Synbiotics Alleviate the Gut Indole Load and Dysbiosis in Chronic Kidney Disease
Abstract
1. Introduction
2. Materials and Methods
2.1. Human Blood Metabolite Analysis
2.2. Human Fecal Microbiota Analysis
2.3. The CKD Animal Model
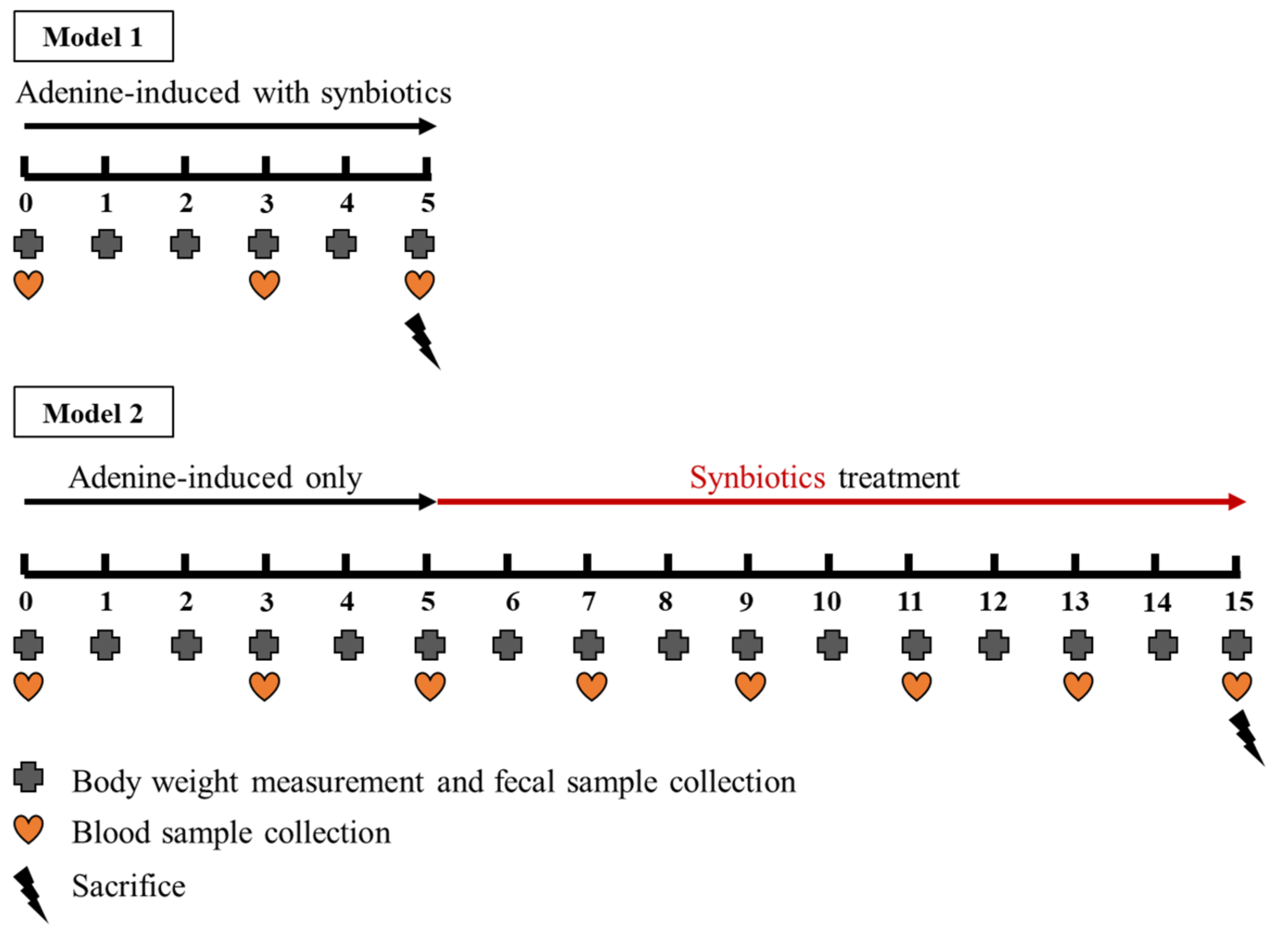
2.4. The Animal Model 1: Concomitant Five-Week Adenine Diet and Five-Week Synbiotic Treatment
2.5. The Animal Model 2: Five-Week Adenine Diet Followed by 10-Week Synbiotic Treatment
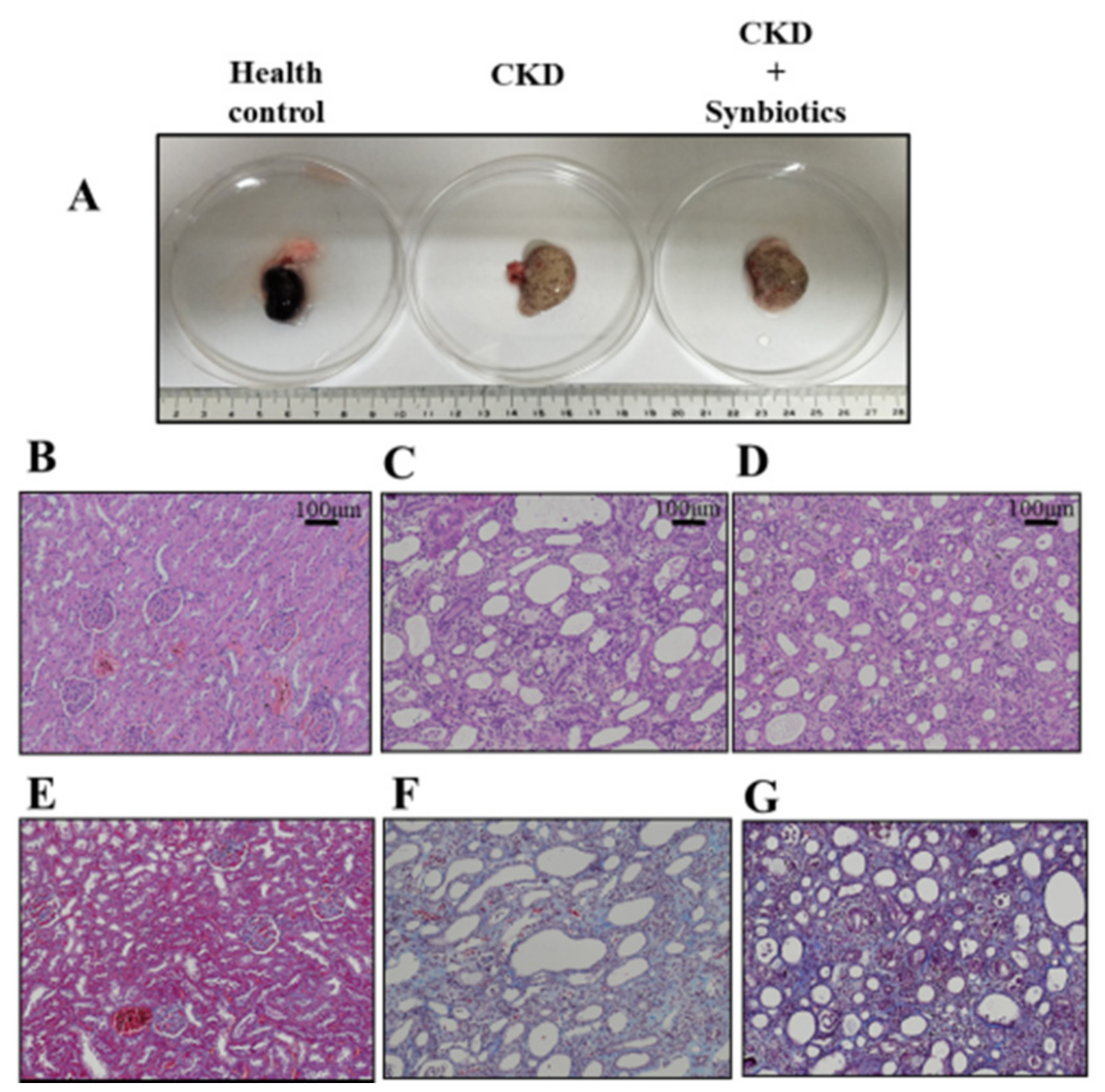
2.6. Rat Serum Biochemistry and Kidney Pathology
2.7. Rat Fecal Microbiota Analysis
2.8. Kovács Analysis for Rat and Human Fecal Indole Quantification
2.9. Statistical Analysis
3. Results
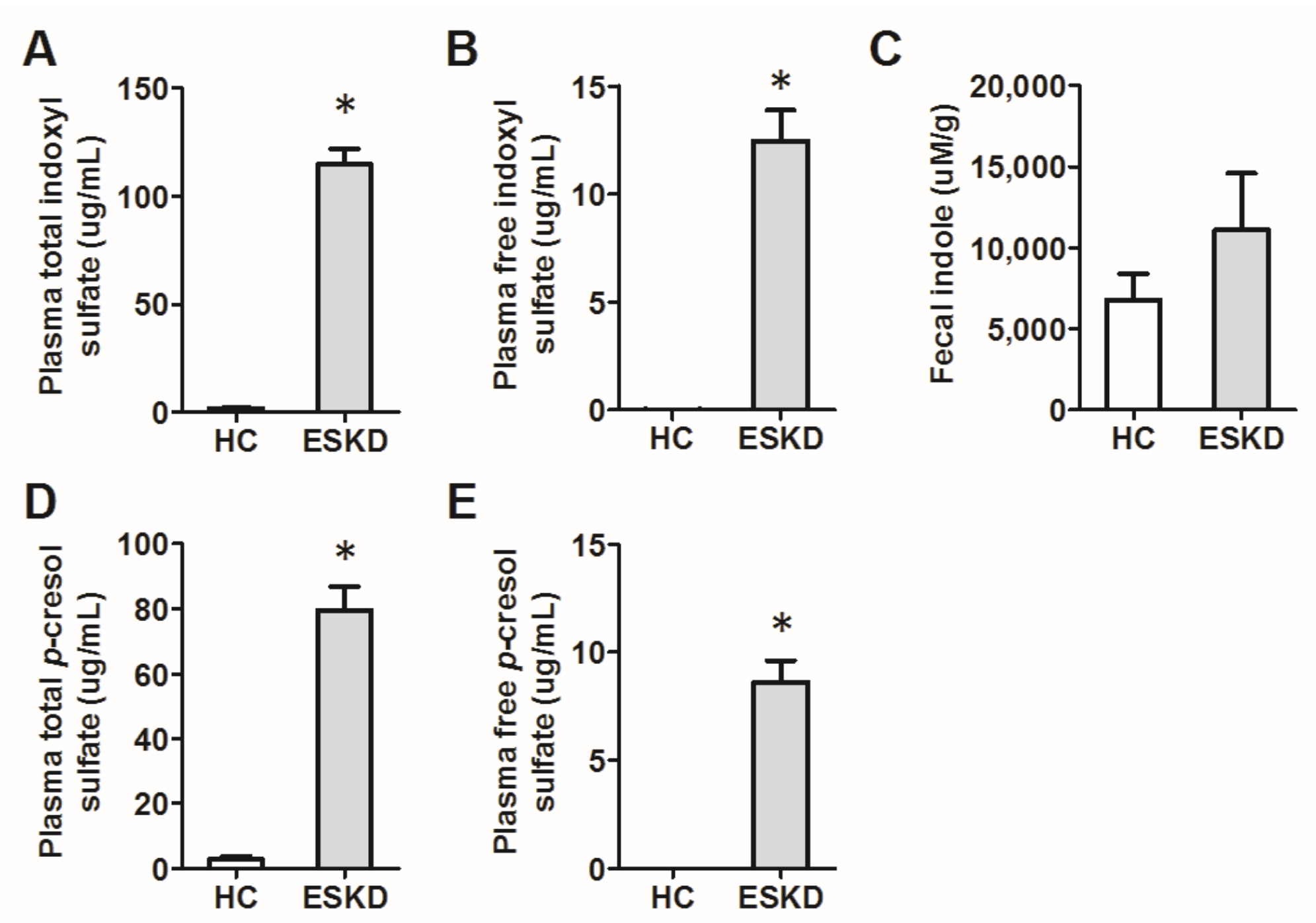
3.1. Human Samples
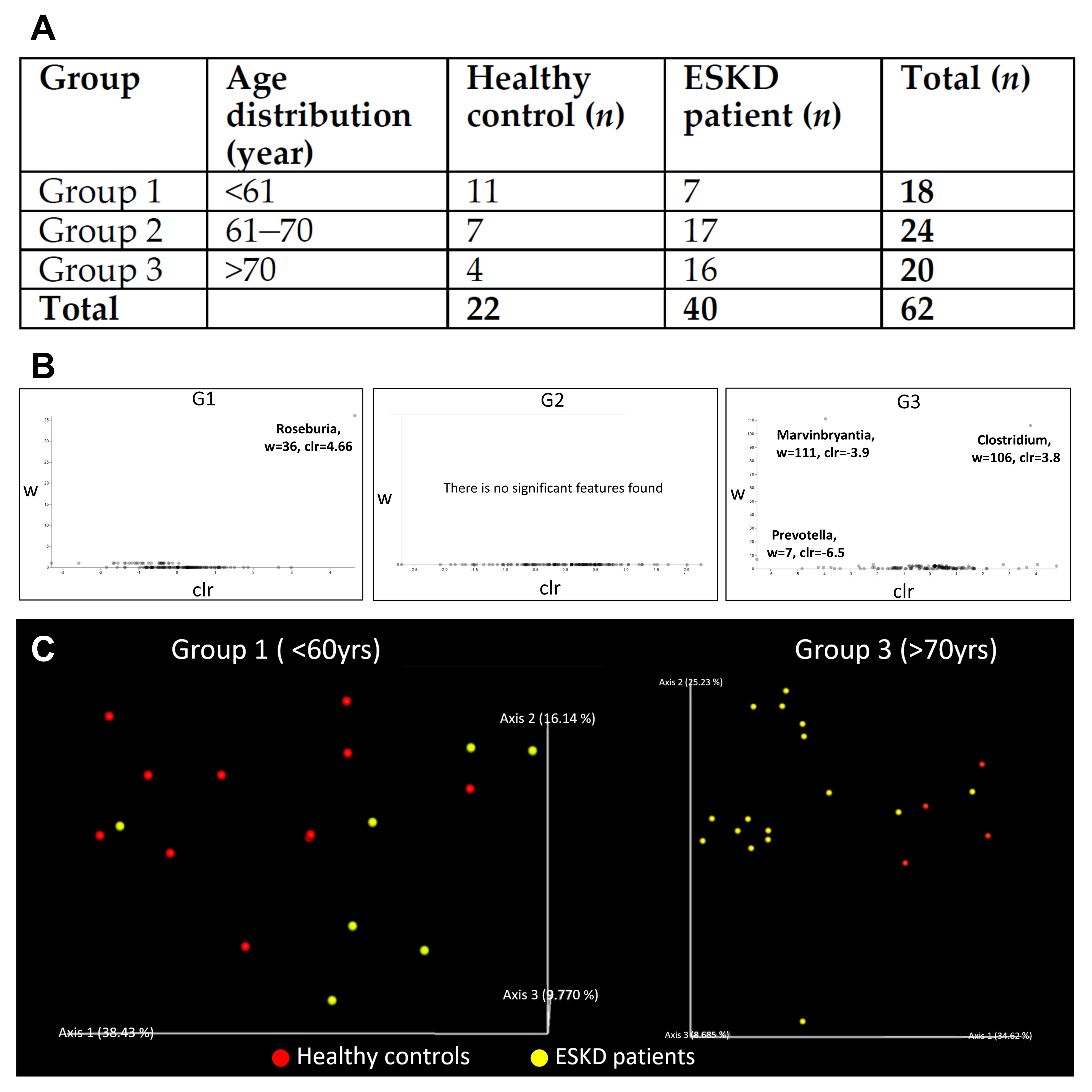
3.1.1. Demographic Characteristics of Human Study Participants
3.1.2. Characteristics of the 16s Microbiome in ESKD Patients
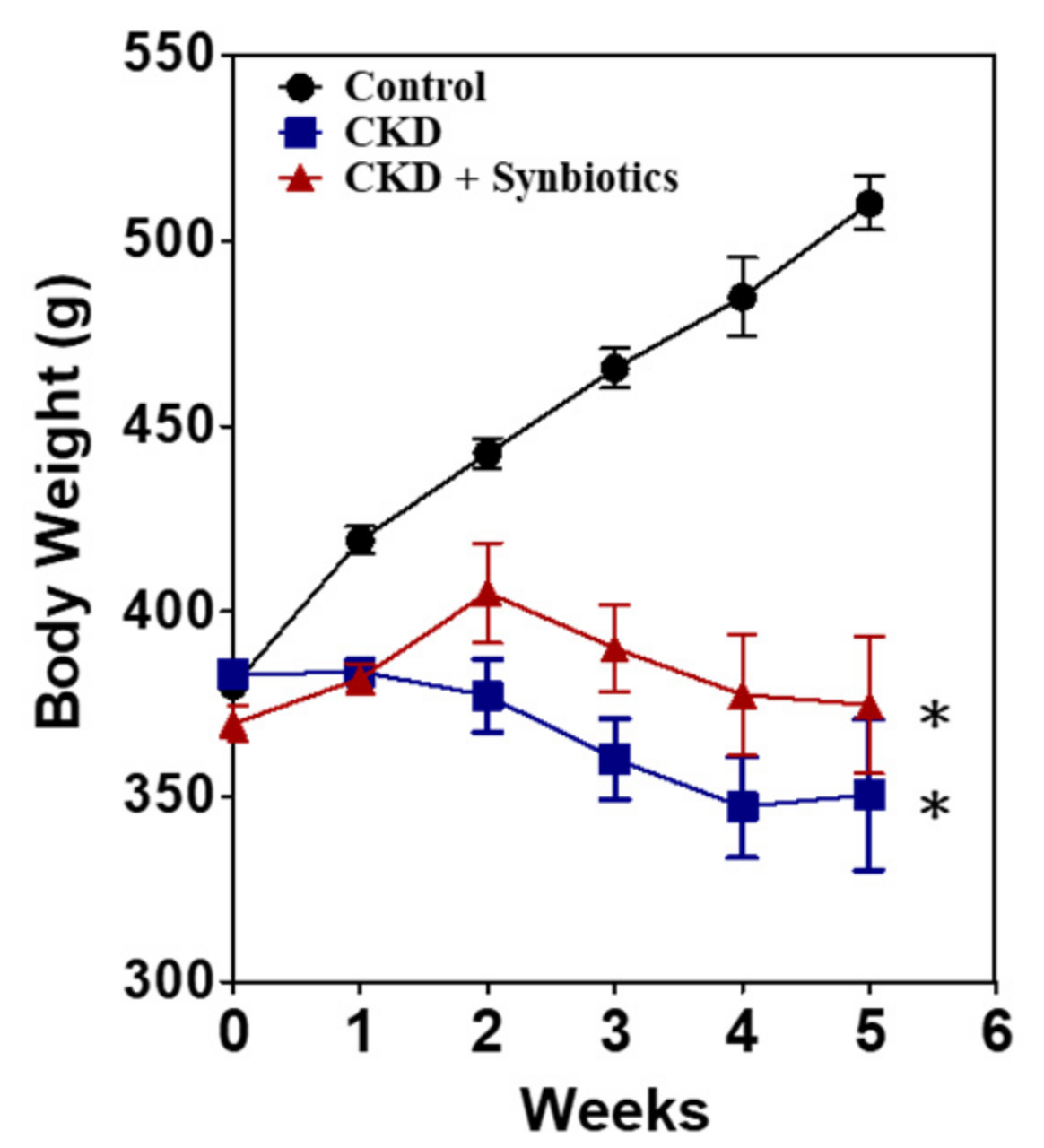
3.2. The Animal Model 1: Concomitant Five-Week Adenine Diet and Five-Week Synbiotic Treatment
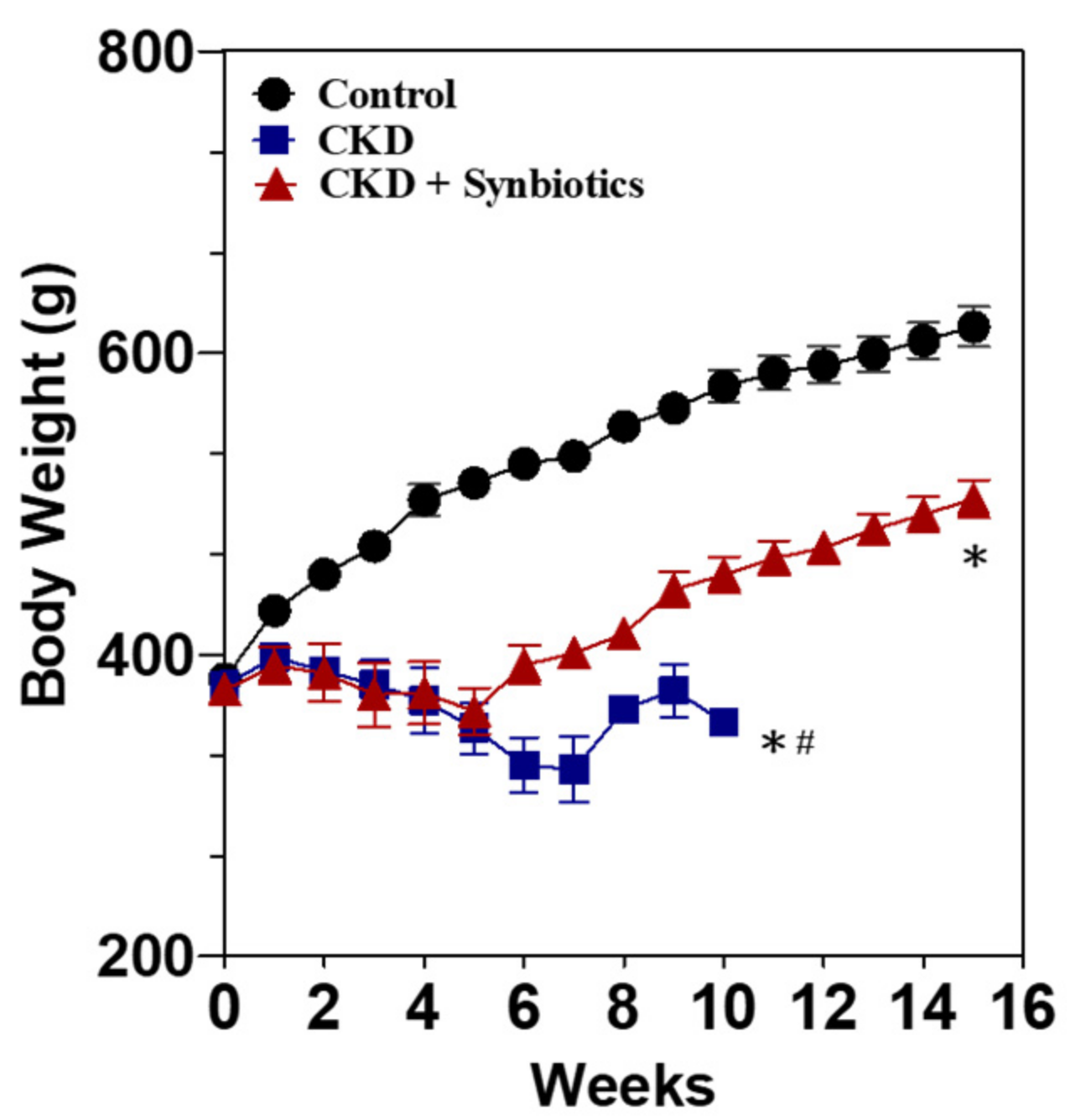
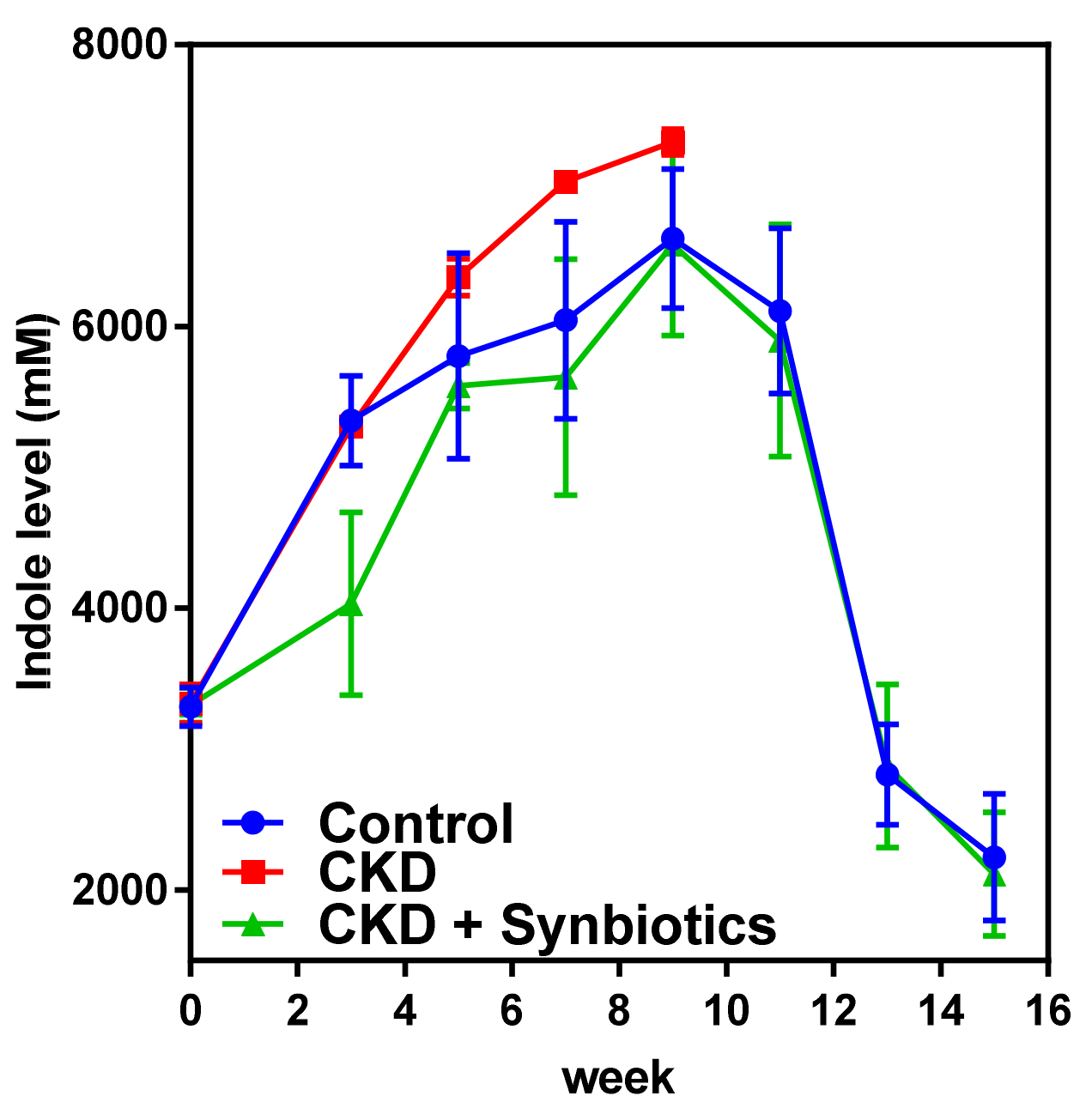
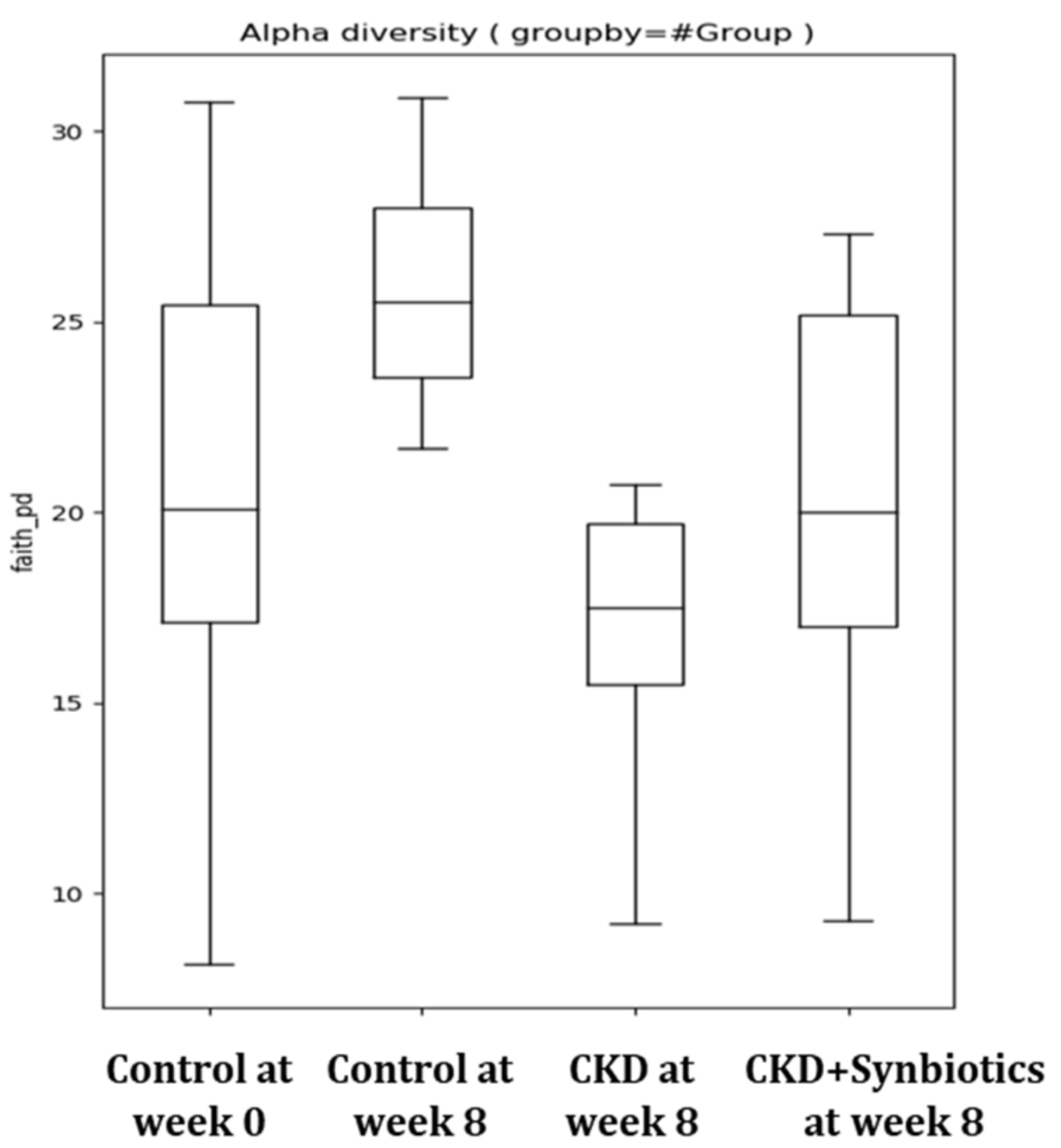
3.3. The Animal Model 2: Five-Week Adenine Diet Followed by 10-Week Synbiotic Treatment
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Wen, C.P.; Cheng, T.Y.; Tsai, M.K.; Chang, Y.C.; Chan, H.T.; Tsai, S.P.; Chiang, P.H.; Hsu, C.C.; Sung, P.K.; Hsu, Y.H.; et al. All-cause mortality attributable to chronic kidney disease: A prospective cohort study based on 462 293 adults in Taiwan. Lancet 2008, 371, 2173–2182. [Google Scholar] [CrossRef]
- Jazani, N.H.; Savoj, J.; Lustgarten, M.; Lau, W.L.; Vaziri, N.D. Impact of Gut Dysbiosis on Neurohormonal Pathways in Chronic Kidney Disease. Diseases 2019, 7, 21. [Google Scholar] [CrossRef] [PubMed]
- Mafra, D.; Borges, N.; Alvarenga, L.; Esgalhado, M.; Cardozo, L.; Lindholm, B.; Stenvinkel, P. Dietary Components That May Influence the Disturbed Gut Microbiota in Chronic Kidney Disease. Nutrients 2019, 11, 496. [Google Scholar] [CrossRef] [PubMed]
- Evenepoel, P.; Dejongh, S.; Verbeke, K.; Meijers, B. The Role of Gut Dysbiosis in the Bone-Vascular Axis in Chronic Kidney Disease. Toxins (Basel) 2020, 12, 285. [Google Scholar] [CrossRef]
