Single-Cell Transcriptomic Landscape of Cervical Cancer Cell Lines Before and After Chemoradiotherapy
Highlights
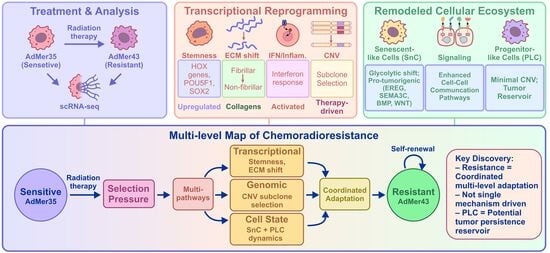
- We established and characterized a unique pair of isogenic cervical cancer cell lines (AdMer35/AdMer43) from a patient with squamous cell carcinoma of the cervix before and after chemoradiotherapy, revealing that resistance is not caused by a single factor but by complex reorganization of cancer cells.
- Single-cell analysis identified key resistance hallmarks: transcriptional reprogramming towards embryonic stemness (HOX, POU5F1), a shift from fibrillar to non-fibrillar collagen expression, and activation of interferon/inflammatory pathways.
- The study maps a remodeled tumor ecosystem with a metabolically reprogrammed, pro-tumorigenic senescent-like cell population and a progenitor-like cell reservoir, identifying them as critical therapeutic targets for eliminating resistant cell pools.
- The identified key pathways, Wnt, TGFβ, BMP, and the specific ligands EREG and SEMA3C, along with the metabolic vulnerabilities of the senescent compartment, provide a robust framework for future therapeutic exploration.
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Lines
2.2. Single-Cell RNA Sequencing
2.3. Single-Cell Transcriptome Analysis
2.4. Cytokine Concentrations in AdMer35 and AdMer43 Cell Media
3. Results and Discussion
3.1. Cervical Cancer Cell Lines AdMer35 and AdMer43 Have Epithelial Cell Morphology
3.2. Single-Cell Transcriptomic Profiling of AdMer35 and AdMer43 Cells
3.3. Differentially Expressed Transcripts in AdMer43 Versus AdMer35 Cervical Cancer Cells
3.4. AdMer35 and AdMer43 Cells Differ in CNV Events in Multiple Chromosomes
3.5. Senescent-like Cells (SnCs) in AdMer43 and AdMer35 Lines
3.6. Cervical Cancer Stem-like Cells in AdMer43 and AdMer35 Lines
3.7. Generalized Data on the Classification of AdMer35 and AdMer43 Cultures
3.8. Validation of Single-Cell Sequencing Data Using an Independent Cytokine/Chemokine Assay in AdMer35 and AdMer43 Cell-Condensed Media
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CC | Cervical Cancer |
| CIN | Cervical Intraepithelial Neoplasia |
| CNV | Copy Number Variation |
| DEG | Differentially Expressed Gene |
| DSB | Double-strand DNA Break |
| ECM | Extracellular Matrix |
| EMT | Epithelial to Mesenchymal Transition |
| HPV | Human Papillomavirus |
| PCA | Principal Component Analysis |
| PLC | Progenitor-like Cell |
| PRC2 | Policomb Repressive Complex 2 |
| ROS | Reactive Oxygen Species |
| RT | Radiotherapy |
| SASP | Senescence Associated Secretory Phenotype |
| SC | Single Cell |
| SLC | Stem-like Cell |
| SnC | Senescent-like Cell |
| TF | Transcription Factor |
| UMAP | Uniform Manifold Approximation and Projection |
| UMI | Unique Molecular Identifier |
| VST | Variance Stabilizing Transformation |
References
- Singh, D.; Vignat, J.; Lorenzoni, V.; Eslahi, M.; Ginsburg, O.; Lauby-Secretan, B.; Arbyn, M.; Basu, P.; Bray, F.; Vaccarella, S. Global Estimates of Incidence and Mortality of Cervical Cancer in 2020: A Baseline Analysis of the WHO Global Cervical Cancer Elimination Initiative. Lancet Glob. Health 2023, 11, e197–e206. [Google Scholar] [CrossRef]
- Wang, M.; Huang, K.; Wong, M.C.S.; Huang, J.; Jin, Y.; Zheng, Z.J. Global Cervical Cancer Incidence by Histological Subtype and Implications for Screening Methods. J. Epidemiol. Glob. Health 2024, 14, 94–101. [Google Scholar] [CrossRef]
- Zhang, Y.; Qiu, K.; Ren, J.; Zhao, Y.; Cheng, P. Roles of Human Papillomavirus in Cancers: Oncogenic Mechanisms and Clinical Use. Signal Transduct. Target. Ther. 2025, 10, 44. [Google Scholar] [CrossRef]
- Łysiak, K.; Łysiak, A. HPV 16 and Its Role in the Development of Cervical Cancer: Prevention, Diagnosis and Treatment. Prospect. Pharm. Sci. 2025. [Google Scholar] [CrossRef]
- Liu, Y.; Ai, H. Comprehensive Insights into Human Papillomavirus and Cervical Cancer: Pathophysiology, Screening, and Vaccination Strategies. Biochim. Biophys. Acta. Rev. Cancer 2024, 1879, 189192. [Google Scholar] [CrossRef]
- Perkins, R.B.; Wentzensen, N.; Guido, R.S.; Schiffman, M. Cervical Cancer Screening: A Review. JAMA 2023, 330, 547–558. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Deng, Y.; Boakye, D.; Tin, M.S.; Lok, V.; Zhang, L.; Lucero-Prisno, D.E.; Xu, W.; Zheng, Z.J.; Elcarte, E.; et al. Global Distribution, Risk Factors, and Recent Trends for Cervical Cancer: A Worldwide Country-Level Analysis. Gynecol. Oncol. 2022, 164, 85–92. [Google Scholar] [CrossRef]
- Yao, G.; Qiu, J.; Zhu, F.; Wang, X. Survival of Patients With Cervical Cancer Treated With Definitive Radiotherapy or Concurrent Chemoradiotherapy According to Histological Subtype: A Systematic Review and Meta-Analysis. Front. Med. 2022, 9, 843262. [Google Scholar] [CrossRef]
