KRAS G12D Mutation Subtype in Pancreatic Ductal Adenocarcinoma: Does It Influence Prognosis or Stage of Disease at Presentation?
Abstract
1. Introduction
2. Methods
2.1. Ethics Statement
2.2. Data and Sample Collection
2.3. Statistical Analysis
3. Results
3.1. Patient Characteristics at Baseline
3.2. Survival Analysis for the Entire Cohort
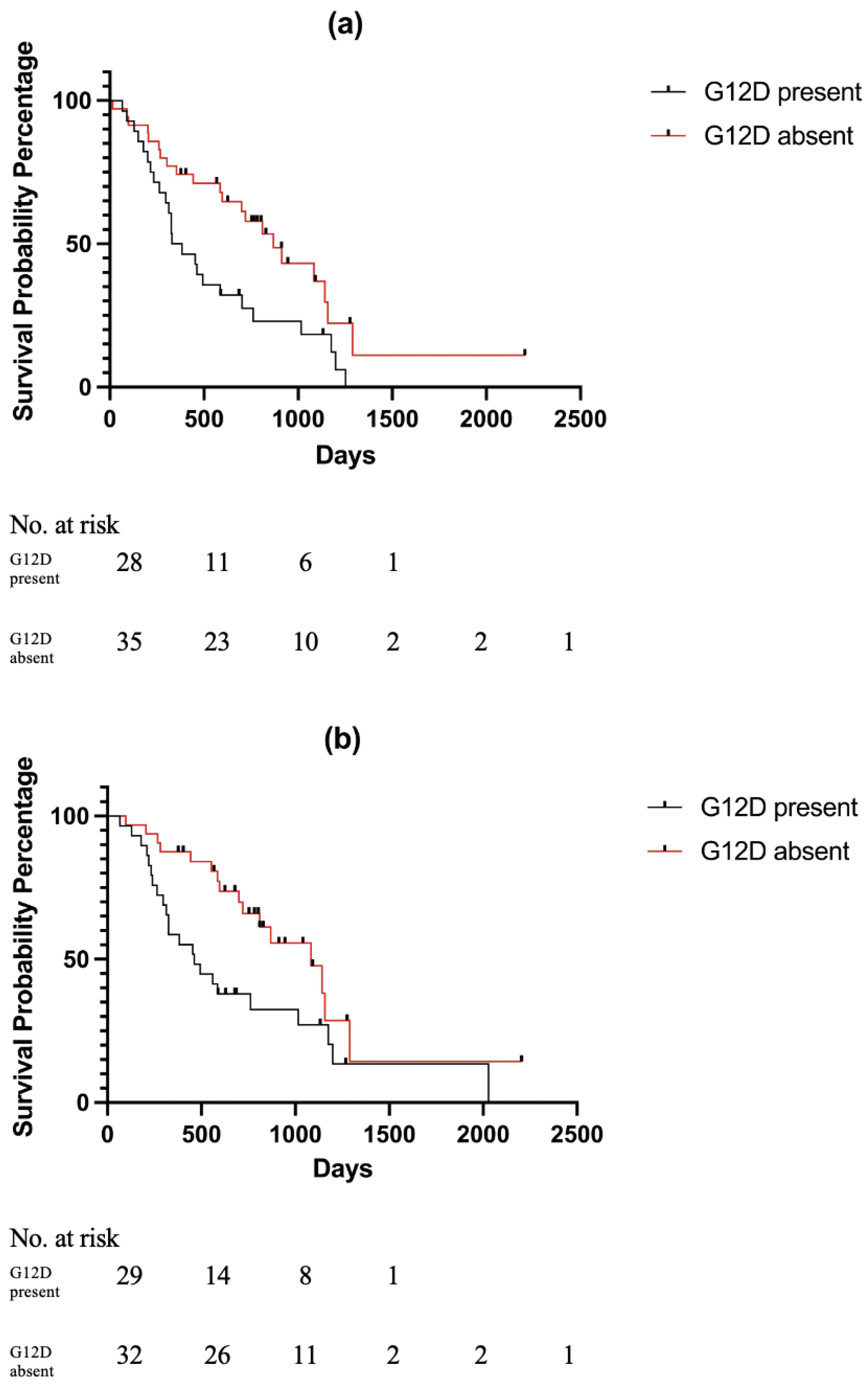
3.3. Survival Analysis by NCCN Stage and Treatment Modality and KRAS Mutation Status
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ferlay, J.E.M.; Lam, F.; Colombet, M.; Mery, L.; Piñeros, M.; Znaor, A.; Soerjomataram, I.; Bray, F. Global Cancer Observatory: Cancer Today; International Agency for Research on Cancer: Lyon, France, 2020. Available online: https://gco.iarc.fr/today (accessed on 6 June 2020).
