Different Approaches to Produce Transgenic Virus B Resistant Chrysanthemum
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Material
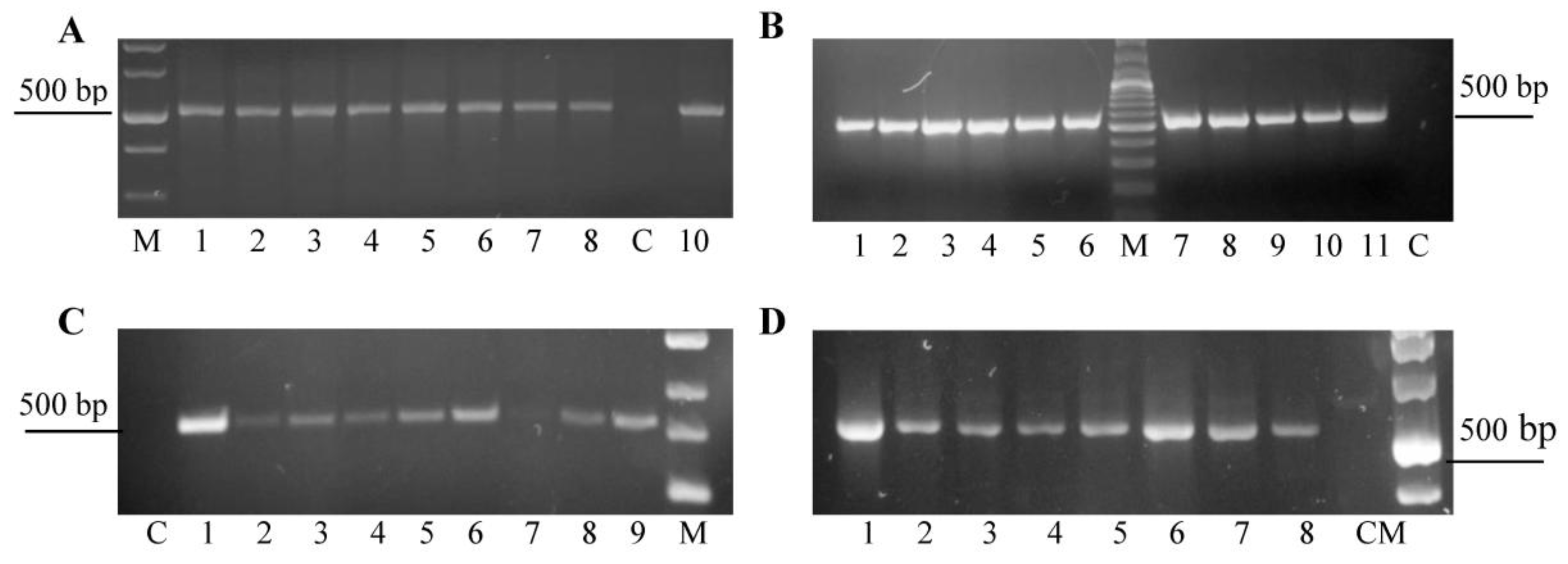
2.2. PCR and RT-PCR Analysis of the Transgenic Plants
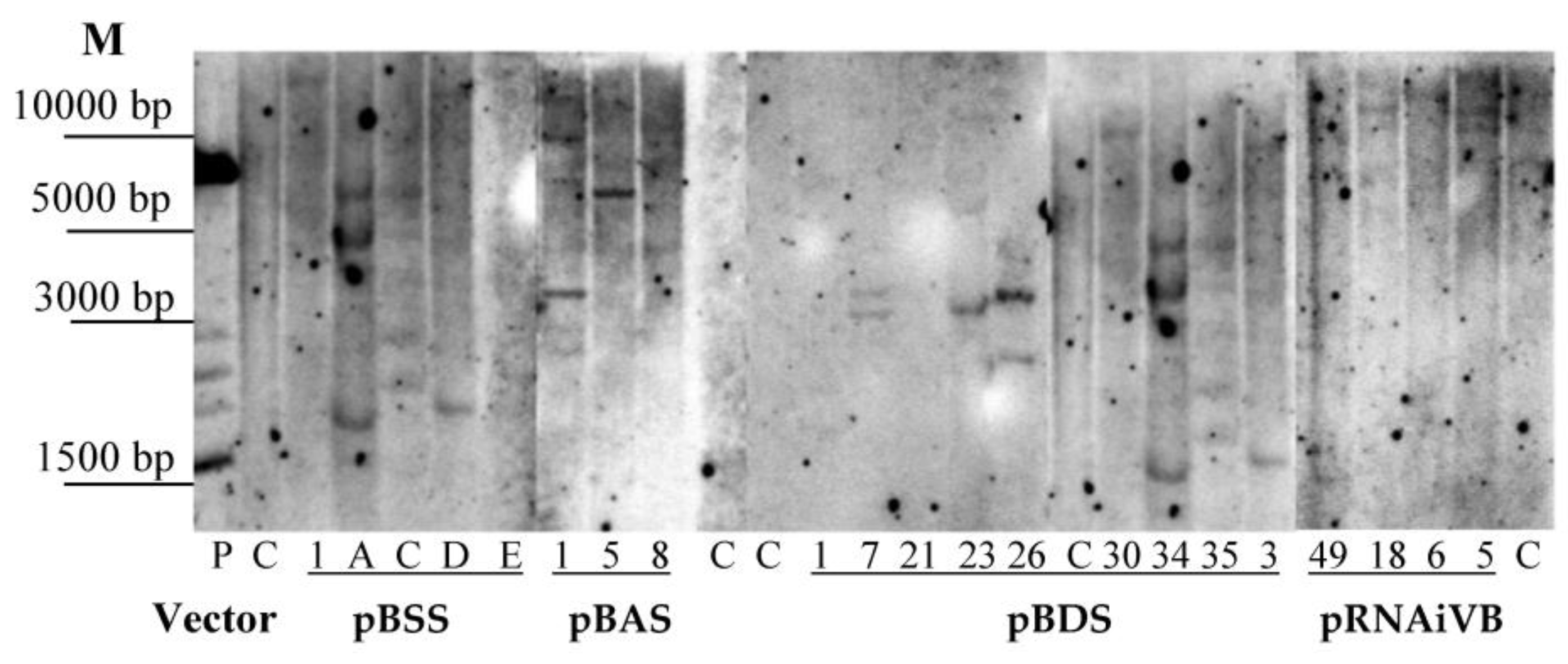
