Epigenetic Regulation Through Histone Deacetylation: Implications and Therapeutic Potential in Hepatocellular Carcinoma
Abstract
1. Introduction
1.1. Epigenetic Landscape
1.2. Post-Translational Histone Modifications
2. Dynamics of Histone Acetylation/Deacetylation
3. HDACs Classification
3.1. Class I HDACs
3.2. Class II HDACs
3.3. Class III HDACs
3.4. Class IV HDACs
4. HDACs Expression Patterns in Hepatocellular Carcinoma
5. Therapeutic Implications: Promising Synthetic HDAC Inhibitors in HCC
5.1. Resminostat
5.2. Vorinostat
5.3. Panobinostat
5.4. Romidepsin
5.5. Belinostat
6. Natural Dietary Components as HDAC Inhibitors
6.1. Short-Chain Fatty Acids (SCFAs)
6.2. Curcumin
6.3. Sulforaphane
6.4. Resveratrol
6.5. Genistein
| Compound and Structure | Molecular Formula (MW) | Source | Study Model | Key Findings | Ref. |
|---|---|---|---|---|---|
Acetic acid | C2H4O2 (60.05 g/mol) | Natural SCFA; Bifidobacteria | In vivo and ex vivo | Anti-tumor activity, efficient immunity | [156] |
Propionic acid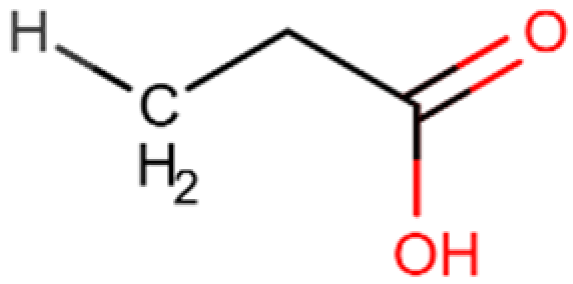 | C3H6O2 (74.08 g/mol) | Natural SCFA; Bacteroidetes | Cell lines; Xenograft mouse model | Tumor regression, enhanced apoptosis and H3 acetylation | [157] |
Butyric acid | C4H8O2 (88.11 g/mol) | Natural SCFA; Firmicutes | Cell lines | Cytotoxic activity, increased acetylation | [159,161,162] |
Valeric acid | C5H10O2 (102.13 g/mol) | Natural SCFA; Clostridia | Cell lines; Xenograft mouse model | Antitumor effects | [165] |
Curcumin | C21H20O6 (368.4 g/mol) | Turmeric (Curcuma longa) | Cell lines | Apoptotic and antiproliferative activity | [169,170] |
Sulforaphane | C6H11NOS2 (177.3 g/mol) | Cruciferous vegetables | Cell lines | Apoptosis induction, gene regulation | [173] |
Resveratrol | C14H12O3 (228.24 g/mol) | Red grapes | In vitro and ex vivo | Antiproliferative effects | [182] |
Genistein | C15H10O5 (270.24 g/mol) | Soy products | Cell lines | Cell proliferation inhibition, increase apoptosis | [187,188] |
7. Future Perspectives and Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CoREST | Corepressor of REST |
| DEN | Diethynitrosamine |
| EMT | Epithelial–mesenchymal transition |
| HATs | Histone acetyltransferases |
| HCC | Hepatocellular carcinomas |
| HIF1α | Hypoxia-inducible factor 1 α |
| HDACi | Histone deacetylase inhibitor |
| HDAC | Histone deacetylase |
| MAD | Matrix associated deacetylase |
| MEF2 | Myocyte enhancer factor 2 |
| mPTP | Mitochondrial permeability transition pore |
| N-CoR | Nuclear receptor co-repressor |
| NaB | Sodium butyrate |
| ncRNA | non-coding RNA |
| NES | Nuclear export signal |
| NLS | Nuclear localization signal |
| NuRD | Nucleosome remodeling and deacetylase complex |
| ODC1 | Ornithine decarboxylase 1 |
| OS | Overall survival |
| PD-L1 | Programmed death ligand 1 |
| PFS | Progression free survival |
| PTM | Post-translational modification |
| Rpd3 | Reduced potassium dependency protein 3 |
| SAHA | Suberoylanilide hydroxamic acid |
| SCFA | Short-chain fatty acid |
| Sir2 | Silent information regulator 2 |
| Treg | Regulatory T cell |
| TSA | Trichostatin A |
| TSG | Tumor suppressor gene |
| VA | Valeric acid |
| ZnF | Zinc finger domain |
References
- Choi, S.-W.; Friso, S. Epigenetics: A New Bridge between Nutrition and Health. Adv. Nutr. 2010, 1, 8–16. [Google Scholar] [CrossRef]
- Handy, D.E.; Castro, R.; Loscalzo, J. Epigenetic Modifications: Basic Mechanisms and Role in Cardiovascular Disease. Circulation 2011, 123, 2145–2156. [Google Scholar] [CrossRef]
- Li, Y. Modern Epigenetics Methods in Biological Research. Methods 2021, 187, 104–113. [Google Scholar] [CrossRef]
- Udali, S.; Guarini, P.; Moruzzi, S.; Ruzzenente, A.; Tammen, S.A.; Guglielmi, A.; Conci, S.; Pattini, P.; Olivieri, O.; Corrocher, R.; et al. Global DNA Methylation and Hydroxymethylation Differ in Hepatocellular Carcinoma and Cholangiocarcinoma and Relate to Survival Rate. Hepatology 2015, 62, 496–504. [Google Scholar] [CrossRef]
- Udali, S.; Guarini, P.; Ruzzenente, A.; Ferrarini, A.; Guglielmi, A.; Lotto, V.; Tononi, P.; Pattini, P.; Moruzzi, S.; Campagnaro, T.; et al. DNA Methylation and Gene Expression Profiles Show Novel Regulatory Pathways in Hepatocellular Carcinoma. Clin. Epigenetics 2015, 7, 43. [Google Scholar] [CrossRef]
- Udali, S.; De Santis, D.; Ruzzenente, A.; Moruzzi, S.; Mazzi, F.; Beschin, G.; Tammen, S.A.; Campagnaro, T.; Pattini, P.; Olivieri, O.; et al. DNA Methylation and Hydroxymethylation in Primary Colon Cancer and Synchronous Hepatic Metastasis. Front. Genet. 2018, 8, 229. [Google Scholar] [CrossRef]
- Friso, S.; Carvajal, C.A.; Fardella, C.E.; Olivieri, O. Epigenetics and Arterial Hypertension: The Challenge of Emerging Evidence. Transl. Res. 2015, 165, 154–165. [Google Scholar] [CrossRef]
- Moruzzi, S.; Guarini, P.; Udali, S.; Ruzzenente, A.; Guglielmi, A.; Conci, S.; Pattini, P.; Martinelli, N.; Olivieri, O.; Tammen, S.A.; et al. One-Carbon Genetic Variants and the Role of MTHFD1 1958G>A in Liver and Colon Cancer Risk According to Global DNA Methylation. PLoS ONE 2017, 12, e0185792. [Google Scholar] [CrossRef]
- Friso, S.; Lotto, V.; Corrocher, R.; Choi, S.W. Vitamin B6 and Cardiovascular Disease. Subcell. Biochem. 2012, 56, 265–290. [Google Scholar] [CrossRef]
- Moruzzi, S.; Udali, S.; Ruzzenente, A.; Guglielmi, A.; Guarini, P.; Martinelli, N.; Conci, S.; Mazzi, F.; Pattini, P.; Tammen, S.A.; et al. The RFC1 80G>A, among Common One-Carbon Polymorphisms, Relates to Survival Rate According to DNA Global Methylation in Primary Liver Cancers. PLoS ONE 2016, 11, e0167534. [Google Scholar] [CrossRef]
- Liu, J.; Huang, B.; Ding, F.; Li, Y. Environment Factors, DNA Methylation, and Cancer. Environ. Geochem. Health 2023, 45, 7543–7568. [Google Scholar] [CrossRef] [PubMed]
- Nishiyama, A.; Nakanishi, M. Navigating the DNA Methylation Landscape of Cancer. Trends Genet. 2021, 37, 1012–1027. [Google Scholar] [CrossRef] [PubMed]
- Deng, S.; Zhang, J.; Su, J.; Zuo, Z.; Zeng, L.; Liu, K.; Zheng, Y.; Huang, X.; Bai, R.; Zhuang, L.; et al. RNA m6A Regulates Transcription via DNA Demethylation and Chromatin Accessibility. Nat. Genet. 2022, 54, 1427–1437. [Google Scholar] [CrossRef]
- Yamaguchi, K.; Chen, X.; Rodgers, B.; Miura, F.; Bashtrykov, P.; Bonhomme, F.; Salinas-Luypaert, C.; Haxholli, D.; Gutekunst, N.; Aygenli, B.Ö.; et al. Non-Canonical Functions of UHRF1 Maintain DNA Methylation Homeostasis in Cancer Cells. Nat. Commun. 2024, 15, 2960. [Google Scholar] [CrossRef]
- Salerno, D.; Peruzzi, G.; Pascucci, G.R.; Levrero, M.; Belloni, L.; Pediconi, N. miRNA-27a-3p Is Involved in the Plasticity of Differentiated Hepatocytes. Gene 2024, 913, 148387. [Google Scholar] [CrossRef]
- Udali, S.; Castagna, A.; Corbella, M.; Ruzzenente, A.; Moruzzi, S.; Mazzi, F.; Campagnaro, T.; De Santis, D.; Franceschi, A.; Pattini, P.; et al. Hepcidin and DNA Promoter Methylation in Hepatocellular Carcinoma. Eur. J. Clin. Investig. 2018, 48, e12870. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Feng, J.; Mi, Y.; Fan, W.; Qin, R.; Mei, Y.; Jin, G.; Mao, J.; Zhang, H. Epigenetic Activation of Cytochrome P450 1A2 Sensitizes Hepatocellular Carcinoma Cells to Sorafenib. Drug Metab. Dispos. 2024, 52, 555–564. [Google Scholar] [CrossRef]
