Diversity and Community Structure of Rhizosphere Arbuscular Mycorrhizal Fungi in Songnen Grassland Saline–Alkali-Tolerant Plants: Roles of Environmental Salinity and Plant Species Identity
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Site and Sample Collection
2.2. Laboratory Analyses
2.3. Statistical Analysis
3. Results
3.1. AMF Species in the Rhizosphere of Eight Salt-Tolerant Plants
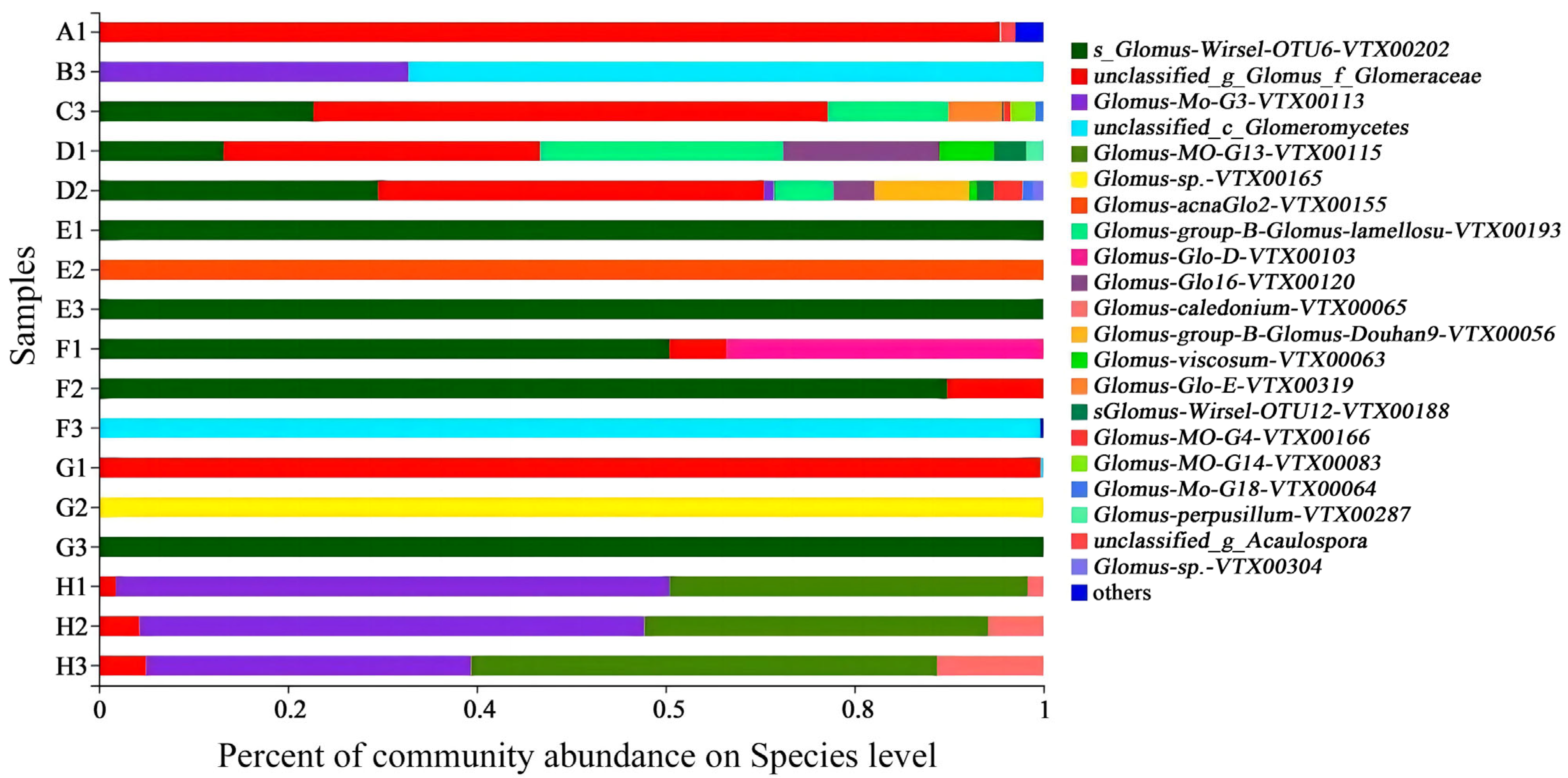
3.2. Distribution of Arbuscular Mycorrhizal Fungi in the Rhizosphere of Eight Salt-Tolerant and Alkali-Resistant Plant Species
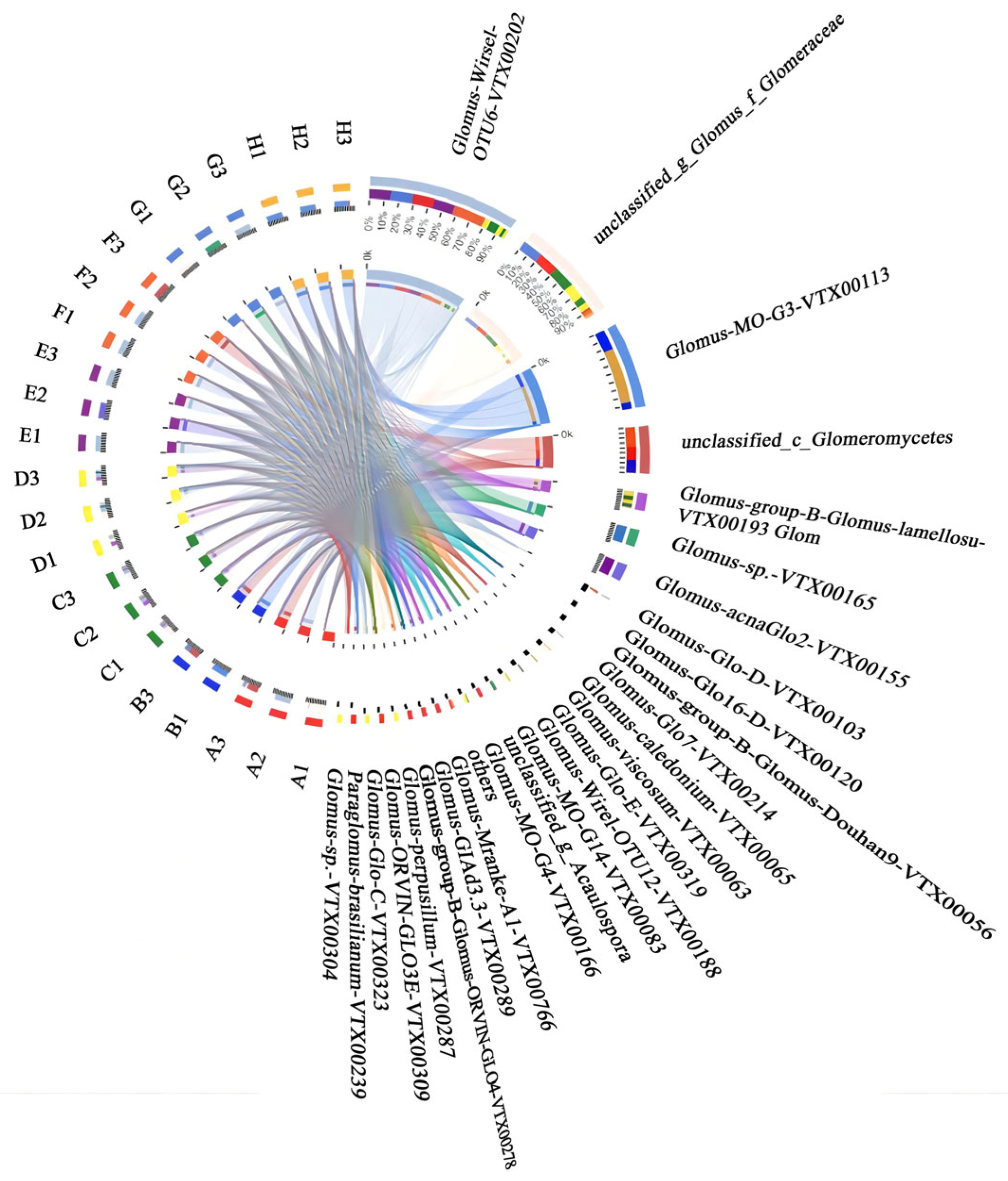
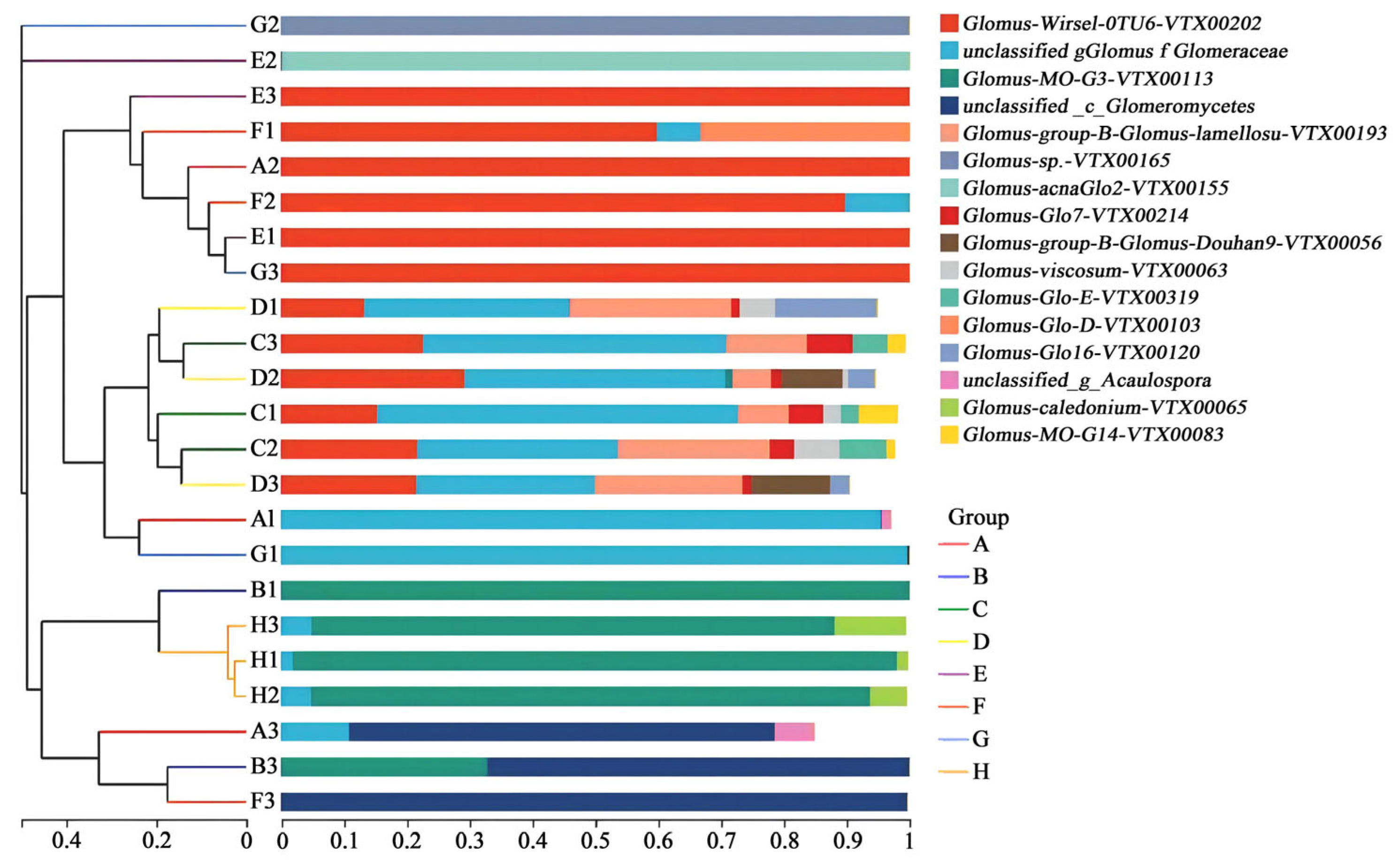
3.3. Relationship Between AMF and Plant Species
3.4. Analysis of Rhizosphere pH, EC, and AMF Diversity in Eight Different Plant Species
4. Discussion
4.1. Species and Distribution of Arbuscular Mycorrhizal Fungi (AMF) in the Rhizosphere of Eight Salt-Tolerant Plants in the Songnen Grassland
4.2. Adaptive Strategies of AMF–Plant Interactions in Rhizospheres of Different Plants in Heterogeneous Habitats
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Luo, S.; Yuan, J.; Song, Y.; Ren, J.; Qi, J.; Zhu, M.; Feng, Y.; Li, M.; Wang, B.; Li, X.; et al. Elevated salinity decreases microbial communities complexity and carbon, nitrogen and phosphorus metabolism in the Songnen Plain wetlands of China. Water Res. 2025, 276, 123285. [Google Scholar] [CrossRef]
- Chang, C.; Tian, L.; Tian, Z.; McLaughlin, N.; Tian, C. Change of soil microorganism communities under saline-sodic land degradation on the Songnen Plain in northeast China. J. Plant Nutr. Soil Sci. 2022, 185, 297–307. [Google Scholar] [CrossRef]
- Tarolli, P.; Luo, J.; Park, E.; Barcaccia, G.; Masin, R. Soil salinization in agriculture: Mitigation and adaptation strategies combining nature-based solutions and bioengineering. iScience 2024, 27, 108830. [Google Scholar] [CrossRef]
- Yin, X.; Ma, C.; He, H.; Wang, Z.; Li, X.; Fu, G.; Liu, J.; Zheng, Y. Distribution and diversity patterns of soil fauna in different salinization habitats of Songnen Grasslands, China. Appl. Soil Ecol. 2018, 123, 375–383. [Google Scholar] [CrossRef]
- Wang, L.; Seki, K.; Miyazaki, T.; Ishihama, Y. The causes of soil alkalinization in the Songnen Plain of Northeast China. Paddy Water Environ. 2009, 7, 259–270. [Google Scholar] [CrossRef]
- Tan, Z.X.; Lal, R.; Wiebe, K.D. Global soil nutrient depletion and yield reduction. J. Sustain. Agric. 2005, 26, 123–146. [Google Scholar] [CrossRef]
- Arif, Y.; Singh, P.; Siddiqui, H.; Bajguz, A.; Hayat, S. Salinity induced physiological and biochemical changes in plants: An omic approach towards salt stress tolerance. Plant Physiol. Biochem. 2020, 156, 64–77. [Google Scholar] [CrossRef]
- Dere, S. Mitigating the adverse effects of salt stress on pepper plants through arbuscular mycorrhizal fungi (AMF) and beneficial bacterial (PGPR) inoculation. Horticulturae 2024, 10, 1150. [Google Scholar] [CrossRef]
- Yadav, R.S.; Mahatma, M.K.; Thirumalaisamy, P.P.; Meena, H.N.; Bhaduri, D.; Arora, S.; Panwar, J. Arbuscular Mycorrhizal Fungi (AMF) for Sustainable Soil and Plant Health in Salt-Affected Soils. In Bioremediation of Salt Affected Soils: An Indian Perspective; Springer: Cham, Switzerland, 2017; pp. 133–156. [Google Scholar]
