Automatic Segmentation and Counting of Aphid Nymphs on Leaves Using Convolutional Neural Networks
Abstract
1. Introduction
- We took a CNN-based approach to segment aphid nymphs on leaves. This method achieved significant performance improvement compared to traditional methods using hand-crafted features. Moreover, the proposed CNN were trained from scratch with limited training data (68 images).
- We measured the quantity of aphid nymphs using segmentation results. The results show low error, and the correlation between automatic counting and manual counting is high (R2 = 0.99).
- We evaluated the CNN architecture with different capacities and show that satisfying performance can be achieved by a relatively small network.
2. Materials and Methods
2.1. Dataset
2.1.1. Aphids Preparation and Image Acquisition
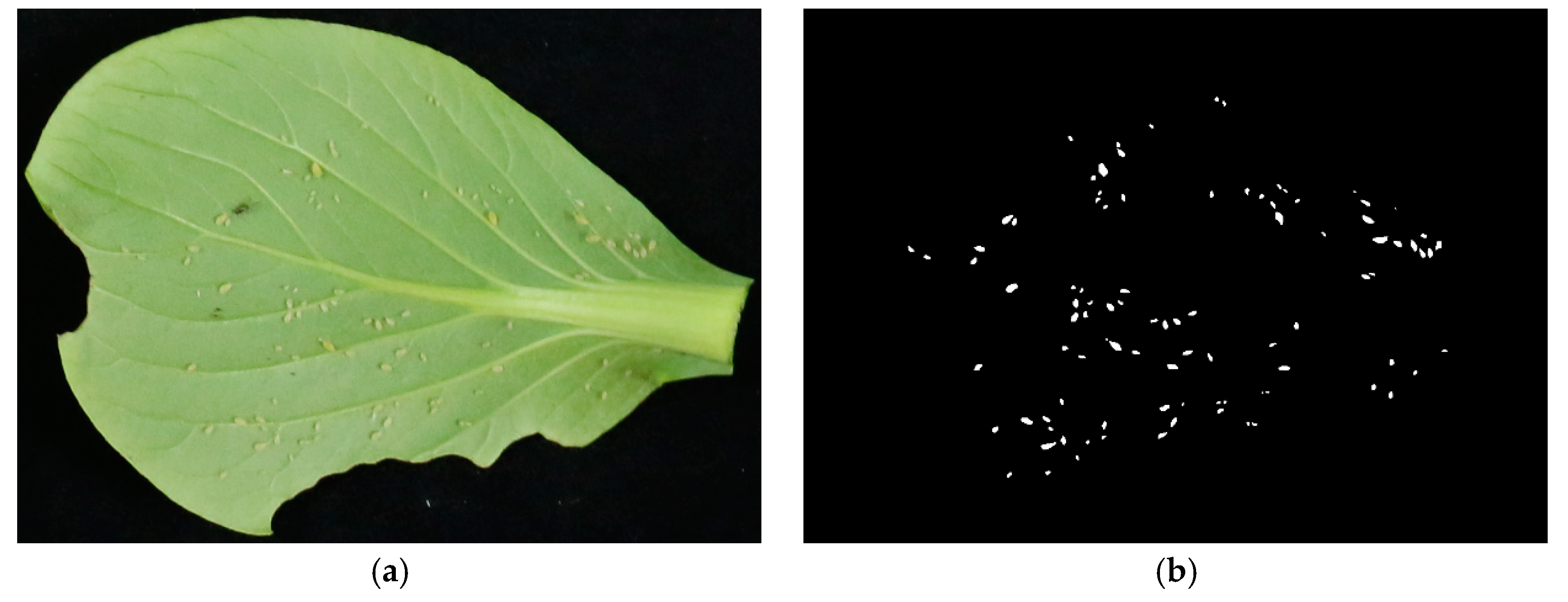
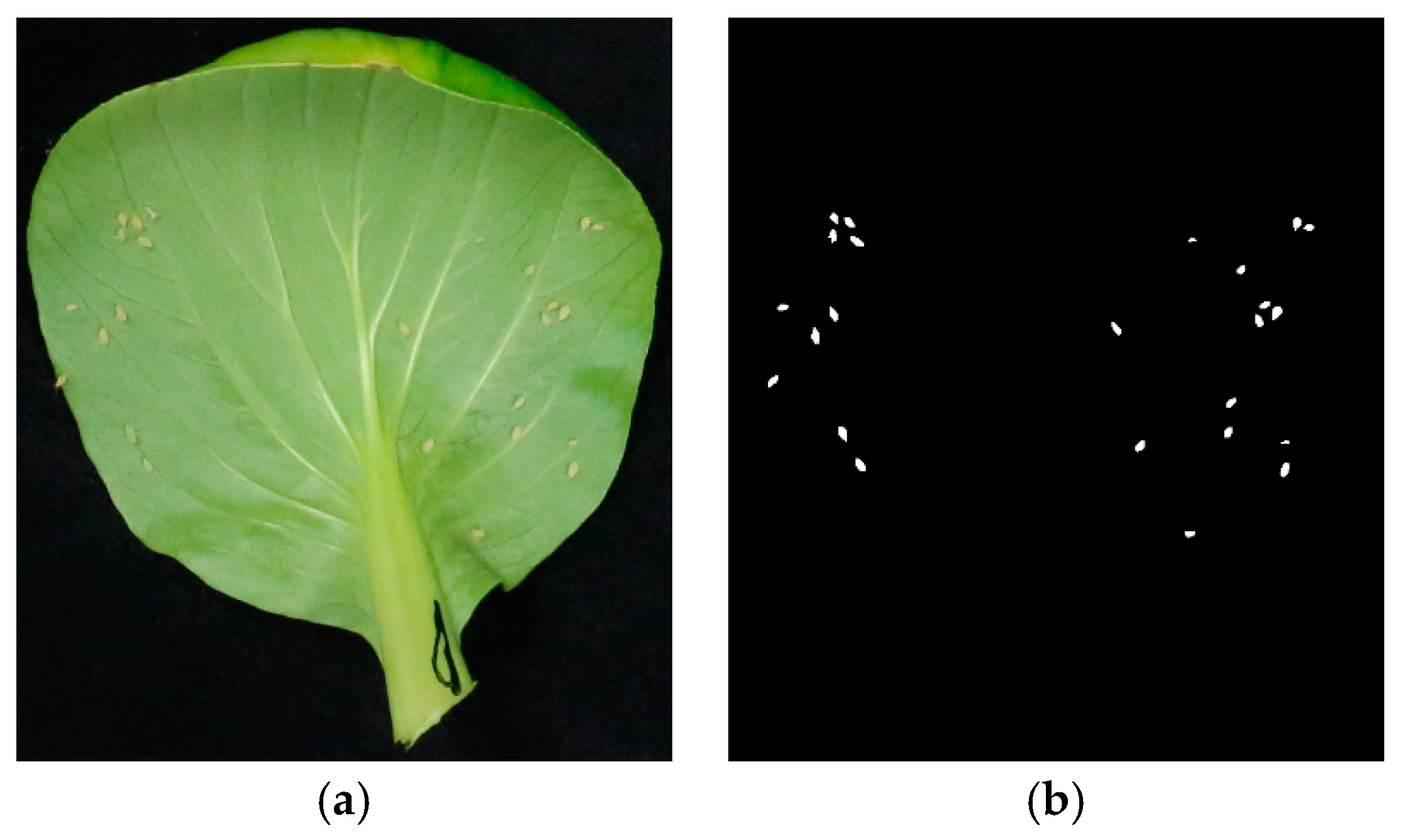
2.1.2. Image Annotation
2.2. Proposed CNN for Segmentation
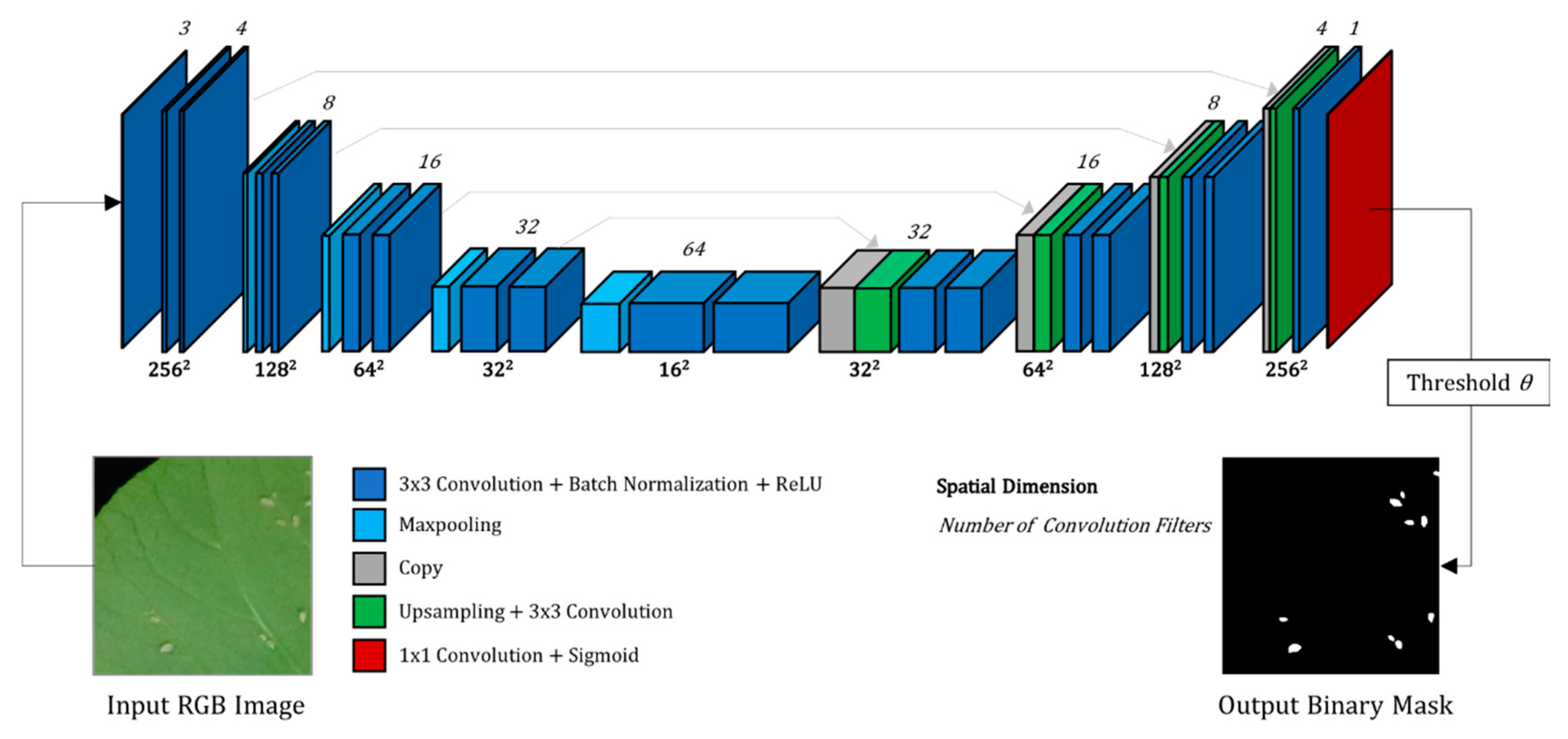
2.2.1. The CNN Architecture
2.2.2. Transfer Learning with Pretrained Contracting Path
2.2.3. CNN Training
2.2.4. CNN Prediction
