Characterizing Microglial Morphology: Methodological Advances in Confocal Imaging and Analysis
Abstract
1. Introduction
2. Materials and Methods
2.1. Animals
2.2. Immunostaining
2.3. Data Analysis
3. Results
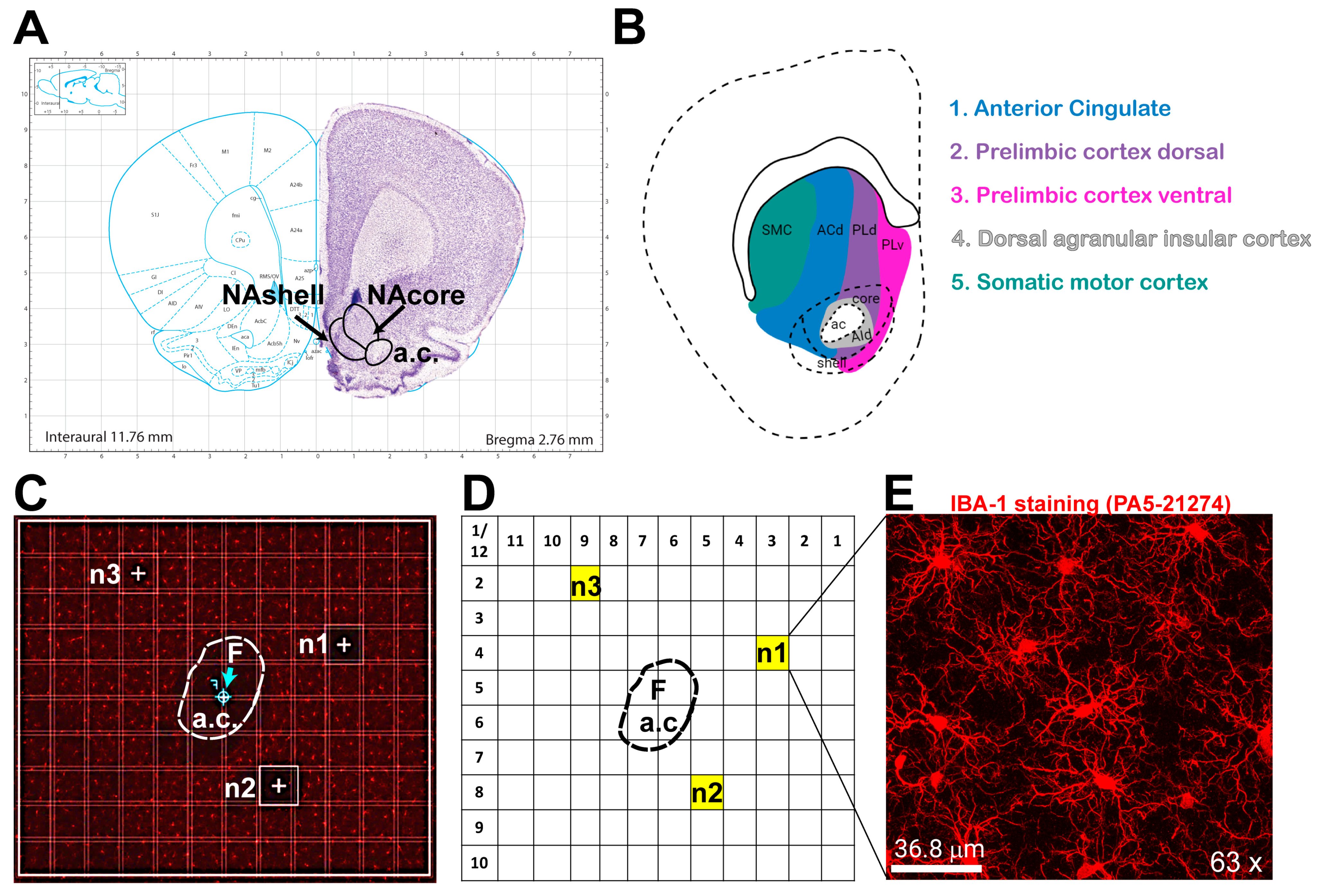
3.1. Unbiased Image Acquisition Methodology
- A.
- Visualization of microglia
- B.
- Optimizing microglia visualization: slices thickness
- C.
- Reference coordinates for imaging the NAcore
- D.
