The Emergence of Senescent Surface Biomarkers as Senotherapeutic Targets
Abstract
:1. Introduction
2. Markers of Cellular Senescence
2.1. Morphological Changes
2.2. Senescence-Associated β-Galactosidase Activity
2.3. Cell Cycle Regulators
2.4. SASP Levels
2.5. DNA Damage and Telomere Shortening
2.6. Novel RNA- and Protein-Markers of Cellular Senescence
3. Molecular Pathways in Cellular Senescence
3.1. p53/p21 Pathway
3.2. p16/RB Pathway
3.3. mTOR Pathway
4. Challenges and Limitations Associated with the Identification of Senescent Cells
4.1. Identification of Senescence in Tissues Ex Vivo
4.2. Identification of Senescent Cells In Vivo
5. The Emergence of Surface Proteins as Biomarkers and Therapeutic Targets
5.1. Novel Biomarkers on Senescent Cell Surface
5.1.1. DPP4
5.1.2. SCAMP4
5.1.3. DEP1
5.1.4. B2MG
5.1.5. CD264
5.1.6. CD36
5.1.7. ICAM-1
5.1.8. MDA-VIMENTIN
5.1.9. NOTCH1 and NOTCH3
5.1.10. MICA/B and ULBP2
5.1.11. uPAR
6. Implications of the Surfaceome in Senescence
6.1. Senescence Surfaceome and Immune Surveillance
6.2. Senescence Surfaceome and SASP
7. Clearance of Senescent Cells
7.1. Pharmacological and Genetic Approaches
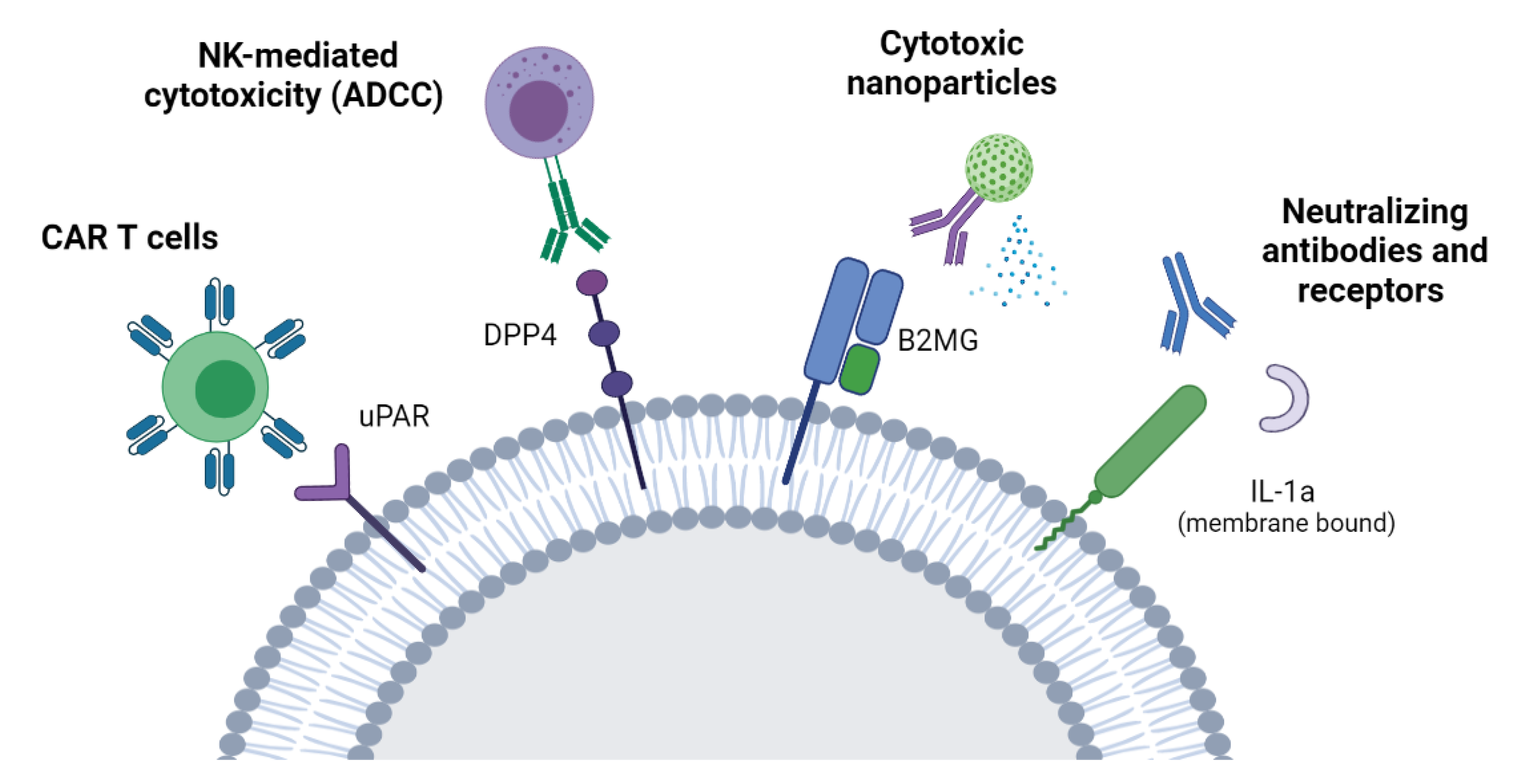
7.2. Targeting the Senescent Surfaceome
7.2.1. ADCC
7.2.2. CAR-T Cells
7.2.3. Neutralizing Antibodies/Nanoparticles
8. Closing Remarks and Future Perspectives
Funding
Conflicts of Interest
References
- Hayflick, L. The Limited In Vitro Lifetime of Human Diploid Cell Strains. Exp. Cell Res. 1965, 37, 614–636. [Google Scholar] [CrossRef]
- Kuilman, T.; Michaloglou, C.; Mooi, W.J.; Peeper, D.S. The Essence of Senescence. Genes Dev. 2010, 24, 2463–2479. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hernandez-Segura, A.; Nehme, J.; Demaria, M. Hallmarks of Cellular Senescence. Trends Cell Biol. 2018, 28, 436–453. [Google Scholar] [CrossRef]
- Kwon, S.M.; Hong, S.M.; Lee, Y.-K.; Min, S.; Yoon, G. Metabolic Features and Regulation in Cell Senescence. BMB Rep. 2019, 52, 5–12. [Google Scholar] [CrossRef] [Green Version]
- Terzi, M.Y.; Izmirli, M.; Gogebakan, B. The Cell Fate: Senescence or Quiescence. Mol. Biol. Rep. 2016, 43, 1213–1220. [Google Scholar] [CrossRef] [PubMed]
- Demaria, M.; Ohtani, N.; Youssef, S.A.; Rodier, F.; Toussaint, W.; Mitchell, J.R.; Laberge, R.-M.; Vijg, J.; Van Steeg, H.; Dollé, M.E.T.; et al. An Essential Role for Senescent Cells in Optimal Wound Healing through Secretion of PDGF-AA. Dev. Cell 2014, 31, 722–733. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Muñoz-Espín, D.; Serrano, M. Cellular Senescence: From Physiology to Pathology. Nat. Rev. Mol. Cell Biol. 2014, 15, 482–496. [Google Scholar] [CrossRef]
- He, S.; Sharpless, N.E. Senescence in Health and Disease. Cell 2017, 169, 1000–1011. [Google Scholar] [CrossRef]
- Freund, A.; Orjalo, A.V.; Desprez, P.-Y.; Campisi, J. Inflammatory Networks during Cellular Senescence: Causes and Consequences. Trends Mol. Med. 2010, 16, 238–246. [Google Scholar] [CrossRef] [Green Version]
- Franceschi, C.; Campisi, J. Chronic Inflammation (Inflammaging) and Its Potential Contribution to Age-Associated Diseases. J. Gerontol. A Biol. Sci. Med. Sci. 2014, 69 (Suppl. 1), S4–S9. [Google Scholar] [CrossRef]
