Genetic Diversity Analysis and Core Collection Construction of the Actinidia chinensis Complex (Kiwifruit) Based on SSR Markers
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials and DNA Extraction
2.2. SSR Primer Screening and Genotyping
2.3. Analysis of Genetic Diversity and Genetic Structure
2.4. Construction and Evaluation of the Core Collection
3. Results
3.1. Genetic Diversity and Variation of the A. chinensis Complex
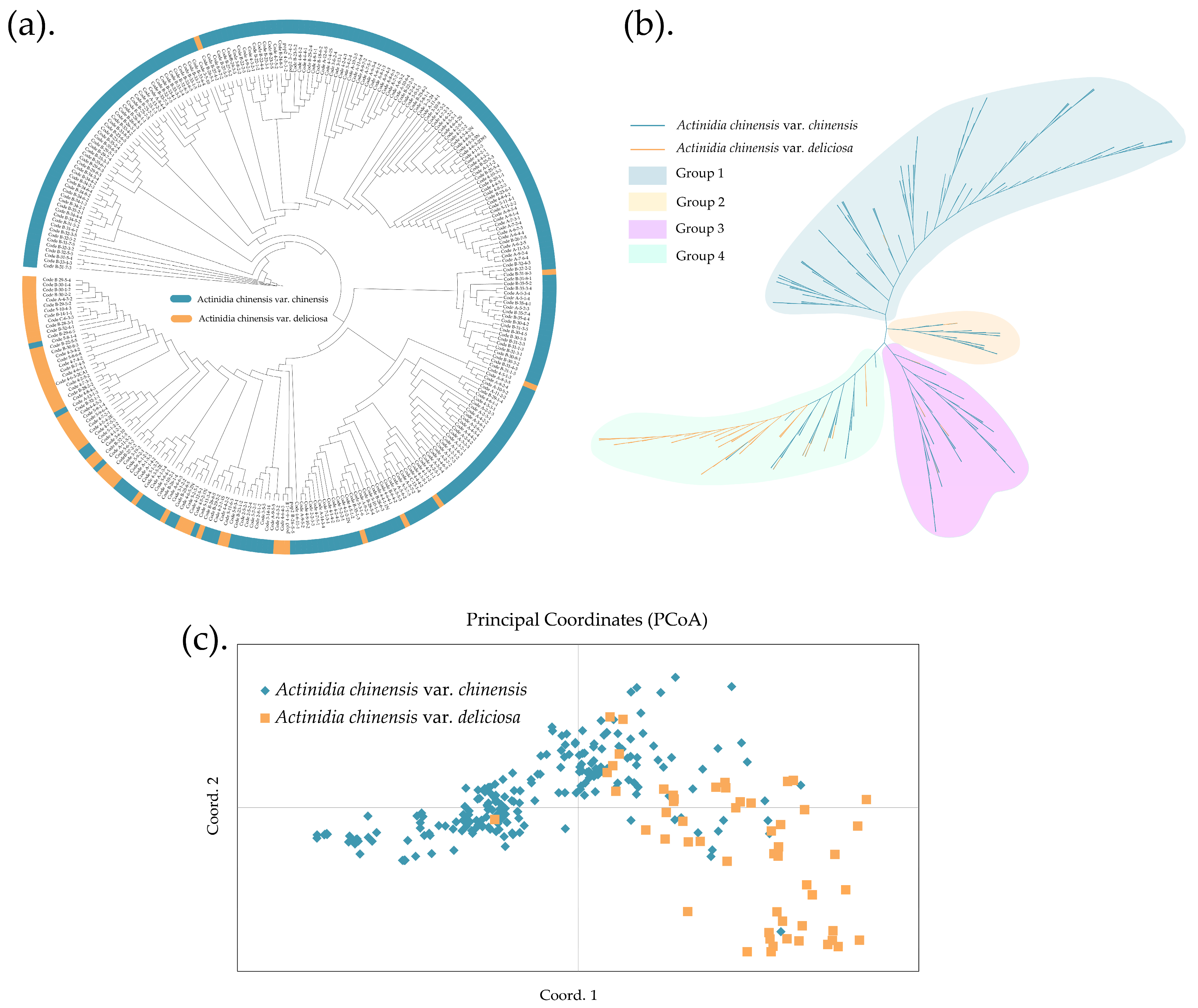
3.2. Cluster Analysis of the A. chinensis Complex
3.3. Genetic Structure of the A. chinensis Complex
3.4. Core Collection of the A. chinensis Complex
3.4.1. Construction of the Core Collection
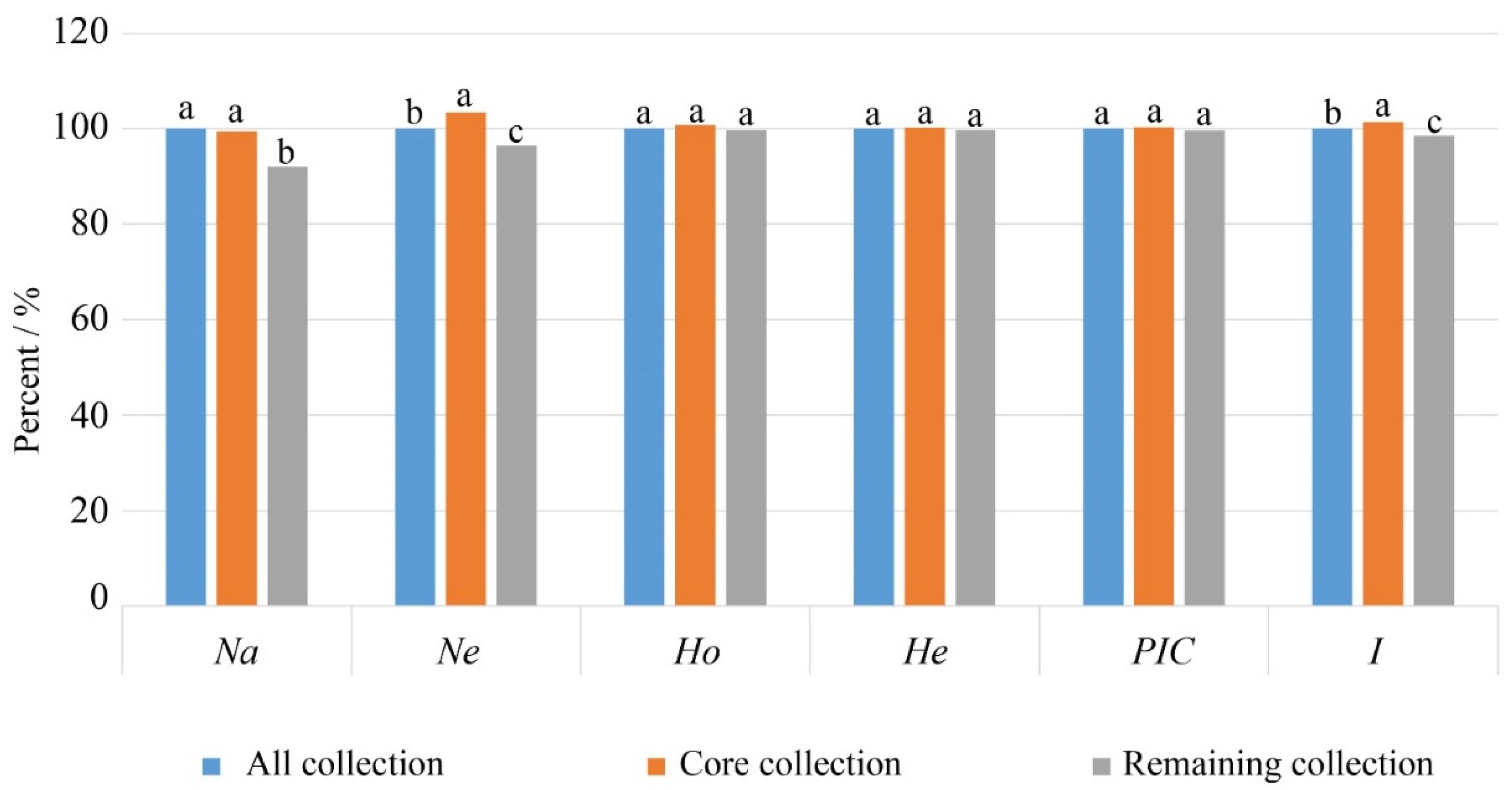
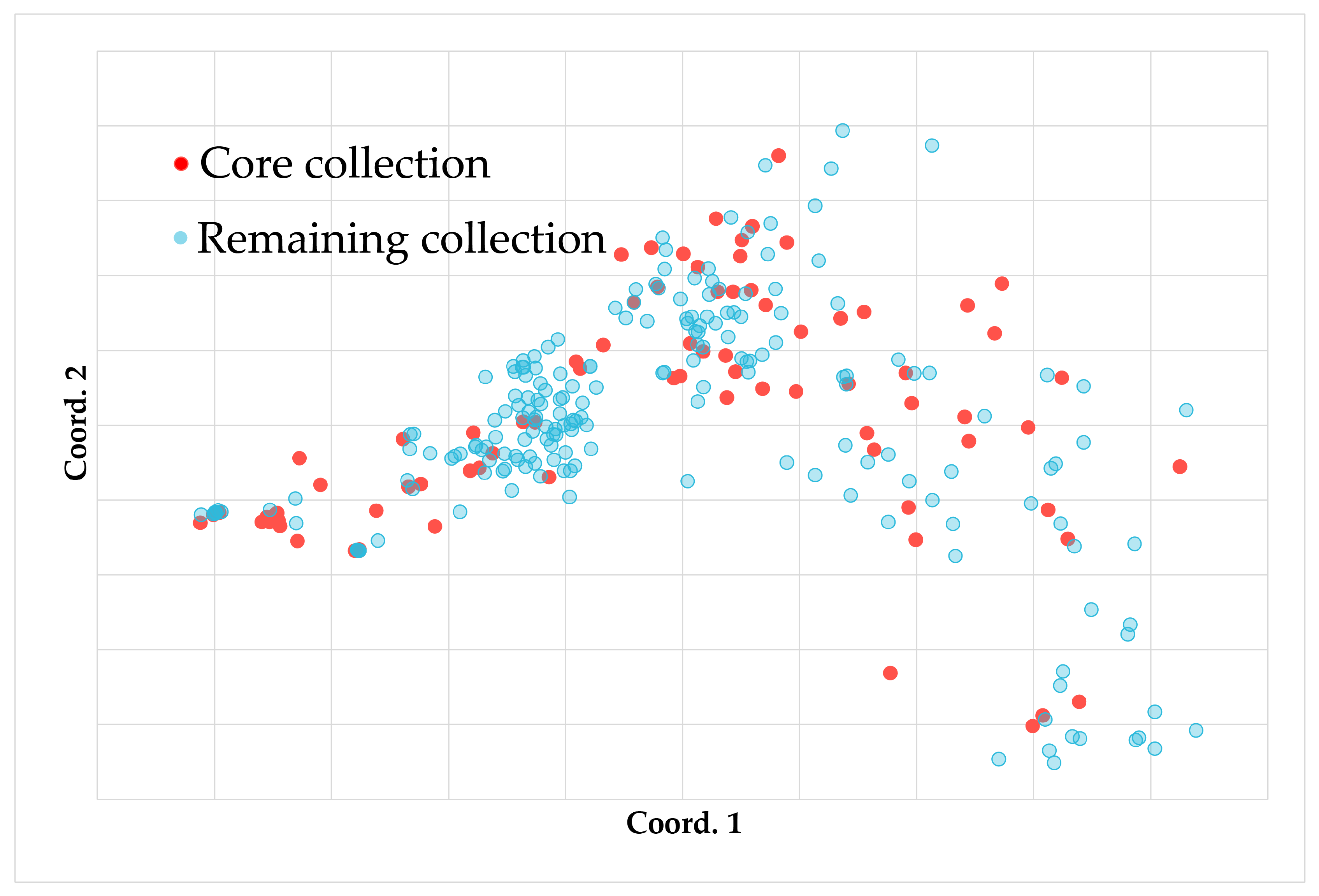
3.4.2. Verification of the Core Collection
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Liu, X.Y.; Wu, R.M.; Bulley, S.M.; Zhong, C.H.; Li, D.W. Kiwifruit MYBS1-like and GBF3 transcription factors influence l-ascorbic acid biosynthesis by activating transcription of GDP-L-galactose phosphorylase 3. New Phytol. 2022, 234, 1782–1800. [Google Scholar] [CrossRef] [PubMed]
- Sanz, V.; López-Hortas, L.; Torres, M.D.; Domínguez, H. Trends in kiwifruit and byproducts valorization. Trends Food Sci. Technol. 2020, 107, 401–414. [Google Scholar] [CrossRef]
- Zhong, C.H.; Huang, W.J.; Li, D.W.; Zhang, Q.; Li, L. Dynamic analysis of global kiwifruit and industry development and fresh fruit trade. China Fruits 2021, 101–108. (In Chinese) [Google Scholar] [CrossRef]
