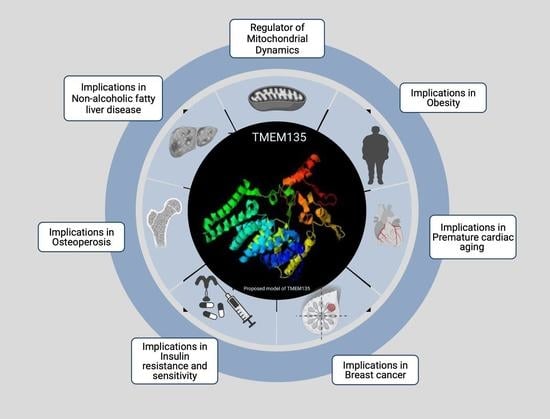
TMEM135 is a Novel Regulator of Mitochondrial Dynamics and Physiology with Implications for Human Health Conditions
Abstract
1. The Structure and Function of Transmembrane Proteins
2. The Discovery of Transmembrane Protein 135 (TMEM135)
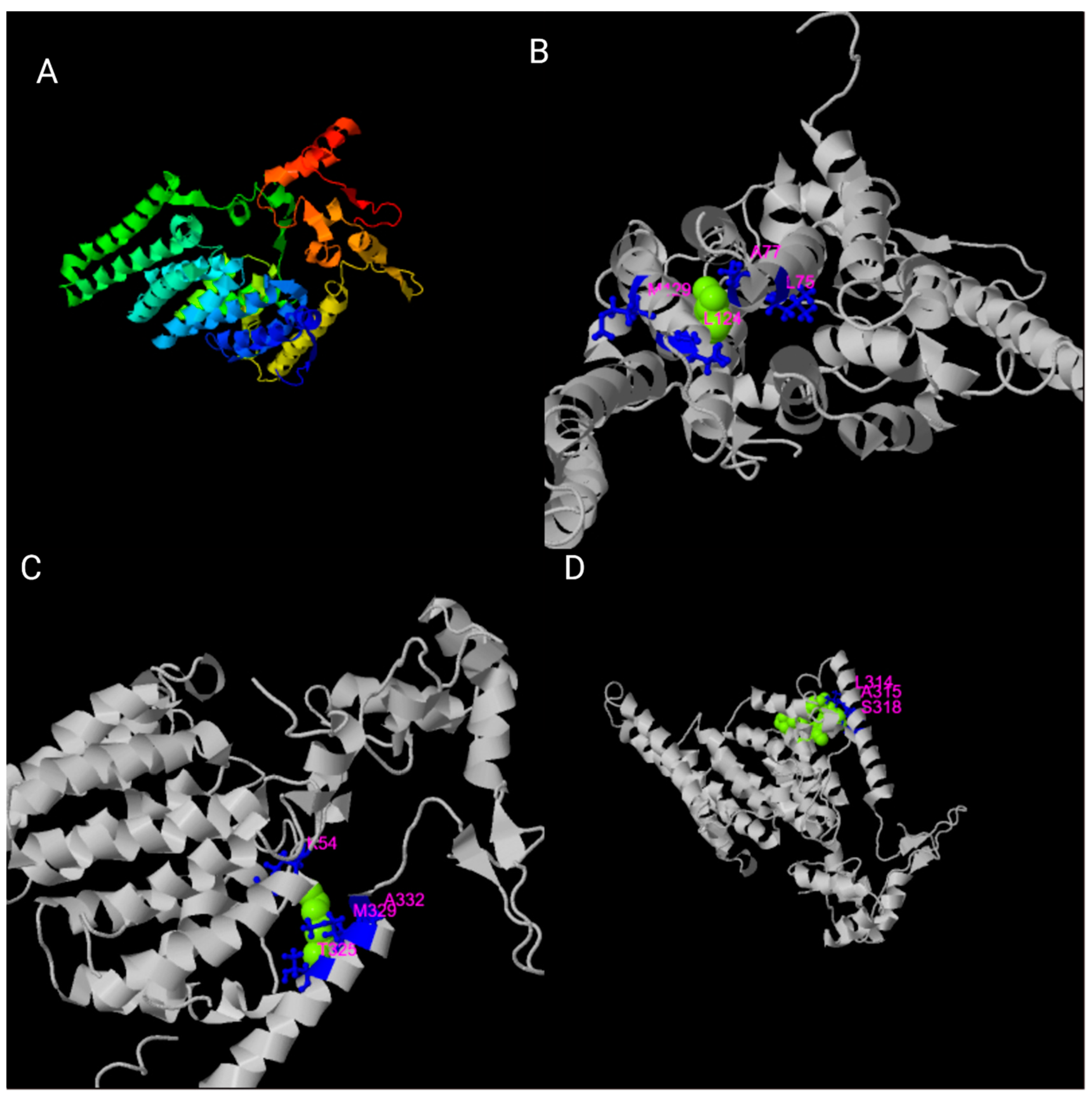
3. Structural Organization of the TMEM135 Gene and Protein
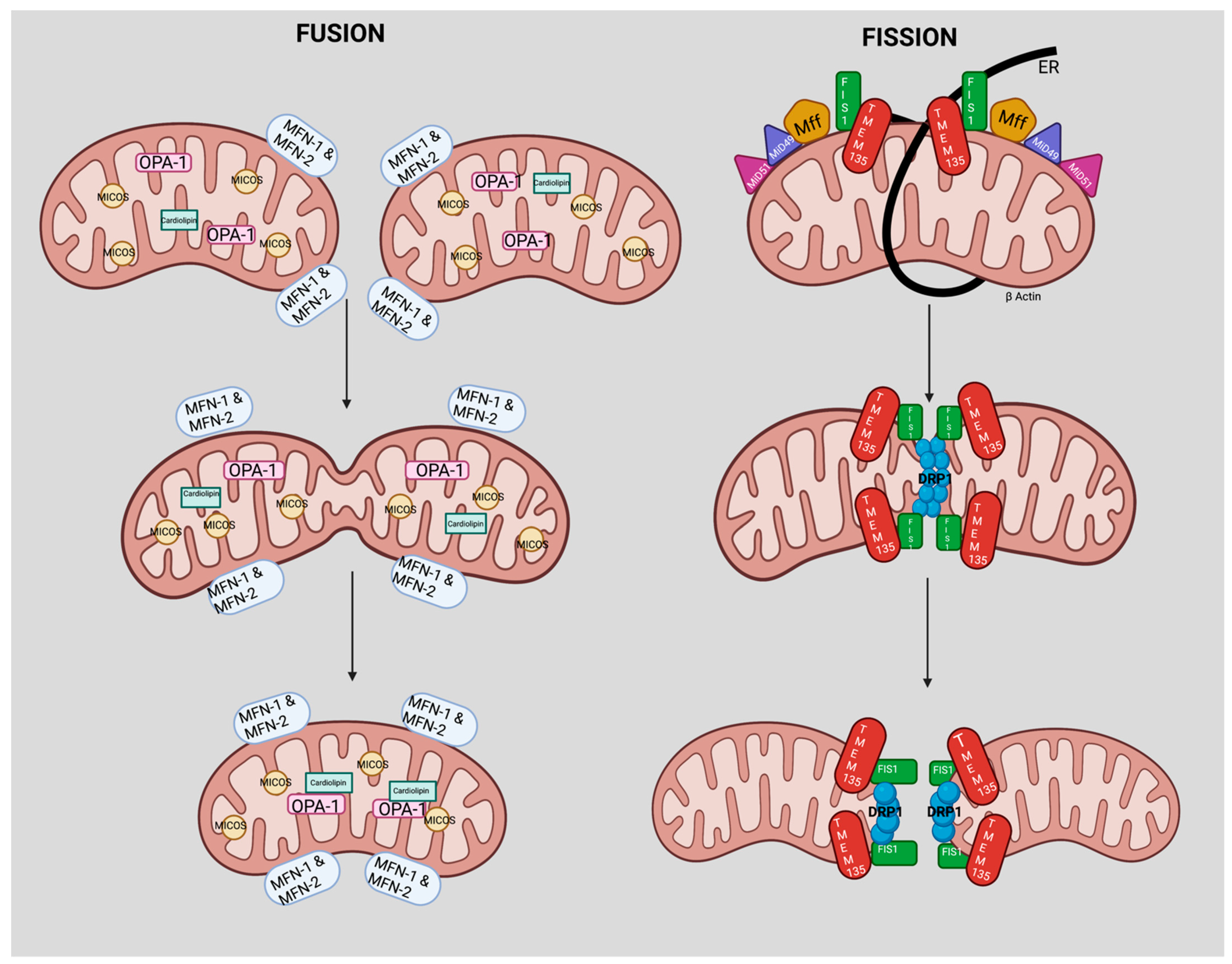
4. TMEM135 is a Regulator of Mitochondrial Dynamics
5. TMEM135 and Peroxisomal Transport
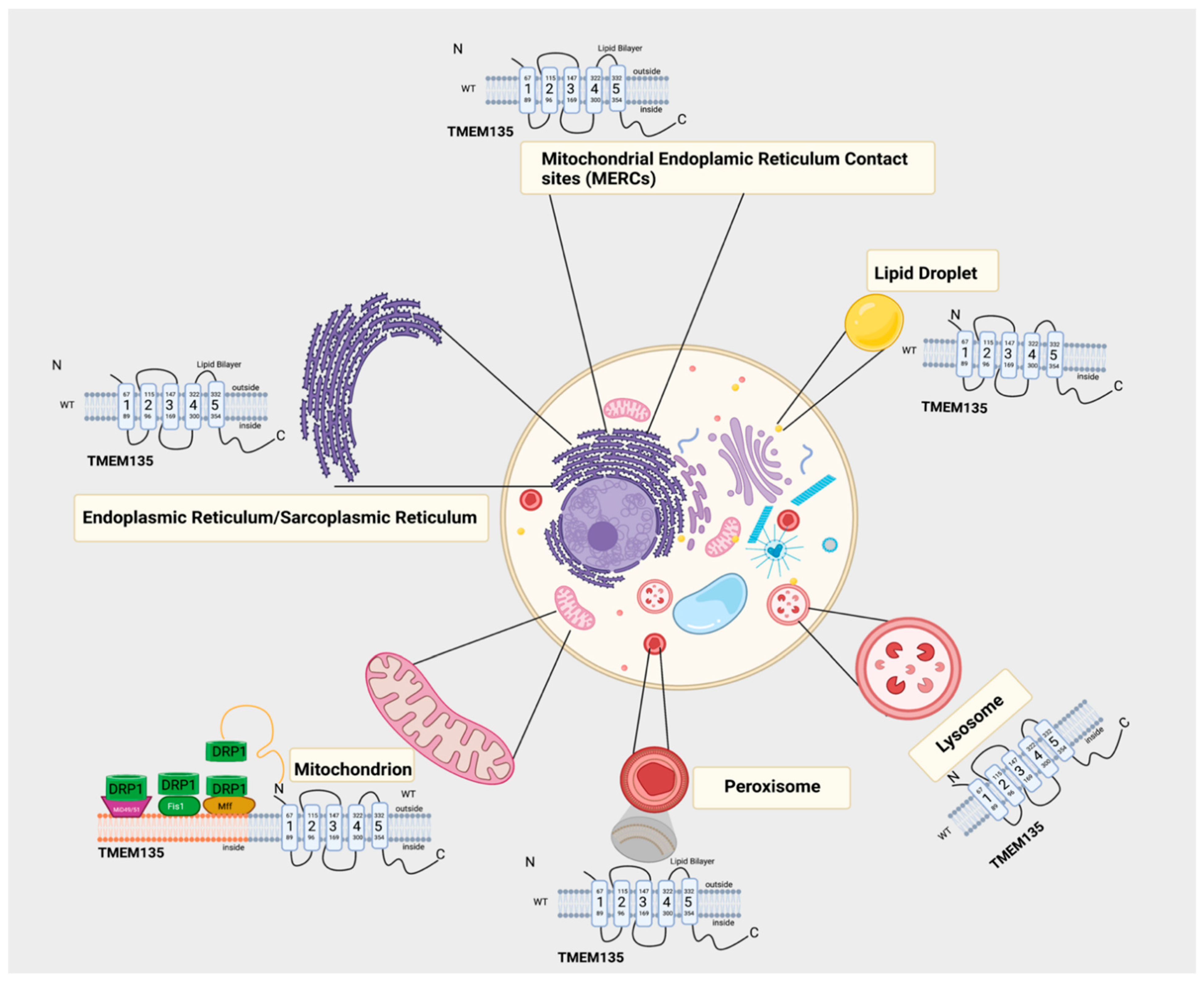
6. Potential Physiological Roles of TMEM135
7. Potential Role of TMEM135 as a Regulator of Calcium Dynamics
8. General Characteristics and Profiling of TMEM135 in Human Diseases
9. Perspective
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Marx, S.; Maso, T.D.; Chen, J.-W.; Bury, M.; Wouters, J.; Michiels, C.; Le Calvé, B. Transmembrane (TMEM) protein family members: Poorly characterized even if essential for the metastatic process. Semin. Cancer Biol. 2020, 60, 96–106. [Google Scholar] [CrossRef]
- Fuller, C.M. Time for TMEM? J. Physiol. 2012, 590, 5931–5932. [Google Scholar] [CrossRef]
- Vinothkumar, K.R.; Henderson, R. Structures of membrane proteins. Q. Rev. Biophys. 2010, 43, 65–158. [Google Scholar] [CrossRef]
- Von Heijne, G. Membrane-protein topology. Nat. Rev. Mol. Cell Biol. 2006, 7, 909–918. [Google Scholar] [CrossRef]
- Alberts, B. (Ed.) Molecular Biology of the Cell, 5th ed.; Alberts, B. (Ed.) Garland Science: New York, NY, USA, 2008; ISBN 978-0-8153-4105-5. [Google Scholar]
- Ruiz, M.D.L.; Kraus, R.L. Voltage-Gated Sodium Channels: Structure, Function, Pharmacology, and Clinical Indications. J. Med. Chem. 2015, 58, 7093–7118. [Google Scholar] [CrossRef]
- Abbott, G.W. KCNQs: Ligand- and Voltage-Gated Potassium Channels. Front. Physiol. 2020, 11, 583. [Google Scholar] [CrossRef] [PubMed]
- Bouza, A.A.; Isom, L.L. Voltage-Gated Sodium Channel β Subunits and Their Related Diseases. In Handbook of Experimental Pharmacology; Springer Science and Business Media LLC: New York, NY, USA, 2017; Volume 246, pp. 423–450. [Google Scholar]
- Kaplan, D.I.; Isom, L.L.; Petrou, S. Role of Sodium Channels in Epilepsy. Cold Spring Harb. Perspect. Med. 2016, 6, a022814. [Google Scholar] [CrossRef]
- Kruger, L.C.; Isom, L.L. Voltage-Gated Na+Channels: Not Just for Conduction. Cold Spring Harb. Perspect. Biol. 2016, 8, a029264. [Google Scholar] [CrossRef]
- Exil, V.J.; Avila, D.S.; Benedetto, A.; Exil, E.A.; Adams, M.R.; Au, C.; Aschner, M. Stressed-Induced TMEM135 Protein Is Part of a Conserved Genetic Network Involved in Fat Storage and Longevity Regulation in C. elegans. PLoS ONE 2010, 5, e14228. [Google Scholar] [CrossRef]
- Lee, W.-H.; Higuchi, H.; Ikeda, S.; Macke, E.L.; Takimoto, T.; Pattnaik, B.; Liu, C.; Chu, L.-F.; Siepka, S.M.; Krentz, K.J.; et al. Mouse Tmem135 mutation reveals a mechanism involving mitochondrial dynamics that leads to age-dependent retinal pathologies. eLife 2016, 5. [Google Scholar] [CrossRef]
- Zaucha, J.; Heinzinger, M.; Kulandaisamy, A.; Kataka, E.; Salvádor Óscar, L.; Popov, P.; Rost, B.; Gromiha, M.M.; Zhorov, B.S.; Frishman, D. Mutations in transmembrane proteins: Diseases, evolutionary insights, prediction and comparison with globular proteins. Briefings Bioinform. 2021, 22. [Google Scholar] [CrossRef]
- Sánchez-Caballero, L.; Elurbe, D.M.; Baertling, F.; Guerrero-Castillo, S.; van der Brand, M.; van Strien, J.; van Dam, T.J.; Rodenburg, R.; Brandt, U.; Huynen, M.A.; et al. TMEM70 functions in the assembly of complexes I and V. Biochim. Biophys. Acta 2020, 1861, 148202. [Google Scholar] [CrossRef]
