Dedifferentiation-Dependent Regeneration of the Biliary Ductal Epithelium in Response to Hepatic Injury in TFF1-Deficient Mice
Abstract
1. Introduction
2. Materials and Methods
2.1. Animals
2.2. Histology and Immunohistochemistry
2.3. Immunofluorescence
2.4. Statistical Analysis
3. Results
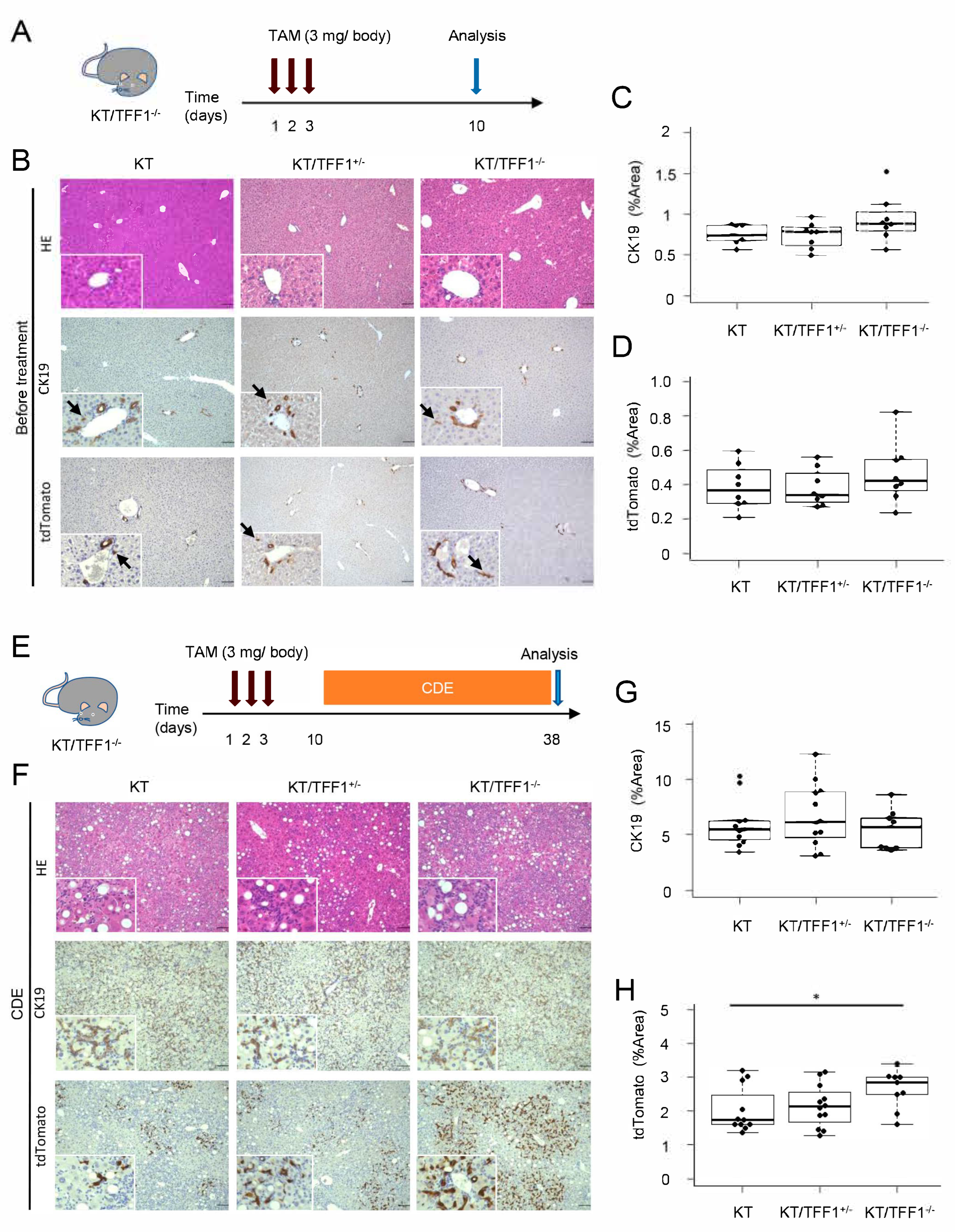
3.1. Loss of TFF1 Resulted in the Proliferation of TdTomato-Labeled Cells
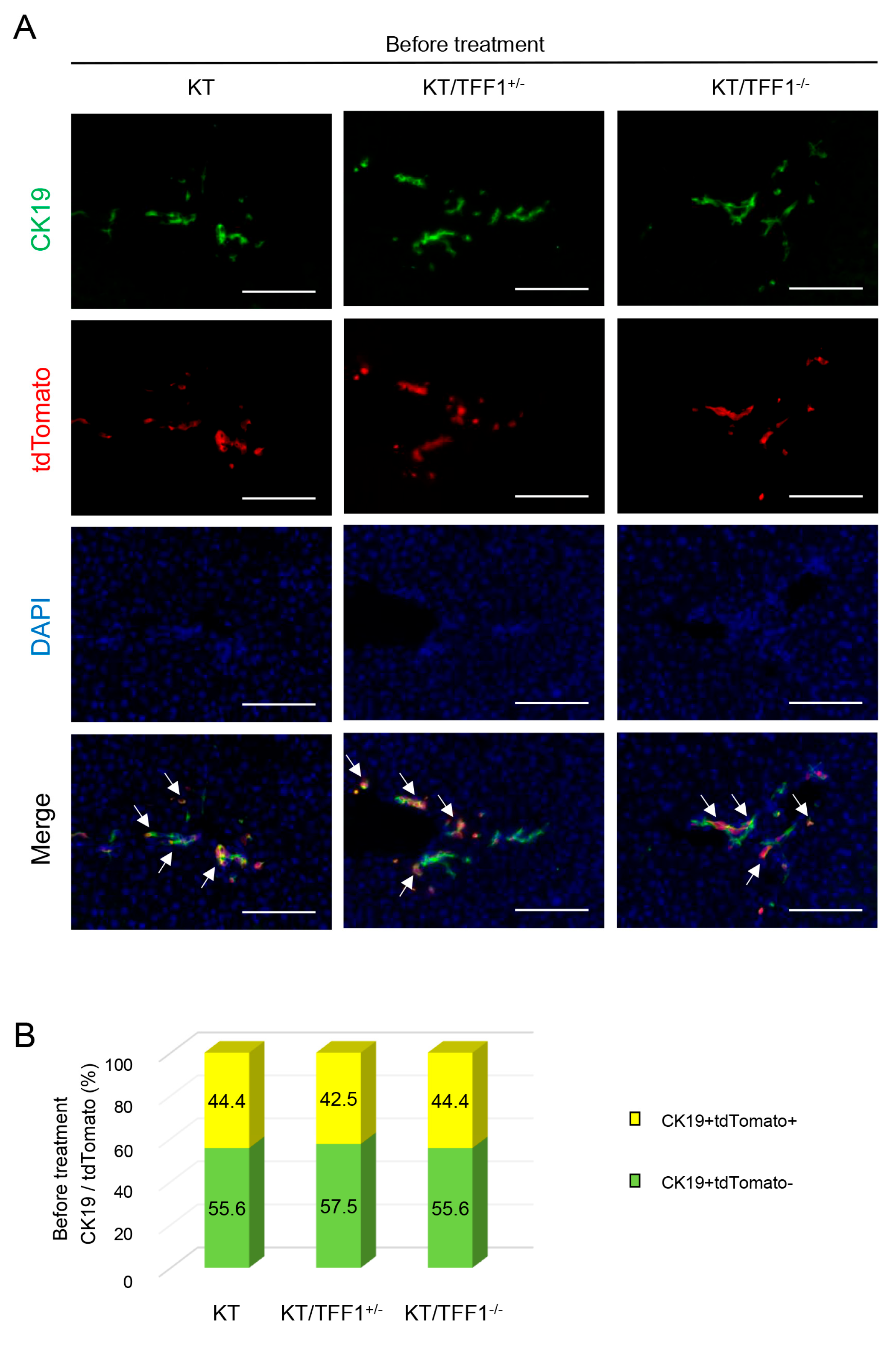
3.2. The Labeling Efficacy Was Equivalent Among the Mice
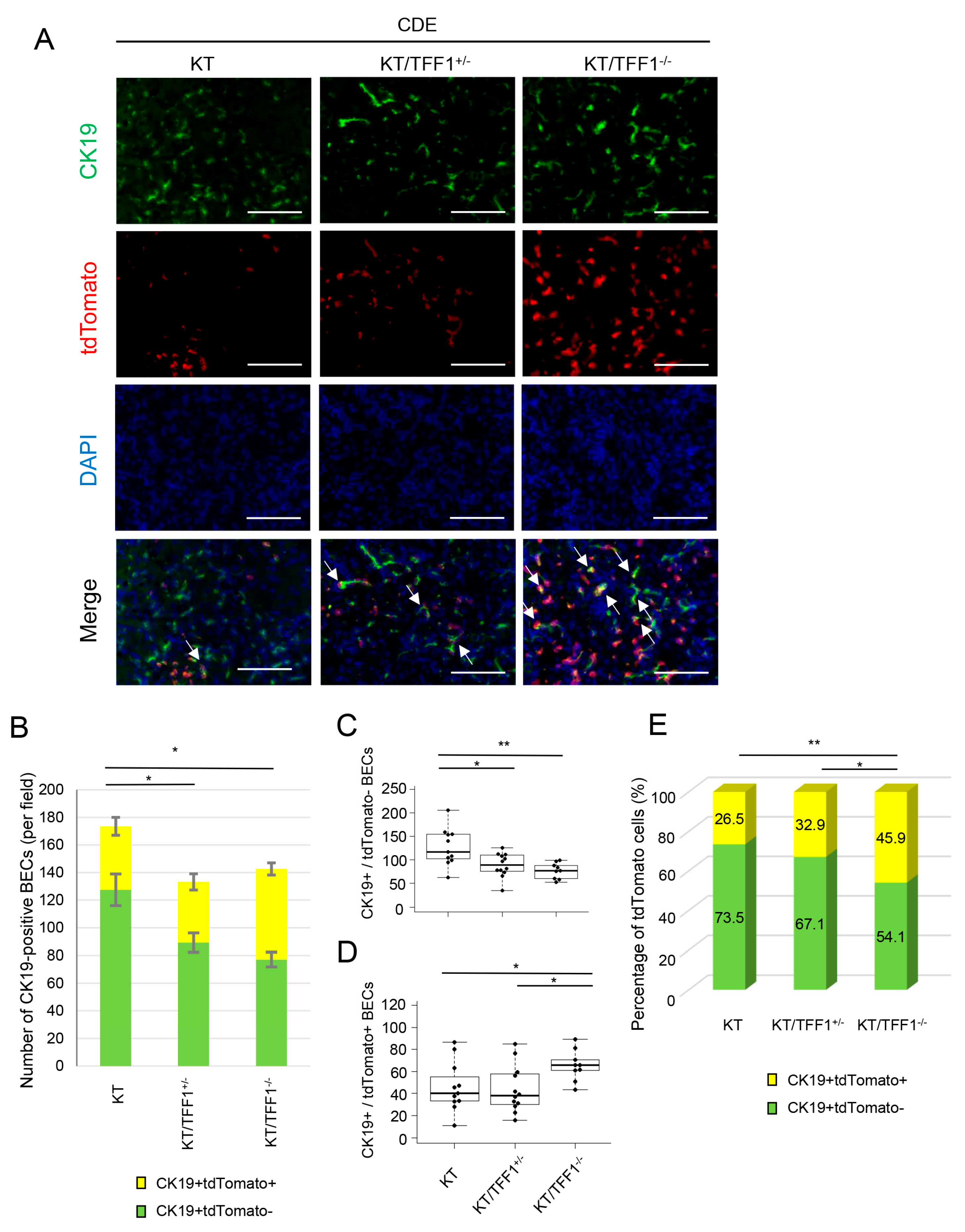
3.3. The Proportion of TdTomato-Positive BECs After CDE Varied Depending on TFF1 Status
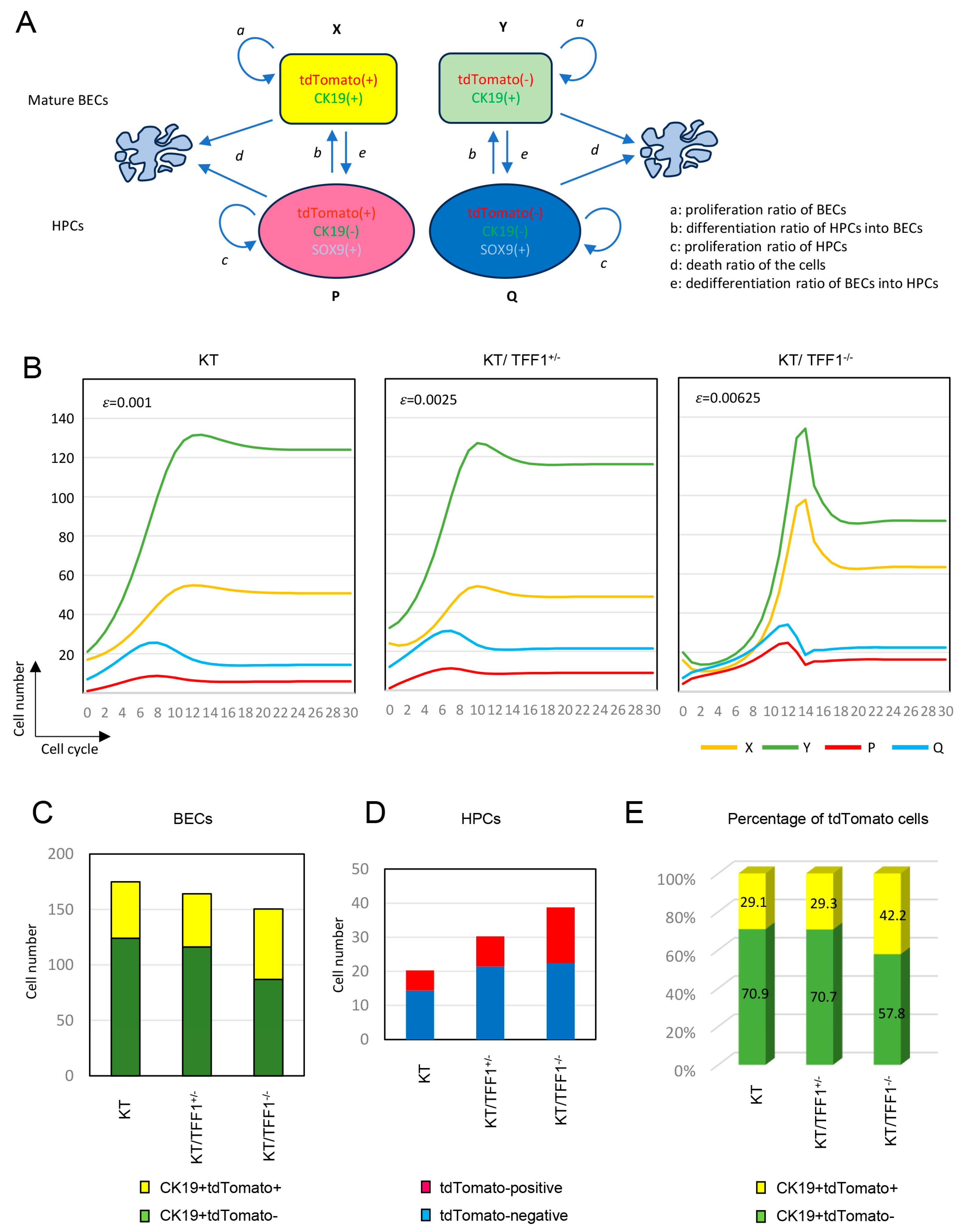
3.4. Model of the Differentiation of HPCs and Biliary Regeneration
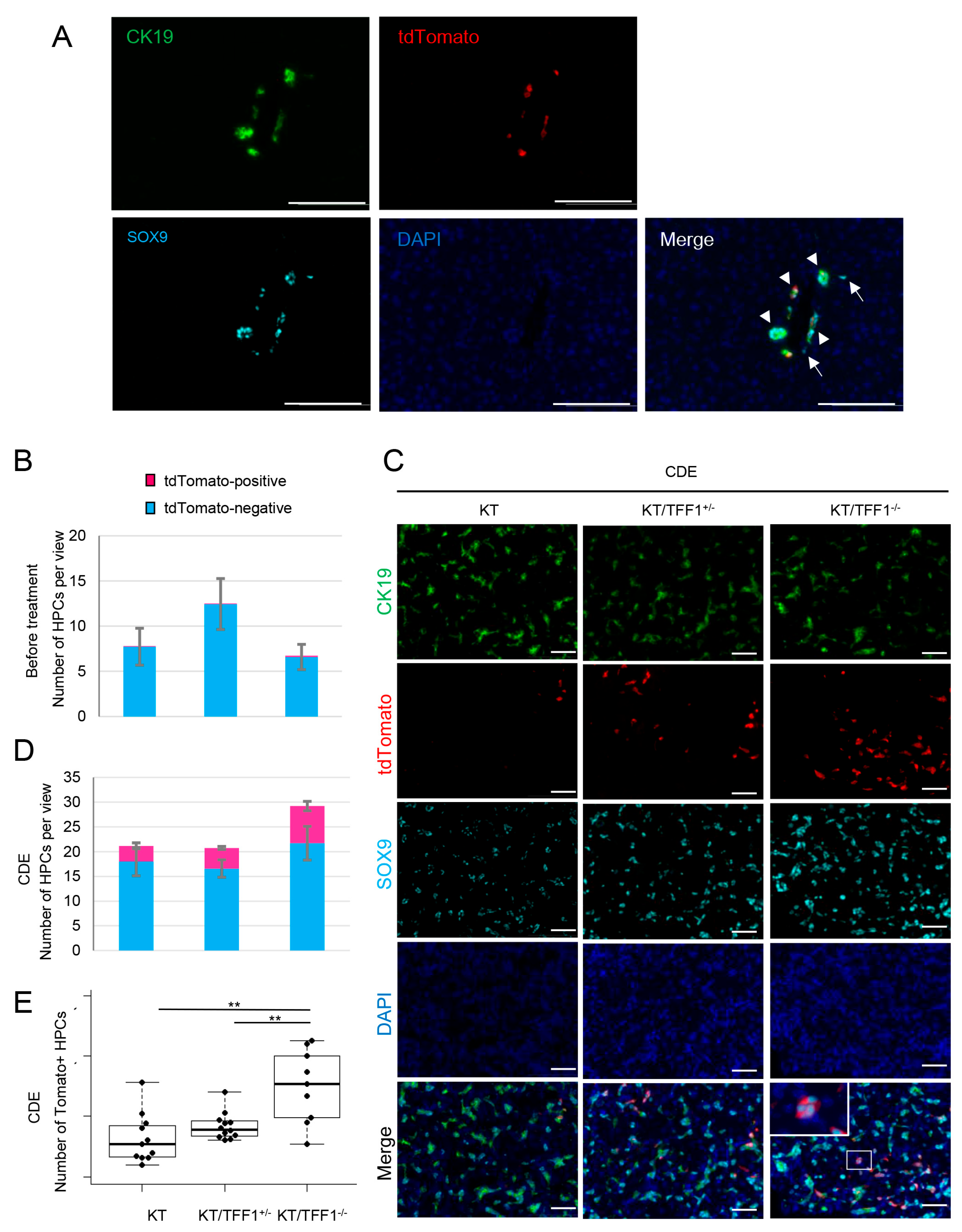
3.5. TdTomato-Labeled BECs Dedifferentiate into HPCs Frequently in KT/TFF1−/− Mice
3.6. Mathematical Model of HPC-Dependent Regeneration of BECs
3.7. dHPC-Dependent Regeneration of BECs in the Mathematical Model
3.8. Many dHPC-Derived Hepatocytes Were Found in TFF1-Deficient Mice
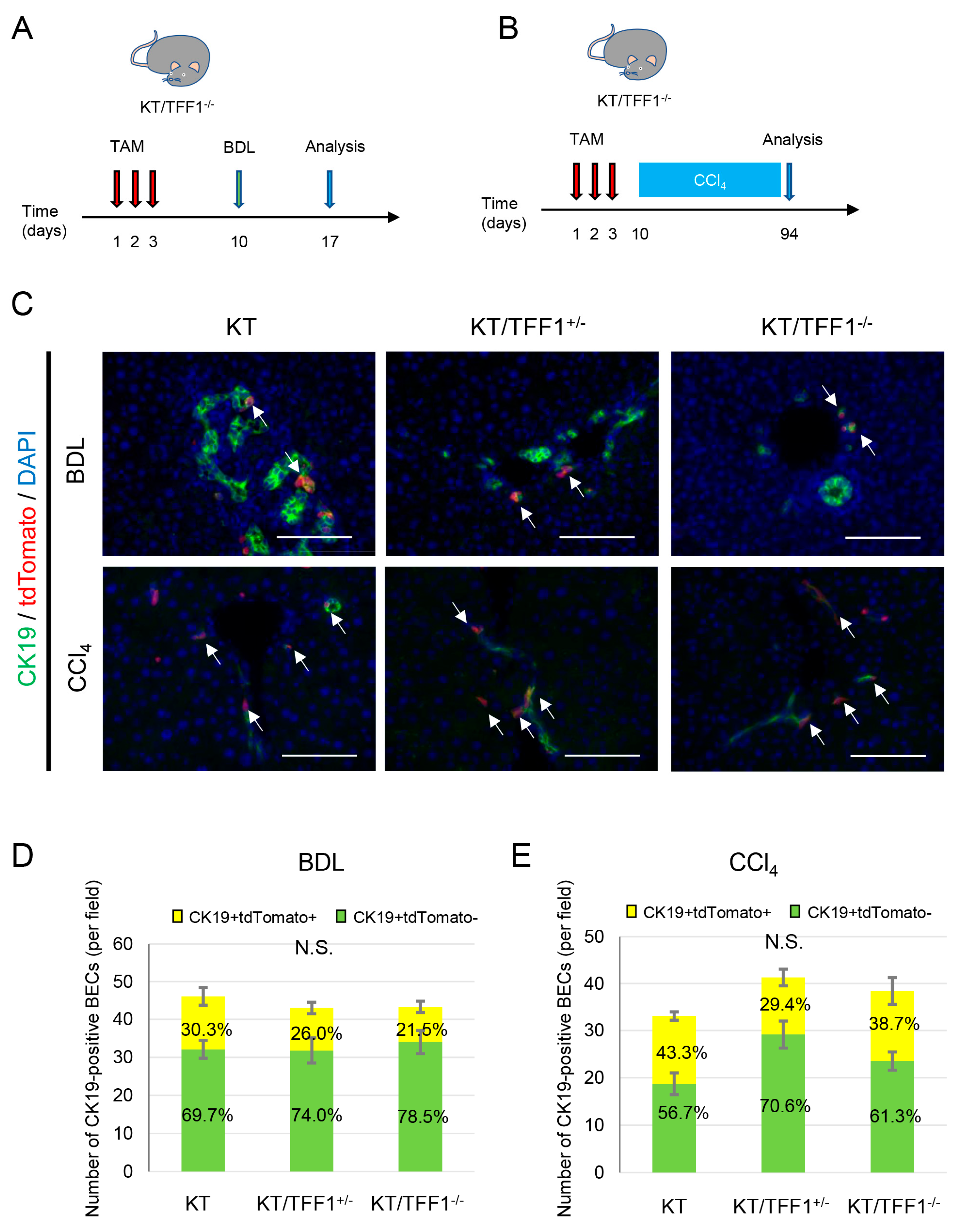
3.9. Different Mechanisms of BEC Regeneration in the Mouse Model of BDL and CCl4
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
References
- Campana, L.; Esser, H.; Huch, M.; Forbes, S. Liver regeneration and inflammation: From fundamental science to clinical applications. Nat. Rev. Mol. Cell Biol. 2021, 22, 608–624. [Google Scholar] [CrossRef] [PubMed]