- McHugh, D.; Gil, J. Senescence and Aging: Causes, Consequences, and Therapeutic Avenues. J. Cell Biol. 2018, 217, 65–77. [Google Scholar] [CrossRef] [PubMed]
- Gorgoulis, V.; Adams, P.D.; Alimonti, A.; Bennett, D.C.; Bischof, O.; Bishop, C.; Campisi, J.; Collado, M.; Evangelou, K.; Ferbeyre, G.; et al. Cellular Senescence: Defining a Path Forward. Cell 2019, 179, 813–827. [Google Scholar] [CrossRef] [PubMed]
- Campisi, J.; Kapahi, P.; Lithgow, G.J.; Melov, S.; Newman, J.C.; Verdin, E. From Discoveries in Ageing Research to Therapeutics for Healthy Ageing. Nature 2019, 571, 183–192. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Druelle, C.; Drullion, C.; Deslé, J.; Martin, N.; Saas, L.; Cormenier, J.; Malaquin, N.; Huot, L.; Slomianny, C.; Bouali, F.; et al. ATF6α Regulates Morphological Changes Associated with Senescence in Human Fibroblasts. Oncotarget 2016, 7, 67699–67715. [Google Scholar] [CrossRef] [Green Version]
- Freund, A.; Laberge, R.-M.; Demaria, M.; Campisi, J. Lamin B1 Loss Is a Senescence-Associated Biomarker. Mol. Biol. Cell 2012, 23, 2066–2075. [Google Scholar] [CrossRef]
- Lee, B.Y.; Han, J.A.; Im, J.S.; Morrone, A.; Johung, K.; Goodwin, E.C.; Kleijer, W.J.; DiMaio, D.; Hwang, E.S. Senescence-Associated Beta-Galactosidase Is Lysosomal Beta-Galactosidase. Aging Cell 2006, 5, 187–195. [Google Scholar] [CrossRef]
- Itahana, K.; Itahana, Y.; Dimri, G.P. Colorimetric Detection of Senescence-Associated β Galactosidase. Methods Mol. Biol. 2013, 965, 143–156. [Google Scholar] [CrossRef]
- Kim, S.R.; Eirin, A.; Zhang, X.; Lerman, A.; Lerman, L.O. Mitochondrial Protection Partly Mitigates Kidney Cellular Senescence in Swine Atherosclerotic Renal Artery Stenosis. Cell. Physiol. Biochem. 2019, 52, 617–632. [Google Scholar] [CrossRef] [Green Version]
- González-Gualda, E.; Baker, A.G.; Fruk, L.; Muñoz-Espín, D. A Guide to Assessing Cellular Senescence in Vitro and in Vivo. FEBS J. 2021, 288, 56–80. [Google Scholar] [CrossRef]
- Coppé, J.-P.; Desprez, P.-Y.; Krtolica, A.; Campisi, J. The Senescence-Associated Secretory Phenotype: The Dark Side of Tumor Suppression. Annu. Rev. Pathol. 2010, 5, 99–118. [Google Scholar] [CrossRef] [Green Version]
- Rea, I.M.; Gibson, D.S.; McGilligan, V.; McNerlan, S.E.; Alexander, H.D.; Ross, O.A. Age and Age-Related Diseases: Role of Inflammation Triggers and Cytokines. Front. Immunol. 2018, 9, 586. [Google Scholar] [CrossRef]
- Kadota, T.; Fujita, Y.; Yoshioka, Y.; Araya, J.; Kuwano, K.; Ochiya, T. Emerging Role of Extracellular Vesicles as a Senescence-Associated Secretory Phenotype: Insights into the Pathophysiology of Lung Diseases. Mol. Aspects Med. 2018, 60, 92–103. [Google Scholar] [CrossRef]
- Kuo, L.J.; Yang, L.-X. Gamma-H2AX—A Novel Biomarker for DNA Double-Strand Breaks. In Vivo 2008, 22, 305–309. [Google Scholar]
- Rodier, F.; Muñoz, D.P.; Teachenor, R.; Chu, V.; Le, O.; Bhaumik, D.; Coppé, J.-P.; Campeau, E.; Beauséjour, C.M.; Kim, S.-H.; et al. DNA-SCARS: Distinct Nuclear Structures That Sustain Damage-Induced Senescence Growth Arrest and Inflammatory Cytokine Secretion. J. Cell Sci. 2011, 124, 68–81. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, R.; Chen, W.; Adams, P.D. Molecular Dissection of Formation of Senescence-Associated Heterochromatin Foci. Mol. Cell Biol. 2007, 27, 2343–2358. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aird, K.M.; Zhang, R. Detection of Senescence-Associated Heterochromatin Foci (SAHF). Methods Mol. Biol. 2013, 965, 185–196. [Google Scholar] [CrossRef] [Green Version]
