Actinobacteria Structure in Autogenic, Hydrogenic and Lithogenic Cultivated and Non-Cultivated Soils: A Culture-Independent Approach
Abstract
1. Introduction
2. Materials and Methods
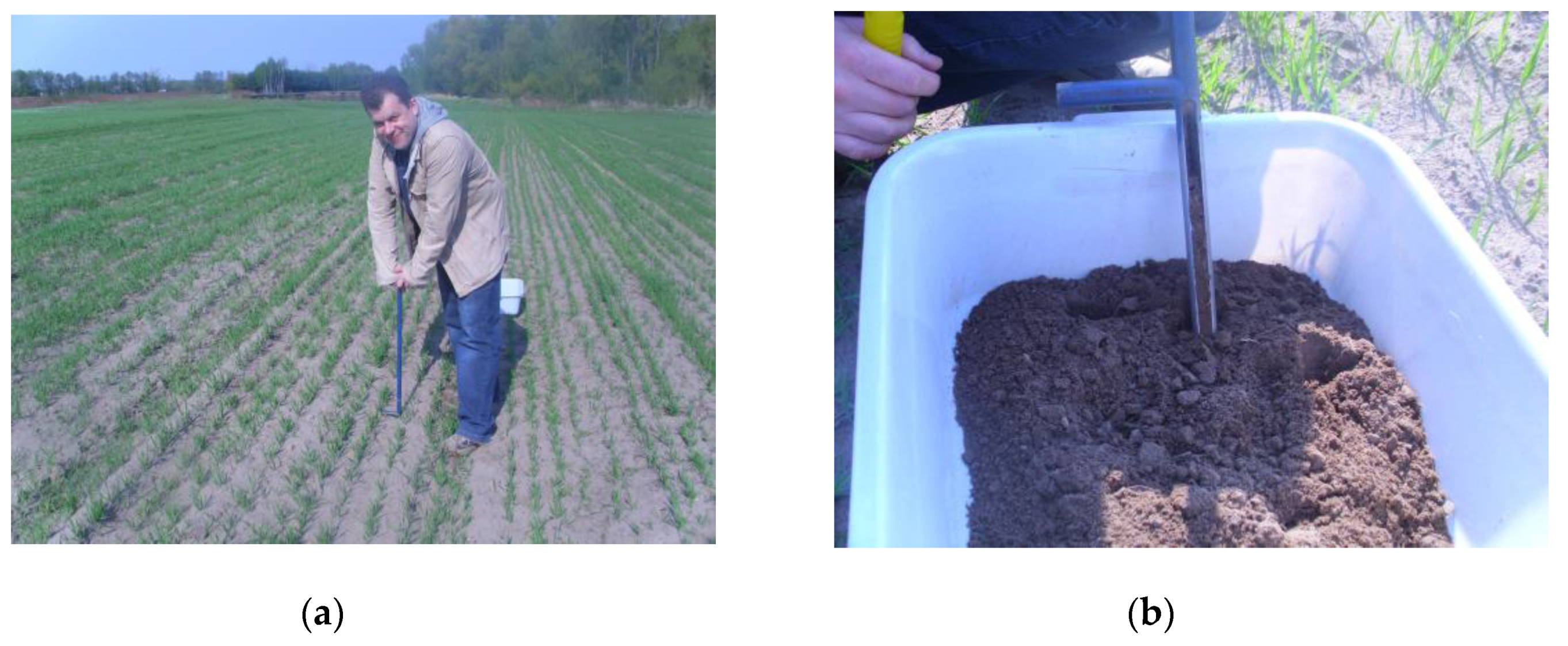
2.1. Soil Sampling
2.2. Soil Chemical Characteristics
2.3. DNA extraction and Next Generation Sequencing
2.4. Bioinformatic Analysis
2.5. Denaturing Gradient Gel Electrophoresis
2.6. Statistical Analysis and Biodiversity Indices Calculation
3. Results
3.1. Chemical Characteristics of Autogenic, Hydrogenic and Lithogenic Soils
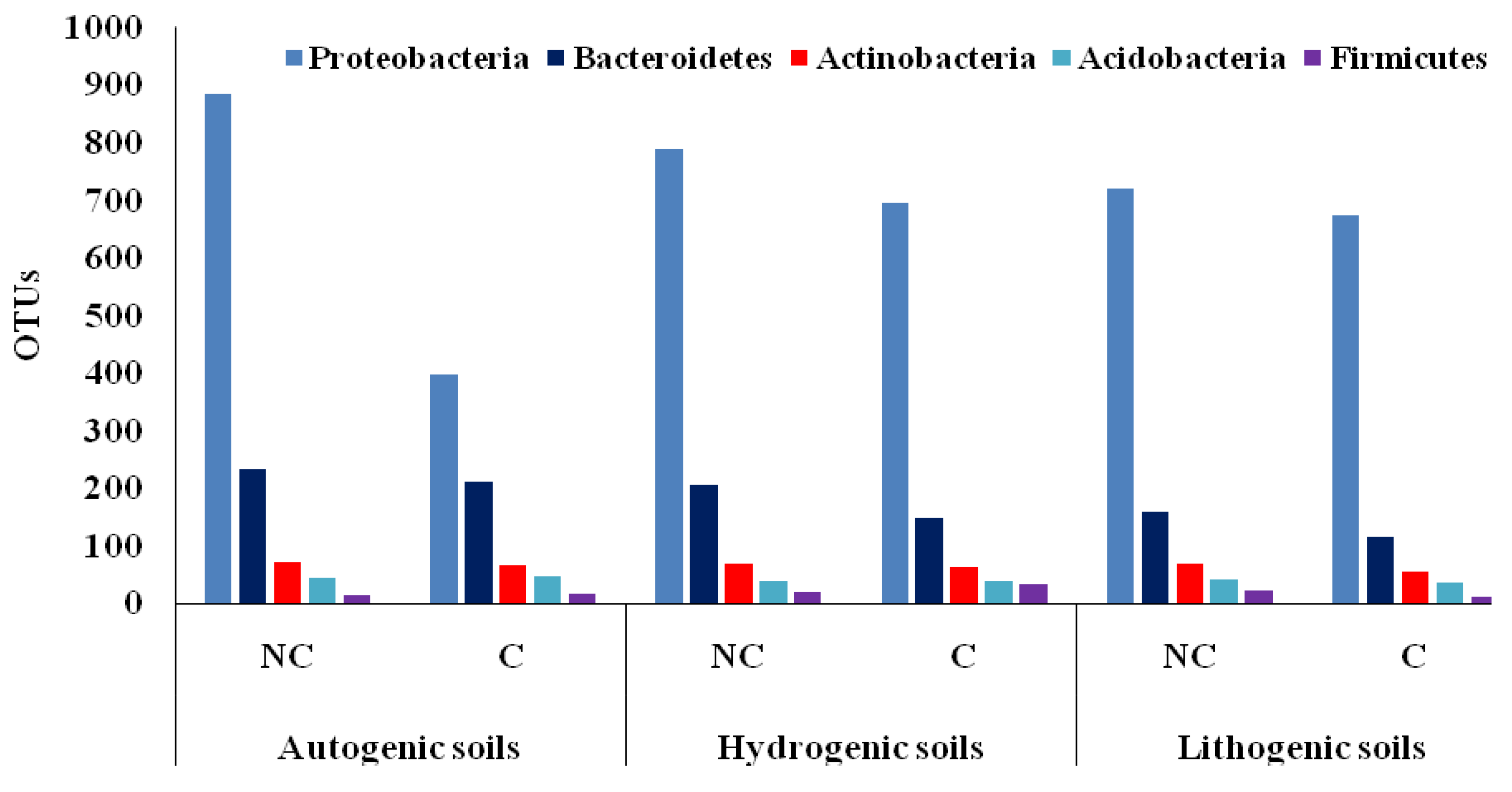
3.2. Biodiversity in Cultivated and Non-Cultivated Soils at the Phylum Level
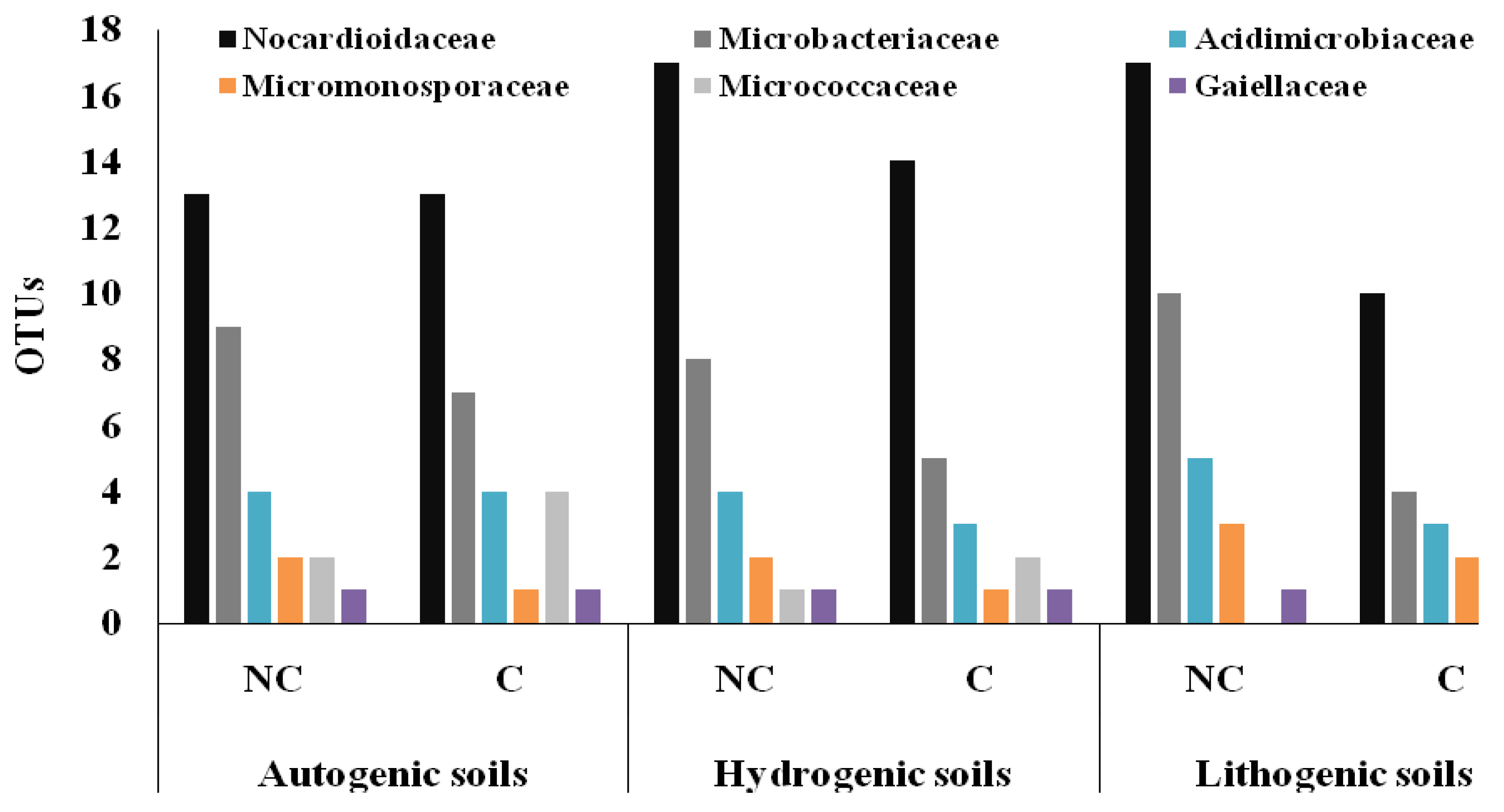
3.3. Structure of Actinobacteria in Arable and Non-Cultivated Soils at the Family Level
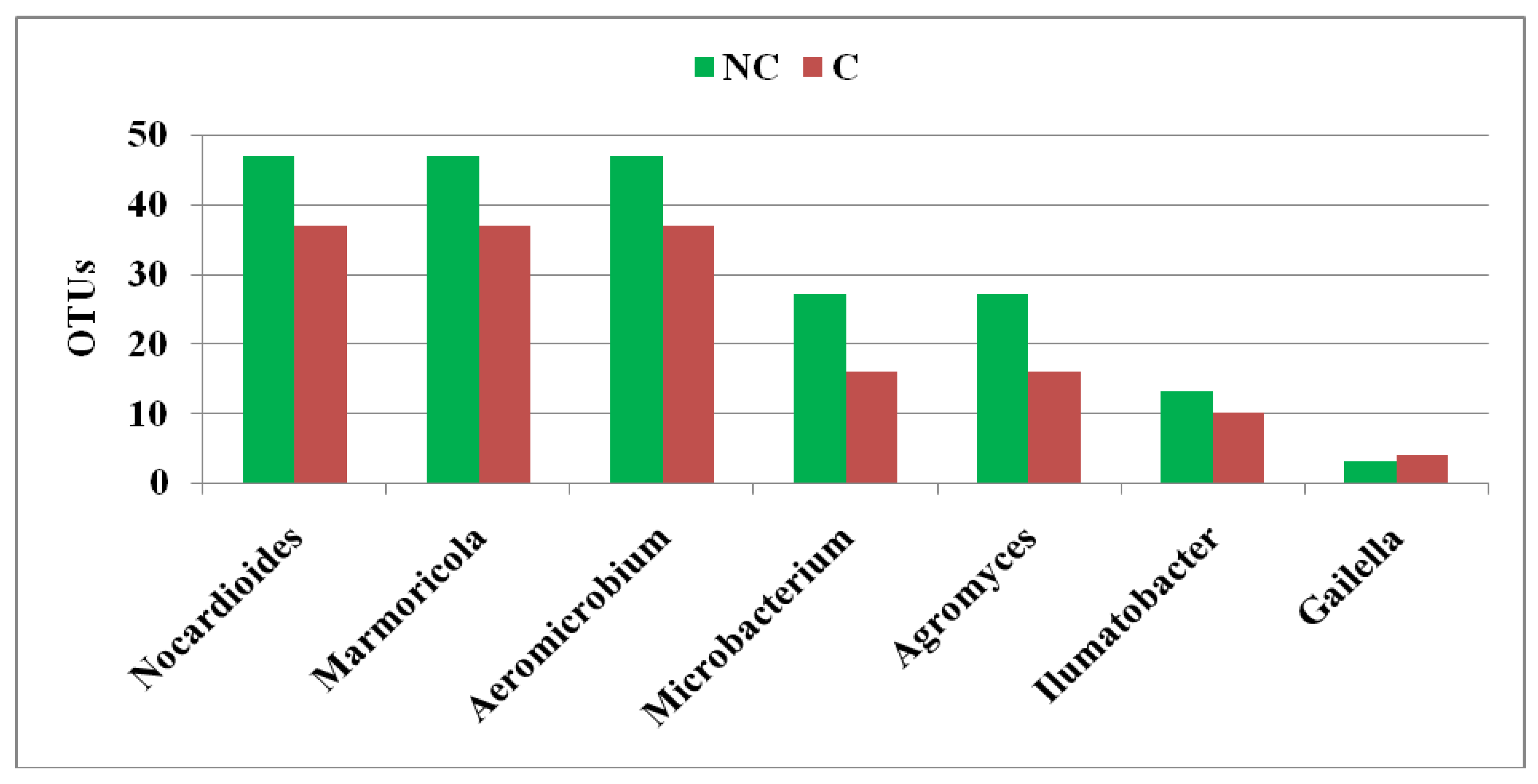
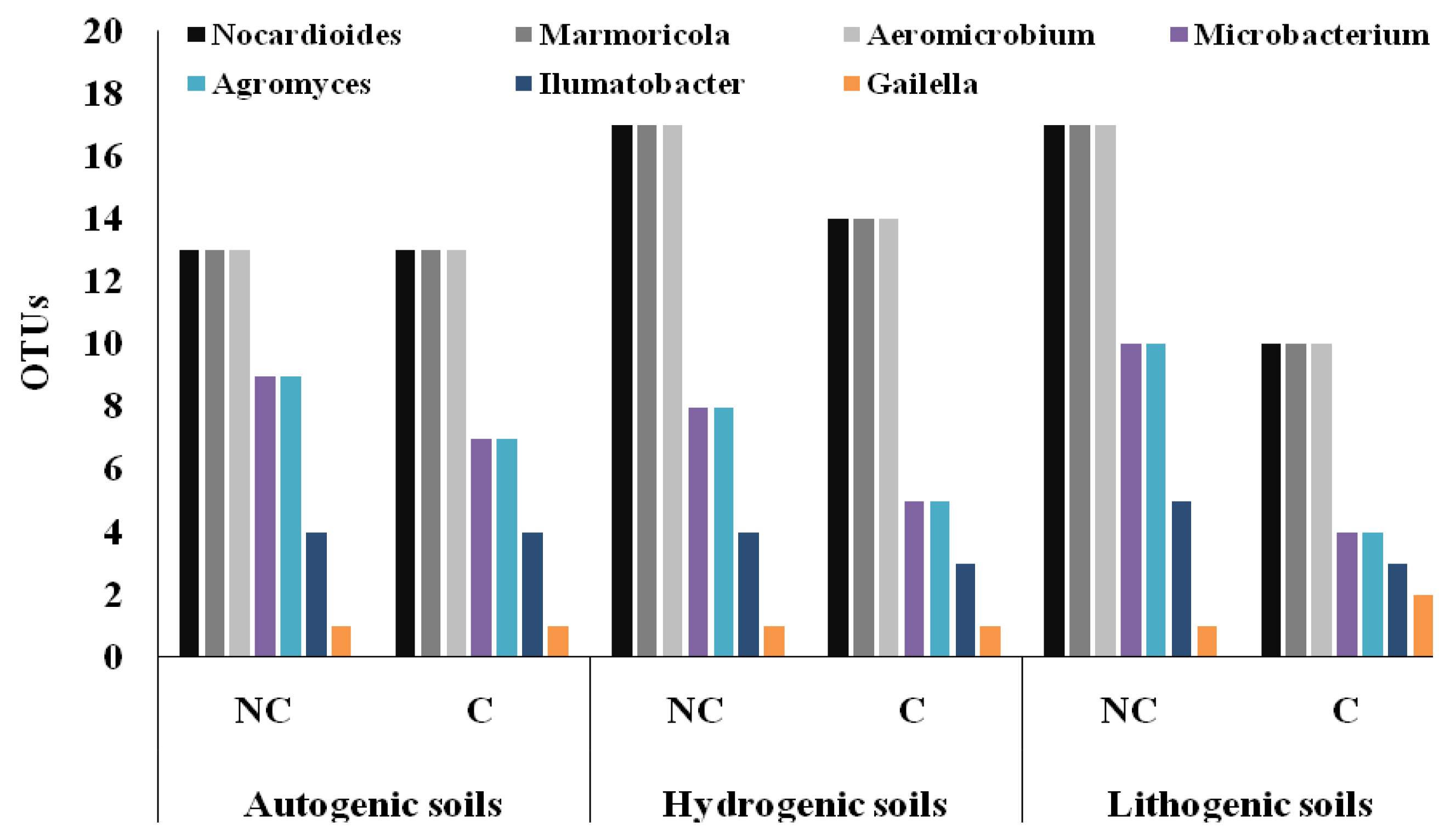
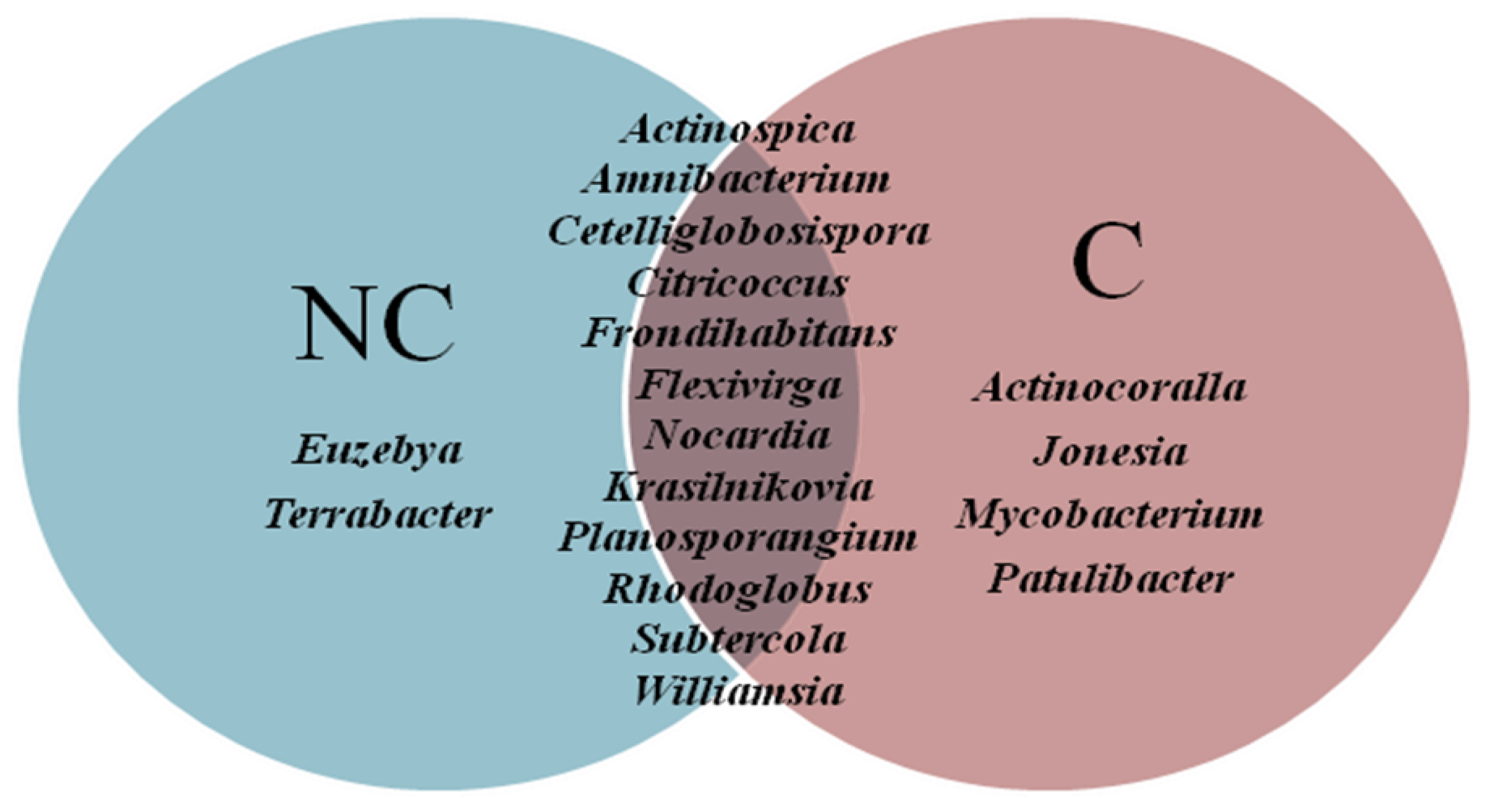
3.4. Structure of Actinobacteria in Arable and Non-Cultivated Soils at the Genus Level
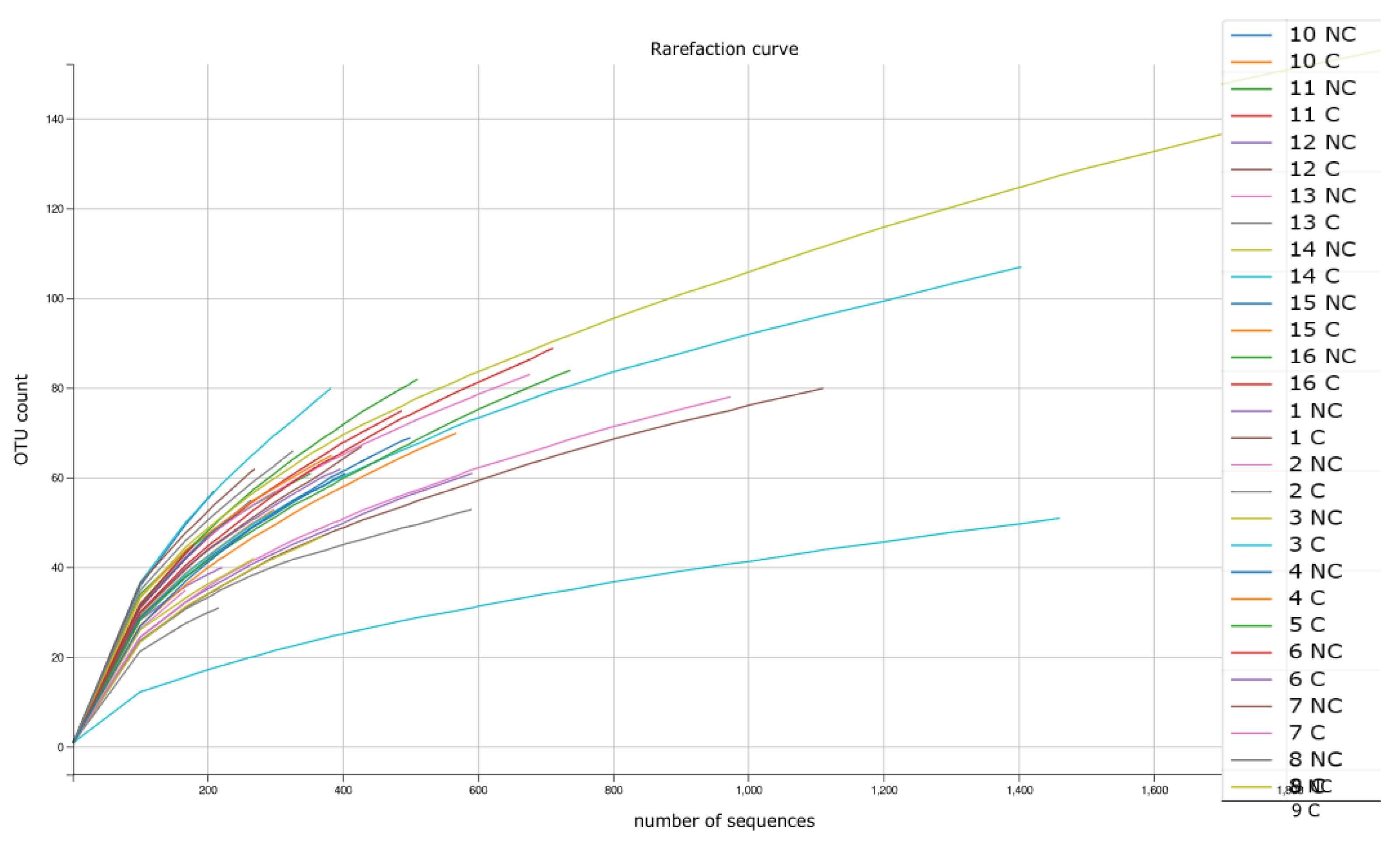
3.5. Biodiversity Indices Calculated Based on NGS Analysis
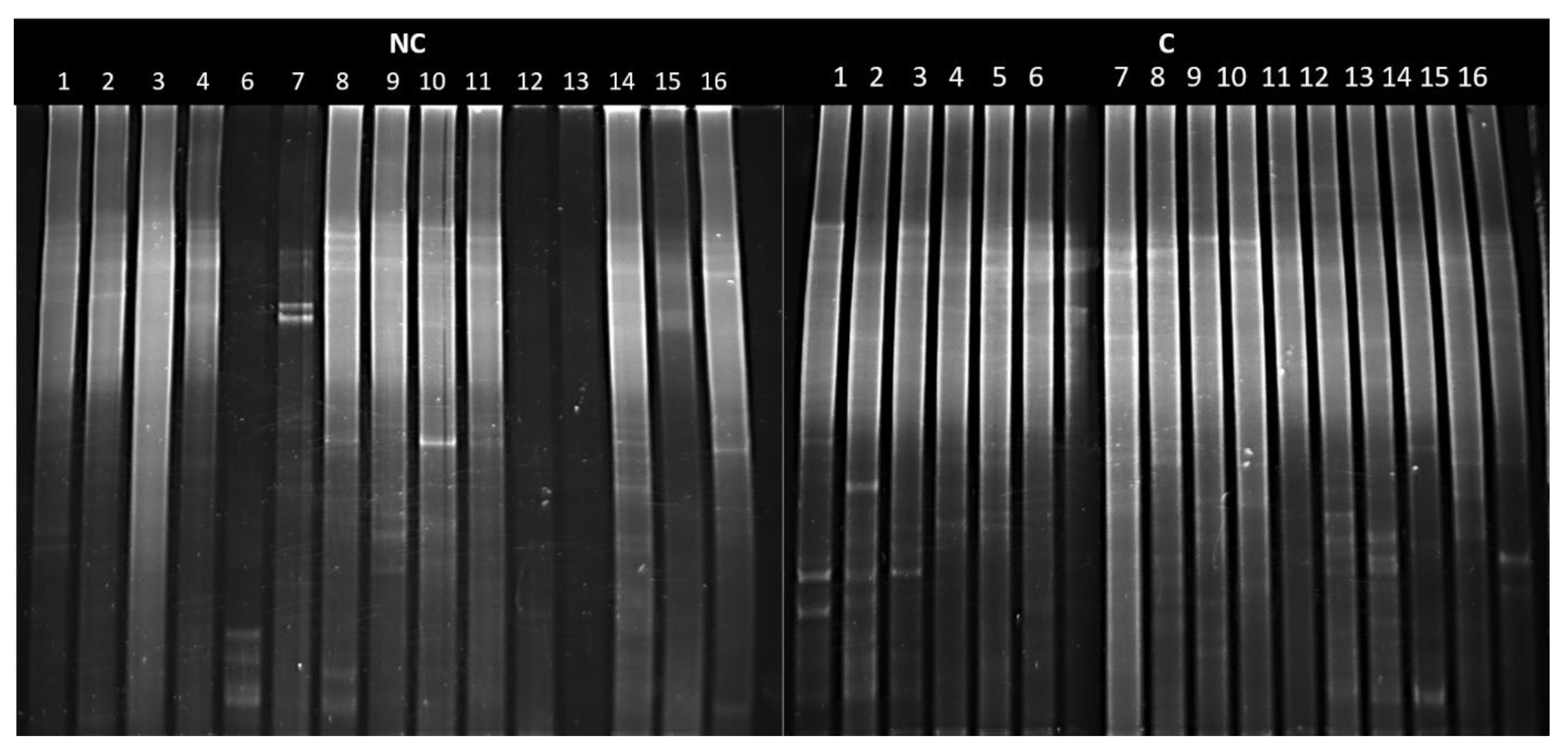
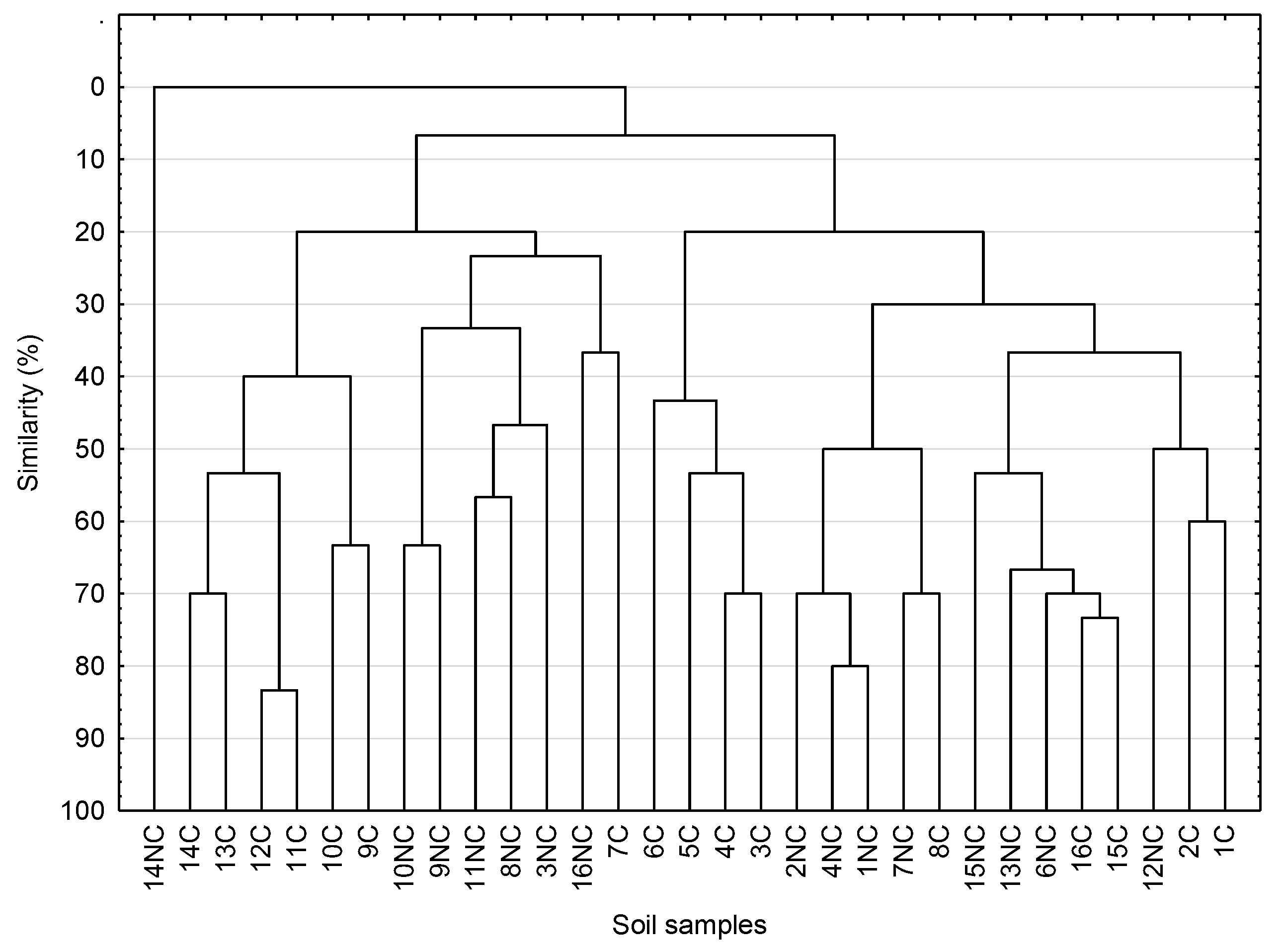
3.6. DGGE analysis of Actinobacteria in Arable and Control Soils with Biodiversity Indices
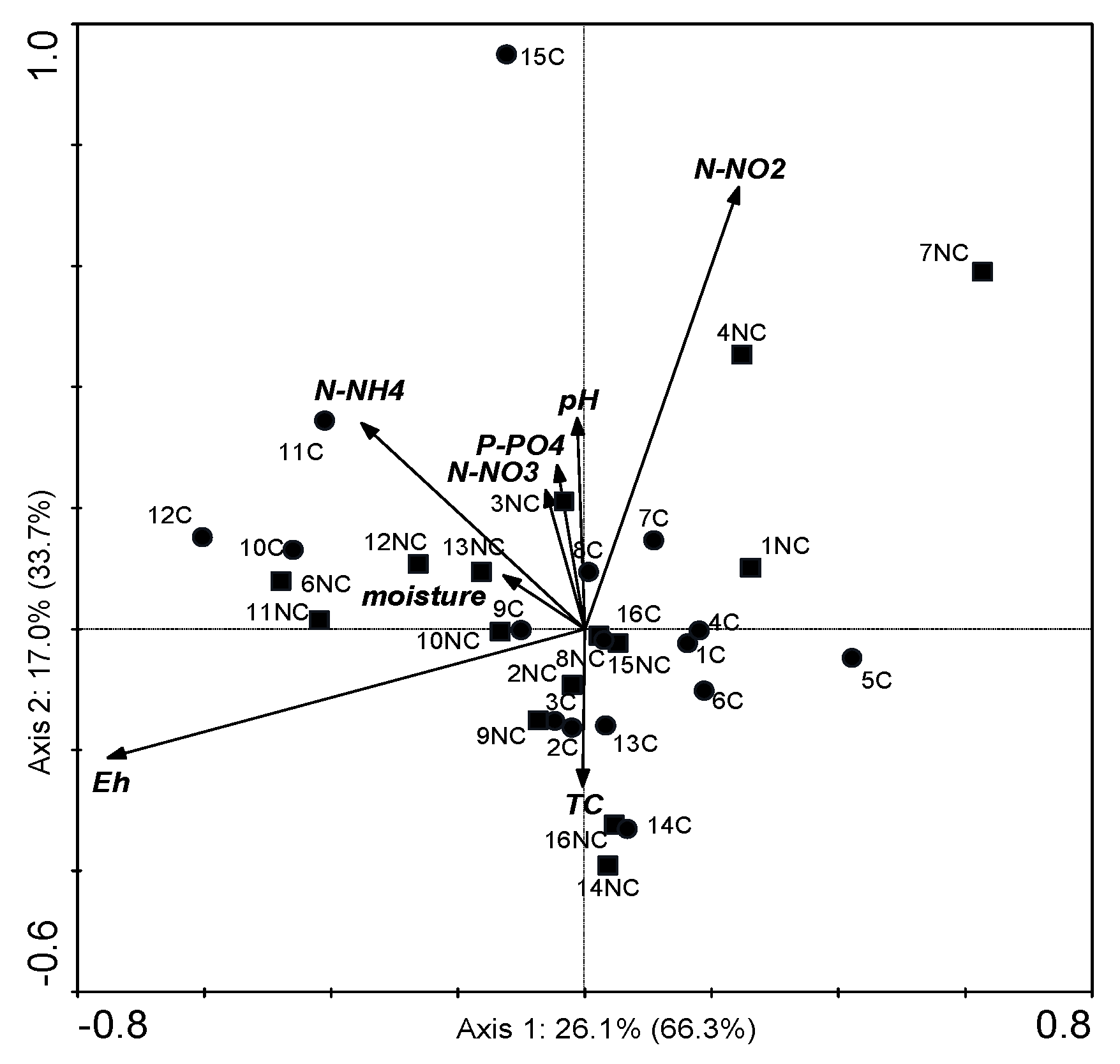
3.7. Canonical Correspondence Analysis for the POSITIONs and Chemical Factors
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
| Factor | Autogenic | Hydrogenic | Lithogenic | |||
|---|---|---|---|---|---|---|
| C | NC | C | NC | C | NC | |
| Moisture a | 11.76 | 13.20 | 7.61 | 10.72 | 11.83 | 15.90 |
| pH | 5.65 | 6.14 | 5.12 | 5.98 | 5.58 | 6.57 |
| Eh b | 486.70 | 445.43 | 547.55 | 536.50 | 496.05 | 469.98 |
| TC a | 1.17 | 2.46 | 1.44 | 2.45 | 1.11 | 3.69 |
| N-NO3 c | 19.83 | 6.55 | 11.31 | 7.02 | 55.07 | 11.97 |
| N-NO2 c | 0.09 | 0.39 | 0.09 | 0.11 | 0.09 | 0.11 |
| N-NH4 c | 0.11 | 0.53 | 0.14 | 1.50 | 0.13 | 1.83 |
| P-PO4 c | 7.41 | 3.25 | 3.38 | 1.46 | 3.93 | 0.80 |
| Factor | Soil Genesis | Land Use | |
|---|---|---|---|
| Family | |||
| Nocardioidaceae | 0.005 | 0.000 | |
| Microbacteriaceae | 0.001 | 0.000 | |
| Acidimicrobiaceae | 0.548 | 0.051 | |
| Micromonosporaceae | 0.148 | 0.053 | |
| Micrococcaceae | 0.000 | 0.024 | |
| Gaiellaceae | 0.549 | 0.445 | |
| Factor | Soil Genesis | Land Use | |
|---|---|---|---|
| Genus | |||
| Nocardioides | 0.005 | 0.000 | |
| Marmoricola | 0.005 | 0.000 | |
| Aeromicrobium | 0.005 | 0.000 | |
| Microbacterium | 0.060 | 0.000 | |
| Agromyces | 0.060 | 0.000 | |
| Ilumatobacter | 0.004 | 0.005 | |
| Gaiella | 0.548 | 0.445 | |
References
- Alvarez, A.; Saez, J.M.; Costa, J.S.D.; Colin, V.L.; Fuentes, M.S.; Cuozzo, C.A.; Benimeli, C.S.; Polti, M.A.; Amoroso, M.J. Actinobacteria: Current research and perspectives for bioremediation of pesticides and heavy metals. Chemosphere 2017, 166, 41–62. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Cong, J.; Lu, H.; Xue, Y.; Wang, X.; Li, D.; Zhang, Y. Community structure and elevational distribution pattern of soil Actinobacteria in alpine grasslands. Acta Ecol. Sin. 2017, 37, 213–218. [Google Scholar] [CrossRef]
- Elbendary, A.A.; Hessain, A.M.; El-Hariri, M.D.; Seida, A.A.; Moussa, I.M.; Mubarak, A.S.; Kabli, S.A.; Hemeg, H.A.; Jakee, J.K.E. Isolation of antimicrobial producing Actinobacteria from soil samples. Saudi J. Biol. Sci. 2018, 25, 44–46. [Google Scholar] [CrossRef] [PubMed]
