Expression of Functional Cannabinoid Type-1 (CB1) Receptor in Mitochondria of White Adipocytes
Abstract
:1. Introduction
2. Materials and Methods
2.1. Animal and Drugs
2.2. Sample Preparation for Morphological Studies
2.3. Peroxidase Immunohistochemistry
2.4. Immunogold Post-Embedding Technique
2.5. Morphometric Analysis
2.6. Homogenate Preparation of eWAT and iBAT Mouse Tissue
2.7. Complex I Assay Activity
2.8. Mitochondrial Isolation from eWAT Rat Tissue
2.9. Mitochondrial Respiration
2.10. Western Blotting
2.11. Statistical Analysis
3. Results
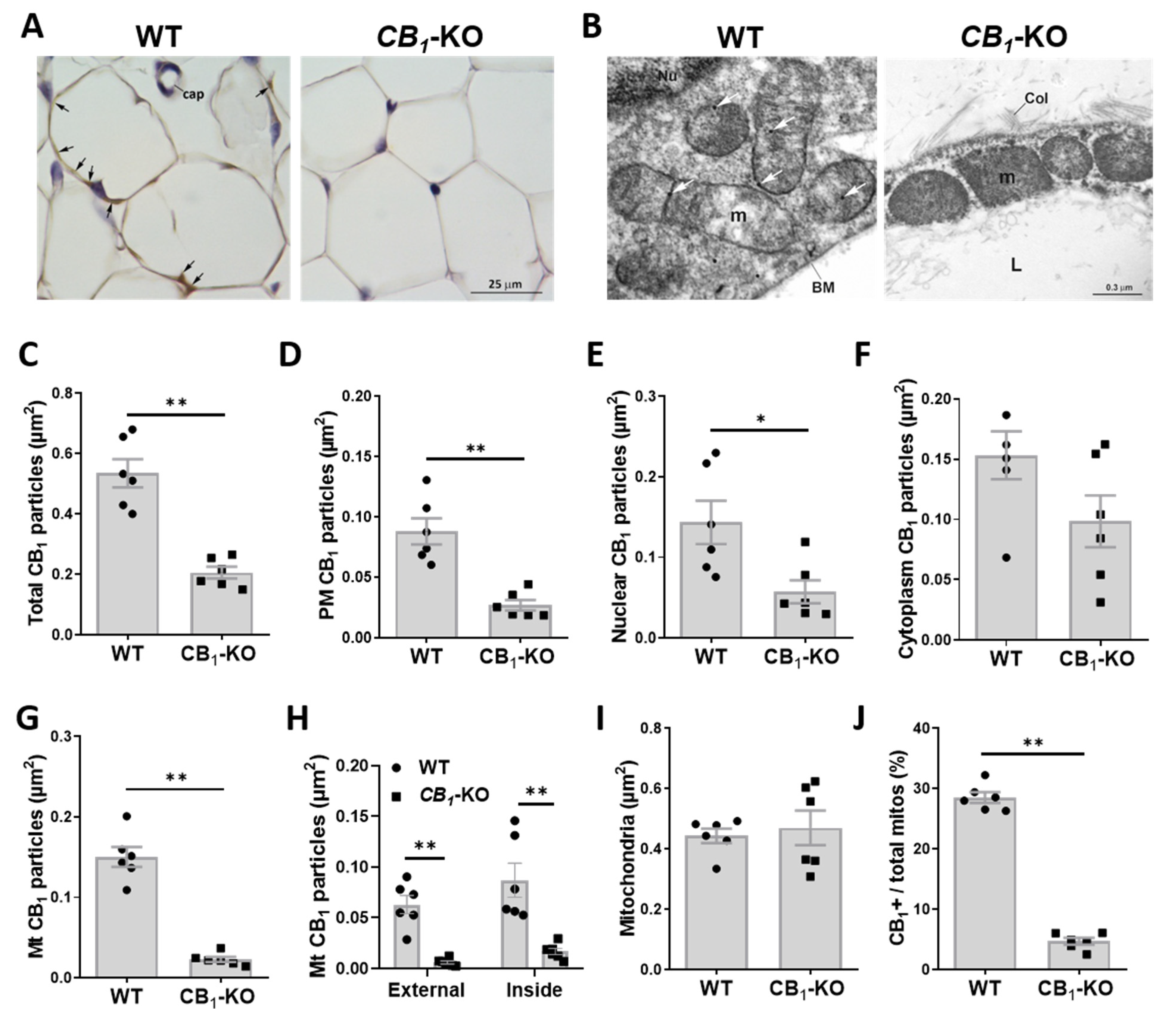
3.1. Immunohistochemical Detection of CB1 in Adipose Tissue
3.2. CB1 Receptor Is Present on Plasma Membrane and Associated to Nucleus and Mitochondria in eWAT Adipocytes
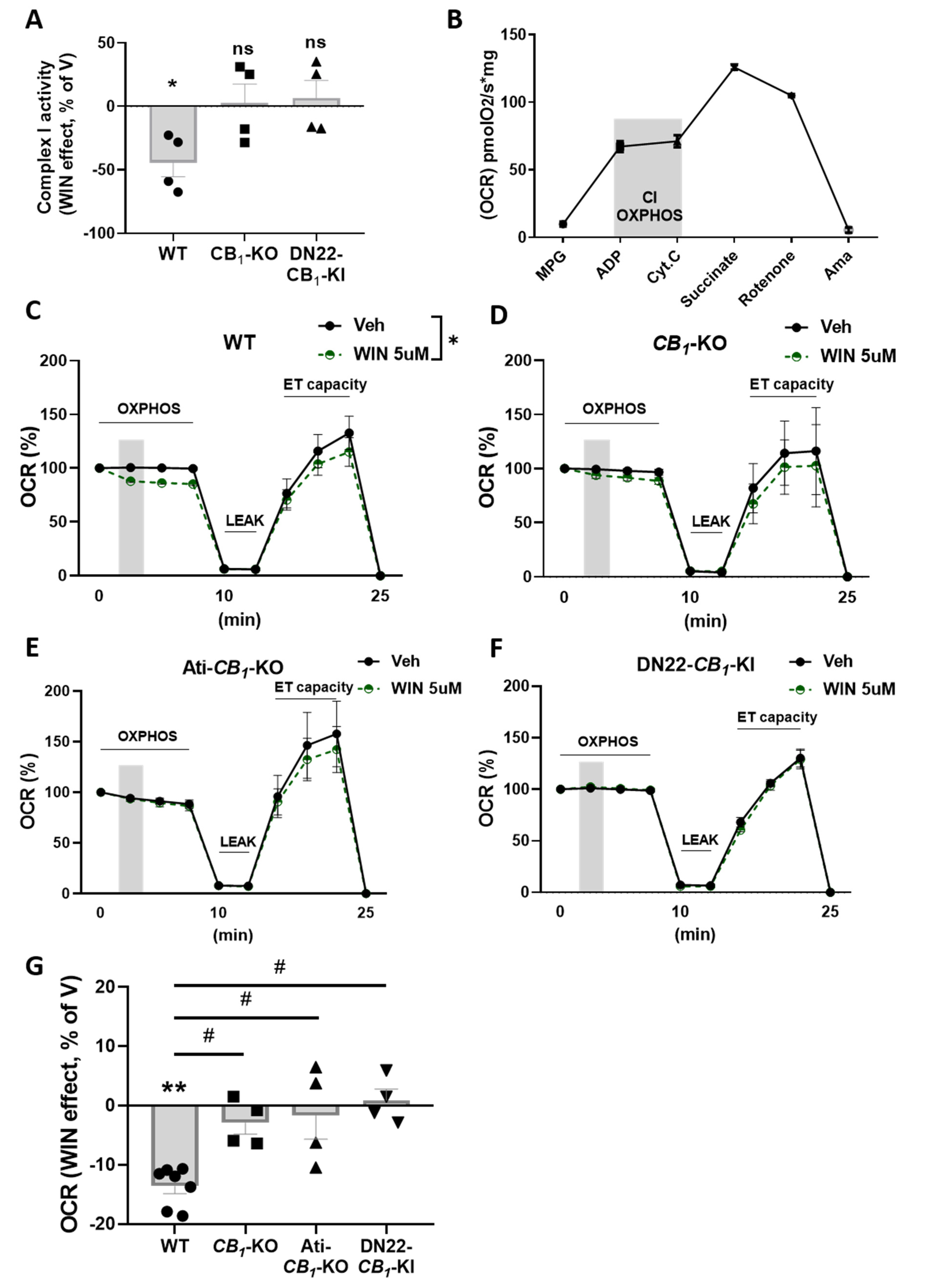
3.3. CB1 Receptor Activation Impacts Mitochondrial Respiration in eWAT
3.4. mtCB1 Reduces Mitochondrial Respiration in eWAT via sAC and PKA Activity
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Pagotto, U.; Marsicano, G.; Cota, D.; Lutz, B.; Pasquali, R. The Emerging Role of the Endocannabinoid System in Endocrine Regulation and Energy Balance. Endocr. Rev. 2006, 27, 73–100. [Google Scholar] [CrossRef] [PubMed]
- Quarta, C.; Mazza, R.; Obici, S.; Pasquali, R.; Pagotto, U. Energy Balance Regulation by Endocannabinoids at Central and Peripheral Levels. Trends Mol. Med. 2011, 17, 518–526. [Google Scholar] [CrossRef] [PubMed]
- Matias, I.; Marzo, V. Di Endocannabinoids and the Control of Energy Balance. Trends Endocrinol. Metab. 2007, 18, 27–37. [Google Scholar] [CrossRef] [PubMed]
- Mazier, W.; Saucisse, N.; Gatta-Cherifi, B.; Cota, D. The Endocannabinoid System: Pivotal Orchestrator of Obesity and Metabolic Disease. Trends Endocrinol. Metab. 2015, 26, 524–537. [Google Scholar] [CrossRef]
- Cinti, S. The Adipose Organ at a Glance. Dis. Model. Mech. 2012, 5, 588–594. [Google Scholar] [CrossRef]
- Giordano, A.; Smorlesi, A.; Frontini, A.; Barbatelli, G.; Cinti, S. MECHANISMS IN ENDOCRINOLOGY: White, Brown and Pink Adipocytes: The Extraordinary Plasticity of the Adipose Organ. Eur. J. Endocrinol. 2014, 170, R159–R171. [Google Scholar] [CrossRef]
- Lee, J.H.; Park, A.; Oh, K.-J.; Kim, W.K.; Bae, K.-H. The Role of Adipose Tissue Mitochondria: Regulation of Mitochondrial Function for the Treatment of Metabolic Diseases. Int. J. Mol. Sci. 2019, 20, 4924. [Google Scholar] [CrossRef]
