IL-3-Induced Immediate Expression of c-fos and c-jun Is Modulated by the IKK2-JNK Axis
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Culture and Cell Stimulation
2.2. Generation of Knockout (KO) Cells by Genome Editing
2.3. RNA Extraction, Reverse Transcription (RT), and Quantitative PCR
2.4. Plasmid Construction and Retroviral Transduction
2.5. Cell Lysis, Immunoprecipitation, and Immunoblot Analyses
2.6. Statistical Analyses
3. Results
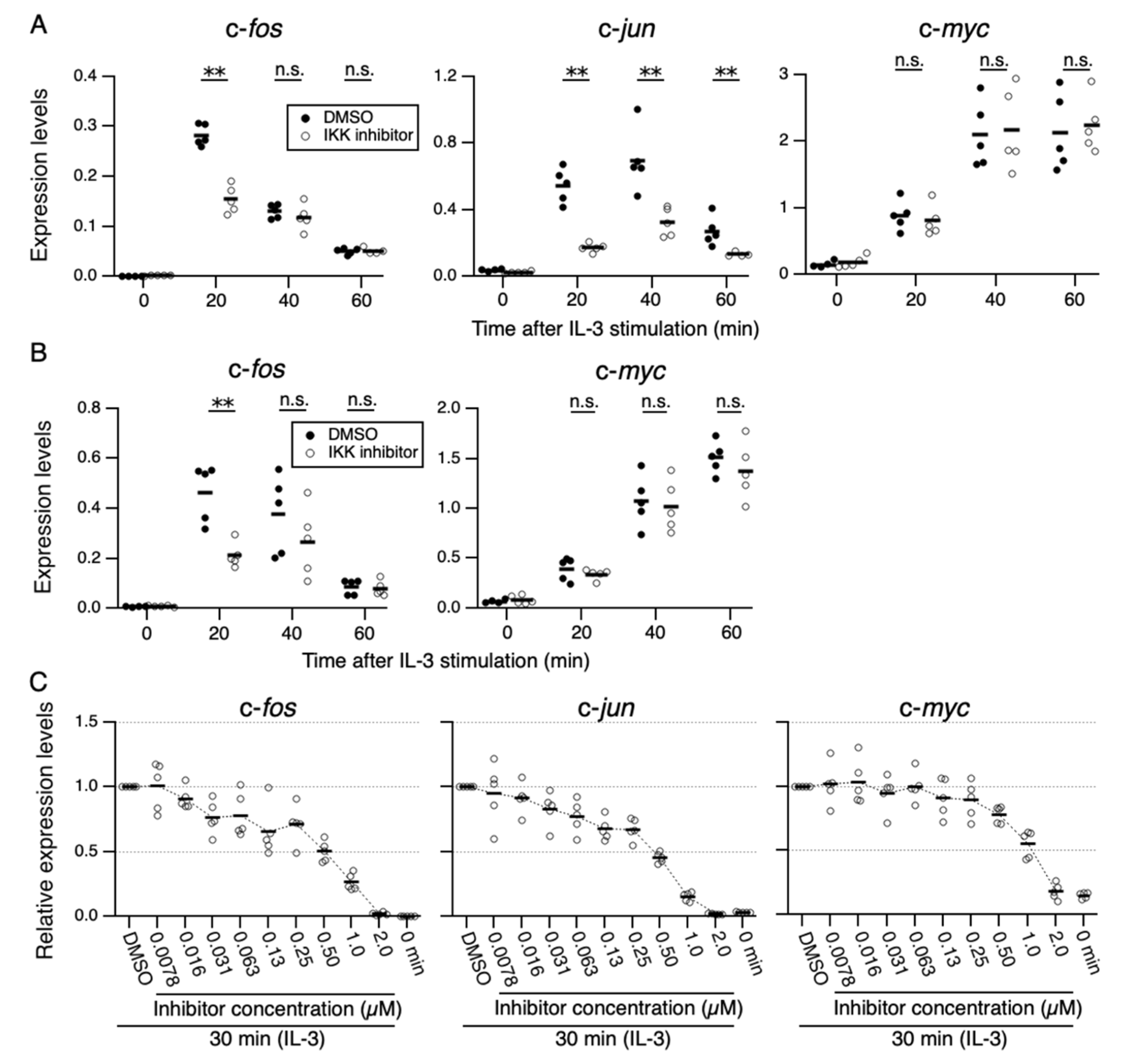
3.1. Suppression of IL-3-Mediated Expression of IEGs by an IKK Inhibitor
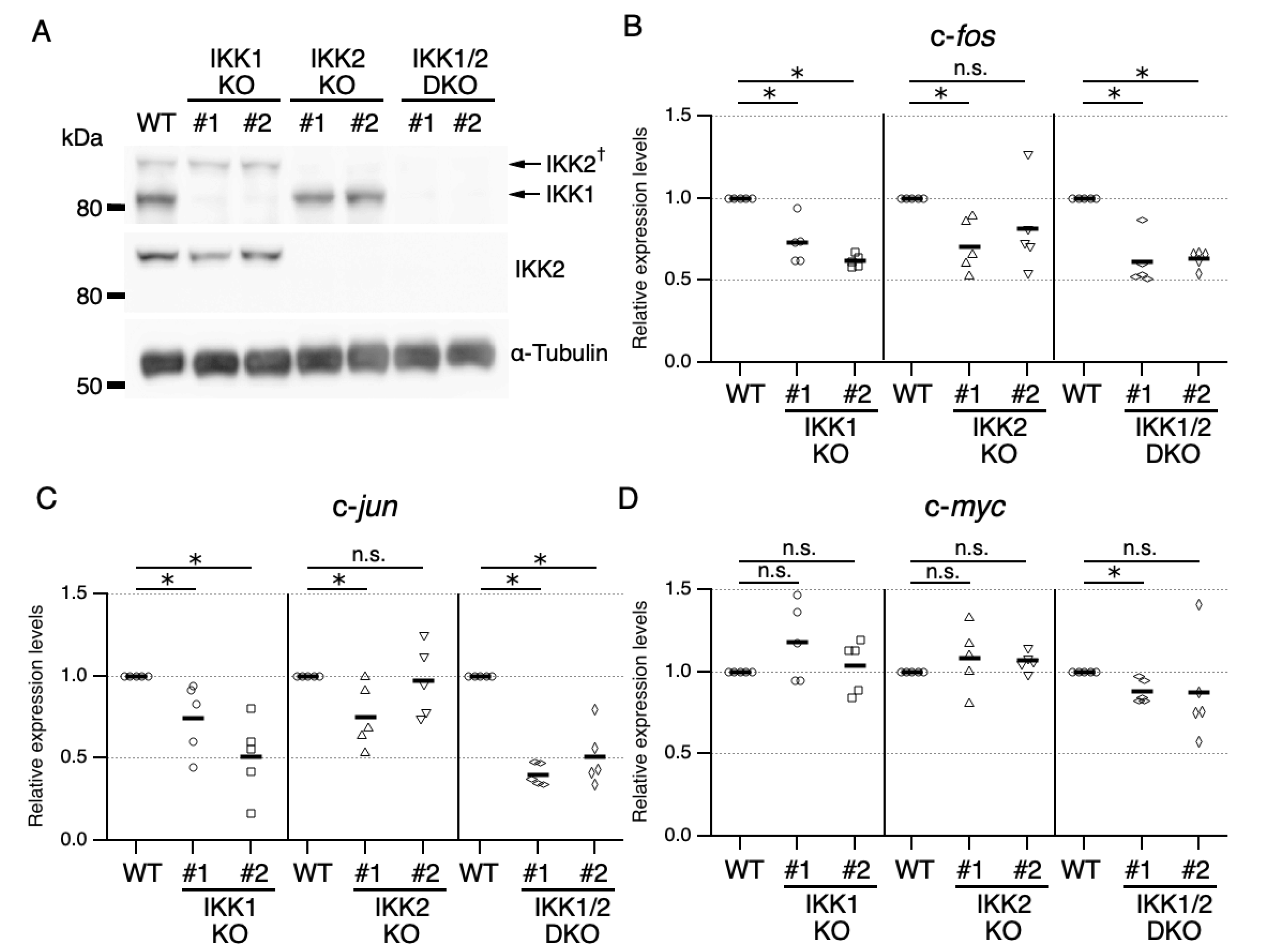
3.2. KO of IKKs Suppressed IL-3-Mediated Expression of IEGs
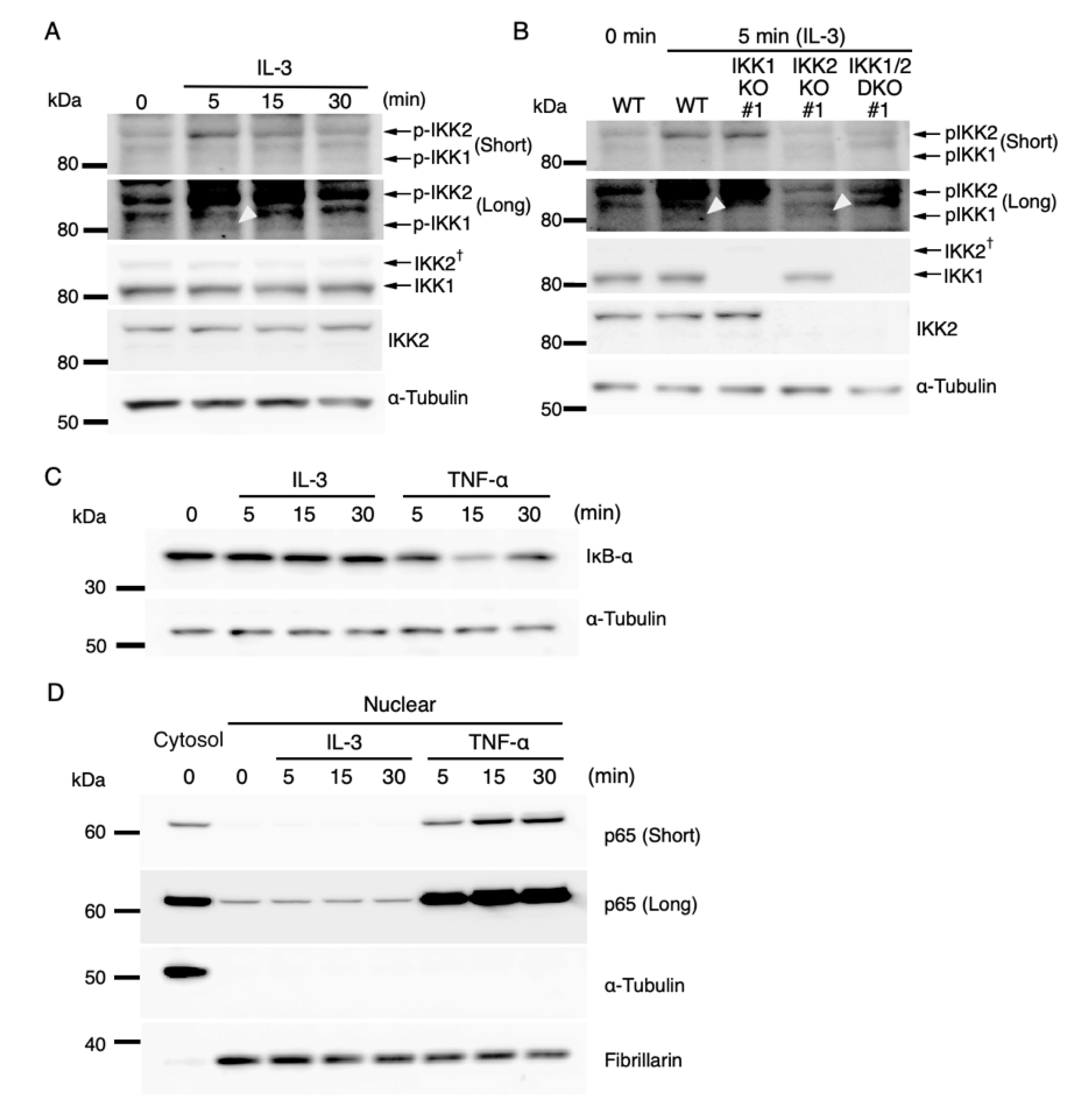
3.3. Regulation of IL-3-Mediated Expression of IEGs by Activated IKKs in a Canonical NF-κB-Independent Manner
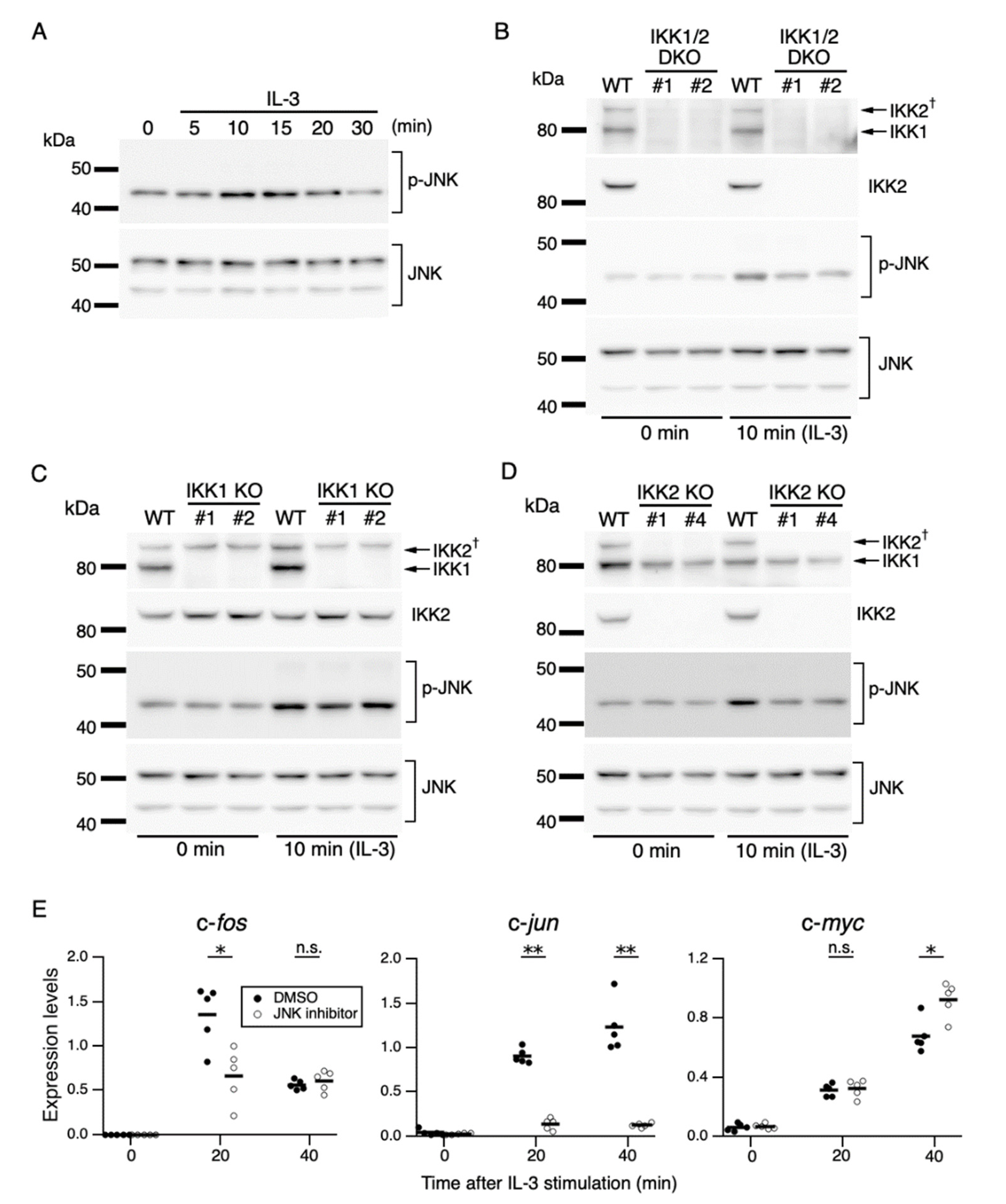
3.4. JNK Mediates c-fos and c-jun Expression Downstream of IKK2
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hercus, T.R.; Kan, W.L.T.; Broughton, S.E.; Tvorogov, D.; Ramshaw, H.S.; Sandow, J.J.; Nero, T.L.; Dhagat, U.; Thompson, E.J.; Shing, K.S.C.T.; et al. Role of the β Common (βc) Family of Cytokines in Health and Disease. Cold Spring Harb. Perspect. Biol. 2018, 10, a028514. [Google Scholar] [CrossRef] [PubMed]
- Dougan, M.; Dranoff, G.; Dougan, S.K. GM-CSF, IL-3, and IL-5 Family of Cytokines: Regulators of Inflammation. Immunity 2019, 50, 796–811. [Google Scholar] [CrossRef] [PubMed]
- Lantz, C.S.; Boesiger, J.; Song, C.H.; Mach, N.; Kobayashi, T.; Mulligan, R.C.; Nawa, Y.; Dranoff, G.; Galli, S.J. Role for Interleukin-3 in Mast-Cell and Basophil Development and in Immunity to Parasites. Nature 1998, 392, 90–93. [Google Scholar] [CrossRef] [PubMed]
- Schroeder, J.T.; Chichester, K.L.; Bieneman, A.P. Human Basophils Secrete IL-3: Evidence of Autocrine Priming for Phenotypic and Functional Responses in Allergic Disease. J. Immunol. 2009, 182, 2432–2438. [Google Scholar] [CrossRef] [PubMed]