- Zhong, C.H.; Huang, W.J.; Wang, Z.P.; Li, L.; Li, D.W.; Zhang, Q.; Zhao, T.T.; Zhang, P. The breeding progress and development status of the kiwifruit industry in China. Acta Hortic. 2022, 1332, 445–454. [Google Scholar] [CrossRef]
- Liu, Y.B.; Yu, W.H.; Wu, B.F.; Li, J.S. Patterns of genomic divergence in sympatric and allopatric speciation of three Mihoutao (Actinidia) species. Hortic. Res. 2022, 9, uhac054. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.F.; Li, D.W.; Zhang, Q.; Song, C.; Zhong, C.H.; Zhang, X.D.; Wang, Y.; Yao, X.H.; Wang, Z.P.; Zeng, S.H.; et al. Rapid radiations of both kiwifruit hybrid lineages and their parents shed light on a two-layer mode of species diversification. New Phytol. 2017, 215, 877–890. [Google Scholar] [CrossRef]
- Huang, H.W.; Liu, Y.F. Natural hybridization, introgression breeding, and cultivar improvement in the genus Actinidia. Tree Genet. Genomes 2014, 10, 1113–1122. [Google Scholar] [CrossRef]
- Li, Y.; Wang, L.R. Genetic resources, breeding programs in China, and gene mining of peach: A review. Hortic. Plant J. 2020, 6, 205–215. [Google Scholar] [CrossRef]
- Huang, H.W. Genetic resources. In The Kiwifruit Genome; Testolin, R., Huang, H.W., Ferguson, A.R., Eds.; Springer International Publishing: Cham, Switzerland, 2016; pp. 15–36. [Google Scholar] [CrossRef]
- Wang, Z.; Hu, G.M.; Li, Z.Z.; Zhong, C.H.; Yao, X.H. Characterizing tetraploid populations of Actinidia chinensis for kiwifruit genetic improvement. Plants 2022, 11, 1154. [Google Scholar] [CrossRef]
- Brown, A.H.D. Core collections: A practical approach to genetic resources management. Genome 1989, 31, 818–824. [Google Scholar] [CrossRef]