- Liu, C.; Liu, L.; Shan, J.; Shen, J.; Xu, Y.; Zhang, Q.; Yang, Z.; Wu, L.; Xia, F.; Bie, P.; et al. Histone Deacetylase 3 Participates in Self-Renewal of Liver Cancer Stem Cells through Histone Modification. Cancer Lett. 2013, 339, 60–69. [Google Scholar] [CrossRef]
- Liu, K.-Y.; Wang, L.-T.; Hsu, S.-H. Modification of Epigenetic Histone Acetylation in Hepatocellular Carcinoma. Cancers 2018, 10, 8. [Google Scholar] [CrossRef]
- Seo, Y.; Kim, D.K.; Park, J.; Park, S.J.; Park, J.J.; Cheon, J.H.; Kim, T.I. A Comprehensive Understanding of Post-Translational Modification of Sox2 via Acetylation and O-GlcNAcylation in Colorectal Cancer. Cancers 2024, 16, 1035. [Google Scholar] [CrossRef]
- Xu, X.; Ding, Y.; Jin, J.; Xu, C.; Hu, W.; Wu, S.; Ding, G.; Cheng, R.; Cao, L.; Jia, S. Post-Translational Modification of CDK1-STAT3 Signaling by Fisetin Suppresses Pancreatic Cancer Stem Cell Properties. Cell Biosci. 2023, 13, 176. [Google Scholar] [CrossRef] [PubMed]
- El Omari, N.; Bakrim, S.; Khalid, A.; Abdalla, A.N.; Almalki, W.H.; Lee, L.-H.; Ardianto, C.; Ming, L.C.; Bouyahya, A. Molecular Mechanisms Underlying the Clinical Efficacy of Panobinostat Involve Stochasticity of Epigenetic Signaling, Sensitization to Anticancer Drugs, and Induction of Cellular Cell Death Related to Cellular Stresses. Biomed. Pharmacother. 2023, 164, 114886. [Google Scholar] [CrossRef] [PubMed]
- Fattahi, S.; Amjadi-Moheb, F.; Tabaripour, R.; Ashrafi, G.H.; Akhavan-Niaki, H. PI3K/AKT/mTOR Signaling in Gastric Cancer: Epigenetics and Beyond. Life Sci. 2020, 262, 118513. [Google Scholar] [CrossRef]
- Wils, L.J.; Bijlsma, M.F. Epigenetic Regulation of the Hedgehog and Wnt Pathways in Cancer. Crit. Rev. Oncol./Hematol. 2018, 121, 23–44. [Google Scholar] [CrossRef]
- Minucci, S.; Pelicci, P.G. Histone Deacetylase Inhibitors and the Promise of Epigenetic (and More) Treatments for Cancer. Nat. Rev. Cancer 2006, 6, 38–51. [Google Scholar] [CrossRef]
- Wu, C.; Song, Q.; Gao, S.; Wu, S. Targeting HDACs for Diffuse Large B-Cell Lymphoma Therapy. Sci. Rep. 2024, 14, 289. [Google Scholar] [CrossRef]
- Kawaf, R.R.; Ramadan, W.S.; El-Awady, R. Deciphering the Interplay of Histone Post-Translational Modifications in Cancer: Co-Targeting Histone Modulators for Precision Therapy. Life Sci. 2024, 346, 122639. [Google Scholar] [CrossRef]
- Kouzarides, T. Chromatin Modifications and Their Function. Cell 2007, 128, 693–705. [Google Scholar] [CrossRef]
- Zhou, V.W.; Goren, A.; Bernstein, B.E. Charting Histone Modifications and the Functional Organization of Mammalian Genomes. Nat. Rev. Genet. 2011, 12, 7–18. [Google Scholar] [CrossRef]
- Allfrey, V.G.; Faulkner, R.; Mirsky, A.E. Acetylation and Methylation of Histones and Their Possible Role in the Regulation of Rna Synthesis. Proc. Natl. Acad. Sci. USA 1964, 51, 786–794. [Google Scholar] [CrossRef] [PubMed]
- Neganova, M.E.; Klochkov, S.G.; Aleksandrova, Y.R.; Aliev, G. Histone Modifications in Epigenetic Regulation of Cancer: Perspectives and Achieved Progress. Semin. Cancer Biol. 2022, 83, 452–471. [Google Scholar] [CrossRef]
- Archana, B.; D’Cruze, L.; Sundaram, S.; Ramanathan, K.; Ganesh, K. Immunohistochemical Expression of Histone Modification Pattern in Adult Glioblastoma. J. Cancer Res. Ther. 2024, 20, 52–56. [Google Scholar] [CrossRef]
- Lu, Y.; Wang, Y.; Zhang, L.; Ma, Z.; Yu, K.; Shu, Y.; Zou, X.; Yang, J.; Liu, X.; Wang, C.; et al. KAT7 Enhances the Proliferation and Metastasis of Head and Neck Squamous Carcinoma by Promoting the Acetylation Level of LDHA. Cancer Lett. 2024, 590, 216869. [Google Scholar] [CrossRef]
- Gu, R.; Kim, T.-D.; Song, H.; Sui, Y.; Shin, S.; Oh, S.; Janknecht, R. SET7/9-Mediated Methylation Affects Oncogenic Functions of Histone Demethylase JMJD2A. JCI Insight 2023, 8, e164990. [Google Scholar] [CrossRef]
- Roy, A.; Niharika; Chakraborty, S.; Mishra, J.; Singh, S.P.; Patra, S.K. Mechanistic Aspects of Reversible Methylation Modifications of Arginine and Lysine of Nuclear Histones and Their Roles in Human Colon Cancer. Prog. Mol. Biol. Transl. Sci. 2023, 197, 261–302. [Google Scholar] [CrossRef]
- Sawesi, S.; Malkaram, S.A.; Abd Elmageed, Z.Y.; Fandy, T.E. Modulation of the Activity of Histone Lysine Methyltransferases and Demethylases by Curcumin Analog in Leukaemia Cells. J. Cell Mol. Med. 2022, 26, 5624–5633. [Google Scholar] [CrossRef]
- Cheng, C.; Yao, H.; Li, H.; Liu, J.; Liu, Z.; Wu, Y.; Zhu, L.; Hu, H.; Fang, Z.; Wu, L. Blockade of the Deubiquitinating Enzyme USP48 Degrades Oncogenic HMGA2 and Inhibits Colorectal Cancer Invasion and Metastasis. Acta Pharm. Sin. B 2024, 14, 1624–1643. [Google Scholar] [CrossRef]
- Cook, K.W.; Xue, W.; Symonds, P.; Daniels, I.; Gijon, M.; Boocock, D.; Coveney, C.; Miles, A.K.; Shah, S.; Atabani, S.; et al. Homocitrullination of Lysine Residues Mediated by Myeloid-Derived Suppressor Cells in the Tumor Environment Is a Target for Cancer Immunotherapy. J. Immunother. Cancer 2021, 9, e001910. [Google Scholar] [CrossRef]
- Liu, L.; Yin, S.; Gan, W. TRAF6 Promotes PRMT5 Activity in a Ubiquitination-Dependent Manner. Cancers 2023, 15, 2501. [Google Scholar] [CrossRef]
- Lyu, Y.; Yang, Y.; Talwar, V.; Lu, H.; Chen, C.; Salman, S.; Wicks, E.E.; Huang, T.Y.-T.; Drehmer, D.; Wang, Y.; et al. Hypoxia-Inducible Factor 1 Recruits FACT and RNF20/40 to Mediate Histone Ubiquitination and Transcriptional Activation of Target Genes. Cell Rep. 2024, 43, 113972. [Google Scholar] [CrossRef]
- Yang, H.; Du, Y.; Fei, X.; Huang, S.; Yimiti, M.; Yang, X.; Ma, J.; Li, S.; Tuoheniyazi, H.; Zhao, Y.; et al. SUMOylation of the Ubiquitin Ligase Component KEAP1 at K39 Upregulates NRF2 and Its Target Function in Lung Cancer Cell Proliferation. J. Biol. Chem. 2023, 299, 105215. [Google Scholar] [CrossRef]
- Marks, P.A.; Rifkind, R.A.; Richon, V.M.; Breslow, R.; Miller, T.; Kelly, W.K. Histone Deacetylases and Cancer: Causes and Therapies. Nat. Rev. Cancer 2001, 1, 194–202. [Google Scholar] [CrossRef]
- Haery, L.; Thompson, R.C.; Gilmore, T.D. Histone Acetyltransferases and Histone Deacetylases in B- and T-Cell Development, Physiology and Malignancy. Genes. Cancer 2015, 6, 184–213. [Google Scholar] [CrossRef]
- Sun, X.-J.; Man, N.; Tan, Y.; Nimer, S.D.; Wang, L. The Role of Histone Acetyltransferases in Normal and Malignant Hematopoiesis. Front. Oncol. 2015, 5, 108. [Google Scholar] [CrossRef]
- Gray, S.G.; Teh, B.T. Histone Acetylation/Deacetylation and Cancer: An “Open” and “Shut” Case? Curr. Mol. Med. 2001, 1, 401–429. [Google Scholar] [CrossRef]
- Wang, X.; Li, N.; Zheng, M.; Yu, Y.; Zhang, S. Acetylation and Deacetylation of Histone in Adipocyte Differentiation and the Potential Significance in Cancer. Transl. Oncol. 2024, 39, 101815. [Google Scholar] [CrossRef] [PubMed]