- Borriello, F.; Longo, M.; Spinelli, R.; Pecoraro, A.; Granata, F.; Staiano, R.I.; Loffredo, S.; Spadaro, G.; Beguinot, F.; Schroeder, J.; et al. IL-3 Synergises with Basophil-Derived IL-4 and IL-13 to Promote the Alternative Activation of Human Monocytes. Eur. J. Immunol. 2015, 45, 2042–2051. [Google Scholar] [CrossRef]
- Esnault, S.; Kelly, E.A.B.; Shen, Z.J.; Johansson, M.W.; Malter, J.S.; Jarjour, N.N. IL-3 Maintains Activation of the P90S6K/RPS6 Pathway and Increases Translation in Human Eosinophils. J. Immunol. 2015, 195, 2529–2539. [Google Scholar] [CrossRef]
- Weber, G.F.; Chousterman, B.G.; He, S.; Fenn, A.M.; Nairz, M.; Anzai, A.; Brenner, T.; Uhle, F.; Iwamoto, Y.; Robbins, C.S.; et al. Interleukin-3 Amplifies Acute Inflammation and Is a Potential Therapeutic Target in Sepsis. Science 2015, 347, 1260–1265. [Google Scholar] [CrossRef]
- Frankel, A.E.; Woo, J.H.; Ahn, C.; Pemmaraju, N.; Medeiros, B.C.; Carraway, H.E.; Frankfurt, O.; Forman, S.J.; Yang, X.A.; Konopleva, M.; et al. Activity of SL-401, a Targeted Therapy Directed to Interleukin-3 Receptor, in Blastic Plasmacytoid Dendritic Cell Neoplasm Patients. Blood 2014, 124, 385–392. [Google Scholar] [CrossRef] [PubMed]
- Jordan, C.T.; Upchurch, D.; Szilvassy, S.J.; Guzman, M.L.; Howard, D.S.; Pettigrew, A.L.; Meyerrose, T.; Rossi, R.; Grimes, B.; Rizzieri, D.A.; et al. The Interleukin-3 Receptor Alpha Chain Is a Unique Marker for Human Acute Myelogenous Leukemia Stem Cells. Leukemia 2000, 14, 1777–1784. [Google Scholar] [CrossRef]
- Nievergall, E.; Ramshaw, H.S.; Yong, A.S.M.; Biondo, M.; Busfield, S.J.; Vairo, G.; Lopez, A.F.; Hughes, T.P.; White, D.L.; Hiwase, D.K. Monoclonal Antibody Targeting of IL-3 Receptor α with CSL362 Effectively Depletes CML Progenitor and Stem Cells. Blood 2014, 123, 1218–1228. [Google Scholar] [CrossRef] [PubMed]
