Mobile Plant Disease Classifier, Trained with a Small Number of Images by the End User
Abstract
:1. Introduction
2. Materials and Methods
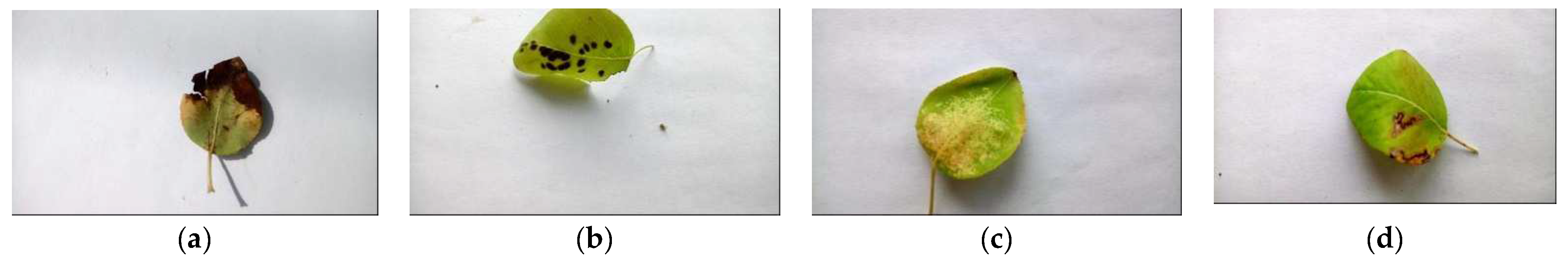
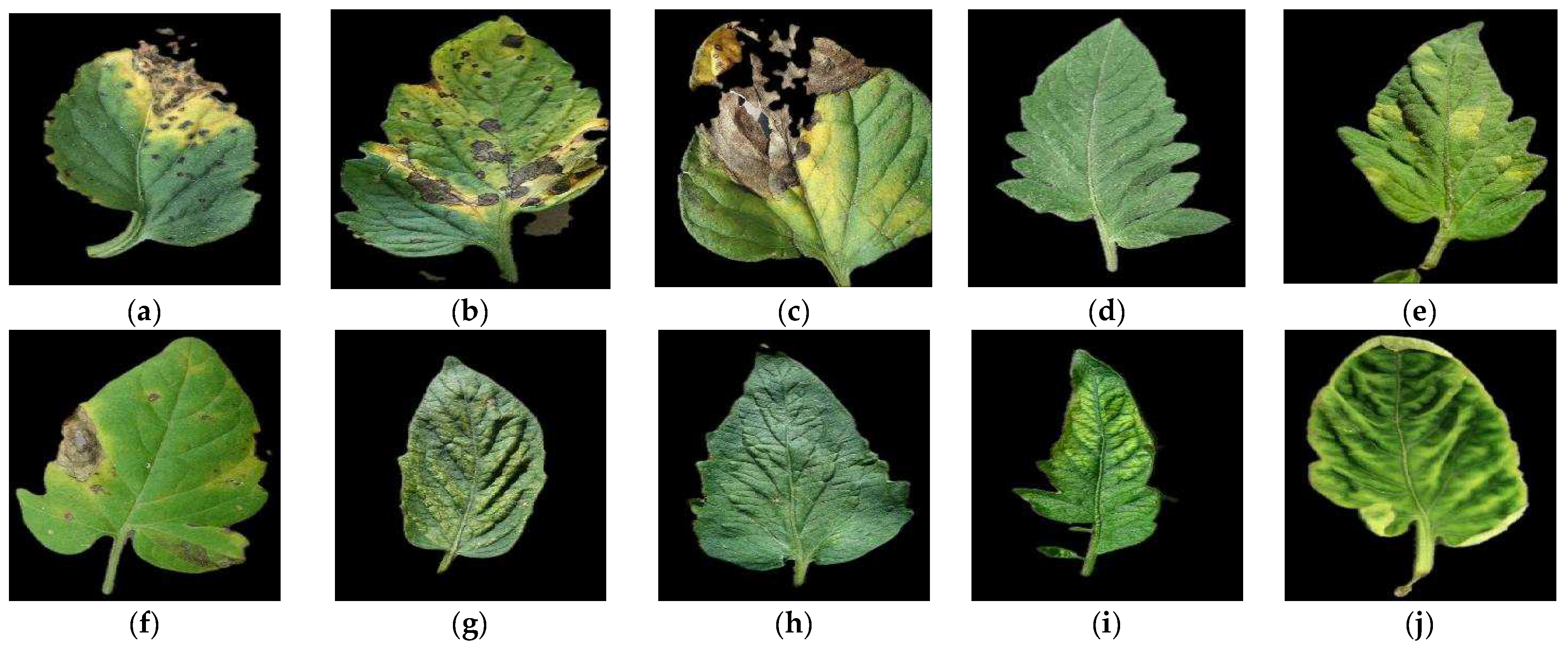
2.1. Dataset and Supported Diseases
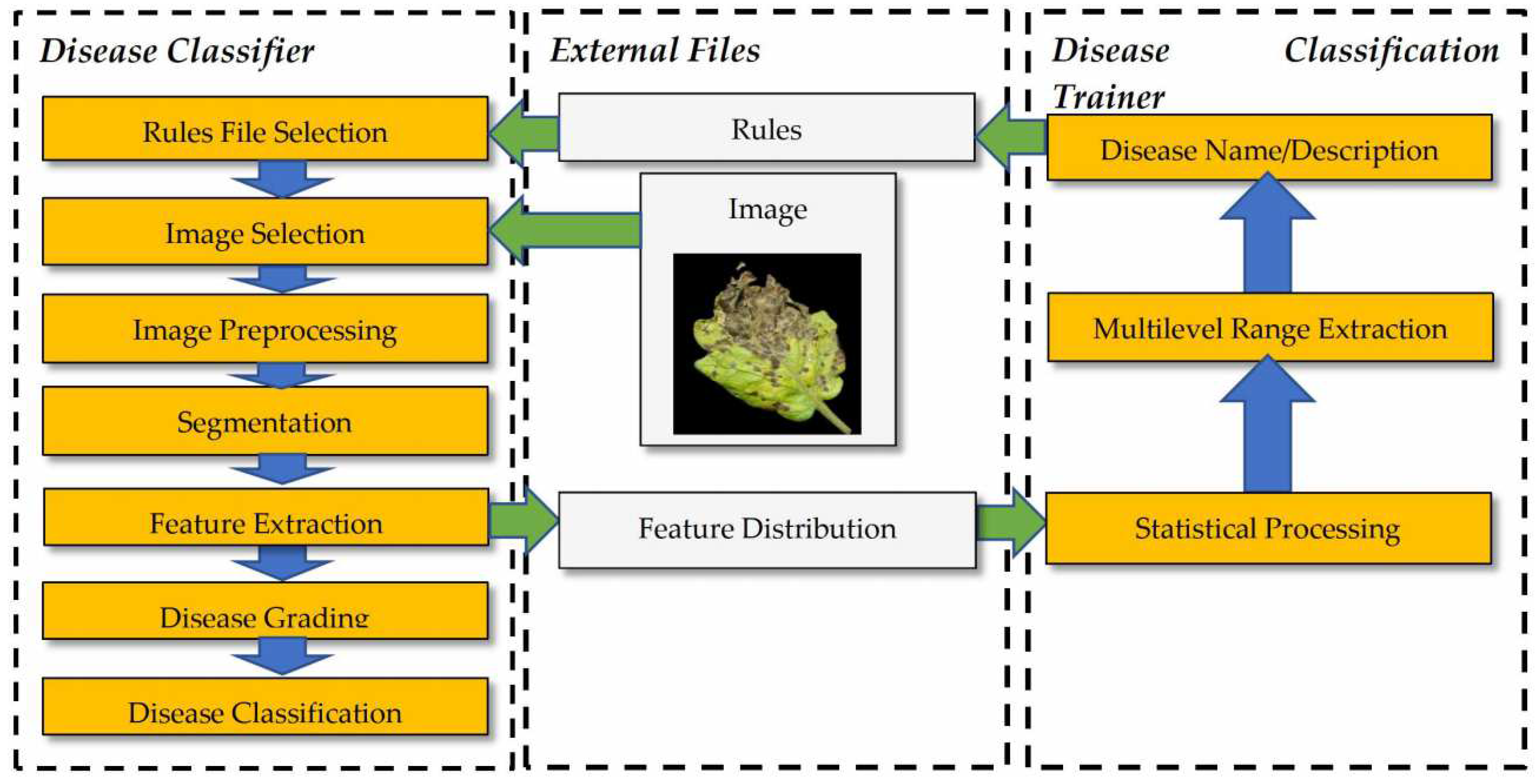
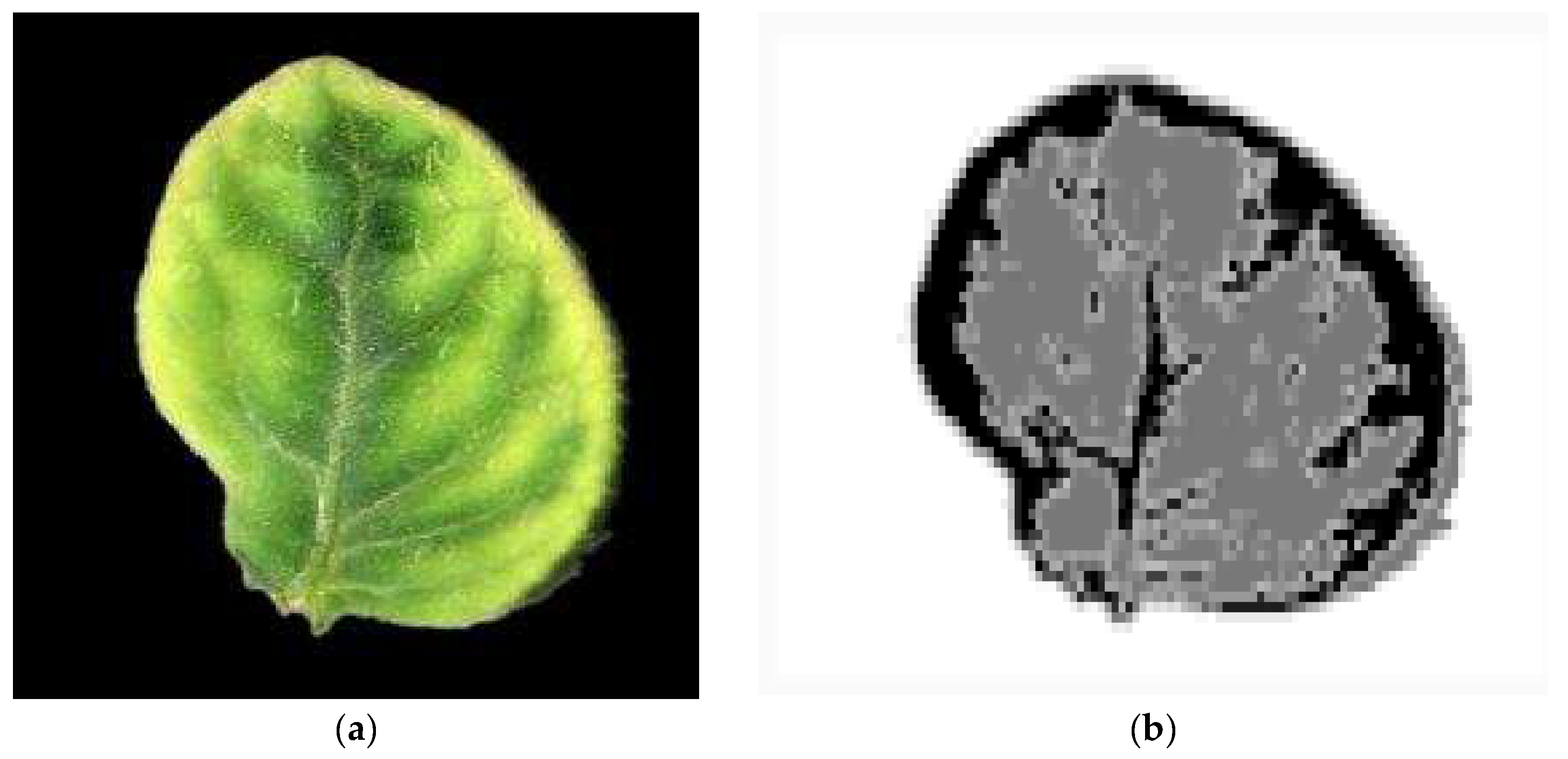
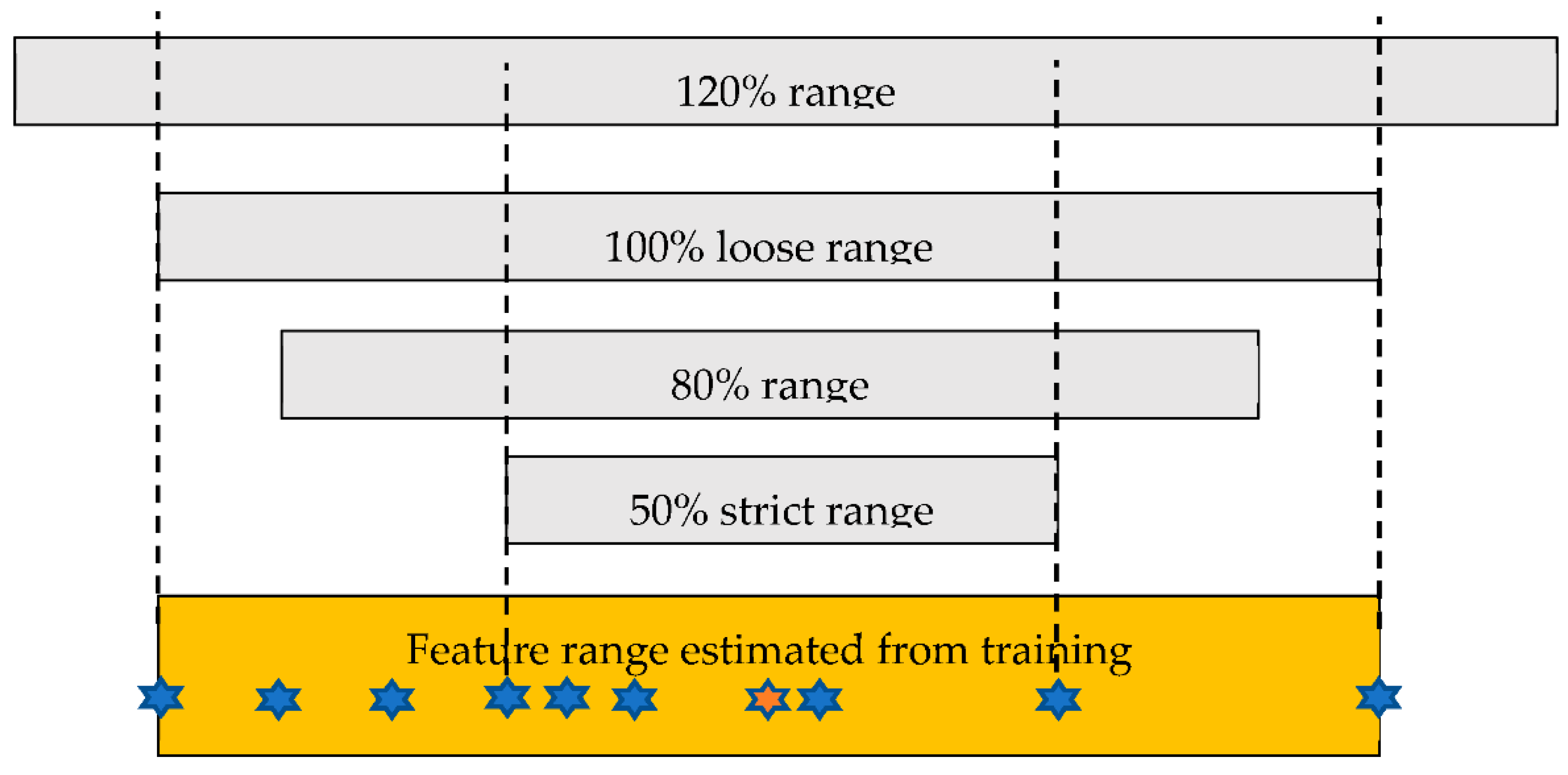
2.2. Image Processing and Classification
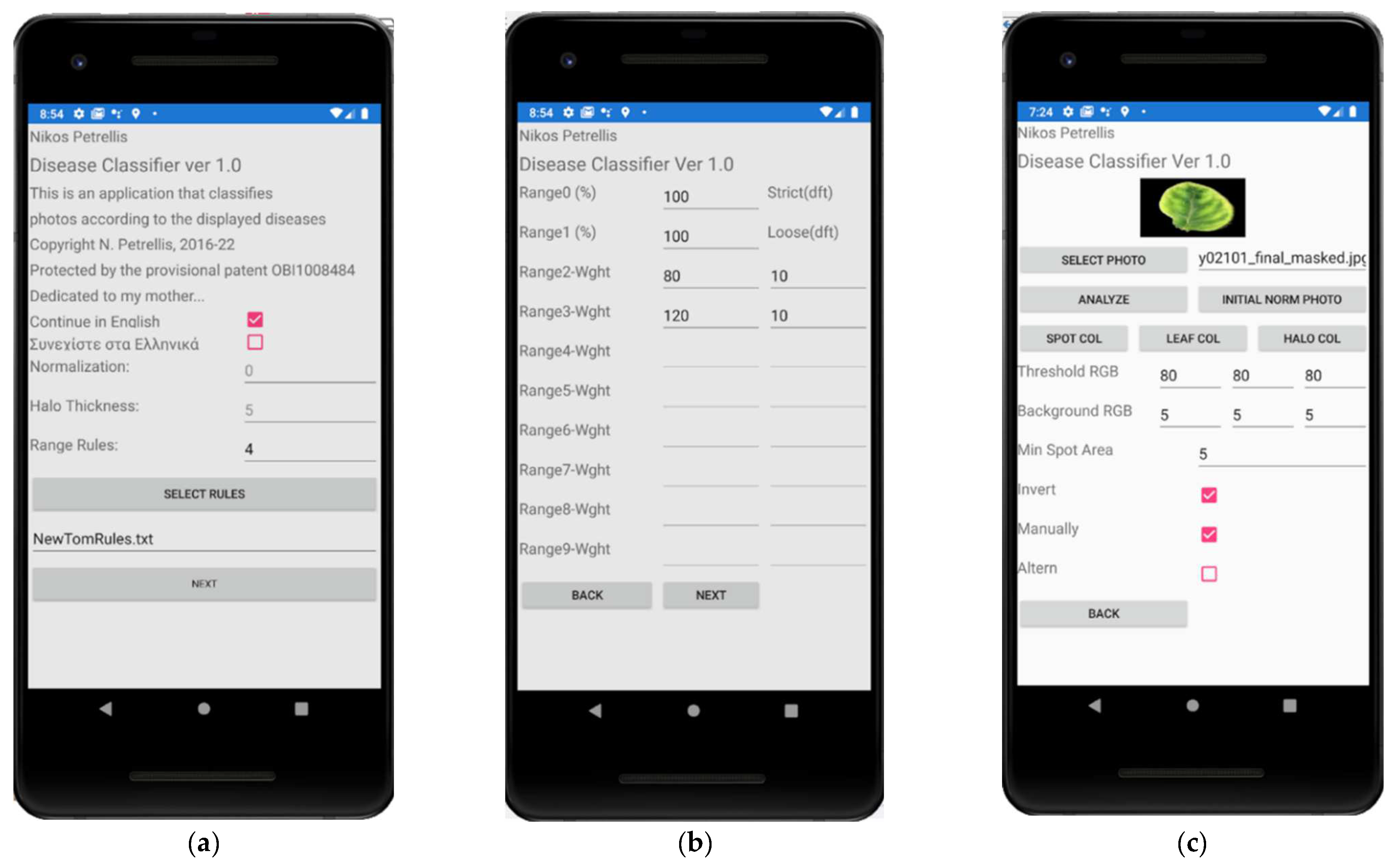
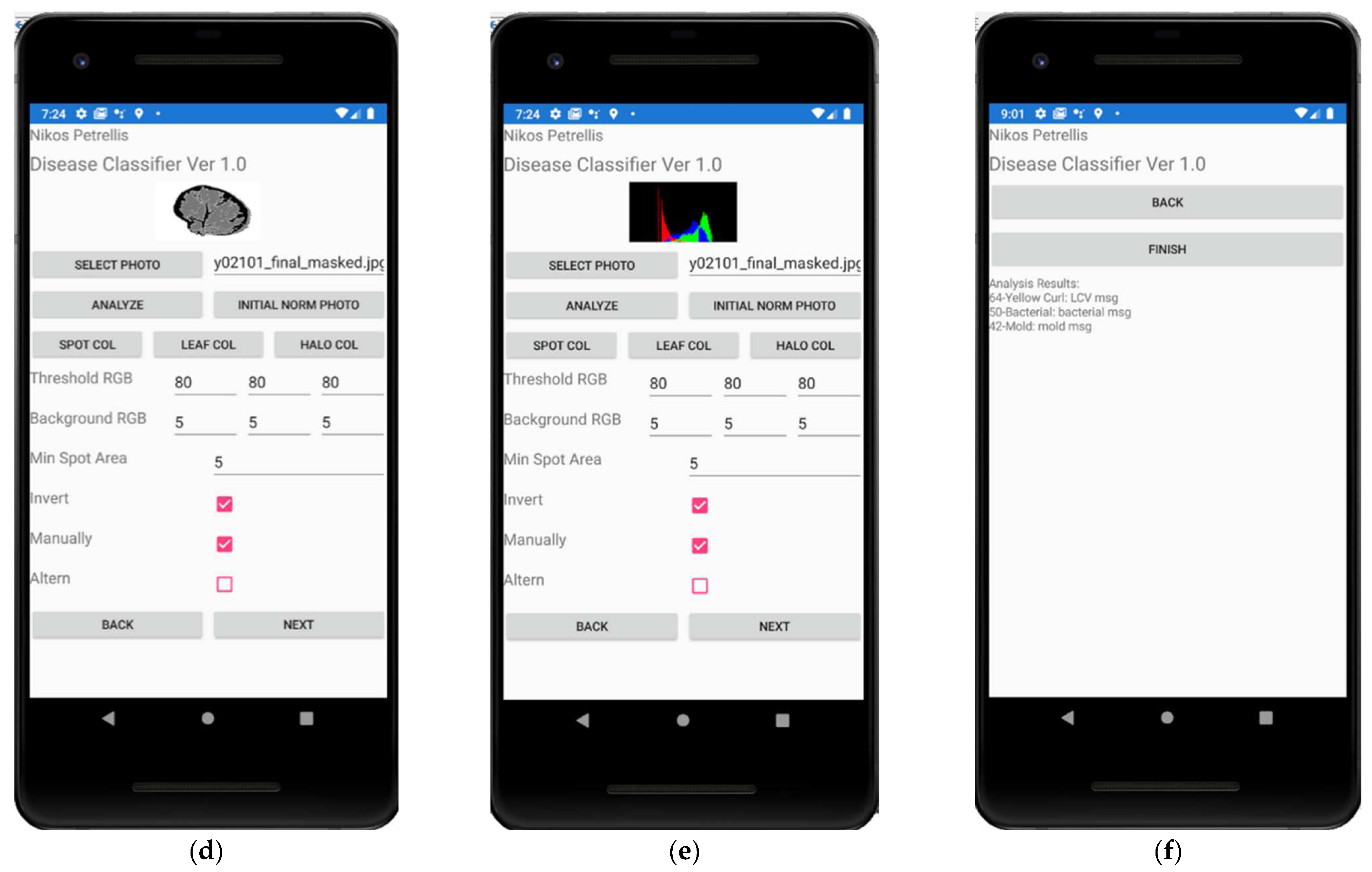
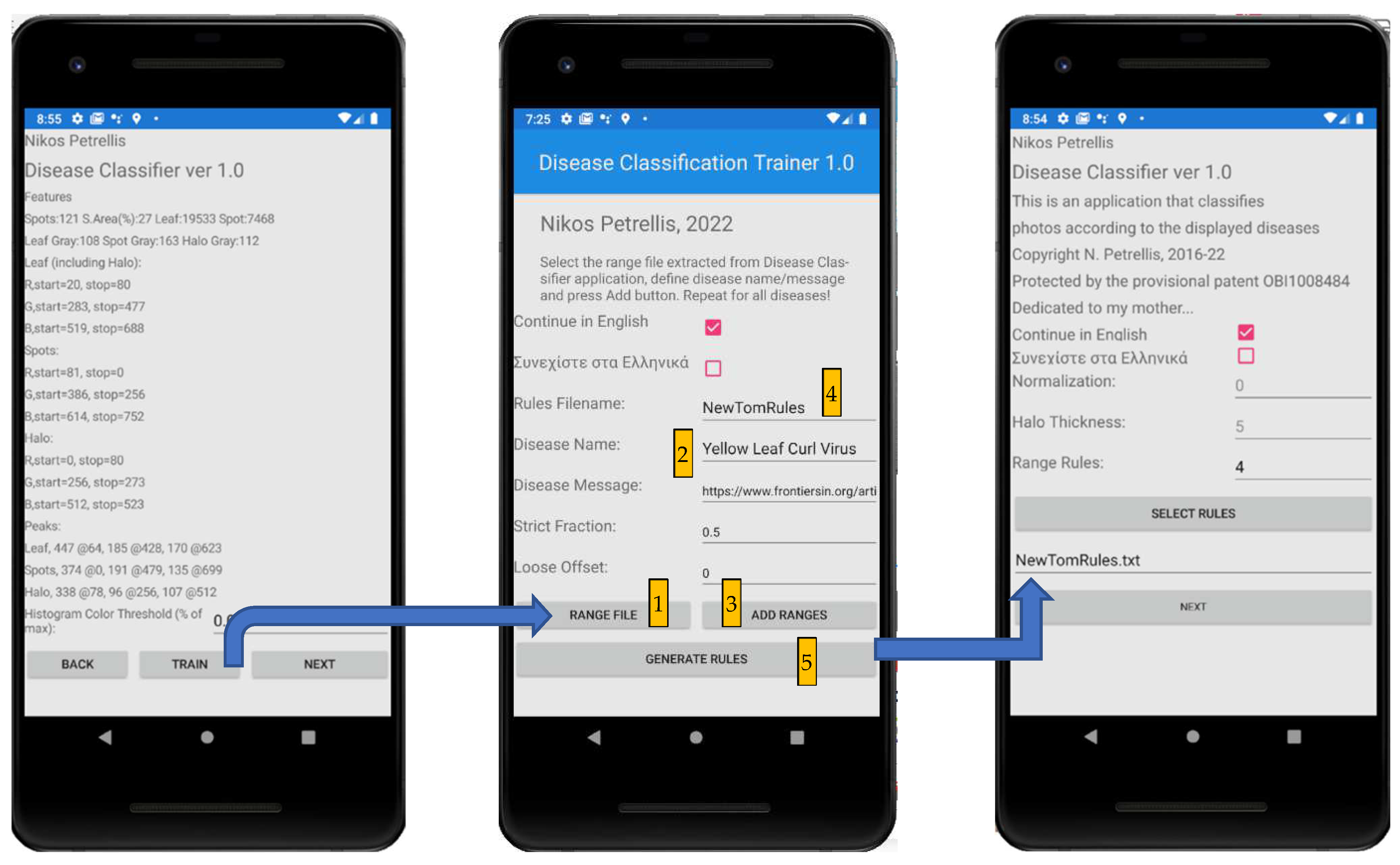
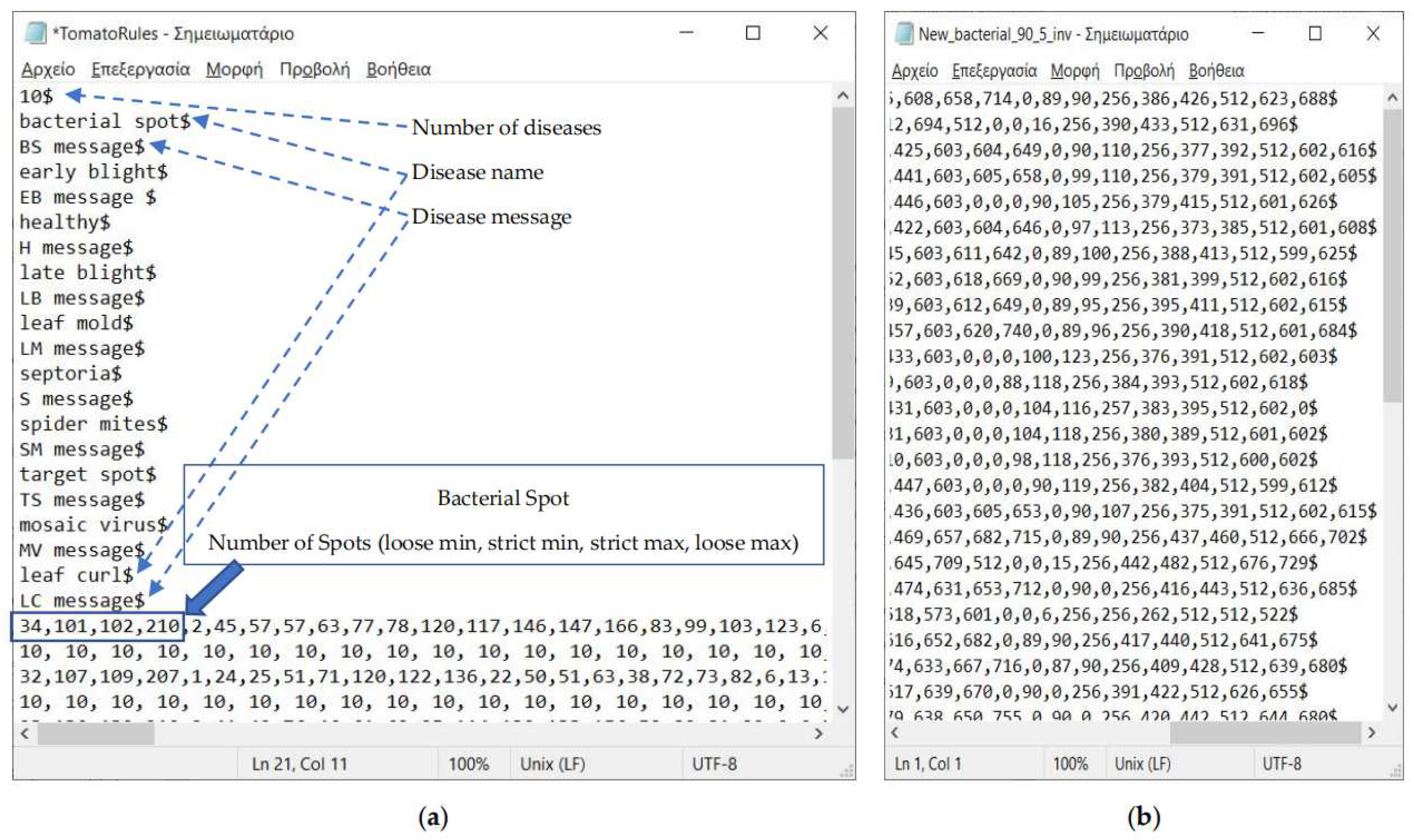
2.3. Description of the Developed Applications
3. Experimental Results
3.1. Use of Small Training Set (T40)
3.2. Use of Large Training Set (T100)
3.3. Testing Pear Diseases with Photographs That Are Not Segmented and Normalized
4. Discussion
5. Conclusions
6. Patents
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sankaran, S.; Mishra, A.; Ehsani, R.; Davis, C. A review of advanced techniques for detecting plant diseases. Comput. Electron. Agric. 2010, 72, 1–13. [Google Scholar] [CrossRef]
- Behmann, J.; Mahlein, A.-K.; Rumpf, T.; Römer, C.; Plümer, L. A review of advanced machine learning methods for the detection of biotic stress in precision crop protection. Precis. Agric. 2014, 16, 239–260. [Google Scholar] [CrossRef]
- Ballesteros, R.; Ortega, J.F.; Hernandez, D.; Moreno, M.A. Applications of georeferenced high-resolution images obtained with unmanned aerial vehicles. Part I: Description of Image Acquisition and Processing. Springer Precis. Agric. 2014, 15, 239–260. [Google Scholar] [CrossRef]
