Multiplex PCR for Discriminating Host Plant Associations of Hyalesthes obsoletus (Hemiptera: Cixiidae), a Key Vector and Driver of ‘Ca. Phytoplasma solani’ Epidemiology
Abstract
1. Introduction
2. Materials and Methods
2.1. Hyalesthes obsoletus Specimens Used for mtDNA COI Genotyping
2.2. PCR Amplification and Sequencing of COI Gene
2.3. COI Sequence Comparison and Haplotype Network Reconstruction
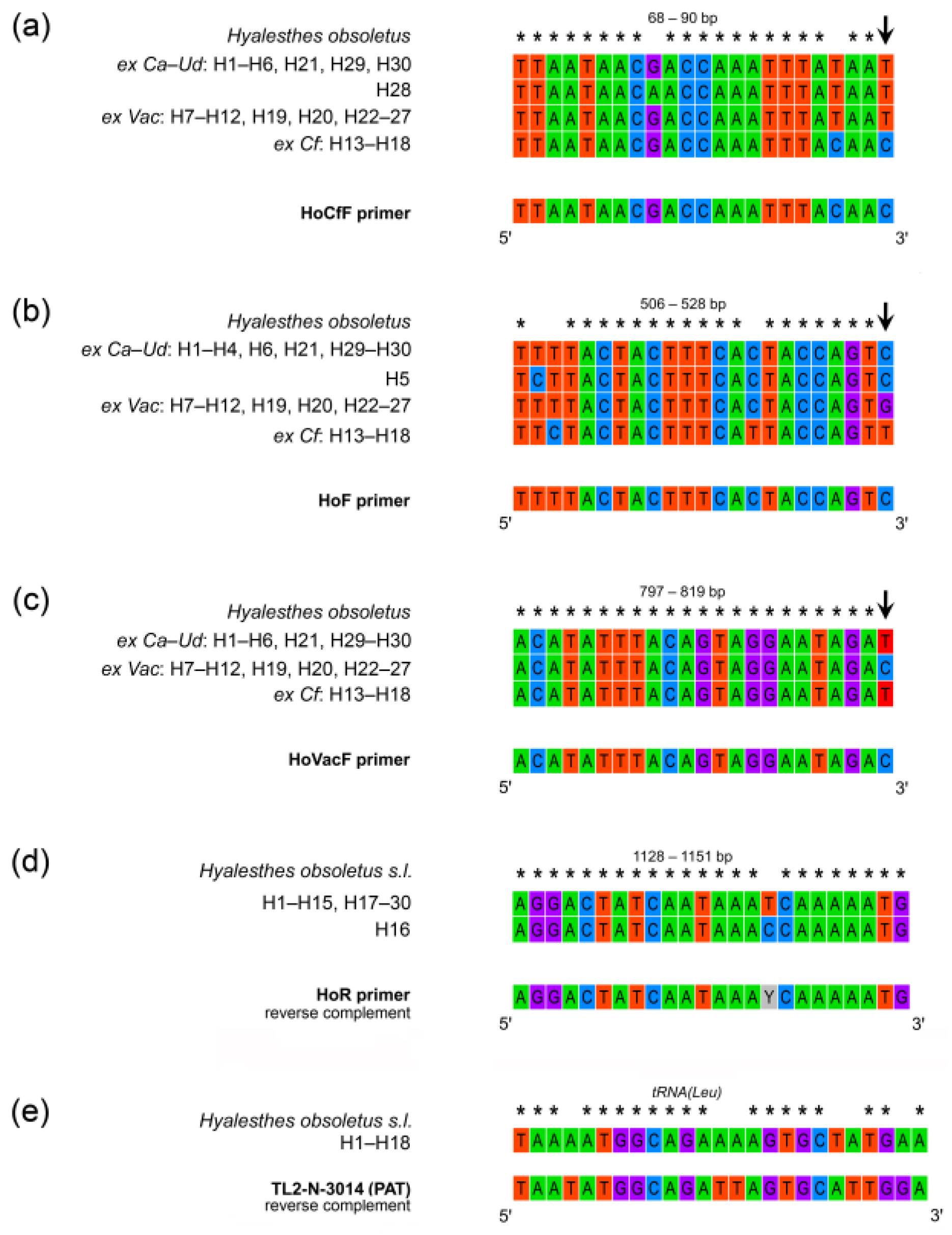
2.4. Design of Multiplex PCR Tool for Specific Identification of Hyalesthes obsoletus Host Plant Associations
2.5. Evaluation of the Strain Diversity of ‘Ca. Phytoplasma solani’ in Hyalesthes obsoletus and Shared Host Plants
3. Results
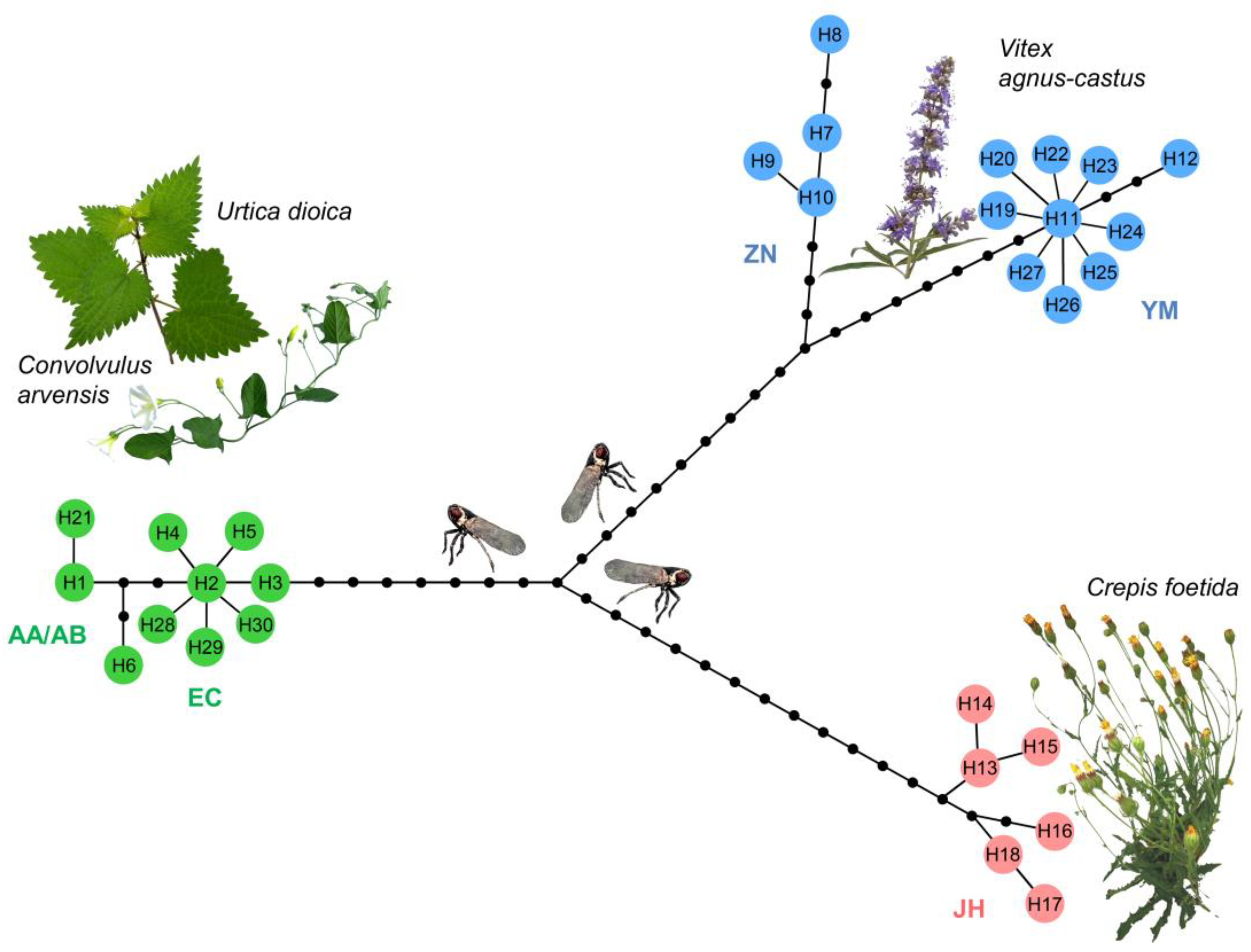
3.1. Haplotype Diversity and Differentiation of Hyalesthes obsoletus Host Plant Associations
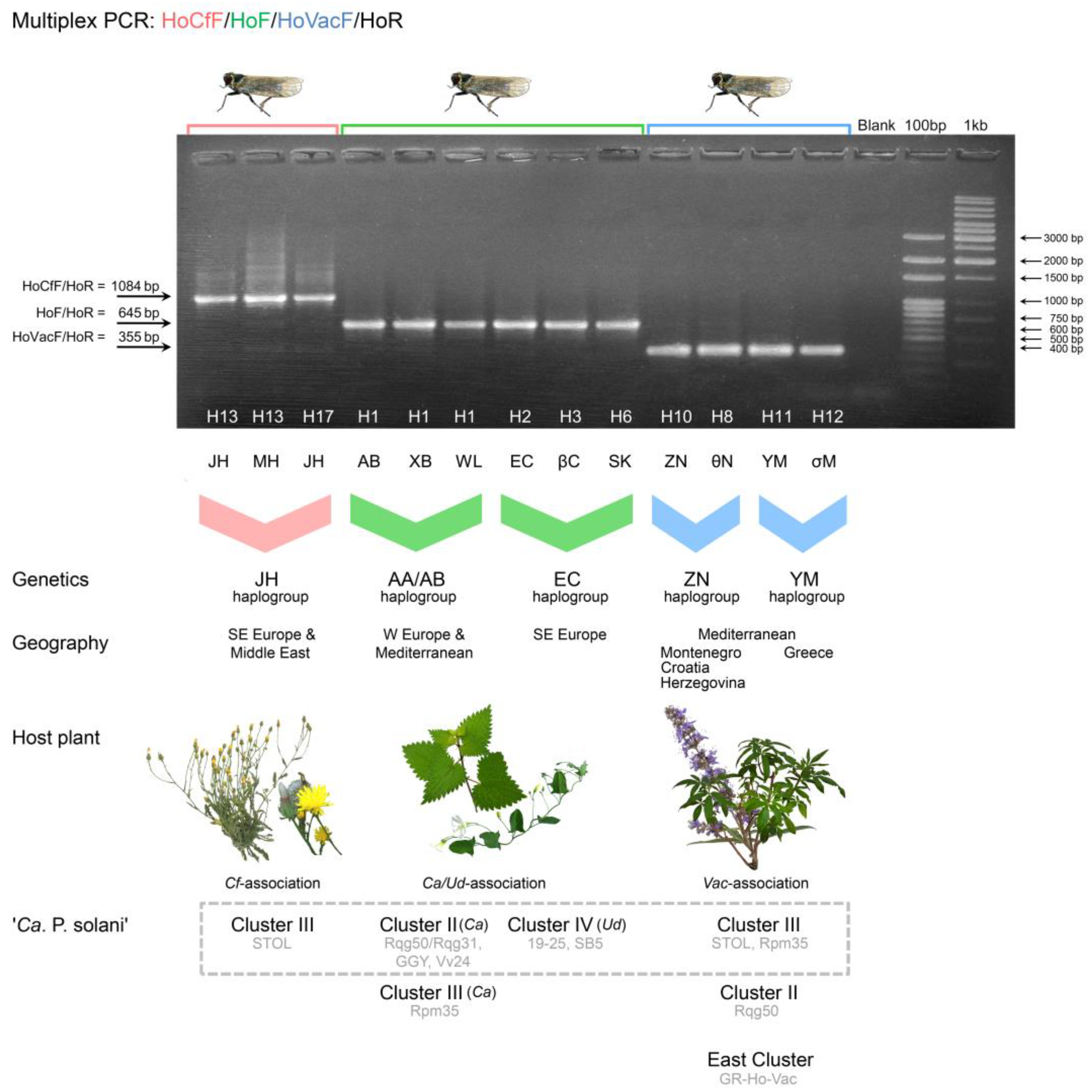
3.2. Multiplex PCR for Discriminating Host Plant Associations of Hyalesthes obsoletus
3.3. Linking ‘Ca. Phytoplasma solani’ Epidemiology with Host Plant-Specialized Hyalesthes obsoletus
4. Discussion
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hebert, P.D.; Cywinska, A.; Ball, S.L.; DeWaard, J.R. Biological identifications through DNA barcodes. Proc. R. Soc. Lond. B 2003, 270, 313–321. [Google Scholar]
- Hebert, P.D.; Gregory, T.R. The promise of DNA barcoding for taxonomy. Syst. Biol. 2005, 54, 852–859. [Google Scholar] [CrossRef]
- Toševski, I.; Caldara, R.; Jović, J.; Baviera, C.; Hernández-Vera, G.; Gassmann, A.; Emerson, B.C. Revision of Mecinus heydenii species complex (Curculionidae): Integrative taxonomy reveals multiple species exhibiting host specialization. Zool. Scr. 2014, 43, 34–51. [Google Scholar]
- Toševski, I.; Caldara, R.; Jović, J.; Hernańdez-Vera, G.; Baviera, C.; Gassmann, A.; Emerson, B.C. Host-associated divergence and taxonomy in the Rhinusa pilosa Gyllenhal species complex: An integrative approach. Syst. Entomol. 2015, 40, 268–287. [Google Scholar]
- Toševski, I.; Caldara, R.; Jović, J.; Hernańdez-Vera, G.; Baviera, C.; Gassmann, A.; Emerson, B.C. Morphological, molecular and biological evidence reveal two cryptic species in Mecinus janthinus Germar (Coleoptera, Curculionidae), a successful biological control agent of Dalmatian toadflax, Linaria dalmatica (Lamiales, Plantaginaceae). Syst. Entomol. 2011, 36, 741–753. [Google Scholar]
- Tay, W.T.; Evans, G.A.; Boykin, L.M.; De Barro, P.J. Will the real Bemisia tabaci please stand up? PLoS ONE 2012, 7, e50550. [Google Scholar]
- Antil, S.; Abraham, J.S.; Sripoorna, S.; Maurya, S.; Dagar, J.; Makhija, S.; Bhagat, P.; Gupta, R.; Sood, U.; Lal, R.; et al. DNA barcoding, an effective tool for species identification: A review. Mol. Biol. Rep. 2023, 50, 761–775. [Google Scholar]
- Guo, M.; Yuan, C.; Tao, L.; Cai, Y.; Zhang, W. Life barcoded by DNA barcodes. Conserv. Genet. Res. 2022, 14, 351–365. [Google Scholar]
- Di Camillo, C.G.; Gravili, C.; De Vito, D.; Pica, D.; Piraino, S.; Puce, S.; Cerrano, C. The importance of applying Standardised Integrative Taxonomy when describing marine benthic organisms and collecting ecological data. Invertebr. Syst. 2018, 32, 794–802. [Google Scholar]
- Bertin, S.; Picciau, L.; Ács, Z.; Alma, A.; Bosco, D. Molecular identification of the Hyalesthes species (Hemiptera: Cixiidae) occurring in vineyard agroecosystems. Ann. Appl. Biol. 2010, 157, 435–445. [Google Scholar]
- Bertin, S.; Picciau, L.; Ács, Z.; Alma, A.; Bosco, D. Molecular differentiation of four Reptalus species (Hemiptera: Cixiidae). Bull. Entomol. Res. 2010, 100, 551–558. [Google Scholar] [PubMed]