- Boudina, S.; Graham, T.E. Mitochondrial Function/Dysfunction in White Adipose Tissue. Exp. Physiol. 2014, 99, 1168–1178. [Google Scholar] [CrossRef]
- Bellocchio, L.; Cervino, C.; Vicennati, V.; Pasquali, R.; Pagotto, U. Cannabinoid Type 1 Receptor: Another Arrow in the Adipocytes’ Bow. J. Neuroendocrinol. 2008, 20, 130–138. [Google Scholar] [CrossRef]
- Bensaid, M.; Gary-Bobo, M.; Esclangon, A.; Maffrand, J.P.; Le Fur, G.; Oury-Donat, F.; Soubrié, P. The Cannabinoid CB 1 Receptor Antagonist SR141716 Increases Acrp30 MRNA Expression in Adipose Tissue of Obese Fa/Fa Rats and in Cultured Adipocyte Cells. Mol. Pharmacol. 2003, 63, 908–914. [Google Scholar] [CrossRef]
- Matias, I.; Gonthier, M.-P.; Orlando, P.; Martiadis, V.; Petrocellis, L.; De Cervino, C.; Petrosino, S.; Hoareau, L.; Festy, F.; Pasquali, R.; et al. Regulation, Function, and Dysregulation of Endocannabinoids in Models of Adipose and β-Pancreatic Cells and in Obesity and Hyperglycemia. J. Clin. Endocrinol. Metab. 2006, 91, 3171–3180. [Google Scholar] [CrossRef] [PubMed]
- Matias, I.; Belluomo, I.; Cota, D. The Fat Side of the Endocannabinoid System: Role of Endocannabinoids in the Adipocyte. Cannabis Cannabinoid Res. 2016, 1, 176–185. [Google Scholar] [CrossRef]
- Tedesco, L.; Valerio, A.; Dossena, M.; Cardile, A.; Ragni, M.; Pagano, C.; Pagotto, U.; Carruba, M.O.; Vettor, R.; Nisoli, E. Cannabinoid Receptor Stimulation Impairs Mitochondrial Biogenesis in Mouse White Adipose Tissue, Muscle, and Liver: The Role of ENOS, P38 MAPK, and AMPK Pathways. Diabetes 2010, 59, 2826–2836. [Google Scholar] [CrossRef] [PubMed]
- Mølhøj, S.; Hansen, H.S.; Schweiger, M.; Zimmermann, R.; Johansen, T.; Malmlöf, K. Effect of the Cannabinoid Receptor-1 Antagonist Rimonabant on Lipolysis in Rats. Eur. J. Pharmacol. 2010, 646, 38–45. [Google Scholar] [CrossRef]
- Bajzer, M.; Olivieri, M.; Haas, M.K.; Pfluger, P.T.; Magrisso, I.J.; Foster, M.T.; Tschöp, M.H.; Krawczewski-Carhuatanta, K.A.; Cota, D.; Obici, S. Cannabinoid Receptor 1 (CB1) Antagonism Enhances Glucose Utilisation and Activates Brown Adipose Tissue in Diet-Induced Obese Mice. Diabetologia 2011, 54, 3121–3131. [Google Scholar] [CrossRef]