- Adams, G.E.; Chandru, A.; Cowley, S.M. Co-Repressor, Co-Activator and General Transcription Factor: The Many Faces of the Sin3 Histone Deacetylase (HDAC) Complex. Biochem. J. 2018, 475, 3921–3932. [Google Scholar] [CrossRef]
- Basta, J.; Rauchman, M. The Nucleosome Remodeling and Deacetylase (NuRD) Complex in Development and Disease. Transl. Res. 2015, 165, 36–47. [Google Scholar] [CrossRef]
- Chen, L.; He, J.; Zhang, Y.; Li, Y.; Zhang, T.; Wang, R.; Bai, L.; Zhao, S.; Liu, E.; Wang, W. Regulation of Endothelial-to-Mesenchymal Transition by Histone Deacetylase 3 Posttranslational Modifications in Neointimal Hyperplasia. Ann. Transl. Med. 2023, 11, 207. [Google Scholar] [CrossRef]
- Zhang, H.; Zhou, P.; Xing, W.; Chen, L.; Zhou, Y.; Yang, H.; Fu, K.; Liu, Z. GLIS2 Prevents Hepatic Fibrosis by Competitively Binding HDAC3 to Inhibit Hepatic Stellate Cell Activation. Cell Mol. Gastroenterol. Hepatol. 2023, 15, 355–372. [Google Scholar] [CrossRef]
- Zhao, H.; Zhang, X.-M.; Xiao, S.; Wu, Z.-R.; Shi, Y.-J.; Xie, M.-J. HDAC11 Is Related to Breast Cancer Prognosis and Inhibits Invasion and Proliferation of Breast Cancer Cells. Int. J. Clin. Exp. Pathol. 2023, 16, 172. [Google Scholar]
- Gray, S.G.; Ekström, T.J. The Human Histone Deacetylase Family. Exp. Cell Res. 2001, 262, 75–83. [Google Scholar] [CrossRef]
- Sacconnay, L.; Carrupt, P.-A.; Nurisso, A. Human Sirtuins: Structures and Flexibility. J. Struct. Biol. 2016, 196, 534–542. [Google Scholar] [CrossRef]
- Cheung, C.H.Y.; Cheng, C.K.; Leung, K.T.; Zhang, C.; Ho, C.Y.; Luo, X.; Kam, A.Y.F.; Xia, T.; Wan, T.S.K.; Pitts, H.A.; et al. C-Terminal Binding Protein (CTBP2) Is a Novel Tumor Suppressor Targeting the MYC-IRF4 Axis in Multiple Myeloma. Blood Adv. 2024, 8, 2217–2234. [Google Scholar] [CrossRef]
- Sanaei, M.; Kavoosi, F. Histone Deacetylase Inhibitors, Intrinsic and Extrinsic Apoptotic Pathways, and Epigenetic Alterations of Histone Deacetylases (HDACs) in Hepatocellular Carcinoma. Iran. J. Pharm. Res. 2021, 20, 324–336. [Google Scholar] [CrossRef] [PubMed]
- Stiegler, P.; De Luca, A.; Bagella, L.; Giordano, A. The COOH-Terminal Region of pRb2/P130 Binds to Histone Deacetylase 1 (HDAC1), Enhancing Transcriptional Repression of the E2F-Dependent Cyclin A Promoter1. Cancer Res. 1998, 58, 5049–5052. [Google Scholar] [PubMed]
- Betz, R.; Gray, S.G.; Ekström, C.; Larsson, C.; Ekström, T.J. Human Histone Deacetylase 2, HDAC2 (Human RPD3), Is Localized to 6q21 by Radiation Hybrid Mapping. Genomics 1998, 52, 245–246. [Google Scholar] [CrossRef] [PubMed]
- Emiliani, S.; Fischle, W.; Van Lint, C.; Al-Abed, Y.; Verdin, E. Characterization of a Human RPD3 Ortholog, HDAC3. Proc. Natl. Acad. Sci. USA 1998, 95, 2795–2800. [Google Scholar] [CrossRef]
- Gao, L.; Cueto, M.A.; Asselbergs, F.; Atadja, P. Cloning and Functional Characterization of HDAC11, a Novel Member of the Human Histone Deacetylase Family. J. Biol. Chem. 2002, 277, 25748–25755. [Google Scholar] [CrossRef] [PubMed]
- Seto, E.; Yoshida, M. Erasers of Histone Acetylation: The Histone Deacetylase Enzymes. Cold Spring Harb. Perspect. Biol. 2014, 6, a018713. [Google Scholar] [CrossRef]
- Yang, X.-J.; Seto, E. The Rpd3/Hda1 Family of Lysine Deacetylases: From Bacteria and Yeast to Mice and Men. Nat. Rev. Mol. Cell Biol. 2008, 9, 206–218. [Google Scholar] [CrossRef] [PubMed]
- Karagianni, P.; Wong, J. HDAC3: Taking the SMRT-N-CoRrect Road to Repression. Oncogene 2007, 26, 5439–5449. [Google Scholar] [CrossRef] [PubMed]
- Buggy, J.J.; Sideris, M.L.; Mak, P.; Lorimer, D.D.; McIntosh, B.; Clark, J.M. Cloning and Characterization of a Novel Human Histone Deacetylase, HDAC8. Biochem. J. 2000, 350 Pt 1, 199–205. [Google Scholar] [CrossRef]
- Takami, Y.; Nakayama, T. N-Terminal Region, C-Terminal Region, Nuclear Export Signal, and Deacetylation Activity of Histone Deacetylase-3 Are Essential for the Viability of the DT40 Chicken B Cell Line. J. Biol. Chem. 2000, 275, 16191–16201. [Google Scholar] [CrossRef]
- Yang, W.-M.; Tsai, S.-C.; Wen, Y.-D.; Fejér, G.; Seto, E. Functional Domains of Histone Deacetylase-3. J. Biol. Chem. 2002, 277, 9447–9454. [Google Scholar] [CrossRef]
- Grozinger, C.M.; Hassig, C.A.; Schreiber, S.L. Three Proteins Define a Class of Human Histone Deacetylases Related to Yeast Hda1p. Proc. Natl. Acad. Sci. USA 1999, 96, 4868–4873. [Google Scholar] [CrossRef]
- Hess, L.; Moos, V.; Lauber, A.A.; Reiter, W.; Schuster, M.; Hartl, N.; Lackner, D.; Boenke, T.; Koren, A.; Guzzardo, P.M.; et al. A Toolbox for Class I HDACs Reveals Isoform Specific Roles in Gene Regulation and Protein Acetylation. PLoS Genet. 2022, 18, e1010376. [Google Scholar] [CrossRef]
- Wright, L.H.; Menick, D.R. A Class of Their Own: Exploring the Nondeacetylase Roles of Class IIa HDACs in Cardiovascular Disease. Am. J. Physiol.-Heart Circ. Physiol. 2016, 311, H199–H206. [Google Scholar] [CrossRef]
- Khochbin, S.; Kao, H.-Y. Histone Deacetylase Complexes: Functional Entities or Molecular Reservoirs. FEBS Lett. 2001, 494, 141–144. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Haberland, M.; Arnold, M.A.; McAnally, J.; Phan, D.; Kim, Y.; Olson, E.N. Regulation of HDAC9 Gene Expression by MEF2 Establishes a Negative-Feedback Loop in the Transcriptional Circuitry of Muscle Differentiation. Mol. Cell Biol. 2007, 27, 518–525. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Marks, P.A.; Rifkind, R.A.; Richon, V.M. Cloning and Characterization of a Histone Deacetylase, HDAC9. Proc. Natl. Acad. Sci. USA 2001, 98, 10572–10577. [Google Scholar] [CrossRef]
- Bertos, N.R.; Wang, A.H.; Yang, X.J. Class II Histone Deacetylases: Structure, Function, and Regulation. Biochem. Cell Biol. 2001, 79, 243–252. [Google Scholar] [CrossRef]
- Curcio, A.; Rocca, R.; Alcaro, S.; Artese, A. The Histone Deacetylase Family: Structural Features and Application of Combined Computational Methods. Pharmaceuticals 2024, 17, 620. [Google Scholar] [CrossRef]
- Liu, P.; Xiao, J.; Wang, Y.; Song, X.; Huang, L.; Ren, Z.; Kitazato, K.; Wang, Y. Posttranslational Modification and beyond: Interplay between Histone Deacetylase 6 and Heat-Shock Protein 90. Mol. Med. 2021, 27, 110. [Google Scholar] [CrossRef]
- Guardiola, A.R.; Yao, T.-P. Molecular Cloning and Characterization of a Novel Histone Deacetylase HDAC10. J. Biol. Chem. 2002, 277, 3350–3356. [Google Scholar] [CrossRef]
- Schwer, B.; Verdin, E. Conserved Metabolic Regulatory Functions of Sirtuins. Cell Metab. 2008, 7, 104–112. [Google Scholar] [CrossRef]
- Khatun, S.; Prasad Bhagat, R.; Dutta, R.; Datta, A.; Jaiswal, A.; Halder, S.; Jha, T.; Amin, S.A.; Gayen, S. Unraveling HDAC11: Epigenetic Orchestra in Different Diseases and Structural Insights for Inhibitor Design. Biochem. Pharmacol. 2024, 225, 116312. [Google Scholar] [CrossRef]
- Tian, Y.; Lv, W.; Li, X.; Wang, C.; Wang, D.; Wang, P.G.; Jin, J.; Shen, J. Stabilizing HDAC11 with SAHA to Assay Slow-Binding Benzamide Inhibitors. Bioorganic Med. Chem. Lett. 2017, 27, 2943–2945. [Google Scholar] [CrossRef] [PubMed]