- Gu, X.; Huang, D.; Ci, L.; Shi, J.; Zhang, M.; Yang, H.; Wang, Z.; Sheng, Z.; Sun, R.; Fei, J. Fate tracing of hepatocytes in mouse liver. Sci. Rep. 2017, 7, 16108. [Google Scholar] [CrossRef]
- Gadd, V.L.; Aleksieva, N.; Forbes, S.J. Epithelial Plasticity during Liver Injury and Regeneration. Cell Stem Cell 2020, 27, 557–573. [Google Scholar] [CrossRef]
- Yanger, K.; Knigin, D.; Zong, Y.; Maggs, L.; Gu, G.; Akiyama, H.; Pikarsky, E.; Stanger, B.Z. Adult hepatocytes are generated by self-duplication rather than stem cell differentiation. Cell Stem Cell 2014, 15, 340–349. [Google Scholar] [CrossRef]
- Wei, Y.; Wang, Y.G.; Jia, Y.; Li, L.; Yoon, J.; Zhang, S.; Wang, Z.; Zhang, Y.; Zhu, M.; Sharma, T.; et al. Liver homeostasis is maintained by midlobular zone 2 hepatocytes. Science 2021, 371, eabb1625. [Google Scholar] [CrossRef]
- Deng, X.; Zhang, X.; Li, W.; Feng, R.X.; Li, L.; Yi, G.R.; Zhang, X.N.; Yin, C.; Yu, H.-Y.; Zhang, J.-P.; et al. Chronic Liver Injury Induces Conversion of Biliary Epithelial Cells into Hepatocytes. Cell Stem Cell 2018, 23, 114–122.e3. [Google Scholar] [CrossRef]
- Russell, J.O.; Lu, W.Y.; Okabe, H.; Abrams, M.; Oertel, M.; Poddar, M.; Singh, S.; Forbes, S.J.; Monga, S.P. Hepatocyte-Specific beta-Catenin Deletion During Severe Liver Injury Provokes Cholangiocytes to Differentiate into Hepatocytes. Hepatology 2019, 69, 742–759. [Google Scholar] [CrossRef]
