Building Pluripotency Identity in the Early Embryo and Derived Stem Cells
Abstract
1. Introduction
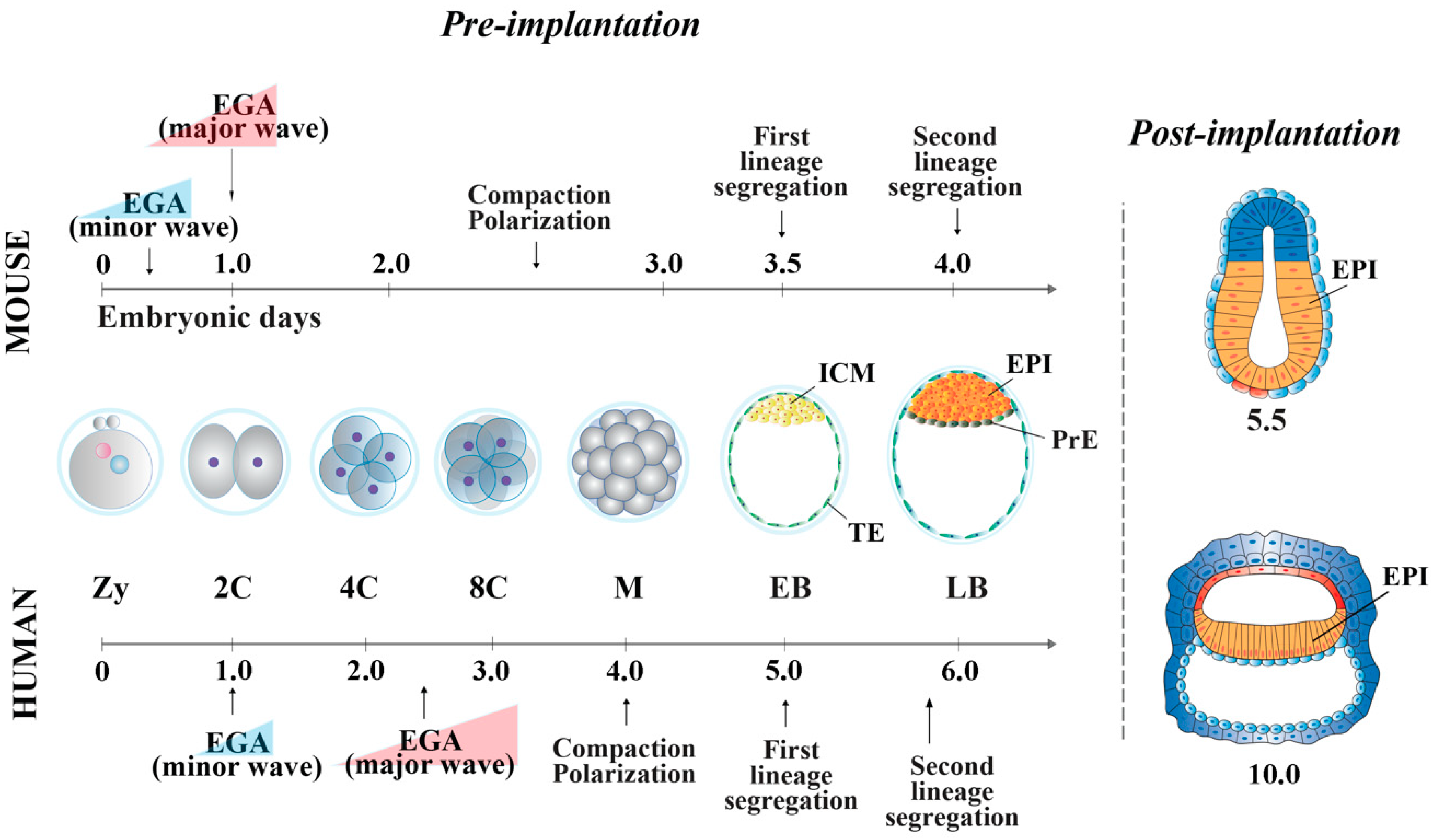
2. Cellular and Molecular Features of Mouse and Human Peri-Implantation Embryos
2.1. Mouse
2.1.1. Embryonic Genome Activation
2.1.2. Embryo Compaction and First Cell Lineage Specification
2.1.3. Second Cell Lineage Specification
2.2. Human
2.2.1. Embryonic Genome Activation
2.2.2. Embryo Compaction and First Cell Lineage Specification
2.2.3. Second Cell Lineage Specification
3. Features Governing Cell Potency
3.1. The Epigenetic Landscape
3.1.1. DNA Methylation and Histone Modifications
Mouse
Human
3.1.2. X Chromosome Inactivation and Reactivation
Mouse
Human
3.2. Non-Coding RNAs
3.2.1. miRNAs
3.2.2. Long Non-Coding RNAs
4. Cellular and Molecular Features of Stem Cell Pluripotency In Vitro
4.1. Cellular Features
4.1.1. Mouse
Stem Cells with Intermediate Pluripotency
Stem Cells with Primed Pluripotency
4.1.2. Human
4.1.3. Mouse and Human Stem Cells with Extended Pluripotency
4.2. Molecular Features
4.2.1. Pluripotency Transcriptional Networks (PTNs)
Mouse PTNs
- Naïve
- Formative
- Primed
Human PTNs
- Naïve and Primed
4.2.2. DNA Methylation and Histone Modification
- Mouse
- Human
4.2.3. X Chromosome Inactivation and Reactivation
4.2.4. Non-Coding RNAs
miRNAs
Long Non-Coding RNAs
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Denker, H.W. Stem cell terminology and ‘synthetic’ embryos: A new debate on totipotency, omnipotency, and pluripotency and how it relates to recent experimental data. Cells Tissues Organs 2014, 199, 221–227. [Google Scholar] [CrossRef]
- Singh, V.K.; Saini, A.; Kalsan, M.; Kumar, N.; Chandra, R. Describing the Stem Cell Potency: The Various Methods of Functional Assessment and In silico Diagnostics. Front. Cell Dev. Biol. 2016, 4, 134. [Google Scholar] [CrossRef]
- Togashi, M.; Suzuki, H.; Miyai, T.; Okamoto, M.T. Production of monozygotic twins by splitting of 2-cell stage embryos in mice. Jpn. J. Anim. Reprod. 1987, 33, 51–57. [Google Scholar] [CrossRef][Green Version]
- Tsunoda, Y.; McLaren, A. Effect of various procedures on the viability of mouse embryos containing half the normal number of blastomeres. J. Reprod. Fertil. 1983, 69, 315–322. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Kato, Y.; Tsunoda, Y. Effects of several factors on the monozygotic twin production in the mouse. J. Reprod. Dev. 1997, 43, 91–95. [Google Scholar] [CrossRef][Green Version]
- Sotomaru, Y.; Kato, Y.; Tsunoda, Y. Production of monozygotic twins after freezing and thawing of bisected mouse embryos. Cryobiology 1998, 37, 139–145. [Google Scholar] [CrossRef]
- Casser, E.; Israel, S.; Witten, A.; Schulte, K.; Schlatt, S.; Nordhoff, V.; Boiani, M. Totipotency segregates between the sister blastomeres of two-cell stage mouse embryos. Sci. Rep. 2017, 7, 8299. [Google Scholar] [CrossRef] [PubMed]
- Casser, E.; Israel, S.; Boiani, M. Multiplying embryos: Experimental monozygotic polyembryony in mammals and its uses. Int. J. Dev. Biol. 2019, 63, 143–155. [Google Scholar] [CrossRef]
- Boiani, M.; Casser, E.; Fuellen, G.; Christians, E.S. Totipotency continuity from zygote to early blastomeres: A model under revision. Reproduction 2019, 158, R49–R65. [Google Scholar] [CrossRef]
- Saiz, N.; Williams, K.M.; Seshan, V.E.; Hadjantonakis, A.K. Asynchronous fate decisions by single cells collectively ensure consistent lineage composition in the mouse blastocyst. Nat. Commun. 2016, 7, 13463. [Google Scholar] [CrossRef]
- Gardner, R.L.; Rossant, J. Investigation of the fate of 4–5 day post-coitum mouse inner cell mass cells by blastocyst injection. J. Embryol. Exp. Morphol. 1979, 52, 141–152. [Google Scholar]
- De Paepe, C.; Krivega, M.; Cauffman, G.; Geens, M.; Van de Velde, H. Totipotency and lineage segregation in the human embryo. Mol. Hum. Reprod. 2014, 20, 599–618. [Google Scholar] [CrossRef]
- Van de Velde, H.; Cauffman, G.; Tournaye, H.; Devroey, P.; Liebaers, I. The four blastomeres of a 4-cell stage human embryo are able to develop individually into blastocysts with inner cell mass and trophectoderm. Hum. Reprod. 2008, 23, 1742–1747. [Google Scholar] [CrossRef]
- Gerri, C.; Menchero, S.; Mahadevaiah, S.K.; Turner, J.M.A.; Niakan, K.K. Human Embryogenesis: A Comparative Perspective. Annu. Rev. Cell Dev. Biol. 2020, 36, 411–440. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.; Yang, M.; Guo, H.; Yang, L.; Wu, J.; Li, R.; Liu, P.; Lian, Y.; Zheng, X.; Yan, J.; et al. Single-cell RNA-Seq profiling of human preimplantation embryos and embryonic stem cells. Nat. Struct. Mol. Biol. 2013, 20, 1131–1139. [Google Scholar] [CrossRef] [PubMed]
- Shahbazi, M.N.; Scialdone, A.; Skorupska, N.; Weberling, A.; Recher, G.; Zhu, M.; Jedrusik, A.; Devito, L.G.; Noli, L.; Macaulay, I.C.; et al. Pluripotent state transitions coordinate morphogenesis in mouse and human embryos. Nature 2017, 552, 239–243. [Google Scholar] [CrossRef]
- Martin, G.R. Isolation of a pluripotent cell line from early mouse embryos cultured in medium conditioned by teratocarcinoma stem cells. Proc. Natl. Acad. Sci. USA 1981, 78, 7634–7638. [Google Scholar] [CrossRef] [PubMed]
- Evans, M.J.; Kaufman, M.H. Establishment in culture of pluripotential cells from mouse embryos. Nature 1981, 292, 154–156. [Google Scholar] [CrossRef]
- Thomson, J.A.; Itskovitz-Eldor, J.; Shapiro, S.S.; Waknitz, M.A.; Swiergiel, J.J.; Marshall, V.S.; Jones, J.M. Embryonic stem cell lines derived from human blastocysts. Science 1998, 282, 1145–1147. [Google Scholar] [CrossRef] [PubMed]
- Tesar, P.J.; Chenoweth, J.G.; Brook, F.A.; Davies, T.J.; Evans, E.P.; Mack, D.L.; Gardner, R.L.; McKay, R.D. New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature 2007, 448, 196–199. [Google Scholar] [CrossRef] [PubMed]
- Brons, I.G.; Smithers, L.E.; Trotter, M.W.; Rugg-Gunn, P.; Sun, B.; Chuva de Sousa Lopes, S.M.; Howlett, S.K.; Clarkson, A.; Ahrlund-Richter, L.; Pedersen, R.A.; et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature 2007, 448, 191–195. [Google Scholar] [CrossRef]
- Osorno, R.; Tsakiridis, A.; Wong, F.; Cambray, N.; Economou, C.; Wilkie, R.; Blin, G.; Scotting, P.J.; Chambers, I.; Wilson, V. The developmental dismantling of pluripotency is reversed by ectopic Oct4 expression. Development 2012, 139, 2288–2298. [Google Scholar] [CrossRef]
- Riveiro, A.R.; Brickman, J.M. From pluripotency to totipotency: An experimentalist’s guide to cellular potency. Development 2020, 147, dev189845. [Google Scholar] [CrossRef]
- Shahbazi, M.N.; Siggia, E.D.; Zernicka-Goetz, M. Self-organization of stem cells into embryos: A window on early mammalian development. Science 2019, 364, 948–951. [Google Scholar] [CrossRef]
- Wu, D.; Dean, J. Maternal factors regulating preimplantation development in mice. Curr. Top. Dev. Biol. 2020, 140, 317–340. [Google Scholar] [PubMed]
- Schulz, K.N.; Harrison, M.M. Mechanisms regulating zygotic genome activation. Nat. Rev. Genet. 2019, 20, 221–234. [Google Scholar] [CrossRef] [PubMed]
- Foygel, K.; Choi, B.; Jun, S.; Leong, D.E.; Lee, A.; Wong, C.C.; Zuo, E.; Eckart, M.; Reijo Pera, R.A.; Wong, W.H.; et al. A novel and critical role for Oct4 as a regulator of the maternal-embryonic transition. PLoS ONE 2008, 3, e4109. [Google Scholar] [CrossRef]
- Wu, G.; Schöler, H.R. Role of Oct4 in the early embryo development. Cell Regen. 2014, 3, 7. [Google Scholar] [CrossRef]
- Peaston, A.E.; Evsikov, A.V.; Graber, J.H.; de Vries, W.N.; Holbrook, A.E.; Solter, D. Knowles, B.B. Retrotransposons regulate host genes in mouse oocytes and preimplantation embryos. Dev. Cell 2004, 7, 597–606. [Google Scholar] [CrossRef]
- Inoue, A.; Matoba, S.; Zhang, Y. Transcriptional activation of transposable elements in mouse zygotes is independent of Tet3-mediated 5-methylcytosine oxidation. Cell Res. 2012, 22, 1640–1649. [Google Scholar] [CrossRef] [PubMed]
- Pikó, L.; Hammons, M.D.; Taylor, K.D. Amounts, synthesis, and some properties of intracisternal A particle-related RNA in early mouse embryos. Proc. Natl. Acad. Sci. USA 1984, 81, 488–492. [Google Scholar] [CrossRef]
- Poznanski, A.A.; Calarco, P.G. The expression of intracisternal A particle genes in the preimplantation mouse embryo. Dev. Biol. 1991, 143, 271–281. [Google Scholar] [CrossRef]
- Macfarlan, T.S.; Gifford, W.D.; Driscoll, S.; Lettieri, K.; Rowe, H.M.; Bonanomi, D.; Firth, A.; Singer, O.; Trono, D.; Pfaff, S.L. Embryonic stem cell potency fluctuates with endogenous retrovirus activity. Nature 2012, 487, 57–63. [Google Scholar] [CrossRef]
- Yartseva, V.; Giraldez, A.J. The Maternal-to-Zygotic Transition during Vertebrate Development: A Model for Reprogramming. Curr. Top. Dev. Biol. 2015, 113, 191–232. [Google Scholar] [CrossRef] [PubMed]
- Bushati, N.; Stark, A.; Brennecke, J.; Cohen, S.M. Temporal reciprocity of miRNAs and their targets during the maternal-to-zygotic transition in Drosophila. Curr. Biol. 2008, 18, 501–506. [Google Scholar] [CrossRef]