- Image acquisition workflow
3.2. Morphological Analysis Tool Comparison
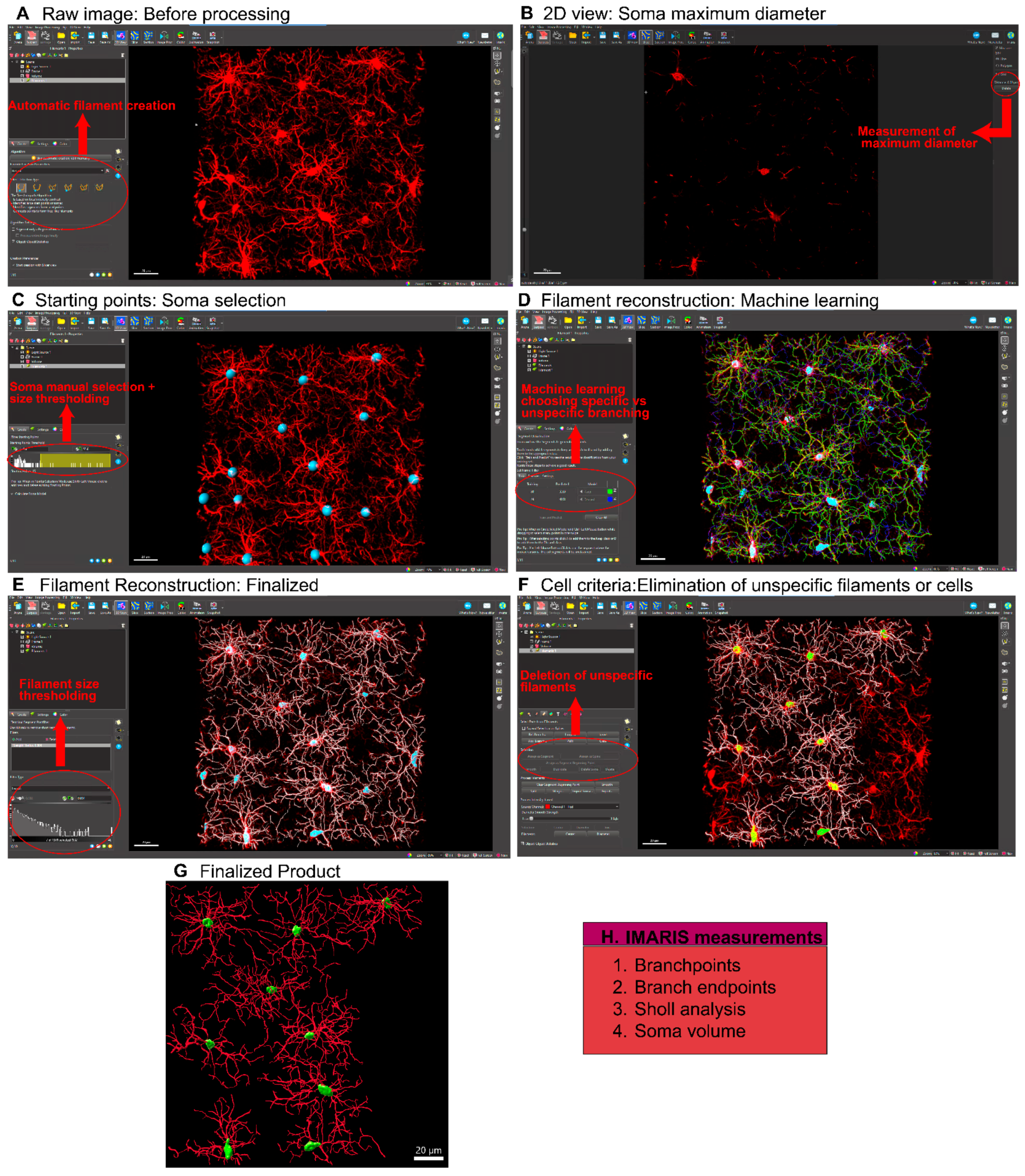
- A.
- Microglial morphological analysis using IMARIS
