Circular RNAs as Targets for Developing Anticancer Therapeutics
Abstract
1. Biogenesis of CircRNAs
2. Regulation of Gene Expression by CircRNAs
3. CircRNAs and RNA-Binding Proteins
4. Roles of CircRNAs in Chemotherapy Resistance
5. Roles of CircRNAs in Immune Checkpoint Inhibitor Resistance
6. CircRNAs May Regulate the Responses to ICIs via Their Effects on Glycolysis
7. CircRNAs as Diagnostic and Prognostic Markers
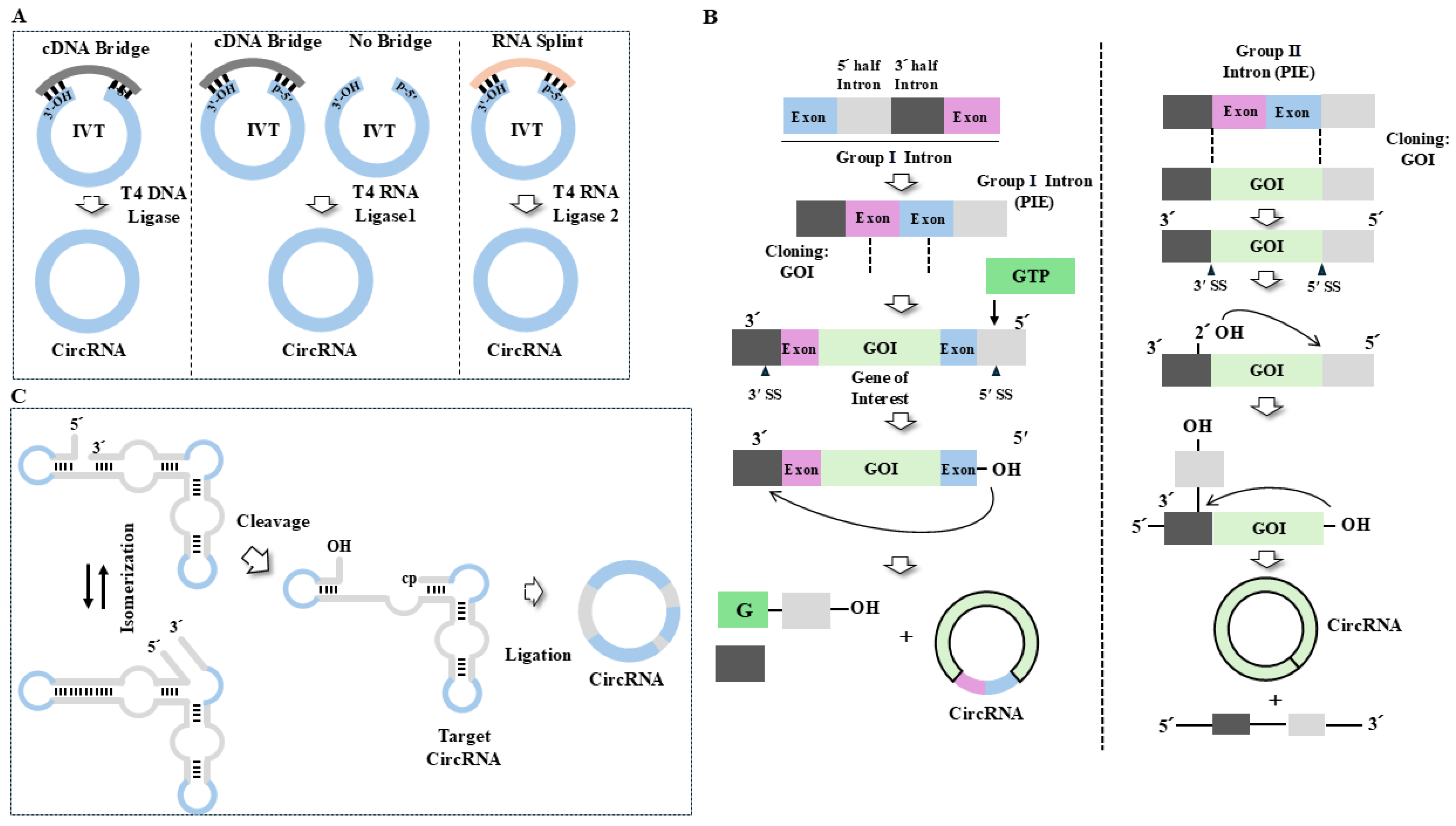
8. Synthesis of CircRNAs
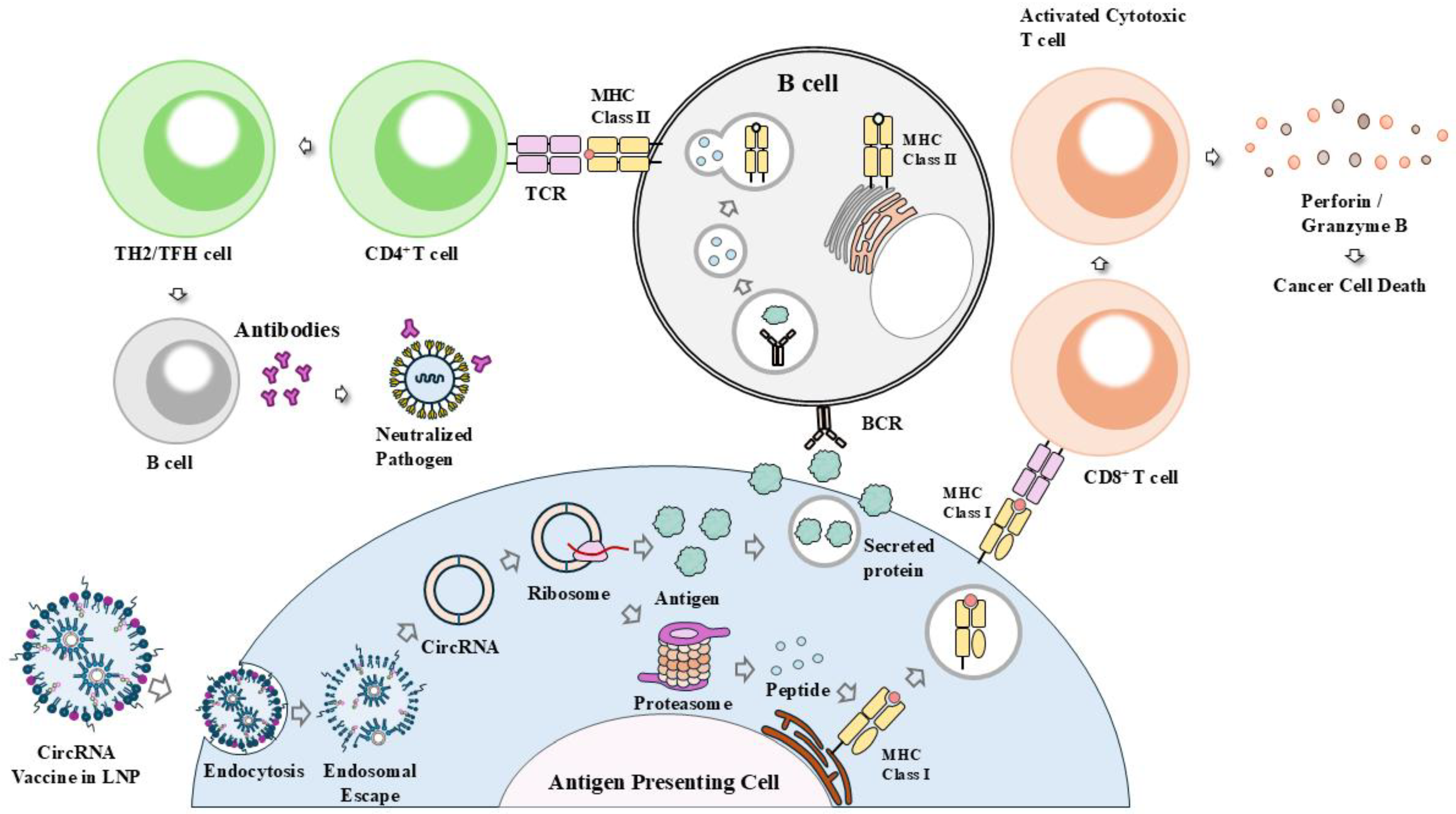
9. Roles of CircRNA Vaccines in Anticancer Immunity
10. Clinical Trials of CircRNA Vaccines and CircRNAs
11. Discussion and Perspectives
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Zhang, H.; Shen, Y.; Li, Z.; Ruan, Y.; Li, T.; Xiao, B.; Sun, W. The biogenesis and biological functions of circular RNAs and their molecular diagnostic values in cancers. J. Clin. Lab. Anal. 2020, 34, e23049. [Google Scholar] [CrossRef] [PubMed]
- Sheng, J.; Zhang, X.; Liang, W.; Lyu, J.; Zhang, B.; Min, J.; Xu, A.; Xu, X.; Li, J.W.; Li, J.L.; et al. The circular RNA circbabo(5,6,7,8S) regulates lipid metabolism and neuronal integrity via TGF-beta/ROS/JNK/SREBP signaling axis in Drosophila. BMC Biol. 2025, 23, 69. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Wang, P.; Wang, S.; Liao, W.; Ouyang, M.; Lin, S.; Lin, R.; Sarris, P.F.; Michalopoulou, V.; Feng, X.; et al. The circular RNA circANK suppresses rice resistance to bacterial blight by inhibiting microRNA398b-mediated defense. Plant Cell 2025, 37, koaf082. [Google Scholar] [CrossRef] [PubMed]
- Palcau, A.C.; Pulito, C.; De Pascale, V.; Casadei, L.; Valerio, M.; Sacconi, A.; Canu, V.; Rutigliano, D.; Donzelli, S.; Sardo, F.L.; et al. CircPVT1 weakens miR-33a-5p unleashing the c-MYC/GLS1 metabolic axis in breast cancer. J. Exp. Clin. Cancer Res. 2025, 44, 100. [Google Scholar] [CrossRef]
- Knupp, D.; Jorgensen, B.G.; Alshareef, H.Z.; Bhat, J.M.; Grubbs, J.J.; Miura, P.; van der Linden, A.M. Loss of circRNAs from the crh-1 gene extends the mean lifespan in Caenorhabditis elegans. Aging Cell 2022, 21, e13560. [Google Scholar] [CrossRef]
- Choi, S.W.; Nam, J.W. Optimal design of synthetic circular RNAs. Exp. Mol. Med. 2024, 56, 1281–1292. [Google Scholar] [CrossRef]
- Misir, S.; Wu, N.; Yang, B.B. Specific expression and functions of circular RNAs. Cell Death Differ. 2022, 29, 481–491. [Google Scholar] [CrossRef]
- Sahoo, B.; Gupta, M.K. Prediction of circRNAs in Mammalian Spermatozoa from RNA Sequencing Data. Methods Mol. Biol. 2025, 2897, 523–529. [Google Scholar]
- Kristensen, L.S.; Jakobsen, T.; Hager, H.; Kjems, J. The emerging roles of circRNAs in cancer and oncology. Nat. Rev. Clin. Oncol. 2022, 19, 188–206. [Google Scholar] [CrossRef]
- Liu, X.; Zhang, Y.; Zhou, S.; Dain, L.; Mei, L.; Zhu, G. Circular RNA: An emerging frontier in RNA therapeutic targets, RNA therapeutics, and mRNA vaccines. J. Control Release 2022, 348, 84–94. [Google Scholar] [CrossRef]
- 1Sur, S.; Pal, J.K.; Shekhar, S.; Bafna, P.; Bhattacharyya, R. Emerging role and clinical applications of circular RNAs in human diseases. Funct. Integr. Genom. 2025, 25, 77. [Google Scholar] [CrossRef]
- Li, L.; Wei, C.; Xie, Y.; Su, Y.; Liu, C.; Qiu, G.; Liu, W.; Liang, Y.; Zhao, X.; Huang, D.; et al. Expanded insights into the mechanisms of RNA-binding protein regulation of circRNA generation and function in cancer biology and therapy. Genes Dis. 2024, 12, 101383. [Google Scholar] [CrossRef] [PubMed]
- Srinivasan, A.; Mroczko-Młotek, E.; Wojciechowska, M. Circular RNA Formation and Degradation Are Not Directed by Universal Pathways. Int. J. Mol. Sci. 2025, 26, 726. [Google Scholar] [CrossRef] [PubMed]
- Chang, S.; Wang, Y.; Wang, X.; Liu, H.; Zhang, T.; Zheng, Y.; Wang, X.; Shan, G.; Chen, L. HNRNPD regulates the biogenesis of circRNAs and the ratio of mRNAs to circRNAs for a set of genes. RNA Biol. 2024, 21, 834–848. [Google Scholar] [CrossRef]
- Tang, L.; Ji, Y.; Ni, C.; Xu, Z.; Shen, Y.; Lu, H.; Zhang, C.; Yang, S. EIF4A3-Mediated Biogenesis of CircFADS1 Promotes the Progression of Hepatocellular Carcinoma via Wnt/beta-Catenin Pathway. Adv. Sci. 2025, 12, e2411869. [Google Scholar] [CrossRef]
- Li, X.; Liu, S.; Zhang, L.; Issaian, A.; Hill, R.C.; Espinosa, S.; Shi, S.; Cui, Y.; Kappel, K.; Das, R.; et al. A unified mechanism for intron and exon definition and back-splicing. Nature 2019, 573, 375–380. [Google Scholar] [CrossRef]
- Yu, H.; Yu, Y.; Xia, Y. circ2LO: Identification of CircRNA Based on the LucaOne Large Model. Genes 2025, 16, 413. [Google Scholar] [CrossRef]
- Du, X.; Liu, Y.; He, X.; Tao, L.; Fang, M.; Chu, M. Identification and expression profile analysis of circRNAs associated with goat uterus with different fecundity during estrous cycle. BMC Genom. 2025, 26, 349. [Google Scholar] [CrossRef]
- Jiang, B.; Li, Y.; Shi, J.; Chalasa, D.D.; Zhang, L.; Wu, S.; Xu, T. Identification and Network Construction of mRNAs, miRNAs, lncRNAs, and circRNAs in Sweetpotato (Ipomoea batatas L.) Adventitious Roots Under Salt Stress via Whole-Transcriptome RNA Sequencing. Int. J. Mol. Sci. 2025, 26, 1660. [Google Scholar] [CrossRef]
- Gu, H.; Yu, W.; Feng, P.; Zeng, C.; Cao, Q.; Chen, F.; Wang, Z.; Shen, H.; Wu, Y.; Wang, S. Circular RNA circSTX12 regulates osteo-adipogenic balance and proliferation of BMSCs in senile osteoporosis. Cell Mol. Life Sci. 2025, 82, 149. [Google Scholar] [CrossRef]
- Li, X.; Zhang, J.L.; Lei, Y.N.; Liu, X.Q.; Xue, W.; Zhang, Y.; Nan, F.; Gao, X.; Zhang, J.; Wei, J.; et al. Linking circular intronic RNA degradation and function in transcription by RNase H1. Sci. China Life Sci. 2021, 64, 1795–1809. [Google Scholar] [CrossRef]
- Sztachera, M.; Wendlandt-Stanek, W.; Serwa, R.A.; Stanaszek, L.; Smuszkiewicz, M.; Wronka, D.; Piwecka, M. Interrogation of RNA-bound proteome with XRNAX illuminates molecular alterations in the mouse brain affected with dysmyelination. Cell Rep. 2025, 44, 115095. [Google Scholar] [CrossRef]
- Chen, R.X.; Xu, S.D.; Deng, M.H.; Hao, S.H.; Chen, J.W.; Ma, X.D.; Zhuang, W.T.; Cao, J.H.; Lv, Y.R.; Lin, J.L.; et al. Mex-3 RNA binding family member A (MEX3A)/circMPP6 complex promotes colorectal cancer progression by inhibiting autophagy. Signal Transduct. Target Ther. 2024, 9, 80. [Google Scholar] [CrossRef]