- Pardanani, A.; Lasho, T.; Chen, D.; Kimlinger, T.K.; Finke, C.; Zblewski, D.; Patnaik, M.M.; Reichard, K.K.; Rowinsky, E.; Hanson, C.A.; et al. Aberrant Expression of CD123 (Interleukin-3 Receptor-α) on Neoplastic Mast Cells. Leukemia 2015, 29, 1605–1608. [Google Scholar] [CrossRef] [PubMed]
- Testa, U.; Riccioni, R.; Militi, S.; Coccia, E.; Stellacci, E.; Samoggia, P.; Latagliata, R.; Mariani, G.; Rossini, A.; Battistini, A.; et al. Elevated Expression of IL-3Ralpha in Acute Myelogenous Leukemia Is Associated with Enhanced Blast Proliferation, Increased Cellularity, and Poor Prognosis. Blood 2002, 100, 2980–2988. [Google Scholar] [CrossRef]
- Jin, L.; Lee, E.M.; Ramshaw, H.S.; Busfield, S.J.; Peoppl, A.G.; Wilkinson, L.; Guthridge, M.A.; Thomas, D.; Barry, E.F.; Boyd, A.; et al. Monoclonal Antibody-Mediated Targeting of CD123, IL-3 Receptor Alpha Chain, Eliminates Human Acute Myeloid Leukemic Stem Cells. Cell Stem Cell 2009, 5, 31–42. [Google Scholar] [CrossRef]
- Quelle, F.W.; Sato, N.; Witthuhn, B.A.; Inhorn, R.C.; Eder, M.; Miyajima, A.; Griffin, J.D.; Ihle, J.N. JAK2 Associates with the Beta c Chain of the Receptor for Granulocyte-Macrophage Colony-Stimulating Factor, and Its Activation Requires the Membrane-Proximal Region. Mol. Cell. Biol. 1994, 14, 4335–4341. [Google Scholar] [CrossRef] [PubMed]
- Hercus, T.R.; Broughton, S.E.; Ekert, P.G.; Ramshaw, H.S.; Perugini, M.; Grimbaldeston, M.; Woodcock, J.M.; Thomas, D.; Pitson, S.; Hughes, T.; et al. The GM-CSF Receptor Family: Mechanism of Activation and Implications for Disease. Growth Factors 2012, 30, 63–75. [Google Scholar] [CrossRef]
- Martinez-Moczygemba, M.; Huston, D.P. Biology of Common Beta Receptor-Signaling Cytokines: IL-3, IL-5, and GM-CSF. J. Allergy Clin. Immunol. 2003, 112, 653–665. [Google Scholar] [CrossRef] [PubMed]
