37] |
| BRCA1 | DNA damage response protein | Crm1/Exportin-1 | H. sapiens | [38] |
| Pho4 | transcription factor | Msn5/Exportin-5 | S. cerevisiae | [39] |
| Srp1 | importin-α | Cse1/CAS | S. cerevisiae | [40] |
2.2. Molecular Gradient of Ran-GTP Navigates Spindle Assembly
| Cargo | Class of Protein | Nuclear Import Receptor | Organism | Reference |
|---|---|---|---|---|
| TPX2 | MAP | Importin-α/β | X. laevis | [51] |
| NuMA | MAP | Importin-α/β | X. laevis | [53,54] |
| HURP | MAP | Importin-β | H. sapiens | [55,56] |
| NuSAP | MAP | Importin-α/β/Importin-7 | X. laevis, H. sapiens | [60] |
| Alp7 (TACC) | MAP | Cut15/Importin-α | S. pombe | [61] |
| Maskin (TACC) | MAP | Importin-β | X. laevis | [62] |
| MCRS1 | MAP | Importin-β | X. laevis | [63] |
| Xnf7 | MAP | Importin-β | X. laevis | [64] |
| Kid | Chromokinesin | Importin-α/β | H. sapiens | [36,65] |
| XCTK2 | Kinesin-14 | Importin-α/β | X. laevis | [66] |
3. Ran-GTP Promotes Formation of Spindle Microtubules in Fission Yeast
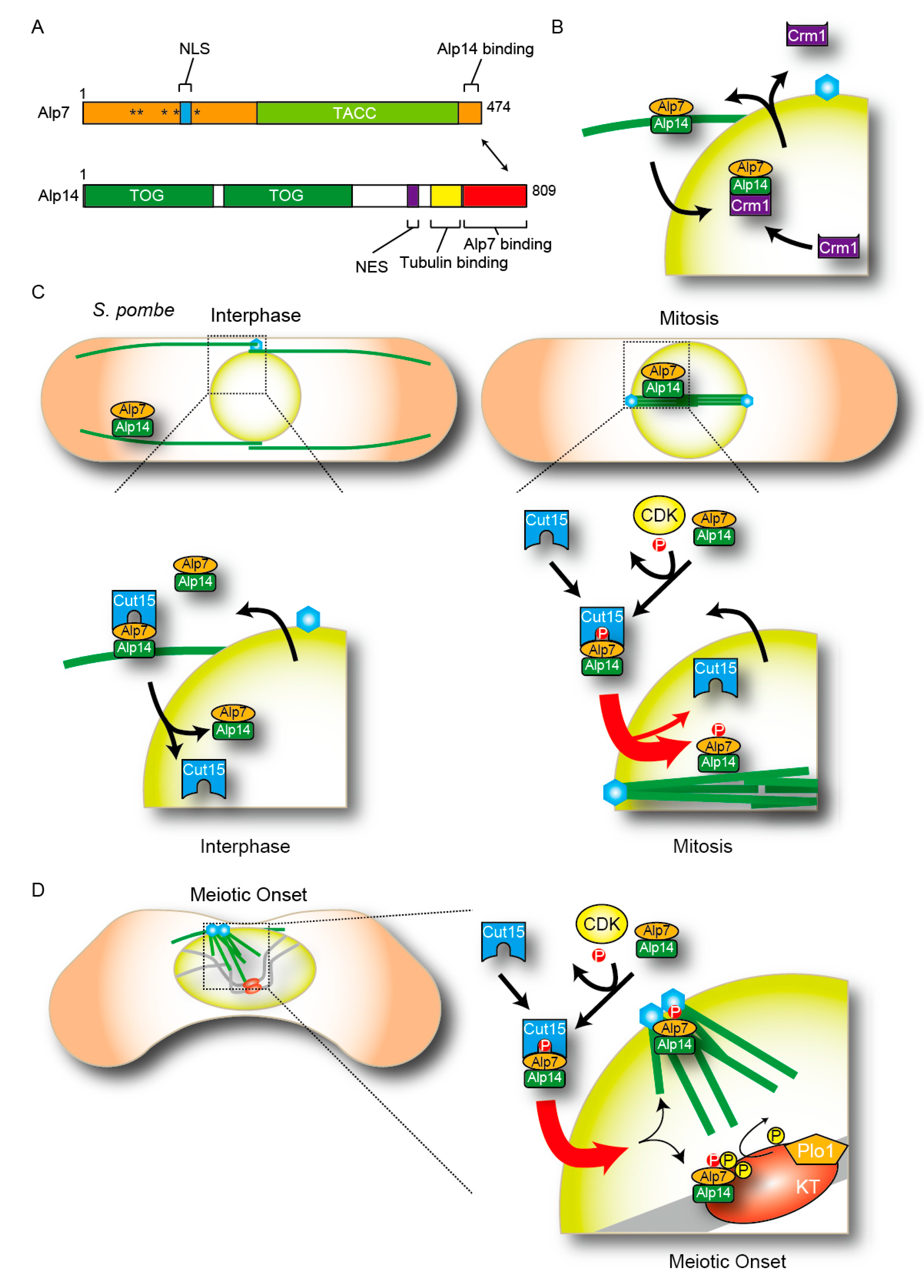
4. Temporal Control of the Nucleocytoplasmic Shuttling
4.1. The Regulation of Temporal Nuclear Accumulation of Shuttling Proteins during the Cell Cycle

4.2. The Nuclear Envelope Permeability Changes during Mitosis in Several Organisms
5. The Regulation of Translocation of MAPs and Spindle Microtubule Organization
6. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Mitchison, T.; Evans, L.; Schulze, E.; Kirschner, M. Sites of microtubule assembly and disassembly in the mitotic spindle. Cell 1986, 45, 515–527. [Google Scholar] [CrossRef]
- Kirschner, M.; Mitchison, T. Beyond self-assembly: From microtubules to morphogenesis. Cell 1986, 45, 329–342. [Google Scholar] [CrossRef]
- Hayles, J.; Nurse, P. Access: A journey into space. Nat. Rev. Mol. Cell Biol. 2001, 2, 647–656. [Google Scholar] [CrossRef] [PubMed]
- Kiyosue, Y.M. Shaping microtubules into diverse patterns: Molecular connections for setting up both ends. Cytoskeleton 2011, 68, 603–618. [Google Scholar] [CrossRef] [PubMed]
- Hagan, I.; Hyams, J. The use of cell division cycle mutants to investigate the control of microtubule distribution in the fission yeast Schizosaccharomyces pombe. J. Cell Sci. 1988, 89, 343–357. [Google Scholar] [PubMed]
- Sato, M.; Toda, T. Space shuttling in the cell: Nucleocytoplasmic transport and microtubule organization during the cell cycle. Nucleus 2010, 1, 231–236. [Google Scholar] [CrossRef] [PubMed]
- Izaurralde, E.; Adam, S. Transport of macromolecules between the nucleus and the cytoplasm. RNA 1998, 4, 351–364. [Google Scholar] [PubMed]
- Ohno, M.; Fornerod, M.; Mattaj, I.W. Nucleocytoplasmic transport: The last 200 nanometers. Cell 1998, 92, 327–336. [Google Scholar] [CrossRef]
- Sazer, S.; Dasso, M. The ran decathlon: Multiple roles of Ran. J. Cell Sci. 2000, 113, 1111–1118. [Google Scholar] [PubMed]
- Moore, J.D. The Ran-GTPase and cell-cycle control. Bioessays 2001, 23, 77–85. [Google Scholar] [CrossRef]
- Dasso, M. Running on Ran: Nuclear transport and the mitotic spindle. Cell 2001, 104, 321–324. [Google Scholar] [CrossRef]