- Toševski, I.; Jović, J.; Krstić, O.; Gassmann, A. PCR-RFLP-based method for reliable discrimination of cryptic species within Mecinus janthinus species complex (Mecinini, Curculionidae) introduced in North America for biological control of invasive toadflaxes. Biol. Control 2013, 58, 563–573. [Google Scholar]
- Andrews, K.J.; Bester, R.; Manrakhan, A.; Maree, H.J. A multiplex PCR assay for the identification of fruit flies (Diptera: Tephritidae) of economic importance in South Africa. Sci. Rep. 2022, 12, 13089. [Google Scholar]
- Gostel, M.R.; Kress, W.J. The Expanding Role of DNA Barcodes: Indispensable Tools for Ecology, Evolution, and Conservation. Diversity 2022, 14, 213. [Google Scholar]
- Hebert, P.D.; Ratnasingham, S.; De Waard, J.R. Barcoding animal life: Cytochrome c oxidase subunit 1 divergences among closely related species. Proc. R. Soc. Lond. B 2003, 270, S96–S99. [Google Scholar]
- Maixner, M. Transmission of German grapevine yellows (Vergilbungskrankheit) by the planthopper Hyalesthes obsoletus (Auchenorrhyncha: Cixiidae). Vitis 1994, 33, 103–104. [Google Scholar]
- Sforza, R.; Clair, D.; Daire, X.; Larrue, J.; Boudon-Padieu, E. The role of Hyalesthes obsoletus (Hemiptera: Cixiidae) in the occurrence of bois noir of grapevines in France. J. Phytopathol. 1998, 146, 549–556. [Google Scholar]
- Quaglino, F.; Zhao, Y.; Casati, P.; Bulgari, D.; Bianco, P.A.; Wei, W.; Davis, R.E. ‘Candidatus Phytoplasma solani’, a novel taxon associated with stolbur- and bois noir-related diseases of plants. Int. J. Syst. Evol. Microbiol. 2013, 63, 2879–2894. [Google Scholar]
- EFSA PLH Panel (EFSA Panel on Plant Health). Scientific Opinion on the pest categorisation of Candidatus Phytoplasma solani. EFSA J. 2014, 12, 3924–3927. [Google Scholar]
- Langer, M.; Maixner, M. Molecular characterisation of grapevine yellows associated phytoplasmas of the stolbur group based on RFLP analysis of non-ribosomal DNA. Vitis 2004, 43, 191–199. [Google Scholar]
- Kosovac, A.; Radonjić, S.; Hrnčić, S.; Krstić, O.; Toševski, I.; Jović, J. Molecular tracing of the transmission routes of bois noir in Mediterranean vineyards of Montenegro and experimental evidence for the epidemiological role of Vitex agnus-castus (Lamiaceae) and associated Hyalesthes obsoletus (Cixiidae). Plant Pathol. 2016, 65, 285–298. [Google Scholar]
- Pierro, R.; Panattoni, A.; Passera, A.; Materazzi, A.; Luvisi, A.; Loni, A.; Ginanni, M.; Lucchi, A.; Bianco, P.A.; Quaglino, F. Proposal of a new Bois noir epidemiological pattern related to ‘Candidatus Phytoplasma solani’ strains characterized by a possiblemoderate virulence in Tuscany. Pathogens 2020, 9, 268. [Google Scholar] [PubMed]
- Jović, J.; Marinković, S.; Jakovljević, M.; Krstić, O.; Cvrković, T.; Mitrović, M.; Toševski, I. Symptomatology, (Co)occurrence and Differential Diagnostic PCR Identification of ‘Ca. Phytoplasma solani’ and ‘Ca. Phytoplasma convolvuli’ in Field Bindweed. Pathogens 2021, 10, 160. [Google Scholar] [PubMed]
- Cvrković, T.; Jović, J.; Krstić, O.; Marinković, S.; Jakovljević, M.; Mitrović, M.; Toševski, I. Epidemiological Role of Dictyophara europaea (Hemiptera: Dictyopharidae) in the Transmission of ‘Candidatus Phytoplasma solani’. Horticulturae 2022, 8, 654. [Google Scholar]
