Comprehensive Molecular Landscape of Cetuximab Resistance in Head and Neck Cancer Cell Lines
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Lines and Cetuximab Resistance Model Development
2.2. Chromosome Preparation and G-Banding
2.3. DNA Isolation and Whole Exome Sequencing (WES) Analysis
2.4. Copy Number Alterations (CNA)
2.5. Cell Surface Markers Screening
2.6. Western Blot, Human RTK, Subcellular Fraction Separation, and Cytokines Arrays
2.7. mRNA NanoStringTM Analysis
2.8. Expression Microarray
2.9. Sanger Sequencing
2.10. Cell Viability Assay
2.11. Morphological Data
2.12. Wound Healing Migration Assay
2.13. Colony Formation Assay Anchorage-Dependent
2.14. Adhesion Assay
2.15. Immunofluorescence Analysis
2.16. Statistical Analysis
3. Results
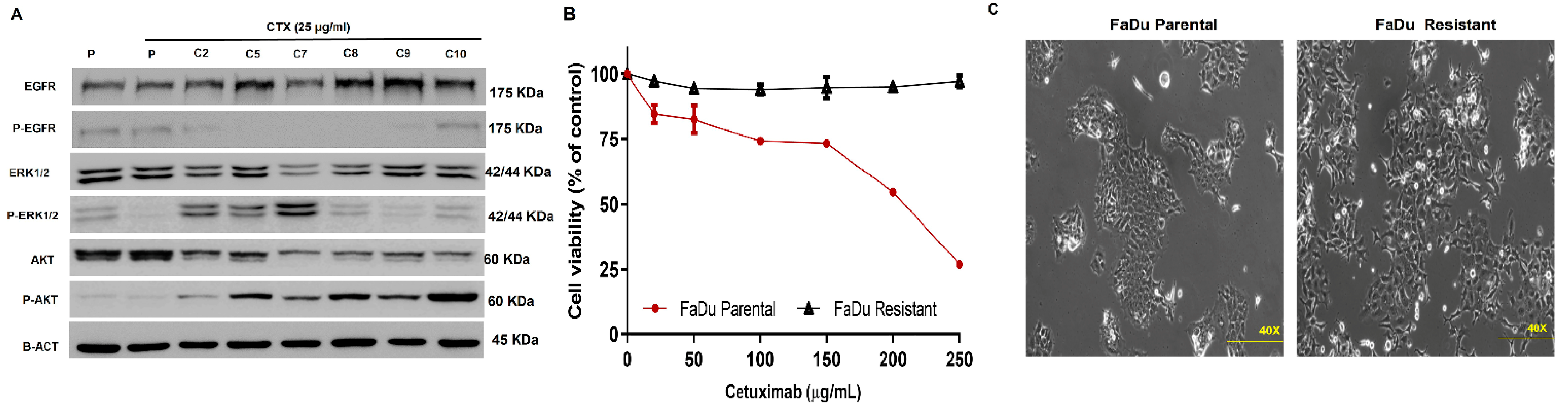
3.1. Cetuximab-Resistance Model Establishment and Characterization
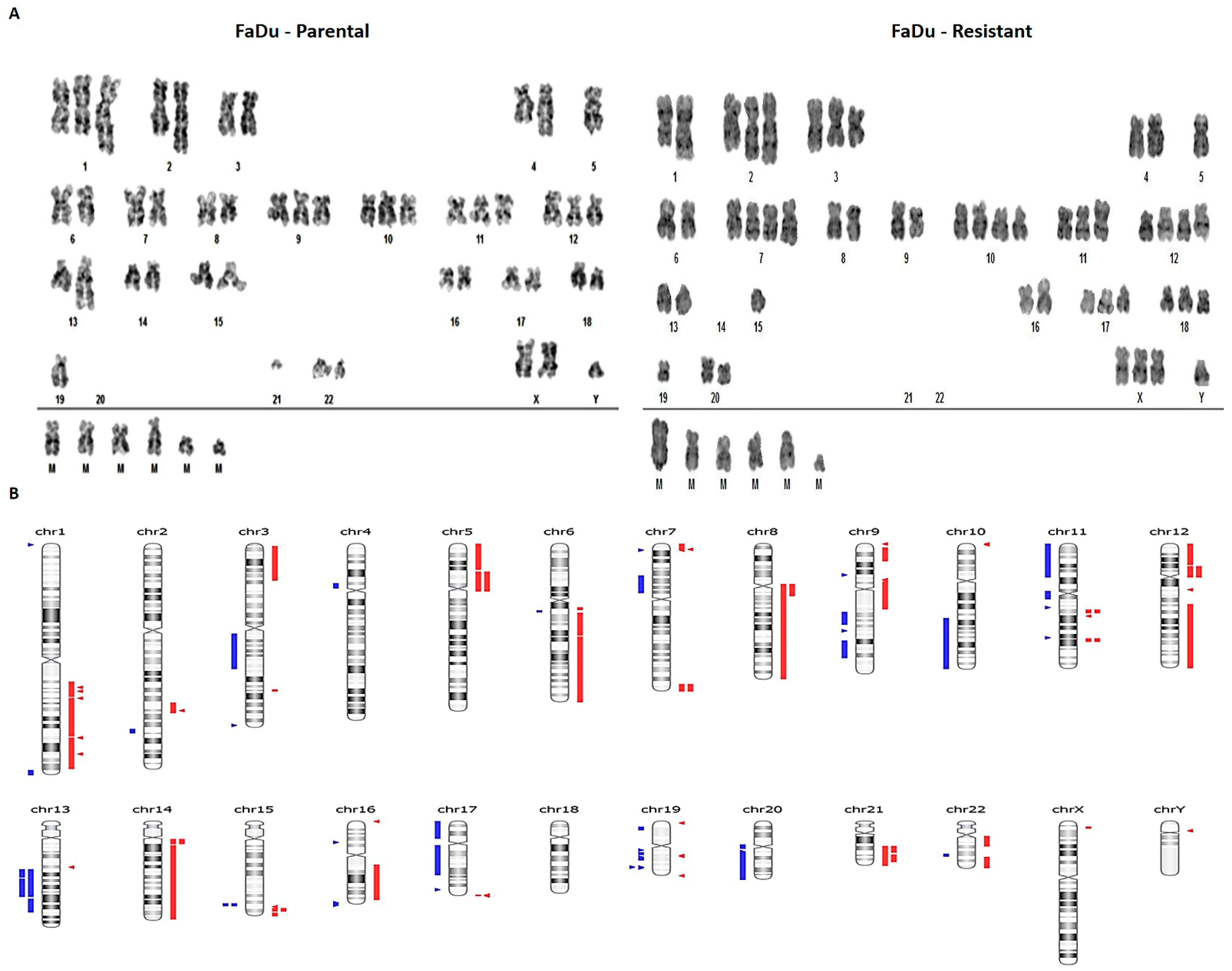
3.2. Cetuximab Resistance Is Associated with Chromosomal Abnormalities
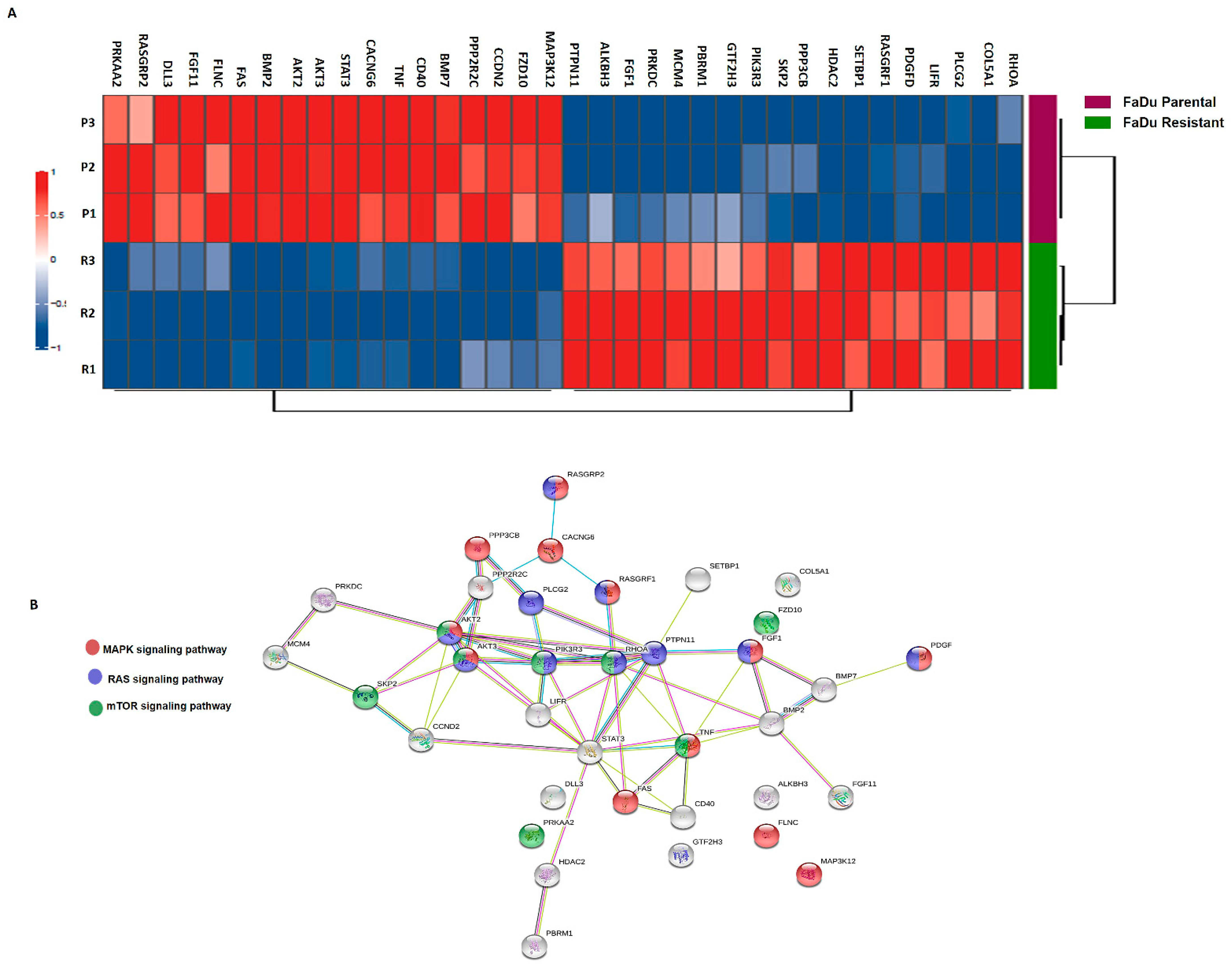
3.3. Differential Gene Expression and Mutation Profile
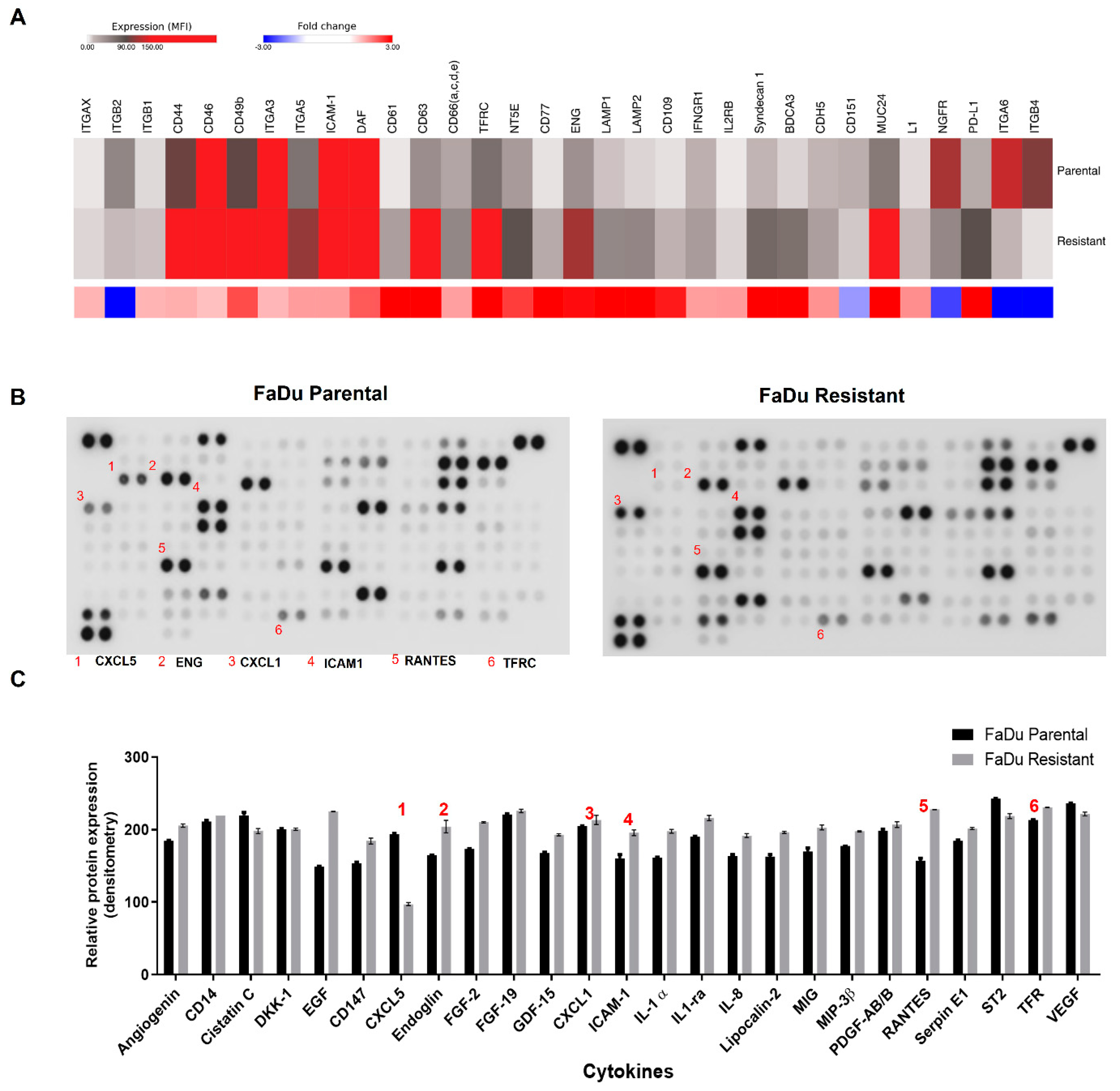
3.4. Differential Protein Profile
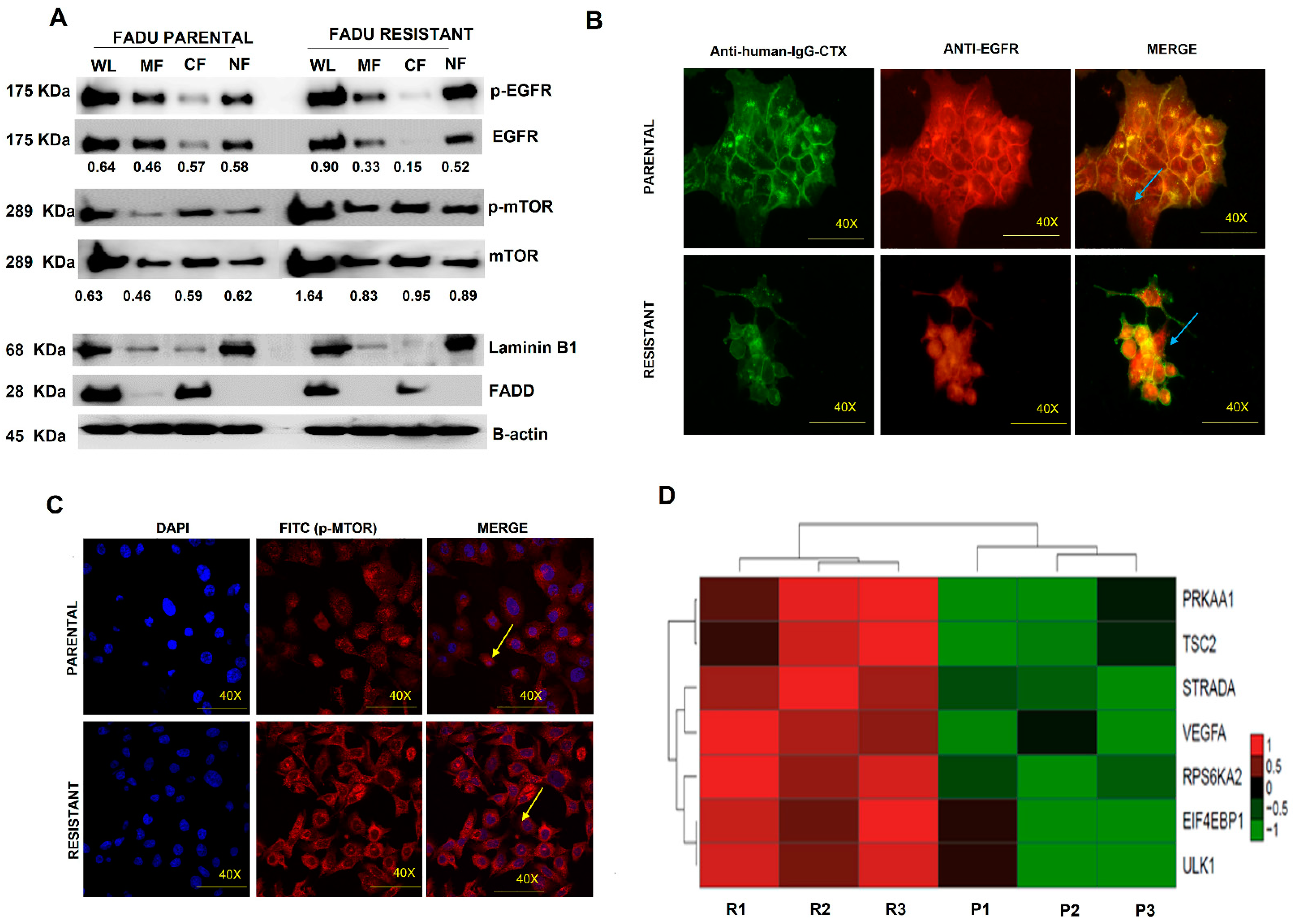
3.5. EGFR Nuclear Translocation and mTOR Overexpression Are Present in Cetuximab-Resistant Cells
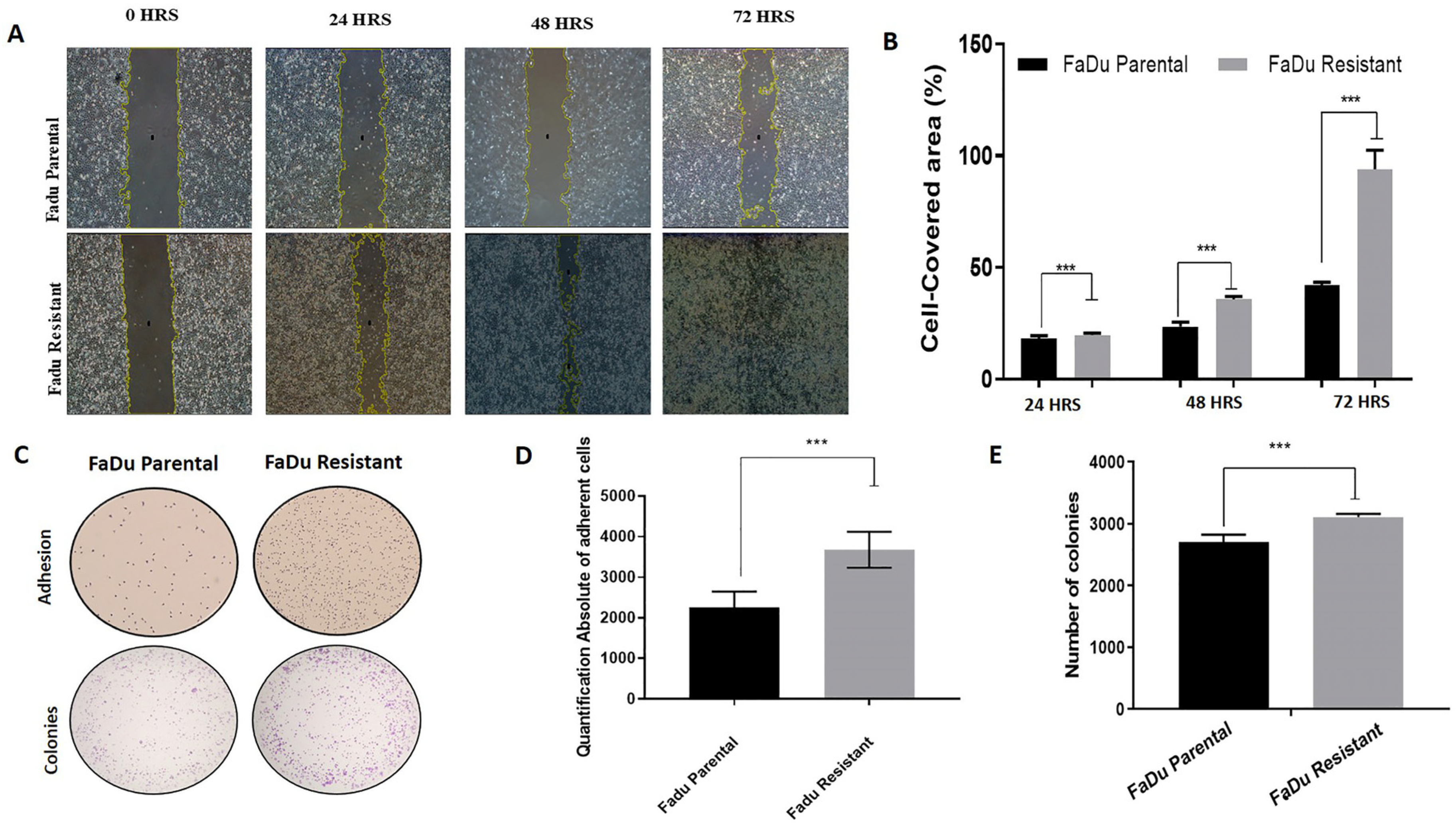
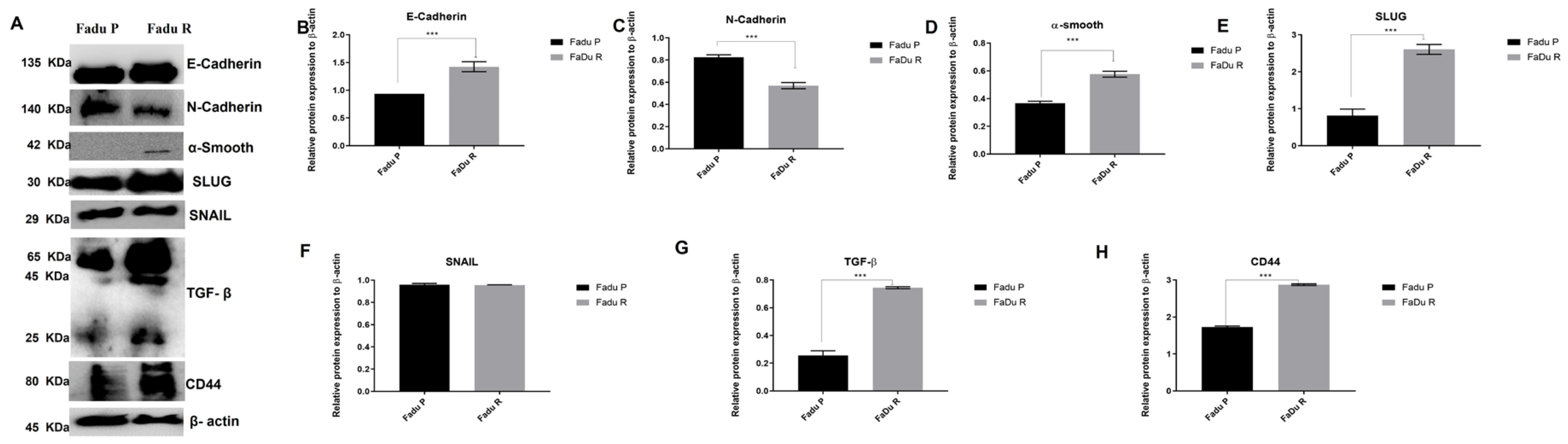
3.6. Increased Aggressiveness Phenotype and Differential Expression of Epithelial–Mesenchymal Transition Markers (EMT) in Cetuximab Resistant Cells
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Cramer, J.D.; Burtness, B.; Le, Q.T.; Ferris, R.L. The changing therapeutic landscape of head and neck cancer. Nat. Rev. Clin. Oncol. 2019, 16, 669–683. [Google Scholar] [CrossRef] [PubMed]
- Alsahafi, E.; Begg, K.; Amelio, I.; Raulf, N.; Lucarelli, P.; Sauter, T.; Tavassoli, M. Clinical update on head and neck cancer: Molecular biology and ongoing challenges. Cell Death Dis. 2019, 10, 1–17. [Google Scholar] [CrossRef]
- Leemans, C.R.; Snijders, P.J.F.; Brakenhoff, R.H. The molecular landscape of head and neck cancer. Nat. Rev. Cancer 2018, 18, 269–282. [Google Scholar] [CrossRef] [PubMed]
- de Carvalho, A.C.; Perdomo, S.; dos Santos, W.; Fernandes, G.C.; de Jesus, L.M.; Carvalho, R.S.; Scapulatempo-Neto, C.; de Almeida, G.C.; Sorroche, B.P.; Arantes, L.M.R.B.; et al. Impact of genetic variants in clinical outcome of a cohort of patients with oropharyngeal squamous cell carcinoma. Sci. Rep. 2020, 10, 1–12. [Google Scholar] [CrossRef]
- Oliveira-Silva, R.J.; De Carvalho, A.C.; Viana, L.D.S.; Carvalho, A.L.; Reis, R. Anti-EGFR Therapy: Strategies in Head and Neck Squamous Cell Carcinoma. Recent Patents Anti-Cancer Drug Discov. 2016, 11, 170–183. [Google Scholar] [CrossRef]
- Brand, T.M.; Iida, M.; Luthar, N.; Starr, M.M.; Huppert, E.J.; Wheeler, D.L. Corrigendum to: “Nuclear EGFR as a molecular target in cancer” [Radiother Oncol 108 (2013) 370–77]. Radiother. Oncol. 2019, 130, 195. [Google Scholar] [CrossRef]