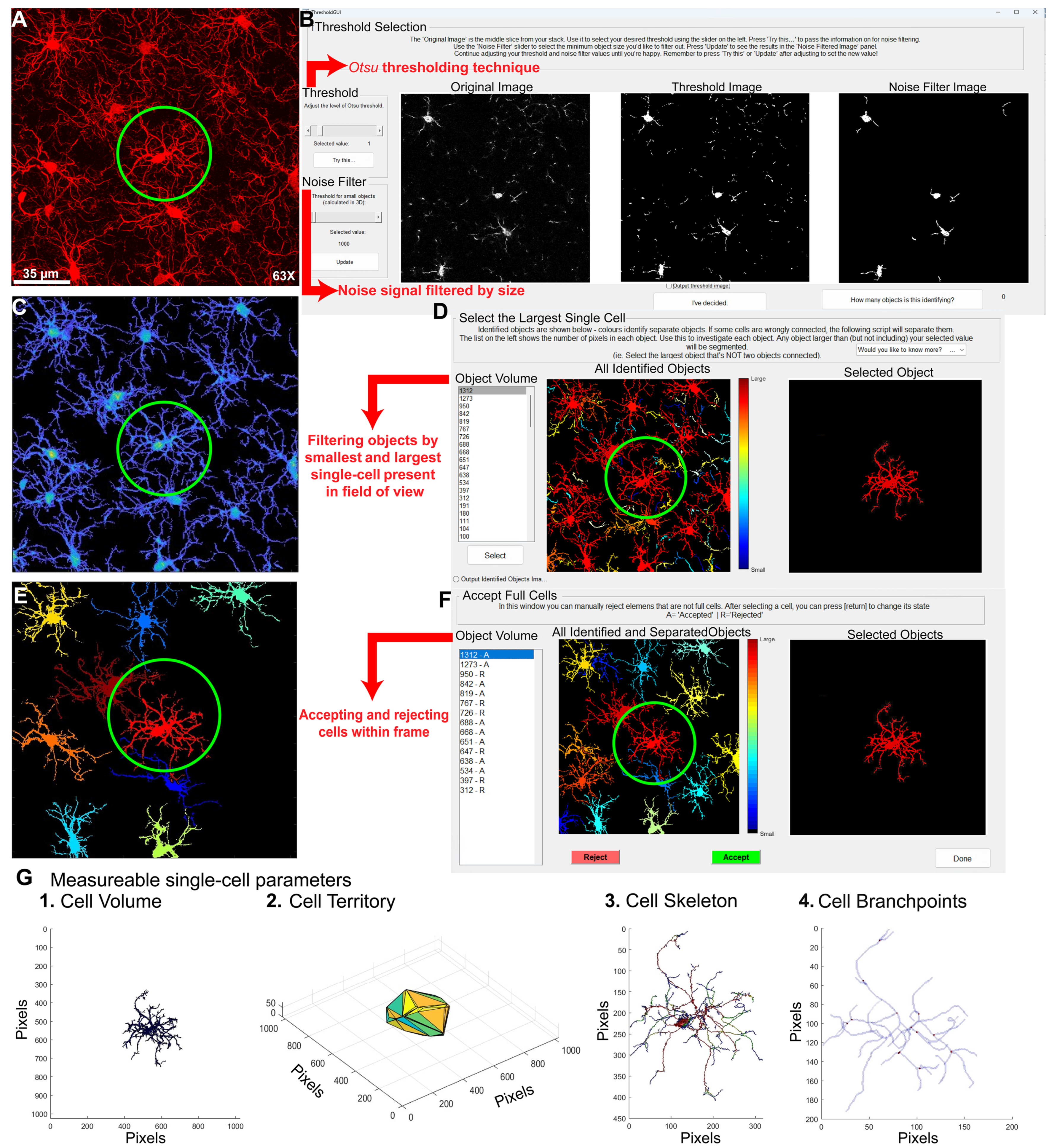
- B.
- Microglial morphological analysis using CellSelect-3DMorph
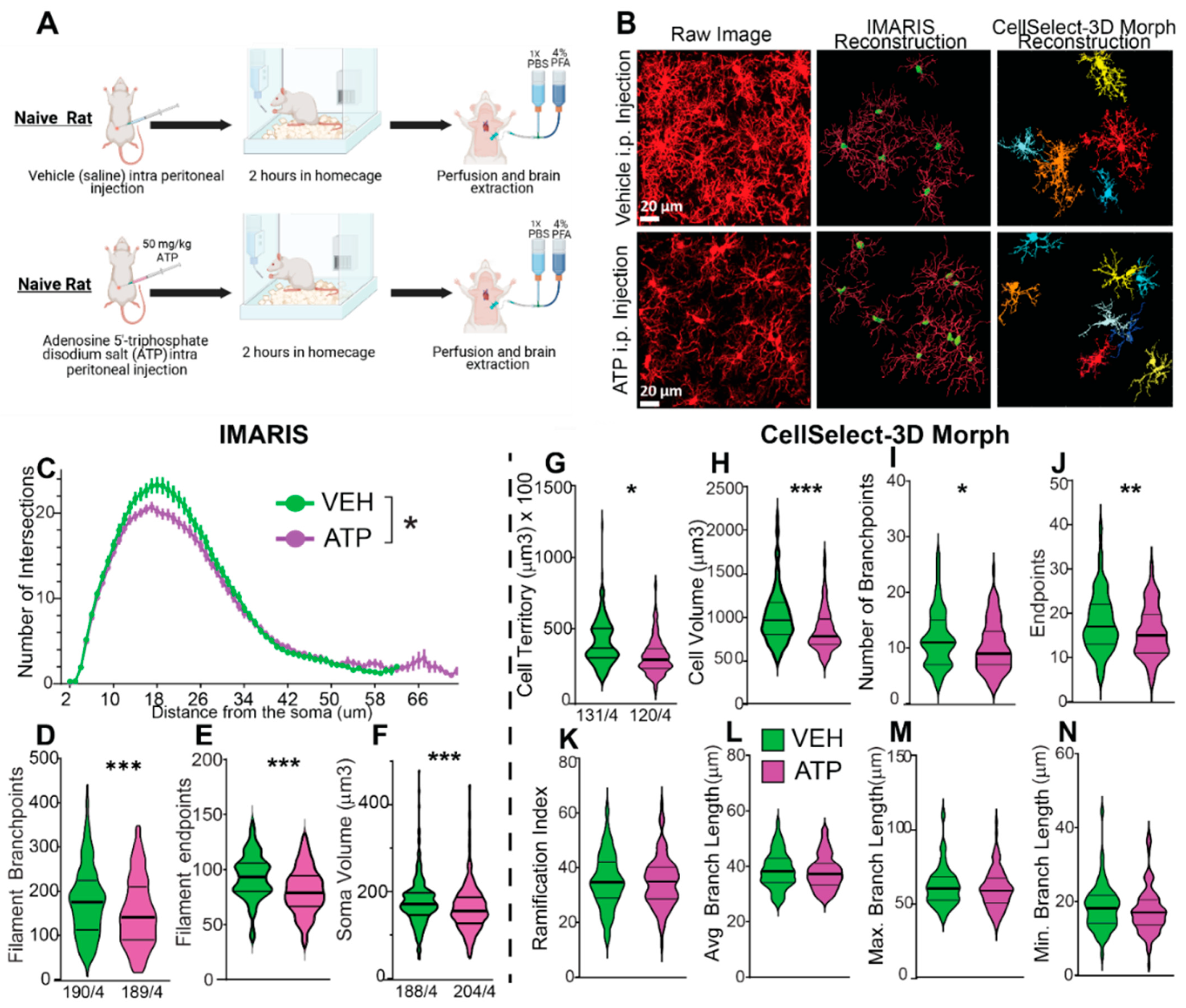