2.2.5. CNN Implementation and Experimental Setup
2.3. Performance Evaluation
2.3.1. Evaluation on Annotated Test Dataset
2.3.2. In Field Evaluation
2.4. CNN Architecture Optimization
3. Results
3.1. Performance of CNN-4-4
3.1.1. Overall Performance
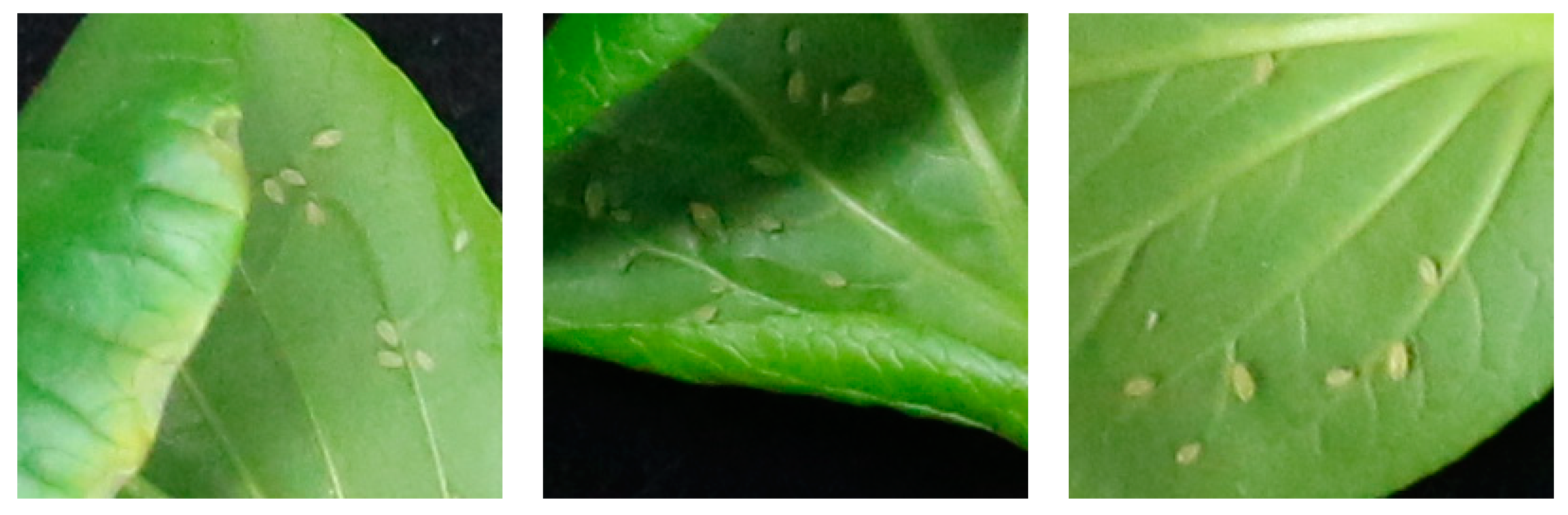
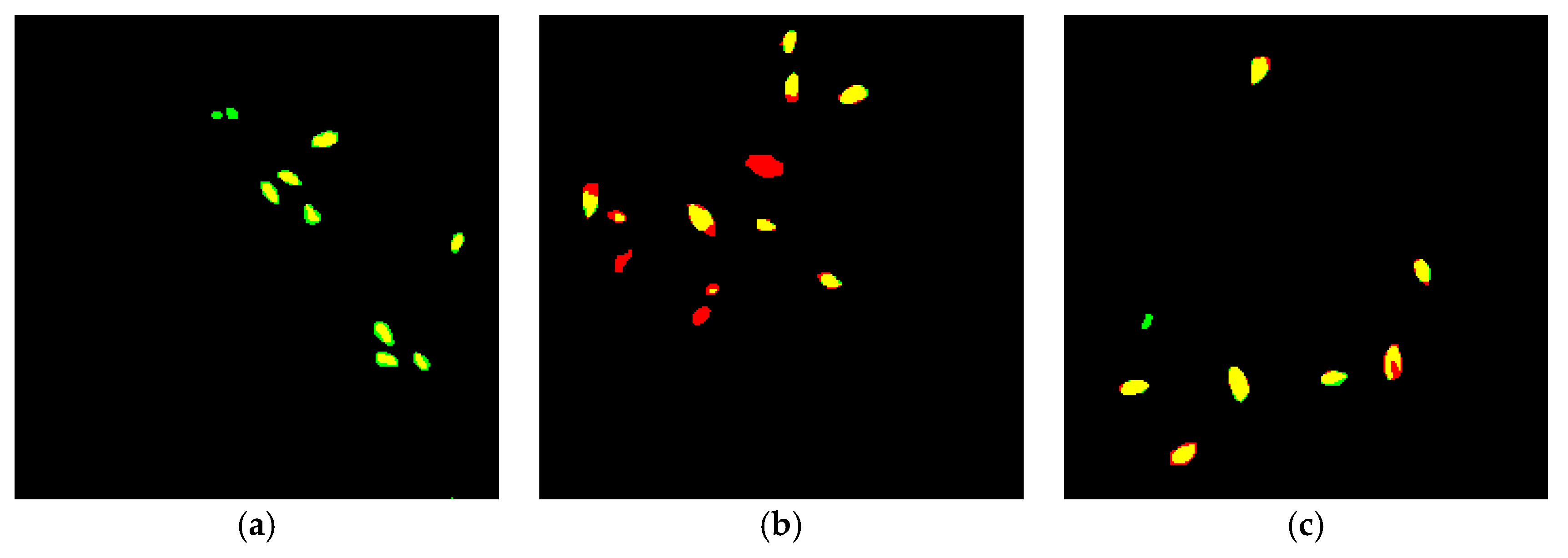
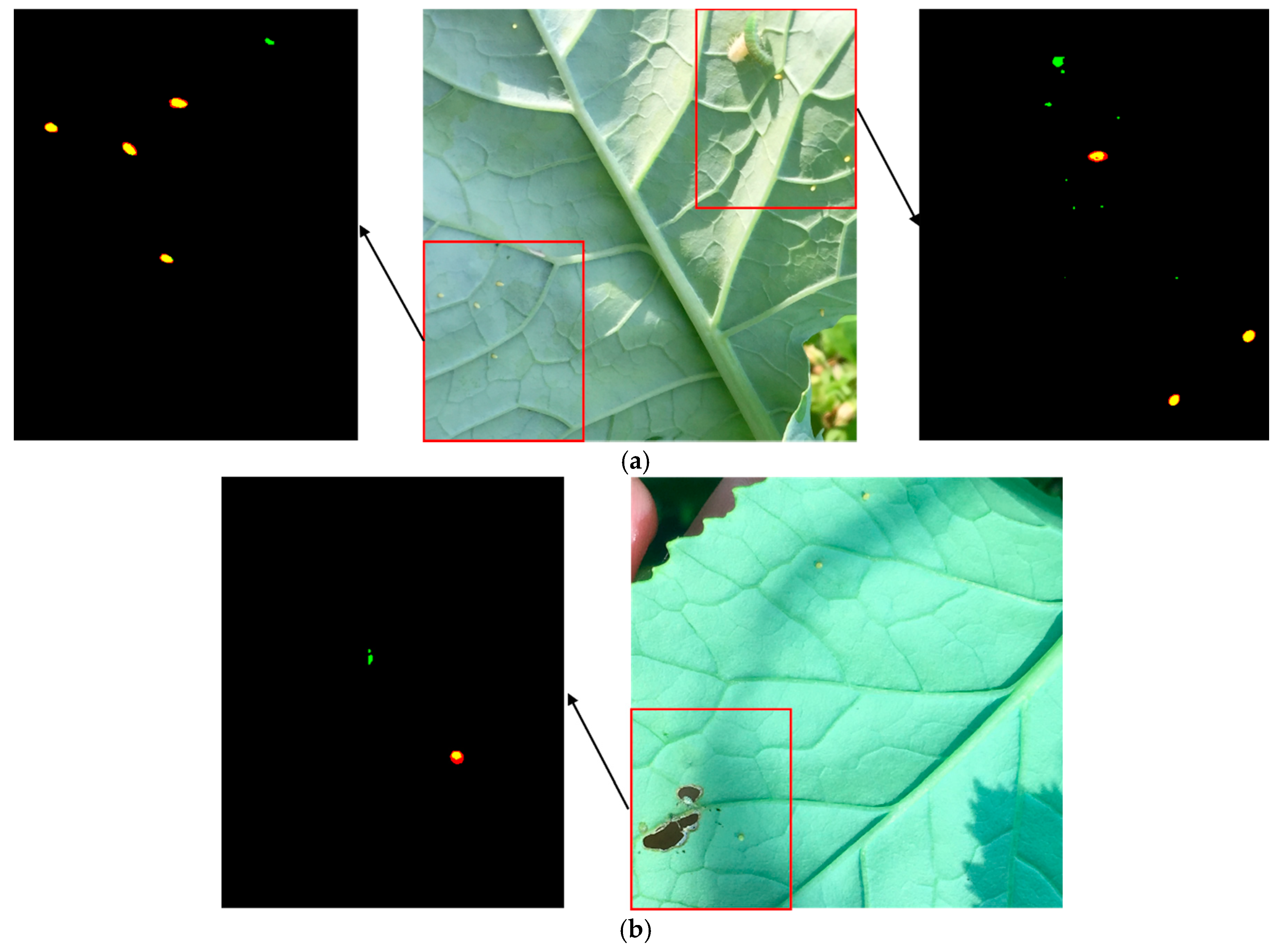
3.1.2. Typical Failure Cases of CNN Segmentation
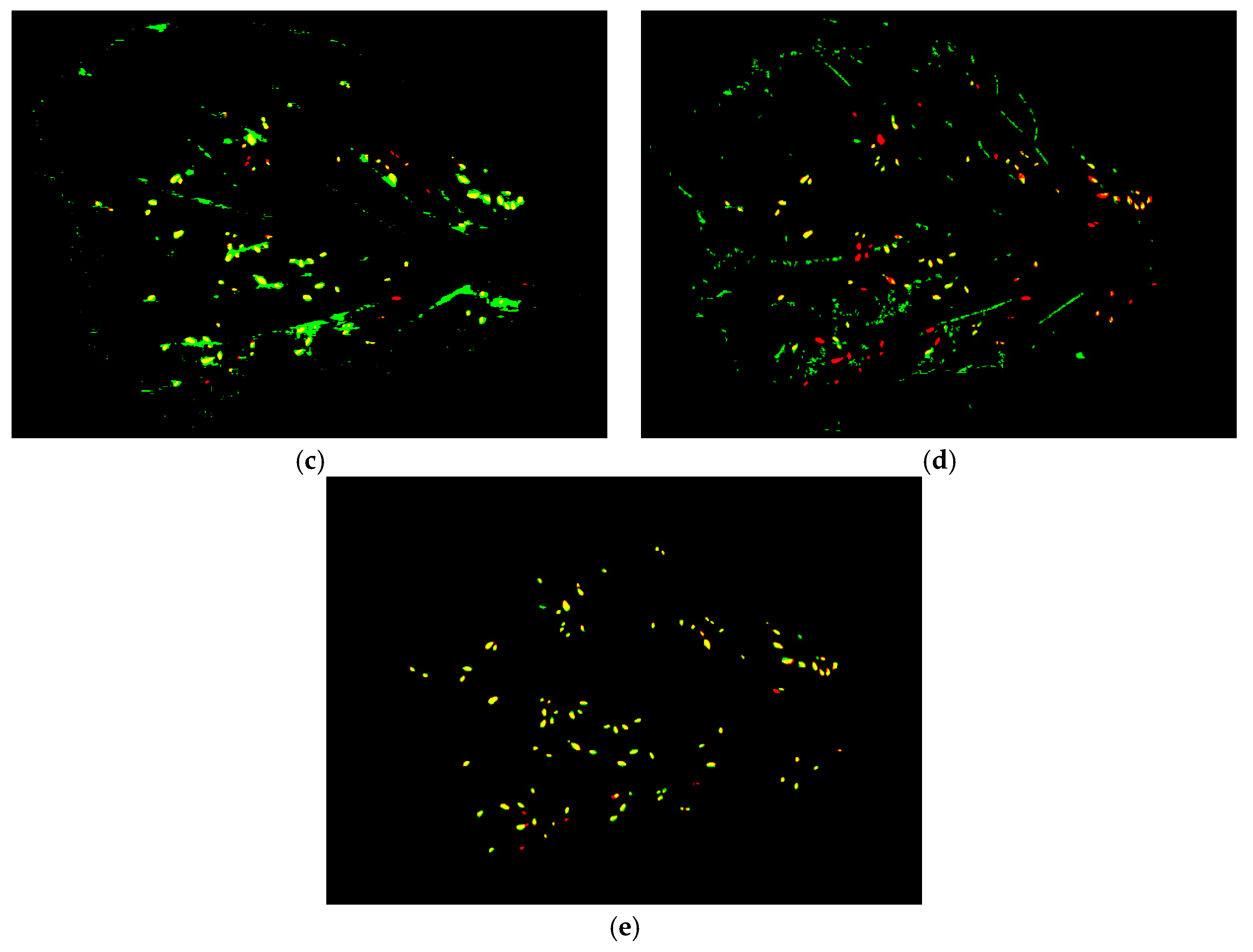
3.2.3. Performance on In-Field Images
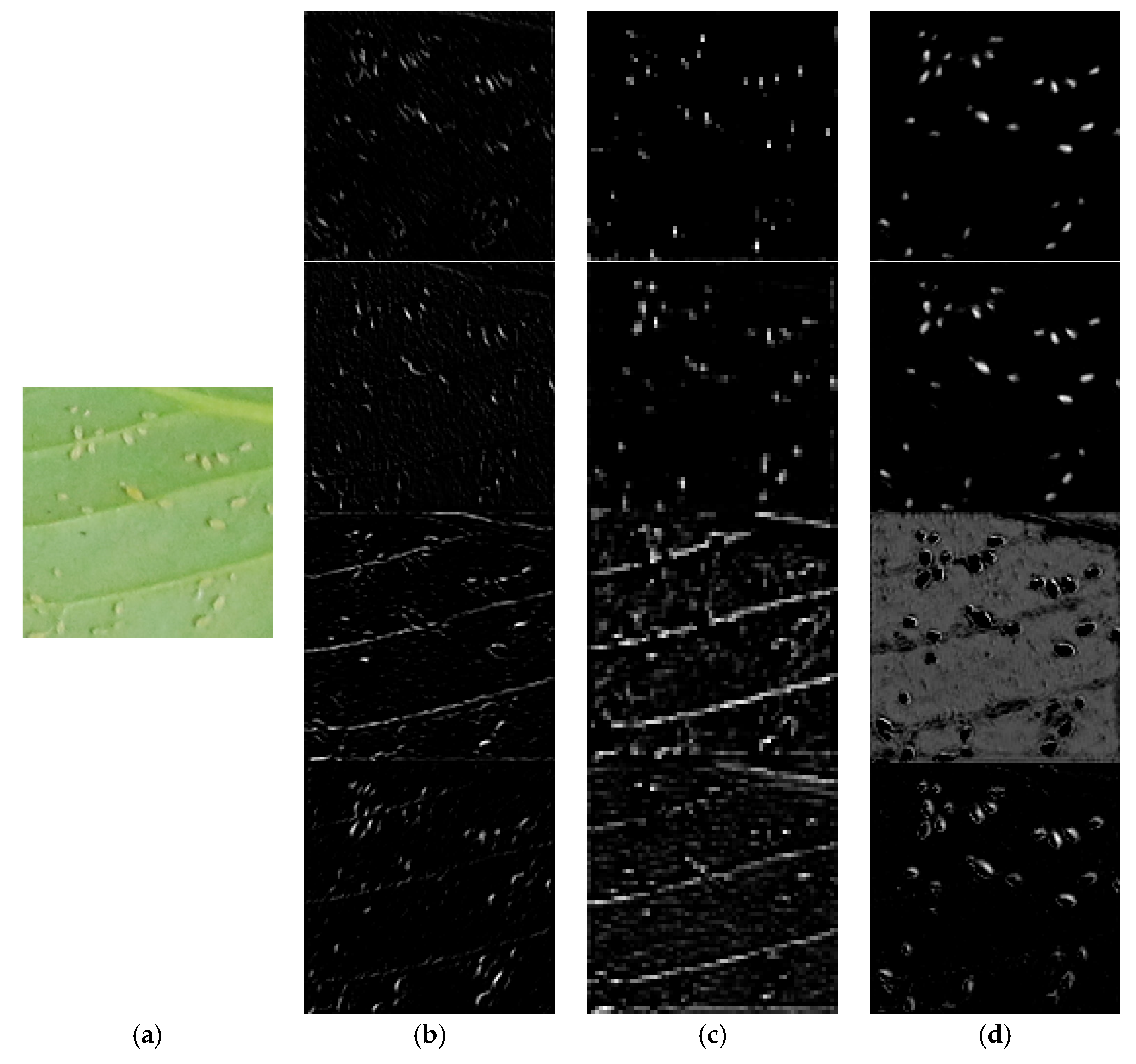
3.2.4. Visualization of CNN Feature Maps
3.2. Performance of Transfer Learning
3.3. Performance of CNNs with Different Sizes
4. Discussion and Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Malais, M.H.; Ravensberg, W.J. Knowing and Recognizing: The Biology of Glasshouse Pests and Their Natural Enemies; Koppert BV: Berkel en Rodenrijs, The Netherlands, 2004. [Google Scholar]
- Sun, Y.; Cheng, H.; Cheng, Q.; Zhou, H.; Li, M.; Fan, Y.; Shan, G.; Damerow, L.; Lammers, P.S.; Jones, S.B. A smart-vision algorithm for counting whiteflies and thrips on sticky traps using two-dimensional Fourier transform spectrum. Biosyst. Eng. 2017, 153, 82–88. [Google Scholar] [CrossRef]