- Moreira, J.; Tobias, A.; O’Brien, M.P.; Agulnik, M. Targeted Therapy in Head and Neck Cancer: An Update on Current Clinical Developments in Epidermal Growth Factor Receptor-Targeted Therapy and Immunotherapies. Drugs 2017, 77, 843–857. [Google Scholar] [CrossRef]
- Azoury, S.C.; Gilmore, R.C.; Shukla, V. Molecularly targeted agents and immunotherapy for the treatment of head and neck squamous cell cancer (HNSCC). Discov. Med. 2016, 21, 507–515. [Google Scholar]
- Vermorken, J.B.; Trigo, J.; Hitt, R.; Koralewski, P.; Diaz-Rubio, E.; Rolland, F.; Knecht, R.; Amellal, N.; Schueler, A.; Baselga, J. Open-Label, Uncontrolled, Multicenter Phase II Study to Evaluate the Efficacy and Toxicity of Cetuximab As a Single Agent in Patients with Recurrent and/or Metastatic Squamous Cell Carcinoma of the Head and Neck Who Failed to Respond to Platinum-Based Therapy. J. Clin. Oncol. 2007, 25, 2171–2177. [Google Scholar] [CrossRef]
- Vermorken, J.B.; Mesia, R.; Rivera, F.; Remenar, E.; Kawecki, A.; Rottey, S.; Erfan, J.; Zabolotnyy, D.; Kienzer, H.-R.; Cupissol, D.; et al. Platinum-Based Chemotherapy plus Cetuximab in Head and Neck Cancer. N. Engl. J. Med. 2008, 359, 1116–1127. [Google Scholar] [CrossRef]
- Bonner, J.A.; Harari, P.M.; Giralt, J.; Azarnia, N.; Shin, D.M.; Cohen, R.B.; Jones, C.U.; Sur, R.; Raben, D.; Jassem, J.; et al. Radiotherapy plus Cetuximab for Squamous-Cell Carcinoma of the Head and Neck. N. Engl. J. Med. 2006, 354, 567–578. [Google Scholar] [CrossRef] [PubMed]
- Leblanc, O.; Vacher, S.; Lecerf, C.; Jeannot, E.; Klijanienko, J.; Berger, F.; Hoffmann, C.; Calugaru, V.; Badois, N.; Chilles, A.; et al. Biomarkers of cetuximab resistance in patients with head and neck squamous cell carcinoma. Cancer Biol. Med. 2020, 17, 208–217. [Google Scholar] [CrossRef] [PubMed]
- Mehra, R.; Cohen, R.B.; Burtness, B.A. The role of cetuximab for the treatment of squamous cell carcinoma of the head and neck. Clin. Adv. Hematol. Oncol. H&O 2008, 6, 742–750. [Google Scholar]
- Silva-Oliveira, R.J.; Melendez, M.; Martinho, O.; Zanon, M.F.; Viana, L.D.S.; Carvalho, A.L.; Reis, A.R.M. AKT can modulate the in vitro response of HNSCC cells to irreversible EGFR inhibitors. Oncotarget 2017, 8, 53288–53301. [Google Scholar] [CrossRef] [PubMed]
- Singchat, W.; Hitakomate, E.; Rerkarmnuaychoke, B.; Suntronpong, A.; Fu, B.; Bodhisuwan, W.; Peyachoknagul, S.; Yang, F.; Koontongkaew, S.; Srikulnath, K. Genomic Alteration in Head and Neck Squamous Cell Carcinoma (HNSCC) Cell Lines Inferred from Karyotyping, Molecular Cytogenetics, and Array Comparative Genomic Hybridization. PLoS ONE 2016, 11, e0160901. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 2010, 26, 589–595. [Google Scholar] [CrossRef] [PubMed]
- Koboldt, D.C.; Larson, D.E.; Wilson, R.K. Using VarScan 2 for Germline Variant Calling and Somatic Mutation Detection. Curr. Protoc. Bioinform. 2013, 44, 15.4.1–15.4.17. [Google Scholar] [CrossRef] [PubMed]
- Forbes, S.A.; Beare, D.; Boutselakis, H.; Bamford, S.; Bindal, N.; Tate, J.; Cole, C.G.; Ward, S.; Dawson, E.; Ponting, L.; et al. COSMIC: Somatic cancer genetics at high-resolution. Nucleic Acids Res. 2017, 45, D777–D783. [Google Scholar] [CrossRef] [PubMed]