- Ruiz de Azua, I.; Mancini, G.; Srivastava, R.K.; Rey, A.A.; Cardinal, P.; Tedesco, L.; Zingaretti, C.M.; Sassmann, A.; Quarta, C.; Schwitter, C.; et al. Adipocyte Cannabinoid Receptor CB1 Regulates Energy Homeostasis and Alternatively Activated Macrophages. J. Clin. Invest. 2017, 127, 4148–4162. [Google Scholar] [CrossRef]
- Bénard, G.; Massa, F.; Puente, N.; Lourenço, J.; Bellocchio, L.; Soria-Gómez, E.; Matias, I.; Delamarre, A.; Metna-Laurent, M.; Cannich, A.; et al. Mitochondrial CB1 Receptors Regulate Neuronal Energy Metabolism. Nat. Neurosci. 2012, 15, 558–564. [Google Scholar] [CrossRef]
- Hebert-Chatelain, E.; Desprez, T.; Serrat, R.; Bellocchio, L.; Soria-Gomez, E.; Busquets-Garcia, A.; Pagano Zottola, A.C.; Delamarre, A.; Cannich, A.; Vincent, P.; et al. A Cannabinoid Link between Mitochondria and Memory. Nature 2016, 539, 555–559. [Google Scholar] [CrossRef]
- Mendizabal-Zubiaga, J.; Melser, S.; Bénard, G.; Ramos, A.; Reguero, L.; Arrabal, S.; Elezgarai, I.; Gerrikagoitia, I.; Suarez, J.; Rodríguez De Fonseca, F.; et al. Cannabinoid CB1 Receptors Are Localized in Striated Muscle Mitochondria and Regulate Mitochondrial Respiration. Front. Physiol. 2016, 7, 476. [Google Scholar] [CrossRef]
- Koch, M.; Varela, L.; Kim, J.G.; Kim, J.D.; Hernández-Nuño, F.; Simonds, S.E.; Castorena, C.M.; Vianna, C.R.; Elmquist, J.K.; Morozov, Y.M.; et al. Hypothalamic POMC Neurons Promote Cannabinoid-Induced Feeding. Nature 2015, 519, 45–50. [Google Scholar] [CrossRef]
- Jimenez-Blasco, D.; Busquets-Garcia, A.; Hebert-Chatelain, E.; Serrat, R.; Vicente-Gutierrez, C.; Ioannidou, C.; Gómez-Sotres, P.; Lopez-Fabuel, I.; Resch-Beusher, M.; Resel, E.; et al. Glucose Metabolism Links Astroglial Mitochondria to Cannabinoid Effects. Nature 2020, 583, 603–608. [Google Scholar] [CrossRef] [PubMed]
- Soria-Gomez, E.; Pagano Zottola, A.C.; Mariani, Y.; Desprez, T.; Barresi, M.; Bonilla-del Río, I.; Muguruza, C.; Le Bon-Jego, M.; Julio-Kalajzić, F.; Flynn, R.; et al. Subcellular Specificity of Cannabinoid Effects in Striatonigral Circuits. Neuron 2021, 109, 1513–1526.e11. [Google Scholar] [CrossRef] [PubMed]
- Aquila, S.; Guido, C.; Santoro, A.; Perrotta, I.; Laezza, C.; Bifulco, M.; Sebastiano, A. Human Sperm Anatomy: Ultrastructural Localization of the Cannabinoid1 Receptor and a Potential Role of Anandamide in Sperm Survival and Acrosome Reaction. Anat. Rec. Adv. Integr. Anat. Evol. Biol. 2010, 293, 298–309. [Google Scholar] [CrossRef] [PubMed]
