Method for the Isolation of “RNA-seq-Quality” RNA from Human Intervertebral Discs after Mortar and Pestle Homogenization
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Intervertebral Disc Sampling
2.3. Mortar and Pestle Homogenization
Grinding Technique
2.4. RNA Isolation
2.5. Sequencing Libraries Preparation
2.6. Quality Control of Sequencing Results
2.7. Statistical Analysis
3. Results
3.1. Sampling and Homogenization
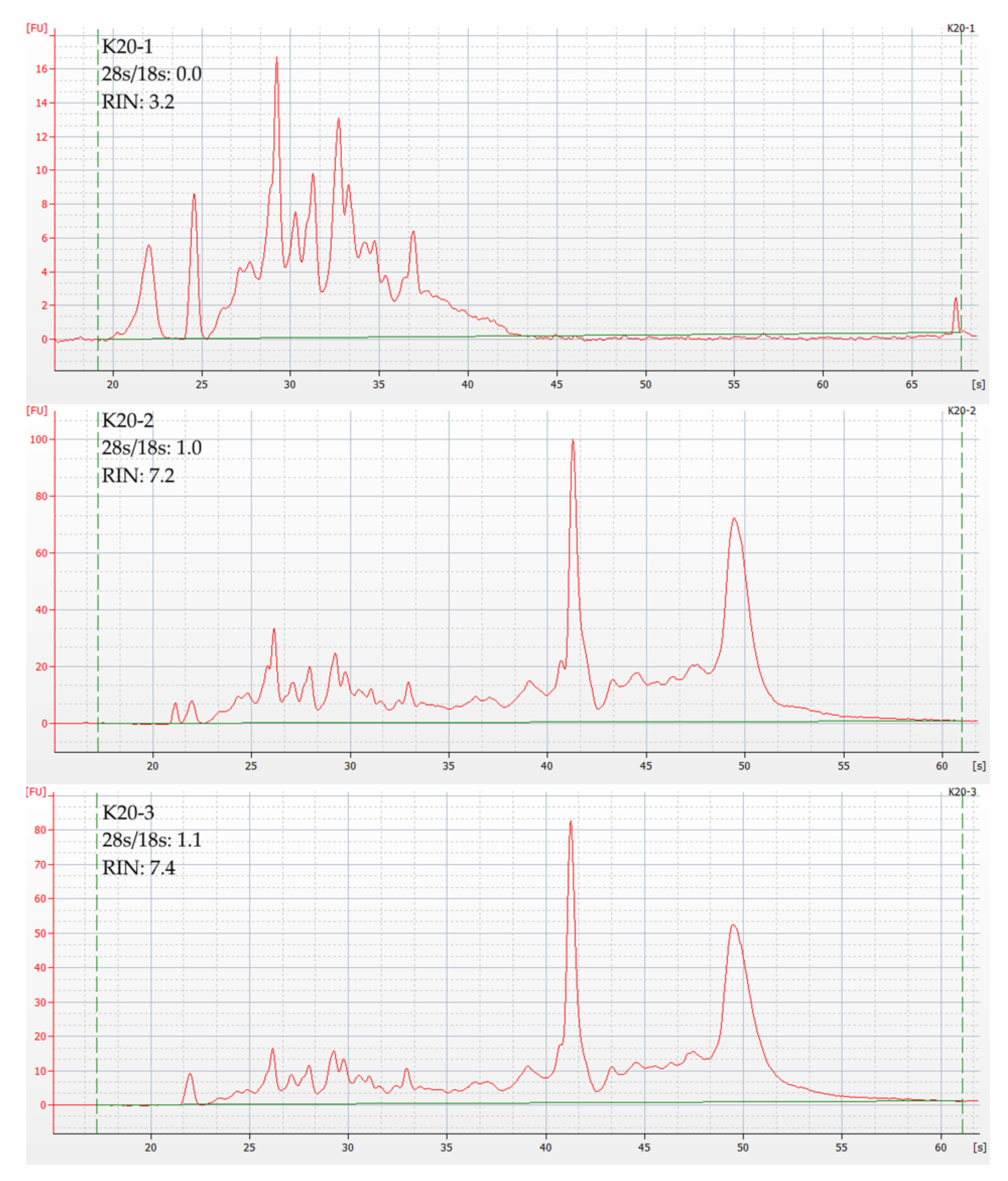
3.2. RNA Quality Control
3.3. Quality Control of RNA-Sequencing Results
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ozsolak, F.; Milos, P.M. RNA sequencing: Advances, challenges and opportunities. Nat. Rev. Genet. 2010, 12, 87–98. [Google Scholar] [CrossRef] [PubMed]
- Ruettger, A.; Neumann, S.; Wiederanders, B.; Huber, R. Comparison of different methods for preparation and characterization of total RNA from cartilage samples to uncover osteoarthritis in vivo. BMC Res. Notes 2010, 3, 7. [Google Scholar] [CrossRef] [PubMed]
- Gan, M.F.; Yang, H.; Qian, J.; Wu, C.; Yuan, C.; Li, X.; Zou, J. Comparison of two methods for RNA extraction from the nucleus pulposus of intervertebral discs. Genet. Mol. Res. 2016, 15, gmr7738. [Google Scholar] [CrossRef] [PubMed]
- le Bleu, H.K.; Kamal, F.A.; Kelly, M.; Ketz, J.P.; Zuscik, M.J.; Elbarbary, A.R. Extraction of high-quality RNA from human articular cartilage. Anal. Biochem. 2017, 518, 134–138. [Google Scholar] [CrossRef] [PubMed]
- Erwin, W.M.; Hood, K.E. The cellular and molecular biology of the intervertebral disc: A clinician’s primer. J. Can. Chiropr. Assoc. 2014, 58, 257. [Google Scholar]
- Dagenais, S.; Caro, J.; Haldeman, S. A systematic review of low back pain cost of illness studies in the United States and internationally. Spine J. 2008, 8, 8–20. [Google Scholar] [CrossRef] [PubMed]