- Oderberg, I.M.; Goessling, W. Biliary epithelial cells are facultative liver stem cells during liver regeneration in adult zebrafish. JCI Insight 2023, 8, e163929. [Google Scholar] [CrossRef]
- Manco, R.; Clerbaux, L.A.; Verhulst, S.; Bou Nader, M.; Sempoux, C.; Ambroise, J.; Bearzatto, B.; Gala, J.L.; Horsmans, Y.; van Grunsven, L.; et al. Reactive cholangiocytes differentiate into proliferative hepatocytes with efficient DNA repair in mice with chronic liver injury. J. Hepatol. 2019, 70, 1180–1191. [Google Scholar] [CrossRef] [PubMed]
- Kitade, M.; Kaji, K.; Yoshiji, H. Relationship between hepatic progenitor cell-mediated liver regeneration and non-parenchymal cells. Hepatol. Res. 2016, 46, 1187–1193. [Google Scholar] [CrossRef] [PubMed]
- Sato, K.; Marzioni, M.; Meng, F.; Francis, H.; Glaser, S.; Alpini, G. Ductular Reaction in Liver Diseases: Pathological Mechanisms and Translational Significances. Hepatology 2019, 69, 420–430. [Google Scholar] [CrossRef] [PubMed]
- Mavila, N.; Siraganahalli Eshwaraiah, M.; Kennedy, J. Ductular Reactions in Liver Injury, Regeneration, and Disease Progression-An Overview. Cells 2024, 13, 579. [Google Scholar] [CrossRef]
- Li, W.; Li, L.; Hui, L. Cell Plasticity in Liver Regeneration. Trends Cell Biol. 2020, 30, 329–338. [Google Scholar] [CrossRef]
- Kjellev, S. The trefoil factor family—Small peptides with multiple functionalities. Cell Mol. Life Sci. 2009, 66, 1350–1369. [Google Scholar] [CrossRef]