- Tang, S.; Cai, L.; Wang, Z.; Pan, D.; Wang, Q.; Shen, Y.; Zhou, Y.; Chen, Q. Emerging roles of circular RNAs in the invasion and metastasis of head and neck cancer: Possible functions and mechanisms. Cancer Innov. 2023, 2, 463–487. [Google Scholar] [CrossRef] [PubMed]
- Neu, C.T.; Gutschner, T.; Haemmerle, M. Post-Transcriptional Expression Control in Platelet Biogenesis and Function. Int. J. Mol. Sci. 2020, 21, 7614. [Google Scholar] [CrossRef] [PubMed]
- Cao, C.; Wang, C.; Dai, Q.; Zou, Q.; Wang, T. CRBPSA: CircRNA-RBP interaction sites identification using sequence structural attention model. BMC Biol. 2024, 22, 260. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Li, C.; Jafari, H.; Yang, G.; Wang, Z.; Lei, C.; Dang, R. CircRNA profiling reveals circSMC1B that promotes bovine male germline stem cells proliferation and apoptosis via sponging let-7i. BMC Genom. 2025, 26, 398. [Google Scholar] [CrossRef]
- Bai, W.; Guo, Z.L.; Guo, J.H.; Li, F.; Bu, P.; Liu, J. Circular RNA contributes to gastric cancer by targeting Wnt family member 2B as a competing endogenous RNA. World J. Gastroenterol. 2025, 31, 99583. [Google Scholar] [CrossRef]
- Liu, R.; Song, Y.; Wang, Z.; Dai, L.; Bai, Q.; Li, Y.; Piao, H.; Wang, C.; Yan, G. circ-0001454 alleviates asthma airway inflammation and remodeling via sponging miR-770-5p and regulating cbl-b. Front. Cell Dev. Biol. 2025, 13, 1566223. [Google Scholar] [CrossRef]
- Bibi, A.; Madè, A.; Greco, S.; Garcia-Manteiga, J.M.; Tascini, A.S.; Tastsoglou, S.; Zaccagnini, G.; Leszek, P.; Gaetano, C.; Martelli, F. Circular PVT1 promotes cardiac fibroblast activation interacting with miR-30a-5p and miR-125b-5p. Cell Death Dis. 2025, 16, 325. [Google Scholar] [CrossRef]
- Lin, W.; Huang, L.; Ou, Z.; Xuan, Y.; Zhu, D.; Zhang, Q.; Xu, E. N6-methyladenosine-modified circ_0000517 promotes non-small cell lung cancer metastasis via miR-1233-3p/CDH6 axis. J. Mol. Histol. 2025, 56, 139. [Google Scholar] [CrossRef]
- Shi, Y.; Li, D.; Xu, Y.; Guo, Y.; Mao, J.; Lu, Y. Circ_RUSC2 Sequesters miR-661 and Elevates TUSC2 Expression to Suppress Colorectal Cancer Progression. Int. J. Mol. Sci. 2025, 26, 2937. [Google Scholar] [CrossRef]
- Gui, C.P.; Liao, B.; Luo, C.G.; Chen, Y.H.; Tan, L.; Tang, Y.M.; Li, J.Y.; Hou, Y.; Song, H.D.; Lin, H.S.; et al. circCHST15 is a novel prognostic biomarker that promotes clear cell renal cell carcinoma cell proliferation and metastasis through the miR-125a-5p/EIF4EBP1 axis. Mol. Cancer 2021, 20, 169. [Google Scholar] [CrossRef] [PubMed]
- Yi, J.; Du, J.; Chen, X.; Nie, R.C.; Hu, G.S.; Wang, L.; Zhang, Y.Y.; Chen, S.; Wen, X.S.; Luo, D.X.; et al. A circRNA-mRNA pairing mechanism regulates tumor growth and endocrine therapy resistance in ER-positive breast cancer. Proc. Natl. Acad. Sci. USA 2025, 122, e2420383122. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Zhao, W.; Wu, X.; Fang, T.; Chen, Z.; Chen, Z.; Su, W.; Zhao, X.; Hu, Y.; Xu, Y.; et al. circFOXK2 Stabilizes STMN1 mRNA via PABPC1 to Promote the Progression of NSCLC. Cancer Med. 2025, 14, e70729. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Li, X.; Xue, W.; Zhang, L.; Yang, L.Z.; Cao, S.M.; Lei, Y.N.; Liu, C.X.; Guo, S.K.; Shan, L.; et al. Screening for functional circular RNAs using the CRISPR-Cas13 system. Nat. Methods 2021, 18, 51–59. [Google Scholar] [CrossRef]
- Peng, M.; Zhang, S.; Wu, P.; Hou, X.; Wang, D.; Ge, J.; Qu, H.; Fan, C.; Zhou, Y.; Xiang, B.; et al. Circular RNA circCLASP2 promotes nasopharyngeal carcinoma progression through binding to DHX9 to enhance PCMT1 translation. Mol. Cancer 2025, 24, 67. [Google Scholar] [CrossRef]
- Li, G.; Zhou, X.; Liu, X.; Gong, L.; Li, W.; Shen, T.; Wu, Q.; Wang, X.; Wang, Z.; Cai, J.; et al. Epithelial splicing regulatory protein 1 promotes peritoneal dissemination of ovarian cancer by inducing the formation of circular RNAs modulating epithelial plasticity. Cell Signal 2025, 125, 111485. [Google Scholar] [CrossRef]
- Abdelmohsen, K.; Panda, A.C.; Munk, R.; Grammatikakis, I.; Dudekula, D.B.; De, S.; Kim, J.; Noh, J.H.; Kim, K.M.; Martindale, J.L.; et al. Identification of HuR target circular RNAs uncovers suppression of PABPN1 translation by CircPABPN1. RNA Biol. 2017, 14, 361–369. [Google Scholar] [CrossRef]
- Schreiner, S.; Didio, A.; Hung, L.H.; Bindereif, A. Design and application of circular RNAs with protein-sponge function. Nucleic Acids Res. 2020, 48, 12326–12335. [Google Scholar] [CrossRef]
- Margvelani, G.; Maquera, K.A.A.; Welden, J.R.; Rodgers, D.W.; Stamm, S. Translation of circular RNAs. Nucleic Acids Res. 2025, 53, gkae1167. [Google Scholar] [CrossRef]
- Zhao, Y.; Liu, H.; Wang, Q.; Li, Z.; Zhao, K.; Sun, H.; Zhang, Y.; Li, N.; Li, W. Circ_0038632 Acts as a Sponge of miR-4306 to Facilitate Breast Cancer Progression Through Regulating CXCR4 Expression. J. Biochem. Mol. Toxicol. 2025, 39, e70249. [Google Scholar] [CrossRef] [PubMed]
- 4Zhang, Y.; Yao, R.; Li, M.; Fang, C.; Feng, K.; Chen, X.; Wang, J.; Luo, R.; Shi, H.; Chen, X.; et al. CircTTC13 promotes sorafenib resistance in hepatocellular carcinoma through the inhibition of ferroptosis by targeting the miR-513a-5p/SLC7A11 axis. Mol. Cancer 2025, 24, 32. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Lan, S.; Duan, Z. circ-TTC17 Promotes Esophagus Squamous Cell Carcinoma Cell Growth, Metastasis, and Inhibits Autophagy-Mediated Radiosensitivity Through miR-145-5p/SIRT1 Axis. Thorac. Cancer 2025, 16, e15494. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Fang, S.; Wu, X.; Fang, T.; Chen, Z.; Su, W.; Zhu, Y.; Zhao, X.; Zhou, C. circZNF707 promoted glycolysis and tumor progression through miR-668-3p-PFKM axis in NSCLC. Eur. J. Med. Res. 2025, 30, 141. [Google Scholar] [CrossRef]
- Sun, J.; Zhang, H.; Wei, W.; Xiao, X.; Huang, C.; Wang, L.; Zhong, H.; Jiang, Y.; Zheng, F.; Yang, H.; et al. Regulation of CD8+ T cells infiltration and immunotherapy by circMGA/HNRNPL complex in bladder cancer. Oncogene 2023, 42, 1247–1262. [Google Scholar] [CrossRef]
- Fang, Z.; Wu, Z.; Yu, C.; Xie, Q.; Zeng, L.; Chen, R. EIF4E-mediated biogenesis of circPHF14 promotes the growth and metastasis of pancreatic ductal adenocarcinoma via Wnt/β-catenin pathway. Mol. Cancer 2025, 24, 56. [Google Scholar] [CrossRef]