- Kamnate, A.; Sirisin, J.; Watanabe, M.; Kondo, H.; Hipkaeo, W.; Chomphoo, S. Mitochondrial Localization of CB1 in Progesterone-Producing Cells of Ovarian Interstitial Glands of Adult Mice. J. Histochem. Cytochem. 2022, 70, 251–257. [Google Scholar] [CrossRef] [PubMed]
- Marsicano, G.; Wotjak, C.T.; Azad, S.C.; Bisogno, T.; Rammes, G.; Cascio, M.G.; Hermann, H.; Tang, J.; Hofmann, C.; Zieglgänsberger, W.; et al. The Endogenous Cannabinoid System Controls Extinction of Aversive Memories. Nature 2002, 418, 530–534. [Google Scholar] [CrossRef]
- Pettit, D.A.D.; Harrison, M.P.; Olson, J.M.; Spencer, R.F.; Cabral, G.A. Immunohistochemical Localization of the Neural Cannabinoid Receptor in Rat Brain. J. Neurosci. Res. 1998, 51, 391–402. [Google Scholar] [CrossRef]
- Phend, K.D.; Rustioni, A.; Weinberg, R.J. An Osmium-Free Method of Epon Embedment That Preserves Both Ultrastructure and Antigenicity for Post-Embedding Immunocytochemistry. J. Histochem. Cytochem. 1995, 43, 283–292. [Google Scholar] [CrossRef]
- Melone, M.; Bellesi, M.; Conti, F. Synaptic Localization of GLT-1a in the Rat Somatic Sensory Cortex. Glia 2009, 57, 108–117. [Google Scholar] [CrossRef]
- Makrecka-Kuka, M.; Krumschnabel, G.; Gnaiger, E. High-Resolution Respirometry for Simultaneous Measurement of Oxygen and Hydrogen Peroxide Fluxes in Permeabilized Cells, Tissue Homogenate and Isolated Mitochondria. Biomolecules 2015, 5, 1319–1338. [Google Scholar] [CrossRef]
- Shabalina, I.G.; Petrovic, N.; de Jong, J.M.A.; Kalinovich, A.V.; Cannon, B.; Nedergaard, J. UCP1 in Brite/Beige Adipose Tissue Mitochondria Is Functionally Thermogenic. Cell Rep. 2013, 5, 1196–1203. [Google Scholar] [CrossRef]
- Blüher, M.; Engeli, S.; Klöting, N.; Berndt, J.; Fasshauer, M.; Bátkai, S.; Pacher, P.; Schön, M.R.; Jordan, J.; Stumvoll, M. Dysregulation of the Peripheral and Adipose Tissue Endocannabinoid System in Human Abdominal Obesity. Diabetes 2006, 55, 3053–3060. [Google Scholar] [CrossRef] [PubMed]
- Rakotoarivelo, V.; Sihag, J.; Flamand, N. Role of the Endocannabinoid System in the Adipose Tissue with Focus on Energy Metabolism. Cells 2021, 10, 1279. [Google Scholar] [CrossRef] [PubMed]
- Quarta, C.; Bellocchio, L.; Mancini, G.; Mazza, R.; Cervino, C.; Braulke, L.J.; Fekete, C.; Latorre, R.; Nanni, C.; Bucci, M.; et al. CB(1) Signaling in Forebrain and Sympathetic Neurons Is a Key Determinant of Endocannabinoid Actions on Energy Balance. Cell Metab. 2010, 11, 273–285. [Google Scholar] [CrossRef] [PubMed]