- Tomita, H.; Takaishi, S.; Menheniott, T.R.; Yang, X.; Shibata, W.; Jin, G.; Betz, K.S.; Kawakami, K.; Minamoto, T.; Tomasetto, C.; et al. Inhibition of gastric carcinogenesis by the hormone gastrin is mediated by suppression of TFF1 epigenetic silencing. Gastroenterology 2011, 140, 879–891. [Google Scholar] [CrossRef] [PubMed]
- Soutto, M.; Belkhiri, A.; Piazuelo, M.B.; Schneider, B.G.; Peng, D.; Jiang, A.; Washington, M.K.; Kokoye, Y.; Crowe, S.E.; Zaika, A.; et al. Loss of TFF1 is associated with activation of NF-kappaB-mediated inflammation and gastric neoplasia in mice and humans. J. Clin. Investig. 2011, 121, 1753–1767. [Google Scholar] [CrossRef]
- Hasebe, K.; Yamazaki, K.; Yamaguchi, J.; Kokuryo, T.; Yokoyama, Y.; Miyata, K.; Fukaya, M.; Nagino, M.; Ebata, T. Trefoil factor 1 inhibits the development of esophageal adenocarcinoma from Barrett’s epithelium. Lab. Investig. 2022, 102, 885–895. [Google Scholar] [CrossRef]
- Hinz, M.; Schwegler, H.; Chwieralski, C.E.; Laube, G.; Linke, R.; Pohle, W.; Hoffmann, W. Trefoil factor family (TFF) expression in the mouse brain and pituitary: Changes in the developing cerebellum. Peptides 2004, 25, 827–832. [Google Scholar] [CrossRef]
- Silva, E.D.; Ulrich, M.; Döring, G.; Botzenhart, K.; Gött, P. Trefoil factor family domain peptides in the human respiratory tract. J. Pathol. 2000, 190, 133–142. [Google Scholar] [CrossRef]