- Bray, F.; Laversanne, M.; Sung, H.; Ferlay, J.; Siegel, R.L.; Soerjomataram, I.; Jemal, A. Global Cancer Statistics 2022: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA A Cancer J. Clin. 2024, 74, 229–263. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.D.; Hainaut, P.; Gores, G.J.; Amadou, A.; Plymoth, A.; Roberts, L.R. A Global View of Hepatocellular Carcinoma: Trends, Risk, Prevention and Management. Nat. Rev. Gastroenterol. Hepatol. 2019, 16, 589–604. [Google Scholar] [CrossRef] [PubMed]
- Sato-Espinoza, K.; Valdivia-Herrera, M.; Chotiprasidhi, P.; Diaz-Ferrer, J. Hepatocellular Carcinoma in Patients without Cirrhosis. World J. Gastroenterol. 2025, 31, 107100. [Google Scholar] [CrossRef] [PubMed]
- Amin, N.; Anwar, J.; Sulaiman, A.; Naumova, N.N.; Anwar, N. Hepatocellular Carcinoma: A Comprehensive Review. Diseases 2025, 13, 207. [Google Scholar] [CrossRef]
- Fan, C.; Kam, S.; Ramadori, P. Metabolism-Associated Epigenetic and Immunoepigenetic Reprogramming in Liver Cancer. Cancers 2021, 13, 5250. [Google Scholar] [CrossRef]
- Kim, H.S.; Shen, Q.; Nam, S.W. Histone Deacetylases and Their Regulatory MicroRNAs in Hepatocarcinogenesis. J. Korean Med. Sci. 2015, 30, 1375–1380. [Google Scholar] [CrossRef] [PubMed]
- Xie, J.; Liu, R.; Cai, Y.; Liu, D. HDAC1: A Promising Target for Cancer Treatment: Insights from a Thorough Analysis of Tumor Functions. Transl. Cancer Res. 2024, 13, 5300–5315. [Google Scholar] [CrossRef]
- Freese, K.; Seitz, T.; Dietrich, P.; Lee, S.M.L.; Thasler, W.E.; Bosserhoff, A.; Hellerbrand, C. Histone Deacetylase Expressions in Hepatocellular Carcinoma and Functional Effects of Histone Deacetylase Inhibitors on Liver Cancer Cells In Vitro. Cancers 2019, 11, 1587. [Google Scholar] [CrossRef]
- Zhou, H.; Xu, J.; Zhang, C.; Wen, Y. Aberrant Histone Deacetylase 1 Expression Upregulates Vimentin Expression via an NF-κB-Dependent Pathway in Hepatocellular Carcinoma. Oncol. Lett. 2019, 18, 339–347. [Google Scholar] [CrossRef]
- Buurman, R.; Gürlevik, E.; Schäffer, V.; Eilers, M.; Sandbothe, M.; Kreipe, H.; Wilkens, L.; Schlegelberger, B.; Kühnel, F.; Skawran, B. Histone Deacetylases Activate Hepatocyte Growth Factor Signaling by Repressing MicroRNA-449 in Hepatocellular Carcinoma Cells. Gastroenterology 2012, 143, 811–820.e15. [Google Scholar] [CrossRef]
- Ler, S.Y.; Leung, C.H.W.; Khin, L.W.; Lu, G.-D.; Salto-Tellez, M.; Hartman, M.; Iau, P.T.C.; Yap, C.T.; Hooi, S.C. HDAC1 and HDAC2 Independently Predict Mortality in Hepatocellular Carcinoma by a Competing Risk Regression Model in a Southeast Asian Population. Oncol. Rep. 2015, 34, 2238–2250. [Google Scholar] [CrossRef] [PubMed]
- Lachenmayer, A.; Toffanin, S.; Cabellos, L.; Alsinet, C.; Hoshida, Y.; Villanueva, A.; Minguez, B.; Tsai, H.-W.; Ward, S.C.; Thung, S.; et al. Combination Therapy for Hepatocellular Carcinoma: Additive Preclinical Efficacy of the HDAC Inhibitor Panobinostat with Sorafenib. J. Hepatol. 2012, 56, 1343–1350. [Google Scholar] [CrossRef]
- Fan, J.; Lou, B.; Chen, W.; Zhang, J.; Lin, S.; Lv, F.; Chen, Y. Down-Regulation of HDAC5 Inhibits Growth of Human Hepatocellular Carcinoma by Induction of Apoptosis and Cell Cycle Arrest. Tumor Biol. 2014, 35, 11523–11532. [Google Scholar] [CrossRef]
- Ye, M.; Fang, Z.; Gu, H.; Song, R.; Ye, J.; Li, H.; Wu, Z.; Zhou, S.; Li, P.; Cai, X.; et al. Histone Deacetylase 5 Promotes the Migration and Invasion of Hepatocellular Carcinoma via Increasing the Transcription of Hypoxia-Inducible Factor-1α under Hypoxia Condition. Tumour Biol. 2017, 39, 1010428317705034. [Google Scholar] [CrossRef]
- Xu, X.; Ding, P.; Shi, L.; Wu, G.; Ma, X. LukS-PV Inhibits Hepatocellular Carcinoma Cells Migration by Downregulating HDAC6 Expression. BMC Cancer 2022, 22, 630. [Google Scholar] [CrossRef] [PubMed]
- Mikami, D.; Kobayashi, M.; Uwada, J.; Yazawa, T.; Kamiyama, K.; Nishimori, K.; Nishikawa, Y.; Nishikawa, S.; Yokoi, S.; Taniguchi, T.; et al. AR420626, a Selective Agonist of GPR41/FFA3, Suppresses Growth of Hepatocellular Carcinoma Cells by Inducing Apoptosis via HDAC Inhibition. Ther. Adv. Med. Oncol. 2020, 12, 1758835920913432. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Du, C.; Lv, Z.; Ding, C.; Cheng, J.; Xie, H.; Zhou, L.; Zheng, S. The Up-Regulation of Histone Deacetylase 8 Promotes Proliferation and Inhibits Apoptosis in Hepatocellular Carcinoma. Dig. Dis. Sci. 2013, 58, 3545–3553. [Google Scholar] [CrossRef]
- Yang, W.; Feng, Y.; Zhou, J.; Cheung, O.K.-W.; Cao, J.; Wang, J.; Tang, W.; Tu, Y.; Xu, L.; Wu, F.; et al. A Selective HDAC8 Inhibitor Potentiates Antitumor Immunity and Efficacy of Immune Checkpoint Blockade in Hepatocellular Carcinoma. Sci. Transl. Med. 2021, 13, eaaz6804. [Google Scholar] [CrossRef]
- Yang, Z.; Zhang, L.; Liu, J.; Yang, L.; Xue, H.; Bai, S.; Wang, K. PD-L1 Combined with HDAC9 Is a Useful Prognostic Predictor in Hepatocellular Carcinoma. Transl. Cancer Res. 2021, 10, 2305. [Google Scholar] [CrossRef] [PubMed]
- Gong, D.; Zeng, Z.; Yi, F.; Wu, J. Inhibition of Histone Deacetylase 11 Promotes Human Liver Cancer Cell Apoptosis. Am. J. Transl. Res. 2019, 11, 983–990. [Google Scholar]
- Bates, S.E. Epigenetic Therapies for Cancer. N. Engl. J. Med. 2020, 383, 650–663. [Google Scholar] [CrossRef]
- Cheng, B.; Pan, W.; Xiao, Y.; Ding, Z.; Zhou, Y.; Fei, X.; Liu, J.; Su, Z.; Peng, X.; Chen, J. HDAC-Targeting Epigenetic Modulators for Cancer Immunotherapy. Eur. J. Med. Chem. 2024, 265, 116129. [Google Scholar] [CrossRef]
- Tiwari, V.; Banerjee, S.K. Therapeutic Potential of HDAC Inhibitors in the Treatment of Cardiac Diseases: A Short Review. Curr. Drug Targets 2023, 24, 718–727. [Google Scholar] [CrossRef]
- Patel, V.K.; Shirbhate, E.; Tiwari, P.; Kore, R.; Veerasamy, R.; Mishra, A.; Rajak, H. Multi-Targeted HDAC Inhibitors as Anticancer Agents: Current Status and Future Prospective. Curr. Med. Chem. 2023, 30, 2762–2795. [Google Scholar] [CrossRef] [PubMed]
- Waitman, K.; Parise-Filho, R. New Kinase and HDAC Hybrid Inhibitors: Recent Advances and Perspectives. Future Med. Chem. 2022, 14, 745–766. [Google Scholar] [CrossRef]
- Shirbhate, E.; Singh, V.; Jahoriya, V.; Mishra, A.; Veerasamy, R.; Tiwari, A.K.; Rajak, H. Dual Inhibitors of HDAC and Other Epigenetic Regulators: A Novel Strategy for Cancer Treatment. Eur. J. Med. Chem. 2024, 263, 115938. [Google Scholar] [CrossRef]
- Karagianni, F.; Piperi, C.; Valero-Diaz, S.; Amato, C.; Vaque, J.P.; Casar, B.; Papadavid, E. Combination of JAKi and HDACi Exerts Antiangiogenic Potential in Cutaneous T-Cell Lymphoma. Cancers 2024, 16, 3176. [Google Scholar] [CrossRef]
- Huang, W.; Zhu, Q.; Shi, Z.; Tu, Y.; Li, Q.; Zheng, W.; Yuan, Z.; Li, L.; Zu, X.; Hao, Y.; et al. Dual Inhibitors of DNMT and HDAC Induce Viral Mimicry to Induce Antitumour Immunity in Breast Cancer. Cell Death Discov. 2024, 10, 143. [Google Scholar] [CrossRef]
- Zhao, J.; Gray, S.G.; Greene, C.M.; Lawless, M.W. Unmasking the Pathological and Therapeutic Potential of Histone Deacetylases for Liver Cancer. Expert Rev. Gastroenterol. Hepatol. 2019, 13, 247–256. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.; Lee, Y.B.; Cho, E.J.; Lee, J.-H.; Yu, S.J.; Kim, Y.J.; Yoon, J.-H. CKD-5, a Novel Pan-Histone Deacetylase Inhibitor, Synergistically Enhances the Efficacy of Sorafenib for Hepatocellular Carcinoma. BMC Cancer 2020, 20, 1001. [Google Scholar] [CrossRef] [PubMed]