- Zhang, N.; Wang, X.; Li, Y.; Lu, Y.; Sheng, C.; Sun, Y.; Ma, N.; Jiao, Y. Mechanisms and therapeutic implications of gene expression regulation by circRNA-protein interactions in cancer. Commun. Biol. 2025, 8, 77. [Google Scholar] [CrossRef]
- Liu, L.; Lei, X.; Wang, Z.; Meng, J.; Song, B. TransRM: Weakly supervised learning of translation-enhancing N6-methyladenosine (m(6)A) in circular RNAs. Int. J. Biol. Macromol. 2025, 306, 141588. [Google Scholar] [CrossRef]
- Song, X.; Wei, Z.; Zhang, C.; Han, D.; Liu, J.; Song, Y.; Xie, X.; Shao, D.; Zhao, M.; Chao, F.; et al. CircAKT3 promotes prostate cancer proliferation and metastasis by enhancing the binding of RPS27A and RPL11. Mol. Cancer 2025, 24, 53. [Google Scholar] [CrossRef]
- Xu, H.; Zhao, Q.; Cai, D.; Chen, X.; Zhou, X.; Gao, Y.; Wu, J.; Yuan, S.; Li, D.; Zhang, R.; et al. o8G-modified circKIAA1797 promotes lung cancer development by inhibiting cuproptosis. J. Exp. Clin. Cancer Res. 2025, 44, 110. [Google Scholar] [CrossRef]
- Yu, M.; Wu, M.; Shen, T.; Qiu, Q.; Luo, A.; Li, J.; Li, X.; Cheng, X.; Lu, B.; Lu, W.; et al. CircMETTL6 Suppresses Ovarian Cancer Cell Growth and Metastasis Through Inhibition of GDF15 Transcription by Disrupting the NONO-POLR2A Complex. Adv. Sci 2025, 12, e2411717. [Google Scholar] [CrossRef] [PubMed]
- Fan, S.; Cui, Y.; Liu, Y.; Li, Y.; Huang, H.; Hu, Z. CircMYH9 promotes the mRNA stability of SPAG6 by recruiting EIF4A3 to facilitate the progression of breast cancer. Epigenetics 2025, 20, 2482382. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Tan, W.; Wang, M.; Wei, Y.; Xie, Z.; Wang, Q.; Zhang, Z.; Zhuang, H.; Ma, X.; Wang, B.; et al. circCCNY enhances lenvatinib sensitivity and suppresses immune evasion in hepatocellular carcinoma by serving as a scaffold for SMURF1 mediated HSP60 degradation. Cancer Lett. 2025, 612, 217470. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Song, X.; Feng, Y.; Qian, Y.; Chen, B.; Zhang, T.; Wang, H.; Chen, Y.; Yu, X.; Ding, H.; et al. The circRNA cEMSY Induces Immunogenic Cell Death and Boosts Immunotherapy Efficacy in Lung Adenocarcinoma. Cancer Res. 2025, 85, 497–514. [Google Scholar] [CrossRef]
- Conn, V.M.; Chinnaiyan, A.M.; Conn, S.J. Circular RNA in cancer. Nat. Rev. Cancer 2024, 24, 597–613. [Google Scholar] [CrossRef]
- Zhou, F.; Zhuo, J.; Xu, X.; Pan, D.; Cai, C.; Huang, J.; Zhao, X.; Mao, Q.; Jiang, X.; Sun, X.U.; et al. Circular RNA hsa_circ_0004650 Enhances 5-Fluorouracil Resistance in Gastric Cancer via Sponging miR-145-5p. Anticancer Res. 2024, 45, 173–187. [Google Scholar] [CrossRef]
- Guo, W.; Liu, H.; Zhong, M.; Qi, Q.; Li, Y. circ_0006528 promotes nonsmall cell lung cancer progression by sponging miR-892a and regulating NRAS expression. Anticancer Drugs 2025, 36, 261–270. [Google Scholar] [CrossRef]
- Hu, S.; Zhang, Q.; Sun, J.; Xue, J.; Wang, C. Circular RNA circ_0000376 promotes paclitaxel resistance and tumorigenesis of non-small cell lung cancer via positively modulating KPNA4 by sponging miR-1298-5p. Thorac. Cancer 2023, 14, 2116–2126. [Google Scholar] [CrossRef]
- Sheng, H.; Wang, X. Knockdown of circ-PIP5K1A overcomes resistance to cisplatin in ovarian cancer by miR-942-5p/NFIB axis. Anticancer Drugs 2023, 34, 214–226. [Google Scholar] [CrossRef]
- Cao, X.; Zhou, X.; Hou, F.; Huang, Y.E.; Yuan, M.; Long, M.; Chen, S.; Lei, W.; Zhu, J.; Chen, J.; et al. ncRNADrug: A database for validated and predicted ncRNAs associated with drug resistance and targeted by drugs. Nucleic Acids Res. 2024, 52, D1393–D1399. [Google Scholar] [CrossRef] [PubMed]
- Su, L.; Gou, J.; Zhou, C.; Li, J.; Wu, J.; Shen, L.; Jia, Y. Knockdown of circ_0076305 decreases the paclitaxel resistance of non-small cell lung cancer cells by regulating TMPRSS4 via miR-936. Toxicol. Res. 2024, 13, tfae102. [Google Scholar] [CrossRef] [PubMed]
- Zhao, T.; Li, K.; Zhang, Y.; Dong, Y.; Li, Y.; Pang, M.; Wei, Y.; Yao, B.; Zhu, Q. N6-methyladenosine-modified circQKI inhibits prostate cancer docetaxel-sensitivity via miR-188-3p/Beclin-1 pathway. Life Sci. 2025, 372, 123646. [Google Scholar] [CrossRef] [PubMed]
- Xuan, X.; Wang, Z.; Wang, Y. Circ_0058608 contributes to the progression and taxol resistance of non-small cell lung cancer by sponging miR-1299 to upregulate GBP1. Anticancer Drugs 2023, 34, 103–114. [Google Scholar] [CrossRef]
- Chen, J.J.; Shi, P.; Cui, Z.C.; Jiang, N.; Ma, J. CircRNA_0044556 affects the sensitivity of triple-negative breast cancer cells to paclitaxel by regulating miR-665. J. Chemother. 2025, 37, 238–246. [Google Scholar] [CrossRef]
- Sun, G.; Tian, J.; Xiao, Y.; Zeng, Y. Circular RNA circ_0005667 promotes cisplatin resistance of endometrial carcinoma cells by regulating IGF2BP1 through miR-145-5p. Anticancer Drugs 2023, 34, 816–826. [Google Scholar] [CrossRef]
- Xu, X.; Peng, Q.; Ren, Z.; Han, Y.; Jiang, X.; Wu, Z.; Tan, S.; Yang, W.; Oyang, L.; Luo, X.; et al. CircRNF13 enhances IGF2BP1 phase separation-mediated ITGB1 mRNA stabilization in an m6A-dependent manner to promote oral cancer cisplatin chemoresistance. Mol. Cancer 2025, 24, 36. [Google Scholar] [CrossRef]
- Li, X.; Deng, Y.; Yin, Z. CircKRT75 augments the cisplatin chemoresistance of nasopharyngeal carcinoma via targeting miR-659/CCAR2 axis. J. Mol. Histol. 2024, 56, 9. [Google Scholar] [CrossRef]
- Ding, M.; Zhao, J.; Li, X. Hsa_circ_0006006 is a potential biomarker for prognosis and cisplatin resistance in non-small cell lung cancer. Hereditas 2025, 162, 32. [Google Scholar] [CrossRef]
- Huang, M.; Sun, J.; Jiang, Q.; Zhao, X.; Huang, H.; Lei, M.; Jiang, S.; Yuan, F.; Liu, Z. CircKIAA0182-YBX1 Axis: A Key Driver of Lung Cancer Progression and Chemoresistance. Cancer Lett. 2025, 612, 217494. [Google Scholar] [CrossRef]
- He, R.; Hu, C.; Yuan, Y.; Li, T.; Tian, Q.; Huang, T.; Lin, Q.; Zheng, S.; Chen, C.; Fu, Z.; et al. Glycolysis reprogramming in CAFs promotes oxaliplatin resistance in pancreatic cancer through circABCC4 mediated PKM2 nuclear translocation. Cell Death Dis. 2025, 16, 126. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Wang, N.; Huang, Y.; He, S.; Bao, M.; Wen, C.; Wu, L. CircMYBL1 suppressed acquired resistance to osimertinib in non-small-cell lung cancer. Cancer Genet. 2024, 284–285, 34–42. [Google Scholar] [CrossRef] [PubMed]