- Clarke, P.R.; Zhang, C. Ran GTPase: A master regulator of nuclear structure and function during the eukaryotic cell division cycle? Trends Cell Biol. 2001, 11, 366–371. [Google Scholar] [CrossRef]
- Drivas, G.; Shih, A.; Coutavas, E.; Rush, M. Characterization of four novel ras-like genes expressed in a human teratocarcinoma cell line. Mol. Cell. Biol. 1990, 10, 1793–1798. [Google Scholar] [PubMed]
- Ohtsubo, M.; Okazaki, H.; Nishimoto, T. The RCC1 protein, a regulator for the onset of chromosome condensation locates in the nucleus and binds to DNA. J. Cell Biol. 1989, 109, 1389–1397. [Google Scholar] [CrossRef] [PubMed]
- Bischoff, F.R.; Ponstingl, H. Catalysis of guanine nucleotide exchange on Ran by the mitotic regulator RCC1. Nature 1991, 354, 80–82. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, T.; Beach, D. Premature initiation of mitosis in yeast lacking RCC1 or an interacting GTPase. Cell 1991, 66, 347–360. [Google Scholar] [CrossRef]
- Bischoff, F.; Krebber, H.; Smirnova, E.; Dong, W. Co-activation of RanGTPase and inhibition of GTP dissociation by Ran-GTP binding protein RanBP1. EMBO J. 1995, 14, 705–715. [Google Scholar] [PubMed]
- Bischoff, F.; Klebe, C.; Kretschmer, J. RanGAP1 induces GTPase activity of nuclear Ras-related Ran. Proc. Natl. Acad. Sci. USA 1994, 91, 2587–2591. [Google Scholar] [CrossRef] [PubMed]
- Melchior, F.; Paschal, B.; Evans, J.; Gerace, L. Inhibition of nuclear protein import by nonhydrolyzable analogues of GTP and identification of the small GTPase Ran/TC4 as an essential transport factor. J. Cell Biol. 1993, 123, 1649–1659. [Google Scholar] [CrossRef] [PubMed]
- Moore, M.S.; Blobel, G. The GTP-binding protein Ran/TC4 is required for protein import into the nucleus. Nature 1993, 365, 661–663. [Google Scholar] [CrossRef] [PubMed]
- Rexach, M.; Blobel, G. Protein import into nuclei: Association and dissociation reactions involving transport substrate, transport factors, and nucleoporins. Cell 1995, 83, 683–692. [Google Scholar] [CrossRef]
- Görlich, D. Nuclear protein import. Curr. Opin. Cell Biol. 1997, 9, 412–419. [Google Scholar] [CrossRef]
- Ström, A.C.; Weis, K. Importin-ß-like nuclear transport receptors. Genome Biol 2001, 2. [Google Scholar] [CrossRef]
- Fried, H.; Kutay, U. Nucleocytoplasmic transport: Taking an inventory. Cell. Mol. Life Sci. 2003, 60, 1659–1688. [Google Scholar] [CrossRef] [PubMed]
- Wente, S.R.; Rout, M.P. The Nuclear Pore Complex and Nuclear Transport. Cold Spring Harb. Perspect. Biol. 2010, 2. [Google Scholar] [CrossRef] [PubMed]
- Chi, N.; Adam, E.; Visser, G. RanBP1 stabilizes the interaction of Ran with p97 nuclear protein import. J. Cell Biol. 1996, 135, 559–569. [Google Scholar] [CrossRef] [PubMed]
- Görlich, D.; Pante, N.; Kutay, U.; Aebi, U. Identification of different roles for RanGDP and RanGTP in nuclear protein import. EMBO J. 1996, 15, 5584–5594. [Google Scholar] [PubMed]
- Fornerod, M.; Ohno, M.; Yoshida, M.; Mattaj, I. CRM1 is an export receptor for leucine-rich nuclear export signals. Cell 1997, 90, 1051–1060. [Google Scholar] [CrossRef]
- Bischoff, F.R.; Görlich, D. RanBP1 is crucial for the release of RanGTP from importin β-related nuclear transport factors. FEBS Lett. 1997, 419, 249–254. [Google Scholar] [CrossRef]