- Guo, Q.; Liu, J.; Li, J.K.; Cao, S.; Zhang, Z.; Zhang, J.T.; Zhang, Y.S.; Deng, Y.P.; Niu, D.S.; Su, L.Z.; et al. Genetic diversity and core collection extraction of Robinia pseudoacacia L. germplasm resources based on phenotype, physiology, and genotyping markers. Ind. Crop. Prod. 2022, 178, 114627. [Google Scholar] [CrossRef]
- Kumar, A.; Kumar, S.; Singh, K.B.M.; Prasad, M.; Thakur, J.K. Designing a mini-core collection effectively representing 3004 diverse rice accessions. Plant Commun. 2020, 1, 100049. [Google Scholar] [CrossRef] [PubMed]
- Esnault, F.; Pellé, R.; Dantec, J.P.; Bérard, A.; Le Paslier, M.C.; Chauvin, J.E. Development of a potato cultivar (Solanum tuberosum L.) core collection, a valuable tool to prospect genetic variation for novel traits. Potato Res. 2016, 59, 329–343. [Google Scholar] [CrossRef]
- Sokolkova, A.; Burlyaeva, M.; Valiannikova, T.; Vishnyakova, M.; Schafleitner, R.; Lee, C.R.; Ting, C.T.; Nair, R.M.; Nuzhdin, S.; Samsonova, M.; et al. Genome-wide association study in accessions of the mini-core collection of mung bean (Vigna radiata) from the World Vegetable Gene Bank (Taiwan). BMC Plant Biol. 2020, 20, 363. [Google Scholar] [CrossRef] [PubMed]
- Pereira-Lorenzo, S.; Ramos-Cabrer, A.M.; Ferreira, V.; Díaz-Hernández, M.B.; Carnide, V.; Pinto-Carnide, O.; Rodrigues, R.; Velázquez-Barrera, M.E.; Rios-Mesa, D.; Ascasíbar-Errasti, J.; et al. Genetic diversity and core collection of Malus × domestica in northwestern Spain, Portugal and the Canary Islands by SSRs. Sci. Hortic. 2018, 240, 49–56. [Google Scholar] [CrossRef]