- Pahl, M.V.; Vaziri, N.D. The Chronic Kidney Disease—Colonic Axis. Semin. Dial. 2015, 28, 459–463. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.Y.; Tarng, D.C. Diet, gut microbiome and indoxyl sulphate in chronic kidney disease patients. Nephrology 2018, 23 (Suppl. 4), 16–20. [Google Scholar] [CrossRef]
- Hung, S.C.; Kuo, K.L.; Huang, H.L.; Lin, C.C.; Tsai, T.H.; Wang, C.H.; Chen, J.W.; Lin, S.J.; Huang, P.H.; Tarng, D.C. Indoxyl sulfate suppresses endothelial progenitor cell-mediated neovascularization. Kidney Int. 2016, 89, 574–585. [Google Scholar] [CrossRef]
- Wu, C.C.; Hsieh, M.Y.; Hung, S.C.; Kuo, K.L.; Tsai, T.H.; Lai, C.L.; Chen, J.W.; Lin, S.J.; Huang, P.H.; Tarng, D.C. Serum Indoxyl Sulfate Associates with Postangioplasty Thrombosis of Dialysis Grafts. J. Am. Soc. Nephrol. 2016, 27, 1254–1264. [Google Scholar] [CrossRef]
- Lee, J.H.; Lee, J. Indole as an intercellular signal in microbial communities. FEMS Microbiol. Rev. 2010, 34, 426–444. [Google Scholar] [CrossRef]
- Morgan, X.C.; Huttenhower, C. Chapter 12: Human Microbiome Analysis. PLoS Comput. Biol. 2012, 8, e1002808. [Google Scholar] [CrossRef]
- Cox, M.J.; Cookson, W.O.C.M.; Moffatt, M.F. Sequencing the human microbiome in health and disease. Hum. Mol. Genet. 2013, 22, R88–R94. [Google Scholar] [CrossRef]
- Krishnan, K.; Chen, T.; Paster, B.J. A practical guide to the oral microbiome and its relation to health and disease. Oral Dis. 2017, 23, 276–286. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Sun, J.; Lin, C.-C.; Abemayor, E.; Wang, M.B.; Wong, D.T.W. The emerging landscape of salivary diagnostics. Periodontology 2000 2016, 70, 38–52. [Google Scholar] [CrossRef] [PubMed]
- Ramezani, A.; Massy, Z.A.; Meijers, B.; Evenepoel, P.; Vanholder, R.; Raj, D.S. Role of the Gut Microbiome in Uremia: A Potential Therapeutic Target. Am. J. Kidney Dis. 2016, 67, 483–498. [Google Scholar] [CrossRef] [PubMed]
- Rossi, M.; Klein, K.; Johnson, D.W.; Campbell, K.L. Pre-, pro-, and synbiotics: Do they have a role in reducing uremic toxins? A systematic review and meta-analysis. Int. J. Nephrol. 2012, 2012, 673631. [Google Scholar] [CrossRef] [PubMed]