2.3. Southern Blot Analysis
2.4. CVB Resistance Assay
2.5. Isolation of Total Protein and Western Blotting
2.6. DAS-ELISA of CVB
3. Results
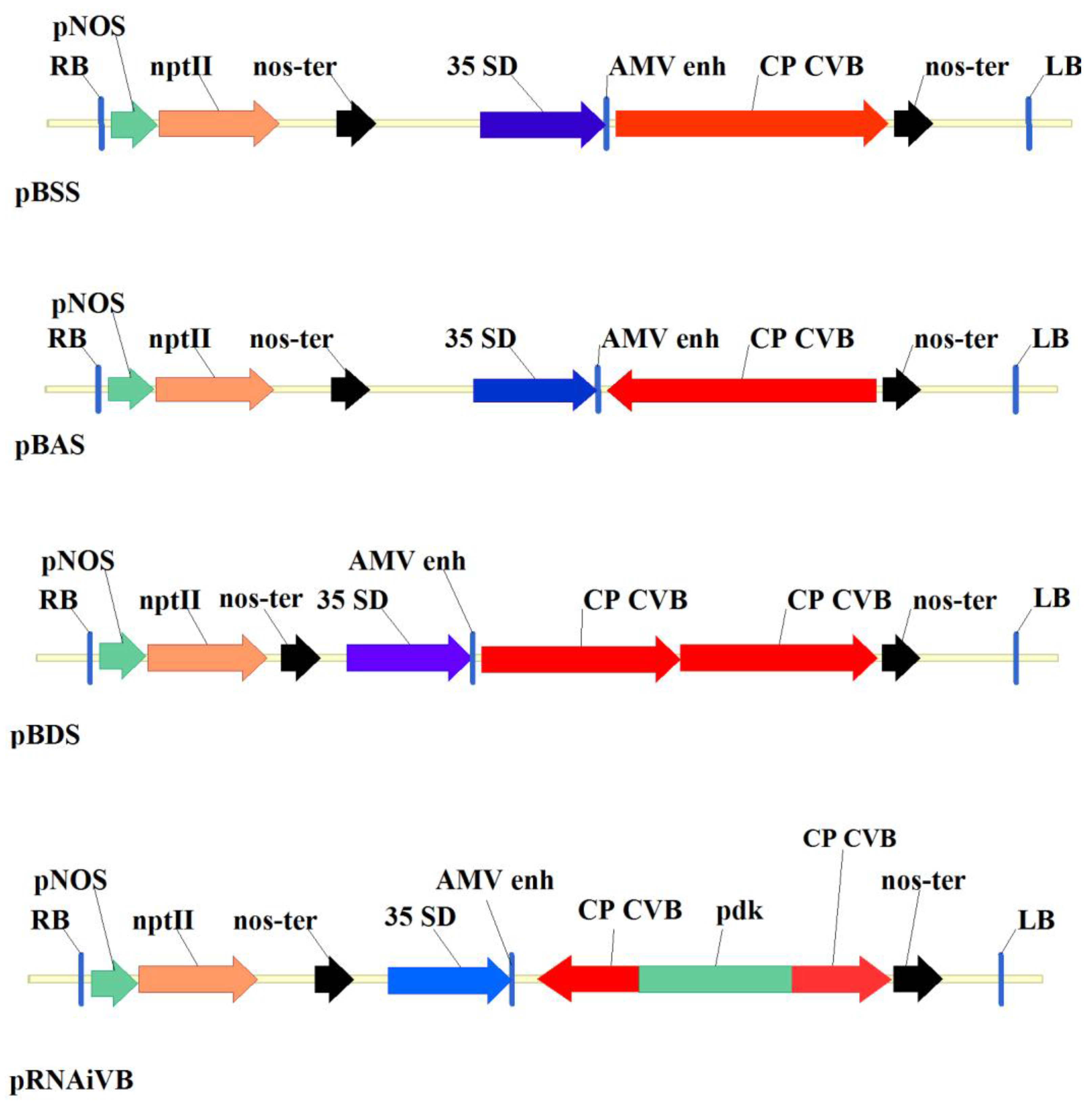
3.1. Agrobacterium-Mediated Transformation of Chrysanthemum Plants with