- Gu, X.Z.; Cao, Y.C.; Xiao, Z.G.; Zhang, Z.H.; Zhang, B.X.; Zhao, H.; Zhang, X.M.; Wang, H.P.; Li, X.X.; Wang, L.H. Genetic diversity and population structure analysis of Capsicum germplasm accessions. J. Integr. Agric. 2019, 18, 1312–1320. [Google Scholar] [CrossRef]
- Han, P.; Tian, X.M.; Wang, Y.; Huang, C.; Ma, Y.Z.; Zhou, X.F.; Yu, Y.; Zhang, D.W.; Xu, H.J.; Cao, Y.; et al. Construction of a core germplasm bank of upland cotton (Gossypium hirsutum L.) based on phenotype, genotype and favorable alleles. Genet. Resour. Crop Evol. 2022, 69, 2399–2411. [Google Scholar] [CrossRef]
- Fickett, N.D.; Ebrahimi, L.; Parco, A.P.; Gutierrez, A.V.; Hale, A.L.; Pontif, M.J.; Todd, J.; Kimbeng, C.A.; Hoy, J.W.; Ayala-Silva, T.; et al. An enriched sugarcane diversity panel for utilization in genetic improvement of sugarcane. Sci. Rep. 2020, 10, 13390. [Google Scholar] [CrossRef]
- Taniguchi, F.; Kimura, K.; Saba, T.; Ogino, A.; Yamaguchi, S.; Tanaka, J. Worldwide core collections of tea (Camellia sinensis) based on SSR markers. Tree Genet. Genomes 2014, 10, 1555–1565. [Google Scholar] [CrossRef]
- Liu, S.L.; Zheng, C.; Xiang, W.; Yi, Z.L.; Xiao, L. A sampling strategy to develop a primary core collection of Miscanthus spp. in China based on phenotypic traits. Agronomy 2022, 12, 678. [Google Scholar] [CrossRef]