- Xin, J.; Wang, Y.; Zheng, Z.; Wang, S.; Na, S.; Zhang, S. Treatment of Intervertebral Disc Degeneration. Orthop. Surg. 2022, 14, 1271–1280. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Li, Y.; Ye, Z.; Cheng, Z.; Huang, J.; Lu, S.; Su, K.; Liang, Y.; Li, M.; Huang, L. Prediction of a Potential Mechanism of Intervertebral Disc Degeneration Based on a Novel Competitive Endogenous RNA Network. BioMed Res. Int. 2021, 2021, 6618834. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.; Mu, K.; Zhang, B.; Sun, C.; Zhao, L.; Dong, Z.-Y.; Cui, Q. The circular RNA FAM169A functions as a competitive endogenous RNA and regulates intervertebral disc degeneration by targeting miR-583 and BTRC. Cell Death Dis. 2020, 11, 315–317. [Google Scholar] [CrossRef] [PubMed]
- Ali, S.A.; Alman, B. RNA extraction from human articular cartilage by chondrocyte isolation. Anal. Biochem. 2012, 429, 39–41. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; McFarland, T.J.; Vartanian, K.; Zhu, Y.; Harrington, C.A.; Chu, C.Q. RNA isolation from micro-quantity of articular cartilage for quantitative gene expression by microarray analysis. Int. J. Med. Sci. 2022, 19, 98. [Google Scholar] [CrossRef] [PubMed]
- Wilson, T.; Kaur, N.; Davis, J.; Al, S.A. Tissue Collection and RNA Extraction from the Human Osteoarthritic Knee Joint. J. Vis. Exp. 2021, 173, e62718. [Google Scholar] [CrossRef] [PubMed]
- Fox, A.J.S.; Bedi, A.; Rodeo, S.A. The Basic Science of Articular Cartilage: Structure, Composition, and Function. Sport. Health 2009, 1, 461. [Google Scholar] [CrossRef]
- Zhang, Y.; Han, S.; Kong, M.; Tu, Q.; Zhang, L.; Ma, X. Single-cell RNA-seq analysis identifies unique chondrocyte subsets and reveals involvement of ferroptosis in human intervertebral disc degeneration. Osteoarthr. Cartil. 2021, 29, 1324–1334. [Google Scholar] [CrossRef] [PubMed]
- Leonova, O.N.; Elgaeva, E.E.; Golubeva, T.S.; Peleganchuk, A.V.; Krutko, A.V.; Aulchenko, Y.S.; Tsepilov, Y.A. A protocol for recruiting and analyzing the disease-oriented Russian disc degeneration study (RuDDS) biobank for functional omics studies of lumbar disc degeneration. PLoS ONE 2022, 17, e0267384. [Google Scholar] [CrossRef] [PubMed]
- Babraham Bioinformatics—FastQC A Quality Control Tool for High Throughput Sequence Data. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 20 August 2022).
