Field Phenotyping and Long-Term Platforms to Characterise How Crop Genotypes Interact with Soil Processes and the Environment
Abstract
:1. Introduction
2. Field Based Assessment of Impacts of Management and Environment on Crop Production
2.1. Long-Term Platforms
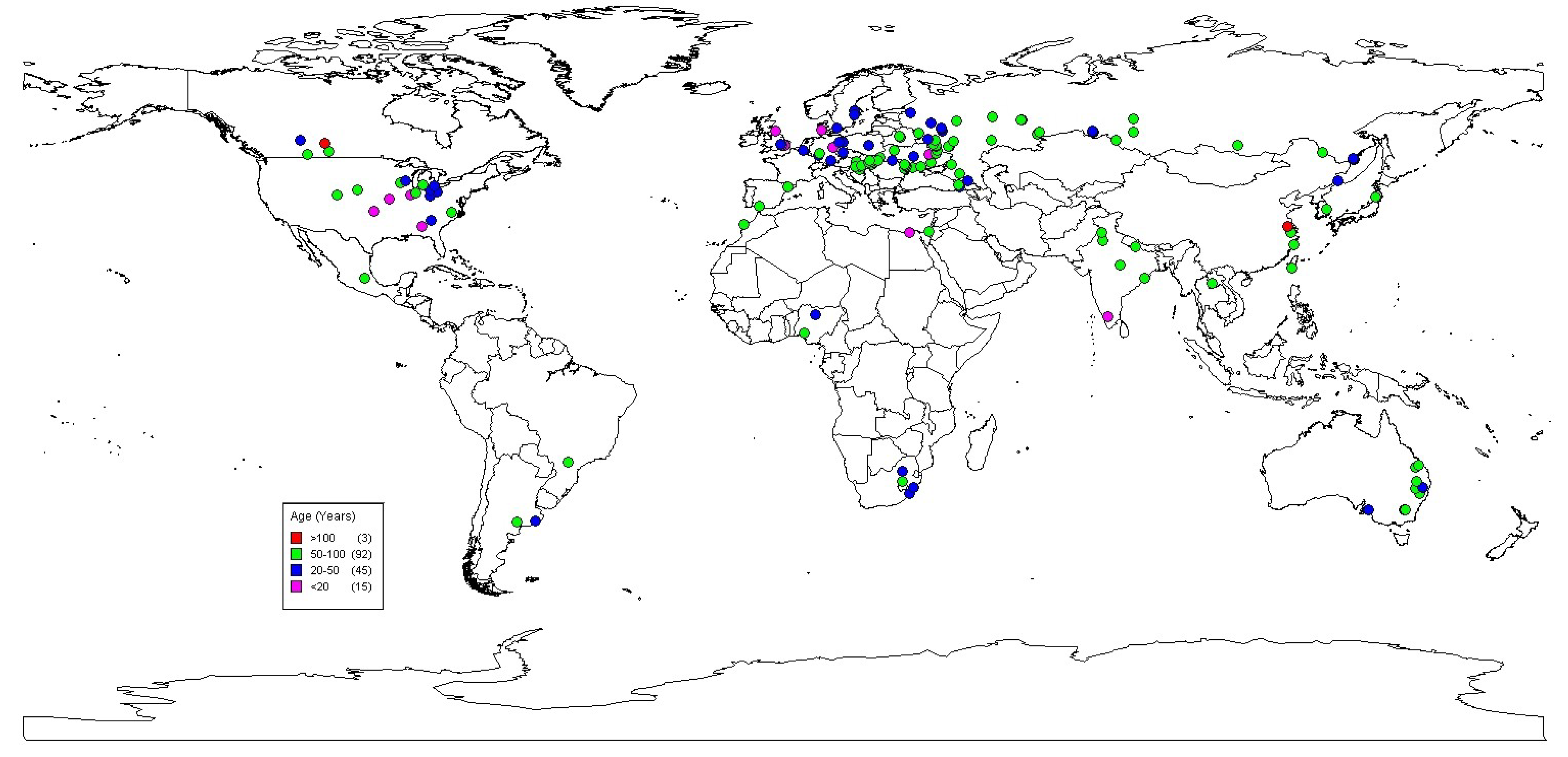
2.1.1. Centre for Sustainable Cropping (CSC), The James Hutton Institute, East Central Scotland
2.1.2. North Wyke Farm Platform (NWFP), Rothamsted, South West England
2.1.3. NIAB (National Institute for Agricultural Botany) New Farm Systems (NFS) and Sustainability Trial in Arable Rotations (STAR), SE England
2.1.4. ORE-ACBB (Observatory for Environmental Research—Agroecosystem Biochemical Cycles and Biodiversity), INRA, France
2.2. Farm Scale Evaluations
3. Methods for Phenotyping Plants and Crop Response to Management and Environment
3.1. Invasive Methods
| Phenotyping Method | Target | Field Applicability | Other Comments |
|---|---|---|---|
| Flat-bed scanner | Roots/leaves | Requires destructive harvesting of sample for field use | Cheap technology and simple to use. Could be used from the back of a trailer in field but will always be destructive for root samples |
| Rhizotron imaging in tubes | Roots | Tubes require installation prior to extensive crop growth, therefore limited to small scale experiments | |
| X-ray/MRI | Roots | Only applicable to samples removed and scanned ex situ | |
| DNA extraction for biomass analysis | Roots | Requires coring for soil samples and destructive extraction of root DNA | Requires extensive calibration of samples to specific soil type and number of sequence copies if multiple genotypes are being investigated |
| Light based imaging | Shoots | Can be done using simple camera systems or via imaging attached to trailers | Can be very simple to use but can suffer from variable light conditions/shadowing and reflectance causing downstream image analysis issues. |
| Infra-Red methodologies | Shoots via root linked physiology | Has been used at a range of scales from individual leaves to field scale. | Is affected by variable temperature When insinuating links to roots size careful interpretation of correlations is need |
| Laser scanning Biomass | Shoot shape/biomass | ||
| Capacitance | Roots | Measures individual plants system requires plant contact | Originally considered to measure root biomass but now shown to measure a cross section of roots diameter at soil contact |
| Electromagnetic | Roots | Requires wires across field but suitable for small plots. Non-contact therefore application to field use |
