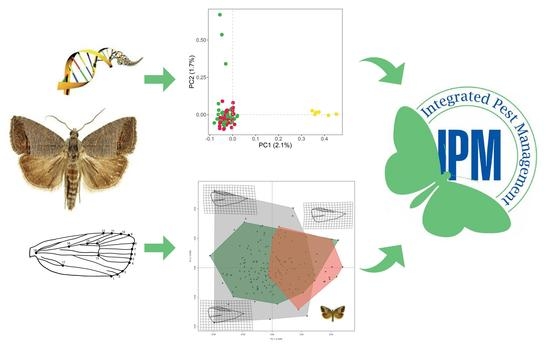
Population Genetic Structure and Geometric Morphology of Codling Moth Populations from Different Management Systems
Abstract
1. Introduction
2. Materials and Methods
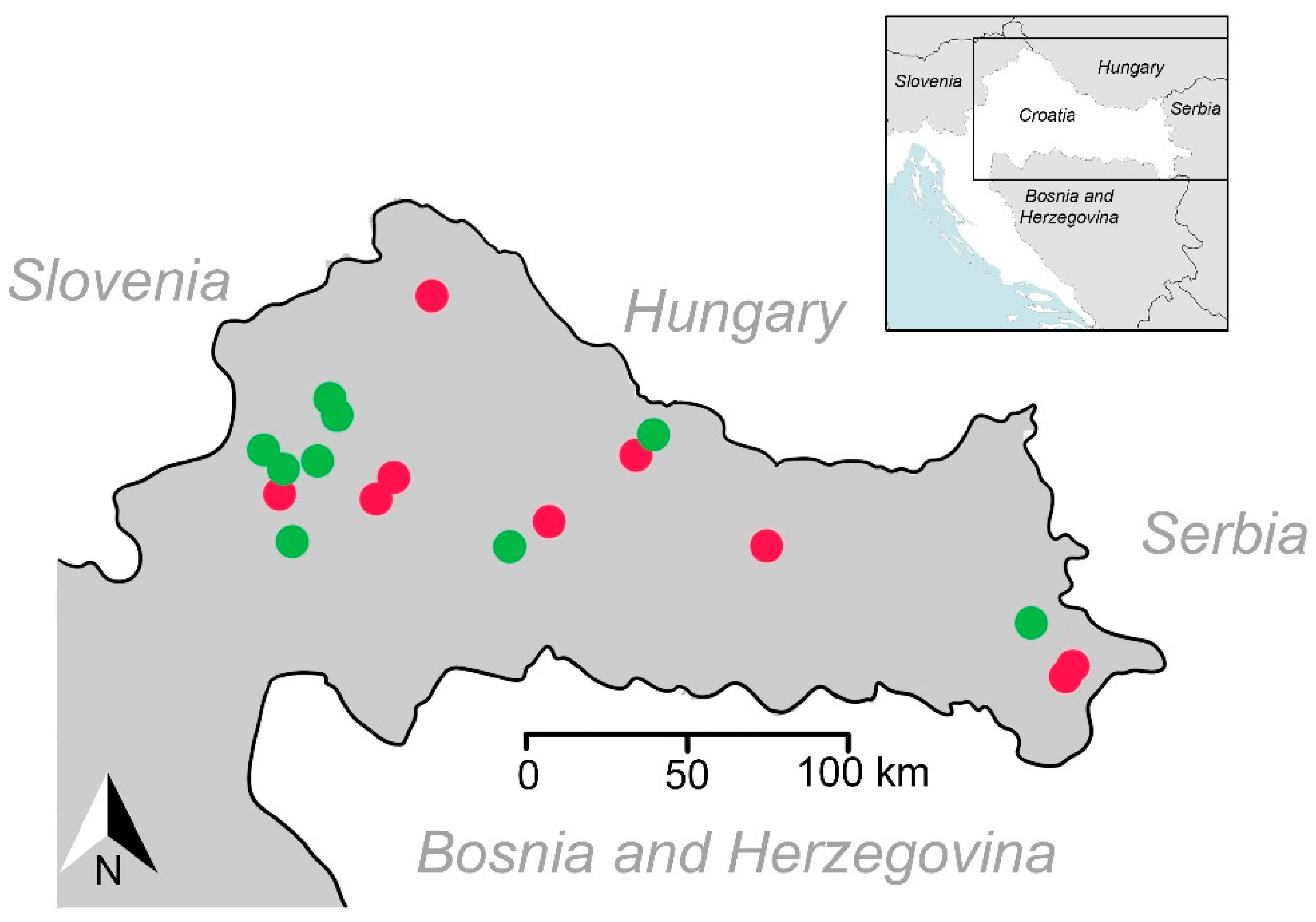
2.1. Collection Sites and Sampling
2.2. DNA Extraction and SNPs Genotyping
2.3. Geometric Morphometric Sample Preparation
2.4. Data Analysis
2.4.1. SNP Quality Control
2.4.2. Population Genetics Analyses
2.4.3. Geometric Morphometrics