- Zhou, J.; Lei, N.; Tian, W.; Guo, R.; Chen, M.; Qiu, L.; Wu, F.; Li, Y.; Chang, L. Recent Progress of the Tumor Microenvironmental Metabolism in Cervical Cancer Radioresistance. Front. Oncol. 2022, 12, 999643. [Google Scholar] [CrossRef] [PubMed]
- Prasanna, P.G.; Citrin, D.E.; Hildesheim, J.; Ahmed, M.M.; Venkatachalam, S.; Riscuta, G.; Xi, D.; Zheng, G.; van Deursen, J.; Goronzy, J.; et al. Therapy-Induced Senescence: Opportunities to Improve Anticancer Therapy. J. Natl. Cancer Inst. 2021, 113, 1285–1298. [Google Scholar] [CrossRef] [PubMed]
- Xing, J.L.; Stea, B. Molecular Mechanisms of Sensitivity and Resistance to Radiotherapy. Clin. Exp. Metastasis 2024, 41, 517–524. [Google Scholar] [CrossRef]
- Liang, M.; Sheng, L.; Ke, Y.; Wu, Z. The Research Progress on Radiation Resistance of Cervical Cancer. Front. Oncol. 2024, 14, 1380448. [Google Scholar] [CrossRef]
- Bhattacharjee, R.; Das, S.S.; Biswal, S.S.; Nath, A.; Das, D.; Basu, A.; Malik, S.; Kumar, L.; Kar, S.; Singh, S.K.; et al. Mechanistic Role of HPV-Associated Early Proteins in Cervical Cancer: Molecular Pathways and Targeted Therapeutic Strategies. Crit. Rev. Oncol. Hematol. 2022, 174, 103675. [Google Scholar] [CrossRef]
- Casamassimi, A.; Federico, A.; Rienzo, M.; Esposito, S.; Ciccodicola, A. Transcriptome Profiling in Human Diseases: New Advances and Perspectives. Int. J. Mol. Sci. 2017, 18, 1652. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Hua, K. Dissecting the Single-Cell Transcriptome Network of Immune Environment Underlying Cervical Premalignant Lesion, Cervical Cancer and Metastatic Lymph Nodes. Front. Immunol. 2022, 13, 897366. [Google Scholar] [CrossRef] [PubMed]
- den Boon, J.A.; Pyeon, D.; Wang, S.S.; Horswill, M.; Schiffman, M.; Sherman, M.; Zuna, R.E.; Wang, Z.; Hewitt, S.M.; Pearson, R.; et al. Molecular Transitions from Papillomavirus Infection to Cervical Precancer and Cancer: Role of Stromal Estrogen Receptor Signaling. Proc. Natl. Acad. Sci. USA 2015, 112, E3255-64. [Google Scholar] [CrossRef]
- Tatarnikova, I.; Talyshev, V.; Sen’kova, A.; Markov, O.; Mikhailova, D.; Pashkovskaya, O.; Krasilnikov, S.; Samoilova, E.; Gainer, T.; Brener, E.; et al. Two paired HPV-59 cervical cancer cell lines with distinct chemoradioresistant phenotypes. Sci. Rep. 2026; in press. [Google Scholar]
- Elias, M.H.; Das, S.; Abdul Hamid, N. Candidate Genes and Pathways in Cervical Cancer: A Systematic Review and Integrated Bioinformatic Analysis. Cancers 2023, 15, 853. [Google Scholar] [CrossRef]
- Chen, T.; Yang, S.; Xu, J.; Lu, W.; Xie, X. Transcriptome Sequencing Profiles of Cervical Cancer Tissues and SiHa Cells. Funct. Integr. Genom. 2020, 20, 211–221. [Google Scholar] [CrossRef]
- Volkova, L.V.; Pashov, A.I.; Omelchuk, N.N. Cervical Carcinoma: Oncobiology and Biomarkers. Int. J. Mol. Sci. 2021, 22, 12571. [Google Scholar] [CrossRef] [PubMed]
- Zuccherato, L.W.; Machado, C.M.T.; Magalhães, W.C.S.; Martins, P.R.; Campos, L.S.; Braga, L.C.; Teixeira-Carvalho, A.; Martins-Filho, O.A.; Franco, T.M.R.F.; Paula, S.O.C.; et al. Cervical Cancer Stem-Like Cell Transcriptome Profiles Predict Response to Chemoradiotherapy. Front. Oncol. 2021, 11, 639339. [Google Scholar] [CrossRef]
- Aibar, S.; González-Blas, C.B.; Moerman, T.; Huynh-Thu, V.A.; Imrichova, H.; Hulselmans, G.; Rambow, F.; Marine, J.-C.; Geurts, P.; Aerts, J.; et al. SCENIC: Single-Cell Regulatory Network Inference and Clustering. Nat. Methods 2017, 14, 1083–1086. [Google Scholar] [CrossRef] [PubMed]
- Jin, S.; Guerrero-Juarez, C.F.; Zhang, L.; Chang, I.; Ramos, R.; Kuan, C.-H.; Myung, P.; Plikus, M.V.; Nie, Q. Inference and Analysis of Cell-Cell Communication Using CellChat. Nat. Commun. 2021, 12, 1088. [Google Scholar] [CrossRef]
- Jin, S.; Plikus, M.V.; Nie, Q. CellChat for Systematic Analysis of Cell-Cell Communication from Single-Cell Transcriptomics. Nat. Protoc. 2025, 20, 180–219. [Google Scholar] [CrossRef]
- Kuleshov, M.V.; Jones, M.R.; Rouillard, A.D.; Fernandez, N.F.; Duan, Q.; Wang, Z.; Koplev, S.; Jenkins, S.L.; Jagodnik, K.M.; Lachmann, A.; et al. Enrichr: A Comprehensive Gene Set Enrichment Analysis Web Server 2016 Update. Nucleic Acids Res. 2016, 44, W90–W97. [Google Scholar] [CrossRef]
- Xie, Z.; Bailey, A.; Kuleshov, M.V.; Clarke, D.J.B.; Evangelista, J.E.; Jenkins, S.L.; Lachmann, A.; Wojciechowicz, M.L.; Kropiwnicki, E.; Jagodnik, K.M.; et al. Gene Set Knowledge Discovery with Enrichr. Curr. Protoc. 2021, 1, e90. [Google Scholar] [CrossRef]
- InferCNV of the Trinity CTAT Project. Available online: https://Github.Com/Broadinstitute/InferCNV (accessed on 20 April 2025).