- Liao, Q.; Deng, J.; Tong, J.; Gan, Y.; Hong, W.; Dong, H.; Cao, M.; Xiong, C.; Chen, Y.; Xie, B.; et al. p53 induces circFRMD4A to suppress cancer development through glycolytic reprogramming and cuproptosis. Mol. Cell 2025, 85, 132–149. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Tan, J.; Qin, F.; Zheng, Y.; Yang, X.; Liu, Z.; Cai, W.; Qin, X.; Liao, H. PKM2 modulates chemotherapy sensitivity by regulating autophagy and predicts the prognosis and immunity in pancancer. Sci. Rep. 2025, 15, 14626. [Google Scholar] [CrossRef]
- Yu, Y.; Liang, Y.; Xie, F.; Zhang, Z.; Zhang, P.; Zhao, X.; Zhang, Z.; Liang, Z.; Li, D.; Wang, L.; et al. Tumor-associated macrophage enhances PD-L1-mediated immune escape of bladder cancer through PKM2 dimer-STAT3 complex nuclear translocation. Cancer Lett. 2024, 593, 216964. [Google Scholar] [CrossRef]
- Oh, W.; Kim, A.M.J.; Dhawan, D.; Knapp, D.W.; Lim, S.O. Lactic acid inhibits the interaction between PD-L1 protein and PD-L1 antibody in the PD-1/PD-L1 blockade therapy-resistant tumor. Mol. Ther. 2025, 33, 723–733. [Google Scholar] [CrossRef]
- Lan, T.; Gao, F.; Cai, Y.; Lv, Y.; Zhu, J.; Liu, H.; Xie, S.; Wan, H.; He, H.; Xie, K.; et al. The protein circPETH-147aa regulates metabolic reprogramming in hepatocellular carcinoma cells to remodel immunosuppressive microenvironment. Nat. Commun. 2025, 16, 333. [Google Scholar] [CrossRef]
- Hong, W.; Xue, M.; Jiang, J.; Zhang, Y.; Gao, X. Circular RNA circ-CPA4/let-7 miRNA/PD-L1 axis regulates cell growth, stemness, drug resistance and immune evasion in non-small cell lung cancer (NSCLC). J. Exp. Clin. Cancer Res. 2020, 39, 149. [Google Scholar] [CrossRef]
- Yang, J.; Jia, Y.; Wang, B.; Yang, S.; Du, K.; Luo, Y.; Li, Y.; Zhu, B. Circular RNA CHST15 Sponges miR-155-5p and miR-194-5p to Promote the Immune Escape of Lung Cancer Cells Mediated by PD-L1. Front. Oncol. 2021, 11, 595609. [Google Scholar] [CrossRef]
- Geng, Z.; Zhang, G. circ_0001006 Promotes Immune Escape in Non-small Cell Lung Cancer by Regulating the miR-320a/PD-L1 Axis. Iran J. Immunol. 2025, 22, 58–69. [Google Scholar]
- Yu, H.; Zhang, Y.; Yang, R.; Xie, C.; Liao, Z.; Zhou, T. CircENTPD7 affects the immune escape of non-small cell lung cancer cells by modulating the IGF2BP2/PD-L1 axis. Oncol. Lett. 2024, 29, 112. [Google Scholar] [CrossRef]
- Zhuang, J.; Zhou, M.; Yu, H.; Zhou, R.; Bai, K.; Lv, J.; Li, K.; Cheng, Y.; Yang, H.; Yang, X.; et al. CircFAM64A(3) promoted bladder cancer proliferation and inhibited CD8+ T cell via sponging to miR-149-5p and activated IL-6/JAK/STAT pathway. Int. Immunopharmacol. 2025, 145, 113762. [Google Scholar] [CrossRef]
- Wang, Y.; Li, S.; Bo, X.; Li, Y.; Wang, C.; Nan, L.; Zhang, D.; Liu, H.; Zhang, J. CircRNome-wide characterisation reveals the promoting role of circAATF in anti-PD-L1 immunotherapy of gallbladder carcinoma. Clin. Transl. Med. 2024, 14, e70060. [Google Scholar] [CrossRef]
- Guan, G.F.; Fu, Z.M.; Zhang, J.; Guo, Y.Y.; Guo, F.; Wan, Y.N.; Bai, J.; Zhao, Y. Interferon Gamma Receptor 2 Collaborates With Circular RNA/MicroRNA to Modulate Programmed Cell Death-Ligand 1 Levels in Nasopharyngeal Carcinoma. World J. Oncol. 2024, 15, 929–941. [Google Scholar] [CrossRef]
- Shan, H.; Zhang, X.; Zhang, X.; Wei, Y.; Meng, L.; Li, J.; Zhang, Z.; Ma, L.; Xiao, Y. CircSCUBE3 Reduces the Anti-gastric Cancer Activity of Anti-PD-L1. Mol. Biotechnol. 2024, 66, 123–137. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.L.; Sheng, H.; Zhang, D.S.; Jin, Y.; Zhao, B.T.; Chen, N.; Song, K.; Xu, R.H. The circular RNA circDLG1 promotes gastric cancer progression and anti-PD-1 resistance through the regulation of CXCL12 by sponging miR-141-3p. Mol. Cancer 2021, 20, 166. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Cai, Z.R.; Huang, R.Z.; Wang, D.S.; Ju, H.Q.; Chen, D.L. Circular RNA circPHLPP2 promotes tumor growth and anti-PD-1 resistance through binding ILF3 to regulate IL36gamma transcription in colorectal cancer. Mol. Cancer 2024, 23, 272. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.L.; Chen, N.; Sheng, H.; Zhang, D.S. Circular RNA circNCOA3 promotes tumor progression and anti-PD-1 resistance in colorectal cancer. Cancer Drug Resist. 2024, 7, 9. [Google Scholar] [CrossRef]
- Ding, N.; You, A.B.; Yang, H.; Hu, G.S.; Lai, C.P.; Liu, W.; Ye, F. A Tumor-suppressive Molecular Axis EP300/circRERE/miR-6837-3p/MAVS Activates Type I IFN Pathway and Antitumor Immunity to Suppress Colorectal Cancer. Clin. Cancer Res. 2023, 29, 2095–2109. [Google Scholar] [CrossRef]
- Wang, F.; Lu, Q.; Yu, H.; Zhang, X.M. The Circular RNA circFGFR4 Facilitates Resistance to Anti-PD-1 of Triple-Negative Breast Cancer by Targeting the miR-185-5p/CXCR4 Axis. Cancer Manag. Res. 2023, 15, 825–835. [Google Scholar] [CrossRef]
- Wang, Z.; Li, Y.; Yang, J.; Sun, Y.; He, Y.; Wang, Y.; Liang, Y.; Chen, X.; Chen, T.; Han, D.; et al. CircCFL1 Promotes TNBC Stemness and Immunoescape via Deacetylation-Mediated c-Myc Deubiquitylation to Facilitate Mutant TP53 Transcription. Adv. Sci. 2024, 11, e2404628. [Google Scholar] [CrossRef]
- Ge, J.; Meng, Y.; Guo, J.; Chen, P.; Wang, J.; Shi, L.; Wang, D.; Qu, H.; Wu, P.; Fan, C.; et al. Human papillomavirus-encoded circular RNA circE7 promotes immune evasion in head and neck squamous cell carcinoma. Nat. Commun. 2024, 15, 8609. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Ju, C.; Du, D.; Zhu, P.; Yin, J.; Jia, J.; Wang, X.; Xu, X.; Zhao, L.; Wan, J.; et al. CircNF1 modulates the progression and immune evasion of esophageal squamous cell carcinoma through dual regulation of PD-L1. Cell Mol. Biol. Lett. 2025, 30, 37. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Wang, Z.; Gao, P.; Cao, D.; Dong, R.; Zhu, M.; Fei, Y.; Zuo, X.; Cai, J. CircRHBDD1 promotes immune escape via IGF2BP2/PD-L1 signaling and acts as a nanotherapeutic target in gastric cancer. J. Transl. Med. 2024, 22, 704. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Xu, X.; Xu, K.; Zhou, X.; Wu, K.; Yao, Y.; Liu, Z.; Chen, C.; Wang, L.; Sun, Z.; et al. N6-methyladenosine-modified circSLCO1B3 promotes intrahepatic cholangiocarcinoma progression via regulating HOXC8 and PD-L1. J. Exp. Clin. Cancer Res. 2024, 43, 119. [Google Scholar] [CrossRef]