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Vincent, A.; Herman, J.; Schulick, R.; Hruban, R.H.; Goggins, M. Pancreatic cancer. Lancet 2011, 378, 607–620. [Google Scholar] [CrossRef]
- Dunne, R.F.; Hezel, A.F. Genetics and Biology of Pancreatic Ductal Adenocarcinoma. Hematol. Clin. N. Am. 2015, 29, 595–608. [Google Scholar] [CrossRef] [PubMed]
- Tempero, M.A.; Malafa, M.P.; Al-Hawary, M.; Asbun, H.; Bain, A.; Behrman, S.W.; Benson, A.B.; Binder, E.; Cardin, D.B.; Cha, C.; et al. Pancreatic Adenocarcinoma, Version 2.2017, NCCN Clinical Practice Guidelines in Oncology. J. Natl. Compr. Cancer Netw. 2017, 15, 1028–1061. [Google Scholar] [CrossRef]
- Fuccio, L.; Hassan, C.; Laterza, L.; Correale, L.; Pagano, N.; Bocus, P.; Fabbri, C.; Maimone, A.; Cennamo, V.; Repici, A.; et al. The role of K-ras gene mutation analysis in EUS-guided FNA cytology specimens for the differential diagnosis of pancreatic solid masses: A meta-analysis of prospective studies. Gastrointest. Endosc. 2013, 78, 596–608. [Google Scholar] [CrossRef]
- Kamisawa, T.; Wood, L.D.; Itoi, T.; Takaori, K. Pancreatic cancer. Lancet 2016, 388, 73–85. [Google Scholar] [CrossRef]
- Cicenas, J.; Kvederaviciute, K.; Meskinyte, I.; Meskinyte-Kausiliene, E.; Skeberdyte, A.; Cicenas, J. KRAS, TP53, CDKN2A, SMAD4, BRCA1, and BRCA2 Mutations in Pancreatic Cancer. Cancers 2017, 9, 42. [Google Scholar] [CrossRef]
- Singhi, A.D.; George, B.; Greenbowe, J.R.; Chung, J.; Suh, J.; Maitra, A.; Klempner, S.J.; Hendifar, A.; Milind, J.M.; Golan, T.; et al. Real-Time Targeted Genome Profile Analysis of Pancreatic Ductal Adenocarcinomas Identifies Genetic Alterations That Might Be Targeted with Existing Drugs or Used as Biomarkers. Gastroenterology 2019, 156, 2242–2253. e4. [Google Scholar] [CrossRef]
- Buscail, L.; Bournet, B.; Cordelier, P. Role of oncogenic KRAS in the diagnosis, prognosis and treatment of pancreatic cancer. Nat. Rev. Gastroenterol. Hepatol. 2020, 17, 153–168. [Google Scholar] [CrossRef]
- Buscail, E.; Maulat, C.; Muscari, F.; Chiche, L.; Cordelier, P.; Dabernat, S.; Alix-Panabières, C.; Buscail, L. Liquid Biopsy Approach for Pancreatic Ductal Adenocarcinoma. Cancers 2019, 11, 852. [Google Scholar] [CrossRef]
- Kawesha, A.; Ghaneh, P.; Andrén-Sandberg, Å.; Ögraed, D.; Skar, R.; Dawiskiba, S.; Evans, J.D.; Campbell, F.; Lemoine, N.; Neoptolemos, J.P. K-ras oncogene subtype mutations are associated with survival but not expression of p53, p16INK4A, p21WAF-1, cyclin D1, erbB-2 and erbB-3 in resected pancreatic ductal adenocarcinoma. Int. J. Cancer 2000, 89, 469–474. [Google Scholar] [CrossRef]
- Shin, S.H.; Kim, S.C.; Hong, S.-M.; Kim, Y.H.; Song, K.-B.; Park, K.-M.; Lee, Y.-J. Genetic Alterations of K-ras, p53, c-erbB-2, and DPC4 in Pancreatic Ductal Adenocarcinoma and Their Correlation with Patient Survival. Pancreas 2013, 42, 216–222. [Google Scholar] [CrossRef] [PubMed]