- Noren Hooten, N.; Evans, M.K. Techniques to Induce and Quantify Cellular Senescence. J. Vis. Exp. 2017. [Google Scholar] [CrossRef]
- Casella, G.; Munk, R.; Kim, K.M.; Piao, Y.; De, S.; Abdelmohsen, K.; Gorospe, M. Transcriptome Signature of Cellular Senescence. Nucleic Acids Res. 2019, 47, 7294–7305. [Google Scholar] [CrossRef] [Green Version]
- Kim, K.M.; Noh, J.H.; Bodogai, M.; Martindale, J.L.; Yang, X.; Indig, F.E.; Basu, S.K.; Ohnuma, K.; Morimoto, C.; Johnson, P.F.; et al. Identification of Senescent Cell Surface Targetable Protein DPP4. Genes Dev. 2017, 31, 1529–1534. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rufini, A.; Tucci, P.; Celardo, I.; Melino, G. Senescence and Aging: The Critical Roles of P53. Oncogene 2013, 32, 5129–5143. [Google Scholar] [CrossRef]
- Mijit, M.; Caracciolo, V.; Melillo, A.; Amicarelli, F.; Giordano, A. Role of P53 in the Regulation of Cellular Senescence. Biomolecules 2020, 10, 420. [Google Scholar] [CrossRef] [Green Version]
- Sammons, M.A.; Nguyen, T.-A.T.; McDade, S.S.; Fischer, M. Tumor Suppressor P53: From Engaging DNA to Target Gene Regulation. Nucleic Acids Res. 2020, 48, 8848–8869. [Google Scholar] [CrossRef]
- Yosef, R.; Pilpel, N.; Papismadov, N.; Gal, H.; Ovadya, Y.; Vadai, E.; Miller, S.; Porat, Z.; Ben-Dor, S.; Krizhanovsky, V. P21 Maintains Senescent Cell Viability under Persistent DNA Damage Response by Restraining JNK and Caspase Signaling. EMBO J. 2017, 36, 2280–2295. [Google Scholar] [CrossRef]
- Sherr, C.J.; Beach, D.; Shapiro, G.I. Targeting CDK4 and CDK6: From Discovery to Therapy. Cancer Discov. 2016, 6, 353–367. [Google Scholar] [CrossRef] [Green Version]
- Weichhart, T. MTOR as Regulator of Lifespan, Aging, and Cellular Senescence: A Mini-Review. Gerontology 2018, 64, 127–134. [Google Scholar] [CrossRef]
- Payea, M.J.; Anerillas, C.; Tharakan, R.; Gorospe, M. Translational Control during Cellular Senescence. Mol. Cell Biol. 2021, 41. [Google Scholar] [CrossRef]
- Yang, N.-C.; Hu, M.-L. The Limitations and Validities of Senescence Associated-Beta-Galactosidase Activity as an Aging Marker for Human Foreskin Fibroblast Hs68 Cells. Exp. Gerontol. 2005, 40, 813–819. [Google Scholar] [CrossRef] [PubMed]
- Kany, S.; Vollrath, J.T.; Relja, B. Cytokines in Inflammatory Disease. Int. J. Mol. Sci. 2019, 20, 6008. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Childs, B.G.; Durik, M.; Baker, D.J.; van Deursen, J.M. Cellular Senescence in Aging and Age-Related Disease: From Mechanisms to Therapy. Nat. Med. 2015, 21, 1424–1435. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Georgakopoulou, E.A.; Tsimaratou, K.; Evangelou, K.; Fernandez Marcos, P.J.; Zoumpourlis, V.; Trougakos, I.P.; Kletsas, D.; Bartek, J.; Serrano, M.; Gorgoulis, V.G. Specific Lipofuscin Staining as a Novel Biomarker to Detect Replicative and Stress-Induced Senescence. A Method Applicable in Cryo-Preserved and Archival Tissues. Aging 2013, 5, 37–50. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Idda, M.L.; McClusky, W.G.; Lodde, V.; Munk, R.; Abdelmohsen, K.; Rossi, M.; Gorospe, M. Survey of Senescent Cell Markers with Age in Human Tissues. Aging 2020, 12, 4052–4066. [Google Scholar] [CrossRef]