- Zhang, Z.; Zhou, Z.; Feng, S.; Guo, P.; Wang, Y.; Hao, B.; Guo, W.; Li, F.Y. Synergistic effects of AMF and PGPR on improving saline-alkaline tolerance of Leymus chinensis by strengthening the link between rhizosphere metabolites and microbiomes. Environ. Technol. Innov. 2024, 36, 103900. [Google Scholar] [CrossRef]
- Klichowska, E.; Nobis, M.; Piszcek, P.; Błaszkowski, J.; Zubek, S. Soil properties rather than topography, climatic conditions, and vegetation type shape AMF–feathergrass relationship in semi-natural European grasslands. Appl. Soil Ecol. 2019, 144, 22–30. [Google Scholar] [CrossRef]
- Wen, Y.; Wu, R.; Qi, D.; Xu, T.; Chang, W.; Li, K.; Fang, X.; Song, F. The effect of AMF combined with biochar on plant growth and soil quality under saline-alkali stress: Insights from microbial community analysis. Ecotoxicol. Environ. Saf. 2024, 281, 116592. [Google Scholar] [CrossRef] [PubMed]
- Ren, L.; Zhang, N.; Wu, P.; Huo, H.; Xu, G.; Wu, G. Arbuscular mycorrhizal colonization alleviates Fusarium wilt in watermelon and modulates the composition of root exudates. Plant Growth Regul. 2015, 77, 77–85. [Google Scholar] [CrossRef]
- Scheublin, T.R.; Van Der Heijden, M.G.A. Arbuscular mycorrhizal fungi colonize nonfixing root nodules of several legume species. New Phytol. 2006, 172, 732–738. [Google Scholar] [CrossRef]
- Shukla, S.; Didwania, N.; Choudhary, R. Arbuscular mycorrhizal fungi (AMF): A pathway to sustainable soil health, carbon sequestration, and greenhouse gas mitigation. J. Saudi Soc. Agric. Sci. 2025, 24, 22. [Google Scholar] [CrossRef]
- Hnini, M.; Rabeh, K.; Oubohssaine, M. Interactions between beneficial soil microorganisms (PGPR and AMF) and host plants for environmental restoration: A systematic review. Plant Stress 2024, 11, 100391. [Google Scholar] [CrossRef]
- Dastogeer, K.M.G.; Zahan, M.I.; Tahjib-Ul-Arif, M.; Akter, M.A.; Okazaki, S. Plant salinity tolerance conferred by arbuscular mycorrhizal fungi and associated mechanisms: A meta-analysis. Front. Plant Sci. 2020, 11, 588550. [Google Scholar] [CrossRef]
- Liu, H.; Tang, H.; Ni, X.; Zhang, Y.; Wang, Y. Interactive Effects of Epichloë Endophytes and Arbuscular Mycorrhizal Fungi on Saline-Alkali Stress Tolerance in Tall Fescue. Front. Microbiol. 2022, 13, 855890. [Google Scholar] [CrossRef]
- Gostinčar, C.; Zalar, P.; Gunde-Cimerman, N. No need for speed: Slow development of fungi in extreme environments. Fungal Biol. Rev. 2022, 39, 1–14. [Google Scholar] [CrossRef]
- Zhang, W.; Huang, C.; Wu, Y.; Rahman, M.A.; Xu, J.; Xiao, Y. Additive and antagonistic interactions between arbuscular mycorrhizal fungi and endophytic fungi dominate effect on plant performance and colonization rate. Plant Soil 2024, 508, 401–416. [Google Scholar] [CrossRef]
- Krishnamoorthy, R.; Kim, K.; Kim, C.; Sa, T. Changes of arbuscular mycorrhizal traits and community structure with respect to soil salinity in a coastal reclamation land. Soil Biol. Biochem. 2014, 72, 1–10. [Google Scholar] [CrossRef]
- Verma, O.; Sharma, S.; Kumar, V.; Singh, T.; Kumar, R.; Auji, R. Salinity stress effect on staple food crops and novel mitigation strategies. Biologia 2024, 79, 2359–2374. [Google Scholar] [CrossRef]