- Kovalčíková, J.; Vrbacký, M.; Pecina, P.; Tauchmannová, K.; Nůsková, H.; Kaplanová, V.; Brázdová, A.; Alán, L.; Eliáš, J.; Čunátová, K.; et al. TMEM70 facilitates biogenesis of mammalian ATP synthase by promoting subunit c incorporation into the rotor structure of the enzyme. FASEB J. 2019, 33, 14103–14117. [Google Scholar] [CrossRef] [PubMed]
- Bahri, H.; Buratto, J.; Rojo, M.; Dompierre, J.P.; Salin, B.; Blancard, C.; Cuvellier, S.; Rose, M.; Elgaaied, A.B.A.; Tetaud, E.; et al. TMEM70 forms oligomeric scaffolds within mitochondrial cristae promoting in situ assembly of mammalian ATP synthase proton channel. Biochim. Biophys. Acta (BBA)-Bioenerg. 2021, 1868, 118942. [Google Scholar] [CrossRef]
- Vrbacky, M.; Kovalčíková, J.; Chawengsaksophak, K.; Beck, I.M.; Mráček, T.; Nůsková, H.; Sedmera, D.; Papoušek, F.; Kolar, F.; Sobol, M.; et al. Knockout of Tmem70 alters biogenesis of ATP synthase and leads to embryonal lethality in mice. Hum. Mol. Genet. 2016, 25, 4674–4685. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kratochvílová, H.; Hejzlarová, K.; Vrbacky, M.; Mráček, T.; Karbanová, V.; Tesarova, M.; Gombitová, A.; Cmarko, D.; Wittig, I.; Zeman, J.; et al. Mitochondrial membrane assembly of TMEM70 protein. Mitochondrion 2014, 15, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Magner, M.; Dvorakova, V.; Tesarova, M.; Mazurova, S.; Hansikova, H.; Zahorec, M.; Brennerova, K.; Bzduch, V.; Spiegel, R.; Horovitz, Y.; et al. TMEM70 deficiency: Long-term outcome of 48 patients. J. Inherit. Metab. Dis. 2014, 38, 417–426. [Google Scholar] [CrossRef]
- Carroll, J.; He, J.; Ding, S.; Fearnley, I.M.; Walker, J.E. TMEM70 and TMEM242 help to assemble the rotor ring of human ATP synthase and interact with assembly factors for complex I. Proc. Natl. Acad. Sci. USA 2021, 118. [Google Scholar] [CrossRef] [PubMed]
- Mathur, A.; Sims, H.F.; Gopalakrishnan, D.; Gibson, B.; Rinaldo, P.; Vockley, J.; Hug, G.; Strauss, A.W. Molecular Heterogeneity in Very-Long-Chain Acyl-CoA Dehydrogenase Deficiency Causing Pediatric Cardiomyopathy and Sudden Death. Circulation 1999, 99, 1337–1343. [Google Scholar] [CrossRef]
- Aoyama, T.; Uchida, Y.; Kelley, R.; Marble, M.; Hofman, K.; Tonsgard, J.; Rhead, W.; Hashimoto, T. A Novel Disease with Deficiency of Mitochondrial Very-Long-Chain Acyl-CoA Dehydrogenase. Biochem. Biophys. Res. Commun. 1993, 191, 1369–1372. [Google Scholar] [CrossRef]
- Aoyama, T.; Souri, M.; Ueno, I.; Kamijo, T.; Yamaguchi, S.; Rhead, W.J.; Tanaka, K.; Hashimoto, T. Cloning of human very-long-chain acyl-coenzyme A dehydrogenase and molecular characterization of its deficiency in two patients. Am. J. Hum. Genet. 1995, 57, 273–283. [Google Scholar] [PubMed]
- Souri, M.; Aoyama, T.; Orii, K.; Yamaguchi, S.; Hashimoto, T. Mutation analysis of very-long-chain acyl-coenzyme A dehydrogenase (VLCAD) deficiency: Identification and characterization of mutant VLCAD cDNAs from four patients. Am. J. Hum. Genet. 1996, 58, 97–106. [Google Scholar] [PubMed]
- Bertrand, C.; Largillière, C.; Zabot, M.T.; Mathieu, M.; Vianey-Saban, C. Very long chain acyl-CoA dehydrogenase deficiency: Identification of a new inborn error of mitochondrial fatty acid oxidation in fibroblasts. Biochim. Biophys. Acta (BBA)-Mol. Basis Dis. 1993, 1180, 327–329. [Google Scholar] [CrossRef]
- Exil, V.J.; Roberts, R.L.; Sims, H.; McLaughlin, J.E.; Malkin, R.A.; Gardner, C.D.; Ni, G.; Rottman, J.N.; Strauss, A.W. Very-Long-Chain Acyl-Coenzyme A Dehydrogenase Deficiency in Mice. Circ. Res. 2003, 93, 448–455. [Google Scholar] [CrossRef] [PubMed]