- Conscience, J.F.; Verrier, B.; Martin, G. Interleukin-3-Dependent Expression of the c-Myc and c-Fos Proto-Oncogenes in Hemopoietic Cell Lines. EMBO J. 1986, 5, 317–323. [Google Scholar] [CrossRef] [PubMed]
- Lord, K.A.; Abdollahi, A.; Hoffman-Liebermann, B.; Liebermann, D.A. Proto-Oncogenes of the Fos/Jun Family of Transcription Factors Are Positive Regulators of Myeloid Differentiation. Mol. Cell. Biol. 1993, 13, 841–851. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.N.; Tuckerman, J.; Nechushtan, H.; Schutz, G.; Razin, E.; Angel, P. c-Fos as a Regulator of Degranulation and Cytokine Production in FcepsilonRI-Activated Mast Cells. J. Immunol. 2004, 173, 2571–2577. [Google Scholar] [CrossRef] [PubMed]
- Shafarenko, M.; Amanullah, A.; Gregory, B.; Liebermann, D.A.; Hoffman, B. Fos Modulates Myeloid Cell Survival and Differentiation and Partially Abrogates the c-Myc Block in Terminal Myeloid Differentiation. Blood 2004, 103, 4259–4267. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, S.; Muto, A.; Yokota, T.; Miyajima, A.; Arai, K. Differential Regulation of Early Response Genes and Cell Proliferation through the Human Granulocyte Macrophage Colony-Stimulating Factor Receptor: Selective Activation of the c-Fos Promoter by Genistein. Mol. Biol. Cell 1993, 4, 983–992. [Google Scholar] [CrossRef][Green Version]
- Mui, A.L.; Wakao, H.; Kinoshita, T.; Kitamura, T.; Miyajima, A. Suppression of Interleukin-3-Induced Gene Expression by a C-Terminal Truncated Stat5: Role of Stat5 in Proliferation. EMBO J. 1996, 15, 2425–2433. [Google Scholar] [CrossRef] [PubMed]