- Rossi, H.L.; Ortiz-Carpena, J.F.; Tucker, D.; Vaughan, A.E.; Mangalmurti, N.S.; Cohen, N.A.; Herbert, D.R. Trefoil Factor Family: A Troika for Lung Repair and Regeneration. Am. J. Respir. Cell Mol. Biol. 2022, 66, 252–259. [Google Scholar] [CrossRef]
- Sunagawa, M.; Yamaguchi, J.; Kokuryo, T.; Ebata, T.; Yokoyama, Y.; Sugawara, G.; Nagino, M. Trefoil factor family 1 expression in the invasion front is a poor prognostic factor associated with lymph node metastasis in pancreatic cancer. Pancreatology 2017, 17, 782–787. [Google Scholar] [CrossRef]
- Hayashi, Y.; Yamaguchi, J.; Kokuryo, T.; Ebata, T.; Yokoyama, Y.; Nagino, M. Loss of trefoil factor 1 inhibits biliary regeneration but accelerates the hepatic differentiation of progenitor cells in mice. Biochem. Biophys. Res. Commun. 2018, 506, 12–19. [Google Scholar] [CrossRef]
- Kornprat, P.; Rehak, P.; Lemmerer, M.; Gogg-Kamerer, M.; Langner, C. Analysis of trefoil factor family protein 1 (TFF1, pS2) expression in chronic cholecystitis and gallbladder carcinoma. Virchows Arch. 2005, 446, 505–510. [Google Scholar] [CrossRef]
- Yamaguchi, J.; Yokoyama, Y.; Kokuryo, T.; Ebata, T.; Enomoto, A.; Nagino, M. Trefoil factor 1 inhibits epithelial-mesenchymal transition of pancreatic intraepithelial neoplasm. J. Clin. Investig. 2018, 128, 3619–3629. [Google Scholar] [CrossRef] [PubMed]
