WiFi Related Radiofrequency Electromagnetic Fields Promote Transposable Element Dysregulation and Genomic Instability in Drosophila melanogaster
Abstract
1. Introduction
2. Materials and Methods
2.1. Drosophila Strains
2.2. RF/EMF Exposure
2.3. Heat Shock Treatment
2.4. Reverse Transcription and Quantitative RT-PCR (qRT-PCR)
| blood F | TGCCACAGTACCTGATTTCG |
| blood R | GATTCGCCTTTTACGTTTGC |
| copia F | TGGAGGTTGTGCCTCCACTT |
| copia R | CAATACCACGCTTAGTGGCATAAA |
| gypsy F | CTTCACGTTCTGCGAGCGGTCT |
| gypsy R | CGCTGCAAGGTTACCAGGTAGGTTC |
| Het-A F | ACTGCTGAAGCTCGGATTCC |
| Het-A R | TGTAGCCGGATTCGTCATATTTC |
| hobo F | AAACTGTTCTGGACGGATGG |
| hobo R | TTATGGCGGGATAAATTGGA |
| I-element F | CAATCACAACAACAAAATCC |
| I-element R | GGTGTTGGTGTGGTTGGTTG |
| R2 F | ATGCTCCCGAAACAACAAAC |
| R2 R | GCACTGCAGACTTGGTTCAA |
| 1360 F | TCGTGCAAGACAATGAGAGG |
| 1360 R | GCAACTGGATCCCTTAGCAA |
| GstD1 F | CGCGCCATCCAGGTGTATTT |
| GstD1 R | CTGGTACAGCGTTCCCATGT |
| catalase F | CAACCCCTTCGATGTCACCA |
| catalase R | TCTGCTCCACCTCAGCAAAG |
| sod1 F | GAACAGGAGAGCAGCGGTA |
| sod1 R | TGCCATACGGATTGAAGTGC |
| sod2 F | CAAACTGCAAGCCTGGCG |
| sod2 R | CTGGTGGTGCTTCTGGTGAT |
| hsp83 F | CAGCTGGTCTCTGTCACCAA |
| hsp83 R | CTCTCGAACTTGGCCTTGTC |
| hsp70 F | CAACCTATCCATCAACCCAGAC |
| hsp70 R | ACGTAGCTCTCCAGAGCATTTC |
| hsr-ω F | TCTGCGACCGTGACTGAGATC |
| hsr-ω R | CAATCCGCACAATCAATCTGA |
| rp49 F | GCGCACCAAGCACTTCATC |
| rp49 R | TTGGGCTTGCGCCATT |
2.5. Western Blotting
2.6. Measurement of Eye Pigment
2.7. Mitotic Chromosome Preparations
2.8. H2DCFDA Staining
2.9. Larval Crawling Assay
2.10. Larval Light Preference Test
2.11. Climbing Assay
3. Results and Discussion
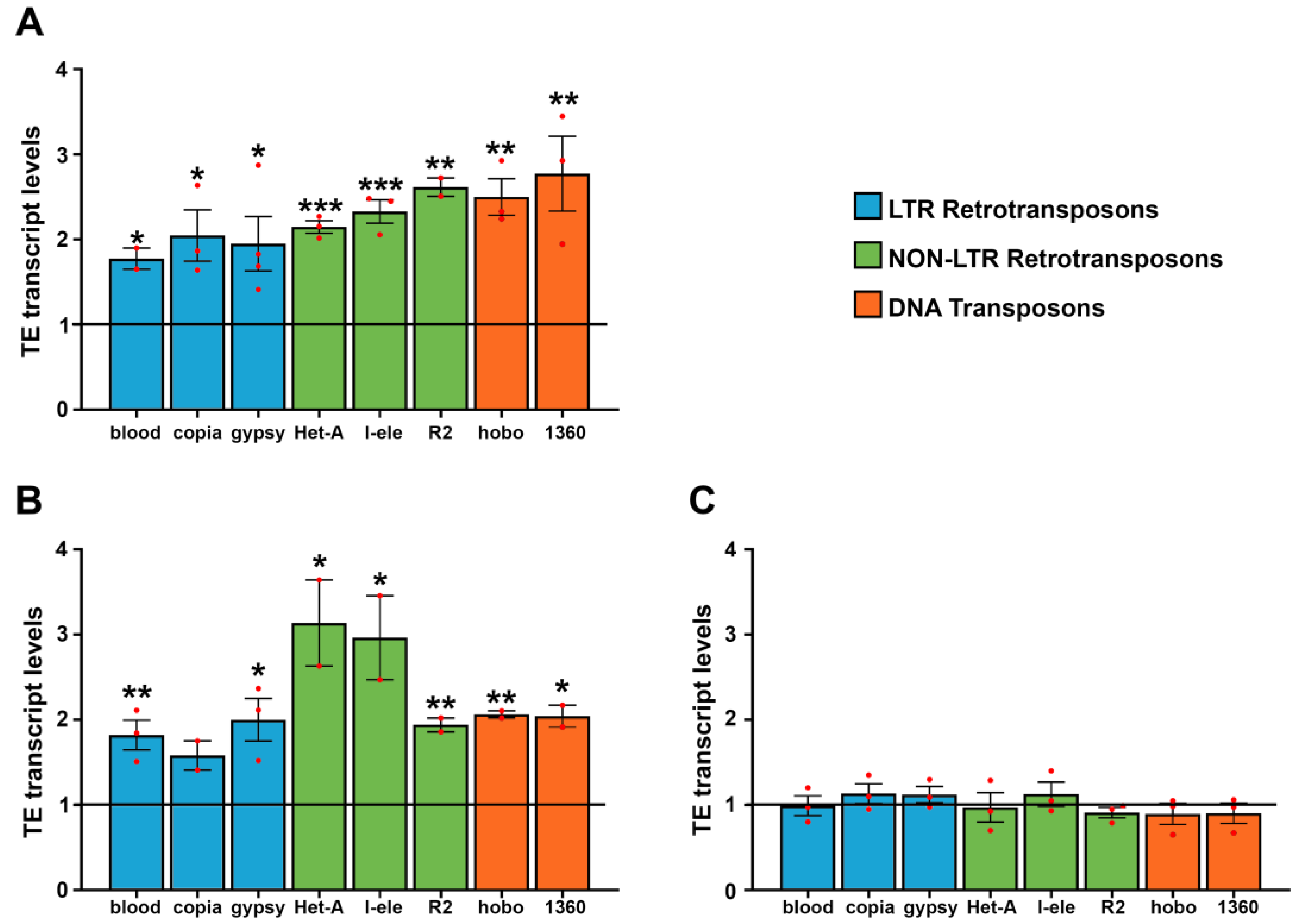
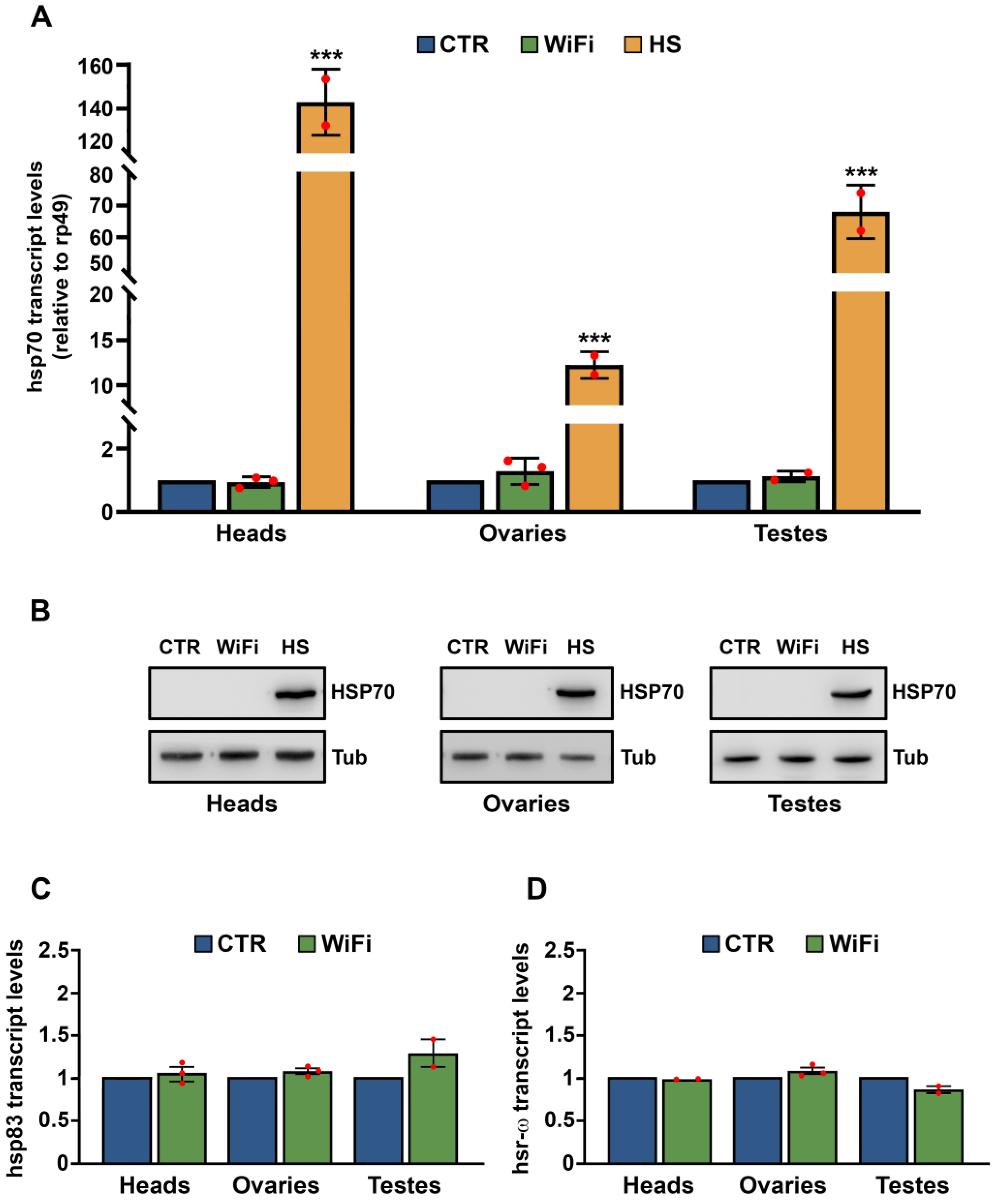
3.1. WiFi Electromagnetic Fields Induce Aberrant Expression of Transposable Elements
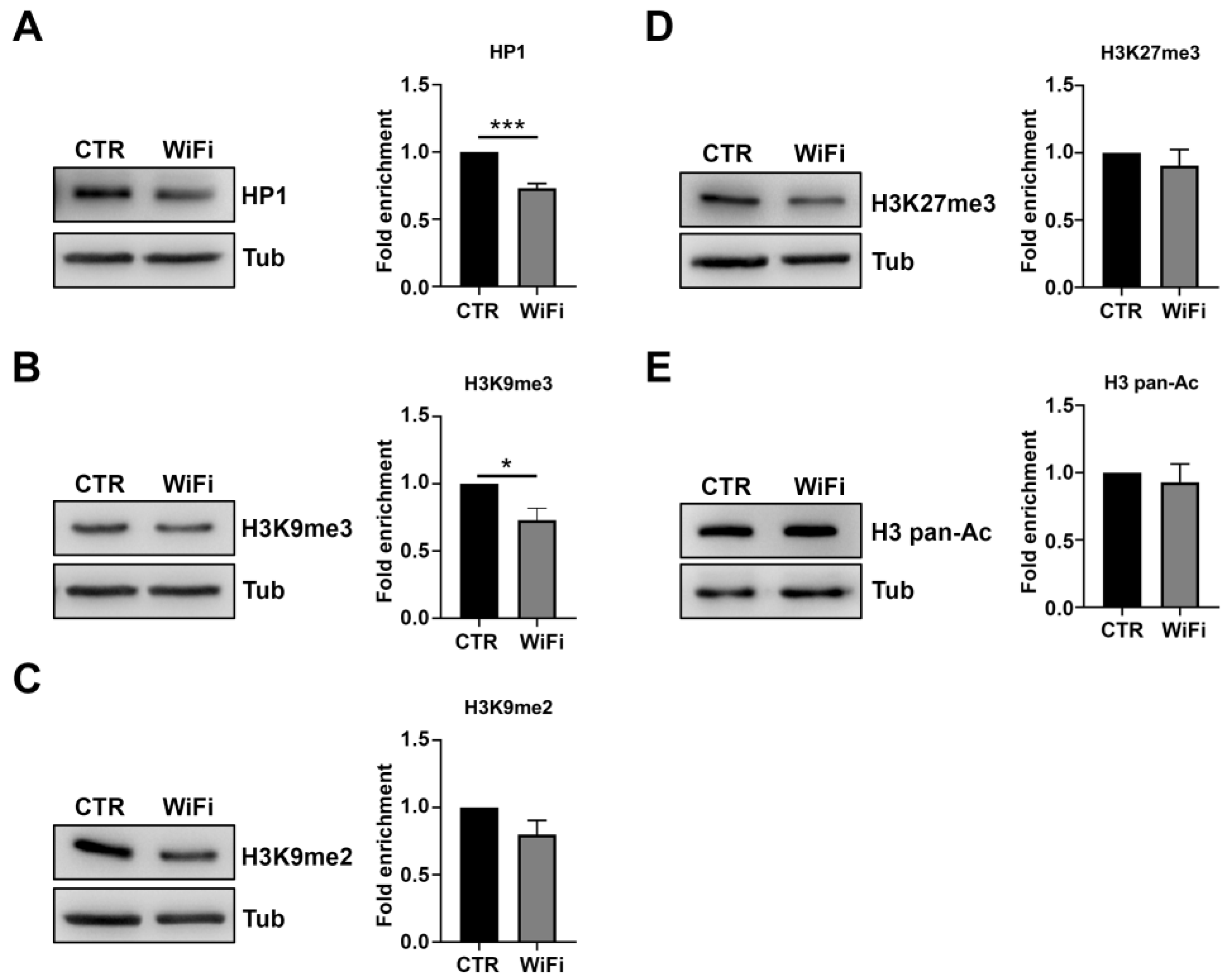
3.2. A Heterochromatin Breakdown Contributes to the Transcriptional Activation of Transposable Elements
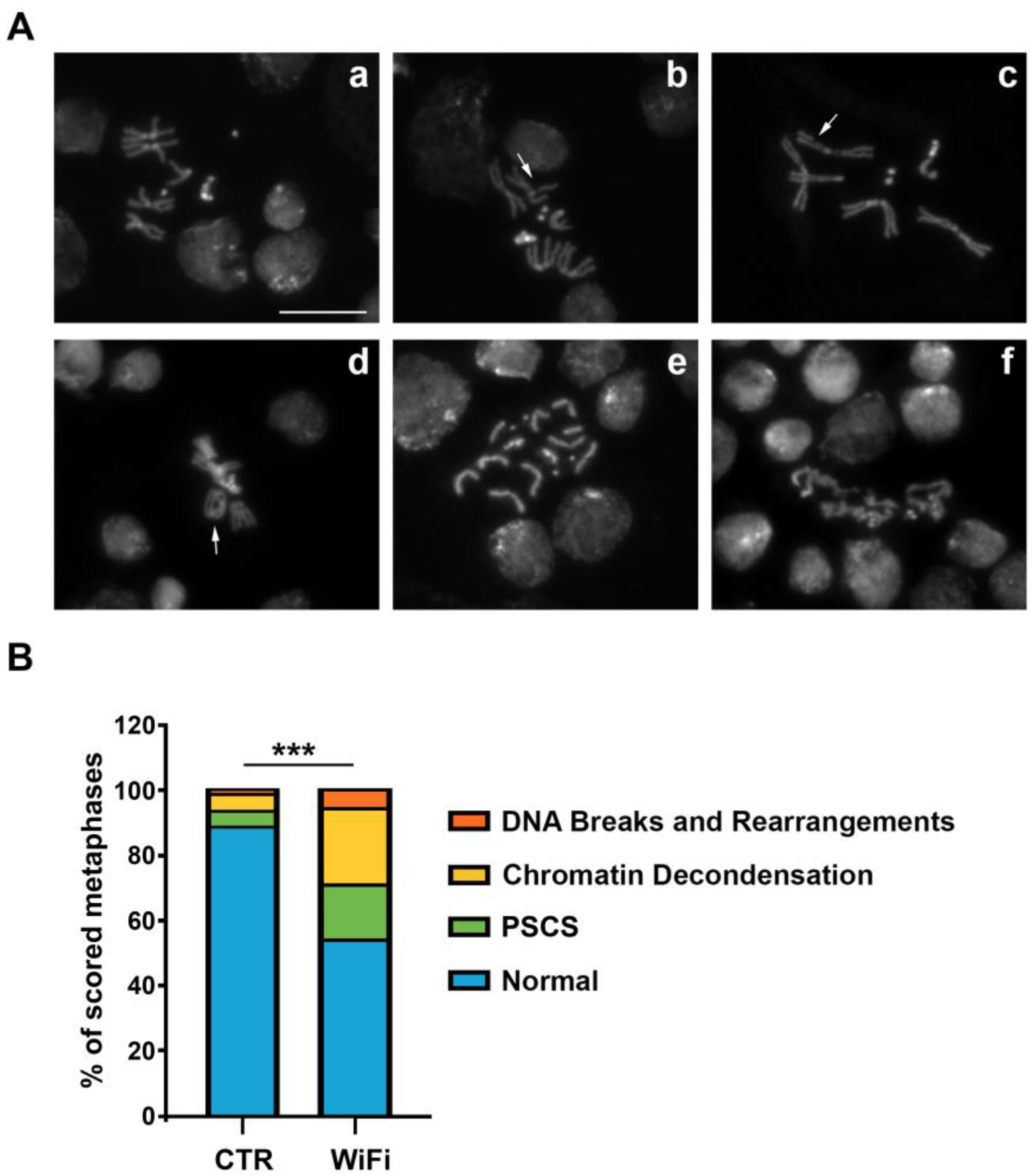
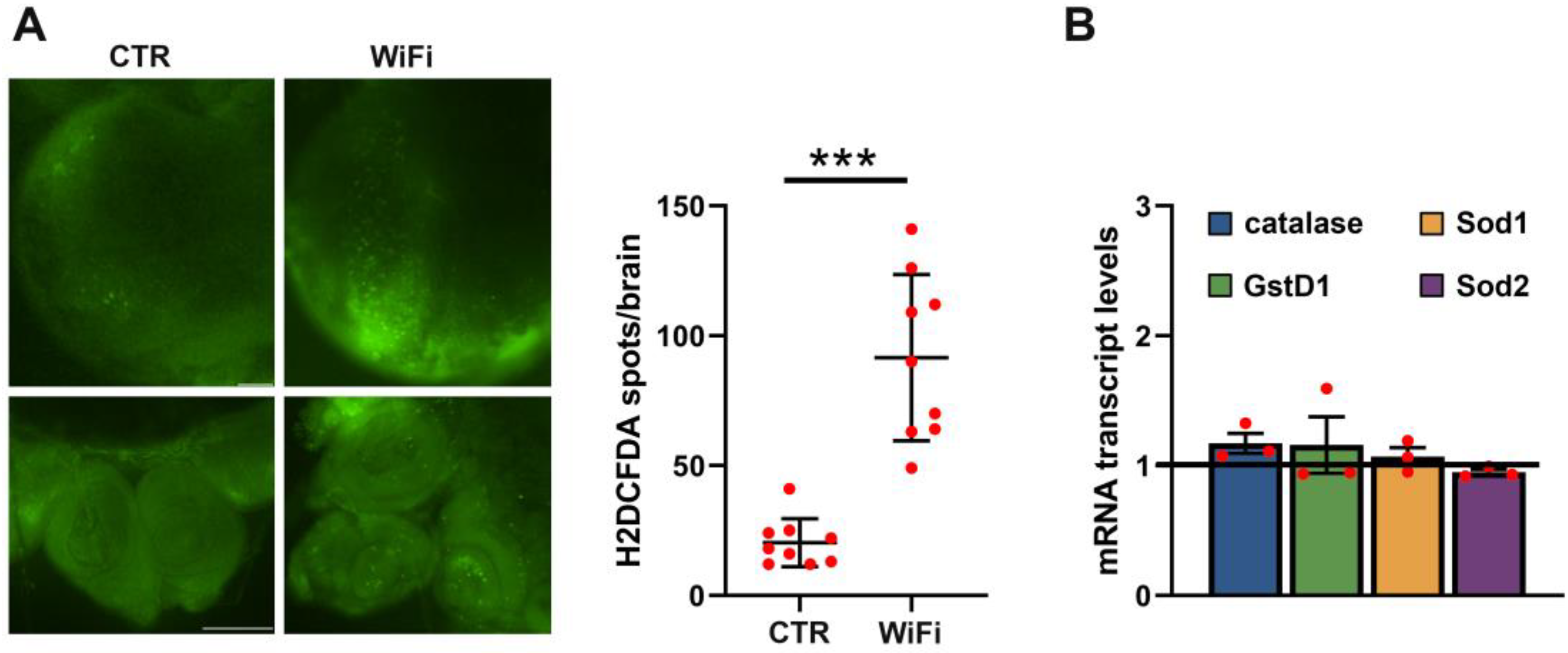
3.3. WiFi Radiation Induces Genome Instability in Larval Brains
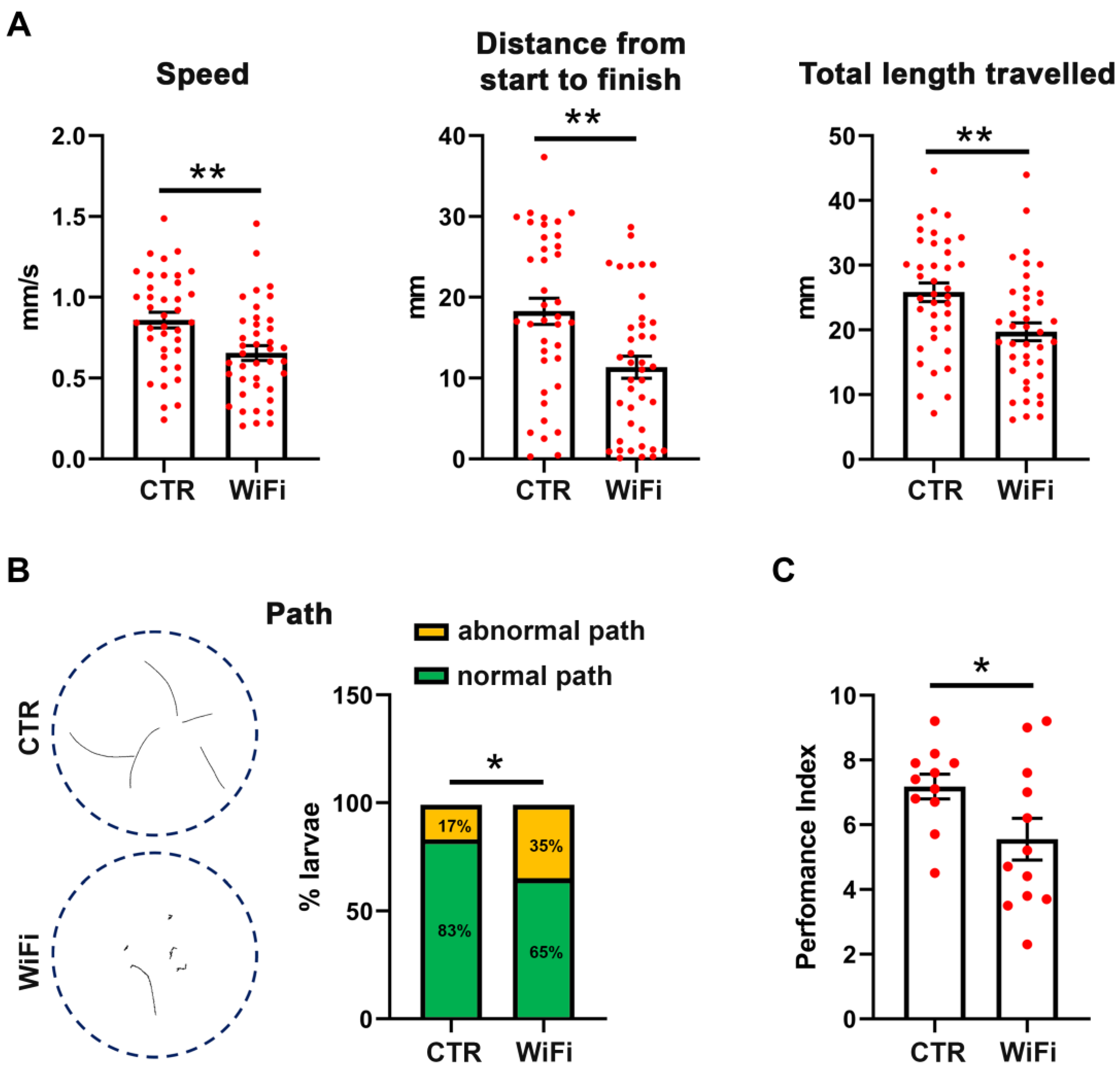
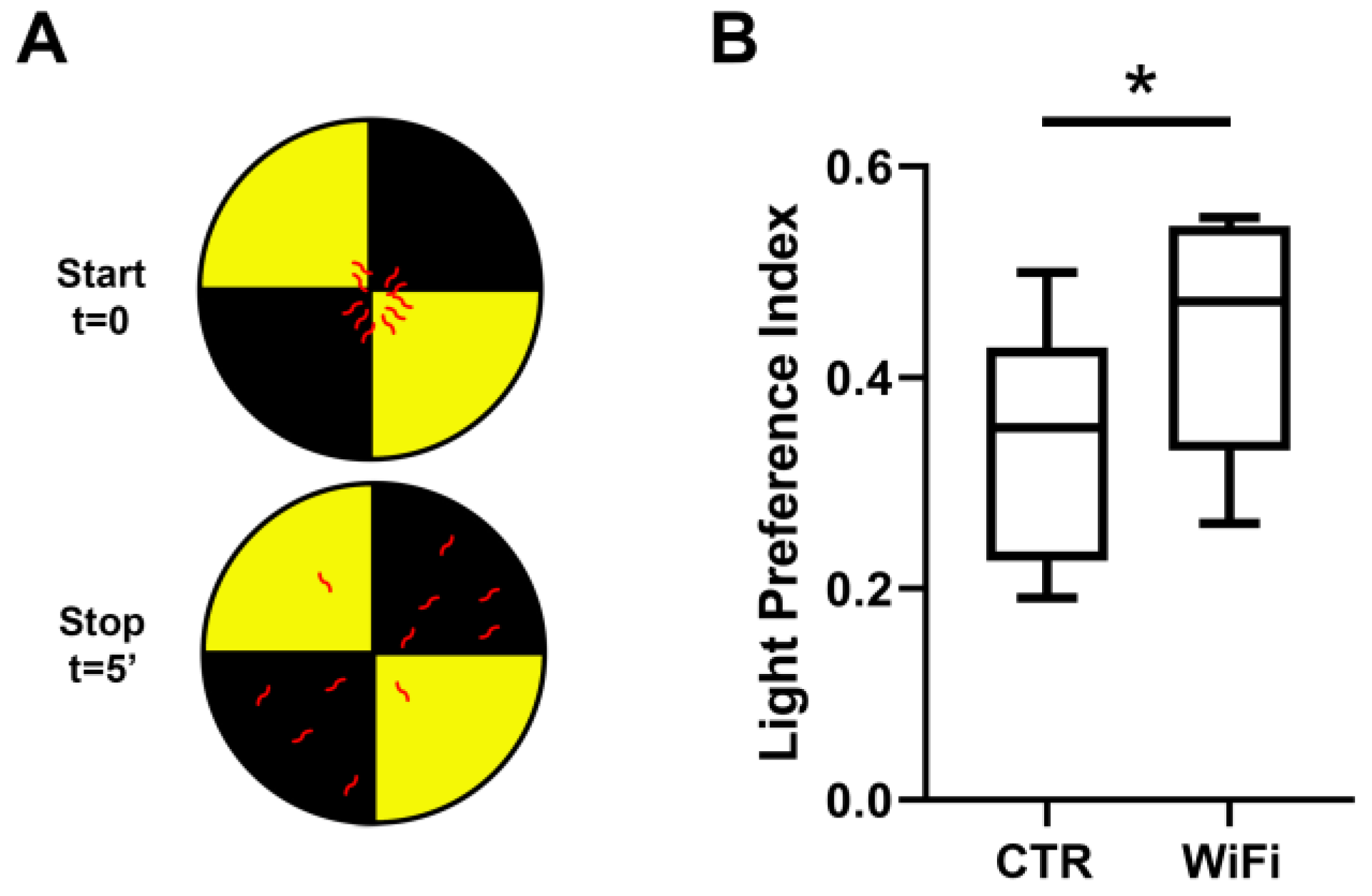
3.4. WiFi Exposure Impairs Locomotor Behaviour in Larvae and Adult Flies
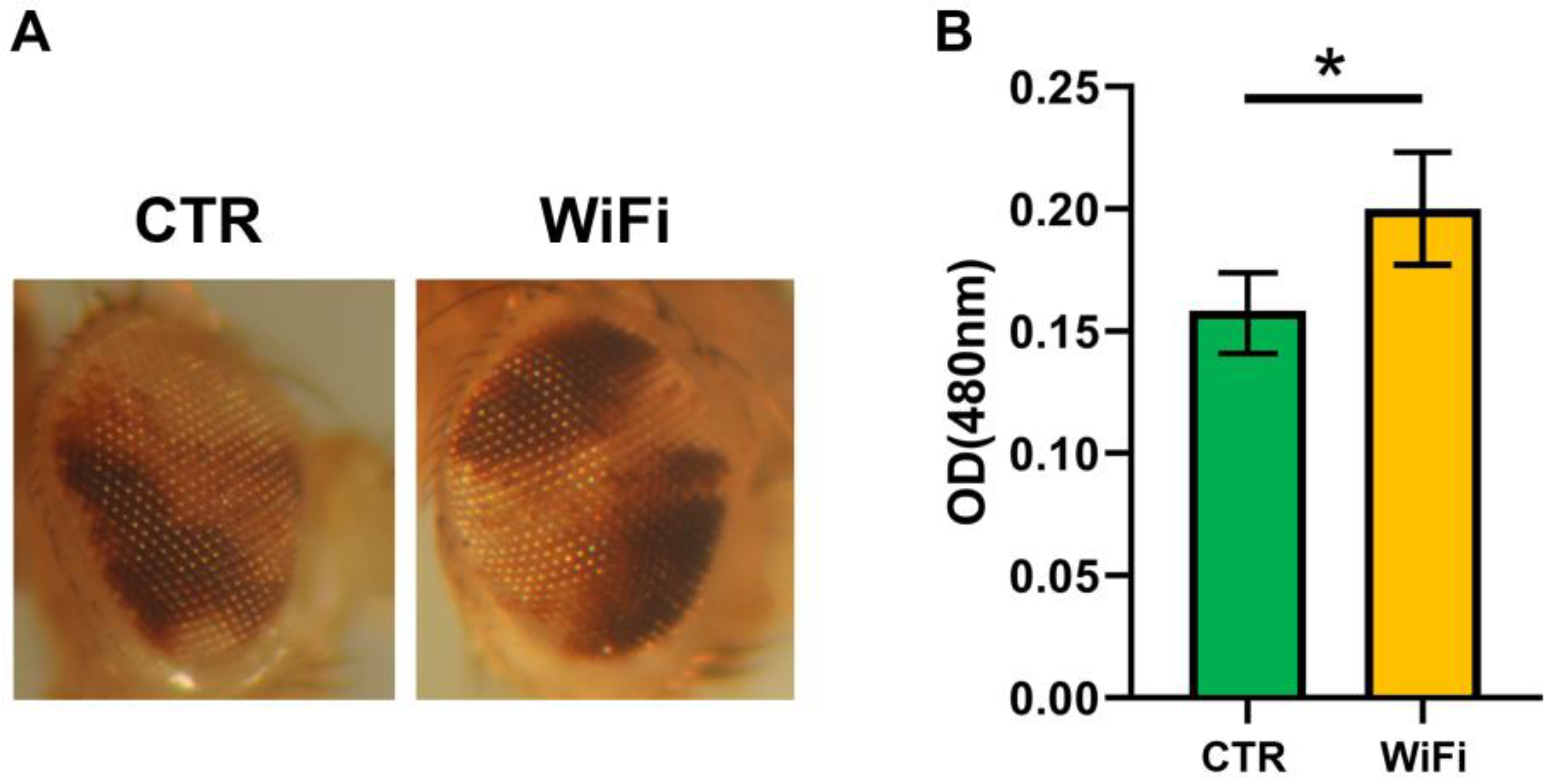
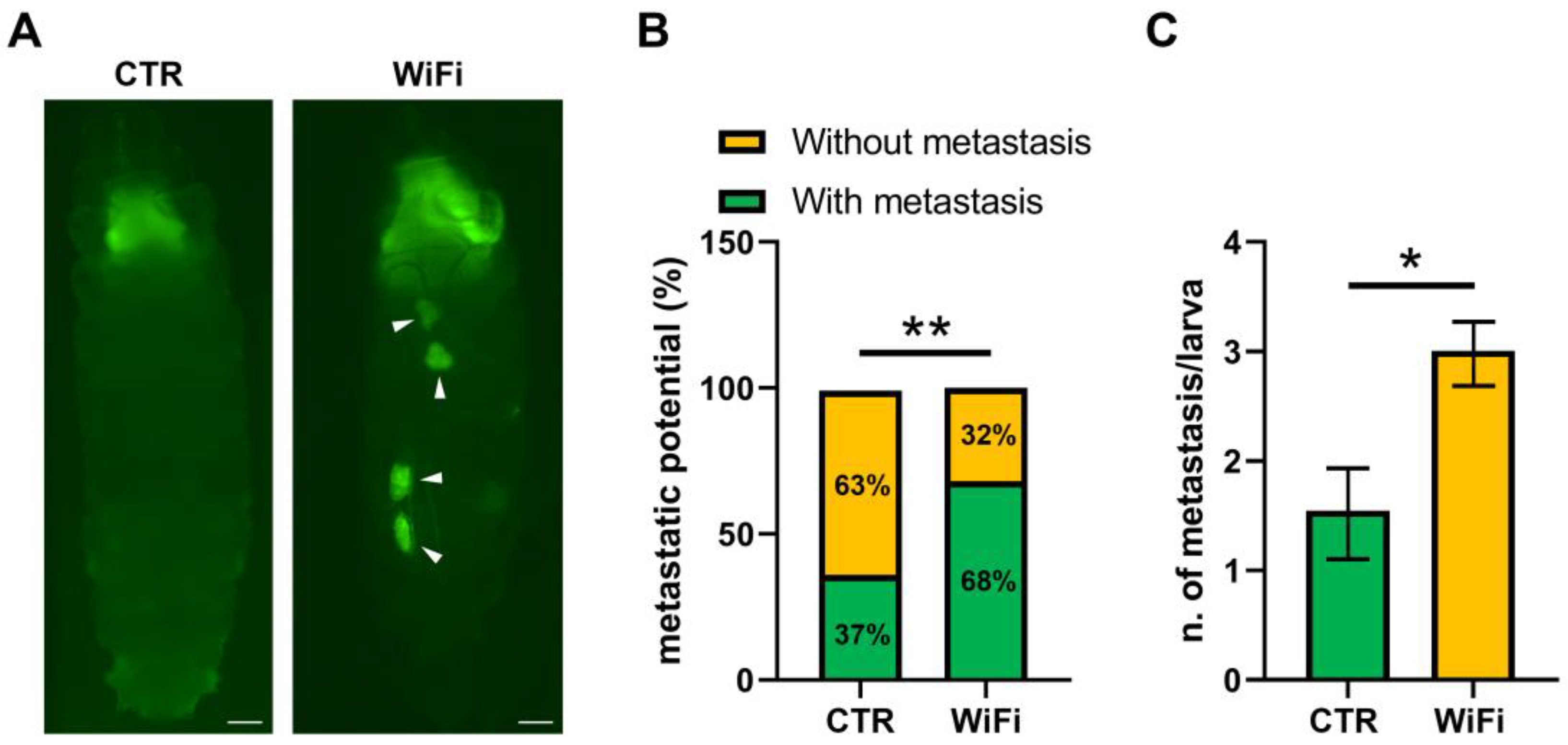
3.5. WiFi Radiation Promotes Metastatic Behaviour of Non-Invasive Eye Disc Clones Expressing the Activated Oncogene RasV12
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Belancio, V.P.; Hedges, D.J.; Deininger, P. Mammalian Non-LTR Retrotransposons: For Better or Worse, in Sickness and in Health. Genome Res. 2008, 18, 343–358. [Google Scholar] [CrossRef] [PubMed]
- Goodier, J.L.; Kazazian, H.H. Retrotransposons Revisited: The Restraint and Rehabilitation of Parasites. Cell 2008, 135, 23–35. [Google Scholar] [CrossRef]
- Finnegan, D.J. Eukaryotic Transposable Elements and Genome Evolution. Trends Genet. 1989, 5, 103–107. [Google Scholar] [CrossRef] [PubMed]
- Orgel, L.; Crick, F. Selfish DNA: The Ultimate Parasite. Nature 1980, 284, 604–607. [Google Scholar] [CrossRef] [PubMed]
- Piacentini, L.; Fanti, L.; Specchia, V.; Bozzetti, M.P.; Berloco, M.; Palumbo, G.; Pimpinelli, S. Transposons, Environmental Changes, and Heritable Induced Phenotypic Variability. Chromosoma 2014, 123, 345–354. [Google Scholar] [CrossRef] [PubMed]
- Fanti, L.; Piacentini, L.; Cappucci, U.; Casale, A.M.; Pimpinelli, S. Canalization by Selection of de Novo Induced Mutations. Genetics 2017, 206, 1995–2006. [Google Scholar] [CrossRef] [PubMed]