- Xenakis, A.; Papastergiou, G.; Gerogiannis, V.C.; Stamoulis, G. Applying a Convolutional Neural Network in an IoT Robotic System for Plant Disease Diagnosis. In Proceedings of the 11th IEEE International Conference on Information, Intelligence, Systems and Applications (IISA), Piraeus, Greece, 11 December 2020. [Google Scholar] [CrossRef]
- Rodríguez-García, M.; García-Sánchez, F.; Valencia-García, R. Knowledge-Based System for Crop Pests and Diseases Recognition. Electronics 2021, 10, 905. [Google Scholar] [CrossRef]
- Liu, H.; Lee, S.-H.; Chahl, J. A review of recent sensing technologies to detect invertebrates on crops. Precis. Agric. 2016, 18, 635–666. [Google Scholar] [CrossRef]
- Amirruddin, A.D.; Muharam, F.M.; Ismail, M.H.; Tan, N.P.; Ismail, M.F. Hyperspectral spectroscopy and imbalance data approaches for classification of oil palm’s macronutrients observed from frond 9 and 17. Comput. Electron. Agric. 2020, 178, 105768. [Google Scholar] [CrossRef]
- Bhujade, V.G.; Sambhe, V. Role of digital, hyper spectral, and SAR images in detection of plant disease with deep learning network. Multimedia Tools Appl. 2022, 1–26. [Google Scholar] [CrossRef]
- Peng, Y.; Zhao, S.; Liu, J. Fused-Deep-Features Based Grape Leaf Disease Diagnosis. Agronomy 2021, 11, 2234. [Google Scholar] [CrossRef]