- Nakabayashi, I.; Nakamura, M.; Kawakami, K.; Ohta, T.; Kato, I.; Uchida, K.; Yoshida, M. Effects of synbiotic treatment on serum level of p-cresol in haemodialysis patients: A preliminary study. Nephrol. Dial. Transplant. 2011, 26, 1094–1098. [Google Scholar] [CrossRef] [PubMed]
- De Preter, V.; Vanhoutte, T.; Huys, G.; Swings, J.; De Vuyst, L.; Rutgeerts, P.; Verbeke, K. Effects of Lactobacillus casei Shirota, Bifidobacterium breve, and oligofructose-enriched inulin on colonic nitrogen-protein metabolism in healthy humans. Am. J. Physiol. Gastrointest. Liver Physiol. 2007, 292, G358–G368. [Google Scholar] [CrossRef]
- De Preter, V.; Raemen, H.; Cloetens, L.; Houben, E.; Rutgeerts, P.; Verbeke, K. Effect of dietary intervention with different pre- and probiotics on intestinal bacterial enzyme activities. Eur. J. Clin. Nutr. 2008, 62, 225–231. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Pena, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods 2010, 7, 335–336. [Google Scholar] [CrossRef]
- Diwan, V.; Small, D.; Kauter, K.; Gobe, G.C.; Brown, L. Gender differences in adenine-induced chronic kidney disease and cardiovascular complications in rats. Am. J. Physiol. Renal Physiol. 2014, 307, F1169–F1178. [Google Scholar] [CrossRef]
- Diwan, V.; Mistry, A.; Gobe, G.; Brown, L. Adenine-induced chronic kidney and cardiovascular damage in rats. J. Pharmacol. Toxicol. Methods 2013, 68, 197–207. [Google Scholar] [CrossRef] [PubMed]
- Diwan, V.; Brown, L.; Gobe, G.C. Adenine-induced chronic kidney disease in rats. Nephrology 2018, 23, 5–11. [Google Scholar] [CrossRef] [PubMed]
- Sueyoshi, M.; Fukunaga, M.; Mei, M.; Nakajima, A.; Tanaka, G.; Murase, T.; Narita, Y.; Hirata, S.; Kadowaki, D. Effects of lactulose on renal function and gut microbiota in adenine-induced chronic kidney disease rats. Clin. Exp. Nephrol. 2019, 23, 908–919. [Google Scholar] [CrossRef] [PubMed]
- Pavan, M. Influence of prebiotic and probiotic supplementation on the progression of chronic kidney disease. Minerva Urol. Nefrol. 2016, 68, 222–226. [Google Scholar]