- Scheideler, M.; Elabd, C.; Zaragosi, L.-E.; Chiellini, C.; Hackl, H.; Sanchez-Cabo, F.; Yadav, S.; Duszka, K.; Friedl, G.; Papak, C.; et al. Comparative transcriptomics of human multipotent stem cells during adipogenesis and osteoblastogenesis. BMC Genom. 2008, 9, 340. [Google Scholar] [CrossRef] [PubMed]
- Antebi, A.; Culotti, J.; Hedgecock, E. daf-12 regulates developmental age and the dauer alternative in C. elegans. Development 1998, 125, 1191–1205. [Google Scholar] [CrossRef] [PubMed]
- Gottlieb, S.; Ruvkun, G. daf-2, daf-16 and daf-23: Genetically interacting genes controlling Dauer formation in C. elegans. Genetics 1994, 137, 107–120. [Google Scholar] [CrossRef] [PubMed]
- Lin, K.; Dorman, J.B.; Rodan, A.; Kenyon, C. daf-16: An HNF-3/forkhead Family Member That Can Function to Double the Life-Span of C. elegans. Science 1997, 278, 1319–1322. [Google Scholar] [CrossRef]
- Ogg, S.; Paradis, S.; Gottlieb, S.; Patterson, G.I.; Lee, L.; Tissenbaum, H.A.; Ruvkun, G. The Fork head transcription factor DAF-16 transduces insulin-like metabolic and longevity signals in C. elegans. Nat. Cell Biol. 1997, 389, 994–999. [Google Scholar] [CrossRef]
- Paradis, S.; Ruvkun, G. C. elegans Akt/PKB transduces insulin receptor-like signals from AGE-1 PI3 kinase to the DAF-16 transcription factor. Genes Dev. 1998, 12, 2488–2498. [Google Scholar] [CrossRef]
- Lee, S.S.; Kennedy, S.; Tolonen, A.; Ruvkun, G. DAF-16 Target Genes That Control C. elegans Life-Span and Metabolism. Science 2003, 300, 644–647. [Google Scholar] [CrossRef]
- McElwee, J.; Bubb, K.; Thomas, J.H. Transcriptional outputs of the C. elegans forkhead protein DAF-16. Aging Cell 2003, 2, 111–121. [Google Scholar] [CrossRef]
- Kimura, K.; Tissenbaum, H.A.; Liu, Y.; Ruvkun, G. daf-2, an Insulin Receptor-Like Gene That Regulates Longevity and Diapause in C. elegans. Science 1997, 277, 942–946. [Google Scholar] [CrossRef] [PubMed]
- Madeira, F.; Park, Y.M.; Lee, J.; Buso, N.; Gur, T.; Madhusoodanan, N.; Basutkar, P.; Tivey, A.R.N.; Potter, S.C.; Finn, R.D.; et al. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res. 2019, 47, W636–W641. [Google Scholar] [CrossRef] [PubMed]
- Rosca, M.; Hoppel, C.L. Mitochondrial dysfunction in heart failure. Hear. Fail. Rev. 2013, 18, 607–622. [Google Scholar] [CrossRef]
- Liesa, M.; Palacín, M.; Zorzano, A. Mitochondrial Dynamics in Mammalian Health and Disease. Physiol. Rev. 2009, 89, 799–845. [Google Scholar] [CrossRef]
- Chen, H.; Chan, D.C. Physiological functions of mitochondrial fusion. Ann. N. Y. Acad. Sci. 2010, 1201, 21–25. [Google Scholar] [CrossRef] [PubMed]
- Naon, D.; Zaninello, M.; Giacomello, M.; Varanita, T.; Grespi, F.; Lakshminaranayan, S.; Serafini, A.; Semenzato, M.; Herkenne, S.; Hernández-Alvarez, M.I.; et al. Critical reappraisal confirms that Mitofusin 2 is an endoplasmic reticulum–mitochondria tether. Proc. Natl. Acad. Sci. USA 2016, 113, 11249–11254. [Google Scholar] [CrossRef]
- Muñoz, J.P.; Ivanova, S.; Wandelmer, J.S.; Martínez-Cristóbal, P.; Noguera, M.; Sancho, A.; Díaz-Ramos, A.; Hernández-Alvarez, M.I.; Sebastián, D.; Mauvezin, C.; et al. Mfn2 modulates the UPR and mitochondrial function via repression of PERK. EMBO J. 2013, 32, 2348–2361. [Google Scholar] [CrossRef]
- Sebastián, D.; Sorianello, E.; Segalés, J.; Irazoki, A.; Ruiz-Bonilla, V.; Sala, D.; Planet, E.; Berenguer-Llergo, A.; Muñoz, J.P.; Sánchez-Feutrie, M.; et al. Mfn2 deficiency links age-related sarcopenia and impaired autophagy to activation of an adaptive mitophagy pathway. EMBO J. 2016, 35, 1677–1693. [Google Scholar] [CrossRef]