- Nübel, U.; Engelen, B.; Felske, A.; Snaidr, J.; Wieshuber, A.; Amman, R.I.; Ludwig, W.; Backhaus, H. Sequence heterogeneities of genes encoding 16S rRNAs in Paenibacillus polymyxa detected by temperature gradient gel electrophoresis. J. Bacteriol. 1996, 178, 5636–5643. [Google Scholar] [CrossRef] [PubMed]
- Heuer, H.; Krsek, M.; Baker, P.; Smalla, K.; Wellington, E.M.H. Analysis of Actinomycete communities by specific amplification of genes encoding 16S rRNA and gel electrophoresis separation in denaturing gradients. Appl. Environ. Microbiol. 1997, 63, 3233–3241. [Google Scholar]
- Ma, T.; Wang, X.W.; Ruan, J.S.; Liu, N.; Huang, Y. Actinomycete flora in soil of differently degraded alpine grassland in Three River Source Area. Microbiology 2008, 35, 1879–1883. [Google Scholar]
- Hussain, E.I.; Jacob, J.H.; Shakhatreh, M.A.K.; Al-Razaq, M.A.A.; Juhmani, A.S.F.; Cornelison, C.T. Detection of antibiotic—Producing Actinobacteria in the sediment and water of Ma’in thermal sprongs (Jordan). GERMS 2018, 8, 191–198. [Google Scholar] [CrossRef]
- Rangseekaew, P.; Pathom-aree, W. Cave Actinobacteria as producers of bioactive metabolites. Front. Microbiol. 2019, 10, 387. [Google Scholar] [CrossRef]
- Assis, D.A.M.; Rezende, R.P.; Dias, J.C.T. Use of metagenomics and isolation of Actinobacteria in Brazil’s atlantic rainforest soil for antimicrobial prospecting. ISRN Biotech. 2014, 2014, 909601. [Google Scholar] [CrossRef]
- Van Dorst, J.M.; Hince, G.; Snape, I.; Ferrari, B.C. Novel culturing techniques select for heterotrophs and hydrocarbon degraders in a subantarctic soil. Sci. Rep. 2016, 6, 36724. [Google Scholar] [CrossRef]
- Adam, D.; Maciejewska, M.; Naome, A.; Martinet, L.; Coppieters, W.; Karim, L.; Baurain, D.; Rigali, S. Isolation, characterization and anitibacterial activity of hard-to-culture Actinobacteria from cave moonmilk deposits. Antibiotics 2018, 7, 28. [Google Scholar] [CrossRef] [PubMed]
- Anandan, R.; Dharumadurai, D.; Manogaran, P.M. An introduction to Actinobacteria. In Actinobacteria-Basics and Biotechnological Applications; Dhanasekaran, D., Jiang, Y., Eds.; InTech Open: London, UK, 2016; Volume 1, pp. 1–37. [Google Scholar]
- Polti, M.; Aparicio, J.D.; Benimeli, C.S.; Amoroso, M.J. Simultaneous bioremediation of Cr(VI) and lindane in soil by Actinobacteria. Int. Biodeter. Biodegr. 2014, 88, 48–55. [Google Scholar] [CrossRef]
- Manivasagan, P.; Venkatesan, J.; Sivakumar, K.; Kim, S.K. Marine actinobacterial metabolites: Current status and future perspectives. Microbiol. Res. 2013, 168, 311–332. [Google Scholar] [CrossRef] [PubMed]
- Carbonetto, B.; Rascovan, N.; Alvarez, N.; Mentaberry, A.; Vazquez, M.P. Structure, composition and metagenomic profile of soil microbiomes associated to agricultural land use and tillage systems in Argentine Pampas. PLoS ONE 2014, 9, e99949. [Google Scholar] [CrossRef] [PubMed]
- Ding, J.J.; Zhang, Y.G.; Deng, Y.; Cong, J.; Lu, H.; Sun, X.; Yang, C.Y.; Yuan, T.; Van Nostrand, J.D.; Li, D.Q.; et al. Integrated metagenomic and network analysis of soil microbial community of the forest timberline. Sci. Rep. 2015, 5, 7994. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Zhang, H.; Sun, L.; Qi, G.; Chen, S.; Zhao, X. Microbial community composition is related to soil biological and chemical properties and bacterial wilt outbreak. Sci. Rep. 2017, 7, 343. [Google Scholar] [CrossRef]
- Wolińska, A.; Kuźniar, A.; Zielenkiewicz, U.; Banach, A.; Izak, D.; Stępniewska, Z.; Błaszczyk, M. Metagenomic analysis of some potential nitrogen-fixing bacteria in arable soils at different formation processes. Microb. Ecol. 2017, 73, 162–176. [Google Scholar] [CrossRef] [PubMed]