- Rachakonda, P.S.; Bauer, A.S.; Xie, H.; Campa, D.; Rizzato, C.; Canzian, F.; Beghelli, S.; Greenhalf, W.; Costello, E.; Schanne, M.; et al. Somatic Mutations in Exocrine Pancreatic Tumors: Association with Patient Survival. PLoS ONE 2013, 8, e60870. [Google Scholar] [CrossRef] [PubMed]
- Qian, Z.R.; Rubinson, D.A.; Nowak, J.A.; Morales-Oyarvide, V.; Dunne, R.F.; Kozak, M.M.; Welch, M.W.; Brais, L.K.; Da Silva, A.; Li, T.; et al. Association of Alterations in Main Driver Genes with Outcomes of Patients with Resected Pancreatic Ductal Adenocarcinoma. JAMA Oncol. 2018, 4, e173420. [Google Scholar] [CrossRef] [PubMed]
- Ogura, T.; Yamao, K.; Hara, K.; Mizuno, N.; Hijioka, S.; Imaoka, H.; Sawaki, A.; Niwa, Y.; Tajika, M.; Kondo, S.; et al. Prognostic value of K-ras mutation status and subtypes in endoscopic ultrasound-guided fine-needle aspiration specimens from patients with unresectable pancreatic cancer. J. Gastroenterol. 2012, 48, 640–646. [Google Scholar] [CrossRef]
- Boeck, S.; Jung, A.; Laubender, R.P.; Neumann, J.; Egg, R.; Goritschan, C.; Ormanns, S.; Haas, M.; Modest, D.P.; Kirchner, T.; et al. KRAS mutation status is not predictive for objective response to anti-EGFR treatment with erlotinib in patients with advanced pancreatic cancer. J. Gastroenterol. 2013, 48, 544–548. [Google Scholar] [CrossRef]
- Bournet, B.; Muscari, F.; Buscail, C.; Assenat, E.; Barthet, M.; Hammel, P.; Selves, J.; Guimbaud, R.; Cordelier, P.; Buscail, L. KRAS G12D Mutation Subtype Is A Prognostic Factor for Advanced Pancreatic Adenocarcinoma. Clin. Transl. Gastroenterol. 2016, 7, e157. [Google Scholar] [CrossRef]
- Gu, Y.; Ji, Y.; Jiang, H.; Qiu, G. Clinical Effect of Driver Mutations of KRAS, CDKN2A/P16, TP53, and SMAD4 in Pancreatic Cancer: A Meta-Analysis. Genet. Test. Mol. Biomarkers 2020, 24, 777–788. [Google Scholar] [CrossRef]
- Hendifar, A.E.; Blais, E.M.; Ng, C.; Thach, D.; Gong, J.; Sohal, D.; Chung, V.; Sahai, V.; Fountzilas, C.; Mikhail, S.; et al. Comprehensive analysis of KRAS variants in patients (pts) with pancreatic cancer (PDAC): Clinical/molecular correlations and real-world outcomes across standard therapies. J. Clin. Oncol. 2020, 38, 4641. [Google Scholar] [CrossRef]
- Philip, P.A.; Azar, I.; Xiu, J.; Hall, M.J.; Hendifar, A.E.; Lou, E.; Hwang, J.J.; Gong, J.; Feldman, R.; Ellis, M.; et al. Molecular Characterization of KRAS Wild-type Tumors in Patients with Pancreatic Adenocarcinoma. Clin. Cancer Res. 2022, 28, 2704–2714. [Google Scholar] [CrossRef]
- Lundy, J.; Harris, M.; Zalcberg, J.; Zimet, A.; Goldstein, D.; Gebski, V.; Borsaru, A.; Desmond, C.; Swan, M.; Jenkins, B.J.; et al. EUS-FNA Biopsies to Guide Precision Medicine in Pancreatic Cancer: Results of a Pilot Study to Identify KRAS Wild-Type Tumours for Targeted Therapy. Front. Oncol. 2021, 11, 770022. [Google Scholar] [CrossRef] [PubMed]
- Tempero, M.A.; Arnoletti, J.P.; Behrman, S.W.; Ben-Josef, E.; Benson Iii, A.B.; Casper, E.S.; Cohen, S.J.; Czito, B.; Ellenhorn, J.D.I.; Hawkins, W.G.; et al. Pancreatic adenocarcinoma, version 2.2012: Featured updates to the NCCN guidelines. JNCCN J. Natl. Compr. Cancer Netw. 2012, 10, 703–713. [Google Scholar] [CrossRef] [PubMed]