- Streubel, G.; Schrepfer, S.; Kallus, H.; Parnitzke, U.; Wulff, T.; Hermann, F.; Borgmann, M.; Hamm, S. Histone Deacetylase Inhibitor Resminostat in Combination with Sorafenib Counteracts Platelet-Mediated pro-Tumoral Effects in Hepatocellular Carcinoma. Sci. Rep. 2021, 11, 9587. [Google Scholar] [CrossRef]
- Wu, B.; Tapadar, S.; Ruan, Z.; Sun, C.Q.; Arnold, R.S.; Olugbami, J.O.; Johnston, A.; Arunsi, U.; Gaul, D.A.; Petros, J.A.; et al. A Novel Liver Cancer-Selective Histone Deacetylase Inhibitor Is Effective Against Hepatocellular Carcinoma and Induces Durable Responses with Immunotherapy. bioRxiv 2024, 7, 3155–3169. [Google Scholar] [CrossRef]
- Tu, Y.; Wu, H.; Zhong, C.; Liu, Y.; Xiong, Z.; Chen, S.; Wang, J.; Wong, P.P.-C.; Yang, W.; Liang, Z.; et al. Pharmacological Activation of STAT1-GSDME Pyroptotic Circuitry Reinforces Epigenetic Immunotherapy for Hepatocellular Carcinoma. Gut 2024, 74, 613–627. [Google Scholar] [CrossRef] [PubMed]
- Lin, Z.-Z.; Hu, M.C.-T.; Hsu, C.; Wu, Y.-M.; Lu, Y.-S.; Ho, J.-A.A.; Yeh, S.-H.; Chen, P.-J.; Cheng, A.-L. Synergistic Efficacy of Telomerase-Specific Oncolytic Adenoviral Therapy and Histone Deacetylase Inhibition in Human Hepatocellular Carcinoma. Cancer Lett. 2023, 556, 216063. [Google Scholar] [CrossRef]
- Xiao, R.; Jin, H.; Huang, F.; Huang, B.; Wang, H.; Wang, Y.-G. Oncolytic Virotherapy for Hepatocellular Carcinoma: A Potent Immunotherapeutic Landscape. World J. Gastrointest. Oncol. 2024, 16, 2867–2876. [Google Scholar] [CrossRef]
- Lai, S.-Y.; Zhu, X.-J.; Sun, W.-D.; Bi, S.-Z.; Zhang, C.-Y.; Liu, A.; Li, J.-H. Nicotinamide N-Methyltransferase (NNMT) and Liver Cancer: From Metabolic Networks to Therapeutic Targets. Biomolecules 2025, 15, 719. [Google Scholar] [CrossRef]
- Ulanovskaya, O.A.; Zuhl, A.M.; Cravatt, B.F. NNMT Promotes Epigenetic Remodeling in Cancer by Creating a Metabolic Methylation Sink. Nat. Chem. Biol. 2013, 9, 300–306. [Google Scholar] [CrossRef]
- Shen, C.; Li, M.; Duan, Y.; Jiang, X.; Hou, X.; Xue, F.; Zhang, Y.; Luo, Y. HDAC Inhibitors Enhance the Anti-Tumor Effect of Immunotherapies in Hepatocellular Carcinoma. Front. Immunol. 2023, 14, 1170207. [Google Scholar] [CrossRef]
- Eckschlager, T.; Plch, J.; Stiborova, M.; Hrabeta, J. Histone Deacetylase Inhibitors as Anticancer Drugs. Int. J. Mol. Sci. 2017, 18, 1414. [Google Scholar] [CrossRef]
- Bitzer, M.; Horger, M.; Giannini, E.G.; Ganten, T.M.; Wörns, M.A.; Siveke, J.T.; Dollinger, M.M.; Gerken, G.; Scheulen, M.E.; Wege, H.; et al. Resminostat plus Sorafenib as Second-Line Therapy of Advanced Hepatocellular Carcinoma—The SHELTER Study. J. Hepatol. 2016, 65, 280–288. [Google Scholar] [CrossRef] [PubMed]
- Fu, M.; Shi, W.; Li, Z.; Liu, H. Activation of mPTP-Dependent Mitochondrial Apoptosis Pathway by a Novel Pan HDAC Inhibitor Resminostat in Hepatocellular Carcinoma Cells. Biochem. Biophys. Res. Commun. 2016, 477, 527–533. [Google Scholar] [CrossRef]
- Soukupova, J.; Bertran, E.; Peñuelas-Haro, I.; Urdiroz-Urricelqui, U.; Borgman, M.; Kohlhof, H.; Fabregat, I. Resminostat Induces Changes in Epithelial Plasticity of Hepatocellular Carcinoma Cells and Sensitizes Them to Sorafenib-Induced Apoptosis. Oncotarget 2017, 8, 110367–110379. [Google Scholar] [CrossRef] [PubMed]
- Yan, S.; Chen, L.; Zhuang, H.; Yang, H.; Yang, Y.; Zhang, N.; Liu, R. HDAC Inhibition Sensitize Hepatocellular Carcinoma to Lenvatinib via Suppressing AKT Activation. Int. J. Biol. Sci. 2024, 20, 3046–3060. [Google Scholar] [CrossRef] [PubMed]
- Salama, B.M.; Helmy, M.W.; Fouad, H.; Shamaa, M.M.; Houssen, M.E. The Synergistic Antitumor Effect of Decitabine and Vorinostat Combination on HepG2 Human Hepatocellular Carcinoma Cell Line via Epigenetic Modulation of Autophagy–Apoptosis Molecular Crosstalk. Curr. Issues Mol. Biol. 2023, 45, 5935–5949. [Google Scholar] [CrossRef] [PubMed]
- Li, J.-Y.; Tian, T.; Han, B.; Yang, T.; Guo, Y.-X.; Wu, J.-Y.; Chen, Y.-S.; Yang, Q.; Xie, R.-J. Suberoylanilide Hydroxamic Acid Upregulates Reticulophagy Receptor Expression and Promotes Cell Death in Hepatocellular Carcinoma Cells. World J. Gastroenterol. 2023, 29, 5038–5053. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Xu, L.; Yue, S.; Su, H.; Chen, X.; Liu, Q.; Li, H.; Liang, H.; Chen, X.; He, J.; et al. Targeting N4-Acetylcytidine Suppresses Hepatocellular Carcinoma Progression by Repressing eEF2-Mediated HMGB2 mRNA Translation. Cancer Commun. 2024, 44, 1018–1041. [Google Scholar] [CrossRef]
- Choi, C.; Lee, G.H.; Son, A.; Yoo, G.S.; Yu, J.I.; Park, H.C. Downregulation of Mcl-1 by Panobinostat Potentiates Proton Beam Therapy in Hepatocellular Carcinoma Cells. Cells 2021, 10, 554. [Google Scholar] [CrossRef]
- Di Fazio, P.; Schneider-Stock, R.; Neureiter, D.; Okamoto, K.; Wissniowski, T.; Gahr, S.; Quint, K.; Meissnitzer, M.; Alinger, B.; Montalbano, R.; et al. The Pan-Deacetylase Inhibitor Panobinostat Inhibits Growth of Hepatocellular Carcinoma Models by Alternative Pathways of Apoptosis. Cell Oncol. 2010, 32, 285–300. [Google Scholar] [CrossRef]
- Sun, W.-J.; Huang, H.; He, B.; Hu, D.-H.; Li, P.-H.; Yu, Y.-J.; Zhou, X.-H.; Lv, Z.; Zhou, L.; Hu, T.-Y.; et al. Romidepsin Induces G2/M Phase Arrest via Erk/cdc25C/Cdc2/cyclinB Pathway and Apoptosis Induction through JNK/c-Jun/Caspase3 Pathway in Hepatocellular Carcinoma Cells. Biochem. Pharmacol. 2017, 127, 90–100. [Google Scholar] [CrossRef]
- Afaloniati, H.; Angelopoulou, K.; Giakoustidis, A.; Hardas, A.; Pseftogas, A.; Makedou, K.; Gargavanis, A.; Goulopoulos, T.; Iliadis, S.; Papadopoulos, V.; et al. HDAC1/2 Inhibitor Romidepsin Suppresses DEN-Induced Hepatocellular Carcinogenesis in Mice. Onco Targets Ther. 2020, 13, 5575–5588. [Google Scholar] [CrossRef]
- Sun, L.; Liu, Y.; Guo, X.; Cui, T.; Wu, C.; Tao, J.; Cheng, C.; Chu, Q.; Ji, C.; Li, X.; et al. Acetylation-Dependent Regulation of Core Spliceosome Modulates Hepatocellular Carcinoma Cassette Exons and Sensitivity to PARP Inhibitors. Nat. Commun. 2024, 15, 5209. [Google Scholar] [CrossRef]
- Spratlin, J.L.; Pitts, T.M.; Kulikowski, G.N.; Morelli, M.P.; Tentler, J.J.; Serkova, N.J.; Eckhardt, S.G. Synergistic Activity of Histone Deacetylase and Proteasome Inhibition Against Pancreatic and Hepatocellular Cancer Cell Lines. Anticancer. Res. 2011, 31, 1093–1103. [Google Scholar]
- Ma, B.B.Y.; Sung, F.; Tao, Q.; Poon, F.F.; Lui, V.W.; Yeo, W.; Chan, S.L.; Chan, A.T.C. The Preclinical Activity of the Histone Deacetylase Inhibitor PXD101 (Belinostat) in Hepatocellular Carcinoma Cell Lines. Investig. New Drugs 2010, 28, 107–114. [Google Scholar] [CrossRef]
- Llopiz, D.; Ruiz, M.; Villanueva, L.; Iglesias, T.; Silva, L.; Egea, J.; Lasarte, J.J.; Pivette, P.; Trochon-Joseph, V.; Vasseur, B.; et al. Enhanced Anti-Tumor Efficacy of Checkpoint Inhibitors in Combination with the Histone Deacetylase Inhibitor Belinostat in a Murine Hepatocellular Carcinoma Model. Cancer Immunol. Immunother. 2018, 68, 379–393. [Google Scholar] [CrossRef] [PubMed]