3.2. Non-Invasive Methods
3.3. Translation of Methods for in Field Phenotyping
3.4. Characterising Soil Conditions for Plant Growth in the Field
4. Specific Crop Trait Requirements of Different Management Systems
4.1. Organic and Low-Input Systems
4.2. High-Tech Systems including Precision Agriculture and Controlled Traffic
4.3. Polyculture vs. Monoculture
5. How Do We Deal with G × E × M in the Field?
5.1. Timescales for Establishment and Assessment
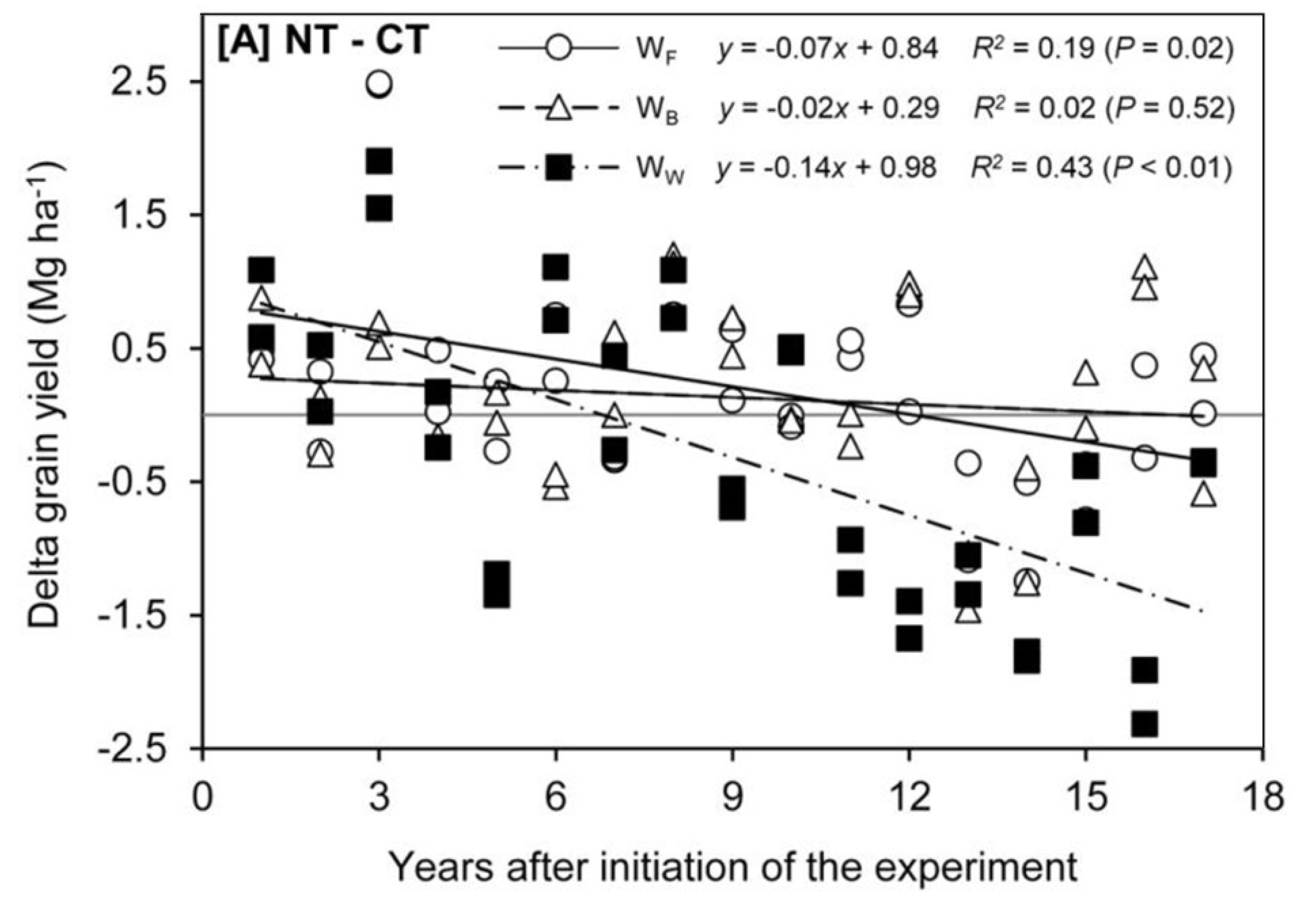
5.2. Requirement for Crop Testing in Multiple Environments
6. Modelling for Better Management of Plants in the Field
7. Translation of Results from LTEs and Phenotyping Platforms to Sustainable Production in Farmers’ Fields
8. Conclusions
- There is a need for a second green revolution where food security is achieved in a sustainable way.
- Testing genotypes in realistic field environments is critical to achieving translation of the understanding of traits at the genetic level to sustainable production in the real world.
- There is a large number of LTEs available to researchers in a range of production systems. However, these are mostly in the developed world. With the major global issues of food security and population pressure likely to be played out in the developing world it is important to ask whether we have enough information available from LTEs in the systems where the real pressures will be felt.
- There is a whole range of high- and low-tech options available for field phenotyping of plants to allow selection of appropriate traits and genotypes for a range of systems. However, systems need to be put in place that allow the integration of phenotyping data and environmental data including soil and climatic conditions from different platforms and this poses a data handling challenge which is yet to be solved.
- There is a whole range of agricultural systems which are poised to respond to environmental change through replacing conventional agriculture with high-tech solutions, low-input systems and polycultures. However, a debate needs to be had as to whether separate breeding programmes are needed to design genotypes (or ideotypes) of crops for these specific environments. This is where field phenotyping and long-term platforms that take account of seasonal and environmental variation have a role to play.
- Modelling has a role to play in both predicting the impact of changes to agricultural systems within the landscape of future environmental change and in the design of crop ideotypes for future agricultural sustainability.
- The translation of agronomic practices and beneficial genotypes from research stations to real world farmers’ fields is often poor and represents a significant bottleneck in improving agricultural sustainability practice. It is vital that we understand fully the G × E × M interaction so that the advances being made in plant genetics can be translated into a benefit in the real world.
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Lynch, J.P. Roots of the second green revolution. Aust. J. Bot. 2007, 55, 493–512. [Google Scholar] [CrossRef]
- Swaminathan, M.S. Can science and technology feed the world in 2025? Field Crops Res. 2007, 104, 3–9. [Google Scholar] [CrossRef]
- Tester, M.; Langridge, P. Breeding technologies to increase crop production in a changing world. Science 2010, 327, 818–822. [Google Scholar] [CrossRef]
- White, P.J.; George, T.S.; Gregory, P.J.; Bengough, A.G.; Hallett, P.D.; McKenzie, B.M. Matching roots to their environment. Ann. Bot. 2013, 112, 207–222. [Google Scholar] [CrossRef]
- White, P.J.; George, T.S.; Dupuy, L.X.; Karley, A.J.; Valentine, T.A.; Wiesel, L.; Wishart, J. Root traits for infertile soils. Front. Plant Sci. 2013, 4, 193–193. [Google Scholar]
- Pinheiro de Carvalho, M.A.A.; Bebeli, P.J.; Bettencourt, E.; Costa, G.; Dias, S.; dos Santos, T.M.M.; Slaski, J.J. Cereal landraces genetic resources in worldwide genebanks. A review. Agron. Sustain. Dev. 2013, 33, 177–203. [Google Scholar] [CrossRef]
- Monaco, M.K.; Stein, J.; Naithani, S.; Wei, S.; Dharmawardhana, P.; Kumari, S.; Amarasinghe, V.; Youens-Clark, K.; Thomason, J.; Preece, J.; et al. Gramene 2013: Comparative plant genomics resources. Nucleic Acids Res. 2014, 42, D1193–D1199. [Google Scholar] [CrossRef]
- Mujeeb-Kazi, A.; Kazi, A.G.; Dundas, I.; Rasheed, A.; Ogbonnaya, F.; Kishii, M.; Bonnett, D.; Wang, R.R.C.; Xu, S.; Chen, P.; et al. Genetic diversity for wheat improvement as a conduit to food security. Adv. Agron. 2013, 122, 179–257. [Google Scholar] [CrossRef]
- Chochois, V.; Vogel, J.P.; Watt, M. Application of brachypodium to the genetic improvement of wheat roots. J. Exp. Bot. 2012, 63, 3467–3474. [Google Scholar] [CrossRef]
- Gregory, P.J.; Atkinson, C.J.; Bengough, A.G.; Else, M.A.; Fernandez-Fernandez, F.; Harrison, R.J.; Schmidt, S. Contributions of roots and rootstocks to sustainable, intensified crop production. J. Exp. Bot. 2013, 64, 1209–1222. [Google Scholar] [CrossRef]
- Wurschum, T. Mapping QTL for agronomic traits in breeding populations. Theor. Appl. Genet. 2012, 125, 201–210. [Google Scholar] [CrossRef]
- Paulus, S.; Dupuis, J.; Mahlein, A.-K.; Kuhlmann, H. Surface feature based classification of plant organs from 3D laserscanned point clouds for plant phenotyping. BMC Bioinform. 2013, 14. [Google Scholar] [CrossRef]
- Mahlein, A.-K.; Oerke, E.-C.; Steiner, U.; Dehne, H.-W. Recent advances in sensing plant diseases for precision crop protection. Eur. J. Plant Pathol. 2012, 133, 197–209. [Google Scholar] [CrossRef]
- Mahlein, A.-K.; Steiner, U.; Hillnhuetter, C.; Dehne, H.-W.; Oerke, E.-C. Hyperspectral imaging for small-scale analysis of symptoms caused by different sugar beet diseases. Plant Methods 2012, 8. [Google Scholar] [CrossRef]
- Broadley, M.R.; White, P.J.; Hammond, J.P.; Zelko, I.; Lux, A. Zinc in plants. New Phytol. 2007, 173, 677–702. [Google Scholar] [CrossRef]
- Ismail, A.M.; Heuer, S.; Thomson, M.J.; Wissuwa, M. Genetic and genomic approaches to develop rice germplasm for problem soils. Plant Mol. Biol. 2007, 65, 547–570. [Google Scholar] [CrossRef]
- White, P.J.; Brown, P.H. Plant nutrition for sustainable development and global health. Ann. Bot. 2010, 105, 1073–1080. [Google Scholar] [CrossRef]
- Vonuexkull, H.R.; Mutert, E. Global extent, development and economic-impact of acid soils. Plant Soil 1995, 171, 1–15. [Google Scholar] [CrossRef]
- Sumner, M.E.; Noble, A.D. Soil Acidification: The World Story. In Handbook of Soil Acidity; Rengel, Z., Ed.; Marcel Dekker Inc.: New York, NY, USA, 2003; pp. 1–28. [Google Scholar]
- White, P.J.; Broadley, M.R. Biofortification of crops with seven mineral elements often lacking in human diets—Iron, zinc, copper, calcium, magnesium, selenium and iodine. New Phytol. 2009, 182, 49–84. [Google Scholar] [CrossRef]