- Sforza, R.; Bourgoin, T.; Wilson, S.W.; Boudon-Padieu, E. Field observations, laboratory rearing and descriptions of immatures of the planthopper Hyalesthes obsoletus (Hemiptera: Cixiidae). Eur. J. Entomol. 1999, 96, 409–418. [Google Scholar]
- Jović, J.; Cvrković, T.; Mitrović, M.; Krnjajić, S.; Redinbaugh, M.G.; Pratt, R.C.; Gingery, R.E.; Hogenhout, S.A.; Toševski, I. Roles of stolbur phytoplasma and Reptalus panzeri (Cixiinae, Auchenorrhyncha) in the epidemiology of Maize redness in Serbia. Eur. J. Plant Pathol. 2007, 118, 85–89. [Google Scholar]
- Jović, J.; Cvrković, T.; Mitrović, M.; Krnjanjić, S.; Petrović, A.; Redinbaugh, M.G.; Pratt, R.C.; Hogenhout, S.A.; Toševski, I. Stolbur phytoplasma transmission to maize by Reptalus panzeri and the disease cycle of maize redness in Serbia. Phytopathology 2009, 99, 1053–1061. [Google Scholar]
- Radonjić, S.; Hrnčić, S.; Jović, J.; Cvrković, T.; Krstić, O.; Krnjajić, S.; Toševski, I. Occurrence and distribution of grapevine yellows caused by stolbur phytoplasma in Montenegro. J. Phytopathol. 2009, 157, 682–685. [Google Scholar]
- Cvrković, T.; Jović, J.; Mitrović, M.; Krstić, O.; Toševski, I. Experimental and molecular evidence of Reptalus panzeri as a natural vector of bois noir. Plant Pathol. 2014, 63, 42–53. [Google Scholar]
- Atanasova, B.; Jakovljević, M.; Spasov, D.; Jović, J.; Mitrović, M.; Toševski, I.; Cvrković, T. The molecular epidemiology of bois noir grapevine yellows caused by ‘Candidatus Phytoplasma solani’ in the Republic of Macedonia. Eur. J. Plant Pathol. 2015, 142, 759–770. [Google Scholar]
- Delić, D.; Balech, B.; Radulović, M.; Lolić, B.; Karačić, A.; Vukosavljević, V.; Đurić, G.; Cvetković, T.J. Vmp1 and stamp genes variability of ‘Candidatus phytoplasma solani’ in Bosnian and Herzegovinian grapevine. Eur. J. Plant Pathol. 2016, 145, 221–225. [Google Scholar]
- Balakishiyeva, G.; Bayramova, J.; Mammadov, A.; Salar, P.; Danet, J.L.; Ember, I.; Verdin, E.; Foissac, X.; Huseynova, I. Important genetic diversity of “Candidatus Phytoplasma solani” related strains associated with Bois noir grapevine yellows and planthoppers in Azerbaijan. Eur. J. Plant. Pathol. 2018, 151, 937–946. [Google Scholar]
- Sémétey, O.; Gaudin, J.; Danet, J.L.; Salar, P.; Theil, S.; Fontaine, M.; Krausz, M.; Chaisse, E.; Eveillard, S.; Verdin, E.; et al. Lavender decline in France is associated with chronic infection by lavender-specific strains of “Candidatus Phytoplasma solani”. Appl. Environ. Microbiol. 2018, 84, e01507–e01518. [Google Scholar]
- Quaglino, F.; Passera, A.; Faccincani, M.; Moussa, A.; Pozzebon, A.; Sanna, F.; Casati, P.; Bianco, P.A.; Mori, N. Molecular and spatial analyses reveal new insights on Bois noir epidemiology in Franciacorta vineyards. Ann. Appl. Biol. 2021, 179, 151–168. [Google Scholar]
- Mehle, N.; Kavčič, S.; Mermal, S.; Vidmar, S.; Pompe Novak, M.; Riedle-Bauer, M.; Brader, G.; Kladnik, A.; Dermastia, M. Geographical and Temporal Diversity of ‘Candidatus Phytoplasma solani’ in Wine-Growing Regions in Slovenia and Austria. Front. Plant Sci. 2022, 13, 889675. [Google Scholar]
- Çağlar, B.K.; Şimşek, E. Detection and Multigene Typing of ‘Candidatus Phytoplasma solani’-Related Strains Infecting Tomato and Potato Plants in Different Regions of Turkey. Pathogens 2022, 11, 1031. [Google Scholar] [PubMed]
- Usta, M.; Guller, A.; Sipahioglu, H.M. Detection, in silico analysis and molecular diversity of phytoplasmas from solanaceous crops in Turkey. Plant Prot. Sci. 2022, 58, 31–39. [Google Scholar]