3.3. Comparison Between IMARIS and CellSelect-3DMorph Outputs
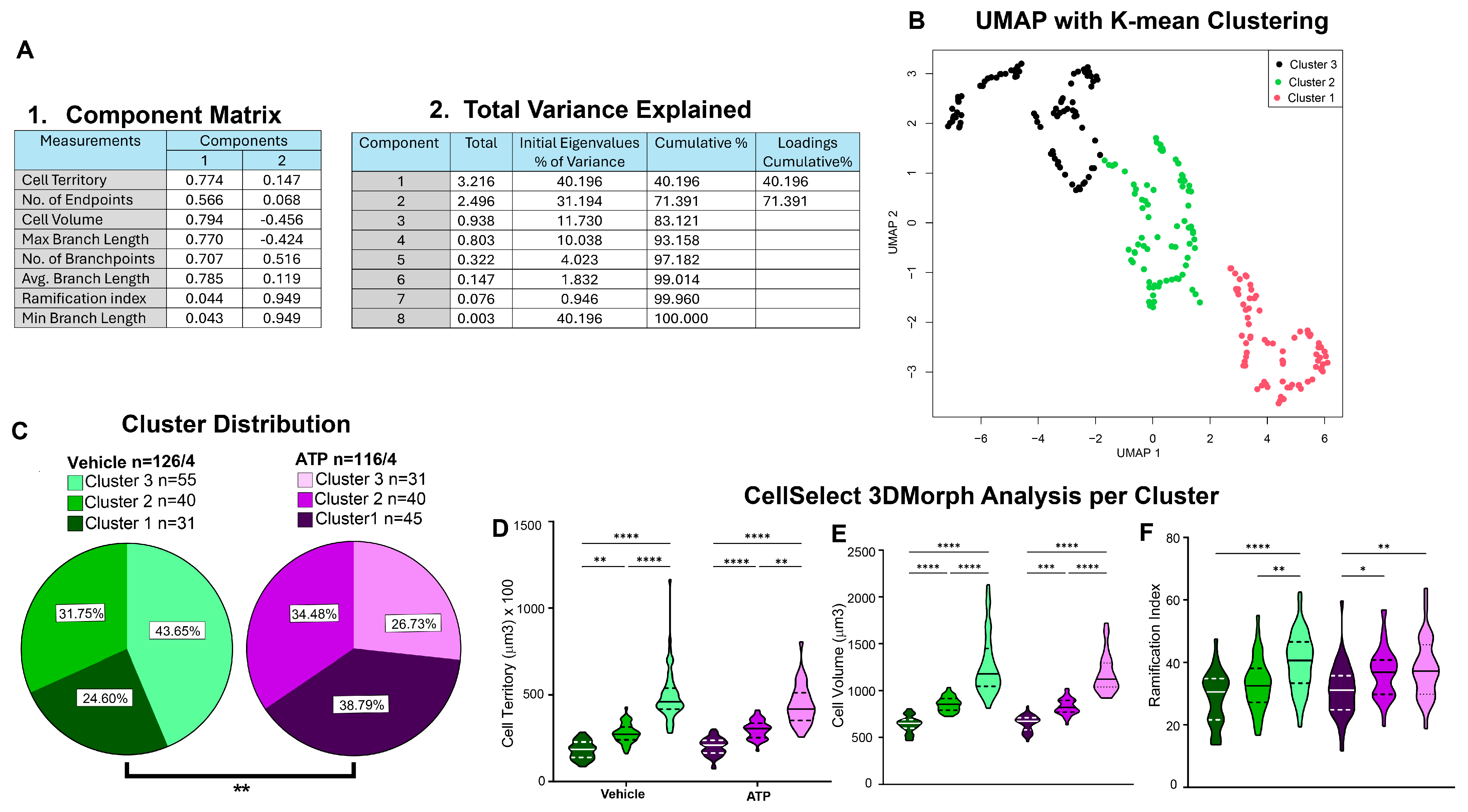
3.4. Cluster Analysis Pipeline
- A.