- Puram, S.V.; Tirosh, I.; Parikh, A.S.; Patel, A.P.; Yizhak, K.; Gillespie, S.; Rodman, C.; Luo, C.L.; Mroz, E.A.; Emerick, K.S.; et al. Single-Cell Transcriptomic Analysis of Primary and Metastatic Tumor Ecosystems in Head and Neck Cancer. Cell 2017, 171, 1611–1624.e24. [Google Scholar] [CrossRef]
- Ono, K.; Fong, D.; Gao, C.; Churas, C.; Pillich, R.; Lenkiewicz, J.; Pratt, D.; Pico, A.R.; Hanspers, K.; Xin, Y.; et al. Cytoscape Web: Bringing Network Biology to the Browser. Nucleic Acids Res. 2025, 53, W203–W212. [Google Scholar] [CrossRef]
- Rosty, C.; Sheffer, M.; Tsafrir, D.; Stransky, N.; Tsafrir, I.; Peter, M.; de Crémoux, P.; de La Rochefordière, A.; Salmon, R.; Dorval, T.; et al. Identification of a Proliferation Gene Cluster Associated with HPV E6/E7 Expression Level and Viral DNA Load in Invasive Cervical Carcinoma. Oncogene 2005, 24, 7094–7104. [Google Scholar] [CrossRef] [PubMed]
- Hao, Y.; Stuart, T.; Kowalski, M.H.; Choudhary, S.; Hoffman, P.; Hartman, A.; Srivastava, A.; Molla, G.; Madad, S.; Fernandez-Granda, C.; et al. Dictionary Learning for Integrative, Multimodal and Scalable Single-Cell Analysis. Nat. Biotechnol. 2024, 42, 293–304. [Google Scholar] [CrossRef]
- Yu, X.; Li, Z.; Bai, R.; Tang, F. Transcriptional Factor 3 Binds to Sirtuin 1 to Activate the Wnt/β-Catenin Signaling in Cervical Cancer. Bioengineered 2022, 13, 12516–12531. [Google Scholar] [CrossRef] [PubMed]
- Shen, L.; Huang, X.; Xie, X.; Su, J.; Yuan, J.; Chen, X. High Expression of SOX2 and OCT4 Indicates Radiation Resistance and an Independent Negative Prognosis in Cervical Squamous Cell Carcinoma. J. Histochem. Cytochem. 2014, 62, 499–509. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Zhang, F.; Li, J.; Yang, D.; Zhao, H.; Yuan, J.; Jia, Y.; Yu, W.; Guo, W.; Zou, L.; et al. GATA2 Promotes Cervical Cancer Progression under the Transcriptional Activation of TRIP4. Cell. Signal. 2025, 132, 111778. [Google Scholar] [CrossRef]
- Lee, T.I.; Jenner, R.G.; Boyer, L.A.; Guenther, M.G.; Levine, S.S.; Kumar, R.M.; Chevalier, B.; Johnstone, S.E.; Cole, M.F.; Isono, K.; et al. Control of Developmental Regulators by Polycomb in Human Embryonic Stem Cells. Cell 2006, 125, 301–313. [Google Scholar] [CrossRef]
- Saha, S.S.; Chowdhury, R.R.; Mondal, N.R.; Roy, S.; Sengupta, S. Expression Signatures of HOX Cluster Genes in Cervical Cancer Pathogenesis: Impact of Human Papillomavirus Type 16 Oncoprotein E7. Oncotarget 2017, 8, 36591–36602. [Google Scholar] [CrossRef]
- Xu, X.; Gong, C.; Wang, Y.; Yin, Z.; Wang, X.; Wu, X.; Fang, Z.; Wei, S. FOXF1 Promotes Ovarian Cancer Metastasis by Facilitating HMGA2-Mediated USP30-Dependent S100A6 Deubiquitination. Biochim. Biophys. Acta Mol. Basis Dis. 2025, 1871, 167633. [Google Scholar] [CrossRef]
- Babadei, O.; Strobl, B.; Müller, M.; Decker, T. Transcriptional Control of Interferon-Stimulated Genes. J. Biol. Chem. 2024, 300, 107771. [Google Scholar] [CrossRef] [PubMed]
- Al Mamun, M.; Mannoor, K.; Cao, J.; Qadri, F.; Song, X. SOX2 in Cancer Stemness: Tumor Malignancy and Therapeutic Potentials. J. Mol. Cell Biol. 2020, 12, 85–98. [Google Scholar] [CrossRef]
- Paskeh, M.D.A.; Mirzaei, S.; Gholami, M.H.; Zarrabi, A.; Zabolian, A.; Hashemi, M.; Hushmandi, K.; Ashrafizadeh, M.; Aref, A.R.; Samarghandian, S. Cervical Cancer Progression Is Regulated by SOX Transcription Factors: Revealing Signaling Networks and Therapeutic Strategies. Biomed. Pharmacother. 2021, 144, 112335. [Google Scholar] [CrossRef]
- Yamaguchi, N.; Wu, Y.G.; Ravetch, E.; Takahashi, M.; Khan, A.G.; Hayashi, A.; Mei, W.; Hsu, D.; Umeda, S.; de Stanchina, E.; et al. A Targetable Secreted Neural Protein Drives Pancreatic Cancer Metastatic Colonization and HIF1α Nuclear Retention. Cancer Discov. 2024, 14, 2489–2508. [Google Scholar] [CrossRef]