- Kosovac, A.; Jakovljević, M.; Krstić, O.; Cvrković, T.; Mitrović, M.; Toševski, I.; Jović, J. Role of plant-specialized Hyalesthes obsoletus associated with Convolvulus arvensis and Crepis foetida in the transmission of ‘Candidatus Phytoplasma solani’-inflicted bois noir disease of grapevine in Serbia. Eur. J. Plant Pathol. 2019, 153, 183–195. [Google Scholar]
- Bressan, A.; Sémétey, O.; Nusillard, B.; Clair, D.; Boudon-Padieu, E. Insect vectors (Hemiptera: Cixiidae) and pathogens associated with the disease syndrome “basses richesses” of sugar beet in France. Plant Dis. 2008, 92, 113–119. [Google Scholar]
- Mitrović, M.; Jakovljević, M.; Jović, J.; Krstić, O.; Kosovac, A.; Trivellone, V.; Jermini, M.; Toševski, I.; Cvrković, T. ‘Candidatus Phytoplasma solani’ genotypes associated with potato stolbur in Serbia and the role of Hyalesthes obsoletus and Reptalus panzeri (Hemiptera, Cixiidae) as natural vectors. Eur. J. Plant Pathol. 2016, 144, 619–630. [Google Scholar]
- Aleksić, Ž.; Šutić, D.; Aleksić, D. Transmission intensity of stolbur virus by means of Hyalesthes obsoletus Sign. on some host plants. Plant Prot. 1967, 93–95, 67–73. [Google Scholar]
- Emeljanov, A.F. Nomenclatorial changes in the family Cixiidae (Homoptera, Auchenorrhyncha, Fulgoroidea), with fixation of type species of the genus Reptalus Emeljanov, 1971 and description of a new subgenus. Zootaxa 2020, 4780, 197–200. [Google Scholar]
- Gatineau, F.; Larrue, J.; Clair, D.; Lorton, F.; Richard-Molard, M.; Boudon-Padieu, E. A new natural planthopper vector of “stolbur” phytoplasma in the genus Pentastiridius (Hemiptera: Cixiidae). Eur. J. Plant Pathol. 2001, 107, 263–271. [Google Scholar] [CrossRef]
- Trivellone, V.; Pinzauti, F.; Bagnoli, B. Reptalus quinquecostatus (Dufour) (Auchenorrhyncha Cixiidae) as a possible vector of “stolbur”-phytoplasma in a vineyard in Tuscany. Redia 2005, 88, 103–108. [Google Scholar]
- Pinzauti, F.; Trivellone, V.; Bagnoli, B. Ability of Reptalus quinquecostatus (Hemiptera: Cixiidae) to inoculate “stolbur” phytoplasma to artificial feeding medium. Ann. Appl. Biol. 2008, 153, 299–305. [Google Scholar]
- Chuche, J.; Danet, J.L.; Salar, P.; Foissac, X.; Thiéry, D. Transmission of ‘Candidatus Phytoplasma solani’ by Reptalus quinquecostatus (Hemiptera: Cixiidae). Ann. Appl. Biol. 2016, 169, 214–223. [Google Scholar]
- Jović, J.; Riedle-Bauer, M.; Chuche, J. Vector Role of Cixiids and Other Planthopper Species. In Phytoplasmas: Plant Pathogenic Bacteria-II: Transmission and Management of Phytoplasma Associated Diseases; Bertaccini, A., Weintraub, P.G., Rao, G.P., Mori, N., Eds.; Springer: Singapore, 2019; Volume 2, pp. 79–113. [Google Scholar]
- Pfitzer, R.; Varrelmann, M.; Hesse, G.; Eini, O. Molecular Detection of Pentastiridius leporinus, the Main Vector of the Syndrome ‘Basses Richesses’ in Sugar Beet. Insects 2022, 13, 992. [Google Scholar] [PubMed]
- Holzinger, W.E.; Kammerlander, I.; Nickel, H. The Auchenorrhyncha of Central Europe, Fulgoromorpha, Cicadomorpha Excl. Cicadellidae; Brill Academic Publishers: Leiden, The Netherlands, 2003; pp. 1–673. [Google Scholar]
- Emeljanov, A.F. Planthoppers of the Family Cixiidae of Russia and Adjacent Territories; KMK Scientific Press Ltd.: St.-Petersburg, Moscow, Russia, 2015; Volume 177, pp. 1–253. Keys to species of the fauna of Russia published by Zoological Institute of Russian Academy of Sciences. (In Russian) [Google Scholar]
- Sharon, R.; Soroker, V.; Wesley, S.D.; Zahavi, T.; Harari, A.; Weintraub, P.G. Vitex agnus-castus is a preferred host plant for Hyalesthes obsoletus. J. Chem. Ecol. 2005, 31, 1051–1063. [Google Scholar]