- Wingett, S.W.; Andrews, S. FastQ Screen: A tool for multi-genome mapping and quality control. F1000Research 2018, 7, 1338. [Google Scholar] [CrossRef] [PubMed]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Alvarez, R.V.; Pongor, L.S.; Mariño-Ramírez, L.; Landsman, D. TPMCalculator: One-step software to quantify mRNA abundance of genomic features. Bioinformatics 2019, 35, 1960–1962. [Google Scholar] [CrossRef] [PubMed]
- Jew, B.; Alvarez, M.; Rahmani, E.; Miao, Z.; Ko, A.; Garske, K.M.; Sul, J.H.; Pietiläinen, K.H.; Pajukanta, P.; Halperin, E. Accurate estimation of cell composition in bulk expression through robust integration of single-cell information. Nat. Commun. 2020, 11, 1971. [Google Scholar] [CrossRef] [PubMed]
- Gan, Y.; He, J.; Zhu, J.; Xu, Z.; Wang, Z.; Yan, J.; Hu, O.; Bai, Z.; Chen, L.; Xie, Y.; et al. Spatially defined single-cell transcriptional profiling characterizes diverse chondrocyte subtypes and nucleus pulposus progenitors in human intervertebral discs. Bone Res. 2021, 9, 37. [Google Scholar] [CrossRef] [PubMed]

| Methodology for Lumbar Disc RNA Isolation after Mortar and Pestle Homogenization |
|---|
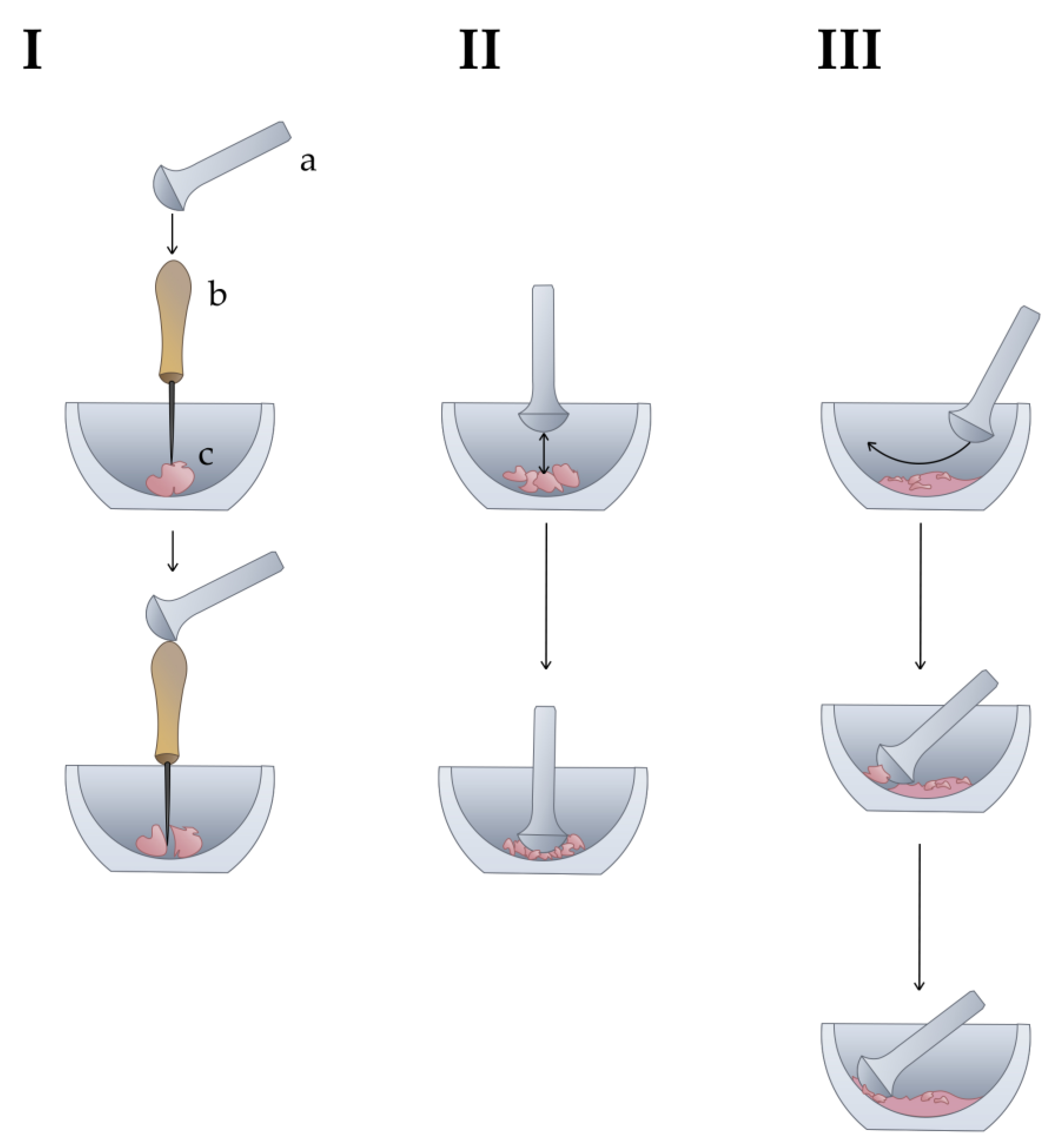
| 1. Grind the sample using a mortar and pestle (Figure 1). |
| 2. Add QIAzol* to the mortar at a rate of 0.5 mL per 300 mg of frozen sample and grind to homogeneity in liquid nitrogen. |
| 3. Pour into a 25 mL Eppendorf test tube, allow the nitrogen to evaporate, then close and mix by turning the tube. The powder should evenly coat the sides of it. |
| 4. Allow it to remain on the surface of the ice until thawed. Vortex once every 5 min. |
| 5. After thawing to viscosity, add another 0.5 mL of QIAzol for every 300 mg of the frozen sample. Vortex for 30 s, then incubate on ice for 5 min. |
| 6. Remove the tip of the automatic pipette spout with sterile scissors. Pour 1 mL lysate into 1.5 mL microcentrifuge tubes. Add 0.2 mL of chloroform to each tube. |
| 7. Stir by turning the tubes for 30 s, then vortex for 1–2 s. Incubate for 2 min on ice. |