- Cappucci, U.; Noro, F.; Casale, A.M.; Fanti, L.; Berloco, M.; Alagia, A.A.; Grassi, L.; Le Pera, L.; Piacentini, L.; Pimpinelli, S. The Hsp70 Chaperone Is a Major Player in Stress-Induced Transposable Element Activation. Proc. Natl. Acad. Sci. USA 2019, 116, 17943–17950. [Google Scholar] [CrossRef]
- Friedli, M.; Trono, D. The Developmental Control of Transposable Elements and the Evolution of Higher Species. Annu. Rev. Cell Dev. Biol. 2015, 31, 429–451. [Google Scholar] [CrossRef]
- Hayward, A.; Gilbert, C. Transposable Elements. Curr. Biol. 2022, 32, R904–R909. [Google Scholar] [CrossRef]
- Feschotte, C.; Pritham, E.J. DNA Transposons and the Evolution of Eukaryotic Genomes. Annu. Rev. Genet. 2007, 41, 331–368. [Google Scholar] [CrossRef]
- Jurka, J.; Kapitonov, V.V.; Kohany, O.; Jurka, M.V. Repetitive Sequences in Complex Genomes: Structure and Evolution. Annu. Rev. Genom. Hum. Genet. 2007, 8, 241–259. [Google Scholar] [CrossRef] [PubMed]
- Naville, M.; Warren, I.A.; Haftek-Terreau, Z.; Chalopin, D.; Brunet, F.; Levin, P.; Galiana, D.; Volff, J.-N. Not so Bad after All: Retroviruses and Long Terminal Repeat Retrotransposons as a Source of New Genes in Vertebrates. Clin. Microbiol. Infect. 2016, 22, 312–323. [Google Scholar] [CrossRef] [PubMed]
- Joly-Lopez, Z.; Bureau, T.E. Exaptation of Transposable Element Coding Sequences. Curr. Opin. Genet. Dev. 2018, 49, 34–42. [Google Scholar] [CrossRef] [PubMed]
- Casacuberta, E.; González, J. The Impact of Transposable Elements in Environmental Adaptation. Mol. Ecol. 2013, 22, 1503–1517. [Google Scholar] [CrossRef] [PubMed]
- Horváth, V.; Merenciano, M.; González, J. Revisiting the Relationship between Transposable Elements and the Eukaryotic Stress Response. Trends Genet. 2017, 33, 832–841. [Google Scholar] [CrossRef]
- Pimpinelli, S.; Piacentini, L. Environmental Change and the Evolution of Genomes: Transposable Elements as Translators of Phenotypic Plasticity into Genotypic Variability. Funct. Ecol. 2019, 34, 428–441. [Google Scholar] [CrossRef]
- McClintock, B. The Significance of Responses of the Genome to Challenge. Science 1984, 226, 792–801. [Google Scholar] [CrossRef]
- Nätt, D.; Thorsell, A. Stress-Induced Transposon Reactivation: A Mediator or an Estimator of Allostatic Load? Environ. Epigenet. 2016, 2, dvw015. [Google Scholar] [CrossRef]
- Cappucci, U.; Torromino, G.; Casale, A.M.; Camon, J.; Capitano, F.; Berloco, M.; Mele, A.; Pimpinelli, S.; Rinaldi, A.; Piacentini, L. Stress-Induced Strain and Brain Region-Specific Activation of LINE-1 Transposons in Adult Mice. Stress 2018, 21, 575–579. [Google Scholar] [CrossRef]
- Wheeler, B.S. Small RNAs, Big Impact: Small RNA Pathways in Transposon Control and Their Effect on the Host Stress Response. Chromosome Res. 2013, 21, 587–600. [Google Scholar] [CrossRef]
- Kapusta, A.; Kronenberg, Z.; Lynch, V.J.; Zhuo, X.; Ramsay, L.; Bourque, G.; Yandell, M.; Feschotte, C. Transposable Elements Are Major Contributors to the Origin, Diversification, and Regulation of Vertebrate Long Noncoding RNAs. PLoS Genet. 2013, 9, e1003470. [Google Scholar] [CrossRef] [PubMed]
- Jönsson, M.E.; Garza, R.; Johansson, P.A.; Jakobsson, J. Transposable Elements: A Common Feature of Neurodevelopmental and Neurodegenerative Disorders. Trends Genet. 2020, 36, 610–623. [Google Scholar] [CrossRef] [PubMed]