- Baker, D.J.; Wijshake, T.; Tchkonia, T.; LeBrasseur, N.K.; Childs, B.G.; van de Sluis, B.; Kirkland, J.L.; van Deursen, J.M. Clearance of P16Ink4a-Positive Senescent Cells Delays Ageing-Associated Disorders. Nature 2011, 479, 232–236. [Google Scholar] [CrossRef] [PubMed]
- Burd, C.E.; Sorrentino, J.A.; Clark, K.S.; Darr, D.B.; Krishnamurthy, J.; Deal, A.M.; Bardeesy, N.; Castrillon, D.H.; Beach, D.H.; Sharpless, N.E. Monitoring Tumorigenesis and Senescence in Vivo with a P16(INK4a)-Luciferase Model. Cell 2013, 152, 340–351. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ekpenyong-Akiba, A.E.; Poblocka, M.; Macip, S. Targeted Senolytic Strategies Based on the Senescent Surfaceome. In Senolytics in Disease, Ageing and Longevity; Muñoz-Espin, D., Demaria, M., Eds.; Springer International Publishing: Cham, Switzerland, 2020; pp. 103–130. ISBN 978-3-030-44903-2. [Google Scholar]
- Kim, K.M.; Noh, J.H.; Gorospe, M. Senolysis and Senostasis Through the Plasma Membrane. In Senolytics in Disease, Ageing and Longevity; Muñoz-Espin, D., Demaria, M., Eds.; Springer International Publishing: Cham, Switzerland, 2020; pp. 131–143. ISBN 978-3-030-44903-2. [Google Scholar]
- Althubiti, M.; Lezina, L.; Carrera, S.; Jukes-Jones, R.; Giblett, S.M.; Antonov, A.; Barlev, N.; Saldanha, G.S.; Pritchard, C.A.; Cain, K.; et al. Characterization of Novel Markers of Senescence and Their Prognostic Potential in Cancer. Cell Death Dis. 2014, 5, e1528. [Google Scholar] [CrossRef] [Green Version]
- Li, L.; van Breugel, P.C.; Loayza-Puch, F.; Ugalde, A.P.; Korkmaz, G.; Messika-Gold, N.; Han, R.; Lopes, R.; Barbera, E.P.; Teunissen, H.; et al. LncRNA-OIS1 Regulates DPP4 Activation to Modulate Senescence Induced by RAS. Nucleic Acids Res. 2018, 46, 4213–4227. [Google Scholar] [CrossRef]
- Johmura, Y.; Yamanaka, T.; Omori, S.; Wang, T.-W.; Sugiura, Y.; Matsumoto, M.; Suzuki, N.; Kumamoto, S.; Yamaguchi, K.; Hatakeyama, S.; et al. Senolysis by Glutaminolysis Inhibition Ameliorates Various Age-Associated Disorders. Science 2021, 371, 265–270. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Yu, J.; Fu, M.; Dong, R.; Yang, Y.; Luo, J.; Hu, S.; Li, W.; Xu, X.; Tu, L. Dipeptidyl Peptidase-4 Inhibition Improves Endothelial Senescence by Activating AMPK/SIRT1/Nrf2 Signaling Pathway. Biochem. Pharmacol. 2020, 177, 113951. [Google Scholar] [CrossRef] [PubMed]
- Worthen, C.A.; Cui, Y.; Orringer, J.S.; Johnson, T.M.; Voorhees, J.J.; Fisher, G.J. CD26 Identifies a Subpopulation of Fibroblasts That Produce the Majority of Collagen during Wound Healing in Human Skin. J. Investig. Dermatol. 2020, 140, 2515–2524.e3. [Google Scholar] [CrossRef]
- Wang, T.; Yang, L.; Liang, Z.; Wang, L.; Su, F.; Wang, X.; You, X.; He, C. Targeting Cellular Senescence Prevents Glucocorticoid-Induced Bone Loss through Modulation of the DPP4-GLP-1 Axis. Signal Transduct Target Ther. 2021, 6, 143. [Google Scholar] [CrossRef]