- Vettor, R.; Pagano, C. The Role of the Endocannabinoid System in Lipogenesis and Fatty Acid Metabolism. Best Pract. Res. Clin. Endocrinol. Metab. 2009, 23, 51–63. [Google Scholar] [CrossRef]
- Stephens, M.; Ludgate, M.; Rees, D.A. Brown Fat and Obesity: The next Big Thing? Clin. Endocrinol. 2011, 74, 661–670. [Google Scholar] [CrossRef]
- Fišar, Z.; Singh, N.; Hroudová, J. Cannabinoid-Induced Changes in Respiration of Brain Mitochondria. Toxicol. Lett. 2014, 231, 62–71. [Google Scholar] [CrossRef]
- Cinti, S. The Adipose Organ. Prostaglandins, Leukot. Essent. Fat. Acids 2005, 73, 9–15. [Google Scholar] [CrossRef]
- Hebert-Chatelain, E.; Reguero, L.; Puente, N.; Lutz, B.; Chaouloff, F.; Rossignol, R.; Piazza, P.-V.; Benard, G.; Grandes, P.; Marsicano, G. Cannabinoid Control of Brain Bioenergetics: Exploring the Subcellular Localization of the CB1 Receptor. Mol. Metab. 2014, 3, 495–504. [Google Scholar] [CrossRef]
- Micakovic, T.; Banczyk, W.; Clark, E.; Kränzlin, B.; Peters, J.; Hoffmann, S. Isolation of Pure Mitochondria from Rat Kidneys and Western Blot of Mitochondrial Respiratory Chain Complexes. BIO-PROTOCOL 2019, 9, e3379. [Google Scholar] [CrossRef]
- Melser, S.; Pagano Zottola, A.C.; Serrat, R.; Puente, N.; Grandes, P.; Marsicano, G.; Hebert-Chatelain, E. Functional Analysis of Mitochondrial CB1 Cannabinoid Receptors (MtCB1) in the Brain. Methods Enzymol. 2017, 593, 143–174. [Google Scholar]
- Valsecchi, F.; Ramos-Espiritu, L.S.; Buck, J.; Levin, L.R.; Manfredi, G. CAMP and Mitochondria. Physiology 2013, 28, 199–209. [Google Scholar] [CrossRef] [PubMed]
- Ould Amer, Y.; Hebert-Chatelain, E. Mitochondrial CAMP-PKA Signaling: What Do We Really Know? Biochim. Biophys. Acta-Bioenerg. 2018, 1859, 868–877. [Google Scholar] [CrossRef] [PubMed]
- Acin-Perez, R.; Salazar, E.; Kamenetsky, M.; Buck, J.; Levin, L.R.; Manfredi, G. Cyclic AMP Produced inside Mitochondria Regulates Oxidative Phosphorylation. Cell Metab. 2009, 9, 265–276. [Google Scholar] [CrossRef] [PubMed]
- Di Benedetto, G.; Scalzotto, E.; Mongillo, M.; Pozzan, T. Mitochondrial Ca2+ Uptake Induces Cyclic AMP Generation in the Matrix and Modulates Organelle ATP Levels. Cell Metab. 2013, 17, 965–975. [Google Scholar] [CrossRef] [PubMed]