- Ravel-Godreuil, C.; Znaidi, R.; Bonnifet, T.; Joshi, R.L.; Fuchs, J. Transposable Elements as New Players in Neurodegenerative Diseases. FEBS Lett. 2021, 595, 2733–2755. [Google Scholar] [CrossRef] [PubMed]
- Muotri, A.R.; Marchetto, M.C.N.; Coufal, N.G.; Oefner, R.; Yeo, G.; Nakashima, K.; Gage, F.H. L1 Retrotransposition in Neurons Is Modulated by MeCP2. Nature 2010, 468, 443–446. [Google Scholar] [CrossRef] [PubMed]
- Barlow, C.; Hirotsune, S.; Paylor, R.; Liyanage, M.; Eckhaus, M.; Collins, F.; Shiloh, Y.; Crawley, J.N.; Ried, T.; Tagle, D.; et al. Atm-Deficient Mice: A Paradigm of Ataxia Telangiectasia. Cell 1996, 86, 159–171. [Google Scholar] [CrossRef]
- Coufal, N.G.; Garcia-Perez, J.L.; Peng, G.E.; Marchetto, M.C.N.; Muotri, A.R.; Mu, Y.; Carson, C.T.; Macia, A.; Moran, J.V.; Gage, F.H. Ataxia Telangiectasia Mutated (ATM) Modulates Long Interspersed Element-1 (L1) Retrotransposition in Human Neural Stem Cells. Proc. Natl. Acad. Sci. USA 2011, 108, 20382–20387. [Google Scholar] [CrossRef]
- Kaneko, H.; Dridi, S.; Tarallo, V.; Gelfand, B.D.; Fowler, B.J.; Cho, W.G.; Kleinman, M.E.; Ponicsan, S.L.; Hauswirth, W.W.; Chiodo, V.A.; et al. DICER1 Deficit Induces Alu RNA Toxicity in Age-Related Macular Degeneration. Nature 2011, 471, 325–330. [Google Scholar] [CrossRef] [PubMed]
- Lathe, R.; Harris, A. Differential Display Detects Host Nucleic Acid Motifs Altered in Scrapie-Infected Brain. J. Mol. Biol. 2009, 392, 813–822. [Google Scholar] [CrossRef] [PubMed]
- Douville, R.; Liu, J.; Rothstein, J.; Nath, A. Identification of Active Loci of a Human Endogenous Retrovirus in Neurons of Patients with Amyotrophic Lateral Sclerosis. Ann. Neurol. 2011, 69, 141–151. [Google Scholar] [CrossRef]
- Krug, L.; Chatterjee, N.; Borges-Monroy, R.; Hearn, S.; Liao, W.-W.; Morrill, K.; Prazak, L.; Rozhkov, N.; Theodorou, D.; Hammell, M.; et al. Retrotransposon Activation Contributes to Neurodegeneration in a Drosophila TDP-43 Model of ALS. PLoS Genet. 2017, 13, e1006635. [Google Scholar] [CrossRef]
- Li, P.; Du, J.; Goodier, J.L.; Hou, J.; Kang, J.; Kazazian, H.H., Jr.; Zhao, K.; Yu, X.-F. Aicardi–Goutières Syndrome Protein TREX1 Suppresses L1 and Maintains Genome Integrity through Exonuclease-Independent ORF1p Depletion. Nucleic Acids Res. 2017, 45, 4619–4631. [Google Scholar] [CrossRef] [PubMed]
- Guo, C.; Jeong, H.H.; Hsieh, Y.C.; Klein, H.U.; Bennett, D.A.; De Jager, P.L.; Liu, Z.; Shulman, J.M. Tau Activates Transposable Elements in Alzheimer’s Disease. Cell Rep. 2018, 23, 2874–2880. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Samimi, H.; Gamez, M.; Zare, H.; Frost, B. Pathogenic Tau-Induced PiRNA Depletion Promotes Neuronal Death through Transposable Element Dysregulation in Neurodegenerative Tauopathies. Nat. Neurosci. 2018, 21, 1038–1048. [Google Scholar] [CrossRef]
- Maggiore, A.; Casale, A.M.; Toscanelli, W.; Cappucci, U.; Rotili, D.; Grieco, M.; Gagné, J.P.; Poirier, G.G.; d’Erme, M.; Piacentini, L. Neuroprotective Effects of PARP Inhibitors in Drosophila Models of Alzheimer’s Disease. Cells 2022, 11, 1284. [Google Scholar] [CrossRef] [PubMed]
- Casale, A.M.; Liguori, F.; Ansaloni, F.; Cappucci, U.; Finaurini, S.; Spirito, G.; Persichetti, F.; Sanges, R.; Gustincich, S.; Piacentini, L. Transposable Element Activation Promotes Neurodegeneration in a Drosophila Model of Huntington’s Disease. iScience 2022, 25, 103702. [Google Scholar] [CrossRef]
- Bundo, M.; Toyoshima, M.; Okada, Y.; Akamatsu, W.; Ueda, J.; Nemoto-Miyauchi, T.; Sunaga, F.; Toritsuka, M.; Ikawa, D.; Kakita, A.; et al. Increased L1 Retrotransposition in the Neuronal Genome in Schizophrenia. Neuron 2014, 81, 306–313. [Google Scholar] [CrossRef]