- Basham, B.; Sathe, M.; Grein, J.; McClanahan, T.; D’Andrea, A.; Lees, E.; Rascle, A. In Vivo Identification of Novel STAT5 Target Genes. Nucleic Acids Res. 2008, 36, 3802–3818. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.X.; Watanabe, S.; Muto, A.; Miyajima, A.; Yokota, T.; Arai, K. Activation of Early Response Genes and Cell Proliferation by Human Interleukin-3, Granulocyte-Macrophage Colony-Stimulating Factor, and Interleukin-5 Receptors: Comparison with Human Interleukin-4 Receptor Signaling. J. Allergy Clin. Immunol. 1994, 94, 605–611. [Google Scholar] [CrossRef]
- Itoh, T.; Muto, A.; Watanabe, S.; Miyajima, A.; Yokota, T.; Arai, K. Granulocyte-Macrophage Colony-Stimulating Factor Provokes RAS Activation and Transcription of c-Fos through Different Modes of Signaling. J. Biol. Chem. 1996, 271, 7587–7592. [Google Scholar] [CrossRef] [PubMed]
- Karin, M. The Regulation of AP-1 Activity by Mitogen-Activated Protein Kinases. J. Biol. Chem. 1995, 270, 16483–16486. [Google Scholar] [CrossRef] [PubMed]
- O’Donnell, A.; Odrowaz, Z.; Sharrocks, A.D. Immediate-Early Gene Activation by the MAPK Pathways: What Do and Don’t We Know? Biochem. Soc. Trans. 2012, 40, 58–66. [Google Scholar] [CrossRef]
- Taniguchi, K.; Karin, M. NF-κB, Inflammation, Immunity and Cancer: Coming of Age. Nat. Rev. Immunol. 2018, 18, 309–324. [Google Scholar] [CrossRef] [PubMed]
- Sun, S.C. The Non-Canonical NF-κB Pathway in Immunity and Inflammation. Nat. Rev. Immunol. 2017, 17, 545–558. [Google Scholar] [CrossRef]
- Hinz, M.; Scheidereit, C. The IκB Kinase Complex in NF-κB Regulation and Beyond. EMBO Rep. 2014, 15, 46–61. [Google Scholar] [CrossRef]
- Antonia, R.J.; Hagan, R.S.; Baldwin, A.S. Expanding the View of IKK: New Substrates and New Biology. Trends Cell Biol. 2021, 31, 166–178. [Google Scholar] [CrossRef] [PubMed]
- Sandow, J.J.; Jabbour, A.M.; Condina, M.R.; Daunt, C.P.; Stomski, F.C.; Green, B.D.; Riffkin, C.D.; Hoffmann, P.; Guthridge, M.A.; Silke, J.; et al. Cytokine Receptor Signaling Activates an IKK-Dependent Phosphorylation of PUMA to Prevent Cell Death. Cell Death Differ. 2012, 19, 633–641. [Google Scholar] [CrossRef]