3. Results
3.1. Genetic Data
3.1.1. Population Diversity Metrics
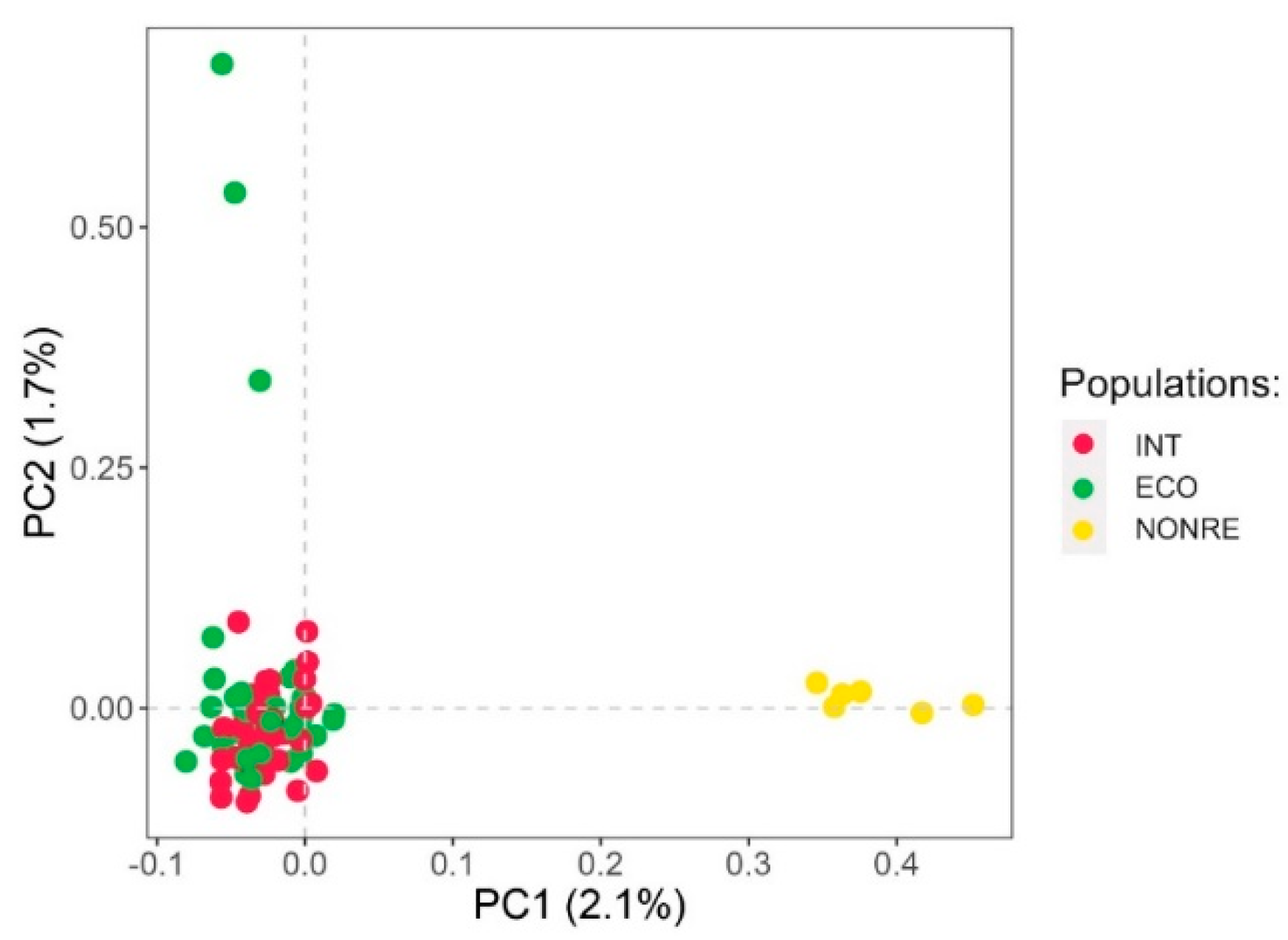
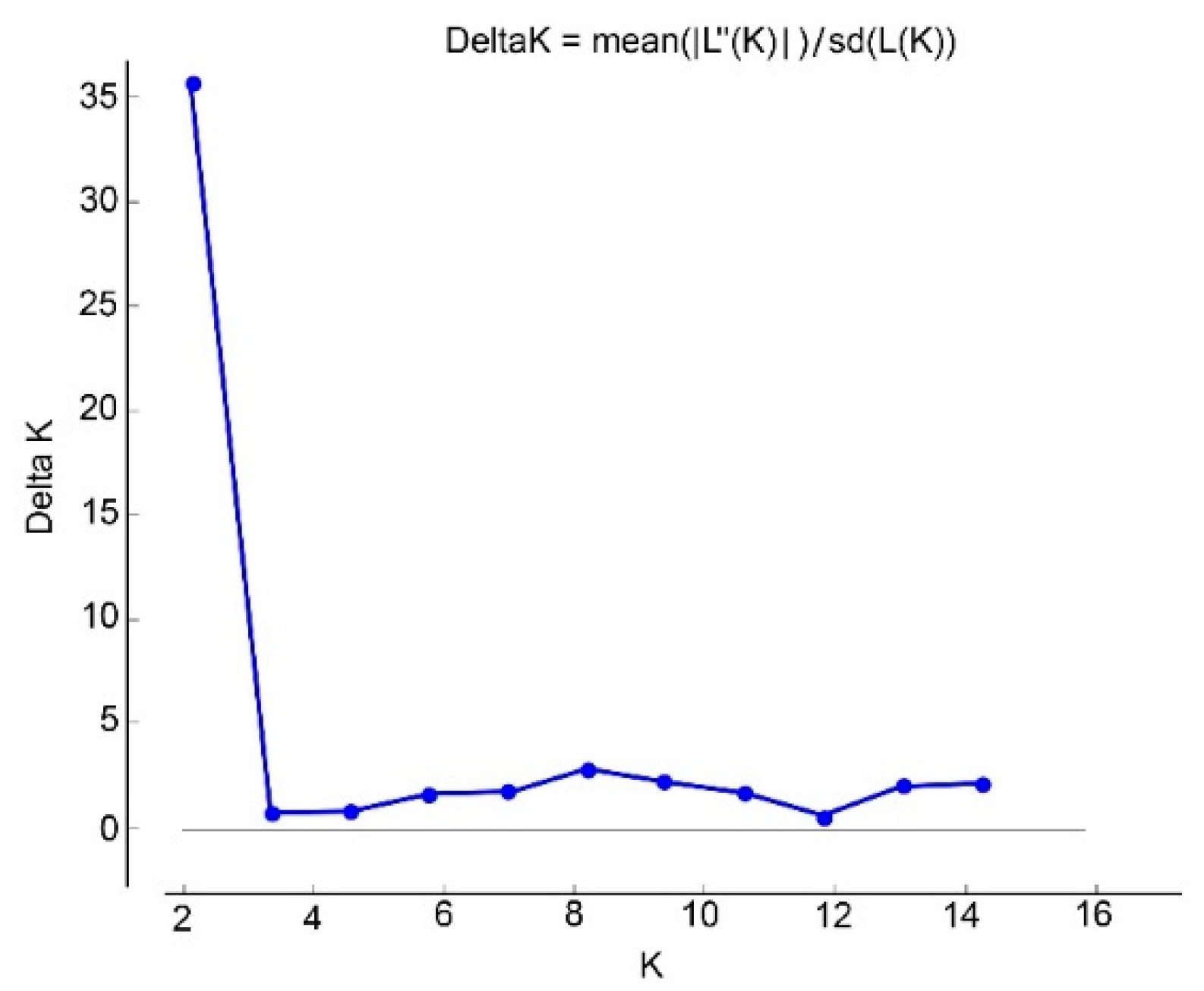
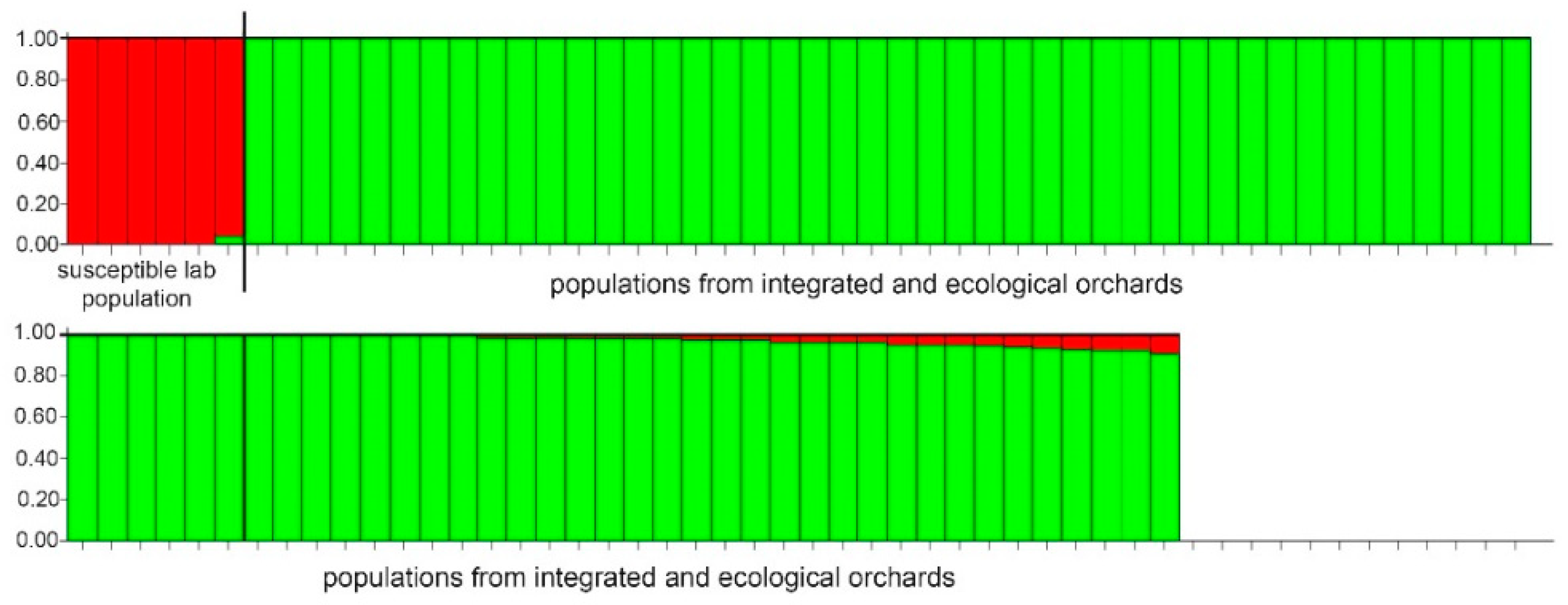
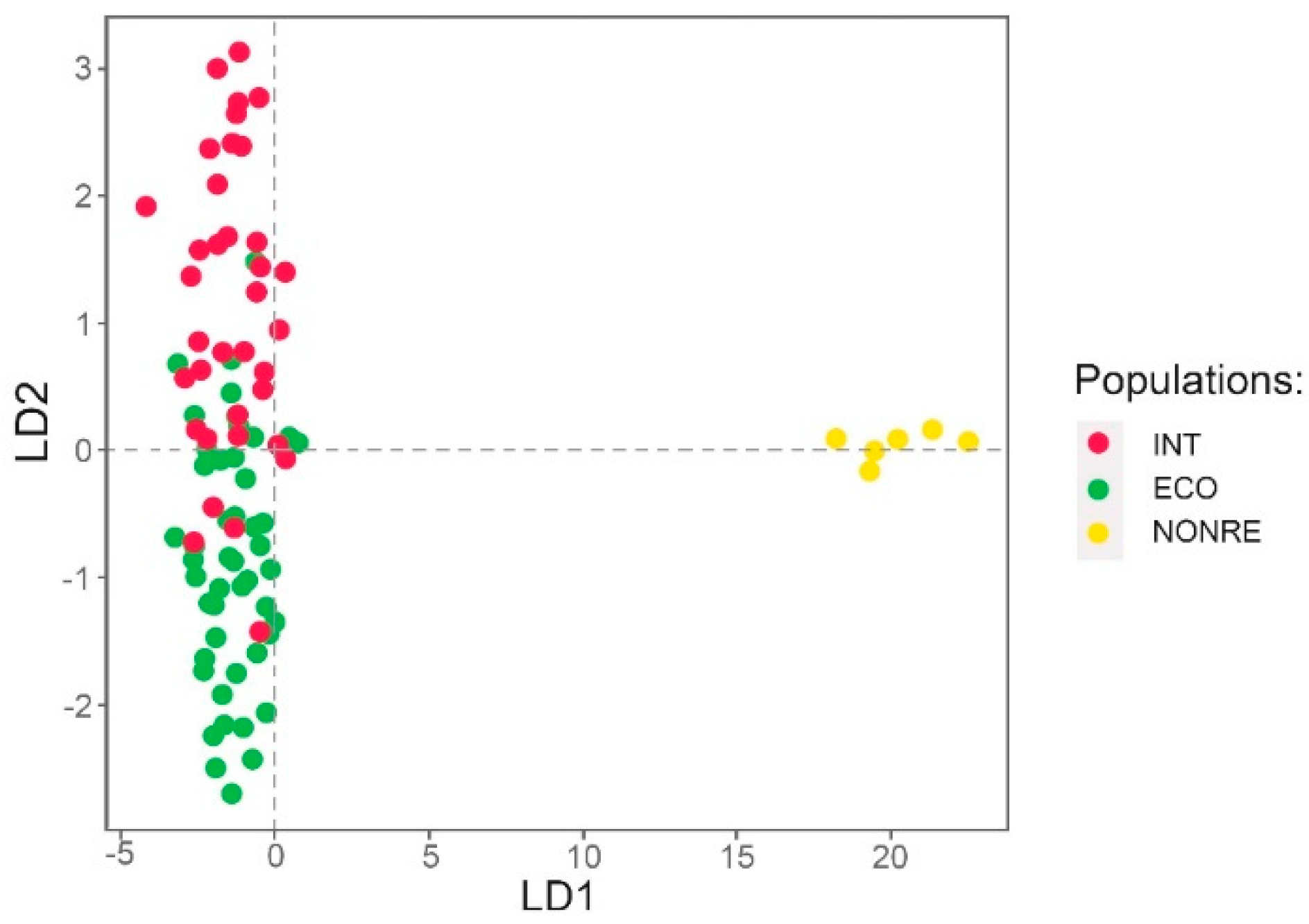
3.1.2. Genetic Structure
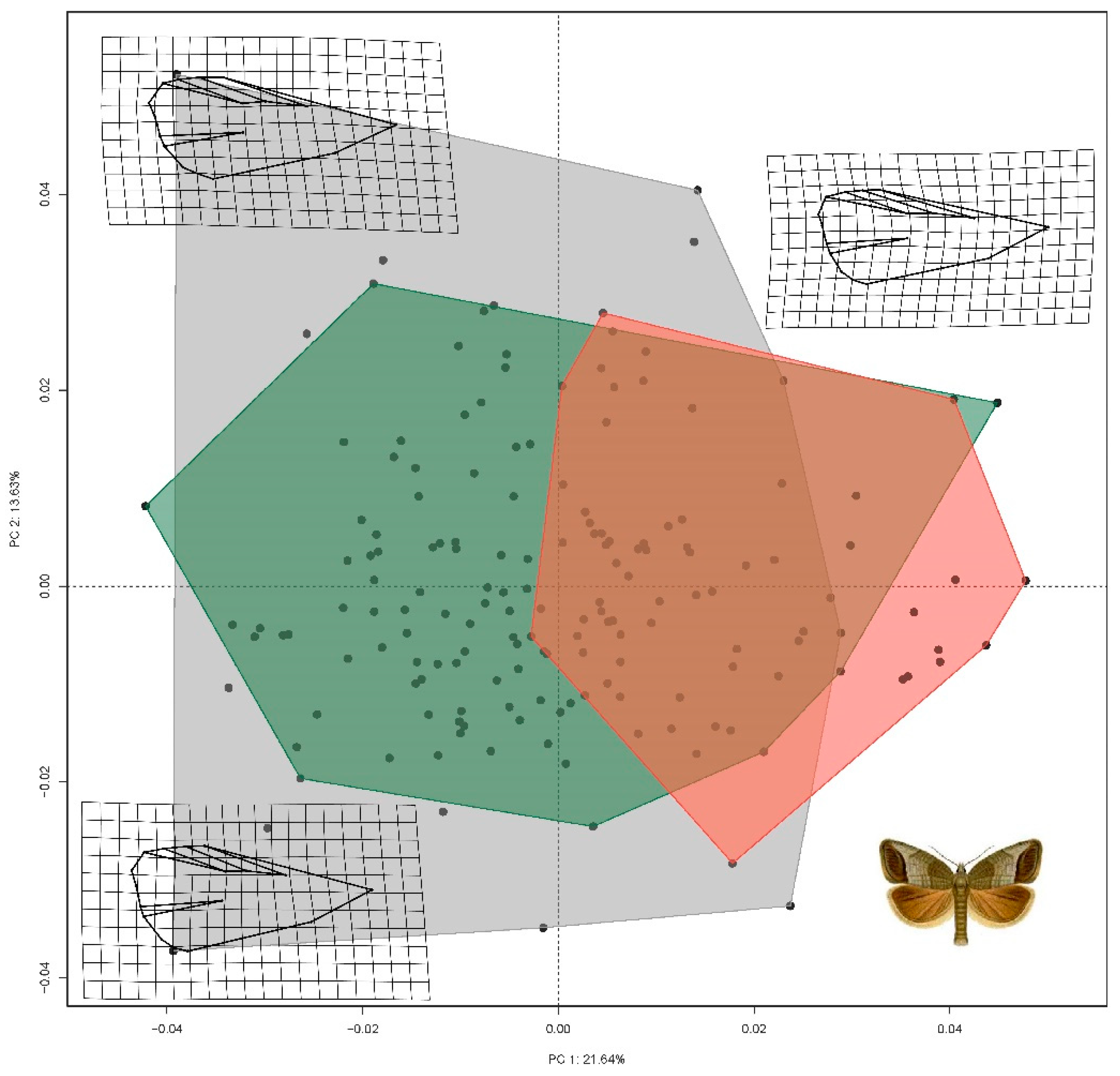
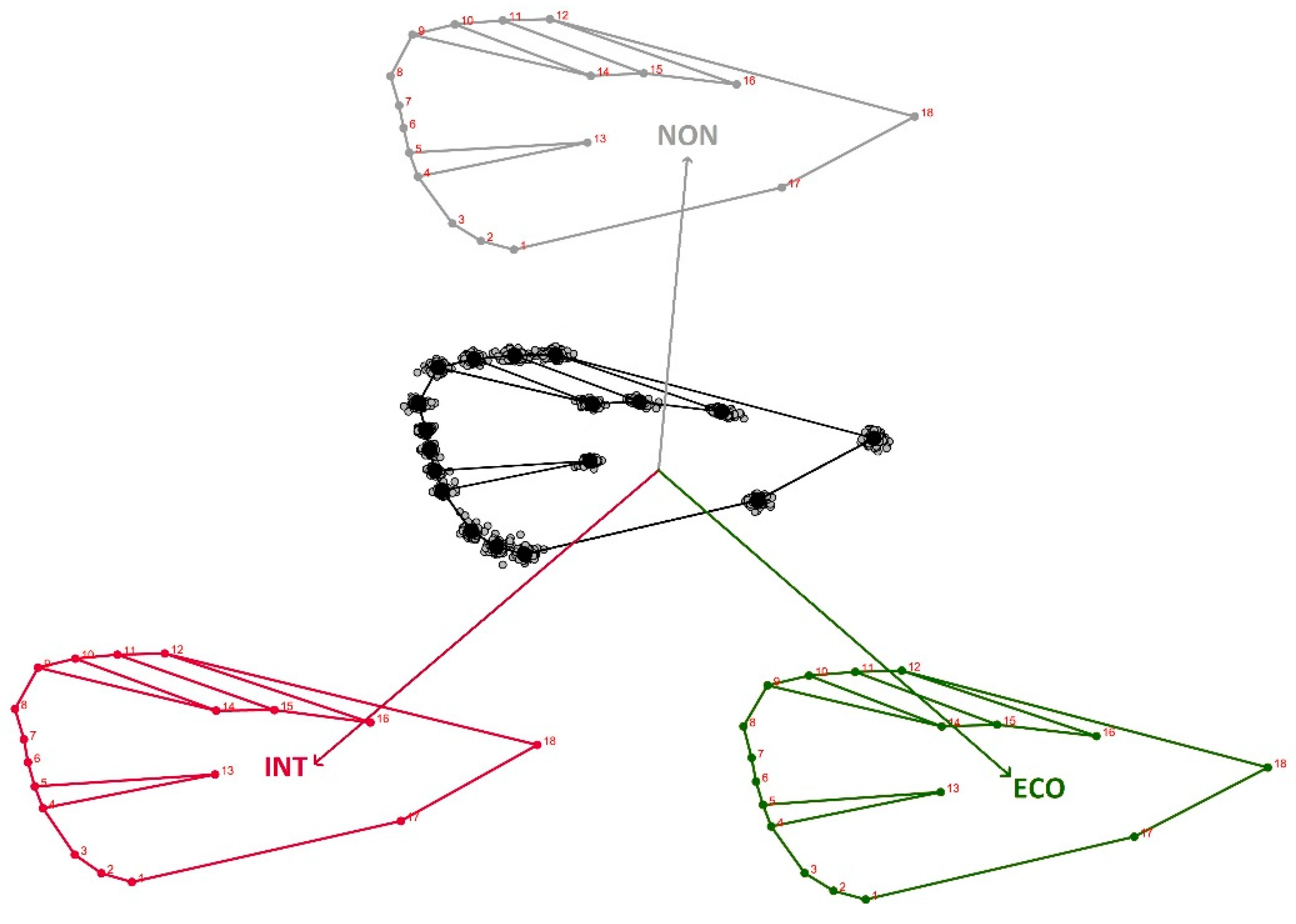
3.2. Geometric Morphometrics
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ciglar, I. Integrirana Zaštita Voćaka i Vinove Loze, 1st ed.; Zrinski: Čakovec, Croatia, 1998; pp. 82–87. [Google Scholar]
- Ciglar, I.; Barić, B.; Tomšić, T.; Šubić, M. Suzbijanje jabukovog savijača (Cydia pomonella) metodom konfuztje. Agron. Glas. Glas. Hrvat. Agron. Društva 2000, 62, 85–93. [Google Scholar]
- Franck, P.; Reyes, M.; Olivares, J.; Sauphanor, B. Genetic architecture in codling moth populations: Comparison between microsatellite and insecticide resistance markers. Mol. Ecol. 2007, 16, 3554–3564. [Google Scholar] [CrossRef]
- Voudouris, C.C.; Franck, P.; Olivares, J.; Sauphanor, B.; Mamuris, Z.; Tsitsipis, J.A.; Margaritopoulos, J.T. Comparing the genetic structure of codling moth Cydia pomonella (L.) from Greece and France: Long distance gene–flow in a sedentary pest species. Bull. Entomol. Res. 2012, 102, 185–198. [Google Scholar] [CrossRef]
- Food and Agriculture Organization of the United Nations. FAO STAT. Available online: http://www.fao.org/faostat/en/#data/QC/visualize (accessed on 31 March 2022).