- Kessler, S.; Schaerer, S.; Delabays, N.; Turlings, T.C.; Trivellone, V.; Kehrli, P. Host plant preferences of Hyalesthes obsoletus, the vector of the grapevine yellows disease ‘bois noir’, in Switzerland. Entomol. Exp. Appl. 2011, 139, 60–67. [Google Scholar]
- Aryan, A.; Brader, G.; Mörtel, J.; Pastar, M.; Riedle-Bauer, M. An abundant ‘Candidatus Phytoplasma solani’ tuf b strain is associated with grapevine, stinging nettle and Hyalesthes obsoletus. Eur. J. Plant Pathol. 2014, 140, 213–227. [Google Scholar]
- Johannesen, J.; Lux, B.; Michel, K.; Seitz, A.; Maixner, M. Invasion biology and host specificity of the grapevine yellows disease vector Hyalesthes obsoletus in Europe. Entomol. Exp. Appl. 2008, 126, 217–227. [Google Scholar]
- Imo, M.; Maixner, M.; Johannesen, J. Sympatric diversification vs. immigration: Deciphering host-plant specialization in a polyphagous insect, the stolbur phytoplasma vector Hyalesthes obsoletus (Cixiidae). Mol. Ecol. 2013, 22, 2188–2203. [Google Scholar] [PubMed]
- Kosovac, A.; Johannesen, J.; Krstić, O.; Mitrović, M.; Cvrković, T.; Toševski, I.; Jović, J. Widespread plant specialization in the polyphagous planthopper Hyalesthes obsoletus (Cixiidae), a major vector of stolbur phytoplasma: Evidence of cryptic speciation. PLoS ONE 2018, 13, e0196969. [Google Scholar]
- Cimerman, A.; Pacifico, D.; Salar, P.; Marzachi, C.; Foissac, X. Striking diversity of vmp1, a variable gene encoding a putative membrane protein of the stolbur phytoplasma. Appl. Environ. Microbiol. 2009, 75, 2951–2957. [Google Scholar]
- Fialová, R.; Válová, P.; Balakishiyeva, G.; Danet, J.L.; Šafárová, D.; Foissac, X.; Navrátil, M. Genetic variability of stolbur phytoplasma in annual crop and wild plant species in south Moravia. J. Plant Pathol. 2009, 91, 411–416. [Google Scholar]
- Fabre, A.; Danet, J.L.; Foissac, X. The stolbur phytoplasma antigenic membrane protein gene stamp is submitted to diversifying positive selection. Gene 2011, 472, 37–41. [Google Scholar]
- Johannesen, J.; Foissac, X.; Kehrli, P.; Maixner, M. Impact of vector dispersal and host-plant fidelity on the dissemination of an emerging plant pathogen. PLoS ONE 2012, 7, e51809. [Google Scholar]
- Johannesen, J.; Riedle-Bauer, M. Origin of a sudden mass occurrence of the stolbur phytoplasma vector Hyalesthes obsoletus (Cixiidae) in Austria. Ann. Appl. Biol. 2014, 165, 488–495. [Google Scholar]
- Chuche, J.; Danet, J.L.; Rivoal, J.B.; Arricau-Bouvery, N.; Thiéry, D. Minor cultures as hosts for vectors of extensive crop diseases: Does Salvia sclarea act as a pathogen and vector reservoir for lavender decline? J. Pest Sci. 2018, 91, 145–155. [Google Scholar]
- DeSalle, R.; Goldstein, P. Review and interpretation of trends in DNA barcoding. Front. Ecol. Evol. 2019, 7, 302. [Google Scholar]
- Simon, C.; Frati, F.; Beckenbach, A.; Crespi, B.; Liu, H.; Flook, P. Evolution, weighting, and phylogenetic utility of mitochondrial gene sequences and a compilation of conserved polymerase chain reaction primers. Ann. Entomol. Soc. Am. 1994, 87, 651–701. [Google Scholar] [CrossRef]
- Folmer, O.; Black, M.; Hoeh, W.; Lutz, R.; Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Marine Biol. Biotechnol. 1994, 3, 294–299. [Google Scholar]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Clement, M.; Posada, D.; Crandall, K. TCS: A computer program to estimate gene genealogies. Mol. Ecol. 2000, 9, 1657–1660. [Google Scholar]