- Benjamin, D.; Sato, T.; Cibulskis, K.; Getz, G.; Stewart, C.; Lichtenstein, L. Calling Somatic SNVs and Indels with Mutect. BioRxiv 2019, 861054. [Google Scholar] [CrossRef]
- McLaren, W.; Gil, L.; Hunt, S.E.; Riat, H.S.; Ritchie, G.R.S.; Thormann, A.; Flicek, P.; Cunningham, F. The Ensembl Variant Effect Predictor. Genome Biol. 2016, 17, 1–14. [Google Scholar] [CrossRef]
- 1000 Genomes Project Consortium; Auton, A.; Brooks, L.D.; Durbin, R.M.; Garrison, E.P.; Kang, H.M.; Korbel, J.O.; Marchini, J.L.; McCarthy, S.; McVean, G.A.; et al. A global reference for human genetic variation. Nature 2015, 526, 68–74. [Google Scholar]
- Lek, M.; Karczewski, K.J.; Minikel, E.V.; Samocha, K.E.; Banks, E.; Fennell, T.; O’Donnell-Luria, A.H.; Ware, J.S.; Hill, A.J.; Cummings, B.B.; et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 2016, 536, 285–291. [Google Scholar] [CrossRef]
- Robinson, J.T.; Thorvaldsdóttir, H.; Winckler, W.; Guttman, M.; Lander, E.S.; Getz, G.; Mesirov, J.P. Integrative genomics viewer. Nat. Biotechnol. 2011, 29, 24–26. [Google Scholar] [CrossRef] [PubMed]
- Rosa, M.N.; Evangelista, A.F.; Leal, L.F.; De Oliveira, C.M.; Silva, V.A.O.; Munari, C.C.; Munari, F.F.; Matsushita, G.D.M.; Dos Reis, R.; Andrade, C.E.; et al. Establishment, molecular and biological characterization of HCB-514: A novel human cervical cancer cell line. Sci. Rep. 2019, 9, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Waggott, D.; Chu, K.; Yin, S.; Wouters, B.G.; Liu, F.-F.; Boutros, P.C. NanoStringNorm: An extensible R package for the pre-processing of NanoString mRNA and miRNA data. Bioinformatics 2012, 28, 1546–1548. [Google Scholar] [CrossRef]
- Gu, Z.; Eils, R.; Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 2016, 32, 2847–2849. [Google Scholar] [CrossRef] [PubMed]
- Vicente, A.L.S.; Crovador, C.S.; Macedo, G.; Scapulatempo-Neto, C.; Reis, R.M.; Vazquez, V.L. Mutational Profile of Driver Genes in Brazilian Melanomas. J. Glob. Oncol. 2019, 5, 1–14. [Google Scholar] [CrossRef]
- Martinho, O.; Silva-Oliveira, R.; Miranda-Gonçalves, V.; Clara, C.; Almeida, J.R.; Carvalho, A.L.; Barata, J.T.; Reis, R.M. In Vitro and In Vivo Analysis of RTK Inhibitor Efficacy and Identification of Its Novel Targets in Glioblastomas. Transl. Oncol. 2013, 6, 187–196, IN17–IN20. [Google Scholar] [CrossRef]
- Geissmann, Q. OpenCFU, a New Free and Open-Source Software to Count Cell Colonies and Other Circular Objects. PLoS ONE 2013, 8, e54072. [Google Scholar] [CrossRef]
- Salo, T.; Sutinen, M.; Apu, E.H.; Sundquist, E.; Cervigne, N.K.; De Oliveira, C.E.; Akram, S.U.; Ohlmeier, S.; Suomi, F.; Eklund, L.; et al. A novel human leiomyoma tissue derived matrix for cell culture studies. BMC Cancer 2015, 15, 981. [Google Scholar] [CrossRef] [PubMed]
- Suomela, S.; Elomaa, O.; Skoog, T.; Ala-Aho, R.; Jeskanen, L.; Pärssinen, J.; Latonen, L.; Grénman, R.; Kere, J.; Kähäri, V.-M.; et al. CCHCR1 Is Up-Regulated in Skin Cancer and Associated with EGFR Expression. PLoS ONE 2009, 4, e6030. [Google Scholar] [CrossRef] [PubMed]