- Kaur, V.; Aravind, J.; Manju; Jacob, S.R.; Kumari, J.; Panwar, B.S.; Pal, N.; Rana, J.C.; Pandey, A.; Kumar, A. Phenotypic characterization, genetic diversity assessment in 6,778 accessions of barley (Hordeum vulgare L. ssp. vulgare) germplasm conserved in National Genebank of India and development of a core set. Front. Plant Sci. 2022, 13, 771920. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Hu, X.; Wang, X.; Zhang, J.J.; Peng, X.B.; Hu, Z.G.; Liu, Y.F. Constructing a core collection of the medicinal plant Angelica biserrata using genetic and metabolic data. Front. Plant Sci. 2020, 11, 600249. [Google Scholar] [CrossRef] [PubMed]
- Hyun, D.Y.; Gi, G.Y.; Sebastin, R.; Cho, G.T.; Kim, S.H.; Yoo, E.; Lee, S.; Son, D.M.; Lee, K.J. Utilization of phytochemical and molecular diversity to develop a target-oriented core collection in tea germplasm. Agronomy 2020, 10, 1667. [Google Scholar] [CrossRef]
- Alekseeva, M.; Zagorcheva, T.; Rusanova, M.; Rusanov, K.; Atanassov, I. Genetic and flower volatile diversity in natural populations of Origanum vulgare subsp. hirtum (Link) ietsw. in Bulgaria: Toward the development of a core collection. Front. Plant Sci. 2021, 12, 679063. [Google Scholar] [CrossRef]
- Nie, X.H.; Wang, Z.H.; Liu, N.W.; Song, L.; Yan, B.Q.; Xing, Y.; Zhang, Q.; Fang, K.F.; Zhao, Y.L.; Chen, X.; et al. Fingerprinting 146 Chinese chestnut (Castanea mollissima Blume) accessions and selecting a core collection using SSR markers. J. Integr. Agric. 2021, 20, 1277–1286. [Google Scholar] [CrossRef]
- Dervishi, A.; Jakše, J.; Ismaili, H.; Javornik, B.; Štajner, N. Genetic structure and core collection of olive germplasm from Albania revealed by microsatellite markers. Genes 2021, 12, 256. [Google Scholar] [CrossRef]