- Wolińska, A.; Kuźniar, A.; Zielenkiewicz, U.; Izak, D.; Szafranek-Nakonieczna, A.; Banach, A.; Błaszczyk, M. Bacteroidetes as a sensitive indicator of agricultural soil usage revealed by culture-independent approach. Appl. Soil Ecol. 2017, 119, 128–137. [Google Scholar] [CrossRef]
- Wolińska, A.; Kuźniar, A.; Zielenkiewicz, U.; Banach, A.; Błaszczyk, M. Indicators of arable soils fatigue—Bacterial families and genera: A metagenomic approach. Ecol. Ind. 2018, 93, 490–500. [Google Scholar] [CrossRef]
- Wolińska, A.; Górniak, D.; Zielenkiewicz, U.; Goryluk-Salmonowicz, A.; Kuźniar, A.; Stępniewska, Z.; Błaszczyk, M. Microbial biodiversity in arable soils is affected by agricultural practices. Int. Agrophys. 2017, 31, 259–271. [Google Scholar] [CrossRef]
- Cong, J.; Yang, Y.F.; Liu, X.D.; Lu, H.; Liu, X.; Zhou, J.Z.; Li, D.Q.; Yin, H.Q.; Ding, J.J.; Zhang, Y.G. Analyses of soil microbial community compositions and functional genes reveal potential consequences of natural forest succession. Sci. Rep. 2015, 5, 10007. [Google Scholar] [CrossRef] [PubMed]
- Davidson, E.A.; Janssens, I.A. Temperature sensitivity of soil carbon decomposition and feedbacks to climate change. Nature 2006, 440, 165–173. [Google Scholar] [CrossRef] [PubMed]
- Niu, J.; Zhou, X.Q.; Jang, N.; Wang, Y.F. Characteristics of soil microbial communities under dry and wet condition in Zoige alpine wetland. Acta Ecol. Sin. 2011, 31, 474–482. [Google Scholar]
- Liu, L.; Gundersen, P.; Zhang, T.; Mo, M. Effects of phosphorus addition on soil microbial biomass and community composition in three forest types in tropical China. Soil Biol. Biochem. 2012, 44, 31–38. [Google Scholar] [CrossRef]
- Sul, W.J.; Asuming-Brempong, S.; Wang, Q.; Tourlousse, D.M.; Penton, C.R.; Deng, Y.; Rodrigues, J.L.M.; Adiku, S.G.K.; Jones, J.W.; Zhou, J.Z.; et al. Tropical agricultural land management influences on soil microbial communities through its effect on soil organic carbon. Soil Biol. Biochem. 2013, 65, 33–38. [Google Scholar] [CrossRef]
- Liu, J.J.; Sui, Y.Y.; Yu, Z.H.; Shi, Y.; Chu, H.Y.; Jin, J.; Liu, X.B.; Wang, G.H. High throughput sequencing analysis of biogeographical distribution of bacterial communities in the black soils of northeast China. Soil Biol. Biochem. 2014, 70, 113–122. [Google Scholar] [CrossRef]
- Wolińska, A.; Szafranek-Nakonieczna, A.; Banach, A.; Błaszczyk, M.; Stępniewska, Z. The impact of agricultural soil usage on activity and abundance of ammonyfying bacteria in selected soils from Poland. Springer Plus 2016, 5, 565. [Google Scholar] [CrossRef]
- Ramirez, K.S.; Craine, J.M.; Fierer, N. Consistent effects of nitrogen amendments on soil microbial communities and processes across biomes. Glob. Chang. Biol. 2012, 18, 1918–1927. [Google Scholar] [CrossRef]
- Cong, J.; Liu, X.D.; Lu, H.; Xu, H.; Li, Y.D.; Deng, Y.; Li, D.Q.; Zhang, Y.D. Available nitrogen is the key factor influencing soil microbial functional gene diversity in tropical rainforest. BMC Microbiol. 2015, 15, 167. [Google Scholar] [CrossRef] [PubMed]
- Chemical and Agricultural Soil Analyses. Soil Sampling; PN-R-04031; PKN: Warsaw, Poland, 1997. (In Polish)
- Tomczyk-Żak, K.; Kaczanowski, S.; Drewniak, Ł.; Dmoch, Ł.; Skłodowska, A.; Zielenkiewicz, U. Bacteria diversity and arsenic mobilization in rock biofilm from an ancient gold and arsenic mice. Sci. Total Environ. 2013, 461, 330–340. [Google Scholar] [CrossRef]
- Schloss, P.D.; Westcott, S.I.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-source, platform independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef] [PubMed]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, 590–596. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quinee, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef] [PubMed]