- Cluster analysis workflow
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Nowak, D.B.; Taborda-Bejarano, J.P.; Chaure, F.J.; Mantsch, J.R.; Garcia-Keller, C. Understanding Microglia in Mesocorticolimbic Circuits: Implications for the Study of Chronic Stress and Substance Use Disorders. Cells 2025, 14, 1014. [Google Scholar] [CrossRef]
- Paolicelli, R.C.; Sierra, A.; Stevens, B.; Tremblay, M.-E.; Aguzzi, A.; Ajami, B.; Amit, I.; Audinat, E.; Bechmann, I.; Bennett, M.; et al. Microglia states and nomenclature: A field at its crossroads. Neuron 2022, 110, 3458–3483. [Google Scholar] [CrossRef] [PubMed]
- Ball, J.B.; Mcnulty, C.J.; Green-Fulgham, S.M.; Dragavon, J.M.; Correia Rocha, I.R.; Finch, M.R.; Prévost, E.D.; Siddique, I.I.; Woodall, B.J.; Watkins, L.R.; et al. Combining RNAscope and immunohistochemistry to visualize inflammatory gene products in neurons and microglia. Front. Mol. Neurosci. 2023, 16, 1225847. [Google Scholar] [CrossRef]
- Schmid, C.D.; Melchior, B.; Masek, K.; Puntambekar, S.S.; Danielson, P.E.; Lo, D.D.; Gregor Sutcliffe, J.; Carson, M.J. Differential gene expression in LPS/IFNγ activated microglia and macrophages: In vitro versus in vivo. J. Neurochem. 2009, 109, 117–125. [Google Scholar] [CrossRef]
- Liu, Q.-R.; Canseco-Alba, A.; Liang, Y.; Ishiguro, H.; Onaivi, E.S. Low Basal CB2R in Dopamine Neurons and Microglia Influences Cannabinoid Tetrad Effects. Int. J. Mol. Sci. 2020, 21, 9763. [Google Scholar] [CrossRef] [PubMed]
- Olah, M.; Menon, V.; Habib, N.; Taga, M.F.; Ma, Y.; Yung, C.J.; Cimpean, M.; Khairallah, A.; Coronas-Samano, G.; Sankowski, R.; et al. Single cell RNA sequencing of human microglia uncovers a subset associated with Alzheimer’s disease. Nat. Commun. 2020, 11, 6129. [Google Scholar] [CrossRef] [PubMed]
- Kemp, G.M.; Altimimi, H.F.; Nho, Y.; Heir, R.; Klyczek, A.; Stellwagen, D. Sustained TNF signaling is required for the synaptic and anxiety-like behavioral response to acute stress. Mol. Psychiatry. 2022, 27, 4474–4484. [Google Scholar] [CrossRef] [PubMed]
- Guo, S.; Wang, H.; Yin, Y. Microglia Polarization From M1 to M2 in Neurodegenerative Diseases. Front. Aging Neurosci. 2022, 14, 815347. [Google Scholar] [CrossRef]
- Streit, W.J.; Graeber, M.B.; George, W.K. Functional plasticity of microglia: A review. Glia 1988, 1, 301–307. [Google Scholar] [CrossRef]
- Chaure, F.; CGK Laboratory; Taborda Bejarano, J.B. CellSelect-3Dmorph—GitHub Repository. 2023. Available online: https://github.com/CGK-Laboratory/CellSelect-3DMorph (accessed on 11 November 2024).