- Jia, T.; Olauson, H.; Lindberg, K.; Amin, R.; Edvardsson, K.; Lindholm, B.; Andersson, G.; Wernerson, A.; Sabbagh, Y.; Schiavi, S.; et al. A novel model of adenine-induced tubulointerstitial nephropathy in mice. BMC Nephrol. 2013, 14, 116. [Google Scholar] [CrossRef]
- Yang, C.; Liu, C.; Zhou, Q.; Xie, Y.C.; Qiu, X.M.; Feng, X. Effect of atracylodes rhizome polysaccharide in rats with adenine-induced chronic renal failure. Indian J. Pharm. Sci. 2015, 77, 103–107. [Google Scholar] [CrossRef]
- Neven, E.; D’Haese, P.C. Vascular calcification in chronic renal failure: What have we learned from animal studies? Circ. Res. 2011, 108, 249–264. [Google Scholar] [CrossRef]
- Yoshifuji, A.; Wakino, S.; Irie, J.; Tajima, T.; Hasegawa, K.; Kanda, T.; Tokuyama, H.; Hayashi, K.; Itoh, H. Gut Lactobacillus protects against the progression of renal damage by modulating the gut environment in rats. Nephrol. Dial. Transplant. 2016, 31, 401–412. [Google Scholar] [CrossRef]
- Ranganathan, N.; Friedman, E.A.; Tam, P.; Rao, V.; Ranganathan, P.; Dheer, R. Probiotic dietary supplementation in patients with stage 3 and 4 chronic kidney disease: A 6-month pilot scale trial in Canada. Curr. Med. Res. Opin. 2009, 25, 1919–1930. [Google Scholar] [CrossRef]
- Viramontes-Horner, D.; Marquez-Sandoval, F.; Martin-del-Campo, F.; Vizmanos-Lamotte, B.; Sandoval-Rodriguez, A.; Armendariz-Borunda, J.; Garcia-Bejarano, H.; Renoirte-Lopez, K.; Garcia-Garcia, G. Effect of a symbiotic gel (Lactobacillus acidophilus + Bifidobacterium lactis + inulin) on presence and severity of gastrointestinal symptoms in hemodialysis patients. J. Ren. Nutr. 2015, 25, 284–291. [Google Scholar] [CrossRef]
- Meijers, B.K.; De Preter, V.; Verbeke, K.; Vanrenterghem, Y.; Evenepoel, P. p-Cresyl sulfate serum concentrations in haemodialysis patients are reduced by the prebiotic oligofructose-enriched inulin. Nephrol. Dial. Transplant. 2010, 25, 219–224. [Google Scholar] [CrossRef] [PubMed]
- Darkoh, C.; Chappell, C.; Gonzales, C.; Okhuysen, P. A rapid and specific method for the detection of indole in complex biological samples. Appl. Environ. Microbiol. 2015, 81, 8093–8097. [Google Scholar] [CrossRef] [PubMed]