- Crump, B.C.; Hobbie, J.E. Synchrony and seasonality in bacterioplankton communities of two temperate rivers. Limnol. Ocean 2005, 50, 1718–1729. [Google Scholar] [CrossRef]
- Ter Braak, C.J.F.; Šmilauer, P. CANOCO Reference Manual and CanoDraw for Windows User’s Guide: Software for Canonical Community Ordination (Version 4.5); Microcomputer Power: Ithaca, NY, USA, 2002. [Google Scholar]
- Vivas, A.; Moreno, B.; Garcia-Rodriguez, S.; Benitez, E. Assessing the impact of composting and vermicomposting on bacterial community size and structure, and microbial functional diversity of an olive-mill waste. Biores. Technol. 2009, 100, 1319–1326. [Google Scholar] [CrossRef]
- Jia, G.M.; Zhang, P.D.; Wang, G.; Cao, J.; Hang, J.C.; Huang, Y.P. Relationship between microbial community and soil properties during natural succession of abandoned agricultural land. Pedosphere 2010, 20, 352–360. [Google Scholar] [CrossRef]
- Swędrzyńska, D.; Małecka, I.; Blecharczyk, A.; Swędrzyński, A.; Satrzyk, J. Effect of various long-term tillage systems on some chemical and biological properties of soil. Pol. J. Environ. Stud. 2013, 22, 1835–1844. [Google Scholar]
- Vinhal-Freitas, I.C.; Corrêa, G.F.; Wendling, B.; Bobul’ska, L.; Ferreira, A.S. Soil textural class plays a major role in evaluating the effects of land use on soil quality indicators. Ecol. Ind. 2017, 74, 182–190. [Google Scholar] [CrossRef]
- Wolińska, A.; Stępniewska, Z.; Szafranek, A. Effect of selected physical parameters on respiration activities in common Polish mineral soils. Pol. J. Environ. Stud. 2011, 20, 1075–1082. [Google Scholar]
- Ulrich, A.; Becker, R. Soil parent material is a key determinant of the bacterial community structure in arable soils. FEMS Microbiol. Ecol. 2006, 56, 430–443. [Google Scholar] [CrossRef]
- Toth, E.M.; Borsodi, A.K. The family Nocardioidaceae. In The Prokaryotes—Actinobacteria; Rosenberg, E., DeLong, E.F., Lory, S., Stackebrandt, E., Thompson, F., Eds.; Springer: Berlin/Heidelberg, Germany, 2014; Volume 33, pp. 1–44. [Google Scholar] [CrossRef]
- Sukda, P.; Gouda, N.; Ito, E.; Myauchi, K.; Masai, E.; Fukuda, M. Characterization of a transcriptional regulatory gene involved in dibenzofuran degradation by Nocardioides sp. strain DF412. Biosci. Biotechnol. Biochem. 2009, 73, 508–516. [Google Scholar] [CrossRef] [PubMed]
- Vibber, L.L.; Pressler, M.J.; Colores, G.M. Isolation and characterization of novel atrazine-degrading microorganisms from an agricultural soil. Appl. Microbiol. Biotechnol. 2007, 75, 921–928. [Google Scholar] [CrossRef] [PubMed]
- Yoon, J.H.; Park, Y.H. The genus Nocardioides. Prokaryotes 2006, 3, 1099–1113. [Google Scholar] [CrossRef]
- Lee, D.W.; Lee, S.D. Marmoricola scoriae sp. nov., isolated from volcanic ash. Int. J. Syst. Evol. Microbiol. 2010, 60, 2135–2139. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.D. Marmoricolaaequoreus sp. nov., a novel actinobacterium isolated from marine sediment. Int. J. Syst. Evol. Microbiol. 2007, 57, 1391–1395. [Google Scholar] [CrossRef] [PubMed]
- Cui, Y.S.; Im, W.T.; Yin, C.R.; Lee, J.S.; Lee, K.C.; Lee, S.T. Aeromicrobium panaciterrae sp. nov., isolated from soil of a ginseng field in South Korea. Int. J. Syst. Evol. Microbiol. 2007, 57, 687–691. [Google Scholar] [CrossRef]
- Siddiqi, M.Z.; Lee, S.Y.; Choi, K.D.; Im, W.T. Aeromicrobium panacisolisp. nov. isolated from soil of ginseng cultivating field. Curr. Microbiol. 2008, 75, 624–629. [Google Scholar] [CrossRef]