- Yang, G.; Liu, N.; Lu, W.; Wang, S.; Kan, H.; Zhang, Y. The interaction between arbuscular mycorrhizal fungi and soil phosphorus availability influences plant community productivity and ecosystem stability. J. Ecol. 2014, 102, 1072–1082. [Google Scholar] [CrossRef]
- Wu, C.; Bi, Y.; Zhu, W.; Yang, W.; Xue, C.; Li, H.; Liu, T. Arbuscular mycorrhizal inoculation alleviates water competition between adjacent plants in the mixed planting system. Rhizosphere 2023, 27, 100751. [Google Scholar] [CrossRef]
- Fu, H.; Yang, Y. How plants tolerate salt stress. Curr. Issues Mol. Biol. 2023, 45, 5914–5934. [Google Scholar] [CrossRef] [PubMed]
- Ling, L.; An, Y.; Wang, D.; Tang, L.; Du, B.; Shu, Y.; Bai, Y.; Guo, C. Proteomic analysis reveals responsive mechanisms for saline-alkali stress in alfalfa. Plant Physiol. Biochem. 2022, 170, 146–159. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Qu, H.; Liu, X.; Zhang, Y.; Chao, L.; Liu, H.; Bao, Y. Changes of root AMF community structure and colonization levels under distribution pattern of geographical substitute for four Stipa species in arid steppe. Microbiol. Res. 2023, 271, 127371. [Google Scholar] [CrossRef]
- Camuy-Velez, L.; Chakraborty, D.; Young, A.; Paudel, S.; Elvers, R.; Vanderhyde, M.; Walter, K.; Herzog, C.; Banerjee, S. Context-dependent contributions of arbuscular mycorrhizal fungi to host performance under global change factors. Soil Biol. Biochem. 2025, 204, 109707. [Google Scholar] [CrossRef]
- Wahab, A.; Batool, F.; Muhammad, M.; Zaman, W.; Mikhlef, R.M.; Qaddoori, S.M.; Ullah, S.; Abdi, G.; Saqib, S. Unveiling the complex molecular dynamics of arbuscular mycorrhizae: A comprehensive exploration and future perspectives in harnessing phosphate-solubilizing microorganisms for sustainable progress. Environ. Exp. Bot. 2024, 219, 105633. [Google Scholar] [CrossRef]
- Wang, F.; Rengel, Z. Disentangling the contributions of arbuscular mycorrhizal fungi to soil multifunctionality. Pedosphere 2024, 34, 269–278. [Google Scholar] [CrossRef]
- Fan, L.; Zhang, C.; Li, J.; Liu, Y. The use of Arbuscular mycorrhizal fungi to alleviate the growth and photosynthetic characteristics of strawberry under salt stress. Acta Physiol. Plant. 2024, 46, 118. [Google Scholar] [CrossRef]
- Liu, B.; Hu, Y.; Wang, Y.; Xue, H.; Li, Z.; Li, M. Effects of saline-alkali stress on bacterial and fungal community diversity in Leymus chinensis rhizosphere soil. Environ. Sci. Pollut. Res. 2022, 29, 70000–70013. [Google Scholar] [CrossRef]
- Hnini, M.; Homadi, C.; Aurag, J.; Chamkhi, I. Morpho-diversity and abundance of arbuscular mycorrhizal fungi (AMF) colonized the rhizospheric soils of two endemic plants localized in sud of Morocco. Microbe 2024, 4, 100133. [Google Scholar] [CrossRef]
- Mei, L.; Zhang, P.; Cui, G.; Yang, X.; Zhang, T.; Guo, J. Arbuscular mycorrhizal fungi promote litter decomposition and alleviate nutrient limitations of soil microbes under warming and nitrogen application. Appl. Soil Ecol. 2022, 171, 104318. [Google Scholar] [CrossRef]