- Xiang, Y.; Wang, X.; Yan, C.; Gao, Q.; Li, S.A.; Liu, J.; Zhou, K.; Guo, X.; Lee, W.; Zhang, Y. Adenosine-5′-triphosphate (ATP) protects mice against bacterial infection by activation of the NLRP3 inflammasome. PLoS ONE 2013, 8, e63759. [Google Scholar] [CrossRef]
- Scofield, M.D.; Heinsbroek, J.A.; Gipson, C.D.; Kupchik, Y.M.; Spencer, S.; Smith, A.C.W.; Roberts-Wolfe, D.; Kalivas, P.W. The Nucleus Accumbens: Mechanisms of Addiction across Drug Classes Reflect the Importance of Glutamate Homeostasis. Pharmacol. Rev. 2016, 68, 816–871. [Google Scholar] [CrossRef]
- Turner, B.D.; Kashima, D.T.; Manz, K.M.; Grueter, C.A.; Grueter, B.A. Synaptic Plasticity in the Nucleus Accumbens: Lessons Learned from Experience. ACS Chem. Neurosci. 2018, 9, 2114–2126. [Google Scholar] [CrossRef]
- Everitt, B.J.; Robbins, T.W. Neural systems of reinforcement for drug addiction: From actions to habits to compulsion. Nat. Neurosci. 2005, 8, 1481–1489. [Google Scholar] [CrossRef] [PubMed]
- Mulholland, P.J.; Chandler, L.J.; Kalivas, P.W. Signals from the Fourth Dimension Regulate Drug Relapse. Trends Neurosci. 2016, 39, 472–485. [Google Scholar] [CrossRef]
- Garcia-Keller, C.; Carter, J.S.; Kruyer, A.; Kearns, A.M.; Hopkins, J.L.; Hodebourg, R.; Kalivas, P.W.; Reichel, C.M. Behavioral and accumbens synaptic plasticity induced by cues associated with restraint stress. Neuropsychopharmacology 2021, 46, 1848–1856. [Google Scholar] [CrossRef]
- Avalos, M.P.; Guzman, A.S.; Rigoni, D.; Gorostiza, E.A.; Sanchez, M.A.; Mongi-Bragato, B.; Garcia-Keller, C.; Perassi, E.M.; Virgolini, M.B.; Peralta Ramos, J.M.; et al. Minocycline prevents chronic restraint stress-induced vulnerability to developing cocaine self-administration and associated glutamatergic mechanisms: A potential role of microglia. Brain Behav. Immun. 2022, 101, 359–376. [Google Scholar] [CrossRef] [PubMed]
- Rigoni, D.; Avalos, M.P.; Boezio, M.J.; Guzmán, A.S.; Calfa, G.D.; Perassi, E.M.; Pierotti, S.M.; Bisbal, M.; Garcia-Keller, C.; Cancela, L.M.; et al. Stress-induced vulnerability to develop cocaine addiction depends on cofilin modulation. Neurobiol. Stress. 2021, 15, 100349. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Keller, C.; Kupchik, Y.M.; Gipson, C.D.; Brown, R.M.; Spencer, S.; Bollati, F.; Esparza, M.A.; Roberts-Wolfe, D.J.; Heinsbroek, J.A.; Bobadilla, A.-C.; et al. Glutamatergic mechanisms of comorbidity between acute stress and cocaine self-administration. Mol. Psychiatry 2016, 21, 1063–1069. [Google Scholar] [CrossRef]
- Garcia-Keller, C.; Scofield, M.D.; Neuhofer, D.; Varanasi, S.; Reeves, M.T.; Hughes, B.; Anderson, E.; Richie, C.T.; Mejias-Aponte, C.; Pickel, J.; et al. Relapse-Associated Transient Synaptic Potentiation Requires Integrin-Mediated Activation of Focal Adhesion Kinase and Cofilin in D1-Expressing Neurons. J. Neurosci. 2020, 40, 8463–8477. [Google Scholar] [CrossRef]
- Fox, M.E.; Wulff, A.B.; Franco, D.; Choi, E.Y.; Calarco, C.A.; Engeln, M.; Turner, M.D.; Chandra, R.; Rhodes, V.M.; Thompson, S.M.; et al. Adaptations in nucleus accumbens neuron subtypes mediate negative affective behaviors in fentanyl abstinence. Biol. Psychiatry 2023, 93, 489–501. [Google Scholar] [CrossRef]
- Jurga, A.M.; Paleczna, M.; Kuter, K.Z. Overview of General and Discriminating Markers of Differential Microglia Phenotypes. Front. Cell. Neurosci. 2020, 14, 198. [Google Scholar] [CrossRef]
- Jung, S.; Aliberti, J.; Graemmel, P.; Sunshine, M.J.; Kreutzberg, G.W.; Sher, A.; Littman, D.R. Analysis of Fractalkine Receptor CX3CR1 Function by Targeted Deletion and Green Fluorescent Protein Reporter Gene Insertion. Mol. Cell. Biol. 2000, 20, 4106–4114. [Google Scholar] [CrossRef] [PubMed]
- Kaiser, T.; Feng, G. Tmem119-EGFP and Tmem119-CreERT2 Transgenic Mice for Labeling and Manipulating Microglia. eNeuro 2019, 6, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Buttgereit, A.; Lelios, I.; Yu, X.; Vrohlings, M.; Krakoski, N.R.; Gautier, E.L.; Nishinakamura, R.; Becher, B.; Greter, M. Sall1 is a transcriptional regulator defining microglia identity and function. Nat. Immunol. 2016, 17, 1397–1406. [Google Scholar] [CrossRef]
- Vanryzin, J.W.; Arambula, S.E.; Ashton, S.E.; Blanchard, A.C.; Burzinski, M.D.; Davis, K.T.; Edwards, S.; Graham, E.L.; Holley, A.; Kight, K.E.; et al. Generation of an Iba1-EGFP Transgenic Rat for the Study of Microglia in an Outbred Rodent Strain. eNeuro 2021, 8, 1–16. [Google Scholar] [CrossRef]