- Tamborero, D.; Rubio-Perez, C.; Deu-Pons, J.; Schroeder, M.P.; Vivancos, A.; Rovira, A.; Tusquets, I.; Albanell, J.; Rodon, J.; Tabernero, J.; et al. Cancer Genome Interpreter annotates the biological and clinical relevance of tumor alterations. Genome Med. 2018, 10, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [PubMed]
- D’Amato, V.; Rosa, R.; D’Amato, C.; Formisano, L.; Marciano, R.; Nappi, L.; Raimondo, L.; Di Mauro, C.; Servetto, A.; Fusciello, C.; et al. The dual PI3K/mTOR inhibitor PKI-587 enhances sensitivity to cetuximab in EGFR-resistant human head and neck cancer models. Br. J. Cancer 2014, 110, 2887–2895. [Google Scholar] [CrossRef] [PubMed]
- Liu, X. The Epithelial-Mesenchymal Transition and Cancer Stem Cells: Functional and Mechanistic Links. Curr. Pharm. Des. 2015, 21, 1279–1291. [Google Scholar] [CrossRef]
- Byeon, H.K.; Ku, M.; Yang, J. Beyond EGFR inhibition: Multilateral combat strategies to stop the progression of head and neck cancer. Exp. Mol. Med. 2019, 51, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Cheng, K.-Y.; Hao, M. Mammalian Target of Rapamycin (mTOR) Regulates Transforming Growth Factor-β1 (TGF-β1)-Induced Epithelial-Mesenchymal Transition via Decreased Pyruvate Kinase M2 (PKM2) Expression in Cervical Cancer Cells. Med. Sci. Monit. 2017, 23, 2017–2028. [Google Scholar] [CrossRef]
- Krzyszczyk, P.; Acevedo, A.; Davidoff, E.J.; Timmins, L.M.; Marrero-Berrios, I.; Patel, M.; White, C.; Lowe, C.; Sherba, J.J.; Hartmanshenn, C.; et al. The growing role of precision and personalized medicine for cancer treatment. Technology 2018, 6, 79–100. [Google Scholar] [CrossRef] [PubMed]
- Benavente, S.; Huang, S.; Armstrong, E.A.; Chi, A.; Hsu, K.-T.; Wheeler, D.L.; Harari, P.M. Establishment and Characterization of a Model of Acquired Resistance to Epidermal Growth Factor Receptor Targeting Agents in Human Cancer Cells. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2009, 15, 1585–1592. [Google Scholar] [CrossRef] [PubMed]
- Boeckx, C.; Blockx, L.; de Beeck, K.O.; Limame, R.; Van Camp, G.; Peeters, M.; Vermorken, J.B.; Specenier, P.; Wouters, A.; Baay, M.; et al. Establishment and characterization of cetuximab resistant head and neck squamous cell carcinoma cell lines: Focus on the contribution of the AP-1 transcription factor. Am. J. Cancer Res. 2015, 5, 1921–1938. [Google Scholar]
- De Pauw, I.; Lardon, F.; Bossche, J.V.D.; Baysal, H.; Pauwels, P.; Peeters, M.; Vermorken, J.B.; Wouters, A. Overcoming Intrinsic and Acquired Cetuximab Resistance in RAS Wild-Type Colorectal Cancer: An In Vitro Study on the Expression of HER Receptors and the Potential of Afatinib. Cancers 2019, 11, 98. [Google Scholar] [CrossRef] [PubMed]
- Willey, C.D.; Anderson, J.C.; Trummell, H.Q.; Naji, F.; de Wijn, R.; Yang, E.S.; Bredel, M.; Thudi, N.K.; Bonner, J.A. Differential escape mechanisms in cetuximab-resistant head and neck cancer cells. Biochem. Biophys. Res. Commun. 2019, 517, 36–42. [Google Scholar] [CrossRef]