- Kim, K.M.; Noh, J.H.; Bodogai, M.; Martindale, J.L.; Pandey, P.R.; Yang, X.; Biragyn, A.; Abdelmohsen, K.; Gorospe, M. SCAMP4 Enhances the Senescent Cell Secretome. Genes Dev. 2018, 32, 909–914. [Google Scholar] [CrossRef]
- Sagiv, A.; Burton, D.G.A.; Moshayev, Z.; Vadai, E.; Wensveen, F.; Ben-Dor, S.; Golani, O.; Polic, B.; Krizhanovsky, V. NKG2D Ligands Mediate Immunosurveillance of Senescent Cells. Aging 2016, 8, 328–344. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Muñoz, D.P.; Yannone, S.M.; Daemen, A.; Sun, Y.; Vakar-Lopez, F.; Kawahara, M.; Freund, A.M.; Rodier, F.; Wu, J.D.; Desprez, P.-Y.; et al. Targetable Mechanisms Driving Immunoevasion of Persistent Senescent Cells Link Chemotherapy-Resistant Cancer to Aging. JCI Insight 2019, 5. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zingoni, A.; Cecere, F.; Vulpis, E.; Fionda, C.; Molfetta, R.; Soriani, A.; Petrucci, M.T.; Ricciardi, M.R.; Fuerst, D.; Amendola, M.G.; et al. Genotoxic Stress Induces Senescence-Associated ADAM10-Dependent Release of NKG2D MIC Ligands in Multiple Myeloma Cells. J. Immunol. 2015, 195, 736–748. [Google Scholar] [CrossRef] [Green Version]
- Madsen, S.D.; Russell, K.C.; Tucker, H.A.; Glowacki, J.; Bunnell, B.A.; O’Connor, K.C. Decoy TRAIL Receptor CD264: A Cell Surface Marker of Cellular Aging for Human Bone Marrow-Derived Mesenchymal Stem Cells. Stem Cell Res. Ther. 2017, 8, 201. [Google Scholar] [CrossRef]
- Hoare, M.; Ito, Y.; Kang, T.-W.; Weekes, M.P.; Matheson, N.J.; Patten, D.A.; Shetty, S.; Parry, A.J.; Menon, S.; Salama, R.; et al. NOTCH1 Mediates a Switch between Two Distinct Secretomes during Senescence. Nat. Cell Biol. 2016, 18, 979–992. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cui, H.; Kong, Y.; Xu, M.; Zhang, H. Notch3 Functions as a Tumor Suppressor by Controlling Cellular Senescence. Cancer Res. 2013, 73, 3451–3459. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chong, M.; Yin, T.; Chen, R.; Xiang, H.; Yuan, L.; Ding, Y.; Pan, C.C.; Tang, Z.; Alexander, P.B.; Li, Q.-J.; et al. CD36 Initiates the Secretory Phenotype during the Establishment of Cellular Senescence. EMBO Rep. 2018, 19, e45274. [Google Scholar] [CrossRef] [PubMed]
- Frescas, D.; Roux, C.M.; Aygun-Sunar, S.; Gleiberman, A.S.; Krasnov, P.; Kurnasov, O.V.; Strom, E.; Virtuoso, L.P.; Wrobel, M.; Osterman, A.L.; et al. Senescent Cells Expose and Secrete an Oxidized Form of Membrane-Bound Vimentin as Revealed by a Natural Polyreactive Antibody. Proc. Natl. Acad. Sci USA 2017, 114, E1668–E1677. [Google Scholar] [CrossRef] [Green Version]
- Amor, C.; Feucht, J.; Leibold, J.; Ho, Y.-J.; Zhu, C.; Alonso-Curbelo, D.; Mansilla-Soto, J.; Boyer, J.A.; Li, X.; Giavridis, T.; et al. Senolytic CAR T Cells Reverse Senescence-Associated Pathologies. Nature 2020, 583, 127–132. [Google Scholar] [CrossRef]
- Lioulios, G.; Fylaktou, A.; Papagianni, A.; Stangou, M. T Cell Markers Recount the Course of Immunosenescence in Healthy Individuals and Chronic Kidney Disease. Clin. Immunol. 2021, 225, 108685. [Google Scholar] [CrossRef]