- Tap, J.; Mondot, S.; Levenez, F.; Pelletier, E.; Caron, C.; Furet, J.P.; Ugarte, E.; Munoz-Tamayo, R.; Paslier, D.L.; Nalin, R.; et al. Towards the human intestinal microbiota phylogenetic core. Environ. Microbiol. 2009, 11, 2574–2584. [Google Scholar] [CrossRef] [PubMed]
- Cummings, J.H. Fermentation in the human large intestine: Evidence and implications for health. Lancet 1983, 1, 1206–1209. [Google Scholar] [CrossRef]
- Wu, I.W.; Hsu, K.H.; Lee, C.C.; Sun, C.Y.; Hsu, H.J.; Tsai, C.J.; Tzen, C.Y.; Wang, Y.C.; Lin, C.Y.; Wu, M.S. p-Cresyl sulphate and indoxyl sulphate predict progression of chronic kidney disease. Nephrol. Dial. Transplant. 2011, 26, 938–947. [Google Scholar] [CrossRef]
- Lin, C.J.; Chen, H.H.; Pan, C.F.; Chuang, C.K.; Wang, T.J.; Sun, F.J.; Wu, C.J. p-Cresylsulfate and indoxyl sulfate level at different stages of chronic kidney disease. J. Clin. Lab. Anal. 2011, 25, 191–197. [Google Scholar] [CrossRef]
- Lysaght, M.J.; Vonesh, E.F.; Gotch, F.; Ibels, L.; Keen, M.; Lindholm, B.; Nolph, K.D.; Pollock, C.A.; Prowant, B.; Farrell, P.C. The influence of dialysis treatment modality on the decline of remaining renal function. ASAIO Trans. 1991, 37, 598–604. [Google Scholar]
- Motojima, M.; Hosokawa, A.; Yamato, H.; Muraki, T.; Yoshioka, T. Uraemic toxins induce proximal tubular injury via organic anion transporter 1-mediated uptake. Br. J. Pharmacol. 2002, 135, 555–563. [Google Scholar] [CrossRef]
- Motojima, M.; Hosokawa, A.; Yamato, H.; Muraki, T.; Yoshioka, T. Uremic toxins of organic anions up-regulate PAI-1 expression by induction of NF-kappaB and free radical in proximal tubular cells. Kidney Int. 2003, 63, 1671–1680. [Google Scholar] [CrossRef]
- Liang, C.; Tseng, H.C.; Chen, H.M.; Wang, W.C.; Chiu, C.M.; Chang, J.Y.; Lu, K.Y.; Weng, S.L.; Chang, T.H.; Chang, C.H.; et al. Diversity and enterotype in gut bacterial community of adults in Taiwan. BMC Genom. 2017, 18, 932. [Google Scholar] [CrossRef]
- Group, N.H.W.; Peterson, J.; Garges, S.; Giovanni, M.; McInnes, P.; Wang, L.; Schloss, J.A.; Bonazzi, V.; McEwen, J.E.; Wetterstrand, K.A.; et al. The NIH Human Microbiome Project. Genome Res. 2009, 19, 2317–2323. [Google Scholar] [CrossRef] [PubMed]