- Morgillo, F.; Della Corte, C.M.; Fasano, M.; Ciardiello, F. Mechanisms of resistance to EGFR-targeted drugs: Lung cancer. ESMO Open 2016, 1, e000060. [Google Scholar] [CrossRef]
- Lièvre, A.; Bachet, J.-B.; Le Corre, D.; Boige, V.; Landi, B.; Emile, J.-F.; Côté, J.-F.; Tomasic, G.; Penna, C.; Ducreux, M.; et al. KRAS Mutation Status Is Predictive of Response to Cetuximab Therapy in Colorectal Cancer. Cancer Res. 2006, 66, 3992–3995. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Peyrollier, K.; Kilic, G.; Brakebusch, C. Rho GTPases and cancer. BioFactors 2014, 40, 226–235. [Google Scholar] [CrossRef]
- Pan, Q.; Bao, L.W.; Teknos, T.N.; Merajver, S.D. Targeted Disruption of Protein Kinase Cε Reduces Cell Invasion and Motility through Inactivation of RhoA and RhoC GTPases in Head and Neck Squamous Cell Carcinoma. Cancer Res. 2006, 66, 9379–9384. [Google Scholar] [CrossRef] [PubMed]
- Ruihua, H.; Mengyi, Z.; Chong, Z.; Meng, Q.; Xin, M.; Qiulin, T.; Feng, B.; Ming, L. RhoA regulates resistance to irinotecan by regulating membrane transporter and apoptosis signaling in colorectal cancer. Oncotarget 2016, 7, 87136–87146. [Google Scholar] [CrossRef]
- Doublier, S.; Riganti, C.; Voena, C.; Costamagna, C.; Aldieri, E.; Pescarmona, G.P.; Ghigo, D.; Bosia, A. RhoA Silencing Reverts the Resistance to Doxorubicin in Human Colon Cancer Cells. Mol. Cancer Res. MCR 2008, 6, 1607–1620. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Gulhati, P.; Bowen, K.A.; Liu, J.; Stevens, P.D.; Rychahou, P.; Chen, M.; Lee, E.Y.; Weiss, H.L.; O’Connor, K.; Gao, T.; et al. mTORC1 and mTORC2 Regulate EMT, Motility, and Metastasis of Colorectal Cancer via RhoA and Rac1 Signaling Pathways. Cancer Res. 2011, 71, 3246–3256. [Google Scholar] [CrossRef]
- Bhowmick, N.A.; Ghiassi, M.; Bakin, A.; Aakre, M.; Lundquist, C.A.; Engel, M.E.; Arteaga, C.L.; Moses, H.L. Transforming Growth Factor-β1 Mediates Epithelial to Mesenchymal Transdifferentiation through a RhoA-dependent Mechanism. Mol. Biol. Cell 2001, 12, 27–36. [Google Scholar] [CrossRef]
- Kalluri, R.; Weinberg, R.A. The basics of epithelial-mesenchymal transition. J. Clin. Investig. 2009, 119, 1420–1428. [Google Scholar] [CrossRef] [PubMed]
- Lamouille, S.; Connolly, E.; Smyth, J.; Akhurst, R.; Derynck, R. TGF-β-induced activation of mTOR complex 2 drives epithelial–mesenchymal transition and cell invasion. J. Cell Sci. 2012, 125, 1259–1273. [Google Scholar] [CrossRef] [PubMed]
- Prieto-Vila, M.; Takahashi, R.-U.; Usuba, W.; Kohama, I.; Ochiya, T. Drug Resistance Driven by Cancer Stem Cells and Their Niche. Int. J. Mol. Sci. 2017, 18, 2574. [Google Scholar] [CrossRef]
- Thapa, R.; Wilson, G. The Importance of CD44 as a Stem Cell Biomarker and Therapeutic Target in Cancer. Stem Cells Int. 2016, 2016, 2087204. [Google Scholar] [CrossRef] [PubMed]
- Suda, K.; Murakami, I.; Yu, H.; Kim, J.; Tan, A.-C.; Mizuuchi, H.; Rozeboom, L.M.; Ellison, K.; Rivard, C.J.; Mitsudomi, T.; et al. CD44 Facilitates Epithelial-to-Mesenchymal Transition Phenotypic Change at Acquisition of Resistance to EGFR Kinase Inhibitors in Lung Cancer. Mol. Cancer Ther. 2018, 17, 2257–2265. [Google Scholar] [CrossRef]