- Ye, R.; Lu, X.; Liu, J.; Duan, Q.; Xiao, J.; Duan, X.; Yue, Z.; Liu, F. CircSOD2 Contributes to Tumor Progression, Immune Evasion and Anti-PD-1 Resistance in Hepatocellular Carcinoma by Targeting miR-497-5p/ANXA11 Axis. Biochem. Genet. 2023, 61, 597–614. [Google Scholar] [CrossRef]
- Ebrahimi, A.; Barati, T.; Mirzaei, Z.; Fattahi, F.; Mansoori Derakhshan, S.; Shekari Khaniani, M. An overview on the interaction between non-coding RNAs and CTLA-4 gene in human diseases. Med. Oncol. 2024, 42, 13. [Google Scholar] [CrossRef]
- Liu, Z.; Zheng, N.; Li, J.; Li, C.; Zheng, D.; Jiang, X.; Ge, X.; Liu, M.; Liu, L.; Song, Z.; et al. N6-methyladenosine-modified circular RNA QSOX1 promotes colorectal cancer resistance to anti-CTLA-4 therapy through induction of intratumoral regulatory T cells. Drug Resist. Updat. 2022, 65, 100886. [Google Scholar] [CrossRef]
- Zhou, B.; Mo, Z.; Lai, G.; Chen, X.; Li, R.; Wu, R.; Zhu, J.; Zheng, F. Targeting tumor exosomal circular RNA cSERPINE2 suppresses breast cancer progression by modulating MALT1-NF-κB-IL-6 axis of tumor-associated macrophages. J. Exp. Clin. Cancer Res. 2023, 42, 48. [Google Scholar] [CrossRef]
- Hu, Z.; Chen, G.; Zhao, Y.; Gao, H.; Li, L.; Yin, Y.; Jiang, J.; Wang, L.; Mang, Y.; Gao, Y.; et al. Exosome-derived circCCAR1 promotes CD8+ T-cell dysfunction and anti-PD1 resistance in hepatocellular carcinoma. Mol. Cancer 2023, 22, 55. [Google Scholar] [CrossRef]
- Liu, L.; Liao, R.; Wu, Z.; Du, C.; You, Y.; Que, K.; Duan, Y.; Yin, K.; Ye, W. Hepatic stellate cell exosome-derived circWDR25 promotes the progression of hepatocellular carcinoma via the miRNA-4474-3P-ALOX-15 and EMT axes. Biosci. Trends 2022, 16, 267–281. [Google Scholar] [CrossRef]
- Zhang, P.F.; Gao, C.; Huang, X.Y.; Lu, J.C.; Guo, X.J.; Shi, G.M.; Cai, J.B.; Ke, A.W. Cancer cell-derived exosomal circUHRF1 induces natural killer cell exhaustion and may cause resistance to anti-PD1 therapy in hepatocellular carcinoma. Mol. Cancer 2020, 19, 110. [Google Scholar] [CrossRef]
- Chen, Y.; Zhu, H.; Luo, Y.; Xie, T.; Hu, Y.; Yan, Z.; Ji, W.; Wang, Y.; Yin, Q.; Xian, H. ALDOC promotes neuroblastoma progression and modulates sensitivity to chemotherapy drugs by enhancing aerobic glycolysis. Front. Immunol. 2025, 16, 573815. [Google Scholar] [CrossRef] [PubMed]
- Xiong, W.; Yang, J. CircSEC24A induces KLF8 expression to promote the malignant progression of non-small cell lung cancer by regulating miR-1253. Thorac. Cancer 2024, 15, 2461–2473. [Google Scholar] [CrossRef] [PubMed]
- Zuo, Y.; Ren, D.; He, H.; Huang, C.; Zhu, X. CircST6GALNAC6 Inhibits Glycolysis of Bladder Cancer by Regulating PRKN/HK1 Signaling Pathway. Mol. Carcinog. 2025, 64, 870–882. [Google Scholar] [CrossRef] [PubMed]
- Ou, W.; Tan, R.; Zhai, J.; Sun, L.; Quan, Z.; Huang, X.; Xu, F.; Xu, Q.; Zhou, C. Silencing circ_0043256 inhibited CoCl2-induced proliferation, migration, and aerobic glycolysis in gastric cancer cells. Sci. Rep. 2025, 15, 171. [Google Scholar] [CrossRef]
- Qian, L.; Wang, L.; Chen, H.; Wang, S.; Hou, Y.; Xu, L.; Xia, Y.; Xu, M.; Huang, X. Hsa_circ_0001756 drives gastric cancer glycolysis by increasing the expression and stability of PGK1 mRNA. Front. Immunol. 2025, 16, 1511247. [Google Scholar] [CrossRef]
- Li, C.; Tang, Y.; Zhang, R.; Shi, L.; Chen, J.; Zhang, P.; Zhang, N.; Li, W. Inhibiting glycolysis facilitated checkpoint blockade therapy for triple-negative breast cancer. Discov. Oncol. 2025, 16, 550. [Google Scholar] [CrossRef]
- Shen, C.; Suo, Y.; Guo, J.; Su, W.; Zhang, Z.; Yang, S.; Wu, Z.; Fan, Z.; Zhou, X.; Hu, H. Development and validation of a glycolysis-associated gene signature for predicting the prognosis, immune landscape, and drug sensitivity in bladder cancer. Front. Immunol. 2025, 15, 1430583. [Google Scholar] [CrossRef]
- Bai, Y.; Chen, D.; Cheng, C.; Li, Z.; Chi, H.; Zhang, Y.; Zhang, X.; Tang, S.; Zhao, Q.; Ang, B.; et al. Immunosuppressive landscape in hepatocellular carcinoma revealed by single-cell sequencing. Front. Immunol. 2022, 13, 950536. [Google Scholar] [CrossRef]
- Wang, Y.; Yuan, J.; Liu, J.; Li, X.; Zhou, C.; Qian, M.; Zou, Z.; Lu, C.; Huang, G.; Jin, M. Melittin suppresses aerobic glycolysis by regulating HSF1/PDK3 to increase chemosensitivity of NSCLC. Eur. J. Pharmacol. 2025, 986, 177084. [Google Scholar] [CrossRef]
- Tang, F.; Cui, Q. Diverse roles of aldolase enzymes in cancer development, drug resistance and therapeutic approaches as moonlighting enzymes. Med. Oncol. 2024, 41, 224. [Google Scholar] [CrossRef]
- Wang, K.X.; Shi, D.M.; Shi, X.L.; Wang, J.Y.; Ai, X.H. Obesity promotes immunotherapy efficacy by up-regulating the glycolytic-mediated histone lactacylation modification of CD8+ T cells. Front. Pharmacol. 2025, 16, 1533464. [Google Scholar] [CrossRef]
- Tu, X.; Lin, W.; Zhai, X.; Liang, S.; Huang, G.; Wang, J.; Jia, W.; Li, S.; Li, B.; Cheng, B. Oleanolic acid inhibits M2 macrophage polarization and potentiates anti-PD-1 therapy in hepatocellular carcinoma by targeting miR-130b-3p-PTEN-PI3K-Akt signaling and glycolysis. Phytomedicine 2025, 141, 156750. [Google Scholar] [CrossRef] [PubMed]
- Ye, X.; Lin, J.; Chen, Y.; Wang, X. IGF2BP1 accelerates the aerobic glycolysis to boost its immune escape in hepatocellular carcinoma microenvironment. Front. Immunol. 2024, 15, 1480834. [Google Scholar] [CrossRef] [PubMed]
- Ding, C.H.; Yan, F.Z.; Xu, B.N.; Qian, H.; Hong, X.L.; Liu, S.Q.; Luo, Y.Y.; Wu, S.H.; Cai, L.Y.; Zhang, X.; et al. PRMT3 drives PD-L1-mediated immune escape through activating PDHK1-regulated glycolysis in hepatocellular carcinoma. Cell Death Dis. 2025, 16, 158. [Google Scholar] [CrossRef]
- Zhu, Q.; Li, J.; Sun, H.; Fan, Z.; Hu, J.; Chai, S.; Lin, B.; Wu, L.; Qin, W.; Wang, Y.; et al. O-GlcNAcylation of enolase 1 serves as a dual regulator of aerobic glycolysis and immune evasion in colorectal cancer. Proc. Natl. Acad. Sci. USA 2024, 121, e2408354121. [Google Scholar] [CrossRef]
- Xu, X.; Li, J.; Li, R.; Tan, Y.; Lu, Z. circPOLA2 promotes proliferation, invasion, migration, and epithelial-mesenchymal transition in breast cancer via the miR-1224-5p/HMGA2 axis. Clinics 2025, 80, 100653. [Google Scholar] [CrossRef]