- Bennett, S.J.; Barrett-Lennard, E.G.; Colmer, T.D. Salinity and waterlogging as constraints to saltland pasture production: A review. Agric. Ecosyst. Environ. 2009, 129, 349–360. [Google Scholar] [CrossRef]
- Munns, R.; Tester, M. Mechanisms of salinity tolerance. Annu. Rev. Plant Biol. 2008, 59, 651–681. [Google Scholar] [CrossRef]
- White, P.J.G.; Greenwood, D.J. Properties and management of cationic elements for crop growth. In Russell’s Soil Conditions and Plant Growth, 12th ed.; Gregory, P.J., Nortcliff, S., Eds.; Wiley-Blackwell: Oxford, UK, 2013. [Google Scholar]
- Lynch, J.P. Root phenes for enhanced soil exploration and phosphorus acquisition: Tools for future crops. Plant Physiol. 2011, 156, 1041–1049. [Google Scholar] [CrossRef]
- Fageria, N.K.; Baligar, V.C.; Jones, C.A. Growth and Mineral Nutrition of Field Crops; CRC Press: Boca Raton, FL, USA, 2011. [Google Scholar]
- Mueller, N.D.; Gerber, J.S.; Johnston, M.; Ray, D.K.; Ramankutty, N.; Foley, J.A. Closing yield gaps through nutrient and water management. Nature 2012, 490, 254–257. [Google Scholar] [CrossRef]
- Bengough, A.G.; Bransby, M.F.; Hans, J.; McKenna, S.J.; Roberts, T.J.; Valentine, T.A. Root responses to soil physical conditions; growth dynamics from field to cell. J. Exp. Bot. 2006, 57, 437–447. [Google Scholar]
- Valentine, T.A.; Hallett, P.D.; Binnie, K.; Young, M.W.; Squire, G.R.; Hawes, C.; Bengough, A.G. Soil strength and macropore volume limit root elongation rates in many UK agricultural soils. Ann. Bot. 2012, 110, 259–270. [Google Scholar] [CrossRef]
- Hallett, P.D.; Bengough, A.G. Managing the soil physical environment for plants. In Russell’s Soil Conditions and Plant Growth, 12th ed.; Gregory, P.J., Nortcliff, S., Eds.; Wiley-Blackwell: Oxford, UK, 2013. [Google Scholar]
- Richards, R.A.; Watt, M.; Rebetzke, G.J. Physiological traits and cereal germplasm for sustainable agricultural systems. Euphytica 2007, 154, 409–425. [Google Scholar] [CrossRef]
- Jackson, M.B.; Colmer, T.D. Response and adaptation by plants to flooding stress—Preface. Ann. Bot. 2005, 96, 501–505. [Google Scholar] [CrossRef]
- Abdalla, M.; Osborne, B.; Lanigan, G.; Forristal, D.; Williams, M.; Smith, P.; Jones, M.B. Conservation tillage systems: A review of its consequences for greenhouse gas emissions. Soil Use Manag. 2013, 29, 199–209. [Google Scholar] [CrossRef]
- Farage, P.K.; Ardoe, J.; Olsson, L.; Rienzi, E.A.; Ball, A.S.; Pretty, J.N. The potential for soil carbon sequestration in three tropical dryland farming systems of Africa and Latin America: A modelling approach. Soil Tillage Res. 2007, 94, 457–472. [Google Scholar] [CrossRef]
- Llewellyn, R.S.; D’Emden, F.H.; Kuehne, G. Extensive use of no-tillage in grain growing regions of Australia. Field Crops Res. 2012, 132, 204–212. [Google Scholar] [CrossRef]
- McKenzie, B.M.; Bengough, A.G.; Hallett, P.D.; Thomas, W.T.B.; Forster, B.; McNicol, J.W. Deep rooting and drought screening of cereal crops: A novel field-based method and its application. Field Crops Res. 2009, 112, 165–171. [Google Scholar] [CrossRef]
- Watt, M.; Kirkegaard, J.A.; Rebetzke, G.J. A wheat genotype developed for rapid leaf growth copes well with the physical and biological constraints of unploughed soil. Funct. Plant Biol. 2005, 32, 695–706. [Google Scholar] [CrossRef]
- White, J.W.; Andrade-Sanchez, P.; Gore, M.A.; Bronson, K.F.; Coffelt, T.A.; Conley, M.M.; Feldmann, K.A.; French, A.N.; Heun, J.T.; Hunsaker, D.J.; et al. Field-based phenomics for plant genetics research. Field Crops Res. 2012, 133, 101–112. [Google Scholar] [CrossRef]
- Staedler, Y.M.; Masson, D.; Schoenenberger, J. Plant tissues in 3D via X-ray tomography: Simple contrasting methods allow high resolution imaging. PLoS One 2013, 8, e75295. [Google Scholar]
- Shi, L.; Shi, T.X.; Broadley, M.R.; White, P.J.; Long, Y.; Meng, J.L.; Xu, F.S.; Hammond, J.P. High-throughput root phenotyping screens identify genetic loci associated with root architectural traits in brassica napus under contrasting phosphate availabilities. Ann. Bot. 2013, 112, 381–389. [Google Scholar] [CrossRef]
- Petersen, J.; Thomsen, I.K.; Mattsson, L.; Hansen, E.M.; Christensen, B.T. Crop response to sustained reductions in annual nitrogen fertilizer rates using long-term experiments as research platform. In Proceedings of NJF Seminar 407, Askov Experimental Station and Sandbjerg Estate, Jutland, Denmark, 16–18 June 2008; pp. 36–39.
- Horn, R. Division s-1—Soil physics—Time dependence of soil mechanical properties and pore functions for arable soils. Soil Sci. Soc. Am. J. 2004, 68, 1131–1137. [Google Scholar] [CrossRef]
- Powlson, D.S.; Poulton, P.R.; Macdonald, A.J. Long-term experiments as unique resources for research on local and global carbon cycling. In Proceedings of NJF Seminar 407, Askov Experimental Station and Sandbjerg Estate, Jutland, Denmark, 16–18 June 2008; pp. 20–23.
- Johnston, A.E.; Poulton, P.R.; Coleman, K. Soil organic matter: Its importance in sustainable agriculture and carbon dioxide fluxes. Adv. Agron. 2009, 101, 1–57. [Google Scholar] [CrossRef]
- Hofmann, A.; Heim, A.; Christensen, B.T.; Gioacchini, P.; Miltner, A.; Gehre, M.; Schmidt, M.W.I. Tracking the fate of lignin in 13C-labelled arable soils. In Proceedings of NJF Seminar 407, Askov Experimental Station and Sandbjerg Estate, Jutland, Denmark, 16–18 June 2008; pp. 28–31.
- Peterson, M.J. Introduction to the biological monitoring and abatement program. Environ. Manag. 2011, 47, 1005–1009. [Google Scholar] [CrossRef]
- Goulding, K. Long-term research in the UK—Lessons learned from the rothamsted classical experiments. Kungl. Skogs Lantbr. Tidskr. 2007, 146, 8–12. [Google Scholar]
- Korschens, M. The importance of long-term field experiments for soil science and environmental research—A review. Plant Soil Environ. 2006, 52, 1–8. [Google Scholar] [CrossRef]
- Rasmussen, P.E.; Goulding, K.W.T.; Brown, J.R.; Grace, P.R.; Janzen, H.H.; Korschens, M. Agroecosystem—Long-term agroecosystem experiments: Assessing agricultural sustainability and global change. Science 1998, 282, 893–896. [Google Scholar] [CrossRef]
- Franko, U.; Schramm, G.; Rodionova, V.; Korschens, M.; Smith, P.; Coleman, K.; Romanenkov, V.; Shevtsova, L. Eurosomnet—A database for long-term experiments on soil organic matter in Europe. Comput. Electron. Agric. 2002, 33, 233–239. [Google Scholar] [CrossRef]
- LTSE Website, Global Inventory of Long-Term Soil-Ecosystem Experiments. Available online: http://nicholas.duke.edu/ltse/ (accessed on 4 February 2014).
- Petersen, J.; Mattsson, L.; Riley, H.; Salo, T.; Thorvaldsson, G.; Christensen, B.T. An inventory of nordic long continued agricultural soil experiments. In Long-Term Field Experiments—A Unique Research Platform, Proceedings of NJF Seminar 407, Askov Experimental Station and Sandbjerg Estate, Denmark, 16–18 June 2008; pp. 76–79.