- Maceljski, M. Poljoprivredna Entomologija, 2nd ed.; Zrinski: Čakovec, Croatia, 2002; pp. 302–309. [Google Scholar]
- Kovačević, Ž. Applied Entomology; University of Zagreb: Zagreb, Croatia, 1952; pp. 312–319. [Google Scholar]
- Sauer, A.J. Novel Types of Resistance of Codling Moth to Cydia pomonella Granulovirus. Ph.D. Thesis, Technische Universität, Darmstadt, Germany, 2017. [Google Scholar]
- Thaler, R.; Brandstätter, A.; Meraner, A.; Chabicovski, M.; Parson, W.; Zelger, R.; Dalla Via, J.; Dallinger, R. Molecular phylogeny and population structure of the codling moth (Cydia pomonella) in Central Europe: II. AFLP analysis reflects human–aided local adaptation of a global pest species. Mol. Phylogenet. Evol. 2008, 48, 838–849. [Google Scholar] [CrossRef]
- Sauphanor, B.; Brosse, V.; Bouvier, J.C.; Speich, P.; Micoud, A.; Martinet, C. Monitoring resistance to diflubenzuron and deltamethrin in French codling moth populations (Cydia pomonella). Pest Manag. Sci. Former. Pest. Sci. 2000, 56, 74–82. [Google Scholar] [CrossRef]
- Mota-Sanchez, D.; Wise, J.C.; Poppen, R.V.; Gut, L.J.; Hollingworth, R.M. Resistance of codling moth, Cydia pomonella (L.) (Lepidoptera: Tortricidae), larvae in Michigan to insecticides with different modes of action and the impact on field residual activity. Pest Manag. Sci. Former. Pest. Sci. 2008, 64, 881–890. [Google Scholar] [CrossRef]
- Hough, W.S. Relative resistance to arsenical poisoning of two codling moth strains. J. Econ. Entomol. 1928, 21, 325–329. [Google Scholar] [CrossRef]
- Reyes, M.; Franck, P.; Charmillot, P.J.; Ioriatti, C.; Olivares, J.; Pasqualini, E.; Sauphanor, B. Diversity of insecticide resistance mechanisms and spectrum in European populations of the codling moth, Cydia pomonella. Pest Manag. Sci. Former. Pest. Sci. 2007, 63, 890–902. [Google Scholar] [CrossRef]
- Thwaite, W.G.; Williams, D.G.; Hately, A.M. Extent and significance of azinphos–methyl resistance in codling moth in Australia. Pest Control Sustain. Agric. 1993, 93, 166–168. [Google Scholar]
- Sauphanor, B.; Bouvier, J.C.; Brosse, V. Spectrum of insecticide resistance in Cydia pomonella (Lepidoptera: Tortricidae) in Southeastern Franc. J. Econ. Entomol. 1998, 91, 1225–1231. [Google Scholar] [CrossRef]
- Reuveny, H.; Cohen, E. Resistance of the codling moth Cydia pomonella (L.) (Lep Tortricidae) to pesticides in Israel. J. Appl. Entomol. 2004, 128, 645–651. [Google Scholar] [CrossRef]
- Arthropod Pesticide Resistance Database (APRD). Cydia pomonella-Shown Resistance to Active Ingredient(s). Available online: https://www.pesticideresistance.org/display.php?page=species&arId=407 (accessed on 30 March 2022).
- Sauphanor, B.; Cuany, A.; Bouvier, J.C.; Brosse, V.; Amichot, M.; Bergé, J.B. Mechanism of resistance to deltamethrin in Cydia pomonella (L.) (Lepidoptera: Tortricidae). Pest. Biochem. Physiol. 1997, 58, 109–117. [Google Scholar] [CrossRef]
- Stara, J.; Nad’ova, K.; Kocourek, F. Insecticide resistance in the codling moth [Cydia pomonella]. J. Fruit Ornam. Plant Res. 2006, 14, 99–106. [Google Scholar]
- Voudouris, C.C.; Sauphanor, B.; Franck, P.; Reyes, M.; Mamuris, Z.; Tsitsipis, J.A.; Vontas, J.; Margaritopoulos, J.T. Insecticide resistance status of the codling moth Cydia pomonella (Lepidoptera: Tortricidae) from Greece. Pest. Biochem. Physiol. 2011, 100, 229–238. [Google Scholar] [CrossRef]
- Reyes, M.; Barros–Parada, W.; Ramírez, C.C.; Fuentes–Contreras, E. Organophosphate resistance and its main mechanism in populations of codling moth (Lepidoptera: Tortricidae) from Central Chile. J. Econ. Entomol. 2015, 108, 277–285. [Google Scholar] [CrossRef]
- Yang, X.Q.; Zhang, Y.L. Investigation of insecticide–resistance status of Cydia pomonella in Chinese populations. Bull. Entomol. Res. 2015, 105, 316–325. [Google Scholar] [CrossRef]
- Bosch, D.; Rodríguez, M.A.; Avilla, J. Monitoring resistance of Cydia pomonella (L.) Spanish field populations to new chemical insecticides and the mechanisms involved. Pest Manag. Sci. 2018, 74, 933–943. [Google Scholar] [CrossRef]
- Ju, D.; Mota-Sanchez, D.; Fuentes-Contreras, E.; Zhang, Y.L.; Wang, X.Q.; Yang, X.Q. Insecticide resistance in the Cydia pomonella (L.): Global status, mechanisms, and research directions. Pest. Biochem. Physiol. 2021, 178, 104925. [Google Scholar] [CrossRef]
- Dunley, J.E.; Welter, S.C. Correlated insecticide cross–resistance in azinphosmethyl resistant codling moth (Lepidoptera: Tortricidae). J. Econ. Entomol. 2000, 93, 955–962. [Google Scholar] [CrossRef]
- Pajač, I.; Barić, B.; Šimon, S.; Mikac, K.M.; Pejić, I. An initial examination of the population genetic structure of Cydia pomonella (Lepidoptera, Tortricidae) in Croatian apple orchards. J. Food Agric. Environ. 2011, 9, 459–464. [Google Scholar]
- Asser–Kaiser, S.; Fritsch, E.; Undorf–Spahn, K.; Kienzle, J.; Eberle, K.E.; Gund, N.A.; Reineke, A.; Zebitz, C.P.W.; Heckel, D.G.; Huber, J.; et al. Rapid emergence of baculovirus resistance in codling moth due to dominant, sex–linked inheritance. Science 2007, 317, 1916–1918. [Google Scholar] [CrossRef] [PubMed]
- Schmitt, A.; Bisutti, I.L.; Ladurner, E.; Benuzzi, M.; Sauphanor, B.; Kienzle, J.; Zingg, D.; Undorf-Spahn, K.; Fritsch, E.; Huber, J.; et al. The occurrence and distribution of resistance of codling moth to Cydia pomonella granulovirus in Europe. J. Appl. Entomol. 2013, 137, 641–649. [Google Scholar] [CrossRef]