- Yan, Z.F.; Lin, P.; Chu, X.; Kook, M.C.; Li, C.T.; Yi, T.H. Aeromicrobium halotolerans sp. nov., isolated from desert soil sample. Arch. Microbiol. 2016, 198, 423–427. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.H.; Yang, H.O.; Sohn, Y.C.; Kwon, H.C. Aeromicrobium halocynthiae sp. nov., taurocholic acid-producing bacterium isolated from the marine ascidian Halocynthiaroretzi. Int. J. Syst. Evol. Microbiol. 2010, 60, 2793–2798. [Google Scholar] [CrossRef]
- Tang, Y.; Zhou, G.; Zhang, L.; Mao, J.; Luo, X.; Wang, M.; Fang, C. Aeromicrobium flavum sp. nov., isolated from air. Int. J. Syst. Evol. Microbiol. 2008, 58, 1860–1863. [Google Scholar] [CrossRef] [PubMed]
- Niu, L.; Xiong, M.; Tang, T.; Song, L.; Hu, X.; Zhao, M.; Zhang, K. Aeromicrobium camelliae sp. nov., isolated from Pu’er tea. Int. J. Syst. Evol. Microbiol. 2015, 65, 4369–4373. [Google Scholar] [CrossRef] [PubMed]
- Fujinami, S.; Takarada, H.; Kasai, H.; Sekine, M.; Omata, S.; Harada, T.; Fukai, R.; Hosoyama, A.; Horikawa, H.; Kato, Y.; et al. Complete genome sequence of YM16-304T. Stand. Genomic Sci. 2013, 8, 430–440. [Google Scholar] [CrossRef] [PubMed]
- Raaijmakers, J.M.; Paulitz, T.C.; Steinberg, C.; Alabouvette, C.; Moenne-Locoz, Y. The rhizosphere: A playground and battlefield for soilborne pathogens and beneficial microorganisms. Plant Soil 2009, 321, 341–361. [Google Scholar] [CrossRef]
- Rivas, R.; Trujillo, M.E.; Mateos, P.F.; Martinez-Molina, E.; Velazquez, E. Agromyces ulmi sp. nov, a xylanolytic bacterium isolated from Ulmusnigra in Spain. Int. J. Syst. Evol. Microbiol. 2004, 54, 1987–1990. [Google Scholar] [CrossRef][Green Version]
- Evtushenko, L.I.; Takeuchi, M. The family Microbacteriaceae. Prokaryotes 2006, 3, 1020–1098. [Google Scholar] [CrossRef]
- Abdel-Razek, A.S.; Refaat, B.M.; Abdel-Shakour, E.H.; Zaher, R.; Mohamed, M.K. Biodegradation of phenol by Microbacterium terregenes isolated from oil filed NORM soil. J. Appl. Environ. Microbiol. 2015, 3, 63–69. [Google Scholar] [CrossRef]
- Albuquerque, L.; França, L.; Rainey, F.A.; Schumann, P.; Nobre, M.F.; da Costa, M.S. Gaiellaoculta gen. nov., sp. nov., a novel representative of a deep branching phylogenetic lineage within the class Actinobacteria and proposal of Gaiellaceae fam. nov. and Gaiellales ord. nov. Syst. Appl. Microbiol. 2011, 34, 595–599. [Google Scholar] [CrossRef]
- Peng, M.; Zi, X.; Wang, Q. Bacterial community diversity of oil-contaminated soils assessed by high throughput sequencing of 16S rRNA genes. Int. J. Environ. Res. Public Health 2015, 12, 12002–12015. [Google Scholar] [CrossRef]
- Kumar, V.; AlMomin, S.; Al-Aqeel, H.; Al-Salameen, F.; Nair, S.; Shajan, A. Metagenomic analysis of rhizosphere microflora of oil-contaminated soil planted with barley alfalfa. PLoS ONE 2018, 13, e0202127. [Google Scholar] [CrossRef]
- Li, M.; Cheng, X.; Guo, H.; Yang, Z. Biomineralization of carbonate by Terrabacter tumescens for havy metal removal and biogrouting applications. J. Environ. Eng. 2016, 142, 1–6. [Google Scholar] [CrossRef]
- Vaijayanthi, G.; Cholarajan, A.; Vijayakumar, R. Isolation, characterization, and antibacterial activity of terrestrial Actinobacteria in the soils of Thanjavur District, Tamil Nadu, India. Res. J. Sci. Technol. 2012, 4, 132–139. [Google Scholar]
- Nacke, H.; Thűrmer, A.; Wollherr, A.; Will, C.; Hodac, L.; Herold, N.; Schöning, I.; Schrumpf, M.; Daniel, R. Pyrosequencing-based assessment of bacterial community structure along different management types in German forest and grassland soils. PLoS ONE 2011, 6, e0017000. [Google Scholar] [CrossRef]