- Hyvonen, R.; Agren, G.I.; Andren, O. Modelling long-term: Carbon and nitrogen dynamics in an arable soil receiving organic matter. Ecol. Appl. 1996, 6, 1345–1354. [Google Scholar] [CrossRef]
- Powlson, D.S.; Glendining, M.J.; Coleman, K.; Whitmore, A.P. Implications for soil properties of removing cereal straw: Results from long-term studies. Agron. J. 2011, 103, 279–287. [Google Scholar] [CrossRef]
- Paul, E.A.; Follett, R.F.; Leavitt, S.W.; Halvorson, A.; Peterson, G.A.; Lyon, D.J. Radiocarbon dating for determination of soil organic matter pool sizes and dynamics. Soil Sci. Soc. Am. J. 1997, 61, 1058–1067. [Google Scholar] [CrossRef]
- Smith, P.; Smith, J.U.; Powlson, D.S.; McGill, W.B.; Arah, J.R.M.; Chertov, O.G.; Coleman, K.; Franko, U.; Frolking, S.; Jenkinson, D.S.; et al. A comparison of the performance of nine soil organic matter models using datasets from seven long-term experiments. Geoderma 1997, 81, 153–225. [Google Scholar] [CrossRef]
- Andren, O.; Katterer, T. Icbm: The introductory carbon balance model for exploration of soil carbon balances. Ecol. Appl. 1997, 7, 1226–1236. [Google Scholar] [CrossRef]
- Gerzabek, M.H. Soil organic matter research using long-term experiments. Kungl. Skogs Lantbr. Tidskr. 2007, 146, 22–29. [Google Scholar]
- Sun, B.; Hallett, P.D.; Caul, S.; Daniell, T.J.; Hopkins, D.W. Distribution of soil carbon and microbial biomass in arable soils under different tillage regimes. Plant Soil 2011, 338, 17–25. [Google Scholar] [CrossRef]
- Katterer, T.; Andren, O. Long-term agricultural field experiments in northern Europe: Analysis of the influence of management on soil carbon stocks using the ICBM model. Agric. Ecosyst. Environ. 1999, 75, 145–146. [Google Scholar] [CrossRef]
- Petersen, J.; Mattsson, L.; Riley, H.; Salo, T.; Thorvaldsson, G.; Christensen, B.T. Long continued agricultural soil experiments: A nordic research platform—An overview. In Proceedings of NJF Seminar 407, Askov Experimental Station and Sandbjerg Estate, Jutland, Denmark, 16–18 June 2008.
- Riley, H. Long-term fertilizer trials on loam soil at moystad, south-eastern Norway: Crop yields, nutrient balances and soil chemical analyses from 1983 to 2003. Acta Agric. Scand. Sect. BSoil Plant Sci. 2007, 57, 140–154. [Google Scholar]
- Mazzoncini, M.; di Bene, C.; Coli, A.; Risaliti, R.; Bonari, E. Long-term tillage and nitrogen fertilisation effects on maize yield and soil quality under rainfed mediterranean conditions: A critical perspective. DJF Rapp. Markbrug 2008, 7, 13–16. [Google Scholar]
- Saarela, I. Changes of yield responses and soil test values in finnish soils in relation to cumulative phosphorus and potassium balances. DJF Rapp. Markbrug 2008, 4, 52–55. [Google Scholar]
- Jaakkola, A.; Yli-Halla, M. Thirty years of growing cereal without P and K fertilization. DJF Rapp. Markbrug 2008, 4, 17–19. [Google Scholar]
- Magid, J.; Jensen, L.S. The crucial facility: A long-term field trial with urban fertilizers—Is recycling of nutrients from urban areas to peri-urban areas detrimental to the environment or the production system integrity? DJF Rapp. Markbrug 2008, 7, 11–12. [Google Scholar]
- Lessons Learned from Long-Term Soil Fertility Management Experiments in Africa; Bationo, A.; Waswa, B.; Kihara, J.; Adolwa, I.; Vanlauwe, B.; Saidou, K. (Eds.) Springer: Dordrecht, Netherlands, 2012; p. 204.
- Hawes, C.; George, T.; Iannetta, P.P.M.; James, E.K.; Karley, A.J.; McKenzie, B.M.; McNichol, J.W.; Squire, G.R.; Valentine, T.; Young, M. A whole-systems approach for designing sustainable cropping systems in northern Europe. Int. J. Agric. Sustain. 2014, in press. [Google Scholar]
- Griffith, B.A.H.; Hawkins, J.M.B.; Orr, R.J.; Blackwell, M.S.A.; Murray, P.J. The north wyke farm platform: Methodologies used in the remote sensing of the quantity and quality of drainage water. In Proceedings of the 22nd International Grasslands Congress, Sydney, Australia, 15–19 September 2013.
- Hatch, D.; Murray, P.; Orr, R.; Griffith, B. The North Wyke Farm Platform: A UK national capability for agri-environmental research. Asp. Appl. Biol. 2011, 115–118. [Google Scholar] [CrossRef]
- Chabbi, A.; Lemaire, G. Long term agro-ecosystem platforms for assessing biogeochemical cycles, environmental fluxes and biodiversity. In Proceedings of NJF Seminar 407. Askov Experimental Station and Sandbjerg Estate, Jutland, Denmark, 16–18 June 2008; pp. 73–75.
- Firbank, L.G.; Heard, M.S.; Woiwod, I.P.; Hawes, C.; Haughton, A.J.; Champion, G.T.; Scott, R.J.; Hill, M.O.; Dewar, A.M.; Squire, G.R.; et al. An introduction to the farm-scale evaluations of genetically modified herbicide-tolerant crops. J. Appl. Ecol. 2003, 40, 2–16. [Google Scholar] [CrossRef]
- Perry, J.N.; Rothery, P.; Clark, S.J.; Heard, M.S.; Hawes, C. Design, analysis and statistical power of the farm-scale evaluations of genetically modified herbicide-tolerant crops. J. Appl. Ecol. 2003, 40, 17–31. [Google Scholar] [CrossRef]
- Hawes, C.; Squire, G.R.; Hallett, P.D.; Watson, C.A.; Young, M. Arable plant communities as indicators of farming practice. Agric. Ecosyst. Environ. 2010, 138, 17–26. [Google Scholar] [CrossRef]
- Tamis, W.L.M.; van den Brink, W.J. Conventional, integrated and organic winter wheat production in the Netherlands in the period 1993–1997. Agric. Ecosyst. Environ. 1999, 76, 47–59. [Google Scholar] [CrossRef]
- Rutter, M.T.; Fenster, C.B. Testing for adaptation to climate in arabidopsis thaliana: A calibrated common garden approach. Ann. Bot. 2007, 99, 529–536. [Google Scholar] [CrossRef]
- Oleksyn, J.; Modrzynski, J.; Tjoelker, M.G.; Zytkowiak, R.; Reich, P.B.; Karolewski, P. Growth and physiology of picea abies populations from elevational transects: Common garden evidence for altitudinal ecotypes and cold adaptation. Funct. Ecol. 1998, 12, 573–590. [Google Scholar] [CrossRef]
- Tracy, S.R.; Roberts, J.A.; Black, C.R.; McNeill, A.; Davidson, R.; Mooney, S.J. The X-factor: Visualizing undisturbed root architecture in soils using X-ray computed tomography. J. Exp. Bot. 2010, 61, 311–313. [Google Scholar] [CrossRef]
- Clark, R.T.; MacCurdy, R.B.; Jung, J.K.; Shaff, J.E.; McCouch, S.R.; Aneshansley, D.J.; Kochian, L.V. Three-dimensional root phenotyping with a novel imaging and software platform. Plant Physiol. 2011, 156, 455–465. [Google Scholar] [CrossRef]
- De Dorlodot, S.; Bertin, P.; Baret, P.; Draye, X. Scaling up quantitative phenotyping of root system architecture using a combination of aeroponics and image analysis. In Proceedings of “Roots and the Soil Environment II, Nottingham, UK, April 2005; pp. 41–54.