- Atasoy, H.I.; Gunal, M.Y.; Atasoy, P.; Elgun, S.; Bugdayci, G. Immunohistopathologic Demonstration of Deleterious Effects on Growing Rat Testes of Radiofrequency Waves Emitted from Conventional Wi-Fi Devices. J. Pediatr. Urol. 2013, 9, 223–229. [Google Scholar] [CrossRef]
- Özorak, A.; Nazıroğlu, M.; Çelik, Ö.; Yüksel, M.; Özçelik, D.; Özkaya, M.O.; Çetin, H.; Kahya, M.C.; Kose, S.A. Wi-Fi (2.45 GHz)- and Mobile Phone (900 and 1800 MHz)-Induced Risks on Oxidative Stress and Elements in Kidney and Testis of Rats During Pregnancy and the Development of Offspring. Biol. Trace Elem. Res. 2013, 156, 221–229. [Google Scholar] [CrossRef]
- Aynali, G.; Nazıroğlu, M.; Çelik, Ö.; Doğan, M.; Yarıktaş, M.; Yasan, H. Modulation of Wireless (2.45 GHz)-Induced Oxidative Toxicity in Laryngotracheal Mucosa of Rat by Melatonin. Eur. Arch. Oto-Rhino-Laryngol. 2013, 270, 1695–1700. [Google Scholar] [CrossRef]
- Çiftçi, Z.Z.; Kırzıoğlu, Z.; Nazıroğlu, M.; Özmen, Ö. Effects of Prenatal and Postnatal Exposure of Wi-Fi on Development of Teeth and Changes in Teeth Element Concentration in Rats. Biol. Trace Elem. Res. 2015, 163, 193–201. [Google Scholar] [CrossRef]
- Tok, L.; Nazirogiu, M.; Doganan, S.; Kahya, M.; Tok, O. Effects of Melatonin on Wi-Fi-Induced Oxidative Stress in Lens of Rats. Indian J. Ophthalmol. 2014, 62, 12–15. [Google Scholar] [CrossRef] [PubMed]
- Çiğ, B.; Nazıroğlu, M. Investigation of the Effects of Distance from Sources on Apoptosis, Oxidative Stress and Cytosolic Calcium Accumulation via TRPV1 Channels Induced by Mobile Phones and Wi-Fi in Breast Cancer Cells. Biochim. Biophys. Acta Biomembr. 2015, 1848, 2756–2765. [Google Scholar] [CrossRef] [PubMed]
- Ghazizadeh, V.; Nazıroğlu, M. Electromagnetic Radiation (Wi-Fi) and Epilepsy Induce Calcium Entry and Apoptosis through Activation of TRPV1 Channel in Hippocampus and Dorsal Root Ganglion of Rats. Metab. Brain Dis. 2014, 29, 787–799. [Google Scholar] [CrossRef]
- Yüksel, M.; Nazıroğlu, M.; Özkaya, M.O. Long-Term Exposure to Electromagnetic Radiation from Mobile Phones and Wi-Fi Devices Decreases Plasma Prolactin, Progesterone, and Estrogen Levels but Increases Uterine Oxidative Stress in Pregnant Rats and Their Offspring. Endocrine 2016, 52, 352–362. [Google Scholar] [CrossRef] [PubMed]
- Topsakal, S.; Ozmen, O.; Cicek, E.; Comlekci, S. The Ameliorative Effect of Gallic Acid on Pancreas Lesions Induced by 2.45 GHz Electromagnetic Radiation (Wi-Fi) in Young Rats. J. Radiat. Res. Appl. Sci. 2017, 10, 233–240. [Google Scholar] [CrossRef]
- Othman, H.; Ammari, M.; Sakly, M.; Abdelmelek, H. Effects of Repeated Restraint Stress and WiFi Signal Exposure on Behavior and Oxidative Stress in Rats. Metab. Brain Dis. 2017, 32, 1459–1469. [Google Scholar] [CrossRef]
- Shokri, S.; Soltani, A.; Kazemi, M.; Sardari, D.; Mofrad, F.B. Effects of Wi-Fi (2.45 GHz) Exposure on Apoptosis, Sperm Parameters and Testicular Histomorphometry in Rats: A Time Course Study. Cell J. 2015, 17, 322–331. [Google Scholar] [CrossRef]
- Dasdag, S.; Taş, M.; Akdag, M.Z.; Yegin, K. Effect of Long-Term Exposure of 2.4 GHz Radiofrequency Radiation Emitted from Wi-Fi Equipment on Testes Functions. Electromagn. Biol. Med. 2015, 34, 37–42. [Google Scholar] [CrossRef]
- Avendaño, C.; Mata, A.; Sanchez Sarmiento, C.A.; Doncel, G.F. Use of Laptop Computers Connected to Internet through Wi-Fi Decreases Human Sperm Motility and Increases Sperm DNA Fragmentation. Fertil. Steril. 2012, 97, 39–45.e2. [Google Scholar] [CrossRef]