- Walewski, J.; Paszkiewicz-Kozik, E.; Borsaru, G.; Hellmann, A.; Janikova, A.; Warszewska, A.; Mais, A.; Ammendola, A.; Herz, T.; Krauss, B.; et al. Resminostat in Patients with Relapsed or Refractory Hodgkin Lymphoma: Results of the Phase II SAPHIRE Study. Leuk. Lymphoma 2019, 60, 675–684. [Google Scholar] [CrossRef]
- Karagianni, F.; Piperi, C.; Mpakou, V.; Spathis, A.; Foukas, P.G.; Dalamaga, M.; Pappa, V.; Papadavid, E. Ruxolitinib with Resminostat Exert Synergistic Antitumor Effects in Cutaneous T-Cell Lymphoma. PLoS ONE 2021, 16, e0248298. [Google Scholar] [CrossRef]
- Ikeda, M.; Ohno, I.; Ueno, H.; Mitsunaga, S.; Hashimoto, Y.; Okusaka, T.; Kondo, S.; Sasaki, M.; Sakamoto, Y.; Takahashi, H.; et al. Phase I Study of Resminostat, an HDAC Inhibitor, Combined with S-1 in Patients with Pre-Treated Biliary Tract or Pancreatic Cancer. Investig. New Drugs 2019, 37, 109–117. [Google Scholar] [CrossRef]
- Brunetto, A.T.; Ang, J.E.; Lal, R.; Olmos, D.; Molife, L.R.; Kristeleit, R.; Parker, A.; Casamayor, I.; Olaleye, M.; Mais, A.; et al. First-in-Human, Pharmacokinetic and Pharmacodynamic Phase I Study of Resminostat, an Oral Histone Deacetylase Inhibitor, in Patients with Advanced Solid Tumors. Clin. Cancer Res. 2013, 19, 5494–5504. [Google Scholar] [CrossRef] [PubMed]
- Tak, W.Y.; Ryoo, B.-Y.; Lim, H.Y.; Kim, D.-Y.; Okusaka, T.; Ikeda, M.; Hidaka, H.; Yeon, J.-E.; Mizukoshi, E.; Morimoto, M.; et al. Phase I/II Study of First-Line Combination Therapy with Sorafenib plus Resminostat, an Oral HDAC Inhibitor, versus Sorafenib Monotherapy for Advanced Hepatocellular Carcinoma in East Asian Patients. Investig. New Drugs 2018, 36, 1072–1084. [Google Scholar] [CrossRef] [PubMed]
- Peng, X.; Zhang, D.; Li, Z.; Fu, M.; Liu, H. mTOR Inhibition Sensitizes Human Hepatocellular Carcinoma Cells to Resminostat. Biochem. Biophys. Res. Commun. 2016, 477, 556–562. [Google Scholar] [CrossRef]
- Parveen, R.; Harihar, D.; Chatterji, B.P. Recent Histone Deacetylase Inhibitors in Cancer Therapy. Cancer 2023, 129, 3372–3380. [Google Scholar] [CrossRef]
- Sanaei, M.; Kavoosi, F.; Pourahmadi, M. Effect of Decitabine (5-Aza-2′-Deoxycytidine, 5-Aza-CdR) in Comparison with Vorinostat (Suberoylanilide Hydroxamic Acid, SAHA) on DNMT1, DNMT3a and DNMT3b, HDAC 1-3, SOCS 1, SOCS 3, JAK2, and STAT3 Gene Expression in Hepatocellular Carcinoma HLE and LCL-PI 11 Cell Lines. Asian Pac. J. Cancer Prev. 2021, 22, 2089–2098. [Google Scholar] [CrossRef] [PubMed]
- Eleutherakis-Papaiakovou, E.; Kanellias, N.; Kastritis, E.; Gavriatopoulou, M.; Terpos, E.; Dimopoulos, M.A. Efficacy of Panobinostat for the Treatment of Multiple Myeloma. J. Oncol. 2020, 2020, 7131802. [Google Scholar] [CrossRef]
- Sun, H.; Song, J.; Sun, M.; Shan, C. Chapter 3—Histone Deacetylase Inhibitors (HDACi), the Ongoing Epigenetic Agents to Enhance Chemotherapy Sensitization. In Epigenetic Regulation in Overcoming Chemoresistance; Wu, C., Wang, L., Eds.; Cancer Sensitizing Agents for Chemotherapy; Academic Press: Cambridge, MA, USA, 2021; Volume 15, pp. 25–39. [Google Scholar]
- Liu, Y.-F.; Chiang, Y.; Hsu, F.-M.; Tsai, C.-L.; Cheng, J.C.-H. Radiosensitization Effect by HDAC Inhibition Improves NKG2D-Dependent Natural Killer Cytotoxicity in Hepatocellular Carcinoma. Front. Oncol. 2022, 12, 1009089. [Google Scholar] [CrossRef] [PubMed]
- Di Fazio, P.; Waldegger, P.; Jabari, S.; Lingelbach, S.; Montalbano, R.; Ocker, M.; Slater, E.P.; Bartsch, D.K.; Illig, R.; Neureiter, D.; et al. Autophagy-Related Cell Death by Pan-Histone Deacetylase Inhibition in Liver Cancer. Oncotarget 2016, 7, 28998–29010. [Google Scholar] [CrossRef] [PubMed]
- Henrici, A.; Montalbano, R.; Neureiter, D.; Krause, M.; Stiewe, T.; Slater, E.P.; Quint, K.; Ocker, M.; Di Fazio, P. The Pan-Deacetylase Inhibitor Panobinostat Suppresses the Expression of Oncogenic miRNAs in Hepatocellular Carcinoma Cell Lines. Mol. Carcinog. 2015, 54, 585–597. [Google Scholar] [CrossRef] [PubMed]
- Bondarev, A.D.; Attwood, M.M.; Jonsson, J.; Chubarev, V.N.; Tarasov, V.V.; Schiöth, H.B. Recent Developments of HDAC Inhibitors: Emerging Indications and Novel Molecules. Br. J. Clin. Pharmacol. 2021, 87, 4577–4597. [Google Scholar] [CrossRef]
- Zhou, H.; Cai, Y.; Liu, D.; Li, M.; Sha, Y.; Zhang, W.; Wang, K.; Gong, J.; Tang, N.; Huang, A.; et al. Pharmacological or Transcriptional Inhibition of Both HDAC1 and 2 Leads to Cell Cycle Blockage and Apoptosis via p21Waf1/Cip1 and p19INK4d Upregulation in Hepatocellular Carcinoma. Cell Prolif. 2018, 51, e12447. [Google Scholar] [CrossRef]
- Afaloniati, H.; Poutahidis, T.; Giakoustidis, A.; Gargavanis, A.; Giakoustidis, D.; Angelopoulou, K. Romidepsin Hepatocellular Carcinoma Suppression in Mice Is Associated with Deregulated Gene Expression of Bone Morphogenetic Protein and Notch Signaling Pathway Components. Mol. Biol. Rep. 2021, 48, 551–562. [Google Scholar] [CrossRef]
- Li, J.; Xiong, X.; Wang, Z.; Zhao, Y.; Shi, Z.; Zhao, M.; Ren, T. In Vitro High-Throughput Drug Sensitivity Screening with Patient-Derived Primary Cells as a Guide for Clinical Practice in Hepatocellular Carcinoma—A Retrospective Evaluation. Clin. Res. Hepatol. Gastroenterol. 2020, 44, 699–710. [Google Scholar] [CrossRef]
- Xie, L.; Song, D.; Lan, J.; Liu, P.; Qin, S.; Ning, Y.; Liu, Q. Plasma Protein Levels and Hepatocellular Carcinoma: A Mendelian Randomization Study with Drug Screening Implications. Discov. Oncol. 2025, 16, 567. [Google Scholar] [CrossRef]
- Kim, N.; Yang, C. Butyrate as a Potential Modulator in Gynecological Disease Progression. Nutrients 2024, 16, 4196. [Google Scholar] [CrossRef]
- Merarchi, M.; Sethi, G.; Shanmugam, M.K.; Fan, L.; Arfuso, F.; Ahn, K.S. Role of Natural Products in Modulating Histone Deacetylases in Cancer. Molecules 2019, 24, 1047. [Google Scholar] [CrossRef]
- Singh, A.K.; Bishayee, A.; Pandey, A.K. Targeting Histone Deacetylases with Natural and Synthetic Agents: An Emerging Anticancer Strategy. Nutrients 2018, 10, 731. [Google Scholar] [CrossRef]
- Choi, S.-W.; Friso, S. (Eds.) Nutrients and Epigenetics; CRC Press: Boca Raton, FL, USA, 2009; ISBN 978-0-429-14672-5. [Google Scholar]
- Fusco, W.; Lorenzo, M.B.; Cintoni, M.; Porcari, S.; Rinninella, E.; Kaitsas, F.; Lener, E.; Mele, M.C.; Gasbarrini, A.; Collado, M.C.; et al. Short-Chain Fatty-Acid-Producing Bacteria: Key Components of the Human Gut Microbiota. Nutrients 2023, 15, 2211. [Google Scholar] [CrossRef]
- Hu, C.; Xu, B.; Wang, X.; Wan, W.; Lu, J.; Kong, D.; Jin, Y.; You, W.; Sun, H.; Mu, X.; et al. Gut Microbiota–Derived Short-chain Fatty Acids Regulate Group 3 Innate Lymphoid Cells in HCC. Hepatology 2023, 77, 48–64. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, M.; Mikami, D.; Uwada, J.; Yazawa, T.; Kamiyama, K.; Kimura, H.; Taniguchi, T.; Iwano, M. A Short-Chain Fatty Acid, Propionate, Enhances the Cytotoxic Effect of Cisplatin by Modulating GPR41 Signaling Pathways in HepG2 Cells. Oncotarget 2018, 9, 31342–31354. [Google Scholar] [CrossRef] [PubMed]