- Xu, C.; Tian, G.; Jiang, C.; Xue, H.; Kuerbanjiang, M.; Sun, L.; Gu, L.; Zhou, H.; Liu, Y.; Zhang, Z.; et al. NPTX2 Promotes Colorectal Cancer Growth and Liver Metastasis by the Activation of the Canonical Wnt/β-Catenin Pathway via FZD6. Cell Death Dis. 2019, 10, 217. [Google Scholar] [CrossRef] [PubMed]
- Xiang, W.; Han, L.; Mo, G.; Lin, L.; Yu, X.; Chen, S.; Gao, T.; Huang, C. MicroRNA-96 Is a Potential Tumor Repressor by Inhibiting NPTX2 in Renal Cell Carcinoma. J. Cell. Biochem. 2020, 121, 1504–1513. [Google Scholar] [CrossRef]
- Chen, T.; Liu, Y.; Chen, J.; Zheng, H.; Chen, Q.; Zhao, J. Exosomal MiR-3180-3p Inhibits Proliferation and Metastasisof Non-Small Cell Lung Cancer by Downregulating FOXP4. Thorac. Cancer 2021, 12, 372–381. [Google Scholar] [CrossRef]
- Abe, H.; Kamimura, K.; Okuda, S.; Watanabe, Y.; Inoue, J.; Aoyagi, Y.; Wakai, T.; Kominami, R.; Terai, S. BCL11B Expression in Hepatocellular Carcinoma Relates to Chemosensitivity and Clinical Prognosis. Cancer Med. 2023, 12, 15650–15663. [Google Scholar] [CrossRef]
- Kim, O.; Tran, P.T.; Gal, M.; Lee, S.J.; Na, S.H.; Hwangbo, C.; Lee, J.-H. RAS-stimulated Release of Exosomal MiR-494-3p Promotes the Osteolytic Bone Metastasis of Breast Cancer Cells. Int. J. Mol. Med. 2023, 52, 84. [Google Scholar] [CrossRef]
- Bian, P.; Liu, C.; Hu, W.; Ding, Y.; Qiu, S.; Li, L. Echinacoside Suppresses the Progression of Breast Cancer by Downregulating the Expression of MiR-4306 and MiR-4508. Integr. Cancer Ther. 2021, 20, 15347354211062640. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Cai, H.; Chen, L.; Zhao, W.; Li, P.; Wang, Z.; Li, Z. STAT3 Correlates with Stem Cell-Related Transcription Factors in Cervical Cancer. J. Huazhong Univ. Sci. Technol. [Med. Sci.] 2015, 35, 891–897. [Google Scholar] [CrossRef] [PubMed]
- Necula, L.; Matei, L.; Dragu, D.; Pitica, I.; Neagu, A.; Bleotu, C.; Diaconu, C.C.; Chivu-Economescu, M. Collagen Family as Promising Biomarkers and Therapeutic Targets in Cancer. Int. J. Mol. Sci. 2022, 23, 12415. [Google Scholar] [CrossRef] [PubMed]
- Gordon, M.K.; Hahn, R.A. Collagens. Cell Tissue Res. 2010, 339, 247–257. [Google Scholar] [CrossRef]
- Folkman, J. Antiangiogenesis in Cancer Therapy—Endostatin and Its Mechanisms of Action. Exp. Cell Res. 2006, 312, 594–607. [Google Scholar] [CrossRef]
- Xu, F.; Chang, K.; Ma, J.; Qu, Y.; Xie, H.; Dai, B.; Gan, H.; Zhang, H.; Shi, G.; Zhu, Y.; et al. The Oncogenic Role of COL23A1 in Clear Cell Renal Cell Carcinoma. Sci. Rep. 2017, 7, 9846. [Google Scholar] [CrossRef]
- Zhao, Q.; Liu, C.; Cui, Q.; Luan, X.; Wang, Q.; Zhou, C. MiR-190b Promotes Colorectal Cancer Progression through Targeting Forkhead Box Protein P2. Exp. Ther. Med. 2019, 19, 79–84. [Google Scholar] [CrossRef]
- Ajoolabady, A.; Pratico, D.; Bahijri, S.; Tuomilehto, J.; Uversky, V.N.; Ren, J. Hallmarks of Cellular Senescence: Biology, Mechanisms, Regulations. Exp. Mol. Med. 2025, 57, 1482–1491. [Google Scholar] [CrossRef]
- Yosef, R.; Pilpel, N.; Papismadov, N.; Gal, H.; Ovadya, Y.; Vadai, E.; Miller, S.; Porat, Z.; Ben-Dor, S.; Krizhanovsky, V. p21 maintains senescent cell viability under persistent DNA damage response by restraining JNK and caspase signaling. EMBO J. 2017, 36, 2280–2295. [Google Scholar] [CrossRef]
- Hernandez-Segura, A.; Nehme, J.; Demaria, M. Hallmarks of Cellular Senescence. Trends Cell Biol. 2018, 28, 436–453. [Google Scholar] [CrossRef]
- Valieva, Y.; Ivanova, E.; Fayzullin, A.; Kurkov, A.; Igrunkova, A. Senescence-Associated β-Galactosidase Detection in Pathology. Diagnostics 2022, 12, 2309. [Google Scholar] [CrossRef] [PubMed]