- Isaji, S.; Mizuno, S.; Windsor, J.A.; Bassi, C.; Castillo, C.F.-D.; Hackert, T.; Hayasaki, A.; Katz, M.H.; Kim, S.-W.; Kishiwada, M.; et al. International consensus on definition and criteria of borderline resectable pancreatic ductal adenocarcinoma 2017. Pancreatology 2018, 18, 2–11. [Google Scholar] [CrossRef] [PubMed]
- McIntyre, C.A.; Lawrence, S.A.; Richards, A.L.; Chou, J.F.; Wong, W.; Capanu, M.; Berger, M.F.; Donoghue, M.T.A.; Yu, K.H.; Varghese, A.M.; et al. Alterations in driver genes are predictive of survival in patients with resected pancreatic ductal adenocarcinoma. Cancer 2020, 126, 3939–3949. [Google Scholar] [CrossRef]
- Kulkarni, N.M.; Soloff, E.V.; Tolat, P.P.; Sangster, G.P.; Fleming, J.B.; Brook, O.R.; EHecht, l.M.; Kastrinos, F.; Wang, Z.J.; Soloff, E.V.; et al. White paper on pancreatic ductal adenocarcinoma from society of abdominal radiology’s disease-focused panel for pancreatic ductal adenocarcinoma: Part I, AJCC staging system, NCCN guidelines, and borderline resectable disease. Abdom. Radiol. 2020, 45, 716–728. [Google Scholar] [CrossRef] [PubMed]
- National Comprehensive Cancer Network. Pancreatic Adenocarcinoma (Version 1.2022). 2022. Available online: https://www.nccn.org/professionals/physician_gls/pdf/pancreatic.pdf (accessed on 6 September 2022).
- Hong, D.S.; Fakih, M.G.; Strickler, J.H.; Desai, J.; Durm, G.A.; Shapiro, G.I.; Falchook, G.S.; Price, T.J.; Sacher, A.; Denlinger, C.S.; et al. KRAS(G12C) Inhibition with Sotorasib in Advanced Solid Tumors. N. Engl. J. Med. 2020, 383, 1207–1217. [Google Scholar] [CrossRef]
- Skoulidis, F.; Li, B.T.; Dy, G.K.; Price, T.J.; Falchook, G.S.; Wolf, J.; Italiano, A.; Schuler, M.; Borghaei, H.; Barlesi, F.; et al. Sotorasib for Lung Cancers with KRAS p.G12C Mutation. N. Engl. J. Med. 2021, 384, 2371–2381. [Google Scholar] [CrossRef]
- Yuan, T.L.; Fellmann, C.; Lee, C.-S.; Ritchie, C.D.; Thapar, V.; Lee, L.C.; Hsu, D.J.; Grace, D.; Carver, J.O.; Zuber, J.; et al. Development of siRNA Payloads to Target KRAS-Mutant Cancer. Cancer Discov. 2014, 4, 1182–1197. [Google Scholar] [CrossRef]
- Golan, T.; Khvalevsky, E.Z.; Hubert, A.; Gabai, R.M.; Hen, N.; Segal, A.; Domb, A.; Harari, G.; Ben David, E.; Raskin, S.; et al. RNAi therapy targeting KRAS in combination with chemotherapy for locally advanced pancreatic cancer patients. Oncotarget 2015, 6, 24560–24570. [Google Scholar] [CrossRef]
- Kamerkar, S.; LeBleu, V.S.; Sugimoto, H.; Yang, S.; Ruivo, C.; Melo, S.; Lee, J.J.; Kalluri, R. Exosomes facilitate therapeutic targeting of oncogenic KRAS in pancreatic cancer. Nature 2017, 546, 498–503. [Google Scholar] [CrossRef]
- Loveday, B.P.T.; Lipton, L.; Thomson, B.N.J. Pancreatic cancer. Aust. J. Gen. Pract. 2019, 48, 826–831. [Google Scholar] [CrossRef] [PubMed]