- Jakobsen, E.; Andersen, J.V.; Christensen, S.K.; Siamka, O.; Larsen, M.R.; Waagepetersen, H.S.; Aldana, B.I.; Bak, L.K. Pharmacological Inhibition of Mitochondrial Soluble Adenylyl Cyclase in Astrocytes Causes Activation of AMP-activated Protein Kinase and Induces Breakdown of Glycogen. Glia 2021, 69, 2828–2844. [Google Scholar] [CrossRef] [PubMed]
- Tam, J.; Cinar, R.; Liu, J.; Godlewski, G.; Wesley, D.; Jourdan, T.; Szanda, G.; Mukhopadhyay, B.; Chedester, L.; Liow, J.-S.; et al. Peripheral Cannabinoid-1 Receptor Inverse Agonism Reduces Obesity by Reversing Leptin Resistance. Cell Metab. 2012, 16, 167–179. [Google Scholar] [CrossRef]
- Wang, Y.; Adjaye, J. A Cyclic AMP Analog, 8-Br-CAMP, Enhances the Induction of Pluripotency in Human Fibroblast Cells. Stem Cell Rev. Rep. 2011, 7, 331–341. [Google Scholar] [CrossRef]
- Cota, D.; Marsicano, G.; Tschöp, M.; Grübler, Y.; Flachskamm, C.; Schubert, M.; Auer, D.; Yassouridis, A.; Thöne-Reineke, C.; Ortmann, S.; et al. The Endogenous Cannabinoid System Affects Energy Balance via Central Orexigenic Drive and Peripheral Lipogenesis. J. Clin. Invest. 2003, 112, 423–431. [Google Scholar] [CrossRef]
- D’Eon, T.M.; Pierce, K.A.; Roix, J.J.; Tyler, A.; Chen, H.; Teixeira, S.R. The Role of Adipocyte Insulin Resistance in the Pathogenesis of Obesity-Related Elevations in Endocannabinoids. Diabetes 2008, 57, 1262–1268. [Google Scholar] [CrossRef]
- Krott, L.M.; Piscitelli, F.; Heine, M.; Borrino, S.; Scheja, L.; Silvestri, C.; Heeren, J.; Di Marzo, V. Endocannabinoid Regulation in White and Brown Adipose Tissue Following Thermogenic Activation. J. Lipid Res. 2016, 57, 464–473. [Google Scholar] [CrossRef]
- Pagano, C.; Rossato, M.; Vettor, R. Endocannabinoids, Adipose Tissue and Lipid Metabolism. J. Neuroendocrinol. 2008, 20, 124–129. [Google Scholar] [CrossRef] [PubMed]
- Jbilo, O.; Ravinet-Trillou, C.; Arnone, M.; Buisson, I.; Bribes, E.; Péleraux, A.; Pénarier, G.; Soubrié, P.; Le Fur, G.; Galiègue, S.; et al. The CB1 Receptor Antagonist Rimonabant Reverses the Diet-induced Obesity Phenotype through the Regulation of Lipolysis and Energy Balance. FASEB J. 2005, 19, 1567–1569. [Google Scholar] [CrossRef] [PubMed]
- Perwitz, N.; Wenzel, J.; Wagner, I.; Büning, J.; Drenckhan, M.; Zarse, K.; Ristow, M.; Lilienthal, W.; Lehnert, H.; Klein, J. Cannabinoid Type 1 Receptor Blockade Induces Transdifferentiation towards a Brown Fat Phenotype in White Adipocytes. Diabetes, Obes. Metab. 2010, 12, 158–166. [Google Scholar] [CrossRef] [PubMed]