- Lee, M.T.; Bonneau, A.R.; Takacs, C.M.; Bazzini, A.A.; DiVito, K.R.; Fleming, E.S.; Giraldez, A.J. Nanog, Pou5f1 and SoxB1 activate zygotic gene expression during the maternal-to-zygotic transition. Nature 2013, 503, 360–364. [Google Scholar] [CrossRef]
- Hahn, S. Structure and mechanism of the RNA polymerase II transcription machinery. Nat. Struct. Mol. Biol. 2004, 11, 394–403. [Google Scholar] [CrossRef]
- Vinot, S.; Le, T.; Ohno, S.; Pawson, T.; Maro, B.; Louvet-Vallée, S. Asymmetric distribution of PAR proteins in the mouse embryo begins at the 8-cell stage during compaction. Dev. Biol. 2005, 282, 307–319. [Google Scholar] [CrossRef]
- Dard, N.; Louvet-Vallée, S.; Maro, B. Orientation of mitotic spindles during the 8- to 16-cell stage transition in mouse embryos. PLoS ONE 2009, 4, e8171. [Google Scholar] [CrossRef] [PubMed]
- Cockburn, K.; Rossant, J. Making the blastocyst: Lessons from the mouse. J. Clin. Investig. 2010, 120, 995–1003. [Google Scholar] [CrossRef] [PubMed]
- White, M.D.; Plachta, N. How adhesion forms the early mammalian embryo. Curr. Top. Dev. Biol. 2015, 112, 1–17. [Google Scholar] [CrossRef]
- Johnson, M.H.; McConnell, J.M. Lineage allocation and cell polarity during mouse embryogenesis. Semin. Cell Dev. Biol. 2004, 15, 583–597. [Google Scholar] [CrossRef] [PubMed]
- Ben-Tabou de-Leon, S.; Davidson, E.H. Gene regulation: Gene control network in development. Annu. Rev. Biophys. Biomol. Struct. 2007, 36, 191. [Google Scholar] [CrossRef]
- Mallo, M.; Alonso, C.R. The regulation of Hox gene expression during animal development. Development 2013, 140, 3951–3963. [Google Scholar] [CrossRef]
- Yagi, R.; Kohn, M.J.; Karavanova, I.; Kaneko, K.J.; Vullhorst, D.; DePamphilis, M.L.; Buonanno, A. Transcription factor TEAD4 specifies the trophectoderm lineage at the beginning of mammalian development. Development 2007, 134, 3827–3836. [Google Scholar] [CrossRef] [PubMed]
- Nishioka, N.; Yamamoto, S.; Kiyonari, H.; Sato, H.; Sawada, A.; Ota, M.; Nakao, K.; Sasaki, H. Tead4 is required for specification of trophectoderm in pre-implantation mouse embryos. Mech. Dev. 2008, 125, 270–283. [Google Scholar] [CrossRef]
- Nishioka, N.; Inoue, K.; Adachi, K.; Kiyonari, H.; Ota, M.; Ralston, A.; Yabuta, N.; Hirahara, S.; Stephenson, R.O.; Ogonuki, N.; et al. The Hippo signaling pathway components Lats and Yap pattern Tead4 activity to distinguish mouse trophectoderm from inner cell mass. Dev. Cell 2009, 16, 398–410. [Google Scholar] [CrossRef] [PubMed]
- Anani, S.; Bhat, S.; Honma-Yamanaka, N.; Krawchuk, D.; Yamanaka, Y. Initiation of Hippo signaling is linked to polarity rather than to cell position in the pre-implantation mouse embryo. Development 2014, 141, 2813–2824. [Google Scholar] [CrossRef]
- Hirate, Y.; Hirahara, S.; Inoue, K.; Kiyonari, H.; Niwa, H.; Sasaki, H. Par-aPKC-dependent and -independent mechanisms cooperatively control cell polarity, Hippo signaling, and cell positioning in 16-cell stage mouse embryos. Dev. Growth Differ. 2015, 57, 544–556. [Google Scholar] [CrossRef] [PubMed]
- Ng, R.K.; Dean, W.; Dawson, C.; Lucifero, D.; Madeja, Z.; Reik, W.; Hemberger, M. Epigenetic restriction of embryonic cell lineage fate by methylation of Elf5. Nat. Cell Biol. 2008, 10, 1280–1290. [Google Scholar] [CrossRef]
- Blakeley, P.; Fogarty, N.M.; del Valle, I.; Wamaitha, S.E.; Hu, T.X.; Elder, K.; Snell, P.; Christie, L.; Robson, P.; Niakan, K.K. Defining the three cell lineages of the human blastocyst by single-cell RNA-seq. Development 2015, 142, 3151–3165. [Google Scholar] [CrossRef]
- Strumpf, D.; Mao, C.A.; Yamanaka, Y.; Ralston, A.; Chawengsaksophak, K.; Beck, F.; Rossant, J. Cdx2 is required for correct cell fate specification and differentiation of trophectoderm in the mouse blastocyst. Development 2005, 132, 2093–2102. [Google Scholar] [CrossRef]
- Niakan, K.K.; Eggan, K. Analysis of human embryos from zygote to blastocyst reveals distinct gene expression patterns relative to the mouse. Dev. Biol. 2013, 375, 54–64. [Google Scholar] [CrossRef]
- Ralston, A.; Cox, B.J.; Nishioka, N.; Sasaki, H.; Chea, E.; Rugg-Gunn, P.; Guo, G.; Robson, P.; Draper, J.S.; Rossant, J. Gata3 regulates trophoblast development downstream of Tead4 and in parallel to Cdx2. Development 2010, 137, 395–403. [Google Scholar] [CrossRef]
- Noli, L.; Dajani, Y.; Capalbo, A.; Bvumbe, J.; Rienzi, L.; Ubaldi, F.M.; Ogilvie, C.; Khalaf, Y.; Ilic, D. Developmental clock compromises human twin model created by embryo splitting. Hum. Reprod. 2015, 30, 2774–2784. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Deglincerti, A.; Croft, G.F.; Pietila, L.N.; Zernicka-Goetz, M.; Siggia, E.D.; Brivanlou, A.H. Self-organization of the in vitro attached human embryo. Nature 2016, 533, 251–254. [Google Scholar] [CrossRef] [PubMed]
- Menchero, S.; Rollan, I.; Lopez-Izquierdo, A.; Andreu, M.J.; Sainz de Aja, J.; Kang, M.; Adan, J.; Benedito, R.; Rayon, T.; Hadjantonakis, A.K.; et al. Transitions in cell potency during early mouse development are driven by Notch. Elife 2019, 8, e42930. [Google Scholar] [CrossRef] [PubMed]
- Frum, T.; Murphy, T.M.; Ralston, A. HIPPO signaling resolves embryonic cell fate conflicts during establishment of pluripotency in vivo. Elife 2018, 7, e42298. [Google Scholar] [CrossRef]
- Wicklow, E.; Blij, S.; Frum, T.; Hirate, Y.; Lang, R.A.; Sasaki, H.; Ralston, A. HIPPO pathway members restrict SOX2 to the inner cell mass where it promotes ICM fates in the mouse blastocyst. PLoS Genet. 2014, 10, e1004618. [Google Scholar] [CrossRef] [PubMed]
- Gharanfoli, S.; Shahverdi, A.; Dalman, A.; Ghaznavi, P.; Alipour, H.; Eftekhari-Yazdi, P. Effect of Maternal Age on Hippo Pathway Related Gene Expressions and Protein Localization Pattern in Human Embryos. Cell J. 2020, 22, 74–80. [Google Scholar] [CrossRef]
- Chazaud, C.; Yamanaka, Y.; Pawson, T.; Rossant, J. Early lineage segregation between epiblast and primitive endoderm in mouse blastocysts through the Grb2-MAPK pathway. Dev. Cell 2006, 10, 615–624. [Google Scholar] [CrossRef]
- Dietrich, J.E.; Hiiragi, T. Stochastic patterning in the mouse pre-implantation embryo. Development 2007, 134, 4219–4231. [Google Scholar] [CrossRef]
- Guo, G.; Huss, M.; Tong, G.Q.; Wang, C.; Sun, L.; Clarke, N.D.; Robson, P. Resolution of cell fate decisions revealed by single-cell gene expression analysis from zygote to blastocyst. Dev. Cell 2010, 18, 675–685. [Google Scholar] [CrossRef]
- Yamanaka, Y.; Lanner, F.; Rossant, J. FGF signal-dependent segregation of primitive endoderm and epiblast in the mouse blastocyst. Development 2010, 137, 715–724. [Google Scholar] [CrossRef] [PubMed]
- Roode, M.; Blair, K.; Snell, P.; Elder, K.; Marchant, S.; Smith, A.; Nichols, J. Human hypoblast formation is not dependent on FGF signalling. Dev. Biol. 2012, 361, 358–363. [Google Scholar] [CrossRef]
- Petropoulos, S.; Edsgärd, D.; Reinius, B.; Deng, Q.; Panula, S.P.; Codeluppi, S.; Reyes, A.P.; Linnarsson, S.; Sandberg, R.; Lanner, F. Single-Cell RNA-Seq Reveals Lineage and X Chromosome Dynamics in Human Preimplantation Embryos. Cell 2016, 167, 285. [Google Scholar] [CrossRef]
- Artus, J.; Piliszek, A.; Hadjantonakis, A.K. The primitive endoderm lineage of the mouse blastocyst: Sequential transcription factor activation and regulation of differentiation by Sox17. Dev. Biol. 2011, 350, 393–404. [Google Scholar] [CrossRef] [PubMed]
- Cauffman, G.; De Rycke, M.; Sermon, K.; Liebaers, I.; Van de Velde, H. Markers that define stemness in ESC are unable to identify the totipotent cells in human preimplantation embryos. Hum. Reprod. 2009, 24, 63–70. [Google Scholar] [CrossRef]
- Young, J.C.; Major, A.T.; Miyamoto, Y.; Loveland, K.L.; Jans, D.A. Distinct effects of importin α2 and α4 on Oct3/4 localization and expression in mouse embryonic stem cells. FASEB J. 2011, 25, 3958–3965. [Google Scholar] [CrossRef] [PubMed]
- Weinberger, L.; Ayyash, M.; Novershtern, N.; Hanna, J.H. Dynamic stem cell states: Naive to primed pluripotency in rodents and humans. Nat. Rev. Mol. Cell. Biol. 2016, 17, 155–169. [Google Scholar] [CrossRef]
- Pesce, M.; Anastassiadis, K.; Schöler, H.R. Oct-4: Lessons of totipotency from embryonic stem cells. Cells Tissues Organs 1999, 165, 144–152. [Google Scholar] [CrossRef]
- Jaenisch, R.; Young, R. Stem cells, the molecular circuitry of pluripotency and nuclear reprogramming. Cell 2008, 132, 567–582. [Google Scholar] [CrossRef]
- Nichols, J.; Zevnik, B.; Anastassiadis, K.; Niwa, H.; Klewe-Nebenius, D.; Chambers, I.; Schöler, H.; Smith, A. Formation of pluripotent stem cells in the mammalian embryo depends on the POU transcription factor Oct4. Cell 1998, 95, 379–391. [Google Scholar] [CrossRef]
- Okumura-Nakanishi, S.; Saito, M.; Niwa, H.; Ishikawa, F. Oct-3/4 and Sox2 regulate Oct-3/4 gene in embryonic stem cells. J. Biol. Chem. 2005, 280, 5307–5317. [Google Scholar] [CrossRef] [PubMed]
- Orkin, S.H.; Wang, J.; Kim, J.; Chu, J.; Rao, S.; Theunissen, T.W.; Shen, X.; Levasseur, D.N. The transcriptional network controlling pluripotency in ES cells. Cold Spring Harb. Symp. Quant. Biol. 2008, 73, 195–202. [Google Scholar] [CrossRef]