- Coppé, J.-P.; Patil, C.K.; Rodier, F.; Sun, Y.; Muñoz, D.P.; Goldstein, J.; Nelson, P.S.; Desprez, P.-Y.; Campisi, J. Senescence-Associated Secretory Phenotypes Reveal Cell-Nonautonomous Functions of Oncogenic RAS and the p53 Tumor Suppressor. PLoS Biol. 2008, 6, e301. [Google Scholar] [CrossRef]
- Jiang, C.; Liu, G.; Luckhardt, T.; Antony, V.; Zhou, Y.; Carter, A.B.; Thannickal, V.J.; Liu, R. Serpine 1 induces alveolar type II cell senescence through activating p53-p21-Rb pathway in fibrotic lung disease. Aging Cell 2017, 16, 1114–1124. [Google Scholar] [CrossRef] [PubMed]
- Dasgupta, J.; Kar, S.; Liu, R.; Joseph, J.; Kalyanaraman, B.; Remington, S.J.; Chen, C.; Melendez, J.A. Reactive oxygen species control senescence-associated matrix metalloproteinase-1 through c-Jun-N-terminal kinase. J. Cell. Physiol. 2010, 225, 52–62. [Google Scholar] [CrossRef] [PubMed]
- Basisty, N.; Kale, A.; Jeon, O.H.; Kuehnemann, C.; Payne, T.; Rao, C.; Holtz, A.; Shah, S.; Sharma, V.; Ferrucci, L.; et al. A proteomic atlas of senescence-associated secretomes for aging biomarker development. PLoS Biol. 2020, 18, e3000599. [Google Scholar] [CrossRef]
- Acosta, J.C.; Banito, A.; Wuestefeld, T.; Georgilis, A.; Janich, P.; Morton, J.P.; Athineos, D.; Kang, T.-W.; Lasitschka, F.; Andrulis, M.; et al. A complex secretory program orchestrated by the inflammasome controls paracrine senescence. Nat. Cell Biol. 2013, 15, 978–990. [Google Scholar] [CrossRef]
- Elzi, D.J.; Lai, Y.; Song, M.; Hakala, K.; Weintraub, S.T.; Shiio, Y. Plasminogen activator inhibitor 1—Insulin-like growth factor binding protein 3 cascade regulates stress-induced senescence. Proc. Natl. Acad. Sci. USA 2012, 109, 12052–12057. [Google Scholar] [CrossRef]
- Wedel, S.; Martic, I.; Navarro, L.G.; Ploner, C.; Pierer, G.; Jansen-Dürr, P.; Cavinato, M. Depletion of growth differentiation factor 15 (GDF15) leads to mitochondrial dysfunction and premature senescence in human dermal fibroblasts. Aging Cell 2022, 22, e13752. [Google Scholar] [CrossRef] [PubMed]
- Heath, C.G.; Viphakone, N.; Wilson, S.A. The role of TREX in gene expression and disease. Biochem. J. 2016, 473, 2911–2935. [Google Scholar] [CrossRef]
- Darweesh, M.; Younis, S.; Hajikhezri, Z.; Ali, A.; Jin, C.; Punga, T.; Gupta, S.; Essand, M.; Andersson, L.; Akusjärvi, G. ZC3H11A loss of function enhances NF-κB signaling through defective IκBα protein expression. Front. Immunol. 2022, 13, 1002823. [Google Scholar] [CrossRef]
- Ferreira, F.J.; Galhardo, M.; Nogueira, J.M.; Teixeira, J.; Logarinho, E.; Bessa, J. FOXM1 Expression Reverts Aging Chromatin Profiles through Repression of the Senescence-Associated Pioneer Factor AP-1. Nat. Commun. 2025, 16, 2931. [Google Scholar] [CrossRef] [PubMed]
- Chien, Y.; Scuoppo, C.; Wang, X.; Fang, X.; Balgley, B.; Bolden, J.E.; Premsrirut, P.; Luo, W.; Chicas, A.; Lee, C.S.; et al. Control of the Senescence-Associated Secretory Phenotype by NF-ΚB Promotes Senescence and Enhances Chemosensitivity. Genes Dev. 2011, 25, 2125–2136. [Google Scholar] [CrossRef]
- Yıldırım, B.; Kulak, K.; Bilir, A. Midkine: A Cancer Biomarker Candidate and Innovative Therapeutic Approaches. Eur. J. Breast Health 2024, 20, 167–177. [Google Scholar] [CrossRef]
- Sorrelle, N.; A Dominguez, A.T.; A Brekken, R. From top to bottom: midkine and pleiotrophin as emerging players in immune regulation. J. Leukoc. Biol. 2017, 102, 277–286. [Google Scholar] [CrossRef]
- Neufeld, G.; Mumblat, Y.; Smolkin, T.; Toledano, S.; Nir-Zvi, I.; Ziv, K.; Kessler, O. The role of the semaphorins in cancer. Cell Adhes. Migr. 2016, 10, 652–674. [Google Scholar] [CrossRef]