- Hruska, K.; Kaevska, M. Mycobateria in water, soil, plants and air: A review. Vet. Med. 2012, 57, 623–679. [Google Scholar] [CrossRef]
- Qin, S.; Li, J.; Chen, H.H.; Zhao, G.Z.; Zhu, W.Y.; Jiang, C.L.; Xu, L.H.; Li, W.J. Isolation, diversity, and antimicrobial activity of rare Actinobacteria from medicinal plants of tropical rain forests in Xsishuangbanna, China. App. Environ. Microbiol. 2009, 75, 6176–6186. [Google Scholar] [CrossRef] [PubMed]
- Passari, A.K.; Mishra, V.K.; Singh, G.; Singh, P.; Kumar, B.; Gupta, V.K.; Sarma, R.K.; Saikia, R.; O’Donovan, A.; Singh, B.P. Insight into the functionality of endophytic actinobacteria with a focus on their biosynthetic potential and secondary metabolites production. Sci. Rep. 2017, 7, 11809. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.K.; Lee, K.C.; Lee, J.S. Patulibacter ginsengiterrae sp. nov. isolated from soil of a ginseng field, and an amended description of the genus Patulibacter. Int. J. Syst. Evol. Microbiol. 2012, 62, 563–568. [Google Scholar] [CrossRef]
- Lauber, C.L.; Hamady, M.; Knight, R.; Fierer, N. Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl. Environ. Microbiol. 2009, 75, 5111–5120. [Google Scholar] [CrossRef] [PubMed]
- Dimitriu, P.A.; Grayston, S.J. Relationship between soil properties and patterns of bacterial b-diversity across reclaimed and natural boreal forest soils. Microb. Ecol. 2010, 59, 563–573. [Google Scholar] [CrossRef] [PubMed]
- Fierer, N.; Leff, J.W.; Adams, B.J.; Nielsen, U.N.; Bates, S.T.; Lauber, C.L.; Owens, S.; Gilbert, J.A.; Wall, D.H.; Caporaso, J.G. Cross-biome metagenomic analyses of soil microbial communities and their functional attributes. Proc. Natl. Acad. Sci. USA 2012, 109, 21390–21395. [Google Scholar] [CrossRef]
- Högberg, M.N.; Högberg, P.; Myrold, D.D. Is microbial community composition in boreal forest soils determined by pH, C-to-N ratio, the trees, or all three? Oecologia 2007, 150, 590–599. [Google Scholar] [CrossRef]

| Soil No. | Field Code (Soil Group) | FAO Classification | Geographic Coordinates |
|---|---|---|---|
| 1 | Autogenic (1–10) | Albic Luvisol | 22°10′17,7′’51°26′24,6″ |
| 2 | Albic Luvisol | 22°27′10,3′’51°24′3,8″ | |
| 3 | Albic Luvisol | 22°36′51,8′’51°21′27,0″ | |
| 4 | Haplic Luvisol | 22°06′54,2′’51°21′52,2″ | |
| 5 | BrunicArenosol | 22°15′19,0′’51°23′0,9″ | |
| 6 | BrunicArenosol | 22°15′55,5′’51°23′1,9″ | |
| 7 | Haplic Luvisol | 24°04′0,3′’50°51′15,81 | |
| 8 | Haplic Luvisol | 23°22′52,4′’50°51′14,8″ | |
| 9 | Haplic Luvisol | 22°07′29,9′’51°25′5,5″ | |
| 10 | Haplic Chernozem | 50°44′48,3″23°42′56,6′’ | |
| 11 | Hydrogenic (11–14) | GleyicPhaeozem | 22°06′18,8′’51°22′48,0″ |
| 12 | MollicGleysol | 22°01′25,5′’51°29′15,3″ | |
| 13 | FluvicGleyicPhaeozem | 21°59′10,1′’51°33′47,7″ | |
| 14 | GleyicUmbrisol | 22°16′38,9′’51°25′27,3″ | |
| 15 | Lithogenic (15–16) | CalcaricCambisol | 23°10′58,3′’51°12′22,3″ |
| 16 | CalcaricPhaeozem | 23°11′43,9′’51°12′10,8″ |
| Soil No. | Moisture a | pH | Eh b | TC a | N–NH4 c | N–NO3 c | N–NO2 c | P– PO4 c |
|---|---|---|---|---|---|---|---|---|
| 1C | 8.20 ± 0.20 | 5.23 ± 0.06 | 477.4 ± 0.40 | 0.98 ± 0.002 | 0.01 ± 0.006 | 9.34 ± 0.8 | 0.11 ± 0.003 | 2.56 ± 0.04 |
| 1NC | 9.76 ± 0.11 | 6.27 ± 0.005 | 435.2 ± 0.20 | 1.76 ± 0.12 | 0.09 ± 0.006 | 1.68 ± 0.014 | 0.17 ± 0.001 | 1.77 ± 0.03 |
| 2C | 9.30 ± 0.10 | 4.66 ± 0.02 | 546.73 ± 0.21 | 1.23 ± 0.04 | 0.02 ± 0.001 | 7.37 ± 0.05 | 0.08 ± 0.001 | 1.51 ± 0.01 |
| 2NC | 11.16 ± 0.11 | 5.02 ± 0.02 | 528.40 ± 0.36 | 1.40 ± 0.05 | 0.04 ± 0.014 | 5.84 ± 0.03 | 0.10 ± 0.001 | 1.01 ± 0.01 |