- Ng, H.H.; Surani, M.A. The transcriptional and signalling networks of pluripotency. Nat. Cell Biol. 2011, 13, 490–496. [Google Scholar] [CrossRef] [PubMed]
- Niwa, H. The pluripotency transcription factor network at work in reprogramming. Curr. Opin. Genet. Dev. 2014, 28, 25–31. [Google Scholar] [CrossRef] [PubMed]
- Elling, U.; Klasen, C.; Eisenberger, T.; Anlag, K.; Treier, M. Murine inner cell mass-derived lineages depend on Sall4 function. Proc. Natl. Acad. Sci. USA 2006, 103, 16319–16324. [Google Scholar] [CrossRef]
- Tang, F.; Barbacioru, C.; Bao, S.; Lee, C.; Nordman, E.; Wang, X.; Lao, K.; Surani, M.A. Tracing the derivation of embryonic stem cells from the inner cell mass by single-cell RNA-Seq analysis. Cell Stem Cell 2010, 6, 468–478. [Google Scholar] [CrossRef]
- Nishimoto, M.; Katano, M.; Yamagishi, T.; Hishida, T.; Kamon, M.; Suzuki, A.; Hirasaki, M.; Nabeshima, Y.; Nabeshima, Y.; Katsura, Y.; et al. In vivo function and evolution of the eutherian-specific pluripotency marker UTF1. PLoS ONE 2013, 8, e68119. [Google Scholar] [CrossRef]
- Madan, B.; Madan, V.; Weber, O.; Tropel, P.; Blum, C.; Kieffer, E.; Viville, S.; Fehling, H.J. The pluripotency-associated gene Dppa4 is dispensable for embryonic stem cell identity and germ cell development but essential for embryogenesis. Mol. Cell. Biol. 2009, 29, 3186–3203. [Google Scholar] [CrossRef] [PubMed]
- Mitsui, K.; Tokuzawa, Y.; Itoh, H.; Segawa, K.; Murakami, M.; Takahashi, K.; Maruyama, M.; Maeda, M.; Yamanaka, S. The homeoprotein Nanog is required for maintenance of pluripotency in mouse epiblast and ES cells. Cell 2003, 113, 631–642. [Google Scholar] [CrossRef]
- Casanova, E.A.; Okoniewski, M.J.; Cinelli, P. Cross-species genome wide expression analysis during pluripotent cell determination in mouse and rat preimplantation embryos. PLoS ONE 2012, 7, e47107. [Google Scholar] [CrossRef]
- Taylor, D.M.; Ray, P.F.; Ao, A.; Winston, R.M.; Handyside, A.H. Paternal transcripts for glucose-6-phosphate dehydrogenase and adenosine deaminase are first detectable in the human preimplantation embryo at the three- to four-cell stage. Mol. Reprod. Dev. 1997, 48, 442–448. [Google Scholar] [CrossRef]
- Vassena, R.; Boué, S.; González-Roca, E.; Aran, B.; Auer, H.; Veiga, A.; Izpisua Belmonte, J.C. Waves of early transcriptional activation and pluripotency program initiation during human preimplantation development. Development 2011, 138, 3699–3709. [Google Scholar] [CrossRef]
- Braude, P.; Bolton, V.; Moore, S. Human gene expression first occurs between the four- and eight-cell stages of preimplantation development. Nature 1988, 332, 459–461. [Google Scholar] [CrossRef] [PubMed]
- Theunissen, T.W.; Friedli, M.; He, Y.; Planet, E.; O’Neil, R.C.; Markoulaki, S.; Pontis, J.; Wang, H.; Iouranova, A.; Imbeault, M.; et al. Molecular criteria for defining the naive human pluripotent state. Cell Stem Cell 2016, 19, 502–515. [Google Scholar] [CrossRef] [PubMed]
- Grow, E.J.; Flynn, R.A.; Chavez, S.L.; Bayless, N.L.; Wossidlo, M.; Wesche, D.J.; Martin, L.; Ware, C.B.; Blish, C.A.; Chang, H.Y.; et al. Intrinsic retroviral reactivation in human preimplantation embryos and pluripotent cells. Nature 2015, 522, 221–225. [Google Scholar] [CrossRef]
- Shahbazi, M.N. Mechanisms of human embryo development: From cell fate to tissue shape and back. Development 2020, 147, dev190629. [Google Scholar] [CrossRef] [PubMed]
- Alikani, M. Epithelial cadherin distribution in abnormal human pre-implantation embryos. Hum. Reprod. 2005, 20, 3369–3375. [Google Scholar] [CrossRef] [PubMed]
- Nikas, G.; Ao, A.; Winston, R.M.; Handyside, A.H. Compaction and surface polarity in the human embryo in vitro. Biol. Reprod. 1996, 55, 32–37. [Google Scholar] [CrossRef]
- De Paepe, C.; Cauffman, G.; Verloes, A.; Sterckx, J.; Devroey, P.; Tournaye, H.; Liebaers, I.; Van de Velde, H. Human trophectoderm cells are not yet committed. Hum. Reprod. 2013, 28, 740–749. [Google Scholar] [CrossRef]
- Guo, S.; Cui, X.; Jiang, X.; Duo, S.; Li, S.; Gao, F.; Wang, H. Tracing the origin of the placental trophoblast cells in mouse embryo development†. Biol. Reprod. 2020, 102, 598–606. [Google Scholar] [CrossRef] [PubMed]
- Royer, C.; Leonavicius, K.; Kip, A.; Fortin, D.; Nandi, K.; Vincent, A.; Jones, C.; Child, T.; Coward, K.; Graham, C.; et al. Establishment of a relationship between blastomere geometry and YAP localisation during compaction. Development 2020, 147, dev189449. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Yabuuchi, A.; Eminli, S.; Takeuchi, A.; Lu, C.W.; Hochedlinger, K.; Daley, G.Q. Cross-regulation of the Nanog and Cdx2 promoters. Cell Res. 2009, 19, 1052–1061. [Google Scholar] [CrossRef]
- Hyslop, L.; Stojkovic, M.; Armstrong, L.; Walter, T.; Stojkovic, P.; Przyborski, S.; Herbert, M.; Murdoch, A.; Strachan, T.; Lako, M. Downregulation of NANOG induces differentiation of human embryonic stem cells to extraembryonic lineages. Stem Cells 2005, 23, 1035–1043. [Google Scholar] [CrossRef] [PubMed]
- Zhou, F.; Wang, R.; Yuan, P.; Ren, Y.; Mao, Y.; Li, R.; Lian, Y.; Li, J.; Wen, L.; Yan, L.; et al. Reconstituting the transcriptome and DNA methylome landscapes of human implantation. Nature 2019, 572, 660–664. [Google Scholar] [CrossRef]
- Xiang, L.; Yin, Y.; Zheng, Y.; Ma, Y.; Li, Y.; Zhao, Z.; Guo, J.; Ai, Z.; Niu, Y.; Duan, K.; et al. A developmental landscape of 3D-cultured human pre-gastrulation embryos. Nature 2020, 577, 537–542. [Google Scholar] [CrossRef]
- Messerschmidt, D.M. Should I stay or should I go: Protection and maintenance of DNA methylation at imprinted genes. Epigenetics 2012, 7, 969–975. [Google Scholar] [CrossRef]
- Denomme, M.M.; Mann, M.R. Maternal control of genomic imprint maintenance. Reprod. Biomed. Online 2013, 27, 629–636. [Google Scholar] [CrossRef]
- Oswald, J.; Engemann, S.; Lane, N.; Mayer, W.; Olek, A.; Fundele, R.; Dean, W.; Reik, W.; Walter, J. Active demethylation of the paternal genome in the mouse zygote. Curr. Biol. 2000, 10, 475–478. [Google Scholar] [CrossRef]
- Santos, F.; Hendrich, B.; Reik, W.; Dean, W. Dynamic reprogramming of DNA methylation in the early mouse embryo. Dev. Biol. 2002, 241, 172–182. [Google Scholar] [CrossRef] [PubMed]
- Santos, F.; Peat, J.; Burgess, H.; Rada, C.; Reik, W.; Dean, W. Active demethylation in mouse zygotes involves cytosine deamination and base excision repair. Epigenet. Chromatin 2013, 6, 39. [Google Scholar] [CrossRef]
- Abdalla, H.; Yoshizawa, Y.; Hochi, S. Active demethylation of paternal genome in mammalian zygotes. J. Reprod. Dev. 2009, 55, 356–360. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Puschendorf, M.; Terranova, R.; Boutsma, E.; Mao, X.; Isono, K.; Brykczynska, U.; Kolb, C.; Otte, A.P.; Koseki, H.; Orkin, S.H.; et al. PRC1 and Suv39h specify parental asymmetry at constitutive heterochromatin in early mouse embryos. Nat. Genet. 2008, 40, 411–420. [Google Scholar] [CrossRef] [PubMed]
- Lepikhov, K.; Walter, J. Differential dynamics of histone H3 methylation at positions K4 and K9 in the mouse zygote. BMC Dev. Biol. 2004, 4, 12. [Google Scholar] [CrossRef]
- Santos, F.; Dean, W. Epigenetic reprogramming during early development in mammals. Reproduction 2004, 127, 643–651. [Google Scholar] [CrossRef]
- Sarmento, O.F.; Digilio, L.C.; Wang, Y.; Perlin, J.; Herr, J.C.; Allis, C.D.; Coonrod, S.A. Dynamic alterations of specific histone modifications during early murine development. J. Cell Sci. 2004, 117, 4449–4459. [Google Scholar] [CrossRef]
- Hales, B.F.; Grenier, L.; Lalancette, C.; Robaire, B. Epigenetic programming: From gametes to blastocyst. Birth Defects Res. A Clin. Mol. Teratol. 2011, 91, 652–665. [Google Scholar] [CrossRef]
- Nakamura, T.; Arai, Y.; Umehara, H.; Masuhara, M.; Kimura, T.; Taniguchi, H.; Sekimoto, T.; Ikawa, M.; Yoneda, Y.; Okabe, M.; et al. PGC7/Stella protects against DNA demethylation in early embryogenesis. Nat. Cell Biol. 2007, 9, 64–71. [Google Scholar] [CrossRef]
- Lepikhov, K.; Zakhartchenko, V.; Hao, R.; Yang, F.; Wrenzycki, C.; Niemann, H.; Wolf, E.; Walter, J. Evidence for conserved DNA and histone H3 methylation reprogramming in mouse, bovine and rabbit zygotes. Epigenet. Chromatin 2008, 1, 8. [Google Scholar] [CrossRef]
- Zhang, W.; Xia, W.; Wang, Q.; Towers, A.J.; Chen, J.; Gao, R.; Zhang, Y.; Yen, C.A.; Lee, A.Y.; Li, Y.; et al. Isoform Switch of TET1 Regulates DNA Demethylation and Mouse Development. Mol. Cell 2016, 64, 1062–1073. [Google Scholar] [CrossRef]
- Ancelin, K.; Syx, L.; Borensztein, M.; Ranisavljevic, N.; Vassilev, I.; Briseño-Roa, L.; Liu, T.; Metzger, E.; Servant, N.; Barillot, E.; et al. Maternal LSD1/KDM1A is an essential regulator of chromatin and transcription landscapes during zygotic genome activation. Elife 2016, 5, e08851. [Google Scholar] [CrossRef]
- Kourmouli, N.; Jeppesen, P.; Mahadevhaiah, S.; Burgoyne, P.; Wu, R.; Gilbert, D.M.; Bongiorni, S.; Prantera, G.; Fanti, L.; Pimpinelli, S.; et al. Heterochromatin and tri-methylated lysine 20 of histone H4 in animals. J. Cell Sci. 2004, 117, 2491–2501. [Google Scholar] [CrossRef] [PubMed]