- Drube, S.; Weber, F.; Loschinski, R.; Beyer, M.; Rothe, M.; Rabenhorst, A.; Göpfert, C.; Meininger, I.; Diamanti, M.A.; Stegner, D.; et al. Subthreshold IKK Activation Modulates the Effector Functions of Primary Mast Cells and Allows Specific Targeting of Transformed Mast Cells. Oncotarget 2015, 6, 5354–5368. [Google Scholar] [CrossRef] [PubMed]
- Greenberger, J.S.; Sakakeeny, M.A.; Humphries, R.K.; Eaves, C.J.; Eckner, R.J. Demonstration of Permanent Factor-Dependent Multipotential (Erythroid/Neutrophil/Basophil) Hematopoietic Progenitor Cell Lines. Proc. Natl. Acad. Sci. USA 1983, 80, 2931–2935. [Google Scholar] [CrossRef]
- Palacios, R.; Steinmetz, M. Il-3-Dependent Mouse Clones That Express B-220 Surface Antigen, Contain Ig Genes in Germ-Line Configuration, and Generate B Lymphocytes In Vivo. Cell 1985, 41, 727–734. [Google Scholar] [CrossRef]
- Morita, S.; Kojima, T.; Kitamura, T. Plat-E: An Efficient and Stable System for Transient Packaging of Retroviruses. Gene Ther. 2000, 7, 1063–1066. [Google Scholar] [CrossRef]
- Naito, Y.; Hino, K.; Bono, H.; Ui-Tei, K. CRISPRdirect: Software for Designing CRISPR/Cas Guide RNA with Reduced off-Target Sites. Bioinformatics 2015, 31, 1120–1123. [Google Scholar] [CrossRef]
- Fujita, T.; Yuno, M.; Fujii, H. Allele-Specific Locus Binding and Genome Editing by CRISPR at the P16INK4a Locus. Sci. Rep. 2016, 6, 30485. [Google Scholar] [CrossRef]
- Nakano, H.; Shindo, M.; Sakon, S.; Nishinaka, S.; Mihara, M.; Yagita, H.; Okumura, K. Differential Regulation of IkappaB Kinase Alpha and Beta by Two Upstream Kinases, NF-KappaB-Inducing Kinase and Mitogen-Activated Protein Kinase/ERK Kinase Kinase-1. Proc. Natl. Acad. Sci. USA 1998, 95, 3537–3542. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing. Available online: http://www.R-project.org (accessed on 29 April 2019).