- Heinz, K.M.; Parrella, M.P.; Newman, J.P. Time-efficient use of yellow sticky traps in monitoring insect populations. J. Econ. Entomol. 1992, 85, 2263–2269. [Google Scholar] [CrossRef]
- Park, J.-J.; Kim, J.-K.; Park, H.; Cho, K. Development of time-efficient method for estimating aphids density using yellow sticky traps in cucumber greenhouses. J. Asia-Pac. Entomol. 2001, 4, 143–148. [Google Scholar] [CrossRef]
- Espinoza, K.; Valera, D.L.; Torres, J.A.; López, A.; Molina-Aiz, F.D. Combination of image processing and artificial neural networks as a novel approach for the identification of Bemisia tabaci and Frankliniella occidentalis on sticky traps in greenhouse agriculture. Comput. Electron. Agric. 2016, 127, 495–505. [Google Scholar] [CrossRef]
- Xia, C.; Chon, T.-S.; Ren, Z.; Lee, J.-M. Automatic identification and counting of small size pests in greenhouse conditions with low computational cost. Ecol. Inform. 2015, 29, 139–146. [Google Scholar] [CrossRef]
- Cho, J.; Choi, J.; Qiao, M.; Ji, C.; Kim, H.; Uhm, K.; Chon, T. Automatic identification of whiteflies, aphids and thrips in greenhouse based on image analysis. Red 2007, 346, 244. [Google Scholar]