- Daujat, S.; Weiss, T.; Mohn, F.; Lange, U.C.; Ziegler-Birling, C.; Zeissler, U.; Lappe, M.; Schübeler, D.; Torres-Padilla, M.E.; Schneider, R. H3K64 trimethylation marks heterochromatin and is dynamically remodeled during developmental reprogramming. Nat. Struct. Mol. Biol. 2009, 16, 777–781. [Google Scholar] [CrossRef]
- Becker, J.S.; Nicetto, D.; Zaret, K.S. H3K9me3-Dependent Heterochromatin: Barrier to Cell Fate Changes. Trends Genet. 2016, 32, 29–41. [Google Scholar] [CrossRef]
- Santos, F.; Peters, A.H.; Otte, A.P.; Reik, W.; Dean, W. Dynamic chromatin modifications characterise the first cell cycle in mouse embryos. Dev. Biol. 2005, 280, 225–236. [Google Scholar] [CrossRef] [PubMed]
- Stein, P.; Worrad, D.M.; Belyaev, N.D.; Turner, B.M.; Schultz, R.M. Stage-dependent redistributions of acetylated histones in nuclei of the early preimplantation mouse embryo. Mol. Reprod. Dev. 1997, 47, 421–429. [Google Scholar] [CrossRef]
- Auclair, G.; Guibert, S.; Bender, A.; Weber, M. Ontogeny of CpG island methylation and specificity of DNMT3 methyltransferases during embryonic development in the mouse. Genome Biol. 2014, 15, 545. [Google Scholar] [CrossRef]
- Ivanova, E.; Canovas, S.; Garcia-Martínez, S.; Romar, R.; Lopes, J.S.; Rizos, D.; Sanchez-Calabuig, M.J.; Krueger, F.; Andrews, S.; Perez-Sanz, F.; et al. DNA methylation changes during preimplantation development reveal inter-species differences and reprogramming events at imprinted genes. Clin. Epigenet. 2020, 12, 64. [Google Scholar] [CrossRef] [PubMed]
- Lessard, J.A.; Crabtree, G.R. Chromatin regulatory mechanisms in pluripotency. Annu. Rev. Cell Dev. Biol. 2010, 26, 503–532. [Google Scholar] [CrossRef] [PubMed]
- Zylicz, J.J.; Dietmann, S.; Günesdogan, U.; Hackett, J.A.; Cougot, D.; Lee, C.; Surani, M.A. Chromatin dynamics and the role of G9a in gene regulation and enhancer silencing during early mouse development. Elife 2015, 4, e09571. [Google Scholar] [CrossRef]
- Liu, X.; Wang, C.; Liu, W.; Li, J.; Li, C.; Kou, X.; Chen, J.; Zhao, Y.; Gao, H.; Wang, H.; et al. Distinct features of H3K4me3 and H3K27me3 chromatin domains in pre-implantation embryos. Nature 2016, 537, 558–562. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, L.P.; VerMilyea, M.D.; Turner, B.M. Epigenetic characterization of the early embryo with a chromatin immunoprecipitation protocol applicable to small cell populations. Nat. Genet. 2006, 38, 835–841. [Google Scholar] [CrossRef]
- VerMilyea, M.D.; O’Neill, L.P.; Turner, B.M. Transcription-independent heritability of induced histone modifications in the mouse preimplantation embryo. PLoS ONE 2009, 4, e6086. [Google Scholar] [CrossRef] [PubMed]
- Heintzman, N.D.; Stuart, R.K.; Hon, G.; Fu, Y.; Ching, C.W.; Hawkins, R.D.; Barrera, L.O.; Van Calcar, S.; Qu, C.; Ching, K.A.; et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat. Genet. 2007, 39, 311–318. [Google Scholar] [CrossRef]
- Barski, A.; Cuddapah, S.; Cui, K.; Roh, T.Y.; Schones, D.E.; Wang, Z.; Wei, G.; Chepelev, I.; Zhao, K. High-resolution profiling of histone methylations in the human genome. Cell 2007, 129, 823–837. [Google Scholar] [CrossRef]
- Vastenhouw, N.L.; Schier, A.F. Bivalent histone modifications in early embryogenesis. Curr. Opin. Cell Biol. 2012, 24, 374–386. [Google Scholar] [CrossRef] [PubMed]
- Rugg-Gunn, P.J.; Cox, B.J.; Ralston, A.; Rossant, J. Distinct histone modifications in stem cell lines and tissue lineages from the early mouse embryo. Proc. Natl. Acad. Sci. USA 2010, 107, 10783–10790. [Google Scholar] [CrossRef]
- Fulka, H.; Mrazek, M.; Tepla, O.; Fulka, J., Jr. DNA methylation pattern in human zygotes and developing embryos. Reproduction 2004, 128, 703–708. [Google Scholar] [CrossRef]
- Okae, H.; Chiba, H.; Hiura, H.; Hamada, H.; Sato, A.; Utsunomiya, T.; Kikuchi, H.; Yoshida, H.; Tanaka, A.; Suyama, M.; et al. Genome-wide analysis of DNA methylation dynamics during early human development. PLoS Genet. 2014, 10, e1004868. [Google Scholar] [CrossRef]
- Smith, Z.D.; Chan, M.M.; Humm, K.C.; Karnik, R.; Mekhoubad, S.; Regev, A.; Eggan, K.; Meissner, A. DNA methylation dynamics of the human preimplantation embryo. Nature 2014, 511, 611–615. [Google Scholar] [CrossRef]
- Guo, H.; Zhu, P.; Yan, L.; Li, R.; Hu, B.; Lian, Y.; Yan, J.; Ren, X.; Lin, S.; Li, J.; et al. The DNA methylation landscape of human early embryos. Nature 2014, 511, 606–610. [Google Scholar] [CrossRef]
- Zhu, P.; Guo, H.; Ren, Y.; Hou, Y.; Dong, J.; Li, R.; Lian, Y.; Fan, X.; Hu, B.; Gao, Y.; et al. Single-cell DNA methylome sequencing of human preimplantation embryos. Nat. Genet. 2018, 50, 12–19. [Google Scholar] [CrossRef]
- Xia, W.; Xu, J.; Yu, G.; Yao, G.; Xu, K.; Ma, X.; Zhang, N.; Liu, B.; Li, T.; Lin, Z.; et al. Resetting histone modifications during human parental-to-zygotic transition. Science 2019, 365, 353–360. [Google Scholar] [CrossRef] [PubMed]
- Xu, R.; Li, C.; Liu, X.; Gao, S. Insights into epigenetic patterns in mammalian early embryos. Protein Cell 2021, 12, 7–28. [Google Scholar] [CrossRef] [PubMed]
- Saha, B.; Home, P.; Ray, S.; Larson, M.; Paul, A.; Rajendran, G.; Behr, B.; Paul, S. EED and KDM6B coordinate the first mammalian cell lineage commitment to ensure embryo implantation. Mol. Cell. Biol. 2013, 33, 2691–2705. [Google Scholar] [CrossRef]
- Inoue, A.; Jiang, L.; Lu, F.; Zhang, Y. Genomic imprinting of Xist by maternal H3K27me3. Genes Dev. 2017, 31, 1927–1932. [Google Scholar] [CrossRef]
- Smith, Z.D.; Chan, M.M.; Mikkelsen, T.S.; Gu, H.; Gnirke, A.; Regev, A.; Meissner, A. A unique regulatory phase of DNA methylation in the early mammalian embryo. Nature 2012, 484, 339–344. [Google Scholar] [CrossRef] [PubMed]
- Rebuzzini, P.; Zuccotti, M.; Garagna, S. X-Chromosome Inactivation during Preimplantation Development and in Pluripotent Stem Cells. Cytogenet. Genome Res. 2020, 160, 283–294. [Google Scholar] [CrossRef]
- Disteche, C.M. Dosage compensation of the sex chromosomes and autosomes. Semin. Cell Dev. Biol. 2016, 56, 9–18. [Google Scholar] [CrossRef]
- Jeon, Y.; Sarma, K.; Lee, J.T. New and Xisting regulatory mechanisms of X chromosome inactivation. Curr. Opin. Genet. Dev. 2012, 22, 62–71. [Google Scholar] [CrossRef]
- Mak, W.; Nesterova, T.B.; de Napoles, M.; Appanah, R.; Yamanaka, S.; Otte, A.P.; Brockdorff, N. Reactivation of the paternal X chromosome in early mouse embryos. Science 2004, 303, 666–669. [Google Scholar] [CrossRef]
- Borensztein, M.; Syx, L.; Ancelin, K.; Diabangouaya, P.; Picard, C.; Liu, T.; Liang, J.B.; Vassilev, I.; Galupa, R.; Servant, N.; et al. Xist-dependent imprinted X inactivation and the early developmental consequences of its failure. Nat. Struct. Mol. Biol. 2017, 24, 226–233. [Google Scholar] [CrossRef] [PubMed]
- Pasque, V.; Plath, K. X chromosome reactivation in reprogramming and in development. Curr. Opin. Cell Biol. 2015, 37, 75–83. [Google Scholar] [CrossRef]
- Talon, I.; Janiszewski, A.; Chappell, J.; Vanheer, L.; Pasque, V. Recent Advances in Understanding the Reversal of Gene Silencing During X Chromosome Reactivation. Front. Cell Dev. Biol. 2019, 7, 169. [Google Scholar] [CrossRef] [PubMed]
- Williams, L.H.; Kalantry, S.; Starmer, J.; Magnuson, T. Transcription precedes loss of Xist coating and depletion of H3K27me3 during X-chromosome reprogramming in the mouse inner cell mass. Development 2011, 138, 2049–2057. [Google Scholar] [CrossRef] [PubMed]
- Okamoto, I.; Otte, A.P.; Allis, C.D.; Reinberg, D.; Heard, E. Epigenetic dynamics of imprinted X inactivation during early mouse development. Science 2004, 303, 644–649. [Google Scholar] [CrossRef] [PubMed]
- Maherali, N.; Sridharan, R.; Xie, W.; Utikal, J.; Eminli, S.; Arnold, K.; Stadtfeld, M.; Yachechko, R.; Tchieu, J.; Jaenisch, R.; et al. Directly reprogrammed fibroblasts show global epigenetic remodeling and widespread tissue contribution. Cell Stem Cell 2007, 1, 55–70. [Google Scholar] [CrossRef]
- Pasque, V.; Tchieu, J.; Karnik, R.; Uyeda, M.; Sadhu Dimashkie, A.; Case, D.; Papp, B.; Bonora, G.; Patel, S.; Ho, R.; et al. X chromosome reactivation dynamics reveal stages of reprogramming to pluripotency. Cell 2014, 159, 1681–1697. [Google Scholar] [CrossRef]
- Lyon, M.F. Gene action in the X-chromosome of the mouse (Mus musculus L.). Nature 1961, 190, 372–373. [Google Scholar] [CrossRef]
- Saiba, R.; Arava, M.; Gayen, S. Dosage compensation in human pre-implantation embryos: X-chromosome inactivation or dampening? EMBO Rep. 2018, 19, e46294. [Google Scholar] [CrossRef] [PubMed]
- Patrat, C.; Ouimette, J.F.; Rougeulle, C. X chromosome inactivation in human development. Development 2020, 147, dev183095. [Google Scholar] [CrossRef] [PubMed]
- Sahakyan, A.; Kim, R.; Chronis, C.; Sabri, S.; Bonora, G.; Theunissen, T.W.; Kuoy, E.; Langerman, J.; Clark, A.T.; Jaenisch, R.; et al. Human Naive Pluripotent Stem Cells Model X Chromosome Dampening and X Inactivation. Cell Stem Cell 2017, 20, 87–101. [Google Scholar] [CrossRef] [PubMed]