- Kumar, M.; Kaur, R.; Kanthaje, S.; Dhiman, R.K.; Chakraborti, A. Bacterial Metabolite Butyrate in Modulating Sorafenib-Targeted microRNAs to Curtail Its Resistance in Hepatocellular Carcinoma. J. Cancer Res. Clin. Oncol. 2023, 149, 5823–5839. [Google Scholar] [CrossRef]
- Shalhoub, H.; Gonzalez, P.; Dos Santos, A.; Guillermet-Guibert, J.; Moniaux, N.; Dupont, N.; Faivre, J. Simultaneous Activation and Blockade of Autophagy to Fight Hepatocellular Carcinoma. Autophagy Rep. 2024, 3, 2326241. [Google Scholar] [CrossRef] [PubMed]
- Sanaei, M.; Kavoosi, F.; Safari, M. Effect of 5′-Fluoro-2′-Deoxycytidine and Sodium Butyrate on the Gene Expression of the Intrinsic Apoptotic Pathway, P21, P27, and P53 Genes Expression, Cell Viability, and Apoptosis in Human Hepatocellular Carcinoma Cell Lines. Adv. Biomed. Res. 2023, 12, 24. [Google Scholar] [CrossRef]
- Kotantaki, P.; Mosialos, G. The Expression of Tumor Suppressor Gene Cyld Is Upregulated by Histone Deacetylace Inhibitors in Human Hepatocellular Carcinoma Cell Lines. Cell Biochem. Funct. 2016, 34, 465–468. [Google Scholar] [CrossRef]
- Hernández-Oliveras, A.; Izquierdo-Torres, E.; Meneses-Morales, I.; Rodríguez, G.; Zarain-Herzberg, Á.; Santiago-García, J. Histone Deacetylase Inhibitors Promote ATP2A3 Gene Expression in Hepatocellular Carcinoma Cells: P300 as a Transcriptional Regulator. Int. J. Biochem. Cell Biol. 2019, 113, 8–16. [Google Scholar] [CrossRef]
- Cao, S.; Budina, E.; Raczy, M.M.; Solanki, A.; Nguyen, M.; Beckman, T.N.; Reda, J.W.; Hultgren, K.; Ang, P.S.; Slezak, A.J.; et al. A Serine-Conjugated Butyrate Prodrug with High Oral Bioavailability Suppresses Autoimmune Arthritis and Neuroinflammation in Mice. Nat. Biomed. Eng. 2024, 8, 611–627. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Zhao, K.-N.; Vitetta, L. Effects of Intestinal Microbial–Elaborated Butyrate on Oncogenic Signaling Pathways. Nutrients 2019, 11, 1026. [Google Scholar] [CrossRef] [PubMed]
- Han, R.; Nusbaum, O.; Chen, X.; Zhu, Y. Valeric Acid Suppresses Liver Cancer Development by Acting as a Novel HDAC Inhibitor. Mol. Ther. Oncolytics 2020, 19, 8–18. [Google Scholar] [CrossRef]
- Mayorga-Ramos, A.; Barba-Ostria, C.; Simancas-Racines, D.; Guamán, L.P. Protective Role of Butyrate in Obesity and Diabetes: New Insights. Front. Nutr. 2022, 9, 1067647. [Google Scholar] [CrossRef]
- Longtine, A.G.; Greenberg, N.T.; Gonzalez, A.; Lindquist, A.; VanDongen, N.S.; Mahoney, S.A.; Rahman, G.; Clayton, Z.S.; Ziemba, B.P.; Ludwig, K.R.; et al. Oral Supplementation with the Short-Chain Fatty Acid Acetate Ameliorates Age-Related Arterial Dysfunction in Mice. Aging Biol. 2024, 2, 20240033. [Google Scholar] [CrossRef]
- Hirata, S.-I.; Kunisawa, J. Gut Microbiome, Metabolome, and Allergic Diseases. Allergol. Int. 2017, 66, 523–528. [Google Scholar] [CrossRef]
- Marquardt, J.U.; Gomez-Quiroz, L.; Camacho, L.O.A.; Pinna, F.; Lee, Y.-H.; Kitade, M.; Domínguez, M.P.; Castven, D.; Breuhahn, K.; Conner, E.A.; et al. Curcumin Effectively Inhibits Oncogenic NF-kB Signaling and Restrains Stemness Features in Liver Cancer. J. Hepatol. 2015, 63, 661–669. [Google Scholar] [CrossRef]
- Sanaei, M.; Kavoosi, F.; Arabloo, M. Effect of Curcumin in Comparison with Trichostatin A on the Reactivation of Estrogen Receptor Alpha Gene Expression, Cell Growth Inhibition and Apoptosis Induction in Hepatocellular Carcinoma Hepa 1-6 Cell lLine. Asian Pac. J. Cancer Prev. 2020, 21, 1045–1050. [Google Scholar] [CrossRef] [PubMed]
- Anand, P.; Kunnumakkara, A.B.; Newman, R.A.; Aggarwal, B.B. Bioavailability of Curcumin: Problems and Promises. Mol. Pharm. 2007, 4, 807–818. [Google Scholar] [CrossRef] [PubMed]
- Abd El-Hack, M.E.; El-Saadony, M.T.; Swelum, A.A.; Arif, M.; Abo Ghanima, M.M.; Shukry, M.; Noreldin, A.; Taha, A.E.; El-Tarabily, K.A. Curcumin, the Active Substance of Turmeric: Its Effects on Health and Ways to Improve Its Bioavailability. J. Sci. Food Agric. 2021, 101, 5747–5762. [Google Scholar] [CrossRef]
- dos Santos, P.W.D.S.; Machado, A.R.T.; De Grandis, R.A.; Ribeiro, D.L.; Tuttis, K.; Morselli, M.; Aissa, A.F.; Pellegrini, M.; Antunes, L.M.G. Transcriptome and DNA Methylation Changes Modulated by Sulforaphane Induce Cell Cycle Arrest, Apoptosis, DNA Damage, and Suppression of Proliferation in Human Liver Cancer Cells. Food Chem. Toxicol. 2020, 136, 111047. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Shi, C.; Zhang, D.; Zhang, L.; Wang, L.; Gong, Z. Sulforaphane, an NRF2 Agonist, Alleviates Ferroptosis in Acute Liver Failure by Regulating HDAC6 Activity. J. Integr. Med. 2023, 21, 464–473. [Google Scholar] [CrossRef]
- Fantone, S.; Marzioni, D.; Tossetta, G. NRF2/KEAP1 Signaling Inhibitors in Gynecologic Cancers. Expert Rev. Anticancer. Ther. 2024, 24, 1191–1194. [Google Scholar] [CrossRef] [PubMed]
- Schiavoni, V.; Emanuelli, M.; Milanese, G.; Galosi, A.B.; Pompei, V.; Salvolini, E.; Campagna, R. Nrf2 Signaling in Renal Cell Carcinoma: A Potential Candidate for the Development of Novel Therapeutic Strategies. Int. J. Mol. Sci. 2024, 25, 13239. [Google Scholar] [CrossRef] [PubMed]
- Tossetta, G.; Fantone, S.; Togni, L.; Santarelli, A.; Olivieri, F.; Marzioni, D.; Rippo, M.R. Modulation of NRF2/KEAP1 Signaling by Phytotherapeutics in Periodontitis. Antioxidants 2024, 13, 1270. [Google Scholar] [CrossRef]
- Tossetta, G.; Fantone, S.; Piani, F.; Crescimanno, C.; Ciavattini, A.; Giannubilo, S.R.; Marzioni, D. Modulation of NRF2/KEAP1 Signaling in Preeclampsia. Cells 2023, 12, 1545. [Google Scholar] [CrossRef]
- Men, X.; Han, X.; Oh, G.; Im, J.-H.; Lim, J.S.; Cho, G.H.; Choi, S.-I.; Lee, O.-H. Plant Sources, Extraction Techniques, Analytical Methods, Bioactivity, and Bioavailability of Sulforaphane: A Review. Food Sci. Biotechnol. 2023, 33, 539–556. [Google Scholar] [CrossRef]
- Okunade, O.; Niranjan, K.; Ghawi, S.K.; Kuhnle, G.; Methven, L. Supplementation of the Diet by Exogenous Myrosinase via Mustard Seeds to Increase the Bioavailability of Sulforaphane in Healthy Human Subjects after the Consumption of Cooked Broccoli. Mol. Nutr. Food Res. 2018, 62, e1700980. [Google Scholar] [CrossRef]
- Yan, L.; Yan, Y. Therapeutic Potential of Sulforaphane in Liver Diseases: A Review. Front. Pharmacol. 2023, 14, 1256029. [Google Scholar] [CrossRef]
- Venturelli, S.; Berger, A.; Böcker, A.; Busch, C.; Weiland, T.; Noor, S.; Leischner, C.; Schleicher, S.; Mayer, M.; Weiss, T.S.; et al. Resveratrol as a Pan-HDAC Inhibitor Alters the Acetylation Status of Jistone Proteins in Human-Derived Hepatoblastoma Cells. PLoS ONE 2013, 8, e73097. [Google Scholar] [CrossRef]
- Nowrasteh, G.; Zand, A.; Raposa, L.B.; Szabó, L.; Tomesz, A.; Molnár, R.; Kiss, I.; Orsós, Z.; Gerencsér, G.; Gyöngyi, Z.; et al. Fruit Extract, Rich in Polyphenols and Flavonoids, Modifies the Expression of DNMT and HDAC Genes Involved in Epigenetic Processes. Nutrients 2023, 15, 1867. [Google Scholar] [CrossRef]