- Schulze-Bopp, S.; Jehle, J.A. Development of a direct test of baculovirus resistance in wild codling moth populations. J. Appl. Entomol. 2013, 137, 153–160. [Google Scholar] [CrossRef]
- Zichová, T.; Stará, J.; Kundu, J.K.; Eberle, K.E.; Jehle, J.A. Resistance to Cydia pomonella granulovirus follows a geographically widely distributed inheritance type within Europe. Biocontrol 2013, 58, 525–534. [Google Scholar] [CrossRef]
- Sauer, A.J.; Fritsch, E.; Undorf–Spahn, K.; Nguyen, P.; Marec, F.; Heckel, D.G.; Jehle, J.A. Novel resistance to Cydia pomonella granulovirus (CpGV) in codling moth shows autosomal and dominant inheritance and confers cross–resistance to different CpGV genome groups. PLoS ONE 2017, 12, e0179157. [Google Scholar]
- Sauer, A.J.; Schulze–Bopp, S.; Fritsch, E.; Undorf–Spahn, K.; Jehle, J.A. A third type of resistance to Cydia pomonella granulovirus in codling moths shows a mixed Z–linked and autosomal inheritance pattern. Appl. Environ. Microbiol. 2017, 83, e01036-17. [Google Scholar] [CrossRef]
- Blommers, L.H. Integrated pest management in European apple orchards. Ann. Rev. Entomol. 1994, 39, 213–241. [Google Scholar] [CrossRef]
- Fuentes–Contreras, E.; Espinoza, J.L.; Lavandero, B.; Ramírez, C.C. Population genetic structure of codling moth (Lepidoptera: Tortricidae) from apple orchards in central Chile. J. Econ. Entomol. 2008, 101, 190–198. [Google Scholar] [CrossRef]
- Pashley, D.P.; Bush, G.L. The use of allozymes in studying insect movement with special reference to the codling moth Laspeyresia Pomonella. In Movement of Highly Mobile Insects, Concepts and Methodology in Research, 1st ed.; Rabb, R.L., Kennedy, G.G., Eds.; North Carolina State University Press: Raleigh, NC, USA, 1979; pp. 333–341. [Google Scholar]
- Franck, P.; Guérin, F.; Loiseau, A.; Sauphanor, B. Isolation and characterization of microsatellite loci in the codling moth Cydia pomonella L. (Lepidoptera, Tortricidae). Mol. Ecol. Notes 2005, 5, 99–102. [Google Scholar] [CrossRef]
- Zhou, Y.H.; Gu, H.N.; Dorn, S. Isolation of microsatellite loci in the codling moth Cydia pomonella (Lepidoptera, Tortricidae). Mol. Ecol. Notes 2005, 5, 226–227. [Google Scholar] [CrossRef]
- Timm, A.E.; Geertsema, H.; Warnich, L. Gene flow among Cydia pomonella (Lepidoptera, Tortricidae) geographic and host populations in South Africa. J. Econ. Entomol. 2006, 99, 341–348. [Google Scholar] [CrossRef] [PubMed]
- Franck, P.; Timm, A.E. Population genetic structure of Cydia pomonella: A review and case study comparing spatiotemporal variation. J. Appl. Entomol. 2010, 134, 191–200. [Google Scholar] [CrossRef]
- Chen, M.H.; Dorn, S. Microsatellites reveal genetic differentiation among populations in an insect species with high genetic variability in dispersal, the codling moth, Cydia pomonella (L.) (Lepidoptera: Tortricidae). Bull. Entomol. Res. 2010, 100, 75–85. [Google Scholar] [CrossRef] [PubMed]
- Duan, X.; Li, Y.; Men, Q.; Zhang, M.; Qiao, X.; Harari, A.; Chen, M. Limited gene flow among Cydia pomonella (Lepidoptera: Tortricidae) populations in two isolated regions in China: Implications for utilization of the SIT. Fla. Entomol. 2016, 99, 23–29. [Google Scholar] [CrossRef][Green Version]
- Men, Q.L.; Chen, M.H.; Zhang, Y.L.; Feng, J.N. Genetic structure and diversity of a newly invasive species, the codling moth, Cydia pomonella (L.) (Lepidoptera: Tortricidae) in China. Biol. Invasions 2013, 15, 447–458. [Google Scholar] [CrossRef]
- Li, Y.; Duan, X.; Qiao, X.; Li, X.; Wang, K.; Men, Q.; Chen, M. Mitochondrial DNA revealed the extent of genetic diversity and invasion origin of populations from two separate invaded areas of a newly invasive pest, Cydia pomonella (L.) (Lepidoptera: Tortricidae) in China. Bull. Entomol. Res. 2015, 105, 485–496. [Google Scholar] [CrossRef]
- Kuyulu, A.; Genç, H. Genetic diversity of codling moth Cydia pomonella L. (Lepidoptera: Tortricidae) populations in Turkey. Turk. J. Zool. 2020, 44, 462–471. [Google Scholar] [CrossRef]
- Basoalto, A.; Ramírez, C.C.; Lavandero, B.; Devotto, L.; Curkovic, T.; Franck, P.; Fuentes-Contreras, E. Population genetic structure of codling moth, Cydia pomonella (L.) (Lepidoptera: Tortricidae), in different localities and host plants in Chile. Insects 2020, 11, 285. [Google Scholar] [CrossRef]
- Cichón, L.I.; Soleño, J.; Garrido, S.A.; Guiñazú, N.; Montagna, C.M.; Franck, P.; Olivares, J.; Musleh, S.; Rodríguez, M.A.; Fuentes-Contreras, E. Genetic structure of Cydia pomonella populations in Argentina and Chile implies isolating barriers exist between populations. J. Appl. Entomol. 2021, 145, 911–921. [Google Scholar] [CrossRef]
- Hood, C.S. Geometric morphometric approaches to the study of sexual size dimorphism in mammals. Hystrix 2000, 11, 77–90. [Google Scholar]
- Levine, E.; Oloumi–Sadeghi, H. Western corn rootworm (Coleoptera, Chrysomelidae) larval injury to corn grown for seed production following soybeans grown for seed production. J. Econ. Entomol. 1996, 89, 1010–1016. [Google Scholar] [CrossRef]
- Bouyer, J.; Ravel, S.; Dujardin, J.P.; De Meeüs, T.; Vial, L.; Thévenon, S.; Guerrini, L.; Sidibé, I.; Solano, P. Population structuring of Glossina palpalis gambiensis (Diptera: Glossinidae) according to landscape fragmentation in the Mouhoun river, Burkina Faso. J. Med. Entomol. 2007, 44, 788–795. [Google Scholar] [CrossRef] [PubMed]