- Koyama, M.; Matsuura, Y. An allosteric mechanism to displace nuclear export cargo from CRM1 and RanGTP by RanBP1. EMBO J. 2010, 29, 2002–2013. [Google Scholar] [CrossRef] [PubMed]
- Okada, N.; Toda, T.; Yamamoto, M. CDK-dependent phosphorylation of Alp7-Alp14 (TACC-TOG) promotes its nuclear accumulation and spindle microtubule assembly. Mol. Biol. Cell 2014, 25, 1969–1982. [Google Scholar] [CrossRef] [PubMed]
- Unsworth, A.; Masuda, H.; Dhut, S.; Toda, T. Fission yeast kinesin-8 Klp5 and Klp6 are interdependent for mitotic nuclear retention and required for proper microtubule dynamics. Mol. Biol. Cell 2008, 19, 5104–5115. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Bardes, E.; Moore, J.; Brennan, J. Control of cyclin B1 localization through regulated binding of the nuclear export factor CRM1. Genes Dev. 1998, 12, 2131–2143. [Google Scholar] [CrossRef] [PubMed]
- Knauer, S.K.; Bier, C.; Habtemichael, N.; Stauber, R.H. The Survivin-Crm1 interaction is essential for chromosomal passenger complex localization and function. EMBO Rep. 2006, 7, 1259–1265. [Google Scholar] [CrossRef] [PubMed]
- Stauber, R.H.; Rabenhorst, U.; Rekik, A.; Engels, K.; Bier, C.; Knauer, S.K. Nucleocytoplasmic Shuttling and the Biological Activity of Mouse Survivin are Regulated by an Active Nuclear Export Signal. Traffic 2006, 7, 1461–1472. [Google Scholar] [CrossRef] [PubMed]
- Trieselmann, N.; Armstrong, S.; Rauw, J. Ran modulates spindle assembly by regulating a subset of TPX2 and Kid activities including Aurora A activation. J. Cell Sci. 2003, 116, 4791–4798. [Google Scholar] [CrossRef] [PubMed]
- Vos, J.W.; Pieuchot, L.; Evrard, J.L.; Janski, N.; Bergdoll, M.; de Ronde, D.; Perez, L.H.; Sardon, T.; Vernos, I.; Schmit, A.C. The Plant TPX2 Protein Regulates Prospindle Assembly before Nuclear Envelope Breakdown. Plant Cell 2008, 20, 2783–2797. [Google Scholar] [CrossRef] [PubMed]
- Brodie, K.M.; Henderson, B.R. Characterization of BRCA1 Protein Targeting, Dynamics, and Function at the Centrosome: A role for the nuclear export signal, crm1, and aurora a kinase. J. Biol. Chem. 2012, 287, 7701–7716. [Google Scholar] [CrossRef] [PubMed]
- Kaffman, A.; Rank, N.M.; O’Neill, E.M.; Huang, L.S.; O’Shea, E.K. The receptor Msn5 exports the phosphorylated transcription factor Pho4 out of the nucleus. Nature 1998, 396, 482–486. [Google Scholar] [PubMed]
- Solsbacher, J.; Maurer, P.; Bischoff, F.; Schlenstedt, G. Cse1p is involved in export of yeast importin α from the nucleus. Mol. Cell. Biol. 1998, 18, 6805–6815. [Google Scholar] [PubMed]
- Karsenti, E.; Newport, J.; Kirschner, M. Respective roles of centrosomes and chromatin in the conversion of microtubule arrays from interphase to metaphase. J. Cell Biol. 1984, 99, 47s–54s. [Google Scholar] [CrossRef] [PubMed]
- Carazo-Salas, R.E.; Guarguaglini, G.; Gruss, O.J.; Segref, A.; Karsenti, E.; Mattaj, I.W. Generation of GTP-bound Ran by RCC1 is required for chromatin-induced mitotic spindle formation. Nature 1999, 400, 178–181. [Google Scholar] [PubMed]
- Kaláb, P.; Pu, R.T.; Dasso, M. The Ran GTPase regulates mitotic spindle assembly. Curr. Biol. 1999, 9, 481–484. [Google Scholar] [CrossRef]