- Favaro, G.; Romanello, V.; Varanita, T.; Desbats, M.A.; Morbidoni, V.; Tezze, C.; Albiero, M.; Canato, M.; Gherardi, G.; De Stefani, D.; et al. DRP1-mediated mitochondrial shape controls calcium homeostasis and muscle mass. Nat. Commun. 2019, 10, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Dulac, M.; Leduc-Gaudet, J.; Reynaud, O.; Ayoub, M.; Guérin, A.; Finkelchtein, M.; Na Hussain, S.; Gouspillou, G. Drp1 knockdown induces severe muscle atrophy and remodelling, mitochondrial dysfunction, autophagy impairment and denervation. J. Physiol. 2020, 598, 3691–3710. [Google Scholar] [CrossRef] [PubMed]
- Scott, I.; Youle, R.J. Mitochondrial fission and fusion. Essays Biochem. 2010, 47, 85–98. [Google Scholar] [CrossRef] [PubMed]
- Lee, W.-H.; Bhute, V.J.; Higuchi, H.; Ikeda, S.; Palecek, S.P.; Ikeda, A. Metabolic alterations caused by the mutation and overexpression of the Tmem135 gene. Exp. Biol. Med. 2020, 245, 1571–1583. [Google Scholar] [CrossRef]
- Pereira, R.O.; Tadinada, S.M.; Zasadny, F.M.; Oliveira, K.J.; Pires, K.M.P.; Olvera, A.; Jeffers, J.; Souvenir, R.; McGlauflin, R.; Seei, A.; et al. OPA 1 deficiency promotes secretion of FGF 21 from muscle that prevents obesity and insulin resistance. EMBO J. 2017, 36, 2126–2145. [Google Scholar] [CrossRef]
- Ren, L.; Chen, X.; Chen, X.; Li, J.; Cheng, B.; Xia, J. Mitochondrial Dynamics: Fission and Fusion in Fate Determination of Mesenchymal Stem Cells. Front. Cell Dev. Biol. 2020, 8, 580070. [Google Scholar] [CrossRef]
- Cipolat, S.; De Brito, O.M.; Zilio, B.D.; Scorrano, L. OPA1 requires mitofusin 1 to promote mitochondrial fusion. Proc. Natl. Acad. Sci. USA 2004, 101, 15927–15932. [Google Scholar] [CrossRef]
- Varanita, T.; Soriano, M.E.; Romanello, V.; Zaglia, T.; Quintana-Cabrera, R.; Semenzato, M.; Menabò, R.; Costa, V.; Civiletto, G.; Pesce, P.; et al. The Opa1-Dependent Mitochondrial Cristae Remodeling Pathway Controls Atrophic, Apoptotic, and Ischemic Tissue Damage. Cell Metab. 2015, 21, 834–844. [Google Scholar] [CrossRef] [PubMed]
- Dietrich, M.O.; Liu, Z.-W.; Horvath, T.L. Mitochondrial Dynamics Controlled by Mitofusins Regulate Agrp Neuronal Activity and Diet-Induced Obesity. Cell 2013, 155, 188–199. [Google Scholar] [CrossRef] [PubMed]
- Lewis, S.; Takimoto, T.; Mehrvar, S.; Higuchi, H.; Doebley, A.-L.; Stokes, G.; Sheibani, N.; Ikeda, S.; Ranji, M.; Ikeda, A. The effect of Tmem135 overexpression on the mouse heart. PLoS ONE 2018, 13, e0201986. [Google Scholar] [CrossRef]
- Faust, J.; Verma, A.; Peng, C.; McNew, J.A. An Inventory of Peroxisomal Proteins and Pathways in Drosophila melanogaster. Traffic 2012, 13, 1378–1392. [Google Scholar] [CrossRef]
- Cipolla, C.M.; Lodhi, I.J. Peroxisomal Dysfunction in Age-Related Diseases. Trends Endocrinol. Metab. 2017, 28, 297–308. [Google Scholar] [CrossRef] [PubMed]
- Renquist, B.J.; Madanayake, T.W.; Hennebold, J.D.; Ghimire, S.; Geisler, C.E.; Xu, Y.; Bogan, R.L. TMEM135 Is an LXR-Inducible Regulator of Peroxisomal Metabolism. Cell Biology 2018, 334979. [Google Scholar] [CrossRef]
- Teraoka, S.N.; Bernstein, J.L.; Reiner, A.S.; Haile, R.W.; Bernstein, L.; Lynch, C.F.; Malone, K.E.; Stovall, M.; Capanu, M.; Liang, X.; et al. Single nucleotide polymorphisms associated with risk for contralateral breast cancer in the Women’s Environment, Cancer, and Radiation Epidemiology (WECARE) Study. Breast Cancer Res. 2011, 13, R114. [Google Scholar] [CrossRef]