- Gross, N.; Kropp, J.; Khatib, H. MicroRNA Signaling in Embryo Development. Biology 2017, 6, 34. [Google Scholar] [CrossRef]
- Salilew-Wondim, D.; Gebremedhn, S.; Hoelker, M.; Tholen, E.; Hailay, T.; Tesfaye, D. The Role of MicroRNAs in Mammalian Fertility: From Gametogenesis to Embryo Implantation. Int. J. Mol. Sci. 2020, 21, 585. [Google Scholar] [CrossRef]
- Mineno, J.; Okamoto, S.; Ando, T.; Sato, M.; Chono, H.; Izu, H.; Takayama, M.; Asada, K.; Mirochnitchenko, O.; Inouye, M.; et al. The expression profile of microRNAs in mouse embryos. Nucleic Acids Res. 2006, 34, 1765–1771. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Bai, W.; Zhang, L.; Yin, G.; Wang, X.; Wang, J.; Zhao, H.; Han, Y.; Yao, Y.Q. Determination of microRNAs in mouse preimplantation embryos by microarray. Dev. Dyn. 2008, 237, 2315–2327. [Google Scholar] [CrossRef] [PubMed]
- Tang, F.; Kaneda, M.; O’Carroll, D.; Hajkova, P.; Barton, S.C.; Sun, Y.A.; Lee, C.; Tarakhovsky, A.; Lao, K.; Surani, M.A. Maternal microRNAs are essential for mouse zygotic development. Genes Dev. 2007, 21, 644–648. [Google Scholar] [CrossRef]
- Jodar, M.; Selvaraju, S.; Sendler, E.; Diamond, M.P.; Krawetz, S.A. Reproductive Medicine Network. The presence, role and clinical use of spermatozoal RNAs. Hum. Reprod. Update 2013, 19, 604–624. [Google Scholar] [CrossRef]
- Liu, W.M.; Pang, R.T.; Chiu, P.C.; Wong, B.P.; Lao, K.; Lee, K.F.; Yeung, W.S. Sperm-borne microRNA-34c is required for the first cleavage division in mouse. Proc. Natl. Acad. Sci. USA 2012, 109, 490–494. [Google Scholar] [CrossRef]
- Yang, Q.; Lin, J.; Liu, M.; Li, R.; Tian, B.; Zhang, X.; Xu, B.; Liu, M.; Zhang, X.; Li, Y.; et al. Highly sensitive sequencing reveals dynamic modifications and activities of small RNAs in mouse oocytes and early embryos. Sci. Adv. 2016, 2, e1501482. [Google Scholar] [CrossRef]
- García-López, J.; del Mazo, J. Expression dynamics of microRNA biogenesis during preimplantation mouse development. Biochim. Biophys. Acta 2012, 1819, 847–854. [Google Scholar] [CrossRef] [PubMed]
- Tan, M.H.; Au, K.F.; Leong, D.E.; Foygel, K.; Wong, W.H.; Yao, M.W. An Oct4-Sall4-Nanog network controls developmental progression in the pre-implantation mouse embryo. Mol. Syst. Biol. 2013, 9, 632. [Google Scholar] [CrossRef] [PubMed]
- Borges, E., Jr.; Setti, A.S.; Braga, D.P.; Geraldo, M.V.; Figueira, R.C.; Iaconelli, A., Jr. miR-142-3p as a biomarker of blastocyst implantation failure—A pilot study. JBRA Assist. Reprod. 2016, 20, 200–205. [Google Scholar] [CrossRef] [PubMed]
- Capalbo, A.; Ubaldi, F.M.; Cimadomo, D.; Noli, L.; Khalaf, Y.; Farcomeni, A.; Ilic, D.; Rienzi, L. MicroRNAs in spent blastocyst culture medium are derived from trophectoderm cells and can be explored for human embryo reproductive competence assessment. Fertil. Steril. 2016, 105, 225–235. [Google Scholar] [CrossRef]
- Abu-Halima, M.; Khaizaran, Z.A.; Ayesh, B.M.; Fischer, U.; Khaizaran, S.A.; Al-Battah, F.; Hammadeh, M.; Keller, A.; Meese, E. MicroRNAs in combined spent culture media and sperm are associated with embryo quality and pregnancy outcome. Fertil. Steril. 2020, 113, 970–980. [Google Scholar] [CrossRef]
- Bouckenheimer, J.; Assou, S.; Riquier, S.; Hou, C.; Philippe, N.; Sansac, C.; Lavabre-Bertrand, T.; Commes, T.; Lemaître, J.M.; Boureux, A.; et al. Long non-coding RNAs in human early embryonic development and their potential in ART. Hum. Reprod. Update 2016, 23, 19–40. [Google Scholar] [CrossRef]
- Zhang, K.; Huang, K.; Luo, Y.; Li, S. Identification and functional analysis of long non-coding RNAs in mouse cleavage stage embryonic development based on single cell transcriptome data. BMC Genom. 2014, 15, 845. [Google Scholar] [CrossRef]
- Qiu, J.J.; Ren, Z.R.; Yan, J.B. Identification and functional analysis of long non-coding RNAs in human and mouse early embryos based on single-cell transcriptome data. Oncotarget 2016, 7, 61215–61228. [Google Scholar] [CrossRef]
- Yunusov, D.; Anderson, L.; DaSilva, L.F.; Wysocka, J.; Ezashi, T.; Roberts, R.M.; Verjovski-Almeida, S. HIPSTR and thousands of lncRNAs are heterogeneously expressed in human embryos, primordial germ cells and stable cell lines. Sci. Rep. 2016, 6, 32753. [Google Scholar] [CrossRef]
- Wang, J.; Wang, L.; Feng, G.; Wang, Y.; Li, Y.; Li, X.; Liu, C.; Jiao, G.; Huang, C.; Shi, J.; et al. Asymmetric Expression of LincGET Biases Cell Fate in Two-Cell Mouse Embryos. Cell 2018, 175, 1887–1901. [Google Scholar] [CrossRef]
- Brown, C.J.; Hendrich, B.D.; Rupert, J.L.; Lafrenière, R.G.; Xing, Y.; Lawrence, J.; Willard, H.F. The human XIST gene: Analysis of a 17 kb inactive X-specific RNA that contains conserved repeats and is highly localized within the nucleus. Cell 1992, 71, 527–542. [Google Scholar] [CrossRef]
- Clemson, C.M.; Chow, J.C.; Brown, C.J.; Lawrence, J.B. Stabilization and localization of Xist RNA are controlled by separate mechanisms and are not sufficient for X inactivation. J. Cell Biol. 1998, 142, 13–23. [Google Scholar] [CrossRef]
- Lee, J.T. Epigenetic regulation by long noncoding RNAs. Science 2012, 338, 1435–1439. [Google Scholar] [CrossRef]
- McHugh, C.A.; Chen, C.K.; Chow, A.; Surka, C.F.; Tran, C.; McDonel, P.; Pandya-Jones, A.; Blanco, M.; Burghard, C.; Moradian, A.; et al. The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3. Nature 2015, 521, 232–236. [Google Scholar] [CrossRef] [PubMed]
- Minajigi, A.; Froberg, J.; Wei, C.; Sunwoo, H.; Kesner, B.; Colognori, D.; Lessing, D.; Payer, B.; Boukhali, M.; Haas, W.; et al. Chromosomes. A comprehensive Xist interactome reveals cohesin repulsion and an RNA-directed chromosome conformation. Science 2015, 349, aab2276. [Google Scholar] [CrossRef]
- Almeida, M.; Pintacuda, G.; Masui, O.; Koseki, Y.; Gdula, M.; Cerase, A.; Brown, D.; Mould, A.; Innocent, C.; Nakayama, M.; et al. PCGF3/5-PRC1 initiates Polycomb recruitment in X chromosome inactivation. Science 2017, 356, 1081–1084. [Google Scholar] [CrossRef]
- Moindrot, B.; Brockdorff, N. RNA binding proteins implicated in Xist-mediated chromosome silencing. Semin. Cell Dev. Biol. 2016, 56, 58–70. [Google Scholar] [CrossRef]
- Mira-Bontenbal, H.; Gribnau, J. New Xist-Interacting Proteins in X-Chromosome Inactivation. Curr. Biol. 2016, 26, 1383. [Google Scholar] [CrossRef] [PubMed]
- da Rocha, S.T.; Heard, E. Novel players in X inactivation: Insights into Xist-mediated gene silencing and chromosome conformation. Nat. Struct. Mol. Biol. 2017, 24, 197–204. [Google Scholar] [CrossRef]
- Loda, A.; Heard, E. Xist RNA in action: Past, present, and future. PLoS Genet. 2019, 15, e1008333. [Google Scholar] [CrossRef] [PubMed]
- Vallot, C.; Huret, C.; Lesecque, Y.; Resch, A.; Oudrhiri, N.; Bennaceur-Griscelli, A.; Duret, L.; Rougeulle, C. XACT, a long noncoding transcript coating the active X chromosome in human pluripotent cells. Nat. Genet. 2013, 45, 239–241. [Google Scholar] [CrossRef]
- Vallot, C.; Patrat, C.; Collier, A.J.; Huret, C.; Casanova, M.; Liyakat Ali, T.M.; Tosolini, M.; Frydman, N.; Heard, E.; Rugg-Gunn, P.J.; et al. XACT Noncoding RNA Competes with XIST in the Control of X Chromosome Activity during Human Early Development. Cell Stem Cell 2017, 20, 102–111. [Google Scholar] [CrossRef]
- Wang, J.; Li, X.; Wang, L.; Li, J.; Zhao, Y.; Bou, G.; Li, Y.; Jiao, G.; Shen, X.; Wei, R.; et al. A novel long intergenic noncoding RNA indispensable for the cleavage of mouse two-cell embryos. EMBO Rep. 2016, 17, 1452–1470. [Google Scholar] [CrossRef]
- Hupalowska, A.; Jedrusik, A.; Zhu, M.; Bedford, M.T.; Glover, D.M.; Zernicka-Goetz, M. CARM1 and Paraspeckles Regulate Pre-implantation Mouse Embryo Development. Cell 2018, 175, 1902–1916. [Google Scholar] [CrossRef] [PubMed]
- Boutourlinsky, K.; Allègre, N.; Chazaud, C. Ex Vivo Culture for Preimplantation Mouse Embryo to Analyze Pluripotency. Methods Mol Biol. 2021, 2214, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Nichols, J.; Smith, A. Pluripotency in the embryo and in culture. Cold Spring Harb. Perspect. Biol. 2012, 4, a008128. [Google Scholar] [CrossRef] [PubMed]
- Nichols, J.; Smith, A. Naive and primed pluripotent states. Cell Stem Cell 2009, 4, 487–492. [Google Scholar] [CrossRef] [PubMed]
- Devika, A.S.; Wruck, W.; Adjaye, J.; Sudheer, S. The quest for pluripotency: A comparative analysis across mammalian species. Reproduction 2019, 158, R97–R111. [Google Scholar] [CrossRef]
- Boroviak, T.; Loos, R.; Bertone, P.; Smith, A.; Nichols, J. The ability of inner-cell-mass cells to self-renew as embryonic stem cells is acquired following epiblast specification. Nat. Cell Biol. 2014, 16, 516–528. [Google Scholar] [CrossRef]
- Czechanski, A.; Byers, C.; Greenstein, I.; Schrode, N.; Donahue, L.R.; Hadjantonakis, A.K.; Reinholdt, L.G. Derivation and characterization of mouse embryonic stem cells from permissive and nonpermissive strains. Nat. Protoc. 2014, 9, 559–574. [Google Scholar] [CrossRef]