- Downie, H.; Holden, N.; Otten, W.; Spiers, A.J.; Valentine, T.A.; Dupuy, L.X. Transparent soil for imaging the rhizosphere. PLoS One 2012, 7, e44276. [Google Scholar]
- Yang, W.; Duan, L.; Chen, G.; Xiong, L.; Liu, Q. Plant phenomics and high-throughput phenotyping: Accelerating rice functional genomics using multidisciplinary technologies. Curr. Opin. Plant Biol. 2013, 16, 180–187. [Google Scholar] [CrossRef]
- Karley, A.J.; Valentine, T.A.; Squire, G.R. Dwarf alleles differentially affect barley root traits influencing nitrogen acquisition under low nutrient supply. J. Exp. Bot. 2011, 62, 3917–3927. [Google Scholar] [CrossRef]
- Wishart, J.; George, T.S.; Brown, L.K.; Ramsay, G.; Bradshaw, J.E.; White, P.J.; Gregory, P.J. Measuring variation in potato roots in both field and glasshouse: The search for useful yield predictors and a simple screen for root traits. Plant Soil 2013, 368, 231–249. [Google Scholar] [CrossRef]
- Zadoks, J.C.; Chang, T.T.; Konzak, C.F. A decimal code for the growth stages of cereals. Weed Res. 1974, 14, 415–421. [Google Scholar] [CrossRef]
- Tajima, R.; Kato, Y. Comparison of threshold algorithms for automatic image processing of rice roots using freeware ImageJ. Field Crops Res. 2011, 121, 460–463. [Google Scholar] [CrossRef]
- Maloof, J.N.; Nozue, K.; Mumbach, M.R.; Palmer, C.M. LeafJ: An ImageJ plugin for semi-automated leaf shape measurement. J. Vis. Exp. 2013, 71. [Google Scholar] [CrossRef]
- Trachsel, S.; Kaeppler, S.M.; Brown, K.M.; Lynch, J.P. Shovelomics: High throughput phenotyping of maize (Zea mays L.) root architecture in the field. Plant Soil 2011, 341, 75–87. [Google Scholar] [CrossRef]
- Haling, R.E.; Simpson, R.J.; Culvenor, R.A.; Lambers, H.; Richardson, A.E. Field application of a DNA-based assay to the measurement of roots of perennial grasses. Plant Soil 2012, 358, 176–192. [Google Scholar]
- Haling, R.E.; Simpson, R.J.; McKay, A.C.; Hartley, D.; Lambers, H.; Ophel-Keller, K.; Wiebkin, S.; Herdina; Riley, I.T.; Richardson, A.E. Direct measurement of roots in soil for single and mixed species using a quantitative DNA-based method. Plant Soil 2011, 348, 123–137. [Google Scholar] [CrossRef]
- Huang, C.Y.; Kuchel, H.; Edwards, J.; Hall, S.; Parent, B.; Eckermann, P.; Herdina; Hartley, D.M.; Langridge, P.; McKay, A.C. A DNA-based method for studying root responses to drought in field-grown wheat genotypes. Sci. Rep. 2013, 3, 3194. [Google Scholar]
- Roessner, U.; Willmitzer, L.; Fernie, A.R. High-resolution metabolic phenotyping of genetically and environmentally diverse potato tuber systems. Identification of phenocopies. Plant Physiol. 2001, 127, 749–764. [Google Scholar] [CrossRef]
- Roessner, U.; Luedemann, A.; Brust, D.; Fiehn, O.; Linke, T.; Willmitzer, L.; Fernie, A.R. Metabolic profiling allows comprehensive phenotyping of genetically or environmentally modified plant systems. Plant Cell 2001, 13, 11–29. [Google Scholar] [CrossRef]
- Roessner, U.; Willmitzer, L.; Fernie, A.R. Metabolic profiling and biochemical phenotyping of plant systems. Plant Cell Rep. 2002, 21, 189–196. [Google Scholar] [CrossRef]
- Golzarian, M.R.; Frick, R.A. Classification of images of wheat, ryegrass and brome grass species at early growth stages using principal component analysis. Plant Methods 2011, 7, 28. [Google Scholar] [CrossRef]
- Al-Tam, F.M.; Adam, H.; Anjos, A.D.; Lorieux, M.; Larmande, P.; Ghesquiere, A.; Jouannic, S.; Shahbazkia, H.R. P-TRAP: A Panicle Trait Phenotyping tool. BMC Plant Biol. 2013, 13, 122. [Google Scholar] [CrossRef]
- Rousseau, C.; Belin, E.; Bove, E.; Rousseau, D.; Fabre, F.; Berruyer, R.; Guillaumes, J.; Manceau, C.; Jacques, M.-A.; Boureau, T. High throughput quantitative phenotyping of plant resistance using chlorophyll fluorescence image analysis. Plant Methods 2013, 9, 17. [Google Scholar] [CrossRef]
- Mooney, S.J.; Pridmore, T.P.; Helliwell, J.; Bennett, M.J. Developing X-ray computed tomography to non-invasively image 3-D root systems architecture in soil. Plant Soil 2012, 352, 1–22. [Google Scholar] [CrossRef]
- Keyes, S.D.; Daly, K.R.; Gostling, N.J.; Jones, D.L.; Talboys, P.; Pinzer, B.R.; Boardman, R.; Sinclair, I.; Marchant, A.; Roose, T. High resolution synchrotron imaging of wheat root hairs growing in soil and image based modelling of phosphate uptake. New Phytol. 2013, 198, 1023–1029. [Google Scholar] [CrossRef]
- Mairhofer, S.; Zappala, S.; Tracy, S.; Sturrock, C.; Bennett, M.J.; Mooney, S.J.; Pridmore, T.P. Recovering complete plant root system architectures from soil via X-ray μ-Computed Tomography. Plant Methods 2013, 9, 8. [Google Scholar] [CrossRef]
- Miguel Costa, J.; Grant, O.M.; Manuela Chaves, M. Thermography to explore plant-environment interactions. J. Exp. Bot. 2013, 64, 3937–3949. [Google Scholar] [CrossRef]
- Guo, W.; Rage, U.K.; Ninomiya, S. Illumination invariant segmentation of vegetation for time series wheat images based on decision tree model. Comput. Electron. Agric. 2013, 96, 58–66. [Google Scholar] [CrossRef]
- Murchie, E.H.; Lawson, T. Chlorophyll fluorescence analysis: A guide to good practice and understanding some new applications. J. Exp. Bot. 2013, 64, 3983–3998. [Google Scholar] [CrossRef]
- Prashar, A.; Yildiz, J.; McNicol, J.W.; Bryan, G.J.; Jones, H.G. Infra-red thermography for high throughput field phenotyping in Solanum tuberosum. PLoS One 2013, 8, e65816. [Google Scholar]
- Jones, H.G.; Serraj, R.; Loveys, B.R.; Xiong, L.; Wheaton, A.; Price, A.H. Thermal infrared imaging of crop canopies for the remote diagnosis and quantification of plant responses to water stress in the field. Funct. Plant Biol. 2009, 36, 978–989. [Google Scholar] [CrossRef]
- Hackl, H.; Baresel, J.P.; Mistele, B.; Hu, Y.; Schmidhalter, U. A comparison of plant temperatures as measured by thermal imaging and infrared thermometry. J. Agron. Crop Sci. 2012, 198, 415–429. [Google Scholar] [CrossRef]
- Gutierrez, M.; Reynolds, M.P.; Klatt, A.R. Association of water spectral indices with plant and soil water relations in contrasting wheat genotypes. J. Exp. Bot. 2010, 61, 3291–3303. [Google Scholar] [CrossRef]
- Munns, R.; James, R.A.; Sirault, X.R.R.; Furbank, R.T.; Jones, H.G. New phenotyping methods for screening wheat and barley for beneficial responses to water deficit. J. Exp. Bot. 2010, 61, 3499–3507. [Google Scholar] [CrossRef]
- Thoren, D.; Schmidhalter, U. Nitrogen status and biomass determination of oilseed rape by laser-induced chlorophyll fluorescence. Eur. J. Agron. 2009, 30, 238–242. [Google Scholar] [CrossRef]
- Thoren, D.; Thoren, P.; Schmidhalter, U. Influence of ambient light and temperature on laser-induced chlorophyll fluorescence measurements. Eur. J. Agron. 2010, 32, 169–176. [Google Scholar] [CrossRef]
- Dietrich, R.C.; Bengough, A.G.; Jones, H.G.; White, P.J. A new physical interpretation of plant root capacitance. J. Exp. Bot. 2012, 63, 6149–6159. [Google Scholar] [CrossRef]
- Chapuis, R.; Delluc, C.; Debeuf, R.; Tardieu, F.; Welcker, C. Resiliences to water deficit in a phenotyping platform and in the field: How related are they in maize? Eur. J. Agron. 2012, 42, 59–67. [Google Scholar] [CrossRef]