- Addy, C.; Wright, H.; Van Laere, K.; Gantz, I.; Erondu, N.; Musser, B.J.; Lu, K.; Yuan, J.; Sanabria-Bohórquez, S.M.; Stoch, A.; et al. The Acyclic CB1R Inverse Agonist Taranabant Mediates Weight Loss by Increasing Energy Expenditure and Decreasing Caloric Intake. Cell Metab. 2008, 7, 68–78. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez-Rodríguez, A.; Bonilla-Del Río, I.; Puente, N.; Gómez-Urquijo, S.M.; Fontaine, C.J.; Egaña-Huguet, J.; Elezgarai, I.; Ruehle, S.; Lutz, B.; Robin, L.M.; et al. Localization of the Cannabinoid Type-1 Receptor in Subcellular Astrocyte Compartments of Mutant Mouse Hippocampus. Glia 2018, 66, 1417–1431. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez-Rodríguez, A.; Puente, N.; Elezgarai, I.; Ruehle, S.; Lutz, B.; Reguero, L.; Gerrikagoitia, I.; Marsicano, G.; Grandes, P. Anatomical Characterization of the Cannabinoid CB 1 Receptor in Cell-Type-Specific Mutant Mouse Rescue Models. J. Comp. Neurol. 2017, 525, 302–318. [Google Scholar] [CrossRef]
- Lipina, C.; Irving, A.J.; Hundal, H.S. Mitochondria: A Possible Nexus for the Regulation of Energy Homeostasis by the Endocannabinoid System? Am. J. Physiol. Metab. 2014, 307, E1–E13. [Google Scholar] [CrossRef]
- London, E.; Bloyd, M.; Stratakis, C.A. PKA Functions in Metabolism and Resistance to Obesity: Lessons from Mouse and Human Studies. J. Endocrinol. 2020, 246, R51–R64. [Google Scholar] [CrossRef]
- Yehuda-Shnaidman, E.; Buehrer, B.; Pi, J.; Kumar, N.; Collins, S. Acute Stimulation of White Adipocyte Respiration by PKA-Induced Lipolysis. Diabetes 2010, 59, 2474–2483. [Google Scholar] [CrossRef]
- Dickson, L.M.; Gandhi, S.; Layden, B.T.; Cohen, R.N.; Wicksteed, B. Protein Kinase A Induces UCP1 Expression in Specific Adipose Depots to Increase Energy Expenditure and Improve Metabolic Health. Am. J. Physiol. Integr. Comp. Physiol. 2016, 311, R79–R88. [Google Scholar] [CrossRef]
- Boon, M.R.; Kooijman, S.; van Dam, A.D.; Pelgrom, L.R.; Berbée, J.F.P.; Visseren, C.A.R.; van Aggele, R.C.; van den Hoek, A.M.; Sips, H.C.M.; Lombès, M.; et al. Peripheral Cannabinoid 1 Receptor Blockade Activates Brown Adipose Tissue and Diminishes Dyslipidemia and Obesity. FASEB J. 2014, 28, 5361–5375. [Google Scholar] [CrossRef] [PubMed]