- Aung, K.T.; Akiyama, K.; Kunitomo, M.; Mun, A.Y.; Tosa, I.; Nguyen, H.T.T.; Zhang, J.; Kohno, T.; Ono, M.; Hara, E.S.; et al. Aging-Affected MSC Functions and Severity of Periodontal Tissue Destruction in a Ligature-Induced Mouse Periodontitis Model. Int. J. Mol. Sci. 2020, 21, 8103. [Google Scholar] [CrossRef]
- Harada, S.; Mabuchi, Y.; Kohyama, J.; Shimojo, D.; Suzuki, S.; Kawamura, Y.; Araki, D.; Suyama, T.; Kajikawa, M.; Akazawa, C.; et al. FZD5 Regulates Cellular Senescence in Human Mesenchymal Stem/Stromal Cells. Stem Cells 2021, 39, 318–330. [Google Scholar] [CrossRef]
- Nielson, C.M.; Wiedrick, J.; Shen, J.; Jacobs, J.; Baker, E.S.; Baraff, A.; Piehowski, P.; Lee, C.G.; Baratt, A.; Petyuk, V.; et al. Identification of Hip BMD Loss and Fracture Risk Markers Through Population-Based Serum Proteomics. J. Bone Miner. Res. 2017, 32, 1559–1567. [Google Scholar] [CrossRef]
- Qu, A.; Wu, X.; Li, S.; Sun, M.; Xu, L.; Kuang, H.; Xu, C. An NIR-Responsive DNA-Mediated Nanotetrahedron Enhances the Clearance of Senescent Cells. Adv. Mater. 2020, 32, e2000184. [Google Scholar] [CrossRef] [PubMed]
- Lizardo, D.Y.; Lin, Y.-L.; Gokcumen, O.; Atilla-Gokcumen, G.E. Regulation of Lipids Is Central to Replicative Senescence. Mol. Biosyst. 2017, 13, 498–509. [Google Scholar] [CrossRef] [PubMed]
- Gorgoulis, V.G.; Pratsinis, H.; Zacharatos, P.; Demoliou, C.; Sigala, F.; Asimacopoulos, P.J.; Papavassiliou, A.G.; Kletsas, D. P53-Dependent ICAM-1 Overexpression in Senescent Human Cells Identified in Atherosclerotic Lesions. Lab Investig. 2005, 85, 502–511. [Google Scholar] [CrossRef]
- Chia, C.W.; Egan, J.M. Incretin-Based Therapies in Type 2 Diabetes Mellitus. J. Clin. Endocrinol. Metab. 2008, 93, 3703–3716. [Google Scholar] [CrossRef] [Green Version]
- Ahrén, B.; Schmitz, O. GLP-1 Receptor Agonists and DPP-4 Inhibitors in the Treatment of Type 2 Diabetes. Horm. Metab. Res. 2004, 36, 867–876. [Google Scholar] [CrossRef] [PubMed]
- Munk, R.; Kim, K.M.; Gorospe, M.; Abdelmohsen, K. Senescence LncRNAs Govern Cell Surface Components: LncRNA-OIS1 Transcriptionally Elevates DPP4. Non Coding RNA Investig. 2019, 3. [Google Scholar] [CrossRef]
- Xin, M.; Jin, X.; Cui, X.; Jin, C.; Piao, L.; Wan, Y.; Xu, S.; Zhang, S.; Yue, X.; Wang, H.; et al. Dipeptidyl Peptidase-4 Inhibition Prevents Vascular Aging in Mice under Chronic Stress: Modulation of Oxidative Stress and Inflammation. Chem. Biol. Interact. 2019, 314, 108842. [Google Scholar] [CrossRef] [PubMed]
- Ban, T.H.; Kim, E.N.; Kim, M.Y.; Lim, J.H.; Lee, J.H.; Kim, H.D.; Yoon, H.E.; Park, C.W.; Choi, B.S. Renoprotective Effect of a Dipeptidyl Peptidase-4 Inhibitor on Aging Mice. Aging Dis. 2020, 11, 588–602. [Google Scholar] [CrossRef] [PubMed]
- Dave, R.K.; Naylor, A.J.; Young, S.P.; Bayley, R.; Hardie, D.L.; Haworth, O.; Rider, D.A.; Cook, A.D.; Buckley, C.D.; Kellie, S. Differential Expression of CD148 on Leukocyte Subsets in Inflammatory Arthritis. Arthritis Res. Ther. 2013, 15, R108. [Google Scholar] [CrossRef] [Green Version]