- Camara, M.; Caro–Riano, H.; Ravel, S.; Dujardin, J.P.; Hervouet, J.P.; De MeEüs, T.; Bouyer, J.; Solano, P. Genetic and morphometric evidence for population isolation of Glossina palpalis gambiensis (Diptera: Glossinidae) on the Loos islands, Guinea. J. Med. Entomol. 2006, 43, 853–860. [Google Scholar] [CrossRef] [PubMed]
- Lemic, D.; Benítez, H.A.; Püschel, T.A.; Gašparić, H.V.; Šatvar, M.; Bažok, R. Organic morphology of the sugar beet weevil Croatian populations: Evaluating the role of environmental conditions on body shape. J. Comp. Zool. 2016, 260, 25–32. [Google Scholar]
- Benítez, H.A.; Lemic, D.; Püschel, T.A.; Gašparić, H.V.; Kos, T.; Barić, B.; Bažok, R.; Živković, I.P. Fluctuating asymmetry indicates levels of disturbance between agricultural productions: An example in Croatian population of Pterostichus melas melas (Coleptera: Carabidae). Zool. Anz. 2018, 276, 42–49. [Google Scholar] [CrossRef]
- Pajač Živković, I.; Lemic, D.; Mešić, A.; Barić, B.; Órdenes, R.; Benítez, H.A. Effect of fruit host on wing morphology in Drosophila suzukii (Diptera: Drosophilidae): A first view using geometric morphometrics. Entomol. Res. 2018, 48, 262–268. [Google Scholar] [CrossRef]
- Lemic, D.; Benítez, H.A.; Bjeliš, M.; Órdenes–Claveria, R.; Ninčević, P.; Mikac, K.M.; Živković, I.P. Agroorganic effect and sexual shape dimorphism in medfly Ceratitis capitata (Diptera: Tephritidae) an example in Croatian populations. Zool. Anz. 2020, 288, 118–124. [Google Scholar] [CrossRef]
- Lemic, D.; Bjeliš, M.; Ninčević, P.; Živković, I.P.; Popović, L.; Gašparić, H.V.; Benitez, H.A. Medfly phenotypic plasticity as a prerequisite for invasiveness and adaptation. Sustainability 2021, 13, 12510. [Google Scholar] [CrossRef]
- Khaghaninia, S.; Mohammadi, S.A.; Sarafrazi, A.M.; Iraninejad, K.H.; Ebrahimi, E.; Zahiri, R. An analysis of seasonal dimorphism in codling moths, Cydia pomonella, from Iran using geometric morphometrics. Bull. Insectol. 2014, 67, 43–50. [Google Scholar]
- Pajač Živković, I.; Benitez, H.A.; Barić, B.; Drmić, Z.; Kadoić Balaško, M.; Lemic, D.; Dominguez Davila, J.H.; Mikac, K.M.; Bažok, R. Codling moth wing morphology changes due to insecticide resistance. Insects 2019, 10, 310. [Google Scholar] [CrossRef]
- Lemic, D.; Mikac, K.M.; Kozina, A.; Benitez, H.A.; McLean, C.M.; Bažok, R. Monitoring techniques of the western corn rootworm are the precursor to effective IPM strategies. Pest Manag. Sci. 2016, 72, 405–417. [Google Scholar] [CrossRef] [PubMed]
- Garnier, S.; Magniez-Jannin, F.; Rasplus, J.Y.; Alibert, P. When morphometry meets genetics: Inferring the phylogeography of Carabus solieri using Fourier analyses of pronotum and male genitalia. J. Evol. Biol. 2005, 18, 269–280. [Google Scholar] [CrossRef] [PubMed]
- Ortego, J.; Aguirre, M.P.; Cordero, P.J. Fine–scale spatial genetic structure and within population male–biased gene–flow in the grasshopper Mioscirtus wagneri. Evol. Ecol. 2011, 25, 1127–1144. [Google Scholar] [CrossRef]
- Francuski, L.; Milankov, V.; Ludoški, J.; Krtinić, B.; Lundström, J.O.; Kemenesi, G.; Ferenc, J. Genetic and phenotypic variation in central and northern European populations of Aedes (Aedimorphus) vexans (Meigen, 1830) (Diptera, Culicidae). J. Vector Ecol. 2016, 41, 160–171. [Google Scholar] [CrossRef]
- Henriques, D.; Chávez–Galarza, J.; SG Teixeira, J.; Ferreira, H.; Neves, J.C.; Francoy, T.M.; Pinto, M.A. Wing geometric morphometrics of workers and drones and single nucleotide polymorphisms provide similar genetic structure in the Iberian honey bee (Apis mellifera iberiensis). Insects 2020, 11, 89. [Google Scholar] [CrossRef]
- EUR-Lex. Directive 2009/128/EC of the European Parliament and of the Council of 21 October 2009 establishing a framework for Community action to achieve the sustainable use of pesticides (Text with EEA relevance). Off. J. Eur. Union Spec. Ed. Croat. 2009, 15, 253–268. [Google Scholar]
- Reyes, M.; Franck, P.; Olivares, J.; Margaritopoulos, J.; Knight, A.; Sauphanor, B. Worldwide variability of insecticide resistance mechanisms in the codling moth, Cydia pomonella L. (Lepidoptera: Tortricidae). Bull. Entomol. Res. 2009, 99, 359–369. [Google Scholar] [CrossRef]
- Kilian, A.; Wenzl, P.; Huttner, E.; Carling, J.; Xia, L.; Blois, H.; Caig, V.; Heller–Uszynska, K.; Jaccoud, D.; Hopper, C.; et al. Diversity arrays technology. A generic genome profiling technology on open platforms. Methods Mol. Biol. 2012, 888, 67–89. [Google Scholar]
- Jombart, T.; Ahmed, I. Adegenet 1.3–1.new tools for the analysis of genome–wide SNP data. Bioinformatics 2011, 27, 3070–3071. [Google Scholar] [CrossRef]
- Zheng, X.; Levine, D.; Shen, J.; Gogarten, S.M.; Laurie, C.; Weir, B.S. A high–performance computing toolset for relatedness and principal component analysis of SNP data. Bioinformatics 2012, 28, 3326–3328. [Google Scholar] [CrossRef]
- Gruber, B.; Unmack, P.J.; Berry, O.F.; Georges, A. dartr: An r package to facilitate analysis of SNP data generated from reduced representation genome sequencing. Mol. Ecol. Resour. 2018, 18, 691–699. [Google Scholar] [CrossRef] [PubMed]
- R Core Team, R. A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2021. [Google Scholar]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef] [PubMed]
- Earl, D.A.; Vonholdt, B.M. STRUCTURE harvester. A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 2012, 4, 359–361. [Google Scholar] [CrossRef]
- Rohlf, F.J. TpsDig2, Digitize Landmarks and Outlines, Version 2.17 (Program). 2016. Available online: http://life.bio.sunysb.edu/morph (accessed on 20 September 2021).