- Ohba, T.; Nakamura, M.; Nishitani, H.; Nishimoto, T. Self-organization of microtubule asters induced in Xenopus egg extracts by GTP-bound Ran. Science 1999, 284, 1356–1358. [Google Scholar] [CrossRef] [PubMed]
- Wilde, A.; Zheng, Y. Stimulation of microtubule aster formation and spindle assembly by the small GTPase Ran. Science 1999, 284, 1359–1362. [Google Scholar] [CrossRef] [PubMed]
- Kahana, J.A.; Cleveland, D.W. Beyond Nuclear Transport: Ran-GTP as a Determinant of Spindle Assembly. J. Cell Biol. 1999, 146, 1205–1210. [Google Scholar] [CrossRef] [PubMed]
- Carazo-Salas, R.E.; Gruss, O.J.; Mattaj, I.W.; Karsenti, E. Ran-GTP coordinates regulation of microtubule nucleation and dynamics during mitotic-spindle assembly. Nat. Cell Biol. 2001, 3, 228–234. [Google Scholar] [CrossRef] [PubMed]
- Caudron, M.; Bunt, G.; Bastiaens, P.; Karsenti, E. Spatial coordination of spindle assembly by chromosome-mediated signaling gradients. Science 2005, 309, 1373–1376. [Google Scholar] [CrossRef] [PubMed]
- Kaláb, P.; Weis, K.; Heald, R. Visualization of a Ran-GTP gradient in interphase and mitotic Xenopus egg extracts. Science 2002, 295, 2452–2456. [Google Scholar] [CrossRef] [PubMed]
- Kaláb, P.; Pralle, A.; Isacoff, E.Y.; Heald, R.; Weis, K. Analysis of a RanGTP-regulated gradient in mitotic somatic cells. Nat. Cell Biol. 2006, 440, 697–701. [Google Scholar] [CrossRef] [PubMed]
- Gruss, O.J.; Carazo-Salas, R.E.; Schatz, C.A.; Guarguaglini, G.; Kast, J.; Wilm, M.; Le Bot, N.; Vernos, I.; Karsenti, E.; Mattaj, I.W. Ran induces spindle assembly by reversing the inhibitory effect of importin α on TPX2 activity. Cell 2001, 104, 83–93. [Google Scholar] [CrossRef]
- Gruss, O.J.; Vernos, I. The mechanism of spindle assembly functions of Ran and its target TPX2. J. Cell Biol. 2004, 166, 949–955. [Google Scholar] [CrossRef] [PubMed]
- Nachury, M.V.; Maresca, T.J.; Salmon, W.C.; Waterman-Storer, C.M.; Heald, R.; Weis, K. Importin β is a mitotic target of the small GTPase Ran in spindle assembly. Cell 2001, 104, 95–106. [Google Scholar] [CrossRef]
- Wiese, C.; Wilde, A.; Moore, M.; Adam, S.; Merdes, A. Role of importin-β in coupling Ran to downstream targets in microtubule assembly. Science 2001, 291, 653–656. [Google Scholar] [CrossRef] [PubMed]
- Koffa, M.D.; Casanova, C.M.; Santarella, R.; Köcher, T.; Wilm, M.; Mattaj, I.W. HURP is part of a Ran-dependent complex involved in spindle formation. Curr. Biol. 2006, 16, 743–754. [Google Scholar] [CrossRef] [PubMed]
- Silljé, H.H.; Nagel, S.; Körner, R.; Nigg, E.A. HURP is a Ran-importin β-regulated protein that stabilizes kinetochore microtubules in the vicinity of chromosomes. Curr. Biol. 2006, 16, 731–742. [Google Scholar] [CrossRef] [PubMed]
- Askjaer, P.; Galy, V.; Hannak, E. Ran GTPase Cycle and Importins α and β Are Essential for Spindle Formation and Nuclear Envelope Assembly in LivingCaenorhabditis elegans Embryos. Mol. Biol. Cell 2002, 13, 4355–4370. [Google Scholar] [CrossRef] [PubMed]
- Silverman-Gavrila, R.V.; Wilde, A. Ran is required before metaphase for spindle assembly and chromosome alignment and after metaphase for chromosome segregation and spindle midbody organization. Mol. Biol. Cell 2006, 17, 2069–2080. [Google Scholar] [CrossRef] [PubMed]