- Kodors, S.; Lacis, G.; Zhukov, V.; Bartulsons, T. Pear and apple recognition using deep learning and mobile. Eng. Rural. Dev. 2020, 20, 1795–1800. [Google Scholar]
- Yang, F.; Li, F.; Zhang, K.; Zhang, W.; Li, S. Influencing factors analysis in pear disease recognition using deep learning. Peer-to-Peer Netw. Appl. 2020, 14, 1816–1828. [Google Scholar] [CrossRef]
- Shrestha, G.; Deepsikha; Das, M.; Dey, N. Plant Disease Detection Using CNN. In Proceedings of the IEEE Applied Signal Processing Conference (ASPCON), Kolkata, India, 7 December 2020; pp. 109–113. [Google Scholar] [CrossRef]
- Elhassouny, A.; Smarandache, F. Smart mobile application to recognize tomato leaf diseases using Convolutional Neural Networks. In Proceedings of the 2019 International Conference of Computer Science and Renewable Energies (ICCSRE), Agadir, Morocco, 22–24 July 2019; pp. 10–13. [Google Scholar]
- Agarwal, M.; Singh, A.; Arjaria, S.; Sinha, A.; Gupta, S. ToLeD: Tomato Leaf Disease Detection using Convolution Neural Network. Procedia Comput. Sci. 2020, 167, 293–301. [Google Scholar] [CrossRef]
- Rangarajan, A.K.; Purushothaman, R.; Ramesh, A. Tomato crop disease classification using pre-trained deep learning algorithm. Procedia Comput. Sci. 2018, 133, 1040–1047. [Google Scholar] [CrossRef]
- Bir, P.; Kumar, R.; Singh, G. Transfer Learning based Tomato Leaf Disease Detection for mobile applications. In Proceedings of the IEEE International Conference on Computing, Power and Communication Technologies (GUCON), Greater Noida, India, 2–4 October 2020; pp. 34–39. [Google Scholar] [CrossRef]