- Allegra, E.; Trapasso, S. Role of CD44 as a marker of cancer stem cells in head and neck cancer. Biol. Targets Ther. 2012, 6, 379–383. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Wu, K.; Xu, H.; Tian, Y.; Yuan, X.; Wu, H.; Liu, Q.; Pestell, R. The role of CD44 in epithelial–mesenchymal transition and cancer development. OncoTargets Ther. 2015, 8, 3783–3792. [Google Scholar] [CrossRef] [PubMed]
- Straub, M.; Drecoll, E.; Pfarr, N.; Weichert, W.; Langer, R.; Hapfelmeier, A.; Götz, C.; Wolff, K.-D.; Kolk, A.; Specht, K. CD274/PD-L1 gene amplification and PD-L1 protein expression are common events in squamous cell carcinoma of the oral cavity. Oncotarget 2016, 7, 12024–12034. [Google Scholar] [CrossRef]
- Jiang, L.; Guo, F.; Liu, X.; Li, X.; Qin, Q.; Shu, P.; Li, Y.; Wang, Y. Continuous targeted kinase inhibitors treatment induces upregulation of PD-L1 in resistant NSCLC. Sci. Rep. 2019, 9, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Yonesaka, K.; Zejnullahu, K.; Okamoto, I.; Satoh, T.; Cappuzzo, F.; Souglakos, J.; Ercan, D.; Rogers, A.; Roncalli, M.; Takeda, M.; et al. Activation of ERBB2 Signaling Causes Resistance to the EGFR-Directed Therapeutic Antibody Cetuximab. Sci. Transl. Med. 2011, 3, 99ra86. [Google Scholar] [CrossRef] [PubMed]
- Iida, M.; Brand, T.M.; Starr, M.M.; Huppert, E.J.; Luthar, N.; Bahrar, H.; Coan, J.P.; Pearson, H.E.; Salgia, R.; Wheeler, D.L. Overcoming acquired resistance to cetuximab by dual targeting HER family receptors with antibody-based therapy. Mol. Cancer 2014, 13, 242. [Google Scholar] [CrossRef] [PubMed]
- Er, E.E.; Mendoza, M.C.; Mackey, A.M.; Rameh, L.E.; Blenis, J. AKT Facilitates EGFR Trafficking and Degradation by Phosphorylating and Activating PIKfyve. Sci. Signal. 2013, 6, ra45. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Iida, M.; Dunn, E.F.; Ghia, A.J.; Wheeler, D.L. Nuclear EGFR contributes to acquired resistance to cetuximab. Oncogene 2009, 28, 3801–3813. [Google Scholar] [CrossRef] [PubMed]
- Cipponi, A.; Goode, D.L.; Bedo, J.; McCabe, M.J.; Pajic, M.; Croucher, D.R.; Rajal, A.G.; Junankar, S.R.; Saunders, D.N.; Lobachevsky, P.; et al. MTOR signaling orchestrates stress-induced mutagenesis, facilitating adaptive evolution in cancer. Science 2020, 368, 1127–1131. [Google Scholar] [CrossRef] [PubMed]
- Bray, S.M.; Lee, J.; Kim, S.T.; Hur, J.Y.; Ebert, P.J.; Calley, J.N.; Wulur, I.H.; Gopalappa, T.; Wong, S.S.; Qian, H.-R.; et al. Genomic characterization of intrinsic and acquired resistance to cetuximab in colorectal cancer patients. Sci. Rep. 2019, 9, 1–13. [Google Scholar] [CrossRef]
- Zhu, S.; Ward, B.M.; Yu, J.; Matthew-Onabanjo, A.N.; Janusis, J.; Hsieh, C.-C.; Tomaszewicz, K.; Hutchinson, L.; Zhu, L.J.; Kandil, D.; et al. IRS2 mutations linked to invasion in pleomorphic invasive lobular carcinoma. JCI Insight 2018, 3. [Google Scholar] [CrossRef]