- Jefwa, J.M.; Okoth, S.; Wachira, P.; Karanja, N.; Kahindi, J.; Njuguini, S.; Ichami, S.; Mung’atu, J.; Okoth, P.; Huising, J. Impact of land use types and farming practices on occurrence of arbuscular mycorrhizal fungi (AMF) in Taita-Taveta district in Kenya. Agric. Ecosyst. Environ. 2012, 157, 32–39. [Google Scholar] [CrossRef]
- Mei, L.; Yang, X.; Zhang, S.; Zhang, T.; Guo, J. Arbuscular mycorrhizal fungi alleviate phosphorus limitation by reducing plant N:P ratios under warming and nitrogen addition in a temperate meadow ecosystem. Sci. Total Environ. 2019, 686, 1129–1139. [Google Scholar] [CrossRef]
- Mei, L.; Zhang, N.; Wei, Q.; Cao, Y.; Li, D.; Cui, G. Alfalfa modified the effects of degraded black soil cultivated land on the soil microbial community. Front. Plant Sci. 2022, 13, 938187. [Google Scholar] [CrossRef]
- Davison, J.; Moora, M.; Öpik, M.; Adholeya, A.; Ainsaar, L.; Bâ, A.; Burla, S.; Diedhiou, A.G.; Hiiesalu, I.; Jairus, T.; et al. Global assessment of arbuscular mycorrhizal fungus diversity reveals very low endemism. Science 2015, 349, 970–973. [Google Scholar] [CrossRef]
- Fang, L.-L.; Liu, Y.-J.; Wang, Z.-H.; Lu, X.-Y.; Li, J.-H.; Yang, C.-X. Electrical conductivity and pH are two of the main factors influencing the composition of arbuscular mycorrhizal fungal communities in the vegetation succession series of Songnen saline-alkali grassland. J. Fungi 2023, 9, 870. [Google Scholar] [CrossRef] [PubMed]
- Bi, Y.; Ma, W.; Xing, F.; Gao, Y.; Li, Z.; Chen, C.; Mu, X.; Li, X.; Zhu, X. Diversity and specificity of arbuscular mycorrhizal fungi in the rhizosphere of six plants in the Songnen grassland, China. Écoscience 2019, 27, 11–21. [Google Scholar] [CrossRef]
- Koo, C.D.; Kim, T.-H.; Yi, C.-K.; Lee, W.-K.; Kang, C.-H.; Lee, B.-C.; Lee, S.K. Sporocarp-forming arbuscular mycorrhizal fungi, Glomus spp. in forest soils of korea. Korean J. Mycol. 1992, 20, 29–36. [Google Scholar]
- Bharti, N.; Baghel, S.; Barnawal, D.; Yadav, A.; Kalra, A. The greater effectiveness of Glomus mosseae and Glomus intraradices in improving productivity, oil content and tolerance of salt-stressed menthol mint (Mentha arvensis). J. Sci. Food Agric. 2013, 93, 2154–2161. [Google Scholar] [CrossRef]
- Yang, C.; Chen, F.; Li, L.; Li, C.; Yu, D.; Lin, J. Isolation and identification of Glomus intraradices in salt-alkali grassland and its effects on salt tolerance of Trifolium repens. Fresenius Environ. Bull. 2016, 25, 4439–4446. [Google Scholar]
- Borjian, M.; Jafarinia, M.; Ghotbi-Ravandi, A.A. Biofertilizer-mediated salt tolerance in sunflower: Synergistic effecs of Glomus mosseae and Pseudomonas fluorescens. Plant Soil 2024, 511, 713–732. [Google Scholar] [CrossRef]
- Mohammad, A.; Mittra, B. Effects of inoculation with stress-adapted arbuscular mycorrhizal fungus Glomus deserticola on growth of Solanum melogena L. and Sorghum sudanese Staph. seedlings under salinity and heavy metal stress conditions. Arch. Agron. Soil Sci. 2013, 59, 173–183. [Google Scholar] [CrossRef]
- Baldrian, P.; Větrovský, T.; Lepinay, C.; Kohout, P. High-throughput sequencing view on the magnitude of global fungal diversity. Fungal Divers. 2021, 114, 539–547. [Google Scholar] [CrossRef]
- Zehra Khan, U.; Kauser Perveen, U. Evaluation of arbuscular mycorrhizal fungi in cultivated crops. Int. J. Sci. Res. Arch. 2023, 10, 497–501. [Google Scholar] [CrossRef]