- Korpela, K.; Flint, H.J.; Johnstone, A.M.; Lappi, J.; Poutanen, K.; Dewulf, E.; Delzenne, N.; de Vos, W.M.; Salonen, A. Gut microbiota signatures predict host and microbiota responses to dietary interventions in obese individuals. PLoS ONE 2014, 9, e90702. [Google Scholar] [CrossRef] [PubMed]
- Qin, J.; Li, Y.; Cai, Z.; Li, S.; Zhu, J.; Zhang, F.; Liang, S.; Zhang, W.; Guan, Y.; Shen, D.; et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 2012, 490, 55–60. [Google Scholar] [CrossRef] [PubMed]
- Krishna Prasad, R.B.; Sharma, A.; Babu, H.M. An insight into salivary markers in oral cancer. Dent. Res. J. 2013, 10, 287–295. [Google Scholar]
- Wang, X.; Kaczor-Urbanowicz, K.E.; Wong, D.T.W. Salivary biomarkers in cancer detection. Med. Oncol. 2016, 34, 7. [Google Scholar] [CrossRef]
- He, J.; Li, Y.; Cao, Y.; Xue, J.; Zhou, X. The oral microbiome diversity and its relation to human diseases. Folia Microbiol. 2015, 60, 69–80. [Google Scholar] [CrossRef]
- Lun, H.; Yang, W.; Zhao, S.; Jiang, M.; Xu, M.; Liu, F.; Wang, Y. Altered gut microbiota and microbial biomarkers associated with chronic kidney disease. Microbiologyopen 2018, 8, e00678. [Google Scholar] [CrossRef]
- Nallu, A.; Sharma, S.; Ramezani, A.; Muralidharan, J.; Raj, D. Gut microbiome in chronic kidney disease: Challenges and opportunities. Transl. Res. 2017, 179, 24–37. [Google Scholar] [CrossRef]
- Machiels, K.; Joossens, M.; Sabino, J.; De Preter, V.; Arijs, I.; Eeckhaut, V.; Ballet, V.; Claes, K.; Van Immerseel, F.; Verbeke, K.; et al. A decrease of the butyrate-producing species Roseburia hominis and Faecalibacterium prausnitzii defines dysbiosis in patients with ulcerative colitis. Gut 2014, 63, 1275–1283. [Google Scholar] [CrossRef]
- Rios-Covian, D.; Ruas-Madiedo, P.; Margolles, A.; Gueimonde, M.; de Los Reyes-Gavilan, C.G.; Salazar, N. Intestinal Short Chain Fatty Acids and their Link with Diet and Human Health. Front. Microbiol. 2016, 7, 185. [Google Scholar] [CrossRef]
- Morrison, D.J.; Preston, T. Formation of short chain fatty acids by the gut microbiota and their impact on human metabolism. Gut Microbes 2016, 7, 189–200. [Google Scholar] [CrossRef] [PubMed]