- Kunath, T.; Saba-El-Leil, M.K.; Almousailleakh, M.; Wray, J.; Meloche, S.; Smith, A. FGF stimulation of the Erk1/2 signalling cascade triggers transition of pluripotent embryonic stem cells from self-renewal to lineage commitment. Development 2007, 134, 2895–2902. [Google Scholar] [CrossRef]
- Singh, A.M.; Hamazaki, T.; Hankowski, K.E.; Terada, N. A heterogeneous expression pattern for Nanog in embryonic stem cells. Stem Cells 2007, 25, 2534–2542. [Google Scholar] [CrossRef]
- Canham, M.A.; Sharov, A.A.; Ko, M.S.; Brickman, J.M. Functional heterogeneity of embryonic stem cells revealed through translational amplification of an early endodermal transcript. PLoS Biol. 2010, 8, e1000379. [Google Scholar] [CrossRef]
- Lanner, F.; Rossant, J. The role of FGF/Erk signaling in pluripotent cells. Development 2010, 137, 3351–3360. [Google Scholar] [CrossRef]
- Morgani, S.M.; Canham, M.A.; Nichols, J.; Sharov, A.A.; Migueles, R.P.; Ko, M.S.; Brickman, J.M. Totipotent embryonic stem cells arise in ground-state culture conditions. Cell Rep. 2013, 3, 1945–1957. [Google Scholar] [CrossRef] [PubMed]
- Illingworth, R.S.; Hölzenspies, J.J.; Roske, F.V.; Bickmore, W.A.; Brickman, J.M. Polycomb enables primitive endoderm lineage priming in embryonic stem cells. Elife 2016, 5, e14926. [Google Scholar] [CrossRef]
- Li, M.; Izpisua Belmonte, J.C. Deconstructing the pluripotency gene regulatory network. Nat. Cell Biol. 2018, 20, 382–392. [Google Scholar] [CrossRef]
- Fu, X.; Djekidel, M.N.; Zhang, Y. A transcriptional roadmap for 2C-like-to-pluripotent state transition. Sci. Adv. 2020, 6, eaay5181. [Google Scholar] [CrossRef]
- Amano, T.; Hirata, T.; Falco, G.; Monti, M.; Sharova, L.V.; Amano, M.; Sheer, S.; Hoang, H.G.; Piao, Y.; Stagg, C.A.; et al. Zscan4 restores the developmental potency of embryonic stem cells. Nat. Commun. 2013, 4, 1966. [Google Scholar] [CrossRef]
- Bošković, A.; Eid, A.; Pontabry, J.; Ishiuchi, T.; Spiegelhalter, C.; Raghu Ram, E.V.; Meshorer, E.; Torres-Padilla, M.E. Higher chromatin mobility supports totipotency and precedes pluripotency in vivo. Genes Dev. 2014, 28, 1042–1047. [Google Scholar] [CrossRef] [PubMed]
- Ishiuchi, T.; Enriquez-Gasca, R.; Mizutani, E.; Bošković, A.; Ziegler-Birling, C.; Rodriguez-Terrones, D.; Wakayama, T.; Vaquerizas, J.M.; Torres-Padilla, M.E. Early embryonic-like cells are induced by downregulating replication-dependent chromatin assembly. Nat. Struct. Mol. Biol. 2015, 22, 662–671. [Google Scholar] [CrossRef] [PubMed]
- Lu, F.; Liu, Y.; Jiang, L.; Yamaguchi, S.; Zhang, Y. Role of Tet proteins in enhancer activity and telomere elongation. Genes Dev. 2014, 28, 2103–2119. [Google Scholar] [CrossRef]
- Choi, Y.J.; Lin, C.P.; Risso, D.; Chen, S.; Kim, T.A.; Tan, M.H.; Li, J.B.; Wu, Y.; Chen, C.; Xuan, Z.; et al. Deficiency of microRNA miR-34a expands cell fate potential in pluripotent stem cells. Science 2017, 355, eaag1927. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Terrones, D.; Gaume, X.; Ishiuchi, T.; Weiss, A.; Kopp, A.; Kruse, K.; Penning, A.; Vaquerizas, J.M.; Brino, L.; Torres-Padilla, M.E. A molecular roadmap for the emergence of early-embryonic-like cells in culture. Nat. Genet. 2018, 50, 106–119. [Google Scholar] [CrossRef] [PubMed]
- Fu, X.; Wu, X.; Djekidel, M.N.; Zhang, Y. Myc and Dnmt1 impede the pluripotent to totipotent state transition in embryonic stem cells. Nat. Cell Biol. 2019, 21, 835–844. [Google Scholar] [CrossRef]
- Ying, Q.L.; Wray, J.; Nichols, J.; Batlle-Morera, L.; Doble, B.; Woodgett, J.; Cohen, P.; Smith, A. The ground state of embryonic stem cell self-renewal. Nature 2008, 453, 519–523. [Google Scholar] [CrossRef] [PubMed]
- Wray, J.; Kalkan, T.; Gomez-Lopez, S.; Eckardt, D.; Cook, A.; Kemler, R.; Smith, A. Inhibition of glycogen synthase kinase-3 alleviates Tcf3 repression of the pluripotency network and increases embryonic stem cell resistance to differentiation. Nat Cell Biol. 2011, 13, 838–845. [Google Scholar] [CrossRef] [PubMed]
- Wray, J.; Kalkan, T.; Smith, A.G. The ground state of pluripotency. Biochem. Soc. Trans. 2010, 38, 1027–1032. [Google Scholar] [CrossRef]
- Marks, H.; Kalkan, T.; Menafra, R.; Denissov, S.; Jones, K.; Hofemeister, H.; Nichols, J.; Kranz, A.; Stewart, A.F.; Smith, A.; et al. The transcriptional and epigenomic foundations of ground state pluripotency. Cell 2012, 149, 590–604. [Google Scholar] [CrossRef]
- Habibi, E.; Brinkman, A.B.; Arand, J.; Kroeze, L.I.; Kerstens, H.H.; Matarese, F.; Lepikhov, K.; Gut, M.; Brun-Heath, I.; Hubner, N.C.; et al. Whole-genome bisulfite sequencing of two distinct interconvertible DNA methylomes of mouse embryonic stem cells. Cell Stem Cell 2013, 13, 360–369. [Google Scholar] [CrossRef]
- Leitch, H.G.; McEwen, K.R.; Turp, A.; Encheva, V.; Carroll, T.; Grabole, N.; Mansfield, W.; Nashun, B.; Knezovich, J.G.; Smith, A.; et al. Naive pluripotency is associated with global DNA hypomethylation. Nat. Struct. Mol. Biol. 2013, 20, 311–316. [Google Scholar] [CrossRef]
- Boroviak, T.; Loos, R.; Lombard, P.; Okahara, J.; Behr, R.; Sasaki, E.; Nichols, J.; Smith, A.; Bertone, P. Lineage-Specific Profiling Delineates the Emergence and Progression of Naive Pluripotency in Mammalian Embryogenesis. Dev. Cell 2015, 35, 366–382. [Google Scholar] [CrossRef] [PubMed]
- Mulas, C.; Kalkan, T.; Smith, A. NODAL Secures Pluripotency upon Embryonic Stem Cell Progression from the Ground State. Stem Cell Rep. 2017, 9, 77–91. [Google Scholar] [CrossRef]
- Buecker, C.; Srinivasan, R.; Wu, Z.; Calo, E.; Acampora, D.; Faial, T.; Simeone, A.; Tan, M.; Swigut, T.; Wysocka, J. Reorganization of enhancer patterns in transition from naive to primed pluripotency. Cell Stem Cell 2014, 14, 838–853. [Google Scholar] [CrossRef] [PubMed]
- Choi, H.W.; Joo, J.Y.; Hong, Y.J.; Kim, J.S.; Song, H.; Lee, J.W.; Wu, G.; Schöler, H.R.; Do, J.T. Distinct Enhancer Activity of Oct4 in Naive and Primed Mouse Pluripotency. Stem Cell Rep. 2016, 7, 911–926. [Google Scholar] [CrossRef]
- Dunn, S.J.; Martello, G.; Yordanov, B.; Emmott, S.; Smith, A.G. Defining an essential transcription factor program for naïve pluripotency. Science 2014, 344, 1156–1160. [Google Scholar] [CrossRef]
- Fiorenzano, A.; Pascale, E.; D’Aniello, C.; Acampora, D.; Bassalert, C.; Russo, F.; Andolfi, G.; Biffoni, M.; Francescangeli, F.; Zeuner, A.; et al. Cripto is essential to capture mouse epiblast stem cell and human embryonic stem cell pluripotency. Nat. Commun. 2016, 7, 12589. [Google Scholar] [CrossRef]
- Kalkan, T.; Olova, N.; Roode, M.; Mulas, C.; Lee, H.J.; Nett, I.; Marks, H.; Walker, R.; Stunnenberg, H.G.; Lilley, K.S.; et al. Tracking the embryonic stem cell transition from ground state pluripotency. Development 2017, 144, 1221–1234. [Google Scholar] [CrossRef]
- Du, P.; Pirouz, M.; Choi, J.; Huebner, A.J.; Clement, K.; Meissner, A.; Hochedlinger, K.; Gregory, R.I. An Intermediate Pluripotent State Controlled by MicroRNAs Is Required for the Naive-to-Primed Stem Cell Transition. Cell Stem Cell 2018, 22, 851–864. [Google Scholar] [CrossRef]
- Neagu, A.; van Genderen, E.; Escudero, I.; Verwegen, L.; Kurek, D.; Lehmann, J.; Stel, J.; Dirks, R.A.M.; van Mierlo, G.; Maas, A.; et al. In vitro capture and characterization of embryonic rosette-stage pluripotency between naive and primed states. Nat. Cell Biol. 2020, 22, 534–545. [Google Scholar] [CrossRef]
- Smith, A. Formative pluripotency: The executive phase in a developmental continuum. Development 2017, 144, 365–373. [Google Scholar] [CrossRef]
- Kinoshita, M.; Smith, A. Pluripotency Deconstructed. Dev. Growth Differ. 2018, 60, 44–52. [Google Scholar] [CrossRef]
- Wang, X.; Xiang, Y.; Yu, Y.; Wang, R.; Zhang, Y.; Xu, Q.; Sun, H.; Zhao, Z.A.; Jiang, X.; Wang, X.; et al. Formative pluripotent stem cells show features of epiblast cells poised for gastrulation. Cell Res. 2021, 31, 526–541. [Google Scholar] [CrossRef]
- Kinoshita, M.; Barber, M.; Mansfield, W.; Cui, Y.; Spindlow, D.; Stirparo, G.G.; Dietmann, S.; Nichols, J.; Smith, A. Capture of Mouse and Human Stem Cells with Features of Formative Pluripotency. Cell Stem Cell 2021, 28, 453–471. [Google Scholar] [CrossRef]
- Guo, G.; Yang, J.; Nichols, J.; Hall, J.S.; Eyres, I.; Mansfield, W.; Smith, A. Klf4 reverts developmentally programmed restriction of ground state pluripotency. Development 2009, 136, 1063–1069. [Google Scholar] [CrossRef]
- Huang, Y.; Osorno, R.; Tsakiridis, A.; Wilson, V. In Vivo differentiation potential of epiblast stem cells revealed by chimeric embryo formation. Cell Rep. 2012, 2, 1571–1578. [Google Scholar] [CrossRef]
- O’Leary, T.; Heindryckx, B.; Lierman, S.; van Bruggen, D.; Goeman, J.J.; Vandewoestyne, M.; Deforce, D.; de Sousa Lopes, S.M.; De Sutter, P. Tracking the progression of the human inner cell mass during embryonic stem cell derivation. Nat. Biotechnol. 2012, 30, 278–282. [Google Scholar] [CrossRef]
- O’Leary, T.; Heindryckx, B.; Lierman, S.; Van der Jeught, M.; Duggal, G.; De Sutter, P.; Chuva de Sousa Lopes, S.M. Derivation of human embryonic stem cells using a post-inner cell mass intermediate. Nat. Protoc. 2013, 8, 254–264. [Google Scholar] [CrossRef]