- Dietrich, R.C.; Bengough, A.G.; Jones, H.G.; White, P.J. Can root electrical capacitance be used to predict root mass in soil? Ann. Bot. 2013, 112, 457–464. [Google Scholar] [CrossRef]
- D’Andrea, K.E.; Otegui, M.E.; Cirilo, A.G.; Eyherabide, G.H. Ecophysiological traits in maize hybrids and their parental inbred lines: Phenotyping of responses to contrasting nitrogen supply levels. Field Crops Res. 2009, 114, 147–158. [Google Scholar] [CrossRef]
- Montes, J.M.; Technow, F.; Dhillon, B.S.; Mauch, F.; Melchinger, A.E. High-throughput non-destructive biomass determination during early plant development in maize under field conditions. Field Crops Res. 2011, 121, 268–273. [Google Scholar] [CrossRef]
- Busemeyer, L.; Mentrup, D.; Moeller, K.; Wunder, E.; Alheit, K.; Hahn, V.; Maurer, H.P.; Reif, J.C.; Wuerschum, T.; Mueller, J.; et al. Breedvision—A multi-sensor platform for non-destructive field-based phenotyping in plant breeding. Sensors 2013, 13, 2830–2847. [Google Scholar] [CrossRef]
- Granier, C.; Tardieu, F. Multi-scale phenotyping of leaf expansion in response to environmental changes: The whole is more than the sum of parts. Plant Cell Environ. 2009, 32, 1175–1184. [Google Scholar] [CrossRef]
- Rebetzke, G.J.; Chenu, K.; Biddulph, B.; Moeller, C.; Deery, D.M.; Rattey, A.R.; Bennett, D.; Barrett-Lennard, E.G.; Mayer, J.E. A multisite managed environment facility for targeted trait and germplasm phenotyping. Funct. Plant Biol. 2013, 40, 1–13. [Google Scholar] [CrossRef]
- Paproki, A.; Sirault, X.; Berry, S.; Furbank, R.; Fripp, J. A novel mesh processing based technique for 3D plant analysis. BMC Plant Biol. 2012, 12. [Google Scholar] [CrossRef]
- Vankadavath, R.N.; Hussain, A.J.; Bodanapu, R.; Kharshiing, E.; Basha, P.O.; Gupta, S.; Sreelakshmi, Y.; Sharma, R. Computer aided data acquisition tool for high-throughput phenotyping of plant populations. Plant Methods 2009, 5, 18. [Google Scholar] [CrossRef]
- Hallett, P.; Mooney, S.; Whalley, R. Soil physics: New approaches and emerging challenges introduction. Eur. J. Soil Sci. 2013, 64, 277–278. [Google Scholar] [CrossRef]
- Kirkegaard, J.A.; Lilley, J.M. Root penetration rate—A benchmark to identify soil and plant limitations to rooting depth in wheat. Aust. J. Exp. Agric. 2007, 47, 590–602. [Google Scholar] [CrossRef]
- George, T.S.; Brown, L.K.; Newton, A.C.; Hallett, P.D.; Sun, B.H.; Thomas, W.T.B.; White, P.J. Impact of soil tillage on the robustness of the genetic component of variation in phosphorus (P) use efficiency in barley (Hordeum vulgare L.). Plant Soil 2011, 339, 113–123. [Google Scholar] [CrossRef]
- Ball, B.C.; Douglas, J.T. A simple procedure for assessing soil structural, rooting and surface conditions. Soil Use Manag. 2003, 19, 50–56. [Google Scholar] [CrossRef]
- Moebius, B.N.; van Es, H.M.; Schindelbeck, R.R.; Idowu, O.J.; Clune, D.J.; Thies, J.E. Evaluation of laboratory-measured soil properties as indicators of soil physical quality. Soil Sci. 2007, 172, 895–912. [Google Scholar] [CrossRef]
- Silva, G.L.; Lima, H.V.; Campanha, M.M.; Gilkes, R.J.; Oliveira, T.S. Soil physical quality of luvisols under agroforestry, natural vegetation and conventional crop management systems in the Brazilian semi-arid region. Geoderma 2011, 167–168, 61–70. [Google Scholar] [CrossRef]
- Dexter, A.R. Soil physical quality—Part I. Theory, effects of soil texture, density, and organic matter, and effects on root growth. Geoderma 2004, 120, 201–214. [Google Scholar] [CrossRef]
- Czyz, E.A. Effects of traffic on soil aeration, bulk density and growth of spring barley. Soil Tillage Res. 2004, 79, 153–166. [Google Scholar] [CrossRef]
- Bengough, A.G.; McKenzie, B.M.; Hallett, P.D.; Valentine, T.A. Root elongation, water stress, and mechanical impedance: A review of limiting stresses and beneficial root tip traits. J. Exp. Bot. 2011, 62, 59–68. [Google Scholar] [CrossRef]
- DaSilva, A.P.; Kay, B.D. The sensitivity of shoot growth of corn to the least limiting water range of soils. Plant Soil 1996, 184, 323–329. [Google Scholar] [CrossRef]
- Ehlers, W.; Kopke, U.; Hesse, F.; Bohm, W. Penetration resistance and root-growth of oats in tilled and untilled loess soil. Soil Tillage Res 1983, 3, 261–275. [Google Scholar]
- White, R.G.; Kirkegaard, J.A. The distribution and abundance of wheat roots in a dense, structured subsoil—Implications for water uptake. Plant Cell Environ. 2010, 33, 133–148. [Google Scholar] [CrossRef]
- Reynolds, W.D.; Drury, C.F.; Tan, C.S.; Fox, C.A.; Yang, X.M. Use of indicators and pore volume-function characteristics to quantify soil physical quality. Geoderma 2009, 152, 252–263. [Google Scholar] [CrossRef]
- Mueller, L.; Kay, B.D.; Hu, C.; Li, Y.; Schindler, U.; Behrendt, A.; Shepherd, T.G.; Ball, B.C. Visual assessment of soil structure: Evaluation of methodologies on sites in Canada, China and Germany part I: Comparing visual methods and linking them with soil physical data and grain yield of cereals. Soil Tillage Res. 2009, 103, 178–187. [Google Scholar] [CrossRef]
- Ball, B.C.; Munkholm, L.J.; Batey, T. Applications of visual soil evaluation. Soil Tillage Res. 2013, 127, 1–2. [Google Scholar] [CrossRef]
- Aitkenhead, M.J.; Coull, M.C.; Donnelly, D.; Hastings, E.J. Innovations in environmental monitoring using mobile technology—A review. Int. J. Interact. Mob. Technol. 2014, 8, 50–58. [Google Scholar]
- Mairura, F.S.; Mugendi, D.N.; Mwanje, J.I.; Rarnisch, J.J.; Mbugua, P.K.; Chianu, J.N. Integrating scientific and fanners’ evaluation of soil quality indicators in central Kenya. Geoderma 2007, 139, 134–143. [Google Scholar] [CrossRef]
- Fernandez-Ugalde, O.; Virto, I.; Bescansa, P.; Imaz, M.J.; Enrique, A.; Karlen, D.L. No-tillage improvement of soil physical quality in calcareous, degradation-prone, semiarid soils. Soil Tillage Res. 2009, 106, 29–35. [Google Scholar] [CrossRef]
- Celeste Miretti, M.; Imhoff, S.; da Silva, A.P.; Lavado, R. Soil structure degradation in patches of alfalfa fields. Sci. Agric. 2010, 67, 604–610. [Google Scholar] [CrossRef]
- Connor, D.J. Organic agriculture cannot feed the world. Field Crops Res. 2008, 106, 187–190. [Google Scholar] [CrossRef]
- De Ponti, T.; Rijk, B.; van Ittersum, M.K. The crop yield gap between organic and conventional agriculture. Agric. Syst. 2012, 108, 1–9. [Google Scholar] [CrossRef]
- Ball, B.C.; Crawford, C.E. Mechanical weeding effects on soil structure under field carrots (Daucus carota L.) and beans (Vicia faba L.). Soil Use Manag. 2009, 25, 303–310. [Google Scholar] [CrossRef]
- Loeschenberger, F.; Fleck, A.; Grausgruber, H.; Hetzendorfer, H.; Hof, G.; Lafferty, J.; Marn, M.; Neumayer, A.; Pfaffinger, G.; Birschitzky, J. Breeding for organic agriculture: The example of winter wheat in Austria. Euphytica 2008, 163, 469–480. [Google Scholar] [CrossRef]
- Wolfe, M.S.; Baresel, J.P.; Desclaux, D.; Goldringer, I.; Hoad, S.; Kovacs, G.; Loeschenberger, F.; Miedaner, T.; Ostergard, H.; van Bueren, E.T.L. Developments in breeding cereals for organic agriculture. Euphytica 2008, 163, 323–346. [Google Scholar] [CrossRef]
- Casler, M.; Riday, H.; Undersander, D. Organic vs. Conventional fodder crops in the USA: A challenge for breeders? In Proceedings of the EUCARPIA Fodder Crops Meetings, Copenhagen, Denmark, 19–23 August 2007; pp. 17–23.