- Parada Venegas, D.; De la Fuente, M.K.; Landskron, G.; Gonzalez, M.J.; Quera, R.; Dijkstra, G.; Harmsen, H.J.M.; Faber, K.N.; Hermoso, M.A. Short Chain Fatty Acids (SCFAs)-Mediated Gut Epithelial and Immune Regulation and Its Relevance for Inflammatory Bowel Diseases. Front. Immunol. 2019, 10, 277. [Google Scholar] [CrossRef] [PubMed]
- Meijers, B.; Farre, R.; Dejongh, S.; Vicario, M.; Evenepoel, P. Intestinal Barrier Function in Chronic Kidney Disease. Toxins (Basel) 2018, 10, 298. [Google Scholar] [CrossRef] [PubMed]

| Healthy Controls | ESKD Patients | Total | |
|---|---|---|---|
| Patient number | 22 | 40 | 62 |
| Age (year) | 62.3 ± 10.0 | 69.5 ± 13.2 | 68.6 ± 13.0 |
| Male gender (%) | 50 | 50 | 50 |
| Healthy Controls | CKD Patients | ||
|---|---|---|---|
| Top 10 Genus | Percentage | Top 10 Genus | Percentage |
| Bacteroides | 50.85% | Bacteroides | 49.15% |
| Prevotella | 7.51% | Prevotella | 5.54% |
| Clostridium | 4.38% | Clostridium | 3.76% |
| Faecalibacterium | 3.96% | Streptococcus | 3.45% |
| Roseburia | 2.89% | Alistipes | 3.29% |
| Ruminococcus | 2.78% | Parabacteroides | 3.12% |
| Alistipes | 2.73% | Succinispira | 3.05% |
| Megamonas | 2.43% | Ruminococcus | 2.94% |
| Succinispira | 2.30% | Faecalibacterium | 2.73% |
| Arcobacter | 2.21% | Dorea | 2.62% |
| Animal Model 1 | ||||
|---|---|---|---|---|
| Serum Test | Week | Control | CKD | CKD + Synbiotics |
| Blood urea nitrogen (mg/dL) | 0 | 14.2 ± 1.3 | 15.4 ± 0.7 | 15.2 ± 0.4 |
| 3 | 14.1 ± 1.5 | 108.5 ± 0.9 * | 88.4 ± 6.7 *,# | |
| 5 | 15.9 ± 0.6 | 214.0 ± 20.0 * | 124.9 ± 1.0 *,# | |
| Serum creatinine (mg/dL) | 0 | 0.2 ± 0.0 | 0.2 ± 0.1 | 0.3 ± 0.2 |
| 3 | 0.3 ± 0.1 | 2.8 ± 0.3 * | 2.1 ± 0.2 | |
| 5 | 0.4 ± 0.1 | 7.8 ± 0.6 * | 3.6 ± 0.4 *,# | |
| Animal Model 2 | ||||
|---|---|---|---|---|
| Serum Test | Week | Control | CKD | CKD + Synbiotics |
| Blood urea nitrogen (mg/dL) | 0 | 15.8 ± 0.2 | 15.1 ± 0.4 | 15.7 ± 0.4 |
| 3 | 16.0 ± 1.0 | 110.1 ± 11.0 * | 109.7 ± 7.4 * | |
| 5 | 17.8 ± 0.6 | 214.2 ± 21.8 * | 201.2 ± 26.6 * | |
| 7 | 17.8 ± 0.8 | 191.2 ± 20.4 * | 100.2 ± 13.4 *,# | |
| 9 | 16.0 ± 0.5 | 190.4 ± 0 * | 87.0 ± 11.2 *# | |
| 11 | 17.4 ± 1.0 | - | 83.3 ± 5.9 * | |
| 13 | 15.6 ± 0.9 | - | 80.0 ± 6.9 * | |
| 15 | 14.9 ± 1.0 | - | 77.3 ± 9.7 * | |
| Serum creatinine (mg/dL) | 0 | 0.2 ± 0 | 0.2 ± 0 | 0.3 ± 0.1 |