- Waelchli, R.; Bollbuck, B.; Bruns, C.; Buhl, T.; Eder, J.; Feifel, R.; Hersperger, R.; Janser, P.; Revesz, L.; Zerwes, H.G.; et al. Design and Preparation of 2-Benzamido-Pyrimidines as Inhibitors of IKK. Bioorg. Med. Chem. Lett. 2006, 16, 108–112. [Google Scholar] [CrossRef] [PubMed]
- Redner, R.L.; Lee, A.W.; Osawa, G.A.; Nienhuis, A.W. Variable Pattern of Jun and Fos Gene Expression in Different Hematopoietic Cell Lines during Interleukin 3-Induced Entry into the Cell Cycle. Oncogene 1992, 7, 43–50. [Google Scholar] [PubMed]
- Liu, F.; Xia, Y.; Parker, A.S.; Verma, I.M. IKK Biology. Immunol. Rev. 2012, 246, 239–253. [Google Scholar] [CrossRef] [PubMed]
- Anest, V.; Cogswell, P.C.; Baldwin, A.S., Jr. IkappaB Kinase Alpha and p65/RelA Contribute to Optimal Epidermal Growth Factor-Induced c-Fos Gene Expression Independent of IkappaBalpha Degradation. J. Biol. Chem. 2004, 279, 31183–31189. [Google Scholar] [CrossRef] [PubMed]
- Comb, W.C.; Cogswell, P.; Sitcheran, R.; Baldwin, A.S. IKK-Dependent, NF-κB-Independent Control of Autophagic Gene Expression. Oncogene 2011, 30, 1727–1732. [Google Scholar] [CrossRef]
- Nakamura, T.; Ouchida, R.; Kodama, T.; Kawashima, T.; Makino, Y.; Yoshikawa, N.; Watanabe, S.; Morimoto, C.; Kitamura, T.; Tanaka, H. Cytokine Receptor Common Beta Subunit-Mediated STAT5 Activation Confers NF-Kappa B Activation in Murine ProB Cell Line Ba/F3 Cells. J. Biol. Chem. 2002, 277, 6254–6265. [Google Scholar] [CrossRef]
- Zhang, T.; Inesta-Vaquera, F.; Niepel, M.; Zhang, J.; Ficarro, S.B.; Machleidt, T.; Xie, T.; Marto, J.A.; Kim, N.; Sim, T.; et al. Discovery of Potent and Selective Covalent Inhibitors of JNK. Chem. Biol. 2012, 19, 140–154. [Google Scholar] [CrossRef]
- Kettritz, R.; Choi, M.; Rolle, S.; Wellner, M.; Luft, F.C. Integrins and Cytokines Activate Nuclear Transcription Factor-KappaB in Human Neutrophils. J. Biol. Chem. 2004, 279, 2657–2665. [Google Scholar] [CrossRef]
- Meads, M.B.; Li, Z.W.; Dalton, W.S. A Novel TNF Receptor-Associated Factor 6 Binding Domain Mediates NF-Kappa B Signaling by the Common Cytokine Receptor Beta Subunit. J. Immunol. 2010, 185, 1606–1615. [Google Scholar] [CrossRef]
- Claudio, E.; Brown, K.; Park, S.; Wang, H.; Siebenlist, U. BAFF-Induced NEMO-Independent Processing of NF-Kappa B2 in Maturing B Cells. Nat. Immunol. 2002, 3, 958–965. [Google Scholar] [CrossRef]
- Liang, C.; Zhang, M.; Sun, S.C. Beta-TrCP Binding and Processing of NF-KappaB2/P100 Involve Its Phosphorylation at Serines 866 and 870. Cell. Signal. 2006, 18, 1309–1317. [Google Scholar] [CrossRef]
- Shaw, P.E.; Saxton, J. Ternary Complex Factors: Prime Nuclear Targets for Mitogen-Activated Protein Kinases. Int. J. Biochem. Cell Biol. 2003, 35, 1210–1226. [Google Scholar] [CrossRef]
- Nosaka, Y.; Arai, A.; Miyasaka, N.; Miura, O. CrkL Mediates Ras-Dependent Activation of the Raf/ERK Pathway through the Guanine Nucleotide Exchange Factor C3G in Hematopoietic Cells Stimulated with Erythropoietin or Interleukin-3. J. Biol. Chem. 1999, 274, 30154–30162. [Google Scholar] [CrossRef] [PubMed]
- Whitmarsh, A.J.; Yang, S.H.; Su, M.S.; Sharrocks, A.D.; Davis, R.J. Role of P38 and JNK Mitogen-Activated Protein Kinases in the Activation of Ternary Complex Factors. Mol. Cell. Biol. 1997, 17, 2360–2371. [Google Scholar] [CrossRef] [PubMed]