- Templeton, A.R.; Crandall, K.A.; Sing, C.F. A cladistic analysis of phenotypic associations with haplotypes inferred from restriction endonuclease mapping and DNA sequence data. III. Cladogram estimation. Genet 1992, 132, 619–633. [Google Scholar]
- Bandelt, H.J.; Forster, P.; Röhl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 1999, 16, 37–48. [Google Scholar]
- Bressan, A.; Holzinger, W.E.; Nusillard, B.; Sémétey, O.; Gatineau, F.; Simonato, M.; Boudon-Padieu, E. Identification and biological traits of a planthopper from the genus Pentastiridius (Hemiptera: Cixiidae) adapted to an annual cropping rotation. Eur. J. Entomol. 2009, 106, 405–413. [Google Scholar]
- Đurić, Z.; Hrnčić, S.; Delić, D. Morphological and molecular identification of Hyalesthes obsoletus Signoret (Auchenorrhyncha: Cixiidae) in Herzegovina vineyards. Mitt. Klosterneubg. 2017, 67, 177–181. [Google Scholar]
- Koressaar, T.; Remm, M. Enhancements and modifications of primer design program Primer3. Bioinformatics 2007, 23, 1289–1291. [Google Scholar] [CrossRef] [PubMed]
- Untergasser, A.; Cutcutache, I.; Koressaar, T.; Ye, J.; Faircloth, B.C.; Remm, M.; Rozen, S.G. Primer3—New capabilities and interfaces. Nucleic Acids Res. 2012, 40, e115. [Google Scholar] [PubMed]
- Quaglino, F.; Maghradze, D.; Casati, P.; Chkhaidze, N.; Lobjanidze, M.; Ravasio, A.; Passera, A.; Venturini, G.; Failla, O.; Bianco, P.A. Identification and characterization of new ‘Candidatus Phytoplasma solani’ strains associated with bois noir disease in Vitis vinifera L. cultivars showing a range of symptom severity in Georgia, the Caucasus region. Plant Dis. 2016, 100, 904–915. [Google Scholar] [CrossRef] [PubMed]
- Pierro, R.; Passera, A.; Panattoni, A.; Casati, P.; Luvisi, A.; Rizzo, D.; Bianco, P.A.; Quaglino, F.; Materazzi, A. Molecular Typing of Bois Noir Phytoplasma Strains in the Chianti Classico Area (Tuscany, Central Italy) and Their Association with Symptom Severity in Vitis vinifera ‘Sangiovese’. Phytopathology 2018, 108, 362–373. [Google Scholar] [PubMed]
- Donnua, S.; Parkpoom, B.; Thaipong, K. Candidatus phytoplasma solani associated with papaya phytoplasma disease in Thailand. Khon Kaen Agric. J. 2021, 49, 1249–1258. [Google Scholar]
- Jović, J. Use of mitochondrial divergence in plant-specialized populations of Hyalesthes obsoletus for identification of ‘Candidatus Phytoplasma solani’ epidemiology. Phytopathogenic Mollicutes 2019, 9, 91–92. [Google Scholar] [CrossRef]
- Chantaranothai, P. A revision of the genus Vitex (Lamiaceae) in Thailand. Trop. Nat. Hist. 2011, 11, 91–118. [Google Scholar]
- Maixner, M.; Albert, A.; Johannesen, J. Survival relative to new and ancestral host plants, phytoplasma infection, and genetic constitution in host races of a polyphagous insect disease vector. Ecol. Evol. 2014, 4, 3082–3092. [Google Scholar]
- Hoch, H.; Remane, R. Evolution und Speziation der Zikaden-Gattung Hyalesthes SIGNORET, 1865 (Homoptera Auchenorrhyncha Fulgoroidea Cixiidae); Marburger Entomologische Publikationen: Marburg, Germany, 1985; pp. 1–427. [Google Scholar]
- Jamshidi, E.; Murolo, S.; Ravari, S.B.; Salehi, M.; Romanazzi, G. Multilocus Genotyping of ‘Candidatus Phytoplasma Solani’ Associated with Grapevine Bois Noir in Iran. Biology 2022, 11, 835. [Google Scholar]