- Elbein, S.C.; Kern, P.A.; Rasouli, N.; Yao-Borengasser, A.; Sharma, N.K.; Das, S.K. Global Gene Expression Profiles of Subcutaneous Adipose and Muscle From Glucose-Tolerant, Insulin-Sensitive, and Insulin-Resistant Individuals Matched for BMI. Diabetes 2011, 60, 1019–1029. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Li, W.-D.; Glessner, J.; Grant, S.; Hakonarson, H.; Price, R.A. Large Copy-Number Variations Are Enriched in Cases With Moderate to Extreme Obesity. Diabetes 2010, 59, 2690–2694. [Google Scholar] [CrossRef]
- Rieusset, J. The role of endoplasmic reticulum-mitochondria contact sites in the control of glucose homeostasis: An update. Cell Death Dis. 2018, 9, 388. [Google Scholar] [CrossRef] [PubMed]
- Tubbs, E.; Chanon, S.; Robert, M.; Bendridi, N.; Bidaux, G.; Chauvin, M.-A.; Ji-Cao, J.; Durand, C.; Gauvrit-Ramette, D.; Vidal, H.; et al. Disruption of Mitochondria-Associated Endoplasmic Reticulum Membrane (MAM) Integrity Contributes to Muscle Insulin Resistance in Mice and Humans. Diabetes 2018, 67, 636–650. [Google Scholar] [CrossRef]
- Tubbs, E.; Theurey, P.; Vial, G.; Bendridi, N.; Bravard, A.; Chauvin, M.-A.; Ji-Cao, J.; Zoulim, F.; Bartosch, B.; Ovize, M.; et al. Mitochondria-Associated Endoplasmic Reticulum Membrane (MAM) Integrity Is Required for Insulin Signaling and Is Implicated in Hepatic Insulin Resistance. Diabetes 2014, 63, 3279–3294. [Google Scholar] [CrossRef] [PubMed]
- Kuzmicic, J.; del Campo, A.; López-Crisosto, C.; Morales, P.E.; Pennanen, C.; Bravo-Sagua, R.; Hechenleitner, J.; Zepeda, R.; Castro, P.F.; Verdejo, H.E.; et al. Dinámica mitocondrial: Un potencial nuevo blanco terapéutico para la insuficiencia cardiaca. Rev. Española Cardiol. 2011, 64, 916–923. [Google Scholar] [CrossRef] [PubMed]
- Correa-Rodríguez, M.; Viatte, S.; Massey, J.; Schmidt-RioValle, J.; Rueda-Medina, B.; Orozco, G. Analysis of SNP-SNP interactions and bone quantitative ultrasound parameter in early adulthood. BMC Med Genet. 2017, 18, 107. [Google Scholar] [CrossRef]
- Mullin, B.H.; Zhao, J.H.; Brown, S.J.; Perry, J.R.; Luan, J.; Zheng, H.-F.; Langenberg, C.; Dudbridge, F.; Scott, R.; Wareham, N.J.; et al. Genome-wide association study meta-analysis for quantitative ultrasound parameters of bone identifies five novel loci for broadband ultrasound attenuation. Hum. Mol. Genet. 2017, 26, 2791–2802. [Google Scholar] [CrossRef]
- Breckenridge, D.G.; Stojanovic, M.; Marcellus, R.C.; Shore, G.C. Caspase cleavage product of BAP31 induces mitochondrial fission through endoplasmic reticulum calcium signals, enhancing cytochrome c release to the cytosol. J. Cell Biol. 2003, 160, 1115–1127. [Google Scholar] [CrossRef]
- Chu, B.-B.; Liao, Y.-C.; Qi, W.; Xie, C.; Du, X.; Wang, J.; Yang, H.; Miao, H.-H.; Li, B.-L.; Song, B.-L. Cholesterol Transport through Lysosome-Peroxisome Membrane Contacts. Cell 2015, 161, 291–306. [Google Scholar] [CrossRef] [PubMed]
- Uhlén, M.; Fagerberg, L.; Hallström, B.M.; Lindskog, C.; Oksvold, P.; Mardinoglu, A.; Sivertsson, Å.; Kampf, C.; Sjöstedt, E.; Asplund, A.; et al. Tissue-based map of the human proteome. Science 2015, 347, 1260419. [Google Scholar] [CrossRef] [PubMed]
- Silva, B.S.; DiGiovanni, L.; Kumar, R.; Carmichael, R.E.; Kim, P.K.; Schrader, M. Maintaining social contacts: The physiological relevance of organelle interactions. Biochim. Biophys. Acta (BBA) Bioenerg. 2020, 1867, 118800. [Google Scholar] [CrossRef] [PubMed]
- Maharjan, Y.; Lee, J.N.; Kwak, S.A.; Dutta, R.K.; Park, C.; Choe, S.; Park, R. TMEM135 regulates primary ciliogenesis through modulation of intracellular cholesterol distribution. EMBO Rep. 2020, 21, e48901. [Google Scholar] [CrossRef] [PubMed]