- Molaei, P.; Mahdavinezhad, A.; Najafi, R.; Hashemi, M.; Tapak, L.; Afshar, S. Role of hsa_Circ_0001821 in Colorectal Cancer Pathogenesis and Response to 5-Fluorouracil through miR-203a-3p/FGF-2 Axis. Iran Biomed. J. 2025, 29, 82–89. [Google Scholar]
- Han, Z.; Chum, L.T.; Lin, X.; Zeng, T. Exploring the diagnostic potential of plasma circ-CCDC66 in colorectal cancer. Sci. Rep. 2025, 15, 11463. [Google Scholar] [CrossRef]
- Abdellatif, E.M.; Hegazy, N.E.; Nassar, E.S. Evaluation of the role of circular RNA (circ-FAF1) as a diagnostic biomarker for breast cancer in a cohort of Egyptian breast cancer patients. Mol. Biol. Rep. 2025, 52, 174. [Google Scholar] [CrossRef]
- Zhang, F.; Jiang, J.; Qian, H.; Yan, Y.; Xu, W. Exosomal circRNA: Emerging insights into cancer progression and clinical application potential. J. Hematol. Oncol. 2023, 16, 67. [Google Scholar] [CrossRef]
- Chen, H.; Wang, X.; Liu, S.; Tang, Z.; Xie, F.; Yin, J.; Sun, P.; Wang, H. Circular RNA in Pancreatic Cancer: Biogenesis, Mechanism, Function and Clinical Application. Int. J. Med. Sci. 2025, 22, 1612–1629. [Google Scholar] [CrossRef] [PubMed]
- Li, L.X.; Hao, Y.; Dong, L.; Qiao, Z.Q.; Yang, S.C.; Chen, Y.D.; Zhang, K.; Wang, Y.W. Circular RNAs as Biomarkers in Breast Cancer Diagnosis, Prognosis, Molecular Types, Metastasis and Drug Resistance. Technol. Cancer Res Treat. 2025, 24, 15330338251328500. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Liu, X.; Shen, T.; Wang, Q.; Zhou, S.; Yang, S.; Liao, S.; Su, T.; Mei, L.; Zhang, B.; et al. Small circular RNAs as vaccines for cancer immunotherapy. Nat. Biomed. Eng. 2025, 9, 249–267. [Google Scholar] [CrossRef] [PubMed]
- Xia, L.; Li, G.; Zhou, Q.; Feng, Y.U.; Ma, H. CircRNA circ_0015278 induces ferroptosis in lung adenocarcinoma through the miR-1228/P53 axis. Oncol. Res. 2025, 33, 465–475. [Google Scholar] [CrossRef]
- Khan, M.A. alpha-Synuclein Iron-Responsive-Element RNA and Iron Regulatory Protein Affinity Is Specifically Reduced by Iron in Parkinson’s Disease. Biomolecules 2025, 15, 214. [Google Scholar] [CrossRef]
- Wesselhoeft, R.A.; Kowalski, P.S.; Anderson, D.G. Engineering circular RNA for potent and stable translation in eukaryotic cells. Nat. Commun. 2018, 9, 2629. [Google Scholar] [CrossRef]
- Koch, P.; Zhang, Z.; Genuth, N.R.; Susanto, T.T.; Haimann, M.; Khmelinskaia, A.; Byeon, G.W.; Dey, S.; Barna, M.; Leppek, K. A versatile toolbox for determining IRES activity in cells and embryonic tissues. EMBO J 2025, 44, 2695–2724. [Google Scholar] [CrossRef]
- Wei, Z.; Liu, J.; Hui, G.; Luan, X. Circ_0020123 promotes non-small cell lung cancer progression via miR-146a-5p mediated regulation of EIF4G2 expression. Thorac. Cancer 2024, 15, 44–56. [Google Scholar] [CrossRef]
- Zeng, K.; Peng, J.; Xing, Y.; Zhang, L.; Zeng, P.; Li, W.; Zhang, W.; Pan, Z.; Zhou, C.; Lin, J. A positive feedback circuit driven by m(6)A-modified circular RNA facilitates colorectal cancer liver metastasis. Mol. Cancer 2023, 22, 202. [Google Scholar] [CrossRef]
- Obi, P.; Chen, Y.G. The design and synthesis of circular RNAs. Methods 2021, 196, 85–103. [Google Scholar] [CrossRef]
- Nakamoto, K.; Abe, N.; Tsuji, G.; Kimura, Y.; Tomoike, F.; Shimizu, Y.; Abe, H. Chemically synthesized circular RNAs with phosphoramidate linkages enable rolling circle translation. Chem. Commun. 2020, 56, 6217–6220. [Google Scholar] [CrossRef]
- Kim, H.; Kim, D.; Moon, S.; Lee, J.B. Efficient circular RNA synthesis through Gap-DNA splint-mediated ligation. Nanoscale 2024, 16, 15529–15532. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Lu, L.; Liu, L.; Liang, R.; Zhang, Q.; Liu, Z.; An, J.; Liu, Q.; Wu, Q.; Wei, S.; et al. Group IIC self-splicing intron-derived novel circular RNA vaccine elicits superior immune response against RSV. Front. Immunol. 2025, 16, 1574568. [Google Scholar] [CrossRef] [PubMed]
- Su, C.I.; Chuang, Z.S.; Shie, C.T.; Wang, H.I.; Kao, Y.T.; Yu, C.Y. A cis-acting ligase ribozyme generates circular RNA in vitro for ectopic protein functioning. Nat. Commun. 2024, 15, 6607. [Google Scholar] [CrossRef]
- Chen, X.; Lu, Y. Circular RNA: Biosynthesis in vitro. Front. Bioeng. Biotechnol. 2021, 9, 787881. [Google Scholar] [CrossRef]
- Gao, X.; Chen, K.; Wang, H. NicOPURE: Nickless RNA circularization and one-step purification with engineered group II introns and cyclizing UTRs. Nucleic Acids Res. 2025, 53, gkaf310. [Google Scholar] [CrossRef]
- Wang, L.; Dong, C.; Zhang, W.; Ma, X.; Rou, W.; Yang, K.; Cui, T.; Qi, S.; Yang, L.; Xie, J.; et al. Developing an enhanced chimeric permuted intron-exon system for circular RNA therapeutics. Theranostics 2024, 14, 5869–5882. [Google Scholar] [CrossRef]
- Tai, J.; Chen, Y.G. Differences in the immunogenicity of engineered circular RNAs. J. Mol. Cell Biol. 2023, 15, mjad002. [Google Scholar] [CrossRef]
- Niu, D.; Wu, Y.; Lian, J. Circular RNA vaccine in disease prevention and treatment. Signal Transduct. Target Ther. 2023, 8, 341. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Wen, Y.; Yu, W.; Lu, L.; Yang, Y.; Liu, C.; Hu, Z.; Fang, Z.; Huang, S. Optimized circular RNA vaccines for superior cancer immunotherapy. Theranostics 2025, 15, 1420–1438. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Hu, Y.; Li, J.; He, J.; Yu, G.; Wang, J.; Lin, X. Intranasal prime-boost RNA vaccination elicits potent T cell response for lung cancer therapy. Signal Transduct. Target Ther. 2025, 10, 101. [Google Scholar] [CrossRef] [PubMed]
- Alshehry, Y.; Liu, X.; Zhang, Y.; Zhu, G. Investigation of the impact of lipid nanoparticle compositions on the delivery and T cell response of circRNA vaccine. J. Control Release 2025, 381, 113617. [Google Scholar] [CrossRef]
- Amaya, L.; Grigoryan, L.; Li, Z.; Lee, A.; Wender, P.A.; Pulendran, B.; Chang, H.Y. Circular RNA vaccine induces potent T cell responses. Proc. Natl. Acad. Sci. USA 2023, 120, e2302191120. [Google Scholar] [CrossRef]
- Qu, L.; Yi, Z.; Shen, Y.; Lin, L.; Chen, F.; Xu, Y.; Wu, Z.; Tang, H.; Zhang, X.; Tian, F.; et al. Circular RNA vaccines against SARS-CoV-2 and emerging variants. Cell 2022, 185, 1728–1744. [Google Scholar] [CrossRef]