- Salazar, V.S.; Gamer, L.W.; Rosen, V. BMP signalling in skeletal development, disease and repair. Nat. Rev. Endocrinol. 2016, 12, 203–221. [Google Scholar] [CrossRef]
- Han, J.M.; Jung, H.J. Cyclophilin A/CD147 Interaction: A Promising Target for Anticancer Therapy. Int. J. Mol. Sci. 2022, 23, 9341. [Google Scholar] [CrossRef] [PubMed]
- Azamar-Llamas, D.; Hernández-Molina, G.; Ramos-Ávalos, B.; Furuzawa-Carballeda, J. Adipokine Contribution to the Pathogenesis of Osteoarthritis. Mediat. Inflamm. 2017, 2017, 5468023. [Google Scholar] [CrossRef]
- Ahmad, S.; Gonzales, L.J.d.C.; Bowler-Barnett, E.H.; Rice, D.L.; Kim, M.; Wijerathne, S.; Luciani, A.; Kandasaamy, S.; Luo, J.; Watkins, X.; et al. The UniProt website API: facilitating programmatic access to protein knowledge. Nucleic Acids Res. 2025, 53, W547–W553. [Google Scholar] [CrossRef]
- Bellan, M.; Pirisi, M.; Sainaghi, P.P. The Gas6/TAM System and Multiple Sclerosis. Int. J. Mol. Sci. 2016, 17, 1807. [Google Scholar] [CrossRef]
- Philtjens, S.; Van Mossevelde, S.; van der Zee, J.; Wauters, E.; Dillen, L.; Vandenbulcke, M.; Vandenberghe, R.; Ivanoiu, A.; Sieben, A.; Willems, C.; et al. Rare nonsynonymous variants in SORT1 are associated with increased risk for frontotemporal dementia. Neurobiol. Aging 2018, 66, 181.e3–181.e10. [Google Scholar] [CrossRef]
- Kucukefe, Y.; Kaypmaz, A. Delayed Feedback Control as Applied to Active Suspension of a Ground Vehicle. In Proceedings of the IEEE EUROCON 2009, St. Petersburg, Russia, 18–23 May 2009; pp. 916–921. [Google Scholar]
- Ieguchi, K.; Fujita, M.; Ma, Z.; Davari, P.; Taniguchi, Y.; Sekiguchi, K.; Wang, B.; Takada, Y.K.; Takada, Y. Direct Binding of the EGF-like Domain of Neuregulin-1 to Integrins (αvβ3 and α6β4) Is Involved in Neuregulin-1/ErbB Signaling. J. Biol. Chem. 2010, 285, 31388–31398. [Google Scholar] [CrossRef]
- Uhl, G.R.; Martinez, M.J. PTPRD: neurobiology, genetics, and initial pharmacology of a pleiotropic contributor to brain phenotypes. Ann. N. Y. Acad. Sci. 2019, 1451, 112–129. [Google Scholar] [CrossRef]
- Won, S.Y.; Lee, P.; Kim, H.M. Synaptic organizer: Slitrks and type IIa receptor protein tyrosine phosphatases. Curr. Opin. Struct. Biol. 2019, 54, 95–103. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Yang, X.; Li, C. Integrated Analysis of EREG Expression, a Gene Associated with Cervical Cancer Prognosis. Am. J. Cancer Res. 2023, 13, 4644–4660. [Google Scholar] [PubMed]
- Cheng, W.L.; Feng, P.H.; Lee, K.Y.; Chen, K.Y.; Sun, W.L.; Van Hiep, N.; Luo, C.S.; Wu, S.M. The Role of Ereg/Egfr Pathway in Tumor Progression. Int. J. Mol. Sci. 2021, 22, 12828. [Google Scholar] [CrossRef]
- Liu, R.; Shuai, Y.; Luo, J.; Zhang, Z. SEMA3C Promotes Cervical Cancer Growth and Is Associated With Poor Prognosis. Front. Oncol. 2019, 9, 1035. [Google Scholar] [CrossRef]
- Fukuda, T.; Suzuki, E.; Fukuda, R. Bone Morphogenetic Protein Signaling Is a Possible Therapeutic Target in Gynecologic Cancer. Cancer Sci. 2023, 114, 722–729. [Google Scholar] [CrossRef] [PubMed]
- Wei, H.; Wang, N.; Zhang, Y.; Wang, S.; Pang, X.; Zhang, J.; Luo, Q.; Su, Y.; Zhang, S. Clinical Significance of Wnt-11 and Squamous Cell Carcinoma Antigen Expression in Cervical Cancer. Med. Oncol. 2014, 31, 933. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Wang, M.; Li, X.; Xie, Y.; Xia, X.; Tian, J.; Zhang, K.; Tang, A. Wnt Signaling in Cervical Cancer? J. Cancer 2018, 9, 1277–1286. [Google Scholar] [CrossRef] [PubMed]
- Ding, X.-W.; Wu, J.-H.; Jiang, C.-P. ABCG2: A potential marker of stem cells and novel target in stem cell and cancer therapy. Life Sci. 2010, 86, 631–637. [Google Scholar] [CrossRef]