- Yildirim, M.E.; Kaynar, M.; Badem, H.; Cavis, M.; Karatas, O.F.; Cimentepe, E. What Is Harmful for Male Fertility: Cell Phone or the Wireless Internet? Kaohsiung J. Med. Sci. 2015, 31, 480–484. [Google Scholar] [CrossRef]
- Oni, O.M.; Amunda, D.B. Effects of Radiofrequency Radiation from WiFi Devices on Human Ejaculated Semen. Int. J. Res. Rev. Appl. Sci. 2011, 9, 292–294. [Google Scholar]
- Akdag, M.Z.; Dasdag, S.; Canturk, F.; Karabulut, D.; Caner, Y.; Adalier, N. Does Prolonged Radiofrequency Radiation Emitted from Wi-Fi Devices Induce DNA Damage in Various Tissues of Rats? J. Chem. Neuroanat. 2016, 75, 116–122. [Google Scholar] [CrossRef] [PubMed]
- Šimaiová, V.; Almášiová, V.; Holovská, K.; Kisková, T.; Horváthová, F.; Ševčíková, Z.; Tóth, Š.; Raček, A.; Račeková, E.; Beňová, K.; et al. The Effect of 2.45 GHz Non-Ionizing Radiation on the Structure and Ultrastructure of the Testis in Juvenile Rats. Histol. Histopathol. 2019, 34, 391–403. [Google Scholar] [CrossRef] [PubMed]
- Jaffar, F.H.F.; Osman, K.; Ismail, N.H.; Chin, K.-Y.; Ibrahim, S.F. Adverse Effects of Wi-Fi Radiation on Male Reproductive System: A Systematic Review. Tohoku J. Exp. Med. 2019, 248, 169–179. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; She, F.; Li, L.; Chen, C.; Xu, S.; Luo, X.; Li, M.; He, M.; Yu, Z. P25/CDK5 Is Partially Involved in Neuronal Injury Induced by Radiofrequency Electromagnetic Field Exposure. Int. J. Radiat. Biol. 2013, 89, 976–984. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Huang, W.-J.; Chen, W.-W. Microwaves and Alzheimer’s Disease (Review). Exp. Ther. Med. 2016, 12, 1969–1972. [Google Scholar] [CrossRef]
- Othman, H.; Ammari, M.; Rtibi, K.; Bensaid, N.; Sakly, M.; Abdelmelek, H. Postnatal Development and Behavior Effects of In-Utero Exposure of Rats to Radiofrequency Waves Emitted from Conventional WiFi Devices. Environ. Toxicol. Pharmacol. 2017, 52, 239–247. [Google Scholar] [CrossRef]
- Saili, L.; Hanini, A.; Smirani, C.; Azzouz, I.; Azzouz, A.; Sakly, M.; Abdelmelek, H.; Bouslama, Z. Effects of Acute Exposure to WIFI Signals (2.45GHz) on Heart Variability and Blood Pressure in Albinos Rabbit. Environ. Toxicol. Pharmacol. 2015, 40, 600–605. [Google Scholar] [CrossRef]
- Pall, M.L. Wi-Fi Is an Important Threat to Human Health. Environ. Res. 2018, 164, 405–416. [Google Scholar] [CrossRef]
- Pagliarini, R.A.; Xu, T. A Genetic Screen in Drosophila for Metastatic Behavior. Science 2003, 302, 1227–1231. [Google Scholar] [CrossRef]
- Crawford, M.L. Generation of Standard EM Fields Using TEM Transmission Cells. IEEE Trans. Electromagn. Compat. 1974, EMC-16, 189–195. [Google Scholar] [CrossRef]
- Stuchly, M.S.; Stuchly, S.S. Experimental Ratio and Microwave Dosimetry. In Handbook of Biological Effects of Electromagnetic Fields; CRC Press: Boca Raton, FL, USA, 1996; pp. 295–336. [Google Scholar]
- Grudzinski, E.; Trzaska, H. TEM Cell as the EMF Standard and Exposure System. In Proceedings of the Modern Problems of Radio Engineering, Telecommunications and Computer Science (IEEE Cat. No.02EX542), Lviv-Slavsko, Ukraine, 18–23 February 2002; pp. 310–312. [Google Scholar]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Pimpinelli, S.; Bonaccorsi, S.; Fanti, L.; Gatti, M. Preparation and Analysis of Drosophila Mitotic Chromosomes. In Drosophila Protocols; Sullivan, W., Ashburner, M., Hawley, S., Eds.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2000. [Google Scholar]