- Li, H.; Peng, K.; Yang, K.; Ma, W.; Qi, S.; Yu, X.; He, J.; Lin, X.; Yu, G. Circular RNA cancer vaccines drive immunity in hard-to-treat malignancies. Theranostics 2022, 12, 6422–6436. [Google Scholar] [CrossRef]
- Ren, Y.; Manoharan, T.; Liu, B.; Cheng, C.Z.M.; En Siew, B.; Cheong, W.K.; Lee, K.Y.; Tan, I.J.; Lieske, B.; Tan, K.K.; et al. Circular RNA as a source of neoantigens for cancer vaccines. J. Immunother. Cancer 2024, 12, e008402. [Google Scholar] [CrossRef]
- Chen, J.; Gao, Y.; Zhong, J.; Wu, X.; Leng, Z.; Liu, M.; Wang, Y.; Wang, Y.; Yang, X.; Huang, N.; et al. Lnc-H19-derived protein shapes the immunosuppressive microenvironment of glioblastoma. Cell Rep. Med. 2024, 5, 101806. [Google Scholar] [CrossRef]
- Wang, F.; Cai, G.; Wang, Y.; Zhuang, Q.; Cai, Z.; Li, Y.; Gao, S.; Li, F.; Zhang, C.; Zhao, B.; et al. Circular RNA-based neoantigen vaccine for hepatocellular carcinoma immunotherapy. MedComm 2024, 5, e667. [Google Scholar] [CrossRef]
- Cai, Z.; Wuri, Q.; Song, Y.; Qu, X.; Hu, H.; Cao, S.; Wu, H.; Wu, J.; Wang, C.; Yu, X.; et al. CircRNA-loaded DC vaccine in combination with low-dose gemcitabine induced potent anti-tumor immunity in pancreatic cancer model. Cancer Immunol. Immunother. 2025, 74, 68. [Google Scholar] [CrossRef]
- de la Peña, M.; Ceprián, R.; Cervera, A. A Singular and Widespread Group of Mobile Genetic Elements: RNA Circles with Autocatalytic Ribozymes. Cells 2020, 9, 2555. [Google Scholar] [CrossRef]
- Yang, J.; Zhu, J.; Sun, J.; Chen, Y.; Du, Y.; Tan, Y.; Wu, L.; Zhai, M.; Wei, L.; Li, N.; et al. Intratumoral delivered novel circular mRNA encoding cytokines for immune modulation and cancer therapy. Mol. Ther. Nucleic Acids 2022, 30, 184–197. [Google Scholar] [CrossRef] [PubMed]
- Fu, J.; Liu, F.; Bai, S.; Jiang, X.; Song, H.; Zhang, M.; Zhao, R.; Ouyang, T.; Yu, M.; Qian, H.; et al. Circular RNA CDYL facilitates hepatocellular carcinoma stemness and PD-L1(+) exosomes-mediated immunotherapy resistance via stabilizing hornerin protein by blocking synoviolin 1-mediated ubiquitination. Int. J. Biol. Macromol. 2025, 310, 143246. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Liu, W.; Zhang, M.; Li, Y.; Tang, H.; Wang, Y.; Song, C.; Song, B.; Tan, B. Circ_0001947 encapsulated by small extracellular vesicles promotes gastric cancer progression and anti-PD-1 resistance by modulating CD8+ T cell exhaustion. J. Nanobiotechnology 2024, 22, 563. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.; Zhu, X.; Ye, S.; Zhang, J.; Liao, J.; Zhang, N.; Zeng, X.; Wang, J.; Yang, B.; Zhang, Y.; et al. Tumour circular RNAs elicit anti-tumour immunity by encoding cryptic peptides. Nature 2024, 625, 593–602. [Google Scholar] [CrossRef]
- Zhang, X.; Li, M.; Chen, K.; Liu, Y.; Liu, J.; Wang, J.; Huang, H.; Zhang, Y.; Huang, T.; Ma, S. Engineered circular guide RNAs enhance miniature CRISPR/Cas12f-based gene activation and adenine base editing. Nat. Commun. 2025, 16, 3016. [Google Scholar] [CrossRef]
- Jahns, H.; Degaonkar, R.; Podbevsek, P.; Gupta, S.; Bisbe, A.; Aluri, K.; Szeto, J.; Kumar, P.; LeBlanc, S.; Racie, T.; et al. Small circular interfering RNAs (sciRNAs) as a potent therapeutic platform for gene-silencing. Nucleic Acids Res 2021, 49, 10250–10264. [Google Scholar] [CrossRef]
- Wang, J.; Zhou, Y.; Sun, L.; Liu, T.; Qu, H.; Mao, Y.; Zheng, L. Binding-Site Directed Selection and Large-Scale Click-Synthesis of a Coagulation Factor XIa-Inhibiting Circular DNA Aptamer. Chemistry 2025, 31, e202404372. [Google Scholar] [CrossRef]
- Huang, G.; Zhou, H.; Xiang, Q.; Zhang, J.; Hu, X.; Cheng, R.; Lan, L.; Wang, Y.; Shen, Z. Exponential and efficient target-catalyst rolling circle amplification for label-free and ultrasensitive fluorescent detection of miR-21 and p53 gene. Anal. Chim. Acta 2022, 1221, 340132. [Google Scholar] [CrossRef]
- Xiao, M.S.; Wilusz, J.E. An improved method for circular RNA purification using RNase R that efficiently removes linear RNAs containing G-quadruplexes or structured 3′ ends. Nucleic Acids Res. 2019, 47, 8755–8769. [Google Scholar] [CrossRef]
- Li, Q.; Zhao, Y.H.; Xu, C.; Liang, Y.L.; Zhao, Y.; He, Q.M.; Li, J.Y.; Chen, K.L.; Qiao, H.; Liu, N.; et al. Chemotherapy-Induced Senescence Reprogramming Promotes Nasopharyngeal Carcinoma Metastasis by circRNA-Mediated PKR Activation. Adv. Sci. 2023, 10, e2205668. [Google Scholar] [CrossRef] [PubMed]
- Muskan, M.; Abeysinghe, P.; Cecchin, R.; Branscome, H.; Morris, K.V.; Kashanchi, F. Therapeutic potential of RNA-enriched extracellular vesicles: The next generation in RNA delivery via biogenic nanoparticles. Mol. Ther. 2024, 32, 2939–2949. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Xu, S.; Hong, X.; Liu, Y.; Qian, S.; Lou, Y.; Wang, W. ADAR1 prevents ZBP1-dependent PANoptosis via A-to-I RNA editing in developmental sevoflurane neurotoxicity. Cell Biol. Toxicol. 2024, 40, 57. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Wang, X.; Zhang, Y.; Zuo, S.; Chereda, B.; Gregory, P.A.; Zhao, C.X.; Goodall, G.J. A Dual-Selection System for Enhanced Efficiency and Fidelity of Circular RNA Overexpression. J. Mol. Biol. 2025, 437, 169064. [Google Scholar] [CrossRef]
- Chen, J.; Ye, Z.; Huang, C.; Qiu, M.; Song, D.; Li, Y.; Xu, Q. Lipid nanoparticle-mediated lymph node-targeting delivery of mRNA cancer vaccine elicits robust CD8+ T cell response. Proc. Natl. Acad. Sci. USA 2022, 119, e2207841119. [Google Scholar] [CrossRef]
- Qi, Y.; Han, H.; Liu, A.; Zhao, S.; Lawanprasert, A.; Nielsen, J.E.; Choudhary, H.; Liang, D.; Barron, A.E.; Murthy, N. Ethylene oxide graft copolymers reduce the immunogenicity of lipid nanoparticles. RSC Adv. 2024, 14, 30071–30076. [Google Scholar] [CrossRef]
- Kozma, G.T.; Mészáros, T.; Berényi, P.; Facskó, R.; Patkó, Z.; Oláh, C.Z.; Nagy, A.; Fülöp, T.G.; Glatter, K.A.; Radovits, T.; et al. Role of anti-polyethylene glycol (PEG) antibodies in the allergic reactions to PEG-containing COVID-19 vaccines: Evidence for immunogenicity of PEG. Vaccine 2023, 41, 4561–4570. [Google Scholar] [CrossRef]
- Wang, J.H.; Gessler, D.J.; Zhan, W.; Gallagher, T.L.; Gao, G. Adeno-associated virus as a delivery vector for gene therapy of human diseases. Signal Transduct. Target Ther. 2024, 9, 78. [Google Scholar] [CrossRef]