- Ochiai, Y.; Yamaguchi, J.; Kokuryo, T.; Yokoyama, Y.; Ebata, T.; Nagino, M. Trefoil Factor Family 1 Inhibits the Development of Hepatocellular Carcinoma by Regulating beta-Catenin Activation. Hepatology 2020, 72, 503–517. [Google Scholar] [CrossRef]
- Raven, A.; Lu, W.Y.; Man, T.Y.; Ferreira-Gonzalez, S.; O’Duibhir, E.; Dwyer, B.J.; Thomson, J.P.; Meehan, R.R.; Bogorad, R.; Koteliansky, V.; et al. Cholangiocytes act as facultative liver stem cells during impaired hepatocyte regeneration. Nature 2017, 547, 350–354. [Google Scholar] [CrossRef]
- So, J.; Kim, A.; Lee, S.H.; Shin, D. Liver progenitor cell-driven liver regeneration. Exp. Mol. Med. 2020, 52, 1230–1238. [Google Scholar] [CrossRef] [PubMed]
- Pu, W.; Zhu, H.; Zhang, M.; Pikiolek, M.; Ercan, C.; Li, J.; Huang, X.; Han, X.; Zhang, Z.; Lv, Z.; et al. Bipotent transitional liver progenitor cells contribute to liver regeneration. Nat. Genet. 2023, 55, 651–664. [Google Scholar] [CrossRef]
- Akhurst, B.; Croager, E.J.; Farley-Roche, C.A.; Ong, J.K.; Dumble, M.L.; Knight, B.; Yeoh, G.C. A modified choline-deficient, ethionine-supplemented diet protocol effectively induces oval cells in mouse liver. Hepatology 2001, 34, 519–522. [Google Scholar] [CrossRef]
- Zhu, C.; Tabas, I.; Schwabe, R.F.; Pajvani, U.B. Maladaptive regeneration—The reawakening of developmental pathways in NASH and fibrosis. Nat. Rev. Gastroenterol. Hepatol. 2021, 18, 131–142. [Google Scholar] [CrossRef]