pRNAiVB Vector and PCR-Analisis of Transformants
3.2. Analysis of Integration and Expression of Target Sequencesin Transgenic Plants
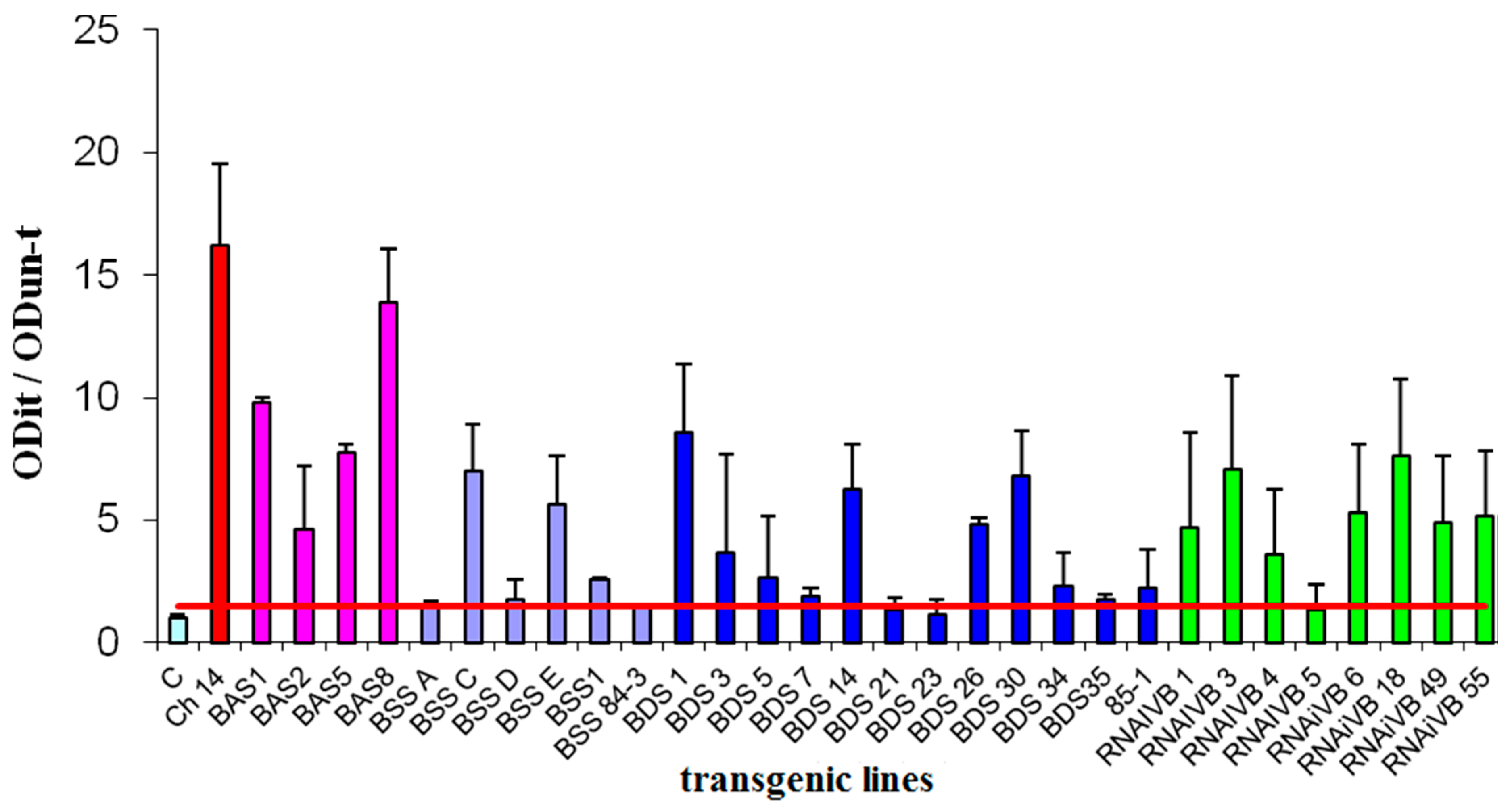
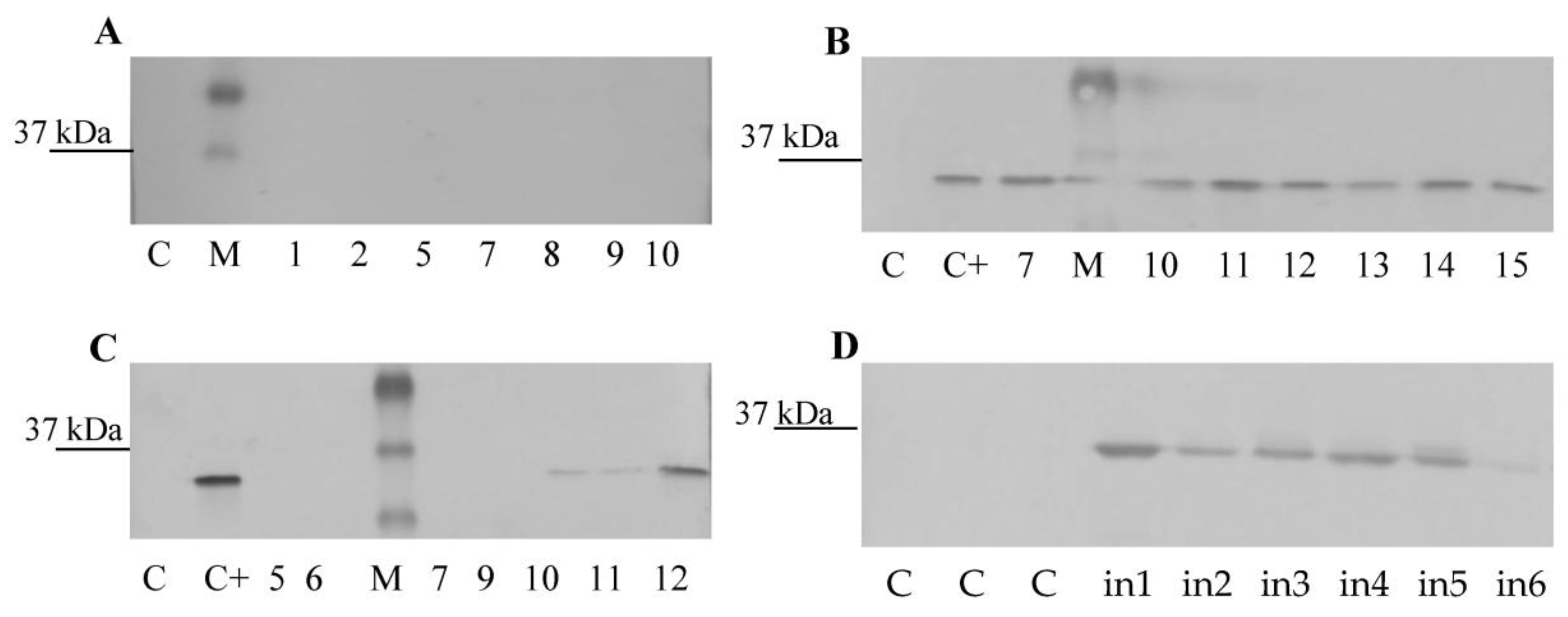
3.3. CVB Resistance Assay and DAS-ELISA and WB Analysis
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Agriculture Victory. Available online: http://agriculture.vic.gov.au/agriculture/pests-diseases-and-weeds/plant-diseases/flowers-and-ornamentals/virus-diseases-of-chrysanthemums (accessed on 5 March 2018).