- Gruss, O.J.; Wittmann, M.; Yokoyama, H.; Pepperkok, R.; Kufer, T.; Sillje, H.; Karsenti, E.; Mattaj, I.W.; Vernos, I. Chromosome-induced microtubule assembly mediated by TPX2 is required for spindle formation in HeLa cells. Nat. Cell Biol. 2002, 4, 871–879. [Google Scholar] [CrossRef] [PubMed]
- Ribbeck, K.; Groen, A.C.; Santarella, R.; Bohnsack, M.T.; Raemaekers, T.; Köcher, T.; Gentzel, M.; Görlich, D.; Wilm, M.; Carmeliet, G.; et al. NuSAP, a Mitotic RanGTP Target That Stabilizes and Cross-links Microtubules. Mol. Biol. Cell 2006, 17, 2646–2660. [Google Scholar] [CrossRef] [PubMed]
- Sato, M.; Toda, T. Alp7/TACC is a crucial target in Ran-GTPase-dependent spindle formation in fission yeast. Nature 2007, 447, 334–337. [Google Scholar] [CrossRef] [PubMed]
- Albee, A.J.; Tao, W.; Wiese, C. Phosphorylation of Maskin by Aurora-A Is Regulated by RanGTP and Importin beta. J. Biol. Chem. 2006, 281, 38293–38301. [Google Scholar] [CrossRef] [PubMed]
- Meunier, S.; Vernos, I. K-fibre minus ends are stabilized by a RanGTP-dependent mechanism essential for functional spindle assembly. Nat. Cell Biol. 2011, 13, 1406–1414. [Google Scholar] [CrossRef] [PubMed]
- Maresca, T.J.; Niederstrasser, H.; Weis, K.; Heald, R. Xnf7 Contributes to Spindle Integrity through Its Microtubule-Bundling Activity. Curr. Biol. 2005, 15, 1755–1761. [Google Scholar] [CrossRef] [PubMed]
- Tahara, K.; Takagi, M.; Ohsugi, M.; Sone, T.; Nishiumi, F.; Maeshima, K.; Horiuchi, Y.; Tokai-Nishizumi, N.; Imamoto, F.; Yamamoto, T.; et al. Importin-β and the small guanosine triphosphatase Ran mediate chromosome loading of the human chromokinesin Kid. J. Cell Biol. 2008, 180, 493–506. [Google Scholar] [CrossRef] [PubMed]
- Ems-McClung, S.; Zheng, Y.; Walczak, C. Importin α/β and Ran-GTP regulate XCTK2 microtubule binding through a bipartite nuclear localization signal. Mol. Biol. Cell 2004, 15, 46–57. [Google Scholar] [CrossRef] [PubMed]
- Hagan, I.M. The fission yeast microtubule cytoskeleton. J. Cell Sci. 1998, 111, 1603–1612. [Google Scholar] [PubMed]
- Peset, I.; Vernos, I. The TACC proteins: TACC-ling microtubule dynamics and centrosome function. Trends Cell Biol. 2008, 18, 379–388. [Google Scholar] [CrossRef] [PubMed]
- Ohkura, H.; Garcia, M.A.; Toda, T. Dis1/TOG universal microtubule adaptors-one MAP for all? J. Cell Sci. 2001, 114, 3805–3812. [Google Scholar] [PubMed]
- Sato, M.; Vardy, L.; Garcia, M.A.; Koonrugsa, N.; Toda, T. Interdependency of fission yeast Alp14/TOG and coiled coil protein Alp7 in microtubule localization and bipolar spindle formation. Mol. Biol. Cell 2004, 15, 1609–1622. [Google Scholar] [CrossRef] [PubMed]
- Al-Bassam, J.; Kim, H.; Flor-Parra, I.; Lal, N.; Velji, H.; Chang, F. Fission yeast Alp14 is a dose-dependent plus end-tracking microtubule polymerase. Mol. Biol. Cell 2012, 23, 2878–2890. [Google Scholar] [CrossRef] [PubMed]
- Brouhard, G.J.; Stear, J.H.; Noetzel, T.L.; Al-Bassam, J.; Kinoshita, K.; Harrison, S.C.; Howard, J.; Hyman, A.A. XMAP215 Is a Processive Microtubule Polymerase. Cell 2008, 132, 79–88. [Google Scholar] [CrossRef] [PubMed]