- De Los Angeles, A.; Ferrari, F.; Xi, R.; Fujiwara, Y.; Benvenisty, N.; Deng, H.; Hochedlinger, K.; Jaenisch, R.; Lee, S.; Leitch, H.G.; et al. Hallmarks of pluripotency. Nature 2015, 525, 469–478. [Google Scholar] [CrossRef]
- De Los Angeles, A. The Pluripotency Continuum and Interspecies Chimeras. Curr. Protoc. Stem Cell Biol. 2019, 50, e87. [Google Scholar] [CrossRef]
- Amit, M.; Carpenter, M.K.; Inokuma, M.S.; Chiu, C.P.; Harris, C.P.; Waknitz, M.A.; Itskovitz-Eldor, J.; Thomson, J.A. Clonally derived human embryonic stem cell lines maintain pluripotency and proliferative potential for prolonged periods of culture. Dev. Biol. 2000, 227, 271–278. [Google Scholar] [CrossRef] [PubMed]
- Vallier, L.; Alexander, M.; Pedersen, R.A. Activin/Nodal and FGF pathways cooperate to maintain pluripotency of human embryonic stem cells. J. Cell Sci. 2005, 118, 4495–4509. [Google Scholar] [CrossRef] [PubMed]
- Hanna, J.; Markoulaki, S.; Mitalipova, M.; Cheng, A.W.; Cassady, J.P.; Staerk, J.; Carey, B.W.; Lengner, C.J.; Foreman, R.; Love, J.; et al. Metastable pluripotent states in NOD-mouse-derived ESCs. Cell Stem Cell 2009, 4, 513–524. [Google Scholar] [CrossRef]
- Hanna, J.; Cheng, A.W.; Saha, K.; Kim, J.; Lengner, C.J.; Soldner, F.; Cassady, J.P.; Muffat, J.; Carey, B.W.; Jaenisch, R. Human embryonic stem cells with biological and epigenetic characteristics similar to those of mouse ESCs. Proc. Natl. Acad. Sci. USA 2010, 107, 9222–9227. [Google Scholar] [CrossRef]
- Takashima, Y.; Guo, G.; Loos, R.; Nichols, J.; Ficz, G.; Krueger, F.; Oxley, D.; Santos, F.; Clarke, J.; Mansfield, W.; et al. Resetting transcription factor control circuitry toward ground-state pluripotency in human. Cell 2014, 158, 1254–1269. [Google Scholar] [CrossRef] [PubMed]
- Theunissen, T.W.; Powell, B.E.; Wang, H.; Mitalipova, M.; Faddah, D.A.; Reddy, J.; Fan, Z.P.; Maetzel, D.; Ganz, K.; Shi, L.; et al. Systematic identification of culture conditions for induction and maintenance of naive human pluripotency. Cell Stem Cell 2014, 15, 471–487. [Google Scholar] [CrossRef]
- Chan, Y.S.; Göke, J.; Ng, J.H.; Lu, X.; Gonzales, K.A.; Tan, C.P.; Tng, W.Q.; Hong, Z.Z.; Lim, Y.S.; Ng, H.H. Induction of a human pluripotent state with distinct regulatory circuitry that resembles preimplantation epiblast. Cell Stem Cell 2013, 13, 663–675. [Google Scholar] [CrossRef]
- Duggal, G.; Warrier, S.; Ghimire, S.; Broekaert, D.; Van der Jeught, M.; Lierman, S.; Deroo, T.; Peelman, L.; Van Soom, A.; Cornelissen, R.; et al. Alternative Routes to Induce Naïve Pluripotency in Human Embryonic Stem Cells. Stem Cells 2015, 33, 2686–2698. [Google Scholar] [CrossRef] [PubMed]
- Gafni, O.; Weinberger, L.; Mansour, A.A.; Manor, Y.S.; Chomsky, E.; Ben-Yosef, D.; Kalma, Y.; Viukov, S.; Maza, I.; Zviran, A.; et al. Derivation of novel human ground state naive pluripotent stem cells. Nature 2013, 504, 282–286. [Google Scholar] [CrossRef] [PubMed]
- Ware, C.B.; Nelson, A.M.; Mecham, B.; Hesson, J.; Zhou, W.; Jonlin, E.C.; Jimenez-Caliani, A.J.; Deng, X.; Cavanaugh, C.; Cook, S.; et al. Derivation of naive human embryonic stem cells. Proc. Natl. Acad. Sci. USA 2014, 111, 4484–4489. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Ryan, D.J.; Wang, W.; Tsang, J.C.; Lan, G.; Masaki, H.; Gao, X.; Antunes, L.; Yu, Y.; Zhu, Z.; et al. Establishment of mouse expanded potential stem cells. Nature 2017, 550, 393–397. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Ryan, D.J.; Lan, G.; Zou, X.; Liu, P. In vitro establishment of expanded-potential stem cells from mouse pre-implantation embryos or embryonic stem cells. Nat. Protoc. 2019, 14, 350–378. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.; Nowak-Imialek, M.; Chen, X.; Chen, D.; Herrmann, D.; Ruan, D.; Chen, A.C.H.; Eckersley-Maslin, M.A.; Ahmad, S.; Lee, Y.L.; et al. Establishment of porcine and human expanded potential stem cells. Nat. Cell Biol. 2019, 21, 687–699. [Google Scholar] [CrossRef]
- Yeo, J.C.; Ng, H.H. The transcriptional regulation of pluripotency. Cell Res. 2013, 23, 20–32. [Google Scholar] [CrossRef]
- Morgani, S.; Nichols, J.; Hadjantonakis, A.K. The many faces of Pluripotency: In vitro adaptations of a continuum of in vivo states. BMC Dev. Biol. 2017, 17, 7. [Google Scholar] [CrossRef] [PubMed]
- Festuccia, N.; Owens, N.; Navarro, P. Esrrb, an estrogen-related receptor involved in early development, pluripotency, and reprogramming. FEBS Lett. 2018, 592, 852–877. [Google Scholar] [CrossRef]
- Messmer, T.; von Meyenn, F.; Savino, A.; Santos, F.; Mohammed, H.; Lun, A.T.L.; Marioni, J.C.; Reik, W. Transcriptional Heterogeneity in Naive and Primed Human Pluripotent Stem Cells at Single-Cell Resolution. Cell Rep. 2019, 26, 815–824. [Google Scholar] [CrossRef]
- Kojima, Y.; Tam, O.H.; Tam, P.P. Timing of developmental events in the early mouse embryo. Semin. Cell Dev. Biol. 2014, 34, 65–75. [Google Scholar] [CrossRef]
- Tsakiridis, A.; Huang, Y.; Blin, G.; Skylaki, S.; Wymeersch, F.; Osorno, R.; Economou, C.; Karagianni, E.; Zhao, S.; Lowell, S.; et al. Distinct Wnt-driven primitive streak-like populations reflect in vivo lineage precursors. Development 2014, 141, 1209–1221. [Google Scholar] [CrossRef] [PubMed]
- Jouneau, A. Heterogeneity in Epiblast Stem Cells. Adv. Exp. Med. Biol. 2019, 1123, 5–17. [Google Scholar] [CrossRef]
- Sim, Y.J.; Kim, M.S.; Nayfeh, A.; Yun, Y.J.; Kim, S.J.; Park, K.T.; Kim, C.H.; Kim, K.S. 2i Maintains a Naive Ground State in ESCs through Two Distinct Epigenetic Mechanisms. Stem Cell Rep. 2017, 8, 1312–1328. [Google Scholar] [CrossRef] [PubMed]
- Shanak, S.; Helms, V. DNA methylation and the core pluripotency network. Dev. Biol. 2020, 464, 145–160. [Google Scholar] [CrossRef] [PubMed]
- Betto, R.M.; Diamante, L.; Perrera, V.; Audano, M.; Rapelli, S.; Lauria, A.; Incarnato, D.; Arboit, M.; Pedretti, S.; Rigoni, G.; et al. Metabolic control of DNA methylation in naive pluripotent cells. Nat. Genet. 2021, 53, 215–229. [Google Scholar] [CrossRef]
- Ficz, G.; Hore, T.A.; Santos, F.; Lee, H.J.; Dean, W.; Arand, J.; Krueger, F.; Oxley, D.; Paul, Y.L.; Walter, J.; et al. FGF signaling inhibition in ESCs drives rapid genome-wide demethylation to the epigenetic ground state of pluripotency. Cell Stem Cell 2013, 13, 351–359. [Google Scholar] [CrossRef] [PubMed]
- Azuara, V.; Perry, P.; Sauer, S.; Spivakov, M.; Jørgensen, H.F.; John, R.M.; Gouti, M.; Casanova, M.; Warnes, G.; Merkenschlager, M.; et al. Chromatin signatures of pluripotent cell lines. Nat. Cell Biol. 2006, 8, 532–538. [Google Scholar] [CrossRef]
- Bernstein, B.E.; Mikkelsen, T.S.; Xie, X.; Kamal, M.; Huebert, D.J.; Cuff, J.; Fry, B.; Meissner, A.; Wernig, M.; Plath, K.; et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 2006, 125, 315–326. [Google Scholar] [CrossRef]
- Langerman, J.; Lopez, D.; Pellegrini, M.; Smale, S.T. Species-Specific Relationships between DNA and Chromatin Properties of CpG Islands in Embryonic Stem Cells and Differentiated Cells. Stem Cell Rep. 2021, 16, 899–912. [Google Scholar] [CrossRef]
- Song, J.; Saha, S.; Gokulrangan, G.; Tesar, P.J.; Ewing, R.M. DNA and chromatin modification networks distinguish stem cell pluripotent ground states. Mol. Cell Proteom. 2012, 11, 1036–1047. [Google Scholar] [CrossRef] [PubMed]
- Factor, D.C.; Corradin, O.; Zentner, G.E.; Saiakhova, A.; Song, L.; Chenoweth, J.G.; McKay, R.D.; Crawford, G.E.; Scacheri, P.C.; Tesar, P.J. Epigenomic comparison reveals activation of "seed" enhancers during transition from naive to primed pluripotency. Cell Stem Cell 2014, 14, 854–863. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Ratanasirintrawoot, S.; Chandrasekaran, S.; Wu, Z.; Ficarro, S.B.; Yu, C.; Ross, C.A.; Cacchiarelli, D.; Xia, Q.; Seligson, M.; et al. LIN28 Regulates Stem Cell Metabolism and Conversion to Primed Pluripotency. Cell Stem Cell 2016, 19, 66–80. [Google Scholar] [CrossRef]
- Xiao, S.; Lu, J.; Sridhar, B.; Cao, X.; Yu, P.; Zhao, T.; Chen, C.C.; McDee, D.; Sloofman, L.; Wang, Y.; et al. SMARCAD1 Contributes to the Regulation of Naive Pluripotency by Interacting with Histone Citrullination. Cell Rep. 2017, 18, 3117–3128. [Google Scholar] [CrossRef]
- Geng, T.; Zhang, D.; Jiang, W. Epigenetic Regulation of Transition among Different Pluripotent States: Concise Review. Stem Cells 2019, 37, 1372–1380. [Google Scholar] [CrossRef] [PubMed]
- Pastor, W.A.; Chen, D.; Liu, W.; Kim, R.; Sahakyan, A.; Lukianchikov, A.; Plath, K.; Jacobsen, S.E.; Clark, A.T. Naive Human Pluripotent Cells Feature a Methylation Landscape Devoid of Blastocyst or Germline Memory. Cell Stem Cell 2016, 18, 323–329. [Google Scholar] [CrossRef]
- Silva, J.; Nichols, J.; Theunissen, T.W.; Guo, G.; van Oosten, A.L.; Barrandon, O.; Wray, J.; Yamanaka, S.; Chambers, I.; Smith, A. Nanog is the gateway to the pluripotent ground state. Cell 2009, 138, 722–737. [Google Scholar] [CrossRef]
- Young, R.A. Control of the embryonic stem cell state. Cell 2011, 144, 940–954. [Google Scholar] [CrossRef]