- Microbial Information Network of China. Available online: http://sdb.im.ac.cn/vide/famly044.htm#Chrysanthemum morifolium (accessed on 5 March 2018).
- Zavriev, S.K.; Kanyuka, K.V.; Levay, K.E. The genome organization of Potato virus M RNA. J. Gen. Virol. 1991, 72, 9–14. [Google Scholar] [CrossRef] [PubMed]
- Lawrence, D.M. Hillman BI Synthesis of infectious transcripts of Blueberry scorch carlavirus in vitro. J. Gen. Virol. 1994, 75, 2509–2512. [Google Scholar] [CrossRef] [PubMed]
- Hataya, T.; Uchino, K.; Arimoto, R.; Suda, N.; Sano, T.; Shikata, E.; Uyeda, I. Molecular characterization of Hop latent virus and phylogenetic relationships among viruses closely related to carlaviruses. Arch. Virol. 2000, 145, 2503–2524. [Google Scholar] [CrossRef] [PubMed]
- Matousek, J.; Schubert, J.; Ptacek, J.; Kozlová, P.; Dědic, P. Complete nucleotide sequence and molecular probing of Potato virus S genome. Acta Virol. 2005, 49, 195–205. [Google Scholar] [PubMed]
- Powell, P.A.; Stark, D.M.; Sanders, P.R.; Beachy, R.N. Protection against tobacco mosaic virus in transgenic plants that express tobacco mosaic virus antisense RNA. Proc. Natl. Acad. Sci. USA 1989, 86, 6949–6952. [Google Scholar] [CrossRef] [PubMed]
- David, C. Baulcombe Mechanisms of Pathogen-Derived Resistance to Viruses in Transgenic Plants. Plant Cell 1996, 8, 1833–1844. [Google Scholar]
- Nelson, A.; Roth, D.A.; Johnson, D. Tobacco mosaic virus infection of transgenic Nicotianatabacum plants is inhibited by antisense constructs directed at the 5′ region of viral RNA. Gene 1993, 127, 227–232. [Google Scholar] [CrossRef]
- Waterhouse, P.M.; Graham, M.W.; Wang, M.-B. Virus resistance and gene silencing in plants can be induced by simultaneous expression of sense and antisense RNA. Proc. Natl. Acad. Sci. USA 1998, 95, 13959–13964. [Google Scholar] [CrossRef] [PubMed]
- Eamens, A.; Wang, M.-B.; Smith, N.A.; Waterhouse, P.M. RNA Silencing in Plants: Yesterday, Today, and Tomorrow. Plant Physiol. 2008, 147, 456–468. [Google Scholar] [CrossRef] [PubMed]
- Agrawal, N.; Dasaradhi, P.V.; Mohmmed, A.; Malhotra, P.; Bhatnagar, R.K.; Mukherjee, S.K. RNA Interference: Biology, Mechanism, and Applications. Microbiol. Mol. Biol. Rev. 2003, 67, 657–685. [Google Scholar] [CrossRef] [PubMed]
- Kurth, E.G.; Peremyslov, V.V.; Prokhnevsky, A.I.; Kasschau, K.D.; Miller, M.; Carrington, J.C.; Dolja, V.V. Virus-Derived Gene Expression and RNA Interference Vector for Grapevine. J. Virol. 2012, 86, 6002–6009. [Google Scholar] [CrossRef] [PubMed]
- Vassilakos, N. Stability of Transgenic Resistance against Plant Viruses. In Transgenic Plants—Advances and Limitations; Çiftçi, Y.O., Ed.; InTech: Rijeka, Croatia; Shanghai, China, 2012. [Google Scholar] [CrossRef]
- Savenkov, E.I.; Valkonen, J.P.T. Coat protein gene-mediated resistance to Potato virus A in transgenic plants is suppressed following infection with another potyvirus. J. Gen. Virol. 2001, 82, 2275–2278. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Gao, S.; Lin, Q.; Wang, H.; Que, Y.; Xu, L. Transgenic Sugarcane Resistant to Sorghum mosaic virus Based on Coat Protein Gene Silencing by RNA Interference. BioMed Res. Int. 2015, 2015, 861907. [Google Scholar] [CrossRef] [PubMed]