- Owusu-Ansah, E.; Yavari, A.; Banerjee, U. A Protocol for in Vivo Detection of Reactive Oxygen Species. Protoc. Exch. 2008. [Google Scholar] [CrossRef]
- Nichols, C.D.; Becnel, J.; Pandey, U.B. Methods to Assay Drosophila Behavior. JoVE 2012, e3795. [Google Scholar] [CrossRef]
- Brooks, D.S.; Vishal, K.; Kawakami, J.; Bouyain, S.; Geisbrecht, E.R. Optimization of WrMTrck to Monitor Drosophila Larval Locomotor Activity. J. Insect Physiol. 2016, 93–94, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Farca Luna, A.J.; von Essen, A.M.H.J.; Widmer, Y.F.; Sprecher, S.G. Light Preference Assay to Study Innate and Circadian Regulated Photobehavior in Drosophila Larvae. JoVE 2013, e50237. [Google Scholar] [CrossRef]
- Laneve, P.; Piacentini, L.; Casale, A.M.; Capauto, D.; Gioia, U.; Cappucci, U.; Di Carlo, V.; Bozzoni, I.; Di Micco, P.; Morea, V.; et al. Drosophila CG3303 Is an Essential Endoribonuclease Linked to TDP-43-Mediated Neurodegeneration. Sci. Rep. 2017, 7, 41559. [Google Scholar] [CrossRef]
- Fedoroff, N. V Transposable Elements, Epigenetics, and Genome Evolution. Science 2012, 338, 758–767. [Google Scholar] [CrossRef]
- Elgin, S.C.R.; Reuter, G. Position-Effect Variegation, Heterochromatin Formation, and Gene Silencing in Drosophila. Cold Spring Harb. Perspect. Biol. 2013, 5, a017780. [Google Scholar] [CrossRef]
- Muller, H.J. Types of Visible Variations Induced by X-Rays InDrosophila. J. Genet. 1930, 22, 299–334. [Google Scholar] [CrossRef]
- Tartof, K.D.; Hobbs, C.; Jones, M. A Structural Basis for Variegating Position Effects. Cell 1984, 37, 869–878. [Google Scholar] [CrossRef]
- Lu, B.Y.; Bishop, C.P.; Eissenberg, J.C. Developmental Timing and Tissue Specificity of Heterochromatin-Mediated Silencing. EMBO J. 1996, 15, 1323–1332. [Google Scholar] [CrossRef] [PubMed]
- Schotta, G.; Ebert, A.; Krauss, V.; Fischer, A.; Hoffmann, J.; Rea, S.; Jenuwein, T.; Dorn, R.; Reuter, G. Central Role of Drosophila SU(VAR)3-9 in Histone H3-K9 Methylation and Heterochromatic Gene Silencing. EMBO J. 2002, 21, 1121–1131. [Google Scholar] [CrossRef] [PubMed]
- Nakayama, J.; Rice, J.C.; Strahl, B.D.; Allis, C.D.; Grewal, S.I.S. Role of Histone H3 Lysine 9 Methylation in Epigenetic Control of Heterochromatin Assembly. Science 2001, 292, 110–113. [Google Scholar] [CrossRef]
- Casale, A.M.; Cappucci, U.; Fanti, L.; Piacentini, L. Heterochromatin Protein 1 (HP1) Is Intrinsically Required for Post-Transcriptional Regulation of Drosophila Germline Stem Cell (GSC) Maintenance. Sci. Rep. 2019, 9, 4372. [Google Scholar] [CrossRef] [PubMed]
- Janssen, A.; Colmenares, S.U.; Karpen, G.H. Heterochromatin: Guardian of the Genome. Annu. Rev. Cell Dev. Biol. 2018, 34, 265–288. [Google Scholar] [CrossRef] [PubMed]
- Price, B.D.; D’Andrea, A.D. Chromatin Remodeling at DNA Double-Strand Breaks. Cell 2013, 152, 1344–1354. [Google Scholar] [CrossRef]
- Ayarpadikannan, S.; Kim, H.-S. The Impact of Transposable Elements in Genome Evolution and Genetic Instability and Their Implications in Various Diseases. Genom. Inform. 2014, 12, 98–104. [Google Scholar] [CrossRef]
- Belgnaoui, S.M.; Gosden, R.G.; Semmes, O.J.; Haoudi, A. Human LINE-1 Retrotransposon Induces DNA Damage and Apoptosis in Cancer Cells. Cancer Cell Int. 2006, 6, 13. [Google Scholar] [CrossRef]
- Gasior, S.L.; Wakeman, T.P.; Xu, B.; Deininger, P.L. The Human LINE-1 Retrotransposon Creates DNA Double-Strand Breaks. J. Mol. Biol. 2006, 357, 1383–1393. [Google Scholar] [CrossRef]