| 8. Centrifuge for 20 min at 15,000× g and 4 °C. At this stage, the white suspension in the water phase is acceptable if one can see through it. If not, repeat the centrifugation. |
| 9. Carefully transfer the aqueous phase (usually ~400 µL) to new 1.5 mL microcentrifuge tubes and add an equal volume of QIAzol. Stir by inverting the tubes. |
| 10. Incubate for 30 min on ice in the dark. |
| 11. Add 0.2 volume of chloroform (usually ~160 µL). Repeat step 7. |
| 12. Centrifuge for 20 min, 15,000× g, 4 °C. Repeat if necessary until a white suspension precipitates. |
| 13. Transfer half of the aqueous phase from each tube (usually ~250 µL) to new 1.5 mL microcentrifuge tubes. To reduce column consumption and increase RNA concentration, it is recommended at this stage to combine the aqueous phase from every two tubes (4 tubes before phase separation–> 2 tubes after, and two RNEasy spin columns respectively). |
| Then the RNEasy plant mini kit (Qiagen) is used. |
| 14. Add 0.5 V of ethanol to each tube and mix by pipetting. Transfer to the RNEasy spin column (pink). |
| 15. Centrifuge for 15 s at 10,000× g and 15 °C. Discard the flow-through. |
| 16. Add 700 µL RW1 buffer. Repeat step 15. |
| 17. Add 500 µL RPE buffer (check that ethanol was added). Repeat step 15. Repeat the washing step with the RPE buffer twice. |
| 18. Transfer each column to a new collection tube. Centrifuge for 1 min at 12,000× g and 15 °C. |
| 19. Place the columns in new 1.5 mL microcentrifuge tubes and gently apply 25 µL of RNAse-free water to the membrane. Incubate for 1 min at room temperature. |
| 20. Centrifuge for 30 s at 10,000× g and 15 °C. |
| 21. Re-apply the eluate to the membrane to increase the yield. Incubate for one minute. Repeat step 20. Collect aliquots for concentration and RIN measurements. |
| 22. Store the isolated RNA at −80 °C. |
| Sample Code | Sample Weight (g) | Derivative Samples * |
|---|---|---|
| K_20_12_25_03 ** | 1.4 | K201 – K203 |
| N_20_12_25_01 ** | 0.7 | N1, N2 |
| 89_RAI ** | 1.8 | R1 – R6 |
| 124_MAA | 1.8 | MAA1-MAA3 |
| 143_KEV | 1.2 | KEV1, KEV2 |
| 134_TSA | 1.2 | TSA1, TSA2 |