- Mitrović, M.; Marinković, S.; Cvrković, T.; Jović, J.; Krstić, O.; Jakovljević, M. Framework for risk assessment of ‘Candidatus Phytoplasma solani’ associated diseases outbreaks in agroecosystems in Serbia. J. Plant Pathol. 2022, 104, 537–552. [Google Scholar]

| COI Marker Gene (1467 bp) | Associated Haplotype COI-COII (16S-ND1) 1 | Country | Locality | Host Plant Association 2 | ||
|---|---|---|---|---|---|---|
| Haplotype | GenBank Acc No | Frequency | ||||
| H1 | MK172874 | 5 | A (B) | Montenegro | Podgorica | Ca |
| Greece | Kilkis | Ca | ||||
| W (L) | Greece | Kilkis | Ca | |||
| W (B) | Greece | Profitis | Ud | |||
| X (B) | Greece | Kilkis | Ca | |||
| H2 | MK172875 | 13 | E (C) | Serbia | Topola | Ca |
| Serbia | Boljetin | Ud | ||||
| Serbia | Knjaževac | Ud | ||||
| Serbia | Bačka Topola | Ud | ||||
| Serbia | Srednjevo | Ud | ||||
| Serbia | Grnčar | Ud | ||||
| Montenegro | Podgorica | Ud | ||||
| Romania | Petrevo selo | Ca | ||||
| π (C) | North Macedonia | Strumica | Ca | |||
| ξ (Q) | Serbia | Zaječar | Ud | |||
| ω (C) | Serbia | Bačka Topola | Ud | |||
| R (C) | Serbia | Gakovo | Ud | |||
| α (C) | Montenegro | Podgorica | Ud | |||
| H3 | MK172876 | 1 | β (C) | Montenegro | Podgorica | Ud |
| H4 | MK172877 | 1 | F (C) | Serbia | Gakovo | Ud |
| H5 | MK172878 | 7 | E (C) | Serbia | Vranje | Ud |
| North Macedonia | Strumica | Ud | ||||
| North Macedonia | Strumica | Ca | ||||
| Greece | Filadelfio | Ud | ||||
| Greece | Profitis | Ud | ||||
| ψ (C) | North Macedonia | Strumica | Ca | |||
| ρ (C) | Greece | Profitis | Ud | |||
| H6 | MK172879 | 1 | S (K) | Turkey | Erzincan | Ca |
| H7 | MK172880 | 1 | η (N) | Montenegro | Ulcinj | Vac |
| H8 | MK172881 | 1 | θ (N) | Montenegro | Ulcinj | Vac |
| H9 | MK172882 | 1 | Z (N) | Montenegro | Kamenari | Vac |
| H10 | MK172883 | 2 | Z (N) | Montenegro | Kamenari | Vac |
| γ (N) | Montenegro | Bar | Vac | |||
| H11 | MK172884 | 4 | Y (M) | Greece | Appolonia | Vac |
| Greece | Arethousa | Vac | ||||
| Greece | Vrasna | Vac | ||||
| Greece | Larisa | Vac | ||||
| H12 | MK172885 | 1 | σ (M) | Greece | Lesbos | Vac |
| H13 | MK172886 | 12 | J (H) | Serbia | Deligrad | Cf |
| Serbia | Porečka reka | Cf | ||||
| Serbia | Negotin | Cf | ||||
| Serbia | Tamnič | Cf | ||||
| Serbia | Pirot | Cf | ||||
| Serbia | Jasenovik | Cf | ||||
| M (H) | Serbia | Negotin | Cf | |||
| Serbia | Tamnič | Cf | ||||
| Romania | Calafat | Cf | ||||
| Bulgaria | Vidin | Cf | ||||
| λ (H) | Serbia | Vranjska banja | Cf | |||
| μ (H) | Serbia | Deligrad | Cf | |||
| H14 | MK172887 | 1 | U (H) | Serbia | Porečka reka | Cf |
| H15 | MK172888 | 1 | V (H) | Serbia | Porečka reka | Cf |
| H16 | MK172889 | 1 | J (H) | Turkey | Kırşehir | Cf |
| H17 | MK172890 | 1 | J (H) | Turkey | Erzincan | Cf |
| H18 | MK172891 | 1 | J (H) | Turkey | Erzincan | Cf |
| COI Marker Gene (1467 bp) | Country | Region | Locality | GPS Coordinates | Host Plant Association 1 | |
|---|---|---|---|---|---|---|
| Haplotype | GenBank Acc No | |||||
| H2 | MK172875 | Greece | Peloponnese (south) | Sparta, Mystras | N37 04.548 E22 21.769 | Ud |
| Greece | Peloponnese (south) | Taygetus, Artemisia | N37 05.818 E22 13.719 | Ud | ||
| H7 | MK172880 | Croatia | Primorje–Gorski Kotar | Krk, Stara Baška | N44 58.138 E14 39.797 | Vac |
| H11 | MK172884 | Greece | Epirus | Igoumenitsa | N39 32.495 E20 17.245 | Vac |
| Greece | Western Greece | Menidi # | N39 03.643 E21 06.175 | Vac | ||
| Greece | Peloponnese (west coast) | Giannitsochori | N37 23.729 E21 42.232 | Vac | ||
| Greece | Peloponnese (west coast) | Kyparissia | N37 15.958 E21 41.007 | Vac | ||