- Zheng, F.; Li, T.; Jin, D.; Syrovatkina, V. Csi1p recruits alp7p/TACC to the spindle pole bodies for bipolar spindle formation. Mol. Biol. Cell 2014, 25, 2750–2760. [Google Scholar] [CrossRef] [PubMed]
- Kakui, Y.; Sato, M.; Okada, N.; Toda, T.; Yamamoto, M. Microtubules and Alp7-Alp14 (TACC-TOG) reposition chromosomes before meiotic segregation. Nat. Cell Biol. 2013, 15, 786–796. [Google Scholar] [CrossRef] [PubMed]
- Sato, M.; Okada, N.; Kakui, Y.; Yamamoto, M.; Yoshida, M.; Toda, T. Nucleocytoplasmic transport of Alp7/TACC organizes spatiotemporal microtubule formation in fission yeast. EMBO Rep. 2009, 10, 1161–1167. [Google Scholar] [CrossRef] [PubMed]
- Tang, N.; Takada, H.; Hsu, K. The internal loop of fission yeast Ndc80 binds Alp7/TACC-Alp14/TOG and ensures proper chromosome attachment. Mol. Biol. Cell 2013, 24, 1122–1133. [Google Scholar] [CrossRef] [PubMed]
- Tang, N.H.; Okada, N.; Fong, C.S.; Arai, K.; Sato, M.; Toda, T. Targeting Alp7/TACC to the spindle pole body is essential for mitotic spindle assembly in fission yeast. FEBS Lett. 2014, 588, 2814–2821. [Google Scholar] [CrossRef] [PubMed]
- Kudo, N.; Wolff, B.; Sekimoto, T.; Schreiner, E.P.; Yoneda, Y.; Yanagida, M.; Horinouchi, S.; Yoshida, M. Leptomycin B inhibition of signal-mediated nuclear export by direct binding to CRM1. Exp. Cell Res. 1998, 242, 540–547. [Google Scholar] [CrossRef] [PubMed]
- Garcia, M.; Vardy, L.; Koonrugsa, N. Fission yeast ch-TOG/XMAP215 homologue Alp14 connects mitotic spindles with the kinetochore and is a component of the Mad2-dependent spindle checkpoint. EMBO J. 2001, 20, 3389–3401. [Google Scholar] [CrossRef] [PubMed]
- Funabiki, H.; Hagan, I.; Uzawa, S. Cell cycle-dependent specific positioning and clustering of centromeres and telomeres in fission yeast. J. Cell Biol. 1993, 121, 961–976. [Google Scholar] [CrossRef] [PubMed]
- Chikashige, Y.; Ding, D.; Funabiki, H.; Haraguchi, T. Telomere-led premeiotic chromosome movement in fission yeast. Science 1994, 264, 270–273. [Google Scholar] [CrossRef] [PubMed]
- Chikashige, Y.; Ding, D.Q.; Imai, Y.; Yamamoto, M.; Haraguchi, T.; Hiraoka, Y. Meiotic nuclear reorganization: Switching the position of centromeres and telomeres in the fission yeast Schizosaccharomyces pombe. EMBO J. 1997, 16, 193–202. [Google Scholar] [CrossRef] [PubMed]
- Komeili, A.; O’Shea, E. Roles of phosphorylation sites in regulating activity of the transcription factor Pho4. Science 1999, 284, 977–980. [Google Scholar] [CrossRef] [PubMed]
- Oshima, Y. The phosphatase system in Saccharomyces cerevisiae. Genes Genet. Syst. 1997, 72, 323–334. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, E.; Kaffman, A.; Jolly, E.; O’Shea, E. Regulation of PHO4 nuclear localization by the PHO80-PHO85 cyclin-CDK complex. Science 1996, 271, 209–212. [Google Scholar] [CrossRef] [PubMed]
- Schneider, K.; Smith, R.; O’Shea, E. Phosphate-regulated inactivation of the kinase PHO80-PHO85 by the CDK inhibitor PHO81. Science 1994, 266, 122–126. [Google Scholar] [CrossRef] [PubMed]
- Kosugi, S.; Hasebe, M.; Tomita, M. Systematic identification of cell cycle-dependent yeast nucleocytoplasmic shuttling proteins by prediction of composite motifs. Proc. Natl. Acad. Sci. USA 2009, 106, 10171–10176. [Google Scholar] [CrossRef] [PubMed]