| 3C | 10.22 ± 0.03 | 4.78 ± 0.02 | 535.7 ± 0.30 | 1.24 ± 0.04 | 0.01 ± 0.001 | 53.32 ± 0.52 | 0.05 ± 0.005 | 19.6 ± 0.98 |
| 3NC | 9.13 ± 0.05 | 6.22 ± 0.09 | 452.86 ± 0.11 | 1.79 ± 0.14 | 0.06 ± 0.006 | 3.58 ± 0.09 | 0.42 ± 0.005 | 1.16 ± 0.05 |
| 4C | 12.56 ± 0.06 | 6.98 ± 0.02 | 450.03 ± 0.32 | 1.96 ± 0.05 | 0.43 ± 0.006 | 18.25 ± 0.06 | 0.10 ± 0.004 | 12.9 ± 0.04 |
| 4NC | 13.50 ± 0.10 | 7.08 ± 0.06 | 419.20 ± 1.11 | 2.52 ± 0.14 | 0.48 ± 0.008 | 7.57 ± 0.32 | 0.53 ± 0.003 | 5.9 ± 0.03 |
| 5C | 6.60 ± 0.10 | 5.45 ± 0.04 | 470.20 ± 17.75 | 1.01 ± 0.04 | 0.07 ± 0.006 | 25.53 ± 0.18 | 0.12 ± 0.001 | 6.88 ± 0.01 |
| 5NC | 8.63 ± 0.15 | 5.58 ± 0.04 | 396.13 ± 0.23 | 2.06 ± 0.19 | 0.69 ± 0.009 | 10.18 ± 0.14 | 0.21 ± 0.002 | 3.52 ± 0.09 |
| 6C | 9.23 ± 0.06 | 4.78 ± 0.006 | 480.60 ± 0.18 | 0.83 ± 0.09 | 0.01 ± 0.007 | 20.26 ± 0.07 | 0.09 ± 0.004 | 4.01 ± 0.01 |
| 6NC | 8.63 ± 0.15 | 5.58 ± 0.04 | 396.13 ± 0.23 | 2.06 ± 0.19 | 0.69 ± 0.009 | 10.18 ± 0.14 | 0.21 ± 0.002 | 3.52 ± 0.09 |
| 7C | 12.13 ± 0.15 | 6.93 ± 0.006 | 403.10 ± 3.64 | 0.97 ± 0.06 | 0.05 ± 0.001 | 14.48 ± 0.04 | 0.04 ± 0.005 | 4.61 ± 0.01 |
| 7NC | 12.76 ± 0.11 | 6.99 ± 0.03 | 400.66 ± 0.15 | 3.49 ± 0.11 | 0.41 ± 0.008 | 5.41 ± 0.14 | 0.87 ± 0.003 | 3.85 ± 0.03 |
| 8C | 19.00 ± 0.17 | 5.96 ± 0.12 | 461.10 ± 0.17 | 0.96 ± 0.11 | 0.36 ± 0.02 | 17.35 ± 0.03 | 0.12 ± 0.002 | 6.81 ± 0.02 |
| 8NC | 20.26 ± 0.63 | 6.06 ± 0.009 | 409.23 ± 0.25 | 2.68 ± 0.07 | 2.61 ± 0.04 | 11.07 ± 0.05 | 0.24 ± 0.02 | 2.94 ± 0.03 |
| 9C | 5.66 ± 0.11 | 5.13 ± 0.006 | 480.73 ± 0.93 | 0.88 ± 0.06 | 0.19 ± 0.009 | 4.96 ± 0.06 | 0.14 ± 0.001 | 13.9 ± 0.24 |
| 9NC | 7.10 ± 0.17 | 5.40 ± 0.006 | 487.23 ± 0.25 | 1.42 ± 0.11 | 0.18 ± 0.001 | 1.76 ± 0.06 | 0.80 ± 0.002 | 7.52 ± 0.02 |
| 10C | 24.66 ± 0.28 | 6.61 ± 0.05 | 561.30 ± 0.36 | 1.64 ± 0.03 | 0.02 ± 0.001 | 27.43 ± 0.08 | 0.09 ± 0.003 | 1.36 ± 0.05 |
| 10NC | 31.03 ± 0.23 | 7.22 ± 0.02 | 529.26 ± 0.23 | 5.43 ± 0.14 | 0.02 ± 0.002 | 8.23 ± 0.02 | 0.44 ± 0.006 | 1.35 ± 0.02 |
| 11C | 12.96 ± 0.28 | 6.73 ± 0.006 | 556.10 ± 0.30 | 1.18 ± 0.02 | 0.41 ± 0.04 | 10.11 ± 0.07 | 0.13 ± 0.004 | 5.77 ± 0.13 |
| 11NC | 14.33 ± 0.57 | 6.76 ± 0.01 | 537.96 ± 0.25 | 3.15 ± 0.29 | 0.78 ± 0.01 | 10.06 ± 0.09 | 0.15 ± 0.001 | 1.09 ± 0.02 |
| 12C | 5.80 ± 0.17 | 4.74 ± 0.02 | 559.36 ± 0.32 | 0.91 ± 0.05 | 0.03 ± 0.004 | 21.90 ± 0.02 | 0.09 ± 0.001 | 2.04 ± 0.03 |
| 12NC | 10.40 ± 0.17 | 6.25 ± 0.03 | 542.90 ± 2.95 | 1.80 ± 0.13 | 4.94 ± 0.08 | 6.75 ± 0.05 | 0.10 ± 0.001 | 1.68 ± 0.008 |
| 13C | 5.20 ± 0.17 | 4.18 ± 0.05 | 551.30 ± 0.30 | 0.98 ± 0.07 | 0.14 ± 0.04 | 2.99 ± 0.03 | 0.09 ± 0.001 | 2.64 ± 0.09 |
| 13NC | 8.86 ± 0.11 | 5.64 ± 0.06 | 545.20 ± 0.40 | 1.23 ± 0.08 | 0.27 ± 0.03 | 2.20 ± 0.05 | 0.13 ± 0.002 | 1.33 ± 0.008 |
| 14C | 6.50 ± 0.10 | 4.85 ± 0.03 | 523.43 ± 0.23 | 2.69 ± 0.19 | 0.01 ± 0.001 | 10.22 ± 0.12 | 0.08 ± 0.002 | 3.09 ± 0.10 |
| 14NC | 9.30 ± 0.20 | 5.27 ± 0.01 | 519.96 ± 0.25 | 3.63 ± 0.14 | 0.02 ± 0.002 | 9.05 ± 0.03 | 0.09 ± 0.001 | 1.74 ± 0.38 |
| 15C | 10.86 ± 0.11 | 5.58 ± 0.06 | 503.90 ± 0.20 | 0.97 ± 0.06 | 0.05 ± 0.01 | 77.17 ± 0.14 | 0.08 ± 0.007 | 6.83 ± 0.19 |
| 15NC | 12.50 ± 0.17 | 5.76 ± 0.01 | 493.80 ± 0.20 | 1.59 ± 0.12 | 3.39 ± 0.06 | 10.12 ± 0.07 | 0.09 ± 0.004 | 0.60 ± 0.007 |
| 16C | 12.80 ± 0.10 | 5.58 ± 0.11 | 488.20 ± 0.20 | 1.25 ± 0.05 | 0.22 ± 0.01 | 32.98 ± 0.27 | 0.09 ± 0.001 | 1.04 ± 0.02 |
| 16NC | 19.30 ± 0.17 | 7.39 ± 0.02 | 446.16 ± 0.47 | 5.80 ± 0.43 | 0.28 ± 0.02 | 13.82 ± 0.5 | 0.13 ± 0.004 | 1.01 ± 0.02 |