- Sharma, K.; Bisht, K.; Eyo, U.B. A Comparative Biology of Microglia Across Species. Front. Cell Dev. Biol. 2021, 9, 652748. [Google Scholar] [CrossRef]
- Franco-Bocanegra, D.K.; Gourari, Y.; Mcauley, C.; Chatelet, D.S.; Johnston, D.A.; Nicoll, J.A.R.; Boche, D. Microglial morphology in Alzheimer’s disease and after Aβ immunotherapy. Sci. Rep. 2021, 11, 1–12. [Google Scholar] [CrossRef]
- Kim, J.; Pavlidis, P.; Ciernia, A.V. Development of a High-Throughput Pipeline to Characterize Microglia Morphological States at a Single-Cell Resolution. eNeuro 2024, 11, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Silburt, J.; Aubert, I. MORPHIOUS: An unsupervised machine learning workflow to detect the activation of microglia and astrocytes. J. Neuroinflamm. 2022, 19, 24. [Google Scholar] [CrossRef]
- Green, T.R.F.; Murphy, S.M.; Rowe, R.K. Comparisons of quantitative approaches for assessing microglial morphology reveal inconsistencies, ecological fallacy, and a need for standardization. Sci. Rep. 2022, 12, 18196. [Google Scholar] [CrossRef]
- Paxinos, G.; Watson, C. Paxinos and Watson’s the Rat Brain in Stereotaxic Coordinates, 7th ed.; Elsevier: Amsterdam, The Netherlands, 2014. [Google Scholar]
- Voorn, P.; Vanderschuren, L.J.M.J.; Groenewegen, H.J.; Robbins, T.W.; Pennartz, C.M.A. Putting a spin on the dorsal–ventral divide of the striatum. Trends Neurosci. 2004, 27, 468–474. [Google Scholar] [CrossRef]
- York, E.M.; Ledue, J.M.; Bernier, L.-P.; Macvicar, B.A. 3DMorph automatic analysis of microglial morphology in three dimensions from ex vivo and in vivo imaging. eNeuro 2018, 5, 1–12. [Google Scholar] [CrossRef]
- Tanaka, K.; Furuyashiki, T.; Kitaoka, S.; Senzai, Y.; Imoto, Y.; Segi-Nishida, E.; Deguchi, Y.; Breyer, R.M.; Breyer, M.D.; Narumiya, S. Prostaglandin E2-mediated attenuation of mesocortical dopaminergic pathway is critical for susceptibility to repeated social defeat stress in mice. J. Neurosci. 2012, 32, 4319–4329. [Google Scholar] [CrossRef]
- Bollinger, J.L.; Dadosky, D.T.; Flurer, J.K.; Rainer, I.L.; Woodburn, S.C.; Wohleb, E.S. Microglial P2Y12 mediates chronic stress-induced synapse loss in the prefrontal cortex and associated behavioral consequences. Neuropsychopharmacology 2023, 48, 1347–1357. [Google Scholar] [CrossRef]
- Reddaway, J.; Richardson, P.E.; Bevan, R.J.; Stoneman, J.; Palombo, M. Microglial morphometric analysis: So many options, so little consistency. Front. Neuroinform. 2023, 17, 1211188. [Google Scholar] [CrossRef] [PubMed]
- Sierra, A.; Paolicelli, R.C.; Kettenmann, H. Cien Años de Microglía: Milestones in a Century of Microglial Research. Trends Neurosci. 2019, 42, 778–792. [Google Scholar] [CrossRef] [PubMed]
- Otsu, N. A Threshold Selection Method from Gray-Level Histograms. IEEE Trans. Syst. Man. Cybern. 1979, 9, 62–66. [Google Scholar] [CrossRef]
- Sáez, P.J.; Shoji, K.F.; Retamal, M.A.; Harcha, P.A.; Ramírez, G.; Jiang, J.X.; Von Bernhardi, R.; Sáez, J.C. ATP Is Required and Advances Cytokine-Induced Gap Junction Formation in Microglia In Vitro. Mediat. Inflamm. 2013, 2013, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Stratoulias, V.; Venero, J.L.; Tremblay, M.È.; Joseph, B. Microglial subtypes: Diversity within the microglial community. EMBO J. 2019, 38, e101997. [Google Scholar] [CrossRef]
- Salamanca, L.; Mechawar, N.; Murai, K.K.; Balling, R.; Bouvier, D.S.; Skupin, A. MIC-MAC: An automated pipeline for high-throughput characterization and classification of three-dimensional microglia morphologies in mouse and human postmortem brain samples. Glia 2019, 67, 1496–1509. [Google Scholar] [CrossRef]
- Heindl, S.; Gesierich, B.; Benakis, C.; Llovera, G.; Duering, M.; Liesz, A. Automated Morphological Analysis of Microglia After Stroke. Front. Cell. Neurosci. 2018, 12, 106. [Google Scholar] [CrossRef] [PubMed]
- Colombo, G.; Cubero, R.J.A.; Kanari, L.; Venturino, A.; Schulz, R.; Scolamiero, M.; Agerberg, J.; Mathys, H.; Tsai, L.-H.; Chachólski, W.; et al. A tool for mapping microglial morphology, morphOMICs, reveals brain-region and sex-dependent phenotypes. Nat. Neurosci. 2022, 25, 1379–1393. [Google Scholar] [CrossRef] [PubMed]
- Young, K.; Morrison, H. Quantifying Microglia Morphology from Photomicrographs of Immunohistochemistry Prepared Tissue Using ImageJ. J. Vis. Exp. 2018, 136, 57648. [Google Scholar] [CrossRef]