- He, F.; Chen, H.; Tang, M. Arbuscular mycorrhizal fungal communities are influenced by host tree species on the Loess Plateau, Northwest China. Forests 2019, 10, 930. [Google Scholar] [CrossRef]
- Bahadur, A.; Batool, A.; Nasir, F.; Jiang, S.; Mingsen, Q.; Zhang, Q.; Pan, J.; Liu, Y.; Feng, H. Mechanistic insights into arbuscular mycorrhizal fungi-mediated drought stress tolerance in plants. Int. J. Mol. Sci. 2019, 20, 4199. [Google Scholar] [CrossRef]
- Chen, J.; Zhou, J.; Smith, P.; Pan, G. Soil organic carbon storage from biochar addition falls over time: A perspective of shifts in soil extracellular enzyme activity. Soil Sci. Environ. 2025, 4, e002. [Google Scholar] [CrossRef]
- Firdu, Z.; Dida, G. Extraction, identification and mass production of arbuscular mycorrhizal fungi (AMF) from faba bean (Vicia faba L.) rhizosphere soils using maize (Zea mays L.) as a host plant. Heliyon 2024, 10, e36838. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Liu, Y.; Zhao, W.; Chen, F.E.I.; Wang, Y.; Yang, C. Isolation and species diversity of arbuscular mycorrhizal fungi in the rhizosphere of Puccinellia tenuiflora of Songnen saline-alkaline grassland, China. Biocell 2022, 46, 2465–2474. [Google Scholar] [CrossRef]
- Wang, J.; Ma, S.; Wang, G.G.; Xu, L.; Fu, Z.; Song, J.; Zhang, J. Arbuscular mycorrhizal fungi communities associated with wild plants in a coastal ecosystem. J. For. Res. 2020, 32, 683–695. [Google Scholar] [CrossRef]
- Schwarzott, D.; Walker, C.; Schüßler, A. Glomus, the largest genus of the arbuscular mycorrhizal fungi (Glomales), is nonmonophyletic. Mol. Phylogenetics Evol. 2001, 21, 190–197. [Google Scholar] [CrossRef]
- Lee, E.-H.; Eo, J.-K.; Ka, K.-H.; Eom, A.-H. Diversity of arbuscular mycorrhizal fungi and their roles in ecosystems. Mycobiology 2013, 41, 121–125. [Google Scholar] [CrossRef]
- Jiang, Y.; Luan, L.; Hu, K.; Liu, M.; Chen, Z.; Geisen, S.; Chen, X.; Li, H.; Xu, Q.; Bonkowski, M.; et al. Trophic interactions as determinants of the arbuscular mycorrhizal fungal community with cascading plant-promoting consequences. Microbiome 2020, 8, 142. [Google Scholar] [CrossRef]
- Malik, J.A.; Dar, B.A.; Alqarawi, A.A.; Assaeed, A.M.; Alotaibi, F.; Alkhasha, A.; Adam, A.M.; Abd-ElGawad, A.M. Species richness of arbuscular mycorrhizal fungi in heterogenous saline environments. Diversity 2025, 17, 183. [Google Scholar] [CrossRef]
- Cao, Y.; Wu, X.; Zhukova, A.; Tang, Z.; Weng, Y.; Li, Z.; Yang, Y. Arbuscular mycorrhizal fungi (AMF) species and abundance exhibit different effects on saline-alkaline tolerance in Leymus chinensis. J. Plant Interact. 2020, 15, 266–279. [Google Scholar] [CrossRef]
- Kumari, C.; Bhardwaj, A.K.; Kumar, R.; Chandra, K.K. Exploring the mutualistic relationship between plants and arbuscular mycorrhizal fungi. J. Adv. Biol. Biotechnol. 2025, 28, 787–800. [Google Scholar] [CrossRef]
- Ephraim Motaroki Menge, U. Investigating the ecological role of arbuscular mycorrhizal fungi (AMF) in natural ecosystems. Int. J. Sci. Res. Arch. 2023, 10, 524–534. [Google Scholar] [CrossRef]
- Ephraim Motaroki Menge, U. Unraveling the genetic diversity and evolution of Arbuscular Mycorrhizal Fungi (AMF). Int. J. Sci. Res. Arch. 2023, 10, 513–523. [Google Scholar] [CrossRef]