- Schmit, K.; Michiels, C. TMEM Proteins in Cancer: A Review. Front. Pharmacol. 2018, 9, 1345. [Google Scholar] [CrossRef] [PubMed]
- Roy, A.; Kucukural, A.; Zhang, Y. I-TASSER: A unified platform for automated protein structure and function prediction. Nat. Protoc. 2010, 5, 725–738. [Google Scholar] [CrossRef]
- Yang, J.; Zhang, Y. I-TASSER server: New development for protein structure and function predictions. Nucleic Acids Res. 2015, 43, W174–W181. [Google Scholar] [CrossRef]
- Hittelman, K.J.; Fairhurst, A.S.; Smith, R.E. Calcium accumulation as a parameter of energy metabolism in mitochondria of brown adipose tissue. Proc. Natl. Acad. Sci. USA 1967, 58, 697–702. [Google Scholar] [CrossRef]
- Golic, I.; Veličković, K.; Markelic, M.; Stancic, A.; Jankovic, A.; Vucetic, M.; Otasevic, V.; Buzadzic, B.; Korac, B.; Korać, A. Calcium-induced alteration of mitochondrial morphology and mitochondrial-endoplasmic reticulum contacts in rat brown adipocytes. Eur. J. Histochem. 2014, 58, 2377. [Google Scholar] [CrossRef]
- Otera, H.; Ishihara, N.; Mihara, K. New insights into the function and regulation of mitochondrial fission. Biochim. Biophys. Acta (BBA)-Bioenerg. 2013, 1833, 1256–1268. [Google Scholar] [CrossRef]
- Giacomello, M.; Pellegrini, L. The coming of age of the mitochondria–ER contact: A matter of thickness. Cell Death Differ. 2016, 23, 1417–1427. [Google Scholar] [CrossRef]
- Vance, J.E. MAM (mitochondria-associated membranes) in mammalian cells: Lipids and beyond. Biochim. Biophys. Acta (BBA)—Mol. Cell Biol. Lipids 2014, 1841, 595–609. [Google Scholar] [CrossRef] [PubMed]
- Rowland, A.A.; Voeltz, G.K. Endoplasmic reticulum–mitochondria contacts: Function of the junction. Nat. Rev. Mol. Cell Biol. 2012, 13, 607–615. [Google Scholar] [CrossRef] [PubMed]
- Dewenter, M.; Von Der Lieth, A.; Katus, H.A.; Backs, J. Calcium Signaling and Transcriptional Regulation in Cardiomyocytes. Circ. Res. 2017, 121, 1000–1020. [Google Scholar] [CrossRef] [PubMed]
- West, A.E.; Chen, W.G.; Dalva, M.B.; Dolmetsch, R.E.; Kornhauser, J.M.; Shaywitz, A.J.; Takasu, M.A.; Tao, X.; Greenberg, M.E. Calcium regulation of neuronal gene expression. Proc. Natl. Acad. Sci. USA 2001, 98, 11024–11031. [Google Scholar] [CrossRef] [PubMed]
- Shambharkar, P.B.; Bittinger, M.; Latario, B.; Xiong, Z.; Bandyopadhyay, S.; Davis, V.; Lin, V.; Yang, Y.; Valdez, R.; Labow, M.A. TMEM203 Is a Novel Regulator of Intracellular Calcium Homeostasis and Is Required for Spermatogenesis. PLoS ONE 2015, 10, e0127480. [Google Scholar] [CrossRef] [PubMed]
- Markin, A.; Khotina, V.; Zabudskaya, X.; Bogatyreva, A.; Starodubova, A.; Ivanova, E.; Nikiforov, N.; Orekhov, A. Disturbance of Mitochondrial Dynamics and Mitochondrial Therapies in Atherosclerosis. Life 2021, 11, 165. [Google Scholar] [CrossRef] [PubMed]
- Landowski, M.; Grindel, S.; Shahi, P.K.; Johnson, A.; Western, D.; Race, A.; Shi, F.; Benson, J.; Gao, M.; Santoirre, E.; et al. Modulation of Tmem135 Leads to Retinal Pigmented Epithelium Pathologies in Mice. Investig. Opthalmology Vis. Sci. 2020, 61, 16. [Google Scholar] [CrossRef]
- Spurthi, K.M.; Sarikhani, M.; Mishra, S.; Desingu, P.A.; Yadav, S.; Rao, S.; Maity, S.; Tamta, A.K.; Kumar, S.; Majumdar, S.; et al. Toll-like receptor 2 deficiency hyperactivates the FoxO1 transcription factor and induces aging-associated cardiac dysfunction in mice. J. Biol. Chem. 2018, 293, 13073–13089. [Google Scholar] [CrossRef] [PubMed]