- Organista-Nava, J.; Garibay-Cerdenares, O.L.; Illades-Aguiar, B.; Gómez-Gómez, Y.; Leyva-Vázquez, M.A. Cervical cancer stem cell-associated genes: Prognostic implications in cervical cancer (Review). Oncol. Lett. 2019, 18, 7–14. [Google Scholar] [CrossRef]
- Di Fiore, R.; Suleiman, S.; Drago-Ferrante, R.; Subbannayya, Y.; Pentimalli, F.; Giordano, A.; Calleja-Agius, J. Cancer Stem Cells and Their Possible Implications in Cervical Cancer: A Short Review. Int. J. Mol. Sci. 2022, 23, 5167. [Google Scholar] [CrossRef]
- Kevat, S.; Mistry, A.; Oza, N.; Majmudar, M.; Patel, N.; Shah, R.; Ramachandran, A.V.; Chauhan, R.; Haque, S.; Parashar, N.C.; et al. Cancer Stem Cell Regulation as a Target of Therapeutic Intervention: Insights into Breast, Cervical and Lung Cancer. Cell Biochem. Biophys. 2025, 83, 1521–1535. [Google Scholar] [CrossRef]
- Nguyen, A.L.; Facey, C.O.B.; Boman, B.M. The Significance of Aldehyde Dehydrogenase 1 in Cancers. Int. J. Mol. Sci. 2024, 26, 251. [Google Scholar] [CrossRef]
- Tsang, J.Y.S.; Huang, Y.-H.; Luo, M.-H.; Ni, Y.-B.; Chan, S.-K.; Lui, P.C.W.; Yu, A.M.C.; Tan, P.H.; Tse, G.M. Cancer stem cell markers are associated with adverse biomarker profiles and molecular subtypes of breast cancer. Breast Cancer Res. Treat. 2012, 136, 407–417. [Google Scholar] [CrossRef]
- Jaggupilli, A.; Elkord, E. Significance of CD44 and CD24 as Cancer Stem Cell Markers: An Enduring Ambiguity. Clin. Dev. Immunol. 2012, 2012, 708036. [Google Scholar] [CrossRef]
- de Sousa, C.; Eksteen, C.; Riedemann, J.; Engelbrecht, A.-M. Highlighting the role of CD44 in cervical cancer progression: immunotherapy’s potential in inhibiting metastasis and chemoresistance. Immunol. Res. 2024, 72, 592–604. [Google Scholar] [CrossRef]
- Liu, H.; Wang, Y.-J.; Bian, L.; Fang, Z.-H.; Zhang, Q.-Y.; Cheng, J.-X. CD44+/CD24+ Cervical Cancer Cells Resist Radiotherapy and Exhibit Properties of Cancer Stem Cells. Eur. Rev. Med. Pharmacol. Sci. 2016, 20, 1745–1754. [Google Scholar] [PubMed]
- Escobar-Hoyos, L.F.; Yang, J.; Zhu, J.; Cavallo, J.-A.; Zhai, H.; Burke, S.; Koller, A.; I Chen, E.; Shroyer, K.R. Keratin 17 in premalignant and malignant squamous lesions of the cervix: proteomic discovery and immunohistochemical validation as a diagnostic and prognostic biomarker. Mod. Pathol. 2014, 27, 621–630. [Google Scholar] [CrossRef]
- Zhang, Q.; Han, Z.; Zhu, Y.; Chen, J.; Li, W. The Role and Specific Mechanism of OCT4 in Cancer Stem Cells: A Review. Int. J. Stem Cells 2020, 13, 312–325. [Google Scholar] [CrossRef]
- Saha, S.K.; Islam, S.M.R.; Kwak, K.-S.; Rahman, M.S.; Cho, S.-G. PROM1 and PROM2 Expression Differentially Modulates Clinical Prognosis of Cancer: A Multiomics Analysis. Cancer Gene Ther. 2020, 27, 147–167. [Google Scholar] [CrossRef]
- Zhu, H.; Luo, H.; Shen, Z.; Hu, X.; Sun, L.; Zhu, X. Transforming Growth Factor-Β1 in Carcinogenesis, Progression, and Therapy in Cervical Cancer. Tumour Biol. 2016, 37, 7075–7083. [Google Scholar] [CrossRef]
- Otani, S.; Fujii, T.; Kukimoto, I.; Yamamoto, N.; Tsukamoto, T.; Ichikawa, R.; Nishio, E.; Iwata, A. Cytokine Expression Profiles in Cervical Mucus from Patients with Cervical Cancer and Its Precursor Lesions. Cytokine 2019, 120, 210–219. [Google Scholar] [CrossRef] [PubMed]
- Gallo, M.; Frezzetti, D.; Roma, C.; Chicchinelli, N.; Barbieri, A.; Arra, C.; Scognamiglio, G.; Botti, G.; De Luca, A.; Normanno, N. RANTES and IL-6 Cooperate in Inducing a More Aggressive Phenotype in Breast Cancer Cells. Oncotarget 2018, 9, 17543–17553. [Google Scholar] [CrossRef] [PubMed]








| Cell Culture | Patient Therapy | Number of Passages of Cultured Cells | Number of Cells in SC Analysis * | Number of Processed Reads * |
|---|---|---|---|---|
| AdMer35 | Three cycles of polychemotherapy according to the scheme paclitaxel combined with a carboplatin. | 12 | 7256 | 183,166,343 |