- Zhu, J.W.; Doan, K.; Park, J.; Chau, A.H.; Zhang, H.; Lowell, C.A.; Weiss, A. Receptor-like Tyrosine Phosphatases CD45 and CD148 Have Distinct Functions in Chemoattractant-Mediated Neutrophil Migration and Response to S. Aureus. Immunity 2011, 35, 757–769. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Takahashi, K.; Kim, R.H.; Pasic, L.; He, L.; Nagasaka, S.; Katagiri, D.; May, T.; Shimizu, A.; Harris, R.C.; Mernaugh, R.L.; et al. Agonistic Anti-CD148 Monoclonal Antibody Attenuates Diabetic Nephropathy in Mice. Am. J. Physiol. Renal Physiol. 2020, 318, F647–F659. [Google Scholar] [CrossRef]
- Pereira, B.I.; Devine, O.P.; Vukmanovic-Stejic, M.; Chambers, E.S.; Subramanian, P.; Patel, N.; Virasami, A.; Sebire, N.J.; Kinsler, V.; Valdovinos, A.; et al. Senescent Cells Evade Immune Clearance via HLA-E-Mediated NK and CD8+ T Cell Inhibition. Nat. Commun. 2019, 10, 2387. [Google Scholar] [CrossRef]
- Orjalo, A.V.; Bhaumik, D.; Gengler, B.K.; Scott, G.K.; Campisi, J. Cell Surface-Bound IL-1alpha Is an Upstream Regulator of the Senescence-Associated IL-6/IL-8 Cytokine Network. Proc. Natl. Acad. Sci. USA 2009, 106, 17031–17036. [Google Scholar] [CrossRef] [Green Version]
- Kim, E.-C.; Kim, J.-R. Senotherapeutics: Emerging Strategy for Healthy Aging and Age-Related Disease. BMB Rep. 2019, 52, 47–55. [Google Scholar] [CrossRef] [Green Version]
- von Kobbe, C. Targeting Senescent Cells: Approaches, Opportunities, Challenges. Aging 2019, 11, 12844–12861. [Google Scholar] [CrossRef]
- Roos, C.M.; Zhang, B.; Palmer, A.K.; Ogrodnik, M.B.; Pirtskhalava, T.; Thalji, N.M.; Hagler, M.; Jurk, D.; Smith, L.A.; Casaclang-Verzosa, G.; et al. Chronic Senolytic Treatment Alleviates Established Vasomotor Dysfunction in Aged or Atherosclerotic Mice. Aging Cell 2016, 15, 973–977. [Google Scholar] [CrossRef] [PubMed]
- Farr, J.N.; Xu, M.; Weivoda, M.M.; Monroe, D.G.; Fraser, D.G.; Onken, J.L.; Negley, B.A.; Sfeir, J.G.; Ogrodnik, M.B.; Hachfeld, C.M.; et al. Targeting Cellular Senescence Prevents Age-Related Bone Loss in Mice. Nat. Med. 2017, 23, 1072–1079. [Google Scholar] [CrossRef]
- Schafer, M.J.; White, T.A.; Iijima, K.; Haak, A.J.; Ligresti, G.; Atkinson, E.J.; Oberg, A.L.; Birch, J.; Salmonowicz, H.; Zhu, Y.; et al. Cellular Senescence Mediates Fibrotic Pulmonary Disease. Nat. Commun. 2017, 8, 14532. [Google Scholar] [CrossRef]
- Justice, J.N.; Nambiar, A.M.; Tchkonia, T.; LeBrasseur, N.K.; Pascual, R.; Hashmi, S.K.; Prata, L.; Masternak, M.M.; Kritchevsky, S.B.; Musi, N.; et al. Senolytics in Idiopathic Pulmonary Fibrosis: Results from a First-in-Human, Open-Label, Pilot Study. EBioMedicine 2019, 40, 554–563. [Google Scholar] [CrossRef] [Green Version]
- Hickson, L.J.; Langhi Prata, L.G.P.; Bobart, S.A.; Evans, T.K.; Giorgadze, N.; Hashmi, S.K.; Herrmann, S.M.; Jensen, M.D.; Jia, Q.; Jordan, K.L.; et al. Senolytics Decrease Senescent Cells in Humans: Preliminary Report from a Clinical Trial of Dasatinib plus Quercetin in Individuals with Diabetic Kidney Disease. EBioMedicine 2019, 47, 446–456. [Google Scholar] [CrossRef] [Green Version]
- Ellison-Hughes, G.M. First Evidence That Senolytics Are Effective at Decreasing Senescent Cells in Humans. EBioMedicine 2020, 56, 102473. [Google Scholar] [CrossRef] [PubMed]
- Thompson, P.J.; Shah, A.; Ntranos, V.; Van Gool, F.; Atkinson, M.; Bhushan, A. Targeted Elimination of Senescent Beta Cells Prevents Type 1 Diabetes. Cell Metab. 2019, 29, 1045–1060.e10. [Google Scholar] [CrossRef] [PubMed]