- Oteyo, I.N.; Marra, M.; Kimani, S.; De Meuter, W.; Boix, E.G. A Survey on Mobile Applications for Smart Agriculture. SN Comput. Sci. 2021, 2, 293. [Google Scholar] [CrossRef]
- Pérez-Castro, A.; Sánchez-Molina, J.; Castilla, M.; Sánchez-Moreno, J.; Moreno-Úbeda, J.; Magán, J. cFertigUAL: A fertigation management app for greenhouse vegetable crops. Agric. Water Manag. 2017, 183, 186–193. [Google Scholar] [CrossRef]
- Che’Ya, N.N.; Mohidem, N.A.; Roslin, N.A.; Saberioon, M.; Tarmidi, M.Z.; Shah, J.A.; Ilahi, W.F.F.; Man, N. Mobile Computing for Pest and Disease Management Using Spectral Signature Analysis: A Review. Agronomy 2022, 12, 967. [Google Scholar] [CrossRef]
- Johannes, A.; Picon, A.; Alvarez-Gila, A.; Echazarra, J.; Rodriguez-Vaamonde, S.; Navajas, A.D.; Ortiz-Barredo, A. Automatic plant disease diagnosis using mobile capture devices, applied on a wheat use case. Comput. Electron. Agric. 2017, 138, 200–209. [Google Scholar] [CrossRef]
- Adam, A.; Kho, P.E.; Sahari, N.; Tida, A.; Chen, Y.S.; Tawie, K.M.; Kamarudin, S.; Mohamad, H. Dr.LADA: Diagnosing Black Pepper Pests and Diseases with Decision Tree. Int. J. Adv. Sci. Eng. Inf. Technol. 2018, 8, 1584–1590. [Google Scholar] [CrossRef] [Green Version]
- Toseef, M.; Khan, M.J. An intelligent mobile application for diagnosis of crop diseases in Pakistan using fuzzy inference system. Comput. Electron. Agric. 2018, 153, 1–11. [Google Scholar] [CrossRef]