- Shih, C.-H.; Chang, Y.-J.; Huang, W.-C.; Jang, T.-H.; Kung, H.-J.; Wang, W.-C.; Yang, M.-H.; Lin, M.-C.; Huang, S.-F.; Chou, S.-W.; et al. EZH2-mediated upregulation of ROS1 oncogene promotes oral cancer metastasis. Oncogene 2017, 36, 6542–6554. [Google Scholar] [CrossRef]
- Velthaus, J.-L.; Iglauer, P.; Simon, R.; Bokemeyer, C.; Bannas, P.; Beumer, N.; Imbusch, C.D.; Goekkurt, E.; Loges, S. Lorlatinib Induces Durable Disease Stabilization in a Pancreatic Cancer Patient with a ROS1 p.L1950F Mutation: Case Report. Oncol. Res. Treat. 2021, 44, 495–502. [Google Scholar] [CrossRef]
- Davies, K.D.; Mahale, S.; Astling, D.P.; Aisner, D.L.; Le, A.T.; Hinz, T.K.; Vaishnavi, A.; Bunn, P.A.; Heasley, L.E.; Tan, A.C.; et al. Resistance to ROS1 Inhibition Mediated by EGFR Pathway Activation in Non-Small Cell Lung Cancer. PLoS ONE 2013, 8, e82236. [Google Scholar] [CrossRef]
| Chromosome | Event | Cytoband | Cancer Gene | Driver Statement by CGI |
|---|---|---|---|---|
| chr 17 | Deletion | p13.3–p11.2 | MAP2K4 | known in: PA; BRCA; COREAD |
| chr 13 | Deletion | q31.1–q32.2 | GPC5 | predicted passenger |
| chr 20 | Deletion | q11.22–q13.33 | EEF1A2 | predicted passenger |
| chr 17 | Deletion | p13.3–p11.2 | MAPK7 | predicted passenger |
| chr 17 | Deletion | p13.3–p11.2 | TP53 | known in: BCL; THYM |
| chr 11 | Deletion | p15.5–p12 | WT1 | known in: WT; DSRCT |
| chr 10 | Deletion | q22.3–q26.3 | PTEN | known in: G; PRAD; ED; CM; TH; BRCA; L; OV; PA |
| chr 7 | Deletion | p14.3–p12.1 | IKZF1 | known in: ALL; DLBCL |
| chr 10 | Deletion | q22.3–q26.3 | SUFU | known in: MB |
| chr 16 | Deletion | q24.3 | FANCA | known in: AML; LK; PRAD |
| chr 17 | Deletion | p13.3–p11.2 | FLCN | known in: TH |
| chr 17 | Deletion | q11.1–q23.1 | NF1 | known in: NF; G; MPN; CM; PLEN; HNC; SG; LK |
| chr 17 | Deletion | q11.1–q23.1 | SUZ12 | known in: CANCER |
| chr 17 | Deletion | q11.1–q23.1 | BRCA1 | known in: OV; BRCA |
| chr 12 | High Amplification | p12.1–q11 | KRAS | predicted driver |
| chr 14 | Amplification | q12–q32.33 | NKX2-1 | known in: NSCLC |
| chr 14 | High Amplification | q11.2–q12 | BCL2L2 | predicted passenger |
| chr 5 | High Amplification | p13.3–q11.2 | SKP2 | predicted passenger |
| chr 22 | Amplification | q11.1–q12.1 | CRKL | predicted passenger |
| chr 11 | High Amplification | q22.1–q22.3 | YAP1, BIRC2 | predicted passenger |
| chr 7 | High Amplification | q36.1–q36.3 | SHH | predicted passenger |
| chr 6 | Amplification | q16.2–q27 | MYB | predicted driver |
| chr 6 | Amplification | q16.2–q27 | MAP3K5 | predicted passenger |
| chr 1 | Amplification | q32.1–q32.2 | MDM4 | known in: GBM; BLCA; RB; S |
| chr 12 | Amplification | q14.3–q24.33 | MDM2, HMGA2 | known in: S; G; COREAD; LIP |
| chr 9 | Amplification | p24.2–p22.1 | JAK2 | known in: BRCA |
| chr 9 | Amplification | p24.2–p22.1 | CD274 | known in: BCC |
| chr 8 | Amplification | q12.1–q24.3 | MYC | known in: BLY; CLL; NB; COREAD; MYMA; PRAD |
| chr 5 | High Amplification | p13.3–q11.2 | RICTOR | known in: L |
| chr 6 | Amplification | q16.2–q27 | ESR1 | known in: UCEC; BRCA; OV |