| Greece | Peloponnese (south coast) | Kalamata | N37 03.761 E22 08.862 | Vac | ||
| H19 | OQ372231 | Greece | Peloponnese (west coast) | Kyparissia | N37 15.958 E21 41.007 | Vac |
| H20 | OQ372232 | Greece | Peloponnese (west coast) | Kyparissia | N37 15.958 E21 41.007 | Vac |
| H21 * | OQ372233 | Serbia | Southeastern Serbia | Pirot | N43 07.878 E22 26.923 | C. cantabrica |
| H22 | OQ372234 | Greece | Epirus | Igoumenitsa | N39 32.495 E20 17.245 | Vac |
| H23 | OQ372235 | Greece | Epirus | Igoumenitsa | N39 32.495 E20 17.245 | Vac |
| H24 | OQ372236 | Greece | Epirus | Igoumenitsa | N39 32.495 E20 17.245 | Vac |
| H25 | OQ372237 | Greece | Western Greece | Menidi | N39 03.643 E21 06.175 | Vac |
| H26 | OQ372238 | Greece | Western Greece | Menidi | N39 03.643 E21 06.175 | Vac |
| H27 | OQ372239 | Greece | Western Greece | Menidi | N39 03.643 E21 06.175 | Vac |
| H28 | OQ372240 | Greece | Peloponnese (south) | Taygetus, Artemisia | N37 05.818 E22 13.719 | Ud |
| H29 | OQ372241 | Greece | Peloponnese (south) | Taygetus, Artemisia | N37 05.818 E22 13.719 | Ud |
| H30 | OQ372242 | Greece | Peloponnese (south) | Taygetus, Artemisia | N37 05.818 E22 13.719 | Ud |
| Host Plant Associated Group | Mean Genetic Distance (SE) | Mean Genetic Diversity (SE) | Coefficient of Differentiation (SE) | |||||
|---|---|---|---|---|---|---|---|---|
| Overall | Within Group | Between Group | Overall | Between Group | ||||
| Ca–Ud | Vac | Cf | ||||||
| Ca–Ud | 0.014 (0.002) | 0.002 (0.001) | – | (0.003) | (0.004) | 0.014 (0.002) | 0.011 (0.002) | 0.773 (0.035) * |
| Vac | 0.005 (0.001) | 0.019 | – | (0.004) | ||||
| Cf | 0.002 (0.001) | 0.017 | 0.022 | – | ||||
| Host Plant Association | Primer Name | Primer Sequence 5′ → 3′ | Primer Position 1 | HoR-Primed PCR Fragment Size | Multiplex PCR Reaction Conditions |
|---|---|---|---|---|---|
| C. foetida | HoCfF | TTAATAACGACCAAATTTACAAC | 68–90 bp | 1084 bp | 94 °C/60 s, 56 °C/60 s, 72 °C/120 s, 30 cycles; high yield reaction buffer, 2.5 mM MgCl2, 0.5 mM each dNTP, 0.5 μM each primer, 1 U Taq DNA polymerase |
| C. arvensis/ U. dioica | HoF | TTTTACTACTTTCACTACCAGTC | 506–528 bp | 645 bp | |
| V. agnus-castus | HoVacF | ACATATTTACAGTAGGAATAGAC | 797–819 bp | 355 bp | |
| Universal for H. obsoletus | HoR | CATTTTTGRTTTATTGATAGTCCT | 1128–1151 bp | – |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jović, J.; Toševski, I. Multiplex PCR for Discriminating Host Plant Associations of Hyalesthes obsoletus (Hemiptera: Cixiidae), a Key Vector and Driver of ‘Ca. Phytoplasma solani’ Epidemiology. Agronomy 2023, 13, 831. https://doi.org/10.3390/agronomy13030831
Jović J, Toševski I. Multiplex PCR for Discriminating Host Plant Associations of Hyalesthes obsoletus (Hemiptera: Cixiidae), a Key Vector and Driver of ‘Ca. Phytoplasma solani’ Epidemiology. Agronomy. 2023; 13(3):831. https://doi.org/10.3390/agronomy13030831
Chicago/Turabian StyleJović, Jelena, and Ivo Toševski. 2023. "Multiplex PCR for Discriminating Host Plant Associations of Hyalesthes obsoletus (Hemiptera: Cixiidae), a Key Vector and Driver of ‘Ca. Phytoplasma solani’ Epidemiology" Agronomy 13, no. 3: 831. https://doi.org/10.3390/agronomy13030831
APA StyleJović, J., & Toševski, I. (2023). Multiplex PCR for Discriminating Host Plant Associations of Hyalesthes obsoletus (Hemiptera: Cixiidae), a Key Vector and Driver of ‘Ca. Phytoplasma solani’ Epidemiology. Agronomy, 13(3), 831. https://doi.org/10.3390/agronomy13030831