- Jia, K.; Chen, D.; Riddle, D.L. The TOR pathway interacts with the insulin signaling pathway to regulate C. elegans larval development, metabolism and life span. Development 2004, 131, 3897–3906. [Google Scholar] [CrossRef]
- Vellai, T.; Takacs-Vellai, K.; Zhang, Y.; Kovacs, A.L.; Orosz, L.; Müller, F. Influence of TOR kinase on lifespan in C. elegans. Nat. Cell Biol. 2003, 426, 620. [Google Scholar] [CrossRef] [PubMed]
- Kondo, M.; Yanase, S.; Ishii, T.; Hartman, P.S.; Matsumoto, K.; Ishii, N. The p38 signal transduction pathway participates in the oxidative stress-mediated translocation of DAF-16 to C. elegans nuclei. Mech. Ageing Dev. 2005, 126, 642–647. [Google Scholar] [CrossRef] [PubMed]
- Fusakio, M.E.; Willy, J.A.; Wang, Y.; Mirek, E.T.; Al Baghdadi, R.J.T.; Adams, C.; Anthony, T.G.; Wek, R.C. Transcription factor ATF4 directs basal and stress-induced gene expression in the unfolded protein response and cholesterol metabolism in the liver. Mol. Biol. Cell 2016, 27, 1536–1551. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [PubMed]
- Stark, C.; Breitkreutz, B.J.; Reguly, T.; Boucher, L.; Breitkreutz, A.; Tyers, M. BioGRID: A general repository for interaction datasets. Nucleic Acids Res. 2006, 34, D535–D539. [Google Scholar] [CrossRef]
- Khavandgar, Z.; Murshed, M. Sphingolipid metabolism and its role in the skeletal tissues. Cell. Mol. Life Sci. 2014, 72, 959–969. [Google Scholar] [CrossRef]
- Pulli, I.; Asghar, M.Y.; Kemppainen, K.; Törnquist, K. Sphingolipid-mediated calcium signaling and its pathological effects. Biochim. Biophys. Acta (BBA)—Bioenerg. 2018, 1865, 1668–1677. [Google Scholar] [CrossRef]
- Marchesini, N.; Osta, W.; Bielawski, J.; Luberto, C.; Obeid, L.M.; Hannun, Y.A. Role for Mammalian Neutral Sphingomyelinase 2 in Confluence-induced Growth Arrest of MCF7 Cells. J. Biol. Chem. 2004, 279, 25101–25111. [Google Scholar] [CrossRef]
- Hofmann, K.; Tomiuk, S.; Wolff, G.; Stoffel, W. Cloning and characterization of the mammalian brain-specific, Mg2+-dependent neutral sphingomyelinase. Proc. Natl. Acad. Sci. USA 2000, 97, 5895–5900. [Google Scholar] [CrossRef] [PubMed]
- Miura, Y.; Gotoh, E.; Nara, F.; Nishijima, M.; Hanada, K. Hydrolysis of sphingosylphosphocholine by neutral sphingomyelinases. FEBS Lett. 2004, 557, 288–292. [Google Scholar] [CrossRef]
- Sehnal, D.; Bittrich, S.; Deshpande, M.; Svobodová, R.; Berka, K.; Bazgier, V.; Velankar, S.; Burley, S.K.; Koča, J.; Rose, A.S. Mol* Viewer: Modern web app for 3D visualization and analysis of large biomolecular structures. Nucleic Acids Res. 2021, 49, W431–W437. [Google Scholar] [CrossRef] [PubMed]
- Airola, M.; Shanbhogue, P.; Shamseddine, A.A.; Guja, K.E.; Senkal, C.E.; Maini, R.; Bartke, N.; Wu, B.X.; Obeid, L.M.; Garcia-Diaz, M.; et al. Structure of human nSMase2 reveals an interdomain allosteric activation mechanism for ceramide generation. Proc. Natl. Acad. Sci. USA 2017, 114, E5549–E5558. [Google Scholar] [CrossRef]
- Natrajan, R.; Mackay, A.; Lambros, M.B.; Weigelt, B.; Wilkerson, P.M.; Manie, E.; Grigoriadis, A.; A’Hern, R.; Van Der Groep, P.; Kozarewa, I.; et al. A whole-genome massively parallel sequencing analysis of BRCA1 mutant oestrogen receptor-negative and -positive breast cancers. J. Pathol. 2012, 227, 29–41. [Google Scholar] [CrossRef]
- Li, J.; Zhong, H.-Y.; Zhang, Y.; Xiao, L.; Bai, L.-H.; Liu, S.-F.; Zhou, G.-B.; Zhang, G.-S. GTF2I-RARAis a novel fusion transcript in a t(7;17) variant of acute promyelocytic leukaemia with clinical resistance to retinoic acid. Br. J. Haematol. 2014, 168, 904–908. [Google Scholar] [CrossRef]