- Silva, J.; Barrandon, O.; Nichols, J.; Kawaguchi, J.; Theunissen, T.W.; Smith, A. Promotion of reprogramming to ground state pluripotency by signal inhibition. PLoS Biol. 2008, 6, e253. [Google Scholar] [CrossRef] [PubMed]
- Ware, C.B. Concise Review: Lessons from Naïve Human Pluripotent Cells. Stem Cells 2017, 35, 35–41. [Google Scholar] [CrossRef] [PubMed]
- Collier, A.J.; Panula, S.P.; Schell, J.P.; Chovanec, P.; Plaza Reyes, A.; Petropoulos, S.; Corcoran, A.E.; Walker, R.; Douagi, I.; Lanner, F.; et al. Comprehensive Cell Surface Protein Profiling Identifies Specific Markers of Human Naive and Primed Pluripotent States. Cell Stem Cell 2017, 20, 874–890. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Zhuang, L.; Lin, C.P. Roles of MicroRNAs in Establishing and Modulating Stem Cell Potential. Int. J. Mol. Sci. 2019, 20, 3643. [Google Scholar] [CrossRef] [PubMed]
- Fico, A.; Fiorenzano, A.; Pascale, E.; Patriarca, E.J.; Minchiotti, G. Long non-coding RNA in stem cell pluripotency and lineage commitment: Functions and evolutionary conservation. Cell. Mol. Life Sci. 2019, 76, 1459–1471. [Google Scholar] [CrossRef]
- Jouneau, A.; Ciaudo, C.; Sismeiro, O.; Brochard, V.; Jouneau, L.; Vandormael-Pournin, S.; Coppée, J.Y.; Zhou, Q.; Heard, E.; Antoniewski, C.; et al. Naive and primed murine pluripotent stem cells have distinct miRNA expression profiles. RNA 2012, 18, 253–264. [Google Scholar] [CrossRef]
- Dodsworth, B.T.; Hatje, K.; Rostovskaya, M.; Flynn, R.; Meyer, C.A.; Cowley, S.A. Profiling of naïve and primed human pluripotent stem cells reveals state-associated miRNAs. Sci. Rep. 2020, 10, 10542. [Google Scholar] [CrossRef]
- Rosa, A.; Brivanlou, A.H. MicroRNAs in early vertebrate development. Cell Cycle 2009, 8, 3513–3520. [Google Scholar] [CrossRef]
- Landgraf, P. UCB transplantation: miRNA involvement. Blood 2009, 113, 6505–6506. [Google Scholar] [CrossRef] [PubMed]
- Lipchina, I.; Elkabetz, Y.; Hafner, M.; Sheridan, R.; Mihailovic, A.; Tuschl, T.; Sander, C.; Studer, L.; Betel, D. Genome-wide identification of microRNA targets in human ES cells reveals a role for miR-302 in modulating BMP response. Genes Dev. 2011, 25, 2173–2186. [Google Scholar] [CrossRef] [PubMed]
- Calabrese, J.M.; Seila, A.C.; Yeo, G.W.; Sharp, P.A. RNA sequence analysis defines Dicer’s role in mouse embryonic stem cells. Proc. Natl. Acad. Sci. USA 2007, 104, 18097–18102. [Google Scholar] [CrossRef]
- Chen, T.; Hao, Y.J.; Zhang, Y.; Li, M.M.; Wang, M.; Han, W.; Wu, Y.; Lv, Y.; Hao, J.; Wang, L.; et al. m6A RNA methylation is regulated by microRNAs and promotes reprogramming to pluripotency. Cell Stem Cell 2015, 16, 289–301. [Google Scholar] [CrossRef]
- Marson, A.; Levine, S.S.; Cole, M.F.; Frampton, G.M.; Brambrink, T.; Johnstone, S.; Guenther, M.G.; Johnston, W.K.; Wernig, M.; Newman, J.; et al. Connecting microRNA genes to the core transcriptional regulatory circuitry of embryonic stem cells. Cell 2008, 134, 521–533. [Google Scholar] [CrossRef]
- Wang, Y.; Blelloch, R. Cell cycle regulation by MicroRNAs in embryonic stem cells. Cancer Res. 2009, 69, 4093–4096. [Google Scholar] [CrossRef] [PubMed]
- Dolezalova, D.; Mraz, M.; Barta, T.; Plevova, K.; Vinarsky, V.; Holubcova, Z.; Jaros, J.; Dvorak, P.; Pospisilova, S.; Hampl, A. MicroRNAs regulate p21(Waf1/Cip1) protein expression and the DNA damage response in human embryonic stem cells. Stem Cells 2012, 30, 1362–1372. [Google Scholar] [CrossRef]
- Card, D.A.; Hebbar, P.B.; Li, L.; Trotter, K.W.; Komatsu, Y.; Mishina, Y.; Archer, T.K. Oct4/Sox2-regulated miR-302 targets cyclin D1 in human embryonic stem cells. Mol. Cell. Biol. 2008, 28, 6426–6438. [Google Scholar] [CrossRef]
- Rodríguez-Varela, C.; Herraiz, S.; Labarta, E. Mitochondrial enrichment in infertile patients: A review of different mitochondrial replacement therapies. Ther. Adv. Reprod. Health 2021, 15, 26334941211023544. [Google Scholar] [CrossRef]
- Jin, S.; Collin, J.; Zhu, L.; Montaner, D.; Armstrong, L.; Neganova, I.; Lako, M. A Novel Role for miR-1305 in Regulation of Pluripotency-Differentiation Balance, Cell Cycle, and Apoptosis in Human Pluripotent Stem Cells. Stem Cells 2016, 34, 2306–2317. [Google Scholar] [CrossRef]
- Mallanna, S.K.; Rizzino, A. Emerging roles of microRNAs in the control of embryonic stem cells and the generation of induced pluripotent stem cells. Dev. Biol. 2010, 344, 16–25. [Google Scholar] [CrossRef]
- Li, N.; Long, B.; Han, W.; Yuan, S.; Wang, K. microRNAs: Important regulators of stem cells. Stem Cell Res. Ther. 2017, 8, 110. [Google Scholar] [CrossRef]
- Stadler, B.; Ivanovska, I.; Mehta, K.; Song, S.; Nelson, A.; Tan, Y.; Mathieu, J.; Darby, C.; Blau, C.A.; Ware, C.; et al. Characterization of microRNAs involved in embryonic stem cell states. Stem Cells Dev. 2010, 19, 935–950. [Google Scholar] [CrossRef]
- Salazar-Roa, M.; Trakala, M.; Álvarez-Fernández, M.; Valdés-Mora, F.; Zhong, C.; Muñoz, J.; Yu, Y.; Peters, T.J.; Graña-Castro, O.; Serrano, R.; et al. Transient exposure to miR-203 enhances the differentiation capacity of established pluripotent stem cells. EMBO J. 2020, 39, e104324. [Google Scholar] [CrossRef] [PubMed]
- Dinger, M.E.; Amaral, P.P.; Mercer, T.R.; Pang, K.C.; Bruce, S.J.; Gardiner, B.B.; Askarian-Amiri, M.E.; Ru, K.; Soldà, G.; Simons, C.; et al. Long noncoding RNAs in mouse embryonic stem cell pluripotency and differentiation. Genome Res. 2008, 18, 1433–1445. [Google Scholar] [CrossRef]
- Guttman, M.; Amit, I.; Garber, M.; French, C.; Lin, M.F.; Feldser, D.; Huarte, M.; Zuk, O.; Carey, B.W.; Cassady, J.P.; et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 2009, 458, 223–227. [Google Scholar] [CrossRef] [PubMed]
- Guttman, M.; Donaghey, J.; Carey, B.W.; Garber, M.; Grenier, J.K.; Munson, G.; Young, G.; Lucas, A.B.; Ach, R.; Bruhn, L.; et al. LincRNAs act in the circuitry controlling pluripotency and differentiation. Nature 2011, 477, 295–300. [Google Scholar] [CrossRef]
- Huo, J.S.; Zambidis, E.T. Pivots of pluripotency: The roles of non-coding RNA in regulating embryonic and induced pluripotent stem cells. Biochim. Biophys. Acta 2013, 1830, 2385–2394. [Google Scholar] [CrossRef]
- Sheik Mohamed, J.; Gaughwin, P.M.; Lim, B.; Robson, P.; Lipovich, L. Conserved long noncoding RNAs transcriptionally regulated by Oct4 and Nanog modulate pluripotency in mouse embryonic stem cells. RNA 2010, 16, 324–337. [Google Scholar] [CrossRef]
- Lin, N.; Chang, K.Y.; Li, Z.; Gates, K.; Rana, Z.A.; Dang, J.; Zhang, D.; Han, T.; Yang, C.S.; Cunningham, T.J.; et al. An evolutionarily conserved long noncoding RNA TUNA controls pluripotency and neural lineage commitment. Mol. Cell 2014, 53, 1005–1019. [Google Scholar] [CrossRef]
- Xu, N.; Papagiannakopoulos, T.; Pan, G.; Thomson, J.A.; Kosik, K.S. MicroRNA-145 regulates OCT4, SOX2, and KLF4 and represses pluripotency in human embryonic stem cells. Cell 2009, 137, 647–658. [Google Scholar] [CrossRef] [PubMed]
- Ng, S.Y.; Johnson, R.; Stanton, L.W. Human long non-coding RNAs promote pluripotency and neuronal differentiation by association with chromatin modifiers and transcription factors. EMBO J. 2012, 31, 522–533. [Google Scholar] [CrossRef]
- Huarte, M.; Guttman, M.; Feldser, D.; Garber, M.; Koziol, M.J.; Kenzelmann-Broz, D.; Khalil, A.M.; Zuk, O.; Amit, I.; Rabani, M.; et al. A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell 2010, 142, 409–419. [Google Scholar] [CrossRef] [PubMed]
- Bao, M.H.; Li, J.M.; Zhou, Q.L.; Li, G.Y.; Zeng, J.; Zhao, J.; Zhang, Y.W. Effects of miR-590 on oxLDL-induced endothelial cell apoptosis: Roles of p53 and NF-κB. Mol. Med. Rep. 2016, 13, 867–873. [Google Scholar] [CrossRef]
- Tsai, M.C.; Manor, O.; Wan, Y.; Mosammaparast, N.; Wang, J.K.; Lan, F.; Shi, Y.; Segal, E.; Chang, H.Y. Long noncoding RNA as modular scaffold of histone modification complexes. Science 2010, 329, 689–693. [Google Scholar] [CrossRef]
- Kaneko, S.; Bonasio, R.; Saldaña-Meyer, R.; Yoshida, T.; Son, J.; Nishino, K.; Umezawa, A.; Reinberg, D. Interactions between JARID2 and noncoding RNAs regulate PRC2 recruitment to chromatin. Mol. Cell 2014, 53, 290–300. [Google Scholar] [CrossRef] [PubMed]
- Dey, B.K.; Mueller, A.C.; Dutta, A. Long non-coding RNAs as emerging regulators of differentiation, development, and disease. Transcription 2014, 5, e944014. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rebuzzini, P.; Zuccotti, M.; Garagna, S. Building Pluripotency Identity in the Early Embryo and Derived Stem Cells. Cells 2021, 10, 2049. https://doi.org/10.3390/cells10082049
Rebuzzini P, Zuccotti M, Garagna S. Building Pluripotency Identity in the Early Embryo and Derived Stem Cells. Cells. 2021; 10(8):2049. https://doi.org/10.3390/cells10082049
Chicago/Turabian StyleRebuzzini, Paola, Maurizio Zuccotti, and Silvia Garagna. 2021. "Building Pluripotency Identity in the Early Embryo and Derived Stem Cells" Cells 10, no. 8: 2049. https://doi.org/10.3390/cells10082049
APA StyleRebuzzini, P., Zuccotti, M., & Garagna, S. (2021). Building Pluripotency Identity in the Early Embryo and Derived Stem Cells. Cells, 10(8), 2049. https://doi.org/10.3390/cells10082049