- Boller, B.; Tanner, P.; Schubiger, F.X. Breeding forage grasses for organic conditions. Euphytica 2008, 163, 459–467. [Google Scholar] [CrossRef]
- Houlbrooke, D.J.; Paton, R.J.; Littlejohn, R.P.; Morton, J.D. Land-use intensification in New Zealand: Effects on soil properties and pasture production. J. Agric. Sci. 2011, 149, 337–349. [Google Scholar] [CrossRef]
- Beutler, A.N.; Centurion, J.F.; da Silva, A.P.; Pessoa da Cruz Centurion, M.A.; Leonel, C.L.; da Freddi, O.S. Soil compaction by machine traffic and least limiting water range related to soybean yield. Pesqui. Agropecu. Bras. 2008, 43, 1591–1600. [Google Scholar] [CrossRef]
- Van Bueren, E.T.L.; Jones, S.S.; Tamm, L.; Murphy, K.M.; Myers, J.R.; Leifert, C.; Messmer, M.M. The need to breed crop varieties suitable for organic farming, using wheat, tomato and broccoli as examples: A review. NJAS—Wagening. J. Life Sci. 2011, 58, 193–205. [Google Scholar] [CrossRef]
- Wojciechowski, T.; Gooding, M.J.; Ramsay, L.; Gregory, P.J. The effects of dwarfing genes on seedling root growth of wheat. J. Exp. Bot. 2009, 60, 2565–2573. [Google Scholar] [CrossRef]
- Lueck, L.; Schmidt, C.S.; Cooper, J.M.; Hall, C.M.; Shotton, P.N.; Leifert, C. Effect of organic, low-input and conventional production systems on yield and quality of winter wheat. In Delivering Sustainability within Profitable Farming Systems—Is It Possible? Andrews, M., Turley, D., Cummings, S., Dale, M.F.B., Rowlinson, P., Eds.; Association of Applied Biologists: Warwickshire, UK, 2006; pp. 135–140. [Google Scholar]
- Dawson, J.C.; Huggins, D.R.; Jones, S.S. Characterizing nitrogen use efficiency in natural and agricultural ecosystems to improve the performance of cereal crops in low-input and organic agricultural systems. Field Crops Res. 2008, 107, 89–101. [Google Scholar] [CrossRef]
- Hoad, S.; Topp, C.; Davies, K. Selection of cereals for weed suppression in organic agriculture: A method based on cultivar sensitivity to weed growth. Euphytica 2008, 163, 355–366. [Google Scholar] [CrossRef]
- Cooper, J.M.; Schmidt, C.S.; Wilkinson, A.; Lueck, L.; Hall, C.M.; Shotton, P.N.; Leifert, C. Effect of organic, “low-input” and conventional production systems on disease incidence and severity in winter wheat. Asp. Appl. Biol. 2006, 80, 121–126. [Google Scholar]
- Makepeace, J.C.; Oxley, S.J.P.; Havis, N.D.; Hackett, R.; Burke, J.I.; Brown, J.K.M. Associations between fungal and abiotic leaf spotting and the presence of mlo alleles in barley. Plant Pathol. 2007, 56, 934–942. [Google Scholar] [CrossRef]
- Simon, M.R.; Worland, A.J.; Struik, P.C. Influence of plant height and heading date on the expression of the resistance to septoria tritici blotch in near isogenic lines of wheat. Crop Sci. 2004, 44, 2078–2085. [Google Scholar] [CrossRef]
- Klahr, A.; Zimmermann, G.; Wenzel, G.; Mohler, V. Effects of environment, disease progress, plant height and heading date on the detection of QTLs for resistance to fusarium head blight in an european winter wheat cross. Euphytica 2007, 154, 17–28. [Google Scholar] [CrossRef]
- Flintham, J.E.; Gale, M.D. The tom thumb dwarfing gene Rht3 in wheat. 2. Effects on height, yield and grain quality. Theor. Appl. Genet. 1983, 66, 249–256. [Google Scholar]
- Beutler, A.N.; Centurion, J.F.; Mengatto, L.H.; Mengatto, L.H.; Alves, J.B.; Cardoso Wagner, G.P. Impact of machine traffic on soil physical quality and corn yield in a kandiustalf. Acta Sci. Agron. 2009, 31, 359–364. [Google Scholar]
- Van Bueren, E.T.L.; Struik, P.C.; Jacobsen, E. Ecological concepts in organic farming and their consequences for an organic crop ideotype. Neth. J. Agric. Sci. 2002, 50, 1–26. [Google Scholar]
- Cassman, K.G. Ecological intensification of cereal production systems: Yield potential, soil quality, and precision agriculture. Proc. Natl. Acad. Sci. USA 1999, 96, 5952–5959. [Google Scholar] [CrossRef]
- Dobermann, A.; Cassman, K.G. Plant nutrient management for enhanced productivity in intensive grain production systems of the United States and Asia. Plant Soil 2002, 247, 153–175. [Google Scholar] [CrossRef]
- Rossel, R.A.V.; McBratney, A.B. Laboratory evaluation of a proximal sensing technique for simultaneous measurement of soil clay and water content. Geoderma 1998, 85, 19–39. [Google Scholar] [CrossRef]
- Van Alphen, B.J.; Stoorvogel, J.J. A functional approach to soil characterization in support of precision agriculture. Soil Sci. Soc. Am. J. 2000, 64, 1706–1713. [Google Scholar] [CrossRef]
- Horn, R.; Fleige, H. Risk assessment of subsoil compaction for arable soils in Northwest Germany at farm scale. Soil Tillage Res. 2009, 102, 201–208. [Google Scholar] [CrossRef]
- Alakukku, L.; Weisskopf, P.; Chamen, W.C.T.; Tijink, F.G.J.; van der Linden, J.P.; Pires, S.; Sommer, C.; Spoor, G. Prevention strategies for field traffic-induced subsoil compaction: A review part 1. Machine/soil interactions. Soil Tillage Res. 2003, 73, 145–160. [Google Scholar] [CrossRef]
- Holpp, M.; Kroulik, M.; Kviv, Z.; Anken, T.; Sauter, M.; Hensel, O. Large-scale field evaluation of driving performance and ergonomic effects of satellite-based guidance systems. Biosyst. Eng. 2013, 116, 190–197. [Google Scholar] [CrossRef]
- Kviv, Z.; Kroulik, M.; Chyba, J. Machinery guidance systems analysis concerning pass-to-pass accuracy as a toll for efficient plant production in fields and for soil damage reduction. Plant Soil Environ. 2014, 60, 36–42. [Google Scholar]
- Betteridge, K.; Schnug, E.; Haneklaus, S. Will site specific nutrient management live up to expectation? Landbauforsch. Volkenrode 2008, 58, 283–293. [Google Scholar]
- Haboudane, D.; Miller, J.R.; Pattery, E.; Zarco-Tejada, P.J.; Strachan, I.B. Hyperspectral vegetation indices and novel algorithms for predicting green LAI of crop canopies: Modeling and validation in the context of precision agriculture. Remote Sens. Environ. 2004, 90, 337–352. [Google Scholar] [CrossRef]
- Gasso, V.; Sorensen, C.A.G.; Oudshoorn, F.W.; Green, O. Controlled traffic farming: A review of the environmental impacts. Eur. J. Agron. 2013, 48, 66–73. [Google Scholar] [CrossRef]
- Newton, A.C.; Guy, D.C. Scale and spatial structure effects on the outcome of barley cultivar mixture trials for disease control. Field Crops Res. 2011, 123, 74–79. [Google Scholar] [CrossRef]
- Creissen, H.E.; Jorgensen, T.H.; Brown, J.K.M. Stabilization of yield in plant genotype mixtures through compensation rather than complementation. Ann. Bot. 2013, 112, 1439–1447. [Google Scholar] [CrossRef]
- Newton, A.C.; Guy, D.C.; Bengough, A.G.; Gordon, D.C.; McKenzie, B.M.; Sun, B.; Valentine, T.A.; Hallett, P.D. Soil tillage effects on the efficacy of cultivars and their mixtures in winter barley. Field Crops Res. 2012, 128, 91–100. [Google Scholar] [CrossRef]
- Locke, M.A.; Zablotowicz, R.M.; Steinriede, R.W.; Testa, S.; Reddy, K.N. Conservation management in cotton production: Long-term soil biological, chemical, and physical changes. Soil Sci. Soc. Am. J. 2013, 77, 974–984. [Google Scholar] [CrossRef]
- Vandenkoornhuyse, P.; Ridgway, K.P.; Watson, I.J.; Fitter, A.H.; Young, J.P.W. Co-existing grass species have distinctive arbuscular mycorrhizal communities. Mol. Ecol. 2003, 12, 3085–3095. [Google Scholar] [CrossRef]