- Müller, G.A.; Herling, A.W.; Wied, S.; Müller, T.D. CB1 Receptor-Dependent and Independent Induction of Lipolysis in Primary Rat Adipocytes by the Inverse Agonist Rimonabant (SR141716A). Molecules 2020, 25, 896. [Google Scholar] [CrossRef] [PubMed]
- Omar, M.H.; Scott, J.D. AKAP Signaling Islands: Venues for Precision Pharmacology. Trends Pharmacol. Sci. 2020, 41, 933–946. [Google Scholar] [CrossRef] [PubMed]
- Bridges, D.; MacDonald, J.A.; Wadzinski, B.; Moorhead, G.B.G. Identification and Characterization of D-AKAP1 as a Major Adipocyte PKA and PP1 Binding Protein. Biochem. Biophys. Res. Commun. 2006, 346, 351–357. [Google Scholar] [CrossRef] [PubMed]
- Pidoux, G.; Witczak, O.; Jarnaess, E.; Myrvold, L.; Urlaub, H.; Stokka, A.J.; Küntziger, T.; Taskén, K. Optic Atrophy 1 Is an A-Kinase Anchoring Protein on Lipid Droplets That Mediates Adrenergic Control of Lipolysis. EMBO J. 2011, 30, 4371–4386. [Google Scholar] [CrossRef]
- Vergnes, L.; Lin, J.Y.; Davies, G.R.; Church, C.D.; Reue, K. Induction of UCP1 and Thermogenesis by a Small Molecule via AKAP1/PKA Modulation. J. Biol. Chem. 2020, 295, 15054–15069. [Google Scholar] [CrossRef]
- Luo, L.; Liu, M. Adipose Tissue in Control of Metabolism. J. Endocrinol. 2016, 231, R77–R99. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pagano Zottola, A.C.; Severi, I.; Cannich, A.; Ciofi, P.; Cota, D.; Marsicano, G.; Giordano, A.; Bellocchio, L. Expression of Functional Cannabinoid Type-1 (CB1) Receptor in Mitochondria of White Adipocytes. Cells 2022, 11, 2582. https://doi.org/10.3390/cells11162582
Pagano Zottola AC, Severi I, Cannich A, Ciofi P, Cota D, Marsicano G, Giordano A, Bellocchio L. Expression of Functional Cannabinoid Type-1 (CB1) Receptor in Mitochondria of White Adipocytes. Cells. 2022; 11(16):2582. https://doi.org/10.3390/cells11162582
Chicago/Turabian StylePagano Zottola, Antonio C., Ilenia Severi, Astrid Cannich, Philippe Ciofi, Daniela Cota, Giovanni Marsicano, Antonio Giordano, and Luigi Bellocchio. 2022. "Expression of Functional Cannabinoid Type-1 (CB1) Receptor in Mitochondria of White Adipocytes" Cells 11, no. 16: 2582. https://doi.org/10.3390/cells11162582
APA StylePagano Zottola, A. C., Severi, I., Cannich, A., Ciofi, P., Cota, D., Marsicano, G., Giordano, A., & Bellocchio, L. (2022). Expression of Functional Cannabinoid Type-1 (CB1) Receptor in Mitochondria of White Adipocytes. Cells, 11(16), 2582. https://doi.org/10.3390/cells11162582