- Yu, Y.-P.; Liu, P.; Nelson, J.; Hamilton, R.L.; Bhargava, R.; Michalopoulos, G.; Chen, Q.; Zhang, J.; Ma, D.; Pennathur, A.; et al. Identification of recurrent fusion genes across multiple cancer types. Sci. Rep. 2019, 9, 1–9. [Google Scholar] [CrossRef]
- Franić, S.; Groen-Blokhuis, M.M.; Dolan, C.V.; Kattenberg, M.V.; Pool, R.; Xiao, X.; Scheet, P.A.; Ehli, E.A.; Davies, G.E.; van der Sluis, S.; et al. Intelligence: Shared genetic basis between Mendelian disorders and a polygenic trait. Eur. J. Hum. Genet. 2015, 23, 1378–1383. [Google Scholar] [CrossRef]
- Schweppe, D.K.; Huttlin, E.; Harper, J.; Gygi, S.P. BioPlex Display: An Interactive Suite for Large-Scale AP–MS Protein–Protein Interaction Data. J. Proteome Res. 2018, 17, 722–726. [Google Scholar] [CrossRef]
- Huttlin, E.L.; Bruckner, R.J.; Paulo, J.A.; Cannon, J.R.; Ting, L.; Baltier, K.; Colby, G.; Gebreab, F.; Gygi, M.P.; Parzen, H.; et al. Architecture of the human interactome defines protein communities and disease networks. Nature 2017, 545, 505–509. [Google Scholar] [CrossRef]
- Huttlin, E.; Ting, L.; Bruckner, R.J.; Gebreab, F.; Gygi, M.P.; Szpyt, J.; Tam, S.; Zarraga, G.; Colby, G.; Baltier, K.; et al. The BioPlex Network: A Systematic Exploration of the Human Interactome. Cell 2015, 162, 425–440. [Google Scholar] [CrossRef]
- Aguayo-Orozco, A.; Bois, F.Y.; Brunak, S.; Taboureau, O. Analysis of Time-Series Gene Expression Data to Explore Mechanisms of Chemical-Induced Hepatic Steatosis Toxicity. Front. Genet. 2018, 9, 396. [Google Scholar] [CrossRef]
- Yang, W.; Li, Y.; He, F.; Wu, H. Microarray profiling of long non-coding RNA (lncRNA) associated with hypertrophic cardiomyopathy. BMC Cardiovasc. Disord. 2015, 15, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Yan, R.; Roy, A.; Xu, D.; Poisson, J.; Zhang, Y. The I-TASSER Suite: Protein structure and function prediction. Nat. Methods 2015, 12, 7–8. [Google Scholar] [CrossRef] [PubMed]
- Hodgkin, A.L.; Huxley, A.F. The components of membrane conductance in the giant axon of Loligo. J. Physiol. 1952, 116, 473–496. [Google Scholar] [CrossRef] [PubMed]
- Kadir, L.A.; Stacey, M.; Barrett-Jolley, R. Emerging Roles of the Membrane Potential: Action Beyond the Action Potential. Front. Physiol. 2018, 9, 1661. [Google Scholar] [CrossRef] [PubMed]
| Predicted Location | Major Expression in Tsue | Known Interactions with Organelles/Vesicles | Physiological Role | Implication in Human Disease |
|---|---|---|---|---|
|
|
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Beasley, H.K.; Rodman, T.A.; Collins, G.V.; Hinton, A., Jr.; Exil, V. TMEM135 is a Novel Regulator of Mitochondrial Dynamics and Physiology with Implications for Human Health Conditions. Cells 2021, 10, 1750. https://doi.org/10.3390/cells10071750
Beasley HK, Rodman TA, Collins GV, Hinton A Jr., Exil V. TMEM135 is a Novel Regulator of Mitochondrial Dynamics and Physiology with Implications for Human Health Conditions. Cells. 2021; 10(7):1750. https://doi.org/10.3390/cells10071750
Chicago/Turabian StyleBeasley, Heather K., Taylor A. Rodman, Greg V. Collins, Antentor Hinton, Jr., and Vernat Exil. 2021. "TMEM135 is a Novel Regulator of Mitochondrial Dynamics and Physiology with Implications for Human Health Conditions" Cells 10, no. 7: 1750. https://doi.org/10.3390/cells10071750
APA StyleBeasley, H. K., Rodman, T. A., Collins, G. V., Hinton, A., Jr., & Exil, V. (2021). TMEM135 is a Novel Regulator of Mitochondrial Dynamics and Physiology with Implications for Human Health Conditions. Cells, 10(7), 1750. https://doi.org/10.3390/cells10071750