- Skiba, M.W.; George, T.S.; Baggs, E.M.; Daniell, T.J. Plant influence on nitrification. Biochem. Soc. Trans. 2011, 39, 275–278. [Google Scholar] [CrossRef]
- Brooker, R.W.; Bennett, A.E.; Cong, W.-F.; Daniel, T.J.; George, T.S.; Hawes, C.; Iannetta, P.P.M.; Jones, H.G.; Karley, A.J.; Li, H.; et al. Improving intercropping: A synthesis of research in agronomy, plant physiology and ecology. New Phytol. 2014, in press. [Google Scholar]
- Trichard, A.; Alignier, A.; Chauvel, B.; Petit, S. Identification of weed community traits response to conservation agriculture. Agric. Ecosyst. Environ. 2013, 179, 179–186. [Google Scholar] [CrossRef]
- Diacono, M.; Montemurro, F. Long-term effects of organic amendments on soil fertility. A review. Agron. Sustain. Dev. 2010, 30, 401–422. [Google Scholar] [CrossRef]
- Amato, G.; Ruisi, P.; Frenda, A.S.; di Miceli, G.; Saia, S.; Plaia, A.; Giambalvo, D. Long-term tillage and crop sequence effects on wheat grain yield and quality. Agron. J. 2013, 105, 1317–1327. [Google Scholar] [CrossRef]
- Arvidsson, J.; Etana, A.; Rydberg, T. Crop yield in swedish experiments with shallow tillage and no-tillage 1983–2012. Eur. J. Agron. 2014, 52Pt B, 307–315. [Google Scholar] [CrossRef]
- Murphy, K.; Jones, S.S. Genetic assessment of the role of breeding wheat for organic systems. 2007, 12, 217–222. [Google Scholar]
- Reid, T.A.; Yang, R.-C.; Salmon, D.F.; Spaner, D. Should spring wheat breeding for organically managed systems be conducted on organically managed land? Euphytica 2009, 169, 239–252. [Google Scholar] [CrossRef]
- Wu, L.; McGechan, M.B.; McRoberts, N.; Baddeley, J.A.; Watson, C.A. Spacsys: Integration of a 3D root architecture component to carbon, nitrogen and water cycling-model description. Ecol. Model. 2007, 200, 343–359. [Google Scholar] [CrossRef]
- Fang, S.; Yan, X.; Liao, H. 3D reconstruction and dynamic modeling of root architecture in situ and its application to crop phosphorus research. Plant J. 2009, 60, 1096–1108. [Google Scholar] [CrossRef]
- El-Sharkawy, M.A. Overview: Early history of crop growth and photosynthesis modeling. Biosystems 2011, 103, 205–211. [Google Scholar] [CrossRef]
- Yin, X.; Struik, P.C. Modelling the crop: From system dynamics to systems biology. J. Exp. Bot. 2010, 61, 2171–2183. [Google Scholar] [CrossRef]
- Brisson, N.; Gary, C.; Justes, E.; Roche, R.; Mary, B.; Ripoche, D.; Zimmer, D.; Sierra, J.; Bertuzzi, P.; Burger, P.; et al. An overview of the crop model stics. Eur. J. Agron. 2003, 18, 309–332. [Google Scholar] [CrossRef]
- Hammer, G.L.; van Oosterom, E.; McLean, G.; Chapman, S.C.; Broad, I.; Harland, P.; Muchow, R.C. Adapting apsim to model the physiology and genetics of complex adaptive traits in field crops. J. Exp. Bot. 2010, 61, 2185–2202. [Google Scholar] [CrossRef]
- Jones, J.W.; Hoogenboom, G.; Porter, C.H.; Boote, K.J.; Batchelor, W.D.; Hunt, L.A.; Wilkens, P.W.; Singh, U.; Gijsman, A.J.; Ritchie, J.T. The dssat cropping system model. Eur. J. Agron. 2003, 18, 235–265. [Google Scholar] [CrossRef]
- Keating, B.A.; Carberry, P.S.; Hammer, G.L.; Probert, M.E.; Robertson, M.J.; Holzworth, D.; Huth, N.I.; Hargreaves, J.N.G.; Meinke, H.; Hochman, Z.; et al. An overview of apsim, a model designed for farming systems simulation. Eur. J. Agron. 2003, 18, 267–288. [Google Scholar] [CrossRef]
- Stockle, C.O.; Donatelli, M.; Nelson, R. Cropsyst, a cropping systems simulation model. Eur. J. Agron. 2003, 18, 289–307. [Google Scholar] [CrossRef]
- Dunbabin, V.M.; Diggle, A.J.; Rengel, Z.; van Hugten, R. Modelling the interactions between water and nutrient uptake and root growth. Plant Soil 2002, 239, 19–38. [Google Scholar] [CrossRef]
- Chen, W.; Dunbabin, V.; Bell, R.; Brennan, R.; Bowden, B. Simulating and understanding root growth using rootmap to guide phosphorus fertiliser placement in wide row lupin cropping systems. In Proceedings of the 12th International Lupin Conference, Fremantle, Western Australia, 14–18 September 2008; pp. 368–372.
- Dunbabin, V.M.; Diggle, A.J.; Rengel, Z. Simulation of field data by a basic three-dimensional model of interactive root growth. Plant Soil 2002, 239, 39–54. [Google Scholar] [CrossRef]
- McCown, R.L.; Hammer, G.L.; Hargreaves, J.N.G.; Holzworth, D.; Huth, N.I. Apsim—An agricultural production system simulation-model for operational-research. Math. Comput. Simul. 1995, 39, 225–231. [Google Scholar] [CrossRef]
- McCown, R.L.; Hammer, G.L.; Hargreaves, J.N.G.; Holzworth, D.P.; Freebairn, D.M. Apsim: A novel software system for model development, model testing and simulation in agricultural systems research. Agric. Syst. 1996, 50, 255–271. [Google Scholar] [CrossRef]
- Boote, K.J.; Jones, J.W.; White, J.W.; Asseng, S.; Lizaso, J.I. Putting mechanisms into crop production models. Plant Cell Environ. 2013, 36, 1658–1672. [Google Scholar] [CrossRef]
- Dunbabin, V.M.; McDermott, S.; Bengough, A.G. Upscaling from rhizosphere to whole root system: Modelling the effects of phospholipid surfactants on water and nutrient uptake. Plant Soil 2006, 283, 57–72. [Google Scholar] [CrossRef]
- Dunbabin, V.M.; Airey, M.; Diggle, A.J.; Renton, M.; Rengel, Z.; Armstrong, R.; Chen, Y.; Siddique, K.H.M. Simulating the interaction between plant roots, soil water and nutrient flows, and barriers and objects in soil using rootmap. In Proceedings of the 19th International Congress on Modelling and Simulation (Modsim 2011), Perth, Western Australia, Australia, 12–16 December 2011; pp. 975–981.
- Chen, Y.L.; Dunbabin, V.M.; Postma, J.A.; Diggle, A.J.; Siddique, K.H.M.; Rengel, Z. Modelling root plasticity and response of narrow-leafed lupin to heterogeneous phosphorus supply. Plant Soil 2013, 372, 319–337. [Google Scholar] [CrossRef]
- Stockle, C.O.; Martin, S.A.; Campbell, G.S. Cropsyst, a cropping systems simulations-model: Water/Nitrogen budgets and crop yield. Agric. Syst. 1994, 46, 335–359. [Google Scholar] [CrossRef]
- Anwar, M.R.; Liu, D.L.; Macadam, I.; Kelly, G. Adapting agriculture to climate change: A review. Theor. Appl. Climatol. 2013, 113, 225–245. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
George, T.S.; Hawes, C.; Newton, A.C.; McKenzie, B.M.; Hallett, P.D.; Valentine, T.A. Field Phenotyping and Long-Term Platforms to Characterise How Crop Genotypes Interact with Soil Processes and the Environment. Agronomy 2014, 4, 242-278. https://doi.org/10.3390/agronomy4020242
George TS, Hawes C, Newton AC, McKenzie BM, Hallett PD, Valentine TA. Field Phenotyping and Long-Term Platforms to Characterise How Crop Genotypes Interact with Soil Processes and the Environment. Agronomy. 2014; 4(2):242-278. https://doi.org/10.3390/agronomy4020242
Chicago/Turabian StyleGeorge, Timothy S., Cathy Hawes, Adrian C. Newton, Blair M. McKenzie, Paul D. Hallett, and Tracy A. Valentine. 2014. "Field Phenotyping and Long-Term Platforms to Characterise How Crop Genotypes Interact with Soil Processes and the Environment" Agronomy 4, no. 2: 242-278. https://doi.org/10.3390/agronomy4020242
APA StyleGeorge, T. S., Hawes, C., Newton, A. C., McKenzie, B. M., Hallett, P. D., & Valentine, T. A. (2014). Field Phenotyping and Long-Term Platforms to Characterise How Crop Genotypes Interact with Soil Processes and the Environment. Agronomy, 4(2), 242-278. https://doi.org/10.3390/agronomy4020242