- Caruana, I.; Diaconu, I.; Dotti, G. From Monoclonal Antibodies to Chimeric Antigen Receptors for the Treatment of Human Malignancies. Semin. Oncol. 2014, 41, 661–666. [Google Scholar] [CrossRef] [PubMed] [Green Version]
| Surface Protein | General Function | Implication in Senescence/Aging | Potential Implication in Senotherapy | Refs. |
|---|---|---|---|---|
| DEP1 PTPRJ CD148 | Negative regulation of growth factor signals and cell proliferation. | Biomarkers of senescence. | [46] | |
| B2MG B2M | Presentation of peptide antigens to the immune system. | Biomarkers of senescence. High levels in serum of elderly population | Target for cytotoxic nanoparticles directed at senescent cells. | [46,65,66] |
| CD264 TNFRSF10D TRAILR4 | Antiapoptotic receptor, decoy receptor for TRAIL. | Markers of senescent hBM-MSCs. | [56] | |
| CD36 | Scavenger receptor with a role in inflammation and lipid metabolism. | Regulation of lipid metabolism. | SASP regulation. | [59,67] |
| ICAM-1 | Glycoprotein which mediates the adhesion between endothelial cells and activated leukocytes. | Marker of senescence Increased expression in atherosclerotic lesions | Oxidized form of vimentin, an intermediate filament. | [68] |
| MDA-VIMENTIN | Oxidized form of vimentin, an intermediate filament | Marker of senescence. Increased expression in plasma of age-accelerated mice | [60] | |
| DPP4 CD26 | Cleavage of several substrates including cytokines and growth factors. Regulation of incretins in glucose homeostasis. | Biomarker of senescence Protective role on the vascular system and kidney of aging mice | Target for ADCC (NK-mediated cytotoxicity) for the clearance of senescent cells. | [29,47,48] |
| NOTCH1 | Member of the NOTCH signaling pathway. | Regulation of different SASP profiles. | SASP regulation. | [57] |
| NOTCH3 | Member of the NOTCH signaling pathway. | Regulation of the onset of cellular senescence. | [58] | |
| SCAMP4 | Secretory protein involved in membrane trafficking. | Regulation of pro-inflammatory SASP. | SASP regulation. | [52] |
| MICA/B ULBP2 | Ligands for the NKG2D receptor. | Regulation of immune surveillance. | Clearance of senescent cells through NK-mediated cytotoxicity. | [53,54,55] |
| uPAR | Regulation of intracellular signaling in response to extracellular components. | Upregulated in senescence. | Target for the elimination of senescent cells through CAR T cells. | [61] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rossi, M.; Abdelmohsen, K. The Emergence of Senescent Surface Biomarkers as Senotherapeutic Targets. Cells 2021, 10, 1740. https://doi.org/10.3390/cells10071740
Rossi M, Abdelmohsen K. The Emergence of Senescent Surface Biomarkers as Senotherapeutic Targets. Cells. 2021; 10(7):1740. https://doi.org/10.3390/cells10071740
Chicago/Turabian StyleRossi, Martina, and Kotb Abdelmohsen. 2021. "The Emergence of Senescent Surface Biomarkers as Senotherapeutic Targets" Cells 10, no. 7: 1740. https://doi.org/10.3390/cells10071740
APA StyleRossi, M., & Abdelmohsen, K. (2021). The Emergence of Senescent Surface Biomarkers as Senotherapeutic Targets. Cells, 10(7), 1740. https://doi.org/10.3390/cells10071740