- Barbedo, J.G.A. Using digital image processing for counting whiteflies on soybean leaves. J. Asia-Pac. Entomol. 2014, 17, 685–694. [Google Scholar] [CrossRef]
- Maharlooei, M.; Sivarajan, S.; Bajwa, S.G.; Harmon, J.P.; Nowatzki, J. Detection of soybean aphids in a greenhouse using an image processing technique. Comput. Electron. Agric. 2017, 132, 63–70. [Google Scholar] [CrossRef]
- Solis-Sánchez, L.; García-Escalante, J.; Castañeda-Miranda, R.; Torres-Pacheco, I.; Guevara-González, R. Machine vision algorithm for whiteflies (Bemisia tabaci Genn.) scouting under greenhouse environment. J. Appl. Entomol. 2009, 133, 546–552. [Google Scholar] [CrossRef]
- Solis-Sánchez, L.O.; Castañeda-Miranda, R.; García-Escalante, J.J.; Torres-Pacheco, I.; Guevara-González, R.G.; Castañeda-Miranda, C.L.; Alaniz-Lumbreras, P.D. Scale invariant feature approach for insect monitoring. Comput. Electron. Agric. 2011, 75, 92–99. [Google Scholar] [CrossRef]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. In Advances in Neural Information Processing Systems 25; Pereira, F., Burges, C.J.C., Bottou, L., Weinberger, K.Q., Eds.; Curran Associates, Inc.: Red Hook, NY, USA, 2012; pp. 1097–1105. [Google Scholar]