- Li, S.; Chi, S.; Lin, C.; Cai, C.; Yang, L.; Peng, K.; Huang, X.; Liu, J. Combination of biochar and AMF promotes phosphorus utilization by stimulating rhizosphere microbial co-occurrence networks and lipid metabolites of Phragmites. Sci. Total Environ. 2022, 845, 157339. [Google Scholar] [CrossRef]
- Yang, Y.; Luo, W.; Xu, J.; Guan, P.; Chang, L.; Wu, X.; Wu, D. Fallow land enhances carbon sequestration in glomalin and soil aggregates through regulating diversity and network complexity of arbuscular mycorrhizal fungi under climate change in relatively high-latitude regions. Front. Microbiol. 2022, 13, 930622. [Google Scholar] [CrossRef] [PubMed]
- Zak, D.R.; Holmes, W.E.; White, D.C.; Peacock, A.D.; Tilman, D. Plant diversity, soil microbial communities, and ecosystem function: Are there any links? Ecology 2003, 84, 2042–2050. [Google Scholar] [CrossRef]
- Jiao, H.; Xu, M.; Ji, J.; Wang, Y.; Zhang, Y.; Su, B.; Zhang, H.; Gao, C. Bacterial and fungal diversity, community composition, functional groups, and co-occurrence network succession in dryland and paddy soils along a 3000-year chronosequence. Catena 2025, 248, 108582. [Google Scholar] [CrossRef]
- Ulbrich, T.C.; Rivas-Ubach, A.; Tiemann, L.K.; Friesen, M.L.; Evans, S.E. Plant root exudates and rhizosphere bacterial communities shift with neighbor context. Soil Biol. Biochem. 2022, 172, 108753. [Google Scholar] [CrossRef]
- Busby, R.R.; Gebhart, D.L.; Stromberger, M.E.; Meiman, P.J.; Paschke, M.W. Early seral plant species’ interactions with an arbuscular mycorrhizal fungi community are highly variable. Appl. Soil Ecol. 2011, 48, 257–262. [Google Scholar] [CrossRef]
- Müller, L.M. Underground connections: Arbuscular mycorrhizal fungi influence on interspecific plant-plant interactions. Plant Physiol. 2021, 187, 1270–1272. [Google Scholar] [CrossRef]
- Vieira, S.; Sikorski, J.; Dietz, S.; Herz, K.; Schrumpf, M.; Bruelheide, H.; Scheel, D.; Friedrich, M.W.; Overmann, J. Drivers of the composition of active rhizosphere bacterial communities in temperate grasslands. ISME J. 2019, 14, 463–475. [Google Scholar] [CrossRef]
- Parasar, B.J.; Sharma, I.; Agarwala, N. Root exudation drives abiotic stress tolerance in plants by recruiting beneficial microbes. Appl. Soil Ecol. 2024, 198, 105351. [Google Scholar] [CrossRef]
- Yang, M.; Shi, Z.; Mickan, B.S.; Zhang, M.; Cao, L. Alterations to arbuscular mycorrhizal fungal community composition is driven by warming at specific elevations. PeerJ 2021, 9, e11792. [Google Scholar] [CrossRef]
- Han, B.; Zhang, W.; Yu, L.; Wei, X.; Geng, Y.; Zhou, H.; Lu, X.; Shao, X. Different response of arbuscular mycorrhizal fungal communities in roots and rhizosphere soil of Elymus nutans to long-term warming in an alpine meadow. J. Soil Sci. Plant Nutr. 2024, 24, 1149–1159. [Google Scholar] [CrossRef]
- Deepika, S.; Kothamasi, D. Soil moisture—A regulator of arbuscular mycorrhizal fungal community assembly and symbiotic phosphorus uptake. Mycorrhiza 2014, 25, 67–75. [Google Scholar] [CrossRef] [PubMed]