- Skachkova, T.S.; Mitiouchkina, T.Y.; Taran, S.A.; Dolgov, S.V. Molecular biology approach for improving chrysanthemum resistance to virus B. Acta Hortic. 2006, 714, 185–192. [Google Scholar] [CrossRef]
- Mitiouchkina, T.; Skachkova, T.; Shulga, O.; Dolgov, S. Expression of TheChrysanthemum Virus BCoat Protein Gene in Transgenic Chrysanthemum. Acta Hortic. 2011, 901, 95–98. [Google Scholar] [CrossRef]
- Van Engelen, F.A.; Molthoff, J.W.; Conner, A.J.; Nap, J.-P.; Pereira, A.; Stiekema, W.J. pBINPLUS: An improved plant transformation vector based on pBIN19. Transgenic Res. 1995, 4, 288–290. [Google Scholar] [CrossRef] [PubMed]
- Mitiouchkina, T.Y.; Dolgov, S.V. Modification of chrysanthemum plant and flower architecture by rolC gene from Agrobacterium rhizogenes introduction. Acta Hortic. 2000, 508, 163–172. [Google Scholar] [CrossRef]
- Rogers, S.O.; Bendich, A.J. Extraction of total cellular DNA from plants, algae and fungi. In Plant Molecular Biology Manual; Gelvin, S.B., Schilperoort, R.A., Eds.; Springer: Dordrecht, The Netherlands, 1994; pp. 183–190. [Google Scholar]
- Kertbundit, S.; Pongtanom, N.; Ruanjan, P.; Chantasingh, D.; Tanwanchai, A.; Panyim, S.; Juříček, M. Resistance of transgenic papaya plants to papaya ringspot virus. Biol. Plant. 2007, 51, 333–339. [Google Scholar] [CrossRef]
- Hefferon, K.L. Transgenic Plants: Biotechnology. In Agricultural Biotechnology; Doelle, H.W., Rokem, S., Eds.; EOLSS Publishers Co. Ltd.: Paris, France, 2009; Volume 8, pp. 144–168. [Google Scholar]
- Mandahar, C.L. Multiplication of RNA Plant Viruses; Springer: Dordrecht, The Netherlands, 2006. [Google Scholar]

| Vector | A Number of the Studied Lines | A Number of Resistant Lines (ODit/ODun-t < 1,4) | A Part of Resistant Lines out of a Total Number of the Studied Lines, % |
|---|---|---|---|
| pBAS | 4 | 0 | 0 |
| pBSS | 6 | 0 | 0 |
| pBDS | 12 | 2 | 16, 7 |
| pRNAiVB | 8 | 1 | 12, 5 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mitiouchkina, T.Y.; Firsov, A.P.; Titova, S.M.; Pushin, A.S.; Shulga, O.A.; Dolgov, S.V. Different Approaches to Produce Transgenic Virus B Resistant Chrysanthemum. Agronomy 2018, 8, 28. https://doi.org/10.3390/agronomy8030028
Mitiouchkina TY, Firsov AP, Titova SM, Pushin AS, Shulga OA, Dolgov SV. Different Approaches to Produce Transgenic Virus B Resistant Chrysanthemum. Agronomy. 2018; 8(3):28. https://doi.org/10.3390/agronomy8030028
Chicago/Turabian StyleMitiouchkina, Tatiana Y., Aleksey P. Firsov, Svetlana M. Titova, Alexander S. Pushin, Olga A. Shulga, and Sergey V. Dolgov. 2018. "Different Approaches to Produce Transgenic Virus B Resistant Chrysanthemum" Agronomy 8, no. 3: 28. https://doi.org/10.3390/agronomy8030028
APA StyleMitiouchkina, T. Y., Firsov, A. P., Titova, S. M., Pushin, A. S., Shulga, O. A., & Dolgov, S. V. (2018). Different Approaches to Produce Transgenic Virus B Resistant Chrysanthemum. Agronomy, 8(3), 28. https://doi.org/10.3390/agronomy8030028