- Oba, A.; Ho, F.; Bao, Q.R.; Al-Musawi, M.H.; Schulick, R.D.; Del Chiaro, M. Neoadjuvant Treatment in Pancreatic Cancer. Front. Oncol. 2020, 10, 245. [Google Scholar] [CrossRef] [PubMed]
- Tummers, W.S.; Groen, J.V.; Mulder, B.G.S.; Farina-Sarasqueta, A.; Morreau, J.; Putter, H.; Van De Velde, C.J.; Vahrmeijer, A.L.; Bonsing, B.A.; Mieog, J.S.; et al. Impact of resection margin status on recurrence and survival in pancreatic cancer surgery. Br. J. Surg. 2019, 106, 1055–1065. [Google Scholar] [CrossRef] [PubMed]
| KRAS | G12D | |||||||
|---|---|---|---|---|---|---|---|---|
| Characteristics | Total | Mutant | WT | p Value | Yes | No | p Value | |
| Total | 231 | 202 | 29 | 93 | 138 | |||
| Sex | Male | 109 | 91 | 18 | 0.1290 | 39 | 70 | 0.2388 |
| Female | 122 | 111 | 11 | 54 | 68 | |||
| Median age | 70.91 | 70.91 | 70.63 | 0.2562 | 71.11 | 70.33 | 0.2533 | |
| Detection method | EUS-FNA | 194 | 169 | 25 | 0.9374 | 75 | 119 | 0.3409 |
| Resection specimen | 37 | 33 | 4 | 18 | 19 | |||
| NCCN stage | Resectable | 63 | 55 | 8 | 0.1627 | 28 | 35 | 0.500 |
| Borderline Resectable | 23 | 23 | 0 | 9 | 14 | |||
| Locally Advanced | 67 | 55 | 12 | 22 | 45 | |||
| Metastatic | 78 | 69 | 9 | 34 | 44 | |||
| Treatment | Surgery | 63 | 52 | 9 | 0.7604 | 29 | 32 | 0.3471 |
| Chemotherapy and/or Radiotherapy | 125 | 111 | 14 | 46 | 79 | |||
| Supportive Care | 43 | 37 | 6 | 16 | 27 |
| Subtype | Number of Patients | Frequency |
|---|---|---|
| G12A | 2 | 0.9% |
| G12C | 4 | 1.7% |
| G12D | 93 | 40.3% |
| G12R | 24 | 10.4% |
| G12V | 64 | 27.7% |
| G13D | 2 | 0.9% |
| Q61H | 11 | 4.8% |
| Q61R | 2 | 0.9% |
| WT | 29 | 12.6% |
| KRAS | G12D | |||||||
|---|---|---|---|---|---|---|---|---|
| Characteristics | Total | Mutant | WT | p Value | Yes | No | p Value | |
| Lymph node status | Negative | 13 | 12 | 1 | 0.2149 | 5 | 8 | 0.7652 |
| Positive | 37 | 33 | 4 | 18 | 19 | |||
| Tumour location | Head | 40 | 33 | 7 | 0.6643 | 18 | 22 | 0.8719 |
| Distal | 13 | 12 | 1 | 5 | 8 | |||
| Surgical Margins | Clear | 27 | 24 | 3 | >0.999 | 12 | 15 | 0.9096 |
| Positive | 24 | 20 | 4 | 11 | 13 | |||
| Neoadjuvant therapy | No | 10 | 36 | 7 | 0.7030 | 18 | 25 | 0.8990 |
| Yes | 43 | 9 | 1 | 5 | 5 | |||
| Adjuvant therapy | No | 12 | 9 | 3 | 0.3720 | 4 | 8 | 0.5095 |
| Yes | 39 | 34 | 5 | 19 | 20 |
| Median Survival (Days) | Number of Patients | Univariate Analysis (p-Value; HR 95% CI) | Multivariate Analysis (p-Value; HR 95% CI) | ||
|---|---|---|---|---|---|
| KRAS | WT | 420.0 | 202 | Reference | Reference |
| Mutant | 296.0 | 29 | 0.2971; | 0.843; | |
| 1.257 | 1.050 | ||||
| (0.8416–1.877) | (0.646–1.709) | ||||
| G12D | No | 296.0 | 93 | Reference | Reference |
| Yes | 313.0 | 138 | 0.2151; | 0.107; | |
| 1.205 | 1.293 | ||||
| (0.8934–1.606) | (0.946–1.767) | ||||
| G12V | No | 301.0 | 167 | Reference | Reference |
| Yes | 295.0 | 64 | 0.5200; | 0.618; | |
| 0.8992 | 0.918 | ||||
| (0.6548–1.235) | (0.656–1.285) | ||||