- Adams, D.C.; Otárola-Castillo, E. Geomorph: An R package for the collection and analysis of geometric morphometric shape data. Methods Ecol. Evol. 2013, 4, 393–399. [Google Scholar] [CrossRef]
- Baken, E.K.; Collyer, M.L.; Kaliontzopoulou, A.; Adams, D.C. geomorph v4.0 and gmShiny: Enhanced analytics and a new graphical interface for a comprehensive morphometric experience. Methods Ecol. Evol. 2021, 12, 2355–2363. [Google Scholar] [CrossRef]
- Klingenberg, C.P. nMorphoJ, An integrated software package for geometric morphometrics. Mol. Ecol. Resour. 2011, 11, 353–357. [Google Scholar] [CrossRef]
- Jolliffe, I.T. Choosing a subset of principal components or variables. Princ. Compon. Anal. 2002, 2, 111–149. [Google Scholar]
- Klingenberg, C.P. Visualizations in geometric morphometrics. How to read and how to make graphs showing shape changes. Hystrix 2013, 24, 15–24. [Google Scholar]
- Monteiro, L.R. Multivariate regression models and geometric morphometrics: The search for causal factors in the analysis of shape. Syst. Biol. 1999, 48, 192–199. [Google Scholar] [CrossRef]
- Jombart, T.; Devillard, S.; Balloux, F. Discriminant analysis of principal components: A new method for the analysis of genetically structured populations. BMC Genet. 2010, 11, 94. [Google Scholar] [CrossRef]
- Pajač, I.; Barić, B.; Mikac, K.M.; Pejić, I. New insights into the biology and ecology of Cydia pomonella from apple orchards in Croatia. Bull. Insectol. 2012, 65, 185–193. [Google Scholar]
- Torres, A.Q.; Valle, D.; Mesquita, R.D.; Schama, R. Gene family evolution and the problem of a functional classification of insect Carboxylesterases. Reference Module in Life Sciences; Elsevier: Amsterdam, The Netherlands, 2018. [Google Scholar]
- Morin, P.A.; Luikart, G.; Wayne, R.K. SNPs in ecology, evolution and conservation. Trends Ecol. Evol. 2004, 19, 208–216. [Google Scholar] [CrossRef]
- Denholm, E.I.; De, G.J.; Williamson, M.S. Insecticide resistance on the move. Science 2002, 297, 2222–2223. [Google Scholar] [CrossRef]
- Belinato, T.A.; Martins, A.J. Insecticide resistance and fitness cost. In Insecticides Resistance; Trdan, S., Ed.; IntechOpen: London, UK, 2016; pp. 243–261. [Google Scholar]
- Grigoraki, L.; Pipini, D.; Labbe, P.; Chaskopoulou, A.; Weill, M.; Vontas, J. Carboxylesterase gene amplifications associated with insecticide resistance in Aedes albopictus: Geographical distribution and evolutionary origin. PLoS Negl. Trop. Dis. 2017, 11, e0005533. [Google Scholar] [CrossRef]
- Saavedra–Rodriguez, K.; Strode, C.; Flores Suarez, A.; Fernandez Salas, I.; Ranson, H.; Hemingway, J.; Black IV, W.C. Quantitative trait loci mapping of genome regions controlling permethrin resistance in the mosquito Aedes aegypti. Genetics 2008, 180, 1137–1152. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Faucon, F.; Dusfour, I.; Gaude, T.; Navratil, V.; Boyer, F.; Chandre, F.; Sirisopa, P.; Thanispong, K.; Juntarajumnong, W.; Poupardin, R.; et al. Identifying genomic changes associated with insecticide resistance in the dengue mosquito Aedes aegypti by deep targeted sequencing. Genome Res. 2015, 25, 1347–1359. [Google Scholar] [CrossRef] [PubMed]
- Adams, D.C.; Rohlf, F.J.; Slice, D.E. A field comes of age: Geometric morphometrics in the 21st century. Hystrix 2013, 24, 7–14. [Google Scholar]
- Benítez, H.A.; Püschel, T.A. Modelando la varianza de la forma: Morfometría geométrica aplicaciones en biología evolutiva. Int. J. Morphol. 2014, 32, 998–1008. [Google Scholar] [CrossRef]
- Mikac, K.M.; Douglas, J.; Spencer, J.L. Wing shape and size of the western corn rootworm (Coleoptera: Chrysomelidae) is related to sex and resistance to soybean–maize crop rotation. J. Econ. Entomol. 2013, 106, 1517–1524. [Google Scholar] [CrossRef]
- Mikac, K.M.; Lemic, D.; Benítez, H.A.; Bažok, R. Changes in corn rootworm wing morphology are related to resistance development. J. Pest Sci. 2019, 92, 443–451. [Google Scholar] [CrossRef]
- DeVries, P.J.; Penz, C.M.; Hill, R.I. Vertical distribution, flight behaviour and evolution of wing morphology in Morpho butterflies. J. Anim. Ecol. 2010, 79, 1077–1085. [Google Scholar] [CrossRef] [PubMed]
- Schumacher, P.; Weyeneth, A.; Weber, D.C.; Dorn, S. Long flights in Cydia pomonella L. (Lepidoptera: Tortricidae) measured by a flight mill: Influence of sex, mated status and age. Physiol. Entomol. 1997, 22, 149–160. [Google Scholar] [CrossRef]
- Varela, L.G.; Welter, S.C.; Jones, V.P.; Brunner, J.F.; Riedl, H. Monitoring and characterization of insecticide resistance Codling moth (Lepidoptera: Tortricidae) in four Western States. J. Econ. Entomol. 1993, 86, 1–10. [Google Scholar] [CrossRef]
- Foster, S. Insecticide Resistance and Its Implications for Potato Production in the UK; British Potato Council. Available online: http://www.potato.org.uk (accessed on 25 November 2021).
- Liu, N. Pyrethroid Resistance in Insects, Genes, Mechanisms, and Regulation. In Insecticides–Advances in Integrated Pest Management, 1st ed.; Perveen, F., Ed.; InTech: Shanghai, China, 2012; pp. 457–468. [Google Scholar]

| Codling Moth Population | Adults Single Nucleotide Polymorphism Genotyped (n) | Geometric Morphometric Wings (n) |
|---|---|---|
| Organic orchards | 44 | 44 |
| Integrated orchards | 44 | 24 |
| Laboratory population | 6 | 99 |
| Population | n | A | p | Ho | He |
|---|---|---|---|---|---|
| Integrated | 44 | 9010 | 1443 | 0.139 | 0.180 |
| Organic | 44 | 9163 | 1931 | 0.147 | 0.180 |
| Laboratory | 6 | 6746 | 187 | 0.104 | 0.118 |
| Overall | 94 | 24919 | 3561 | 0.130 | 0.159 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Balaško, M.K.; Bažok, R.; Mikac, K.M.; Benítez, H.A.; Suazo, M.J.; Viana, J.P.G.; Lemic, D.; Živković, I.P. Population Genetic Structure and Geometric Morphology of Codling Moth Populations from Different Management Systems. Agronomy 2022, 12, 1278. https://doi.org/10.3390/agronomy12061278
Balaško MK, Bažok R, Mikac KM, Benítez HA, Suazo MJ, Viana JPG, Lemic D, Živković IP. Population Genetic Structure and Geometric Morphology of Codling Moth Populations from Different Management Systems. Agronomy. 2022; 12(6):1278. https://doi.org/10.3390/agronomy12061278
Chicago/Turabian StyleBalaško, Martina Kadoić, Renata Bažok, Katarina M. Mikac, Hugo A. Benítez, Manuel J. Suazo, João Paulo Gomes Viana, Darija Lemic, and Ivana Pajač Živković. 2022. "Population Genetic Structure and Geometric Morphology of Codling Moth Populations from Different Management Systems" Agronomy 12, no. 6: 1278. https://doi.org/10.3390/agronomy12061278
APA StyleBalaško, M. K., Bažok, R., Mikac, K. M., Benítez, H. A., Suazo, M. J., Viana, J. P. G., Lemic, D., & Živković, I. P. (2022). Population Genetic Structure and Geometric Morphology of Codling Moth Populations from Different Management Systems. Agronomy, 12(6), 1278. https://doi.org/10.3390/agronomy12061278