- Bhat, A.; Ghatage, T.; Bhan, S.; Lahane, G.P.; Dhar, A.; Kumar, R.; Pandita, R.K.; Bhat, K.M.; Ramos, K.S.; Pandita, T.K. Role of Transposable Elements in Genome Stability: Implications for Health and Disease. Int. J. Mol. Sci. 2022, 23, 7802. [Google Scholar] [CrossRef]
- Allshire, R.C.; Nimmo, E.R.; Ekwall, K.; Javerzat, J.P.; Cranston, G. Mutations Derepressing Silent Centromeric Domains in Fission Yeast Disrupt Chromosome Segregation. Genes Dev. 1995, 9, 218–233. [Google Scholar] [CrossRef] [PubMed]
- Kellum, R.; Alberts, B.M. Heterochromatin Protein 1 Is Required for Correct Chromosome Segregation in Drosophila Embryos. J. Cell Sci. 1995, 108, 1419–1431. [Google Scholar] [CrossRef] [PubMed]
- Ekwall, K.; Nimmo, E.R.; Javerzat, J.P.; Borgstrom, B.; Egel, R.; Cranston, G.; Allshire, R. Mutations in the Fission Yeast Silencing Factors Clr4+ and Rik1+ Disrupt the Localisation of the Chromo Domain Protein Swi6p and Impair Centromere Function. J. Cell Sci. 1996, 109, 2637–2648. [Google Scholar] [CrossRef] [PubMed]
- Ekwall, K.; Cranston, G.; Allshire, R.C. Fission Yeast Mutants That Alleviate Transcriptional Silencing in Centromeric Flanking Repeats and Disrupt Chromosome Segregation. Genetics 1999, 153, 1153–1169. [Google Scholar] [CrossRef]
- Peters, A.H.F.M.; O’Carroll, D.; Scherthan, H.; Mechtler, K.; Sauer, S.; Schöfer, C.; Weipoltshammer, K.; Pagani, M.; Lachner, M.; Kohlmaier, A.; et al. Loss of the Suv39h Histone Methyltransferases Impairs Mammalian Heterochromatin and Genome Stability. Cell 2001, 107, 323–337. [Google Scholar] [CrossRef]
- Nonaka, N.; Kitajima, T.; Yokobayashi, S.; Xiao, G.; Yamamoto, M.; Grewal, S.I.S.; Watanabe, Y. Recruitment of Cohesin to Heterochromatic Regions by Swi6/HP1 in Fission Yeast. Nat. Cell Biol. 2002, 4, 89–93. [Google Scholar] [CrossRef]
- Pidoux, A.L.; Allshire, R.C. Kinetochore and Heterochromatin Domains of the Fission Yeast Centromere. Chromosom Res. 2004, 12, 521–534. [Google Scholar] [CrossRef]
- Inoue, A.; Hyle, J.; Lechner, M.S.; Lahti, J.M. Perturbation of HP1 Localization and Chromatin Binding Ability Causes Defects in Sister-Chromatid Cohesion. Mutat. Res. Toxicol. Environ. Mutagen. 2008, 657, 48–55. [Google Scholar] [CrossRef]
- Yamagishi, Y.; Sakuno, T.; Shimura, M.; Watanabe, Y. Heterochromatin Links to Centromeric Protection by Recruiting Shugoshin. Nature 2008, 455, 251–255. [Google Scholar] [CrossRef]
- Shimura, M.; Toyoda, Y.; Iijima, K.; Kinomoto, M.; Tokunaga, K.; Yoda, K.; Yanagida, M.; Sata, T.; Ishizaka, Y. Epigenetic Displacement of HP1 from Heterochromatin by HIV-1 Vpr Causes Premature Sister Chromatid Separation. J. Cell Biol. 2011, 194, 721–735. [Google Scholar] [CrossRef]
- Frost, B.; Hemberg, M.; Lewis, J.; Feany, M.B. Tau Promotes Neurodegeneration through Global Chromatin Relaxation. Nat. Neurosci. 2014, 17, 357–366. [Google Scholar] [CrossRef] [PubMed]
- Perkins, A.T.; Greig, M.M.; Sontakke, A.A.; Peloquin, A.S.; McPeek, M.A.; Bickel, S.E. Increased Levels of Superoxide Dismutase Suppress Meiotic Segregation Errors in Aging Oocytes. Chromosoma 2019, 128, 215–222. [Google Scholar] [CrossRef] [PubMed]
- Kreuz, S.; Fischle, W. Oxidative Stress Signaling to Chromatin in Health and Disease. Epigenomics 2016, 8, 843–862. [Google Scholar] [CrossRef] [PubMed]
- Welch, G.; Tsai, L.-H. Mechanisms of DNA Damage-Mediated Neurotoxicity in Neurodegenerative Disease. EMBO Rep. 2022, 23, e54217. [Google Scholar] [CrossRef]
- Jakubowski, B.R.; Longoria, R.A.; Shubeita, G.T. A High Throughput and Sensitive Method Correlates Neuronal Disorder Genotypes to Drosophila Larvae Crawling Phenotypes. Fly (Austin) 2012, 6, 303–308. [Google Scholar] [CrossRef]