- Róna, G.; Borsos, M.; Ellis, J.J.; Mehdi, A.M.; Christie, M.; Környei, Z.; Neubrandt, M.; Tóth, J.; Bozóky, Z.; Buday, L. Dynamics of re-constitution of the human nuclear proteome after cell division is regulated by NLS-adjacent phosphorylation. Cell Cycle 2014, 13, 3551–3564. [Google Scholar] [CrossRef] [PubMed]
- Marelli, M.; Aitchison, J.; Wozniak, R. Specific binding of the karyopherin Kap121p to a subunit of the nuclear pore complex containing Nup53p, Nup59p, and Nup170p. J. Cell Biol. 1998, 143, 1813–1830. [Google Scholar] [CrossRef] [PubMed]
- Makhnevych, T.; Lusk, C.P.; Anderson, A.M.; Aitchison, J.D.; Wozniak, R.W. Cell cycle regulated transport controlled by alterations in the nuclear pore complex. Cell 2003, 115, 813–823. [Google Scholar] [CrossRef]
- Wu, Z.; Jiang, Q.; Clarke, P.; Zhang, C. Phosphorylation of Crm1 by CDK1-cyclin-B promotes Ran-dependent mitotic spindle assembly. J. Cell Sci. 2013, 126, 3417–3428. [Google Scholar] [CrossRef] [PubMed]
- Hayashi, H.; Kimura, K.; Kimura, A. Localized accumulation of tubulin during semi-open mitosis in the Caenorhabditis elegans embryo. Mol. Biol. Cell 2012, 23, 1688–1699. [Google Scholar] [CrossRef] [PubMed]
- De Souza, C.P.C.; Osmani, A.H.; Hashmi, S.B.; Osmani, S.A. Partial Nuclear Pore Complex Disassembly during Closed Mitosis in Aspergillus nidulans. Curr. Biol. 2004, 14, 1973–1984. [Google Scholar] [CrossRef] [PubMed]
- Arai, K.; Sato, M.; Tanaka, K.; Yamamoto, M. Nuclear compartmentalization is abolished during fission yeast meiosis. Curr. Biol. 2010, 20, 1913–1918. [Google Scholar] [CrossRef] [PubMed]
- Asakawa, H.; Kojidani, T.; Mori, C.; Osakada, H.; Sato, M.; Ding, D.Q.; Hiraoka, Y.; Haraguchi, T. Virtual Breakdown of the Nuclear Envelope in Fission Yeast Meiosis. Curr. Biol. 2010, 20, 1919–1925. [Google Scholar] [CrossRef] [PubMed]
- Shimoda, C. Forespore membrane assembly in yeast: Coordinating SPBs and membrane trafficking. J. Cell Sci. 2004, 117, 389–396. [Google Scholar] [CrossRef] [PubMed]
- Watson, M. The nuclear envelope: Its structure and relation to cytoplasmic membranes. J. Biophys. Biochem. Cytol. 1955, 1, 257–279. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, Y.; Meerbrey, K.; Chong, J.; Torii, Y. Nuclear shape, growth and integrity in the closed mitosis of fission yeast depend on the Ran-GTPase system, the spindle pole body and the endoplasmic reticulum. J. Cell Sci. 2009, 122, 2464–2472. [Google Scholar] [CrossRef] [PubMed]
- Aoki, K.; Hayashi, H.; Furuya, K.; Sato, M.; Takagi, T.; Osumi, M.; Kimura, A.; Niki, H. Breakage of the nuclear envelope by an extending mitotic nucleus occurs during anaphase in Schizosaccharomyces japonicus. Genes Cells 2011, 16, 911–926. [Google Scholar] [CrossRef] [PubMed]
- O’Day, D.; Budniak, A. Nucleocytoplasmic protein translocation during mitosis in the social amoebozoan Dictyostelium discoideum. Biol. Rev. 2015, 90, 126–141. [Google Scholar] [CrossRef] [PubMed]
- Leo, M.; Santino, D.; Tikhonenko, I.; Magidson, V. Rules of engagement: Centrosome-nuclear connections in a closed mitotic system. Biol. Open 2012, 1, 1111–1117. [Google Scholar] [CrossRef] [PubMed]