- Long, J.; Shelhamer, E.; Darrell, T. Fully convolutional networks for semantic segmentation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 3431–3440. [Google Scholar]
- Ren, S.; He, K.; Girshick, R.; Sun, J. Faster R-CNN: Towards Real-Time Object Detection with Region Proposal Networks. In Advances in Neural Information Processing Systems 28; Cortes, C., Lawrence, N.D., Lee, D.D., Sugiyama, M., Garnett, R., Eds.; Curran Associates, Inc.: Red Hook, NY, USA, 2015; pp. 91–99. [Google Scholar]
- Fischer, P.; Dosovitskiy, A.; Brox, T. Descriptor matching with convolutional neural networks: A comparison to sift. arXiv 2014, arXiv:1405.5769. [Google Scholar]
- Badrinarayanan, V.; Kendall, A.; Cipolla, R. Segnet: A deep convolutional encoder-decoder architecture for image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 39, 2481–2495. [Google Scholar] [CrossRef] [PubMed]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. arXiv 2015, arXiv:1505.04597. [Google Scholar]
- Bai, W.; Sinclair, M.; Tarroni, G.; Oktay, O.; Rajchl, M.; Vaillant, G.; Lee, A.M.; Aung, N.; Lukaschuk, E.; Sanghvi, M.M.; et al. Human-level CMR image analysis with deep fully convolutional networks. arXiv 2017, arXiv:1710.09289. [Google Scholar]
- Chen, H.; Qi, X.; Yu, L.; Dou, Q.; Qin, J.; Heng, P.-A. DCAN: Deep contour-aware networks for object instance segmentation from histology images. Med. Image Anal. 2017, 36, 135–146. [Google Scholar] [CrossRef] [PubMed]
- Saito, S.; Yamashita, T.; Aoki, Y. Multiple Object Extraction from Aerial Imagery with Convolutional Neural Networks. J. Imaging Sci. Technol. 2016, 60, 10402-1–10402-9. [Google Scholar] [CrossRef]
- Tajbakhsh, N.; Shin, J.Y.; Gurudu, S.R.; Hurst, R.T.; Kendall, C.B.; Gotway, M.B.; Liang, J. Convolutional neural networks for medical image analysis: Full training or fine tuning? IEEE Trans. Med. Imaging 2016, 35, 1299–1312. [Google Scholar] [CrossRef] [PubMed]
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. arXiv 2014, arXiv:1409.1556. [Google Scholar]
- Nair, V.; Hinton, G.E. Rectified Linear Units Improve Restricted Boltzmann Machines. In Proceedings of the 27th International Conference on Machine Learning, Haifa, Israel, 21–24 June 2010; pp. 807–814. [Google Scholar]
- Milletari, F.; Navab, N.; Ahmadi, S.-A. V-Net: Fully Convolutional Neural Networks for Volumetric Medical Image Segmentation. arXiv 2016, arXiv:1606.04797. [Google Scholar]
- Ioffe, S.; Szegedy, C. Batch normalization: Accelerating deep network training by reducing internal covariate shift. In Proceedings of the 32nd International Conference on Machine Learning, Lille, France, 6–11 July 2015; pp. 448–456. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Iglovikov, V.; Shvets, A. TernausNet: U-Net with VGG11 Encoder Pre-Trained on ImageNet for Image Segmentation. arXiv 2018, arXiv:1801.05746. [Google Scholar]
- Kingma, D.P.; Ba, J. Adam: A method for stochastic optimization. arXiv 2014, arXiv:1412.6980. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Delving deep into rectifiers: Surpassing human-level performance on imagenet classification. In Proceedings of the IEEE International Conference on Computer Vision, Santiago, Chile, 7–13 December 2015; pp. 1026–1034. [Google Scholar]
- Chollet, F. Keras. Available online: https://keras.io (accessed on 25 July 2018).