- Gegear, R.J.; Casselman, A.; Waddell, S.; Reppert, S.M. Cryptochrome Mediates Light-Dependent Magnetosensitivity in Drosophila. Nature 2008, 454, 1014–1018. [Google Scholar] [CrossRef]
- Yoshii, T.; Ahmad, M.; Helfrich-Förster, C. Cryptochrome Mediates Light-Dependent Magnetosensitivity of Drosophila’s Circadian Clock. PLoS Biol. 2009, 7, e1000086. [Google Scholar] [CrossRef]
- Fedele, G.; Green, E.W.; Rosato, E.; Kyriacou, C.P. An Electromagnetic Field Disrupts Negative Geotaxis in Drosophila via a CRY-Dependent Pathway. Nat. Commun. 2014, 5, 4391. [Google Scholar] [CrossRef]
- Burns, K.H. Transposable Elements in Cancer. Nat. Rev. Cancer 2017, 17, 415–424. [Google Scholar] [CrossRef]
- Ouadah, N.S.; Lecomte, A.; Robidel, F.; Olsson, A.; Deltour, I.; Schüz, J.; Blazy, K.; Villégier, A.-S. Possible Effects of Radiofrequency Electromagnetic Fields on in Vivo C6 Brain Tumors in Wistar Rats. J. Neurooncol. 2018, 140, 539–546. [Google Scholar] [CrossRef]
- Hardell, L.; Carlberg, M.; Hansson Mild, K. Use of Mobile Phones and Cordless Phones Is Associated with Increased Risk for Glioma and Acoustic Neuroma. Pathophysiology 2013, 20, 85–110. [Google Scholar] [CrossRef] [PubMed]
- Mirzoyan, Z.; Sollazzo, M.; Allocca, M.; Valenza, A.M.; Grifoni, D.; Bellosta, P. Drosophila Melanogaster: A Model Organism to Study Cancer. Front. Genet. 2019, 10, 51. [Google Scholar] [CrossRef] [PubMed]
- Saavedra, P.; Perrimon, N. Drosophila as a Model for Tumor-Induced Organ Wasting BT—The Drosophila Model in Cancer; Deng, W.-M., Ed.; Springer International Publishing: Cham, Switzerland, 2019; pp. 191–205. ISBN 978-3-030-23629-8. [Google Scholar]
- Richardson, H.E.; Cordero, J.B.; Grifoni, D. Basic and Translational Models of Cooperative Oncogenesis. Int. J. Mol. Sci. 2020, 21, 5919. [Google Scholar] [CrossRef] [PubMed]
- Miles, W.O.; Dyson, N.J.; Walker, J.A. Modeling Tumor Invasion and Metastasis in Drosophila. Dis. Model. Mech. 2011, 4, 753–761. [Google Scholar] [CrossRef]
- Enomoto, M.; Siow, C.; Igaki, T. Drosophila as a Cancer Model BT—Drosophila Models for Human Diseases; Yamaguchi, M., Ed.; Springer: Singapore, 2018; pp. 173–194. ISBN 978-981-13-0529-0. [Google Scholar]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cappucci, U.; Casale, A.M.; Proietti, M.; Marinelli, F.; Giuliani, L.; Piacentini, L. WiFi Related Radiofrequency Electromagnetic Fields Promote Transposable Element Dysregulation and Genomic Instability in Drosophila melanogaster. Cells 2022, 11, 4036. https://doi.org/10.3390/cells11244036
Cappucci U, Casale AM, Proietti M, Marinelli F, Giuliani L, Piacentini L. WiFi Related Radiofrequency Electromagnetic Fields Promote Transposable Element Dysregulation and Genomic Instability in Drosophila melanogaster. Cells. 2022; 11(24):4036. https://doi.org/10.3390/cells11244036
Chicago/Turabian StyleCappucci, Ugo, Assunta Maria Casale, Mirena Proietti, Fiorenzo Marinelli, Livio Giuliani, and Lucia Piacentini. 2022. "WiFi Related Radiofrequency Electromagnetic Fields Promote Transposable Element Dysregulation and Genomic Instability in Drosophila melanogaster" Cells 11, no. 24: 4036. https://doi.org/10.3390/cells11244036
APA StyleCappucci, U., Casale, A. M., Proietti, M., Marinelli, F., Giuliani, L., & Piacentini, L. (2022). WiFi Related Radiofrequency Electromagnetic Fields Promote Transposable Element Dysregulation and Genomic Instability in Drosophila melanogaster. Cells, 11(24), 4036. https://doi.org/10.3390/cells11244036