- Nigg, E.; Blangy, A.; Lane, H. Dynamic changes in nuclear architecture during mitosis: On the role of protein phosphorylation in spindle assembly and chromosome segregation. Exp. Cell Res. 1996, 229, 174–180. [Google Scholar] [CrossRef] [PubMed]
- Andersen, S.S. Spindle assembly and the art of regulating microtubule dynamics by MAPs and Stathmin/Op18. Trends Cell Biol. 2000, 10, 261–267. [Google Scholar] [CrossRef]
- Sedbrook, J.C. MAPs in plant cells: Delineating microtubule growth dynamics and organization. Curr. Opin. Plant Biol. 2004, 7, 632–640. [Google Scholar] [CrossRef] [PubMed]
- Peterman, E.J.G.; Scholey, J.M. Mitotic Microtubule Crosslinkers: Insights from Mechanistic Studies. Curr. Biol. 2009, 19, R1089–R1094. [Google Scholar] [CrossRef] [PubMed]
- Pellman, D.; Bagget, M.; Tu, Y.; Fink, G.R.; Tu, H. Two microtubule-associated proteins required for anaphase spindle movement in Saccharomyces cerevisiae. J. Cell Biol. 1995, 130, 1373–1385. [Google Scholar] [CrossRef] [PubMed]
- Loiodice, I.; Staub, J.; Setty, T.G.; Nguyen, N.-P.T.; Paoletti, A.; Tran, P. Ase1p organizes antiparallel microtubule arrays during interphase and mitosis in fission yeast. Mol. Biol. Cell 2005, 16, 1756–1768. [Google Scholar] [CrossRef] [PubMed]
- Yamashita, A.; Sato, M.; Fujita, A.; Yamamoto, M.; Toda, T. The roles of fission yeast ase1 in mitotic cell division, meiotic nuclear oscillation, and cytokinesis checkpoint signaling. Mol. Biol. Cell 2005, 16, 1378–1395. [Google Scholar] [CrossRef] [PubMed]
- Fu, C.; Ward, J.J.; Loiodice, I.; Velve-Casquillas, G.; Nedelec, F.J.; Tran, P.T. Phospho-Regulated Interaction between Kinesin-6 Klp9p and Microtubule Bundler Ase1p Promotes Spindle Elongation. Dev. Cell 2009, 17, 257–267. [Google Scholar] [CrossRef] [PubMed]
- Garcia, M.A.; Koonrugsa, N.; Toda, T. Two Kinesin-like Kin I Family Proteins in Fission Yeast Regulate the Establishment of Metaphase and the Onset of Anaphase A. Curr. Biol. 2002, 12, 610–621. [Google Scholar] [CrossRef]
- Chou, H.Y.; Wang, T.H.; Lee, S.C.; Hsu, P.H.; Tsai, M.D.; Chang, C.L.; Jeng, Y.M. Phosphorylation of NuSAP by Cdk1 regulates its interaction with microtubules in mitosis. Cell Cycle 2011, 10, 4083–4089. [Google Scholar] [CrossRef] [PubMed]
- Shim, S.; de Castro, I.; Neumayer, G.; Wang, J. Phosphorylation of Targeting Protein for Xenopus kinesin-like protein 2 (TPX2) at threonine 72 in spindle assembly. J. Biol. Chem. 2015, 290, 9122–9134. [Google Scholar]
© 2015 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Okada, N.; Sato, M. Spatiotemporal Regulation of Nuclear Transport Machinery and Microtubule Organization. Cells 2015, 4, 406-426. https://doi.org/10.3390/cells4030406
Okada N, Sato M. Spatiotemporal Regulation of Nuclear Transport Machinery and Microtubule Organization. Cells. 2015; 4(3):406-426. https://doi.org/10.3390/cells4030406
Chicago/Turabian StyleOkada, Naoyuki, and Masamitsu Sato. 2015. "Spatiotemporal Regulation of Nuclear Transport Machinery and Microtubule Organization" Cells 4, no. 3: 406-426. https://doi.org/10.3390/cells4030406
APA StyleOkada, N., & Sato, M. (2015). Spatiotemporal Regulation of Nuclear Transport Machinery and Microtubule Organization. Cells, 4(3), 406-426. https://doi.org/10.3390/cells4030406