- Moawad, N.; Elsayed, A. Smartphone Application for Diagnosing Maize Diseases in Egypt. In Proceedings of the 14th IEEE International Conference on Innovations in Information Technology (IIT), Al Ain, United Arab Emirates, 25 December 2020; pp. 24–28. [Google Scholar] [CrossRef]
- Moloo, R.K.; Caleechurn, K. An App for Fungal Disease Detection on Plants. In Proceedings of the 2022 International Conference for Advancement in Technology (ICONAT), Goa, India, 21–22 January 2022; pp. 1–5. [Google Scholar] [CrossRef]
- Loyani, L.; Machuve, D. A Deep Learning-based Mobile Application for Segmenting Tuta Absoluta’s Damage on Tomato Plants. Eng. Technol. Appl. Sci. Res. 2021, 11, 7730–7737. [Google Scholar] [CrossRef]
- Shrimali, S. PlantifyAI: A Novel Convolutional Neural Network Based Mobile Application for Efficient Crop Disease Detection and Treatment. Procedia Comput. Sci. 2021, 191, 469–474. [Google Scholar] [CrossRef]
- Petrellis, N. Plant Disease Diagnosis for Smart Phone Applications with Extensible Set of Diseases. Appl. Sci. 2019, 9, 1952. [Google Scholar] [CrossRef] [Green Version]
- Petrellis, N. A Review of Image Processing Techniques Common in Human and Plant Disease Diagnosis. Symmetry 2018, 10, 270. [Google Scholar] [CrossRef] [Green Version]
- PlantVillage Dataset. Available online: https://www.kaggle.com/datasets/abdallahalidev/plantvillage-dataset (accessed on 1 May 2022).