- Cavigelli, M.; Dolfi, F.; Claret, F.X.; Karin, M. Induction of c-Fos Expression through JNK-Mediated TCF/Elk-1 Phosphorylation. EMBO J. 1995, 14, 5957–5964. [Google Scholar] [CrossRef]
- Drobic, B.; Pérez-Cadahía, B.; Yu, J.; Kung, S.K.P.; Davie, J.R. Promoter Chromatin Remodeling of Immediate-Early Genes Is Mediated through H3 Phosphorylation at Either Serine 28 or 10 by the MSK1 Multi-Protein Complex. Nucleic Acids Res. 2010, 38, 3196–3208. [Google Scholar] [CrossRef] [PubMed]
- Josefowicz, S.Z.; Shimada, M.; Armache, A.; Li, C.H.; Miller, R.M.; Lin, S.; Yang, A.; Dill, B.D.; Molina, H.; Park, H.S.; et al. Chromatin Kinases Act on Transcription Factors and Histone Tails in Regulation of Inducible Transcription. Mol. Cell 2016, 64, 347–361. [Google Scholar] [CrossRef] [PubMed]
- Moser, B.; Hochreiter, B.; Basílio, J.; Gleitsmann, V.; Panhuber, A.; Pardo-Garcia, A.; Hoesel, B.; Salzmann, M.; Resch, U.; Noreen, M.; et al. The Inflammatory Kinase IKKα Phosphorylates and Stabilizes c-Myc and Enhances Its Activity. Mol. Cancer 2021, 20, 16. [Google Scholar] [CrossRef] [PubMed]
- Xia, X.; Li, L.; Choi, Y.S. Human Recombinant IL-3 Is a Growth Factor for Normal B Cells. J. Immunol. 1992, 148, 491–497. [Google Scholar] [PubMed]
- Snapper, C.M.; Moorman, M.A.; Rosas, F.R.; Kehry, M.R.; Maliszewski, C.R.; Mond, J.J. IL-3 and Granulocyte-Macrophage Colony-Stimulating Factor Strongly Induce Ig Secretion by Sort-Purified Murine B Cell Activated through the Membrane Ig, but Not the CD40, Signaling Pathway. J. Immunol. 1995, 154, 5842–5850. [Google Scholar] [PubMed]
- Srivastava, R.K.; Tomar, G.B.; Barhanpurkar, A.P.; Gupta, N.; Pote, S.T.; Mishra, G.C.; Wani, M.R. IL-3 Attenuates Collagen-Induced Arthritis by Modulating the Development of Foxp3+ Regulatory T Cells. J. Immunol. 2011, 186, 2262–2272. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Rani, L.; Mhaske, S.T.; Pote, S.T.; Behera, S.; Mishra, G.C.; Wani, M.R. IL-3 Receptor Expression on Activated Human Th Cells Is Regulated by IL-4, and IL-3 Synergizes with IL-4 to Enhance Th2 Cell Differentiation. J. Immunol. 2020, 204, 819–831. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.; Minemoto, Y.; Zhang, J.; Liu, J.; Tang, F.; Bui, T.N.; Xiang, J.; Lin, A. JNK Suppresses Apoptosis via Phosphorylation of the Proapoptotic Bcl-2 Family Protein BAD. Mol. Cell 2004, 13, 329–340. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fujita, H.; Fujita, T.; Fujii, H. IL-3-Induced Immediate Expression of c-fos and c-jun Is Modulated by the IKK2-JNK Axis. Cells 2022, 11, 1451. https://doi.org/10.3390/cells11091451
Fujita H, Fujita T, Fujii H. IL-3-Induced Immediate Expression of c-fos and c-jun Is Modulated by the IKK2-JNK Axis. Cells. 2022; 11(9):1451. https://doi.org/10.3390/cells11091451
Chicago/Turabian StyleFujita, Hirotaka, Toshitsugu Fujita, and Hodaka Fujii. 2022. "IL-3-Induced Immediate Expression of c-fos and c-jun Is Modulated by the IKK2-JNK Axis" Cells 11, no. 9: 1451. https://doi.org/10.3390/cells11091451
APA StyleFujita, H., Fujita, T., & Fujii, H. (2022). IL-3-Induced Immediate Expression of c-fos and c-jun Is Modulated by the IKK2-JNK Axis. Cells, 11(9), 1451. https://doi.org/10.3390/cells11091451