- Liao, X.; Zhao, J.; Xu, L.; Tang, L.; Li, J.; Zhang, W.; Xiao, J.; Xiao, D.; Hu, P.; Nie, Y.; et al. Arbuscular mycorrhizal fungi increase the interspecific competition between two forage plant species and stabilize the soil microbial network during a drought event: Evidence from the field. Appl. Soil Ecol. 2023, 185, 104805. [Google Scholar] [CrossRef]
- Ouhaddou, R.; Anli, M.; Ben-Laouane, R.; Boutasknit, A.; Baslam, M.; Meddich, A. The importance of the Glomus Genus as a potential candidate for sustainable agriculture under arid environments: A review. Int. J. Plant Biol. 2025, 16, 32. [Google Scholar] [CrossRef]
- Ma, X.; Li, J.; Ding, F.; Zheng, Y.; Chao, L.; Liu, H.; Liu, X.; Qu, H.; Bao, Y. Changes of arbuscular mycorrhizal fungal community and glomalin in the rhizosphere along the distribution gradient of zonal Stipa populations across the arid and semiarid steppe. Microbiol. Spectr. 2022, 10, e01489-22. [Google Scholar] [CrossRef]
- Kumar, N.; Haldar, S.; Saikia, R. Root exudation as a strategy for plants to deal with salt stress: An updated review. Environ. Exp. Bot. 2023, 216, 105518. [Google Scholar] [CrossRef]
- Wang, J.; Wang, G.G.; Zhang, B.; Yuan, Z.; Fu, Z.; Yuan, Y.; Zhu, L.; Ma, S.; Zhang, J. Arbuscular mycorrhizal fungi associated with tree species in a planted forest of Eastern China. Forests 2019, 10, 424. [Google Scholar] [CrossRef]
- Perez-Lamarque, B.; Selosse, M.A.; Öpik, M.; Morlon, H.; Martos, F. Cheating in arbuscular mycorrhizal mutualism: A network and phylogenetic analysis of mycoheterotrophy. New Phytol. 2020, 226, 1822–1835. [Google Scholar] [CrossRef]

| Genus | Species |
|---|---|
| Glomus | G. deserticola, G. mosseae, G. pansihalos, G. convolutum, G. magnicaule, G. etunicatum, G. tenebrosum, Glomus sp. 1 |
| Acaulospora | A. foveata, A. laevis, A. lacunosa, A. mellea, A.rehmii |
| Rhizophagus | R. Fasciculatus, R. manihotis, R. intraradices, R. cluram |
| Ambispora | A. Leptoticha, A. gerdemannii |
| Claroideoglomus | C. claroideum |
| Gigaspora | Gi. decipiens |
| Funneliformis | F. constrictum |
| Genus | Virtual Taxa |
|---|---|
| Glomus | Glomus Franke A1 VTX00076, Glomus Franke A1 VTX00269, Glomus Glo C VTX00323, Glomus Glo D VTX00103, Glomus Glo E VTX00319, Glomus Glo7 VTX00214, Glomus Glo16 VTX00120, Glomus PSAMG1 VTX00291, Glomus Yamato2005 D VTX00084, Glomus Yamato2005 D VTX00224, Glomus sp. VTX00165 Glomus sp. VTX00279, Glomus sp. VTX00304, Glomus caledonium VTX00065, Glomus Douhan9 VTX00056, Glomus lamellosu VTX00193, Glomus perpusillum VTX00287 Glomus GlAc3.1 VTX00190, Glomus GlAd3.3 VTX00289, Glomus MO G3 VTX00113 Glomus MO G4 VTX00166, Glomus MO G13 VTX00115, Glomus MO G14 VTX00083 Glomus MO G16 VTX00072, Glomus MO G22 VTX00125, Glomus Wirsel OTU12 VTX00188, Glomus Wirsel OTU6 VTX00202, Glomus acnaGlo2 VTX00155 Glomus acnaGlo7 VTX00057, Glomus viscosum VTX00063, Glomus ORVIN GLO3B VTX00223, Glomus ORVIN GLO3E VTX00309, Glomus ORVIN GLO3D VTX00310 Glomus ORVIN GLO4 VTX00278 |
| Archaeospora | Archaeospora Other1 VTX00005, Archaeospora sp. VTX00009 |
| Paraglomus | Paraglomus brasilianum VTX00239, Paraglomus occultum VTX00238 |
| Diversispora | Diversispora MO GC1 VTX00060 |
| Gigaspora | Gigaspora decipiens VTX00039 |
| Acaulospora | ------ |
| Genus | Specie | A | B | C | D | E | F | G | H |
|---|---|---|---|---|---|---|---|---|---|