- Parraga-Alava, J.; Cusme, K.; Loor, A.; Santander, E. RoCoLe: A robusta coffee leaf images dataset for evaluation of machine learning based methods in plant diseases recognition. Data Brief 2019, 25, 104414. [Google Scholar] [CrossRef] [PubMed]
- Parraga-Alava, J.; Alcivar-Cevallos, R.; Carrillo, J.M.; Castro, M.; Avellán, S.; Loor, A.; Mendoza, F. LeLePhid: An Image Dataset for Aphid Detection and Infestation Severity on Lemon Leaves. Data 2021, 6, 51. [Google Scholar] [CrossRef]
- Petrellis, N. Plant Disease Diagnosis with Color Normalization. In Proceedings of the 8th International Conference on Modern Circuits and Systems Technologies, MOCAST, Thessaloniki, Greece, 13–15 May 2019. [Google Scholar]


| Feature | Notes |
|---|---|
| Number of spots | A spot is defined as an island of adjacent pixels |
| Area of the spots | (lesion spot pixels)/(overall plant part pixels) |
| Average gray level of the spots | The average gray level extracted from the pixels belonging in each one of the normal, spot and halo ROIs |
| Average gray level of the normal plant part | |
| Average gray level of the halo | |
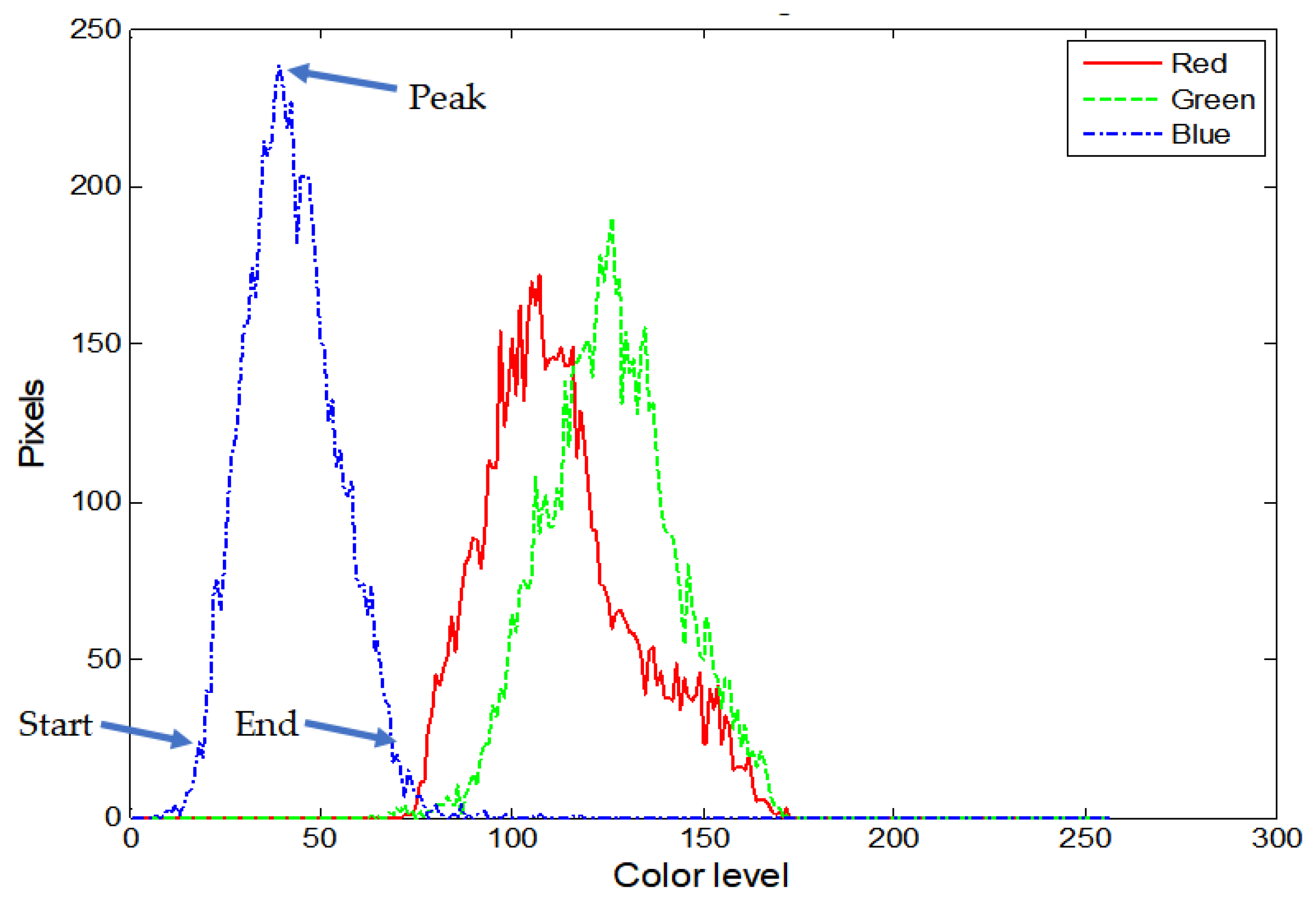
| Histogram lobe start, peak, end | 3 ROIs × 3 colors × 3 parameters = 27 features |
| Average moisture | The average daily moisture, minimum and maximum daily temperatures estimated for a number of dates determined by the user (not tested in this paper) |
| Average minimum temperature | |
| Average maximum temperature |
| Disease | Bacterial Spot | Blight | Healthy | Leaf Mold | Septoria | Spider Mites | Target Spot | Mosaic | Yellow Leaf Curl |
|---|---|---|---|---|---|---|---|---|---|
| Bacterial Spot | 36 | 0 | 0 | 0 | 0 | 4 | 0 | 0 | 0 |
| Blight | 0 | 80 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Healthy | 2 | 0 | 34 | 4 | 0 | 0 | 0 | 0 | 0 |
| Leaf Mold | 6 | 0 | 2 | 30 | 0 | 2 | 0 | 0 | 0 |
| Septoria | 0 | 0 | 0 | 0 | 40 | 0 | 0 | 0 | 0 |
| Spider Mites | 4 | 0 | 2 | 0 | 0 | 34 | 0 | 0 | 0 |
| Target Spot | 0 | 0 | 0 | 0 | 0 | 0 | 40 | 0 | 0 |
| Mosaic | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 40 | 0 |
| Yellow Leaf Curled | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 40 |
| Disease | Sensitivity | Specificity | Precision | Accuracy | F1 Score |
|---|---|---|---|---|---|
| Bacterial Spot | 90.0% | 97.0% | 75.0% | 96.4% | 81.8% |
| Blight | 100.0% | 100.0% | 100.0% | 100.0% | 100.0% |
| Healthy | 85.0% | 99.0% | 89.5% | 97.7% | 87.2% |
| Leaf Mold | 75.0% | 99.0% | 88.2% | 96.8% | 81.1% |
| Septoria | 100.0% | 100.0% | 100.0% | 100.0% | 100.0% |
| Spider Mites | 85.0% | 98.5% | 85.0% | 97.3% | 85.0% |
| Target Spot | 100.0% | 100.0% | 100.0% | 100.0% | 100.0% |
| Mosaic | 100.0% | 100.0% | 100.0% | 100.0% | 100.0% |
| Yellow Leaf Curled | 100.0% | 100.0% | 100.0% | 100.0% | 100.0% |
| Average: | 92.8% | 99.3% | 93.1% | 98.7% | 92.8% |
| Disease | Bacterial Spot | Blight | Healthy | Leaf Mold | Septoria | Spider Mites | Target Spot | Mosaic | Yellow Leaf Curl |
|---|---|---|---|---|---|---|---|---|---|
| Bacterial Spot | 74 | 0 | 2 | 10 | 0 | 0 | 0 | 0 | 14 |
| Blight | 0 | 196 | 0 | 0 | 4 | 0 | 0 | 0 | 0 |
| Healthy | 36 | 0 | 52 | 10 | 0 | 2 | 0 | 0 | 0 |
| Leaf Mold | 18 | 0 | 2 | 52 | 0 | 16 | 0 | 0 | 12 |
| Septoria | 0 | 28 | 0 | 0 | 72 | 0 | 0 | 0 | 0 |
| Spider Mites | 10 | 0 | 14 | 8 | 0 | 68 | 0 | 0 | 0 |
| Target Spot | 6 | 0 | 0 | 6 | 0 | 4 | 70 | 12 | 2 |
| Mosaic | 2 | 0 | 0 | 2 | 0 | 0 | 10 | 82 | 4 |
| Yellow Leaf Curled | 0 | 0 | 6 | 4 | 0 | 0 | 0 | 0 | 90 |
| Disease | Sensitivity | Specificity | Precision | Accuracy | F1 Score |
|---|---|---|---|---|---|
| Bacterial Spot | 74.0% | 92.0% | 50.7% | 90.2% | 60.2% |
| Blight | 98.0% | 96.9% | 87.5% | 97.1% | 92.5% |
| Healthy | 52.0% | 97.3% | 68.4% | 92.8% | 59.1% |
| Leaf Mold | 52.0% | 95.6% | 56.5% | 91.2% | 54.2% |
| Septoria | 72.0% | 99.6% | 94.7% | 96.8% | 81.8% |
| Spider Mites | 68.0% | 97.6% | 75.6% | 94.6% | 71.6% |