- Lavenniah, A.; Luu, T.D.A.; Li, Y.P.; Lim, T.B.; Jiang, J.; Ackers-Johnson, M.; Foo, R.S. Engineered Circular RNA Sponges Act as miRNA Inhibitors to Attenuate Pressure Overload-Induced Cardiac Hypertrophy. Mol. Ther. 2020, 28, 1506–1517. [Google Scholar] [CrossRef]
- Zhao, P.; Yin, C.; Liu, R.; Shao, S.; Ke, W.; Song, Z. Exosome-Delivered circFOXP1 Upregulates Autophagy and Promotes Hepatocellular Carcinoma Progression Through Its Encoded p196 Protein Targeting the KHDRBS3/ULK1 Axis. Int. J. Nanomed. 2025, 20, 5247–5265. [Google Scholar] [CrossRef]
- Jhunjhunwala, S.; Hammer, C.; Delamarre, L. Antigen presentation in cancer: Insights into tumour immunogenicity and immune evasion. Nat. Rev. Cancer 2021, 21, 298–312. [Google Scholar] [CrossRef] [PubMed]
- Sultan, H.; Takeuchi, Y.; Ward, J.P.; Sharma, N.; Liu, T.T.; Sukhov, V.; Firulyova, M.; Song, Y.; Ameh, S.; Brioschi, S.; et al. Neoantigen-specific cytotoxic Tr1 CD4 T cells suppress cancer immunotherapy. Nature 2024, 632, 182–191. [Google Scholar] [CrossRef] [PubMed]
- Bulashevska, A.; Nacsa, Z.; Lang, F.; Braun, M.; Machyna, M.; Diken, M.; Childs, L.; König, R. Artificial intelligence and neoantigens: Paving the path for precision cancer immunotherapy. Front. Immunol. 2024, 15, 1394003. [Google Scholar] [CrossRef] [PubMed]
- Mackensen, A.; Haanen, J.B.A.G.; Koenecke, C.; Alsdorf, W.; Wagner-Drouet, E.; Borchmann, P.; Heudobler, D.; Ferstl, B.; Klobuch, S.; Bokemeyer, C.; et al. CLDN6-specific CAR-T cells plus amplifying RNA vaccine in relapsed or refractory solid tumors: The phase 1 BNT211-01 trial. Nat. Med. 2023, 29, 2844–2853. [Google Scholar] [CrossRef]
- Cai, J.; Liu, Z.; Chen, S.; Zhang, J.; Li, H.; Wang, X.; Yang, F.; Wang, S.; Li, X.; Li, Y.; et al. Engineered circular RNA-based DLL3-targeted CAR-T therapy for small cell lung cancer. Exp. Hematol. Oncol. 2025, 14, 35. [Google Scholar] [CrossRef]
- He, A.T.; Liu, J.; Li, F.; Yang, B.B. Targeting circular RNAs as a therapeutic approach: Current strategies and challenges. Signal Transduct. Target Ther. 2021, 6, 185. [Google Scholar] [CrossRef]
- Du, P.; Xu, X.; Wang, Y. Hsa_circ_0000585 promotes chemoresistance to cis-platin in epithelial cells of ovarian cancer by modulating autophagy. Biochem. Biophys. Res. Commun. 2023, 678, 186–192. [Google Scholar] [CrossRef]






| Circular RNAs | Effect on Drug Resistance | Mechanism | Refs |
|---|---|---|---|
| CircCCNY | Sensitivity to lenvatinib ↑ |
| [54] |
| Circ_0000376 | Docetaxel resistance ↑ |
| [59] |
| Circ_0076305 | Paclitaxel resistance ↑ |
| [62] |
| CircQKI | Docetaxel resistance ↑ |
| [63] |
| Circ_0058608 | Taxol resistance ↑ |
| [64] |
| CircRNA_0044556 | Paclitaxel resistance ↑ |
| [65] |
| Circ_0005667 | Cisplatin resistance ↑ |
| [66] |
| CircKIAA0182 | Cisplatin resistance ↑ |
| [70] |
| CircFRMD4A | Sensitivity to elesclomol ↑ |
| [73] |
| Circular RNAs | Effect on Immune Responses | Mechanism | Refs |
|---|---|---|---|
| CircPETH_147aa | Immune evasion |
| [77] |
| Circ_CPA4 | Immune evasion |
| [78] |
| Circ_CHST15 | Immune evasion |
| [79] |
| CircENTPD7 | Immune evasion |
| [81] |
| CircFAM64A(3) | Immune evasion |
| [82] |
| CircAATF | Immune evasion |
| [83] |
| CircMGA | Immune activation |
| [46] |
| Circ_001377 | Immune evasion |
| [84] |
| CircDLG1 | Immune evasion |
| [86] |
| CircPHLPP2 | Immune evasion |
| [87] |
| CircNCOA3 | Immune activation |
| [88] |
| CircFGFR4 | Immune evasion |
| [90] |
| CircCFL1 | Immune evasion |
| [91] |
| CircNF1 | Immune evasion |
| [93] |
| CircRHBDD1 | Immune evasion |
| [94] |
| CircSOD2 | Immune evasion |
| [96] |
| CircQSOX1 | Immune evasion |
| [98] |
| Exosomal circCCAR1 | Immune evasion |
| [100] |
| Exosomal circWDR25 | Immune evasion |
| [101] |
| Exosomal circUHRF1 | Immune evasion |
| [102] |
| Circular RNAs | Effect on Glycolysis | Mechanism | Refs |
|---|---|---|---|
| CircPVT1 | Enhances glycolytic activity |
| [4] |
| circZNF707 | Enhances glycolytic activity |
| [45] |
| CircRUNX1 | Enhances glycolytic activity |
| [95] |
| CircSEC24A | Enhances glycolytic activity |
| [104] |
| circST6GALNAC6 | Inhibits glycolysis |
| [105] |
| Circ_0043256 | Enhances glycolytic activity |
| [106] |
| Circ_0001756 | Enhances glycolytic activity |
| [107] |
| Title | Phase | Condition or Disease | Prospective Outcome Measures | Dates | ID/Purpose |
|---|---|---|---|---|---|
| A Single Arm Clinical Study of Dendritic Cell Vaccine Loaded With Circular RNA Encoding Cryptic Peptide for Patients With HER2-negative Advanced Breast Cancer | Phase I, not yet recruiting, enrollment 48 | Advanced breast cancer |
| 1 December 2024 (start) 1 January 2027 (completion) | NCT06530082/Treatment |
| CIRcular and Non-coding RNAs as Clinically USeful Biomarkers in Pancreaticobiliary Cancers |
| Pancreatic cancer, Biliary tract cancer |
| 4 October 2020 (start) 5 November 2023 (completion) | NCT04584996/Observational |
| The Value of circRNAs (hsa_circ_0004001) in Early Diagnosis of HCC |
| Hepatocellular carcinoma |
| November 2023 (start) February 2025 (completion) | NCT06042842/Observational |
| Deciphering the Role of Circular RNAs in ALK positive Anaplastic Large-cell Lymphoma (CIRComa) |
| Anaplastic Large-cell Lymphoma |
| 1 January 2023 (start) 31 December 2025 (completion) | NCT05934045/observational |
| The Role of CircDENND4C in Epithelial Ovarian Cancer |
| Epithelial ovarian can cer |
| 1 December 2024 (start) 1 October 2028 (completion) | NCT06617585/observational |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jeoung, J.; Kim, W.; Jo, H.; Jeoung, D. Circular RNAs as Targets for Developing Anticancer Therapeutics. Cells 2025, 14, 1106. https://doi.org/10.3390/cells14141106
Jeoung J, Kim W, Jo H, Jeoung D. Circular RNAs as Targets for Developing Anticancer Therapeutics. Cells. 2025; 14(14):1106. https://doi.org/10.3390/cells14141106
Chicago/Turabian StyleJeoung, Jaewhoon, Wonho Kim, Hyein Jo, and Dooil Jeoung. 2025. "Circular RNAs as Targets for Developing Anticancer Therapeutics" Cells 14, no. 14: 1106. https://doi.org/10.3390/cells14141106
APA StyleJeoung, J., Kim, W., Jo, H., & Jeoung, D. (2025). Circular RNAs as Targets for Developing Anticancer Therapeutics. Cells, 14(14), 1106. https://doi.org/10.3390/cells14141106