| 3 | 0.3 ± 0.1 | 2.2 ± 0.5 * | 2.4 ± 0.3 * | |
| 5 | 0.3 ± 0.1 | 7.1 ± 0.8 * | 7.3 ± 1.6 * | |
| 7 | 0.3 ± 0.1 | 5.5 ± 1.5 * | 3.3 ± 0.5 *,# | |
| 9 | 0.3 ± 0.1 | 4.2 ± 0 * | 1.7 ± 0.3 *,# | |
| 11 | 0.3 ± 0.1 | - | 1.4 ± 0.1 | |
| 13 | 0.3 ± 0.1 | - | 1.3 ± 0.2 | |
| 15 | 0.3 ± 0 | - | 1.3 ± 0.2 | |
| Control | CKD | CKD + Synbiotics | |||
|---|---|---|---|---|---|
| Top 10 Genus | Top 10 Genus | Top 10 Genus | |||
| Ruminococcus | 20.26% | Parabacteroides | 19.52% | Parabacteroides | 18.36% |
| Parabacteroides | 16.16% | Clostridium | 19.22% | Lactobacillus | 16.70% |
| Lactobacillus | 15.50% | Ruminococcus | 17.07% | Ruminococcus | 12.29% |
| Clostridium | 10.07% | Lactobacillus | 11.27% | Clostridium | 10.84% |
| Marvinbryantia | 9.75% | Akkermansia | 8.11% | Allobaculum | 9.60% |
| Prevotella | 8.37% | Anaeroplasma | 5.86% | Alistipes | 8.44% |
| Alistipes | 4.31% | Marvinbryantia | 4.14% | Anaeroplasma | 8.23% |
| Sporobacter | 4.24% | Alistipes | 4.00% | Bifidobacterium | 6.14% |
| Adlercreutzia | 2.15% | Sporobacter | 1.62% | Marvinbryantia | 3.56% |
| Bifidobacterium | 1.54% | Bifidobacterium | 1.13% | rc4-4 | 1.71% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, C.-Y.; Chen, T.-W.; Lu, W.-L.; Liang, S.-S.; Huang, H.-D.; Tseng, C.-P.; Tarng, D.-C. Synbiotics Alleviate the Gut Indole Load and Dysbiosis in Chronic Kidney Disease. Cells 2021, 10, 114. https://doi.org/10.3390/cells10010114
Yang C-Y, Chen T-W, Lu W-L, Liang S-S, Huang H-D, Tseng C-P, Tarng D-C. Synbiotics Alleviate the Gut Indole Load and Dysbiosis in Chronic Kidney Disease. Cells. 2021; 10(1):114. https://doi.org/10.3390/cells10010114
Chicago/Turabian StyleYang, Chih-Yu, Ting-Wen Chen, Wan-Lun Lu, Shih-Shin Liang, Hsien-Da Huang, Ching-Ping Tseng, and Der-Cherng Tarng. 2021. "Synbiotics Alleviate the Gut Indole Load and Dysbiosis in Chronic Kidney Disease" Cells 10, no. 1: 114. https://doi.org/10.3390/cells10010114
APA StyleYang, C.-Y., Chen, T.-W., Lu, W.-L., Liang, S.-S., Huang, H.-D., Tseng, C.-P., & Tarng, D.-C. (2021). Synbiotics Alleviate the Gut Indole Load and Dysbiosis in Chronic Kidney Disease. Cells, 10(1), 114. https://doi.org/10.3390/cells10010114