| Target Spot | 70.0% | 98.9% | 87.5% | 96.0% | 77.8% |
| Mosaic | 82.0% | 98.7% | 87.2% | 97.0% | 84.5% |
| Yellow Leaf Curled | 90.0% | 96.4% | 73.8% | 95.8% | 81.1% |
| Average: | 73.1% | 97.0% | 75.8% | 94.6% | 73.6% |
| Disease | Bacterial Spot | Blight | Healthy | Leaf Mold | Septoria | Spider Mites | Target Spot | Mosaic | Yellow Leaf Curl |
|---|---|---|---|---|---|---|---|---|---|
| Bacterial Spot | 62 | 0 | 2 | 10 | 0 | 2 | 0 | 0 | 24 |
| Blight | 0 | 196 | 0 | 0 | 4 | 0 | 0 | 0 | 0 |
| Healthy | 42 | 0 | 48 | 10 | 0 | 0 | 0 | 0 | 0 |
| Leaf Mold | 18 | 0 | 2 | 56 | 0 | 14 | 0 | 0 | 12 |
| Septoria | 0 | 42 | 0 | 0 | 58 | 0 | 0 | 0 | 0 |
| Spider Mites | 4 | 0 | 12 | 18 | 0 | 64 | 2 | 0 | 0 |
| Target Spot | 6 | 0 | 2 | 2 | 0 | 8 | 70 | 8 | 4 |
| Mosaic | 2 | 0 | 0 | 2 | 0 | 0 | 10 | 74 | 12 |
| Yellow Leaf Curled | 0 | 0 | 8 | 14 | 0 | 2 | 0 | 0 | 76 |
| Disease | Sensitivity | Specificity | Precision | Accuracy | F1 Score |
|---|---|---|---|---|---|
| Bacterial Spot | 62.0% | 92.0% | 46.3% | 89.0% | 53.0% |
| Blight | 98.0% | 95.3% | 82.4% | 95.8% | 89.5% |
| Healthy | 48.0% | 97.1% | 64.9% | 92.2% | 55.2% |
| Leaf Mold | 56.0% | 93.8% | 50.0% | 90.0% | 52.8% |
| Septoria | 58.0% | 99.6% | 93.5% | 95.4% | 71.6% |
| Spider Mites | 64.0% | 97.1% | 71.1% | 93.8% | 67.4% |
| Target Spot | 70.0% | 98.7% | 85.4% | 95.8% | 76.9% |
| Mosaic | 74.0% | 99.1% | 90.2% | 96.6% | 81.3% |
| Yellow Leaf Curled | 76.0% | 94.2% | 59.4% | 92.4% | 66.7% |
| Average: | 67.3% | 96.3% | 71.5% | 93.4% | 68.3% |
| Disease | Bacterial Spot | Blight | Healthy | Leaf Mold | Septoria | Spider Mites | Target Spot | Mosaic | Yellow Leaf Curl |
|---|---|---|---|---|---|---|---|---|---|
| Bacterial Spot | 70 | 0 | 2 | 14 | 0 | 0 | 0 | 0 | 14 |
| Blight | 0 | 200 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Healthy | 36 | 0 | 46 | 12 | 0 | 4 | 0 | 0 | 2 |
| Leaf Mold | 20 | 2 | 4 | 48 | 0 | 8 | 0 | 0 | 18 |
| Septoria | 0 | 32 | 0 | 0 | 68 | 0 | 0 | 0 | 0 |
| Spider Mites | 4 | 0 | 10 | 14 | 0 | 70 | 0 | 0 | 2 |
| Target Spot | 2 | 0 | 0 | 6 | 0 | 4 | 70 | 10 | 8 |
| Mosaic | 2 | 0 | 0 | 4 | 0 | 0 | 10 | 72 | 12 |
| Yellow Leaf Curled | 0 | 0 | 4 | 8 | 0 | 0 | 0 | 0 | 88 |
| Disease | Sensitivity | Specificity | Precision | Accuracy | F1 Score |
|---|---|---|---|---|---|
| Bacterial Spot | 70.0% | 92.9% | 52.2% | 90.6% | 59.8% |
| Blight | 100.0% | 96.2% | 85.5% | 96.9% | 92.2% |
| Healthy | 46.0% | 97.8% | 69.7% | 92.6% | 55.4% |
| Leaf Mold | 48.0% | 93.6% | 45.3% | 89.0% | 46.6% |
| Septoria | 68.0% | 100.0% | 100.0% | 96.8% | 81.0% |
| Spider Mites | 70.0% | 98.2% | 81.4% | 95.4% | 75.3% |
| Target Spot | 70.0% | 98.9% | 87.5% | 96.0% | 77.8% |
| Mosaic | 72.0% | 98.9% | 87.8% | 96.2% | 79.1% |
| Yellow Leaf curled | 88.0% | 93.8% | 61.1% | 93.2% | 72.1% |
| Average: | 70.2% | 96.7% | 74.5% | 94.1% | 71.0% |
| Disease | Bacterial Spot | Blight | Healthy | Leaf Mold | Septoria | Spider Mites | Target Spot | Mosaic | Yellow Leaf Curl |
|---|---|---|---|---|---|---|---|---|---|
| Bacterial Spot | 30 | 0 | 0 | 4 | 0 | 0 | 0 | 0 | 6 |
| Blight | 0 | 80 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Healthy | 6 | 0 | 30 | 2 | 0 | 0 | 0 | 0 | 2 |
| Leaf Mold | 16 | 0 | 0 | 24 | 0 | 0 | 0 | 0 | 0 |
| Septoria | 0 | 14 | 0 | 0 | 26 | 0 | 0 | 0 | 0 |
| Spider Mites | 4 | 0 | 8 | 6 | 0 | 22 | 0 | 0 | 0 |
| Target Spot | 0 | 0 | 0 | 6 | 0 | 2 | 30 | 2 | 0 |
| Mosaic | 0 | 0 | 0 | 0 | 0 | 0 | 8 | 32 | 0 |
| Yellow Leaf Curled | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 40 |
| Disease | Sensitivity | Specificity | Precision | Accuracy | F1 Score |
|---|---|---|---|---|---|
| Bacterial Spot | 75.0% | 92.8% | 53.6% | 91.0% | 62.5% |
| Blight | 100.0% | 95.6% | 85.1% | 96.5% | 92.0% |
| Healthy | 75.0% | 97.8% | 78.9% | 95.5% | 76.9% |
| Leaf Mold | 60.0% | 95.0% | 57.1% | 91.5% | 58.5% |
| Septoria | 65.0% | 100.0% | 100.0% | 96.5% | 78.8% |
| Spider Mites | 55.0% | 99.4% | 91.7% | 95.0% | 68.8% |