- Galiniak, S.; Aebisher, D.; Bartusik-Aebisher, D. Health Benefits of Resveratrol Administration. Acta Biochim. Pol. 2019, 66, 13–21. [Google Scholar] [CrossRef]
- de Vries, K.; Strydom, M.; Steenkamp, V. A Brief Updated Review of Advances to Enhance Resveratrol’s Bioavailability. Molecules 2021, 26, 4367. [Google Scholar] [CrossRef]
- Szymkowiak, I.; Marcinkowska, J.; Kucinska, M.; Regulski, M.; Murias, M. Resveratrol Bioavailability After Oral Administration: A Meta-Analysis of Clinical Trial Data. Phytother. Res. 2025, 39, 453–464. [Google Scholar] [CrossRef] [PubMed]
- Sanaei, M.; Kavoosi, F.; Salehi, H. Genistein and Trichostatin A Induction of Estrogen Receptor Alpha Gene Expression, Apoptosis and Cell Growth Inhibition in Hepatocellular Carcinoma HepG 2 Cells. Asian Pac. J. Cancer Prev. 2017, 18, 3445–3450. [Google Scholar] [CrossRef] [PubMed]
- Sanaei, M.; Kavoosi, F.; Roustazadeh, A.; Golestan, F. Effect of Genistein in Comparison with Trichostatin A on Reactivation of DNMTs Genes in Hepatocellular Carcinoma. J. Clin. Transl. Hepatol. 2018, 6, 141–146. [Google Scholar] [CrossRef]
- Sanaei, M.; Kavoosi, F. Effect of DNA Methyltransferase in Comparison to and in Combination with Histone Deacetylase Inhibitors on Hepatocellular Carcinoma HepG2 Cell Line. Asian Pac. J. Cancer Prev. 2019, 20, 1119–1125. [Google Scholar] [CrossRef] [PubMed]
- El-Far, Y.M.; Khodir, A.E.; Emarah, Z.A.; Ebrahim, M.A.; Al-Gayyar, M.M.H. Chemopreventive and Hepatoprotective Effects of Genistein via Inhibition of Oxidative Stress and the Versican/PDGF/PKC Signaling Pathway in Experimentally Induced Hepatocellular Carcinoma in Rats by Thioacetamide. Redox Rep. 2022, 27, 9–20. [Google Scholar] [CrossRef]
- Sharifi-Rad, J.; Quispe, C.; Imran, M.; Rauf, A.; Nadeem, M.; Gondal, T.A.; Ahmad, B.; Atif, M.; Mubarak, M.S.; Sytar, O.; et al. Genistein: An Integrative Overview of Its Mode of Action, Pharmacological Properties, and Health Benefits. Oxid. Med. Cell Longev. 2021, 2021, 3268136. [Google Scholar] [CrossRef]
- Setchell, K.D.R.; Brown, N.M.; Desai, P.; Zimmer-Nechemias, L.; Wolfe, B.E.; Brashear, W.T.; Kirschner, A.S.; Cassidy, A.; Heubi, J.E. Bioavailability of Pure Isoflavones in Healthy Humans and Analysis of Commercial Soy Isoflavone Supplements. J. Nutr. 2001, 131, 1362S–1375S. [Google Scholar] [CrossRef]
- Bian, X.; Piipponen, M.; Liu, Z.; Luo, L.; Geara, J.; Chen, Y.; Sangsuwan, T.; Maselli, M.; Diaz, C.; Bain, C.A.; et al. Epigenetic Memory of Radiotherapy in Dermal Fibroblasts Impairs Wound Repair Capacity in Cancer Survivors. Nat. Commun. 2024, 15, 9286. [Google Scholar] [CrossRef] [PubMed]




| Protein | Gene ID | Aliases | Chr | HDAC Class | Cellular Localization | Histone Substrates | Non-Histone Substrates |
|---|---|---|---|---|---|---|---|
| HDAC1 | 3065 | GON-10, HD1, KDAC1, RPD3, RPD3L1 | 1p35.2 | I | Nucleus, Cytoplasm | Lysine residues of H2A, H2B, H3, H4 | NR1D2, RELA, SP1, SP3, STAT3 and TSHZ3 |
| HDAC2 | 3066 | HD2, KDAC2, RPD3, YAF1 | 6q21 | I | Nucleus, Cytoplasm | Lysine residues of H2A, H2B, H3, H4 | TSHZ3 |
| HDAC3 | 8841 | HD3, KDAC3, RPD3, RPD3-2 | 5q31.3 | I | Plasma membrane, Nucleus, Cytoplasm | H3K27 | KAT5, MEF2D, MAPK14, STAT3 and RARA |
| HDAC4 | 9759 | AHO3, BDMR, HA6116, HD4, HDAC-4, HDAC-A, HDACA, NEDCHF, NEDCHID | 2q37.3 | IIa | Nucleus, Cytoplasm | Lysine residues of H2A, H2B, H3, H4 | HSPA1A and HSPA1B at Lys-77 |
| HDAC5 | 10014 | HD5, NY-CO-9 | 17q21.31 | IIa | Nucleus, Cytoplasm | Lysine residues of H2A, H2B, H3, H4 | RARA |
| HDAC6 | 10013 | CPBHM, HD6, JM21, KDAC6, PPP1R90 | Xp11.23 | IIb | Plasma membrane, Nucleus, Cytoplasm | Lysine residues of H2A, H2B, H3, H4 | Tubulin, α-tubulin, SQSTM1 and CTTN |
| HDAC7 | 51564 | HD7, HD7AA, HDAC7 | 12q13.11 | IIa | Nucleus, Cytoplasm | Lysine residues of H2A, H2B, H3, H4 | RARA and ALKBH5 |
| HDAC8 | 55869 | CDA07, CDLS5, HD8, HDACL1, KDAC8, MRXS6, RPD3, WTS | Xq13.1 | I | Plasma membrane, Nucleus, Cytoplasm | Lysine residues of H2A, H2B, H3, H4 | SMC3 |
| HDAC9 | 9734 | ARCND4, HD7, HD7b, HD9, HDAC, HDAC7, HDAC7BB, HDAC9FL, HDRP, MITR | 7p21.1 | IIa | Nucleus, Cytoplasm | Lysine residues of H2A, H2B, H3, H4 | - |
| HDAC10 | 83933 | HD10 | 22q13.33 | IIb | Nucleus, Cytoplasm | Lysine residues of H2A, H2B, H3, H4 | MSH2 |
| HDAC11 | 79885 | HD11 | 3p25.1 | IV | Plasma membrane, Nucleus, Cytoplasm | Lysine residues of H2A, H2B, H3, H4 | - |
| HDAC Inhibitor Name (Trade Name) and Structure | Aliases | Molecular Formula (MW) | Study Design/Model | Key Findings | Ref. |
|---|---|---|---|---|---|
Resminostat (Kinselby) | 4SC-201 RAS2410 BYK408740 | C16H19N3O4S (349.4 g/mol) | Phase I/II clinical trial; in vitro studies | Apoptosis induction; enhanced antitumor activity; improved efficacy with Sorafenib | [118,119,120] |
Vorinostat (Zolinza) | SAHA MK0683 | C14H20N2O3 (264.32 g/mol) | In vitro and in vivo studies | Improved efficiency with Lenvatinib; synergistic anticancer effect via dual induction of apoptosis and autophagy | [121,122,123] |
Panobinostat (Farydak) | LBH589 | C21H23N3O2 (349.4 g/mol) | Xenograft mouse models; cell lines | Reduction in lung metastasis; enhanced antitumor effect and apoptosis induction in combination with radiotherapy | [124,125,126] |
Romidepsin (Istodax) | Depsipeptide Chromadax FK228 | C24H36N4O6S2 (540.7 g/mol) | In vitro and in vivo studies | Cell cycle arrest; HCC tumor suppression; enhanced outcome with Olaparib | [127,128,129] |
Belinostat (Beleodaq) | PXD101 | C15H14N2O4S (318.3 g/mol) | Cell lines; murine models | Antiproliferative and cytotoxic effects with Bortezomib; apoptotic induction and cell suppression; enhanced anti-tumor immunity | [130,131,132] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sadia, K.; Castagna, A.; Udali, S.; Ambrosani, F.; Pattini, P.; Beri, R.; Argentino, G.; Masutti, M.; Moruzzi, S.; Friso, S. Epigenetic Regulation Through Histone Deacetylation: Implications and Therapeutic Potential in Hepatocellular Carcinoma. Cells 2025, 14, 1337. https://doi.org/10.3390/cells14171337
Sadia K, Castagna A, Udali S, Ambrosani F, Pattini P, Beri R, Argentino G, Masutti M, Moruzzi S, Friso S. Epigenetic Regulation Through Histone Deacetylation: Implications and Therapeutic Potential in Hepatocellular Carcinoma. Cells. 2025; 14(17):1337. https://doi.org/10.3390/cells14171337
Chicago/Turabian StyleSadia, Khulah, Annalisa Castagna, Silvia Udali, Francesca Ambrosani, Patrizia Pattini, Ruggero Beri, Giuseppe Argentino, Maria Masutti, Sara Moruzzi, and Simonetta Friso. 2025. "Epigenetic Regulation Through Histone Deacetylation: Implications and Therapeutic Potential in Hepatocellular Carcinoma" Cells 14, no. 17: 1337. https://doi.org/10.3390/cells14171337
APA StyleSadia, K., Castagna, A., Udali, S., Ambrosani, F., Pattini, P., Beri, R., Argentino, G., Masutti, M., Moruzzi, S., & Friso, S. (2025). Epigenetic Regulation Through Histone Deacetylation: Implications and Therapeutic Potential in Hepatocellular Carcinoma. Cells, 14(17), 1337. https://doi.org/10.3390/cells14171337