- Yamaguchi, J.; Kokuryo, T.; Yokoyama, Y.; Oishi, S.; Sunagawa, M.; Mizuno, T.; Onoe, S.; Watanabe, N.; Ogura, A.; Ebata, T. Trefoil factor 1 suppresses stemness and enhances chemosensitivity of pancreatic cancer. Cancer Med. 2024, 13, e7395. [Google Scholar] [CrossRef]
- MacParland, S.A.; Liu, J.C.; Ma, X.Z.; Innes, B.T.; Bartczak, A.M.; Gage, B.K.; Manuel, J.; Khuu, N.; Echeverri, J.; Linares, I.; et al. Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat. Commun. 2018, 9, 4383. [Google Scholar] [CrossRef]
- Segal, J.M.; Kent, D.; Wesche, D.J.; Ng, S.S.; Serra, M.; Oules, B.; Kar, G.; Emerton, G.; Blackford, S.J.I.; Darmanis, S.; et al. Single cell analysis of human foetal liver captures the transcriptional profile of hepatobiliary hybrid progenitors. Nat. Commun. 2019, 10, 3350. [Google Scholar] [CrossRef]
- Gribben, C.; Galanakis, V.; Calderwood, A.; Williams, E.C.; Chazarra-Gil, R.; Larraz, M.; Frau, C.; Puengel, T.; Guillot, A.; Rouhani, F.J.; et al. Acquisition of epithelial plasticity in human chronic liver disease. Nature 2024, 630, 166–173. [Google Scholar] [CrossRef]


| Antigen | Company | Clone | Host | Concentration |
|---|---|---|---|---|
| CK19 | Developmental Studies Hybridoma Bank (Iowa, IA, USA) | TROMA-III | rat | 1:100 |
| RFP | ROCKLAND (Limerick, PA, USA) | 600-401-379 | rabbit | 1:1000 |
| SOX9 | EMD Millipore (Burlington, MA, USA) | AB5535 | rabbit | 1:500 |
| HNF4a | Santa Cruz Biotechnology (Dallas, TX, USA) | sc-6556 | goat | 1:50 |