| Target Spot | 75.0% | 97.8% | 78.9% | 95.5% | 76.9% |
| Mosaic | 80.0% | 99.4% | 94.1% | 97.5% | 86.5% |
| Yellow Leaf Curled | 100.0% | 97.8% | 83.3% | 98.0% | 90.9% |
| Average: | 76.1% | 97.3% | 80.3% | 95.2% | 76.9% |
| Disease | Fire Blight | Fusicladium | Podosphaera | Septoria |
|---|---|---|---|---|
| Fire blight | 30 | 0 | 0 | 4 |
| Fusicladium | 0 | 80 | 0 | 0 |
| Podosphaera | 6 | 0 | 30 | 2 |
| Septoria | 16 | 0 | 0 | 24 |
| Disease | Sensitivity | Specificity | Precision | Accuracy | F1 Score |
|---|---|---|---|---|---|
| Blight | 81.7% | 96.7% | 90.7% | 92.4% | 86.0% |
| Fusicladium | 82.5% | 95.4% | 82.5% | 92.7% | 82.5% |
| Podosphaera | 100.0% | 100.0% | 100.0% | 100.0% | 100.0% |
| Septoria | 90.0% | 95.4% | 56.3% | 95.0% | 69.2% |
| Average: | 88.5% | 96.9% | 82.4% | 95.0% | 84.4% |
| Disease | R | G | B | Invert |
|---|---|---|---|---|
| Tomato Bacterial Spot | 90 | 90 | 90 | Yes |
| Tomato Blight | 80 | 80 | 80 | No |
| Tomato Healthy | 90 | 90 | 90 | Yes |
| Tomato Leaf Mold | 90 | 90 | 90 | Yes |
| Tomato Septoria | 100 | 100 | 160 | No |
| Tomato Spider Mites | 90 | 90 | 90 | Yes |
| Tomato Target Spot | 100 | 100 | 100 | Yes |
| Tomato Mosaic | 100 | 100 | 100 | Yes |
| Tomato Yellow Leaf Curled | 80 | 80 | 80 | Yes |
| Pear Fire Blight | 40 | 60 | 60 | No |
| Pear Fusicladium | 20 | 60 | 80 | No |
| Pear Podosphaera | 40 | 60 | 60 | Yes |
| Pear Septoria | 40 | 60 | 60 | No |
| Reference | Accuracy | Notes: Implementation Type, Supported Diseases, Dataset |
|---|---|---|
| [4] | 92.39% (viral), 95.33% (late blight), 98.56% (bacterial) | CNN, 3 disease states, training:test set ratio = 800:200 |
| [9] | 99.55% | ResNet50/101 + SVM classifier, 4 grape diseases (black rot, esca measles, leaf spot, healthy), 423–1383 images/disease |
| [10] | 99.8% | 120 classes, pear and apple diseases testing MobileNet and MobileNetV2 on mobile devices, ~10,000 images training:test ratio = 75%:25% |
| [11] | 99.44%, 98.43% and 97.67% | Septoria, Alternaria and Gymnosporangium pear disease with 6 severity levels 4944 pear images. Architecture: VGG16, Inception V3, ResNet50 and ResNet101 |
| [12] | 88.8% | CNN, 12 diseases on tomato/potato leafs, training:test ratio = 160:40 |
| [13] | 89.2% | MobileNet, 10 tomato leaf diseases 7176 images |
| [14] | 76–100% | CNN, 10 tomato leaf diseases Training:validation:test ratio= 10,000:7000:500 |
| [15] | 97.49% | AlexNet, 7 tomato diseases 13,262 segmented images |
| [16] | Weighted average precision: 96.61% (segmented), 98.61 (unsegmented) | EfficientNet B0, 10 tomato leaf disease 15,000 segmented/unsegmented images |
| [24] | 68.3% | CNN, 5 diseases, training:test ratio= 3000:50 |
| [26] | 95.7% | MobileNetV2 + Canny Edge Detection, 26 crop diseases of 14 different species, 87,860 leaf images |
| This work | 94.4%95% | 10 tomato diseases, (1400 images) training:test ratio = 400:1000 4 pear diseases (382 images) training:test ratio = 80:302 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Petrellis, N.; Antonopoulos, C.; Keramidas, G.; Voros, N. Mobile Plant Disease Classifier, Trained with a Small Number of Images by the End User. Agronomy 2022, 12, 1732. https://doi.org/10.3390/agronomy12081732
Petrellis N, Antonopoulos C, Keramidas G, Voros N. Mobile Plant Disease Classifier, Trained with a Small Number of Images by the End User. Agronomy. 2022; 12(8):1732. https://doi.org/10.3390/agronomy12081732
Chicago/Turabian StylePetrellis, Nikos, Christos Antonopoulos, Georgios Keramidas, and Nikolaos Voros. 2022. "Mobile Plant Disease Classifier, Trained with a Small Number of Images by the End User" Agronomy 12, no. 8: 1732. https://doi.org/10.3390/agronomy12081732
APA StylePetrellis, N., Antonopoulos, C., Keramidas, G., & Voros, N. (2022). Mobile Plant Disease Classifier, Trained with a Small Number of Images by the End User. Agronomy, 12(8), 1732. https://doi.org/10.3390/agronomy12081732