- Abadi, M.; Agarwal, A.; Barham, P.; Brevdo, E.; Chen, Z.; Citro, C.; Corrado, G.S.; Davis, A.; Dean, J.; Devin, M.; et al. TensorFlow: Large-Scale Machine Learning on Heterogeneous Systems. arXiv 2015, arXiv:1603.04467. [Google Scholar]
- Van der Walt, S.; Colbert, S.C.; Varoquaux, G. The NumPy array: A structure for efficient numerical computation. Comput. Sci. Eng. 2011, 13, 22–30. [Google Scholar] [CrossRef]
- Paszke, A.; Gross, S.; Chintala, S.; Chanan, G.; Yang, E.; DeVito, Z.; Lin, Z.; Desmaison, A.; Antiga, L.; Lerer, A. Automatic differentiation in PyTorch. In Proceedings of the 31st Conference on Neural Information Processing Systems (NIPS 2017), Long Beach, CA, USA, 8 December 2017. [Google Scholar]
- Rother, C.; Kolmogorov, V.; Blake, A. “GrabCut”: Interactive Foreground Extraction Using Iterated Graph Cuts. ACM Trans. Graph. 2004, 23, 309–314. [Google Scholar] [CrossRef]





| Dice Score | Mean Count Error | R2 | Precision | Recall | F1 Score | |
|---|---|---|---|---|---|---|
| Method 1 | 0.3683 | 61.5 | 0.50 | 0.8723 | 0.2529 | 0.3969 |
| Method 2 | 0.3271 | 29.4 | 0.56 | 0.5980 | 0.2899 | 0.3905 |
| Proposed method | 0.8207 | 1.2 | 0.99 | 0.9563 | 0.9650 | 0.9606 |
| Weight Initialization | Training Dice Score | Testing Dice Score |
|---|---|---|
| [29] | 0.8620 | 0.8265 |
| VGG-13 with batch normalization | 0.8659 | 0.8453 |
| Architecture | # Parameters (million) | Prediction Time on CPU (seconds) | Training Dice Score | Testing Dice Score |
|---|---|---|---|---|
| CNN-2-4 | 0.026 | 1.05 (±0.128) | 0.8447 | 0.7952 |
| CNN-4-4 | 0.10 | 1.35 (±0.069) | 0.8681 | 0.8242 |
| CNN-8-4 | 0.40 | 1.43 (±0.077) | 0.8759 | 0.8278 |
| CNN-16-4 | 1.60 | 2.77 (±0.371) | 0.8832 | 0.8247 |
| CNN-4-3 | 0.025 | 1.35 (±0.143) | 0.8618 | 0.8185 |
| CNN-4-5 | 0.40 | 1.50 (±0.166) | 0.8702 | 0.8270 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, J.; Fan, Y.; Wang, T.; Zhang, C.; Qiu, Z.; He, Y. Automatic Segmentation and Counting of Aphid Nymphs on Leaves Using Convolutional Neural Networks. Agronomy 2018, 8, 129. https://doi.org/10.3390/agronomy8080129
Chen J, Fan Y, Wang T, Zhang C, Qiu Z, He Y. Automatic Segmentation and Counting of Aphid Nymphs on Leaves Using Convolutional Neural Networks. Agronomy. 2018; 8(8):129. https://doi.org/10.3390/agronomy8080129
Chicago/Turabian StyleChen, Jian, Yangyang Fan, Tao Wang, Chu Zhang, Zhengjun Qiu, and Yong He. 2018. "Automatic Segmentation and Counting of Aphid Nymphs on Leaves Using Convolutional Neural Networks" Agronomy 8, no. 8: 129. https://doi.org/10.3390/agronomy8080129
APA StyleChen, J., Fan, Y., Wang, T., Zhang, C., Qiu, Z., & He, Y. (2018). Automatic Segmentation and Counting of Aphid Nymphs on Leaves Using Convolutional Neural Networks. Agronomy, 8(8), 129. https://doi.org/10.3390/agronomy8080129