| AFP | R and D Systems (Minneapolis, MN, USA) | AF5369 | goat | 1:100 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yamamoto, T.; Yamaguchi, J.; Kokuryo, T.; Yokoyama, Y.; Mizuno, T.; Onoe, S.; Sunagawa, M.; Baba, T.; Ebata, T. Dedifferentiation-Dependent Regeneration of the Biliary Ductal Epithelium in Response to Hepatic Injury in TFF1-Deficient Mice. Cells 2025, 14, 1323. https://doi.org/10.3390/cells14171323
Yamamoto T, Yamaguchi J, Kokuryo T, Yokoyama Y, Mizuno T, Onoe S, Sunagawa M, Baba T, Ebata T. Dedifferentiation-Dependent Regeneration of the Biliary Ductal Epithelium in Response to Hepatic Injury in TFF1-Deficient Mice. Cells. 2025; 14(17):1323. https://doi.org/10.3390/cells14171323
Chicago/Turabian StyleYamamoto, Taisuke, Junpei Yamaguchi, Toshio Kokuryo, Yukihiro Yokoyama, Takashi Mizuno, Shunsuke Onoe, Masaki Sunagawa, Taisuke Baba, and Tomoki Ebata. 2025. "Dedifferentiation-Dependent Regeneration of the Biliary Ductal Epithelium in Response to Hepatic Injury in TFF1-Deficient Mice" Cells 14, no. 17: 1323. https://doi.org/10.3390/cells14171323
APA StyleYamamoto, T., Yamaguchi, J., Kokuryo, T., Yokoyama, Y., Mizuno, T., Onoe, S., Sunagawa, M., Baba, T., & Ebata, T. (2025). Dedifferentiation-Dependent Regeneration of the Biliary Ductal Epithelium in Response to Hepatic Injury in TFF1-Deficient Mice. Cells, 14(17), 1323. https://doi.org/10.3390/cells14171323