- Clarke, D.; Crombag, H.S.; Hall, C.N. An open-source pipeline for analysing changes in microglial morphology. Open Biol. 2021, 11, 210045. [Google Scholar] [CrossRef]
| Tool/Method | Reference/Source | Pros | Cons | Notable Features/Notes |
|---|---|---|---|---|
| IMARIS | Proprietary software | - Robust 3D rendering - Versatile analysis modules - Good documentation | - Expensive - Time/labor-intensive - Requires training | Industry standard for 3D imaging; suited for detailed, high-res analysis |
| CellSelect-3DMorph | In-house | - Open-source - Faster than IMARIS - Includes ramification index | - MATLAB required - Limited customization | Optimized for 3D microglial analysis in confocal z-stacks |
| MIC-MAC | Salamanca et al., 2019 [42] | - Open-source - Combines clustering with morphology - Good for 4D data | - May need classifier training - Less flexible for custom metrics - MATLAB required | Identifies subpopulations based on morphology with clustering |
| 3DMorph | York et al., 2018 [34] | - Easy to use - Open-source - Works on 2D/3D skeletons | - Older MATLAB version required - Assumes well-isolated cells | Widely adopted; suitable for standard morphometric analysis |
| Microglia morphology quantification tool (MMQT) | Heindl et al., 2018 [43] | - High-throughput - Unsupervised and automated analysis | - Time intensive - Image preprocessing needed | Suited for dynamic changes in microglia without introducing bias |
| MORPHIOUS | Silburt & Aubert 2022 [30] | - Measures and classifies microglia on a whole brain region - Scalable and modular | - Uses strict classification and prior machine training - Not beginner-friendly | Classifies activation states based on morphological complexity |
| MorphOMICs | Colombo et al., 2022 [44] | - Broad analysis and classification of microglia - Quantifies complexity and state | - Not a software but a topological data analysis approach - Very time intensive with classifier machine learning - Multiple dependencies and software needed | Ideal for large datasets and custom pipelines |
| MicrogliaMorphology (ImageJ tool) and MicrogliaMorphologyR (R package) | Kim et al., 2024 [29] | - Standardized IHC-based pipeline - User-friendly workflow - Vast diversity in microglial measurements | - Pipeline of analysis instead include multiple software - Manual input for some steps | Good analysis pipeline for classification and identification microglial subpopulations |
| FracLac and AnalyzeSkeleton (ImageJ plugin) | Young & Morrison 2018 [45] | - Fractal dimension analysis - Skeleton-based length/branching - Easy to use | - Only provides complexity (no cell-level metrics) - 2D only | Great for comparing activation via complexity (e.g., resting vs. activated) and measuring branching, length, and endpoints |
| Inflammation-Index | Clarke et al., 2021 [46] | - High-throughput - Motion tracking and morphometry | - Best for time-lapse - Preprocessing and prior machine learning required | Suited for dynamic, in vivo imaging studies |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Taborda-Bejarano, J.P.; Nowak, D.B.; Chaure, F.; Allen, M.L.; Blek, K.A.; Walterhouse, S.; Mantsch, J.R.; Garcia-Keller, C. Characterizing Microglial Morphology: Methodological Advances in Confocal Imaging and Analysis. Cells 2025, 14, 1354. https://doi.org/10.3390/cells14171354
Taborda-Bejarano JP, Nowak DB, Chaure F, Allen ML, Blek KA, Walterhouse S, Mantsch JR, Garcia-Keller C. Characterizing Microglial Morphology: Methodological Advances in Confocal Imaging and Analysis. Cells. 2025; 14(17):1354. https://doi.org/10.3390/cells14171354
Chicago/Turabian StyleTaborda-Bejarano, Juan P., David B. Nowak, Fernando Chaure, Malika L. Allen, Kathryn A. Blek, Stephen Walterhouse, John R. Mantsch, and Constanza Garcia-Keller. 2025. "Characterizing Microglial Morphology: Methodological Advances in Confocal Imaging and Analysis" Cells 14, no. 17: 1354. https://doi.org/10.3390/cells14171354
APA StyleTaborda-Bejarano, J. P., Nowak, D. B., Chaure, F., Allen, M. L., Blek, K. A., Walterhouse, S., Mantsch, J. R., & Garcia-Keller, C. (2025). Characterizing Microglial Morphology: Methodological Advances in Confocal Imaging and Analysis. Cells, 14(17), 1354. https://doi.org/10.3390/cells14171354