| Soil Type | Mode of Land Use | Shannon-Weaver Diversity Index (H’) | Simpson’s Dominance Index (1/D) | Simpson’s Diversity Index (D) |
|---|---|---|---|---|
| Autogenic | NC | 2.87 ± 0.004 | 10.28 ± 0.005 | 0.14 ± 0.004 |
| C | 2.86 ± 0.006 | 9.90 ± 0.004 | 0.11 ± 0.005 | |
| Hydrogenic | NC | 2.99 ± 0.002 | 11.22 ± 0.004 | 0.09 ± 0.006 |
| C | 2.94 ± 0.001 | 10.19 ± 0.002 | 0.10 ± 0.004 | |
| Lithogenic | NC | 3.13 ± 0.003 | 14.00 ± 0.001 | 0.08 ± 0.003 |
| C | 3.08 ± 0.004 | 11.74 ± 0.003 | 0.07 ± 0.004 |
| Non–Cultivated Soils | H’ | 1/D | D | S | Cultivated Soils | Soil Type | H’ | 1/D | D | S |
|---|---|---|---|---|---|---|---|---|---|---|
| 11NC | 3.63 | 6.07 | 0.165 | 16 | 11C | MollicGleysol | 2.44 | 7.37 | 0.136 | 6 |
| 15NC | 2.88 | 7.64 | 0.131 | 8 | 15C | Rendzina Leptosol | 2.44 | 7.37 | 0.136 | 6 |
| 14NC | 3.64 | 6.05 | 0.165 | 22 | 14C | EutricHistosol | 2.65 | 6.80 | 0.147 | 7 |
| 4NC | 3.29 | 6.70 | 0.149 | 11 | 4C | Haplic Luvisols | 2.83 | 6.36 | 0.157 | 8 |
| 16NC | 3.63 | 6.07 | 0.165 | 16 | 16C | Rendzina Leptosol | 2.83 | 6.36 | 0.157 | 8 |
| 10NC | 3.59 | 6.13 | 0.163 | 15 | 10C | Haplic Phaezoem | 2.99 | 6.03 | 0.166 | 9 |
| 12NC | 3.17 | 6.94 | 0.144 | 10 | 12C | MollicGleysol | 2.99 | 6.03 | 0.166 | 9 |
| 8NC | 3.59 | 6.13 | 0.163 | 15 | 8C | Haplic Luvisols | 3.12 | 5.76 | 0.173 | 10 |
| 2NC | 3.17 | 6.94 | 0.144 | 10 | 2C | Albic Luvisols | 3.34 | 5.39 | 0.186 | 12 |
| 13NC | 2.70 | 8.15 | 0.123 | 7 | 13C | EutricFluvisol | 3.34 | 5.39 | 0.186 | 12 |
| 1NC | 3.29 | 6.70 | 0.149 | 11 | 1C | Albic Luvisols | 3.43 | 5.25 | 0.190 | 13 |
| 3NC | 3.63 | 6.07 | 0.165 | 16 | 3C | Albic Luvisols | 3.43 | 5.25 | 0.190 | 13 |
| 9NC | 3.63 | 6.07 | 0.165 | 16 | 9C | Haplic Luvisols | 3.50 | 5.14 | 0.194 | 14 |
| 7NC | 3.04 | 7.25 | 0.138 | 9 | 7C | Haplic Luvisols | 3.56 | 5.06 | 0.198 | 15 |
| 5NC | 3.29 | 6.70 | 0.149 | 11 | 5C | BrunicArenosols | 3.66 | 4.92 | 0.203 | 18 |
| 6NC | 2.25 | 9.77 | 0.102 | 5 | 6C | BrunicArenosols | 3.66 | 4.92 | 0.203 | 18 |
| Mode of Land Use | Shannon-Weaver Diversity Index (H’) | Simpson’s Dominance Index (1/D) | Simpson’s Diversity Index (D) |
|---|---|---|---|
| Non-cultivated (NC) | 9.34 ± 0.06 * | 2.90 ± 0.004 | 0.34 ± 0.007 |
| Cultivated (C) | 8.77 ± 0.06 | 3.35 ± 0.006 | 0.27 ± 0.006 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wolińska, A.; Górniak, D.; Zielenkiewicz, U.; Kuźniar, A.; Izak, D.; Banach, A.; Błaszczyk, M. Actinobacteria Structure in Autogenic, Hydrogenic and Lithogenic Cultivated and Non-Cultivated Soils: A Culture-Independent Approach. Agronomy 2019, 9, 598. https://doi.org/10.3390/agronomy9100598
Wolińska A, Górniak D, Zielenkiewicz U, Kuźniar A, Izak D, Banach A, Błaszczyk M. Actinobacteria Structure in Autogenic, Hydrogenic and Lithogenic Cultivated and Non-Cultivated Soils: A Culture-Independent Approach. Agronomy. 2019; 9(10):598. https://doi.org/10.3390/agronomy9100598
Chicago/Turabian StyleWolińska, Agnieszka, Dorota Górniak, Urszula Zielenkiewicz, Agnieszka Kuźniar, Dariusz Izak, Artur Banach, and Mieczysław Błaszczyk. 2019. "Actinobacteria Structure in Autogenic, Hydrogenic and Lithogenic Cultivated and Non-Cultivated Soils: A Culture-Independent Approach" Agronomy 9, no. 10: 598. https://doi.org/10.3390/agronomy9100598
APA StyleWolińska, A., Górniak, D., Zielenkiewicz, U., Kuźniar, A., Izak, D., Banach, A., & Błaszczyk, M. (2019). Actinobacteria Structure in Autogenic, Hydrogenic and Lithogenic Cultivated and Non-Cultivated Soils: A Culture-Independent Approach. Agronomy, 9(10), 598. https://doi.org/10.3390/agronomy9100598