| 137_SGN | 1.2 | SGH1, SGH2 |
| 131_MTU | 1.9 | MTU1-MTU3 |
| 152_ZAI *** | 1.8 | ZAI1-ZAI3 |
| 140_YAGi | 2.1 | YAG1-YAG3 |
| Sample Code | Sample Weight (g) | Mean C ± SEM (ng/uL) | Mean RIN ± SEM |
|---|---|---|---|
| K_20_12_25_03 | 1.4 | 71.2 ± 17.2 * | 7.3 ± 0.1 * |
| N_20_12_25_01 | 0.7 | 8.5 ± 1.7 | 6.7 ± 0.2 |
| 89_RAI | 1.8 | 49.4 ± 3.4 * | 6.9 ± 0.1 * |
| 124_MAA | 1.8 | 69.1 ± 4.0 | 8.5 ± 0.3 |
| 143_KEV | 1.2 | 14.8 ± 3.6 | 7.0 ± 0.05 |
| 134_TSA | 1.2 | 27.3 ± 2.3 | 6.7 ± 0.1 |
| 137_SGN | 1.2 | 18.5 ± 0.4 | 6.3 ± 0.2 |
| 131_MTU | 1.9 | 12.8 ± 2.5 | 6.5 ± 0.2 |
| 152_ZAI | 1.8 | 6.1 ± 1.4 | N/D ** |
| 140_YAGi | 2.1 | 29.4 ± 7.1 | 7.8 *** |
| All | - | 32.4 ± 4.7 * | 7.1 ± 0.2 * |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ivanov, A.A.; Leonova, O.N.; Wiebe, D.S.; Krutko, A.V.; Gridina, M.M.; Fishman, V.S.; Aulchenko, Y.S.; Tsepilov, Y.A.; Golubeva, T.S. Method for the Isolation of “RNA-seq-Quality” RNA from Human Intervertebral Discs after Mortar and Pestle Homogenization. Cells 2022, 11, 3578. https://doi.org/10.3390/cells11223578
Ivanov AA, Leonova ON, Wiebe DS, Krutko AV, Gridina MM, Fishman VS, Aulchenko YS, Tsepilov YA, Golubeva TS. Method for the Isolation of “RNA-seq-Quality” RNA from Human Intervertebral Discs after Mortar and Pestle Homogenization. Cells. 2022; 11(22):3578. https://doi.org/10.3390/cells11223578
Chicago/Turabian StyleIvanov, Artemii A., Olga N. Leonova, Daniil S. Wiebe, Alexsandr V. Krutko, Mariya M. Gridina, Veniamin S. Fishman, Yurii S. Aulchenko, Yakov A. Tsepilov, and Tatiana S. Golubeva. 2022. "Method for the Isolation of “RNA-seq-Quality” RNA from Human Intervertebral Discs after Mortar and Pestle Homogenization" Cells 11, no. 22: 3578. https://doi.org/10.3390/cells11223578
APA StyleIvanov, A. A., Leonova, O. N., Wiebe, D. S., Krutko, A. V., Gridina, M. M., Fishman, V. S., Aulchenko, Y. S., Tsepilov, Y. A., & Golubeva, T. S. (2022). Method for the Isolation of “RNA-seq-Quality” RNA from Human Intervertebral Discs after Mortar and Pestle Homogenization. Cells, 11(22), 3578. https://doi.org/10.3390/cells11223578