| G12R | No | 307.0 | 207 | Reference | Reference |
| Yes | 268.0 | 24 | 0.3059; | 0.641; | |
| 1.260 | 1.119 | ||||
| (0.7741–2.049) | (0.697–1.797) | ||||
| Sex | Male | 301.0 | 109 | Reference | - |
| Female | 295.0 | 122 | 0.4866; | - | |
| 1.106 | |||||
| (0.8321–1.470) | |||||
| Age | ≤70.91 | 397.0 | 116 | Reference | Reference |
| >70.91 | 228.0 | 115 | 0.0135; | 0.052 | |
| 1.425 | 0.732; | ||||
| (1.070–1.899) | (0.534–1.003) | ||||
| Clinical stage | Resectable | 596.0 | 63 | Reference | Reference |
| Borderline Resectable | 296.0 | 23 | 0.2633; | 0.986; | |
| 1.365 | 1.005 | ||||
| (0.7488–2.487) | (0.558–1.812) | ||||
| Locally Advanced | 316.0 | 67 | <0.0001; | 0.833; | |
| 2.068 | 1.061 | ||||
| (1.383–3.093) | (0.614–1.834) | ||||
| Metastatic | 193.0 | 78 | <0.0001; | 0.047; | |
| 3.277 | 1.775 | ||||
| (2.194–4.894) | (1.007–3.128) | ||||
| Treatment | Surgery | 762.0 | 63 | Reference | Reference |
| Chemo/Radio | 292.0 | 125 | <0.0001; | 0.002; | |
| 2.860 | 2.490 | ||||
| (2.071–3.950) | (1.392–4.453) | ||||
| Supportive care | 139.0 | 43 | <0.0001; | <0.0001; | |
| 4.443 | 5.504 | ||||
| (2.558–7.718) | (3.126–9.689) |
| Median Survival (Days) | Number of Patients | Univariate Analysis (p-Value; HR 95% CI) | Multivariate Analysis (p-Value; HR 95% CI) | ||
|---|---|---|---|---|---|
| Lymph node status | Negative | Undefined | 13 | Reference | Reference |
| Positive | 729.0 | 37 | 0.0176; | 0.543; | |
| 3.235 | 1.621 | ||||
| (1.572–6.656) | (0.342–7.681) | ||||
| Tumour location | Head | 720.0 | 40 | Reference | Reference |
| Distal | Undefined | 13 | 0.0118; | 0.073; | |
| 0.2878 | 0.290 | ||||
| (0.1414–0.5860) | (0.075–1.124) | ||||
| Neoadjuvant therapy | No | 810.0 | 27 | Reference | Reference |
| Yes | 1028 | 24 | 0.2943; | 0.361; | |
| 0.6083 | 2.075 | ||||
| (0.2726–1.357) | (0.433–9.948) | ||||
| Surgical margins | Clear | 1016 | 10 | Reference | Reference |
| Positive | 729.0 | 43 | 0.2018; | 0.156; | |
| 1.532 | 1.849 | ||||
| (0.7855–2.988) | (0.791–4.323) | ||||
| Adjuvant therapy | No | 636.5 | 12 | Reference | Reference |
| Yes | 1016 | 39 | 0.3166; | 0.108; | |
| 0.7894 | 0.360 | ||||
| (0.3225–1.932) | (0.104–1.249) |
| Number of Patients | Median Survival (Days) | Univariate Analysis (p-Value; HR 95% CI) | Multivariate Analysis (p-Value; HR 95% CI) | |||
|---|---|---|---|---|---|---|
| Resectable | KRAS | WT | 8 | 1143 | Reference | Reference |
| Mutant | 55 | 494.0 | 0.1296; | 0.306; | ||
| 1.994 | 1.647 | |||||
| (1.008–4.239) | (0.633–4.285) | |||||
| G12D | No | 35 | 810.0 | Reference | Reference | |
| Yes | 28 | 356.0 | 0.0168; | 0.019; | ||
| 2.053 | 1.991 | |||||
| (1.139–3.702) | (1.121–3.537) | |||||
| Borderline Resectable | KRAS | WT | 0 | - | - | - |
| Mutant | 23 | 296.0 | - | - | ||
| G12D | No | 14 | 286.5 | Reference | Reference | |
| Yes | 9 | 358.0 | 0.3180; | 0.113; | ||
| 0.6083 | 0.377 | |||||
| (0.2347–1.576) | (0.113–1.259) | |||||