| chr 12 | Amplification | p13.33–p12.1 | CCND2 | known in: L |
| chr 12 | Amplification | q14.3–q24.33 | FRS2 | known in: LIP |
| chr 14 | Amplification | q12–q32.33 | FOXA1 | Known in: COREAD |
| chr 14 | Amplification | q12–q32.33 | PAX9 | Known in: NSCLC |
| chr 14 | Amplification | q12–q32.33 | NKX2-8 | predicted passenger |
| Chromosome | cDNA | Protein | Consequence | Gene | Driver Status |
|---|---|---|---|---|---|
| chr 1 | c.2492A>T | p.N831I | Missense | EPHA2 | Tumor Driver |
| chr 2 | c.2074G>T | p.G692W | Missense | ALK | Tumor Driver |
| chr 6 | c.6341A>G | p.Y2114C | Missense | ROS1 | Tumor Driver |
| chr 6 | c.3391A>T | p.K1131* | Nonsense | ZNF292 | Tumor Driver |
| chr 8 | c.1648delT | p.S550Qfs*12 | Frameshift | UBR5 | Tumor Driver |
| chr 9 | c.3127_3129delAGC | p.S1043delS | In Frame Deletion | TSC1 | Tumor Driver |
| chr 9 | c.740delC | p.P247Qfs*30 | Frameshift | NOTCH1 | Tumor Driver |
| chr 9 | c.250_252delGAA | p.E84delE | In Frame Deletion | XPA | Tumor Driver |
| chr 13 | c.3273dupG | p.K1092Efs*233 | Frameshift | IRS2 | Tumor Driver |
| chr 14 | c.928delG | p.E310Kfs*68 | Frameshift | ARID4A | Tumor Driver |
| chr 15 | c.3416delG | p.G1139Efs*25 | Frameshift | FANCI | Tumor Driver |
| chr 16 | c.1183delC | p.H395Tfs*78 | Frameshift | TRAF7 | Tumor Driver |
| chr 17 | c.1420_1422delCAT | p.H474delH | In Frame Deletion | AXIN2 | Tumor Driver |
| chr 19 | c.209A>T | p.N70I | Missense | ARHGAP35 | Tumor Driver |
| chr 21 | c.146delC | p.P49Qfs*4 | Frameshift | RUNX1 | Tumor Driver |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gomes, I.N.F.; da Silva-Oliveira, R.J.; da Silva, L.S.; Martinho, O.; Evangelista, A.F.; van Helvoort Lengert, A.; Leal, L.F.; Silva, V.A.O.; dos Santos, S.P.; Nascimento, F.C.; et al. Comprehensive Molecular Landscape of Cetuximab Resistance in Head and Neck Cancer Cell Lines. Cells 2022, 11, 154. https://doi.org/10.3390/cells11010154
Gomes INF, da Silva-Oliveira RJ, da Silva LS, Martinho O, Evangelista AF, van Helvoort Lengert A, Leal LF, Silva VAO, dos Santos SP, Nascimento FC, et al. Comprehensive Molecular Landscape of Cetuximab Resistance in Head and Neck Cancer Cell Lines. Cells. 2022; 11(1):154. https://doi.org/10.3390/cells11010154
Chicago/Turabian StyleGomes, Izabela N. F., Renato J. da Silva-Oliveira, Luciane Sussuchi da Silva, Olga Martinho, Adriane F. Evangelista, André van Helvoort Lengert, Letícia Ferro Leal, Viviane Aline Oliveira Silva, Stéphanie Piancenti dos Santos, Flávia Caroline Nascimento, and et al. 2022. "Comprehensive Molecular Landscape of Cetuximab Resistance in Head and Neck Cancer Cell Lines" Cells 11, no. 1: 154. https://doi.org/10.3390/cells11010154
APA StyleGomes, I. N. F., da Silva-Oliveira, R. J., da Silva, L. S., Martinho, O., Evangelista, A. F., van Helvoort Lengert, A., Leal, L. F., Silva, V. A. O., dos Santos, S. P., Nascimento, F. C., Lopes Carvalho, A., & Reis, R. M. (2022). Comprehensive Molecular Landscape of Cetuximab Resistance in Head and Neck Cancer Cell Lines. Cells, 11(1), 154. https://doi.org/10.3390/cells11010154