| Glomus | G. deserticola | ✓ | |||||||
| G. mosseae | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | |
| G. pansihalos | ✓ | ✓ | ✓ | ✓ | |||||
| G. convolutum | ✓ | ✓ | ✓ | ✓ | ✓ | ||||
| G. magnicaule | ✓ | ✓ | ✓ | ✓ | |||||
| G. etunicatum | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | |
| G. tenebrosum | ✓ | ✓ | ✓ | ✓ | |||||
| Glomus sp. 1 | ✓ | ✓ | ✓ | ✓ | |||||
| Acaulospora | A. foveata | ✓ | ✓ | ✓ | ✓ | ||||
| A. laevis | ✓ | ✓ | ✓ | ✓ | ✓ | ||||
| A. lacunosa | ✓ | ✓ | ✓ | ✓ | |||||
| A. mellea | ✓ | ✓ | ✓ | ✓ | |||||
| A.rehmii | ✓ | ✓ | ✓ | ||||||
| Rhizophagus | R. fasciculatus | ✓ | ✓ | ✓ | ✓ | ||||
| R. manihotis | ✓ | ✓ | |||||||
| R. intraradices | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | |||
| R. cluram | ✓ | ✓ | ✓ | ||||||
| Ambispora | A. leptoticha | ✓ | ✓ | ✓ | |||||
| A.gerdemannii | ✓ | ✓ | ✓ | ||||||
| Claroideoglomus | C. claroideum | ✓ | ✓ | ✓ | |||||
| Gigaspora | Gi. decipiens | ✓ | ✓ | ✓ | |||||
| Funneliformis | F. constrictum | ✓ | ✓ | ✓ |
| Plants | pH | Electrical Conductivity (mS/cm) | Spore Density (Number per Gram) | Hyphal Length Density (m/g Dry Soil) | Species Richness |
|---|---|---|---|---|---|
| A | 8.79 ± 0.04 ab | 2.61 ± 0.06 a | 12.0 ± 0.69 ab | 6.38 ± 0.37 d | 10 |
| B | 8.76 ± 0.04 ab | 2.49 ± 0.03 ab | 10.67 ± 0.62 ab | 7.22 ± 0.42 c | 12 |
| C | 8.81 ± 0.07 a | 2.43 ± 0.05 b | 15.0 ± 0.87 a | 16.94 ± 0.98 a | 13 |
| D | 8.70 ± 0.06 b | 2.07 ± 0.04 d | 13.33 ± 0.77 ab | 13.01 ± 0.75 b | 15 |
| E | 7.49 ± 0.05 c | 0.48 ± 0.01 e | 8.67 ± 0.50 b | 6.67 ± 0.39 d | 9 |
| F | 7.35 ± 0.04 d | 0.40 ± 0.02 e | 7.67 ± 0.44 b | 8.88 ± 0.51 c | 11 |
| G | 8.77 ± 0.06 ab | 2.30 ± 0.18 c | 4.67 ± 0.27 c | 3.94 ± 0.23 e | 7 |
| H | 7.30 ± 0.07 e | 0.20 ± 0.01 f | 4.0 ± 0.23 c | 5.88 ± 0.34 d | 10 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mei, L.; Liu, Y.; Wang, Z.; Xiong, Z.; Wang, Y.; Jin, T.; Yang, X. Diversity and Community Structure of Rhizosphere Arbuscular Mycorrhizal Fungi in Songnen Grassland Saline–Alkali-Tolerant Plants: Roles of Environmental Salinity and Plant Species Identity. Agronomy 2025, 15, 2070. https://doi.org/10.3390/agronomy15092070
Mei L, Liu Y, Wang Z, Xiong Z, Wang Y, Jin T, Yang X. Diversity and Community Structure of Rhizosphere Arbuscular Mycorrhizal Fungi in Songnen Grassland Saline–Alkali-Tolerant Plants: Roles of Environmental Salinity and Plant Species Identity. Agronomy. 2025; 15(9):2070. https://doi.org/10.3390/agronomy15092070
Chicago/Turabian StyleMei, Linlin, Yingbin Liu, Zixian Wang, Zixuan Xiong, Yuze Wang, Tianqi Jin, and Xuechen Yang. 2025. "Diversity and Community Structure of Rhizosphere Arbuscular Mycorrhizal Fungi in Songnen Grassland Saline–Alkali-Tolerant Plants: Roles of Environmental Salinity and Plant Species Identity" Agronomy 15, no. 9: 2070. https://doi.org/10.3390/agronomy15092070
APA StyleMei, L., Liu, Y., Wang, Z., Xiong, Z., Wang, Y., Jin, T., & Yang, X. (2025). Diversity and Community Structure of Rhizosphere Arbuscular Mycorrhizal Fungi in Songnen Grassland Saline–Alkali-Tolerant Plants: Roles of Environmental Salinity and Plant Species Identity. Agronomy, 15(9), 2070. https://doi.org/10.3390/agronomy15092070