| Locally Advanced | KRAS | WT | 12 | 248.0 | Reference | Reference |
| Mutant | 55 | 316.0 | 0.5046; | 0.217; | ||
| 0.7941 | 0.634 | |||||
| (0.3788–1.665) | (0.307–1.308) | |||||
| G12D | No | 22 | 326.0 | Reference | Reference | |
| Yes | 45 | 316.0 | 0.2745; | 0.216; | ||
| 1.351 | 1.441 | |||||
| (0.7502–2.432) | (0.808–2.568) | |||||
| Metastatic | KRAS | WT | 9 | 196.0 | Reference | Reference |
| Mutant | 69 | 189.0 | 0.5584; | 0.874; | ||
| 1.228 | 1.068 | |||||
| (0.5995–2.579) | (0.475–2.401) | |||||
| G12D | No | 44 | 196.0 | Reference | Reference | |
| Yes | 34 | 190.0 | 0.6529; | 0.815; | ||
| 1.114 | 0.941 | |||||
| (0.6872–1.805) | (0.562–1.574) | |||||
| Surgery | KRAS | WT | 9 | 1143 | Reference | Reference |
| Mutant | 52 | 729.0 | 0.2470; | 0.438; | ||
| 1.717 | 1.459 | |||||
| (0.7920–3.721) | (0.562–3.787) | |||||
| G12D | No | 32 | 1084 | Reference | Reference | |
| Yes | 29 | 462.0 | 0.0221; | 0.254; | ||
| 2.000 | 1.471 | |||||
| (1.079–3.707) | (0.758–2.855) | |||||
| Chemotherapy and/or Radiotherapy | KRAS | WT | 14 | 225.0 | Reference | Reference |
| Mutant | 111 | 290.0 | 0.7631; | 0.782; | ||
| 1.089 | 1.090 | |||||
| (0.6208–1.911) | (0.594–1.998) | |||||
| G12D | No | 79 | 260.0 | Reference | Reference | |
| Yes | 46 | 310.5 | 0.4619; | 0.364; | ||
| 1.156 | 1.207 | |||||
| (0.7723–1.730) | (0.804–1.813) | |||||
| Supportive Care | KRAS | WT | 6 | 97.00 | Reference | Reference |
| Mutant | 37 | 140.0 | 0.3007; | 0.283; | ||
| 0.6427 | 0.591 | |||||
| (0.2310–1.788) | (0.226–1.544) | |||||
| G12D | No | 27 | 138.0 | Reference | Reference | |
| Yes | 17 | 135.0 | 0.6952; | 0.696; | ||
| 1.131 | 1.136 | |||||
| (0.5980–2.138) | (0.599–2.155) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shen, H.; Lundy, J.; Strickland, A.H.; Harris, M.; Swan, M.; Desmond, C.; Jenkins, B.J.; Croagh, D. KRAS G12D Mutation Subtype in Pancreatic Ductal Adenocarcinoma: Does It Influence Prognosis or Stage of Disease at Presentation? Cells 2022, 11, 3175. https://doi.org/10.3390/cells11193175
Shen H, Lundy J, Strickland AH, Harris M, Swan M, Desmond C, Jenkins BJ, Croagh D. KRAS G12D Mutation Subtype in Pancreatic Ductal Adenocarcinoma: Does It Influence Prognosis or Stage of Disease at Presentation? Cells. 2022; 11(19):3175. https://doi.org/10.3390/cells11193175
Chicago/Turabian StyleShen, Henry, Joanne Lundy, Andrew H. Strickland, Marion Harris, Michael Swan, Christopher Desmond, Brendan J. Jenkins, and Daniel Croagh. 2022. "KRAS G12D Mutation Subtype in Pancreatic Ductal Adenocarcinoma: Does It Influence Prognosis or Stage of Disease at Presentation?" Cells 11, no. 19: 3175. https://doi.org/10.3390/cells11193175
APA StyleShen, H., Lundy, J., Strickland, A. H., Harris, M., Swan, M., Desmond, C., Jenkins, B. J., & Croagh, D. (2022). KRAS G12D Mutation Subtype in Pancreatic Ductal Adenocarcinoma: Does It Influence Prognosis or Stage of Disease at Presentation? Cells, 11(19), 3175. https://doi.org/10.3390/cells11193175