| AdMer43 | Chemoradiation with a total focal dose of 50 Gy supplemented with cisplatin administration. Session of intracavitary brachytherapy with an SFD of 5 Gy, external boost to the cervical tumor with 20 Gy, and retroperitoneal lymph node conglomerates with 20 Gy. Gemcitabine therapy. | 12 | 7198 | 295,237,133 |
| Cell Types | PLC | Non-PLC | PLC (%) | Non-PLC (%) |
|---|---|---|---|---|
| SnC | 107 | 1401 | 26.03 | 9.55 |
| Non-SnC | 304 | 13,273 | 73.97 | 90.45 |
| Total | 411 | 14,674 | 100 | 100 |
| Cell Type | Cell Cycle Phase | |||||
|---|---|---|---|---|---|---|
| G1 | S | G2M | G1 (%) | S (%) | G2M (%) | |
| SnC | 703 | 483 | 322 | 47 | 32 | 21 |
| Non-SnC | 4709 | 4333 | 4535 | 35 | 32 | 32 |
| Cell Types | AdMer43 | AdMer35 | AdMer43 (%) | AdMer35 (%) |
|---|---|---|---|---|
| SLC | 508 | 1001 | 6.61 | 13.53 |
| Non-SLC | 7177 | 6399 | 93.39 | 86.47 |
| Total | 7685 | 7400 | 100 | 100 |
| Characteristic | Seurat Clusters | |||||
|---|---|---|---|---|---|---|
| 0 | 1 | 2 | 3 | 4 | 5 | |
| Cell Number | 4422 | 4102 | 2682 | 2677 | 788 | 414 |
| Cell Lines | ||||||
| AdMer35, % | 99.95 | 0.07 | 0.04 | 99.89 | 19.42 | 35.99 |
| AdMer43, % | 0.05 | 99.93 | 99.96 | 0.11 | 80.58 | 64.01 |
| Cell Line Attribution | AdMer35 | AdMer43 | AdMer43 | AdMer35 | Mixed 35/43 | Mixed 35/43 |
| HPV59 Viral Transcripts, % | ||||||
| Median HPV59 Transcript Contribution | 0.061 | 0.074 | 0.075 | 0.071 | 0.109 | 0.001 |
| HPV59 Class | Medium | Medium | Medium | Medium | High | Low |
| Cell Cycle, % | ||||||
| G1 | 47.85 | 62.94 | 5.48 | 2.99 | 39.85 | 41.79 |
| S | 51.18 | 35.79 | 17.04 | 10.09 | 33.88 | 21.98 |
| G2M | 0.97 | 1.27 | 77.48 | 86.93 | 26.27 | 36.23 |
| Cell Cycle Phase | G1/S | G1/S | G2M | G2M | G1/S > G2M | G1/S > G2M |
| Progenitor-Like Cells (PLCs), % | ||||||
| PLC | 0.81 | 1.39 | 1.27 | 0.45 | 1.52 | 62.80 |
| Enriched with PLCs (>10%) | No | No | No | No | No | Yes |
| Cancer Stem-Like Cells (SLCs), % | ||||||
| SLC | 13.89 | 7.09 | 4.40 | 13.56 | 15.23 | 0.72 |
| Enriched with SLCs (>10%) | Yes | No | No | Yes | Yes | No |
| Senescence-Like Cells (SnCs), % | ||||||
| SnCs | 10.33 | 11.38 | 5.03 | 7.88 | 16.12 | 26.81 |
| SnC Contribution | Medium | Medium | Low | Low | High | High |
| Short Description * | ||||||
| Main Cell Line Main Phase of Cell Cycle | AdMer35 G1/S | AdMer43 G1/S | AdMer43 G2M | AdMer35 G2M | Mixed 35/43 Mixed | Mixed 35/43 Mixed |
| Enrichment | SLC | No | No | No | SLC, HPV59, SnC | PLC, SnC (Lowest SLC, HPV59) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Semenov, D.V.; Tatarnikova, I.S.; Chesnokova, A.S.; Talyshev, V.A.; Zenkova, M.A.; Logashenko, E.B. Single-Cell Transcriptomic Landscape of Cervical Cancer Cell Lines Before and After Chemoradiotherapy. Cells 2026, 15, 115. https://doi.org/10.3390/cells15020115
Semenov DV, Tatarnikova IS, Chesnokova AS, Talyshev VA, Zenkova MA, Logashenko EB. Single-Cell Transcriptomic Landscape of Cervical Cancer Cell Lines Before and After Chemoradiotherapy. Cells. 2026; 15(2):115. https://doi.org/10.3390/cells15020115
Chicago/Turabian StyleSemenov, Dmitriy V., Irina S. Tatarnikova, Anna S. Chesnokova, Vadim A. Talyshev, Marina A. Zenkova, and Evgeniya B. Logashenko. 2026. "Single-Cell Transcriptomic Landscape of Cervical Cancer Cell Lines Before and After Chemoradiotherapy" Cells 15, no. 2: 115. https://doi.org/10.3390/cells15020115
APA StyleSemenov, D. V., Tatarnikova, I. S., Chesnokova, A. S., Talyshev, V. A., Zenkova, M. A., & Logashenko, E. B. (2026). Single-Cell Transcriptomic Landscape of Cervical Cancer Cell Lines Before and After Chemoradiotherapy. Cells, 15(2), 115. https://doi.org/10.3390/cells15020115