- Feng, S.J.; Zhang, J.P.; Mu, Z.H.; Wang, Y.J.; Wen, C.L.; Wu, T.; Yu, C.; Li, Z.; Wang, H.S. Recent progress on the molecular breeding of Cucumis sativus L. in China. Theor. Appl. Genet. 2020, 133, 1777–1790. [Google Scholar] [CrossRef]
- Garrido-Cardenas, J.A.; Mesa-Valle, C.; Manzano-Agugliaro, F. Trends in plant research using molecular markers. Planta 2018, 247, 543–557. [Google Scholar] [CrossRef]
- Kim, J.H.; Oh, Y.; Lee, G.A.; Soon Kwon, Y.; Ae Kim, S.; Kwon, S.I.; Do, Y.S.; Choi, C. Genetic diversity, structure, and core collection of Korean apple germplasm using simple sequence repeat markers. Hortic. J. 2019, 88, 329–337. [Google Scholar] [CrossRef]
- Zhong, Y.C.; Wang, Y.; Sun, Z.M.; Niu, J.; Shi, Y.L.; Huang, K.Y.; Chen, J.; Chen, J.H.; Luan, M.B. Genetic diversity of a natural population of Akebia trifoliata (Thunb.) Koidz and extraction of a core collection using simple sequence repeat markers. Front. Genet. 2021, 12, 716498. [Google Scholar] [CrossRef]
- Hu, G.M.; Zhang, Q.; Han, F.; Li, D.W.; Li, Z.Z.; Wang, Z.; Zhao, T.T.; Tian, H.; Liu, X.L.; Zhong, C.H. Screening and application of universal SSR molecular marker primers in Actinidia. Sci. Agric. Sin. 2022, 55, 3411–3425. (In Chinese) [Google Scholar] [CrossRef]
- Liu, C.Y.; Zhang, Q.; Yao, X.H.; Zhong, C.H.; Yan, C.L.; Huang, H.W. Characterization of genome-wide simple sequence repeats and application in interspecific genetic map integration in kiwifruit. Tree Genet. Genomes 2016, 12, 21. [Google Scholar] [CrossRef]
- Yang, X.S.; Su, W.J.; Wang, L.J.; Lei, J.; Chai, S.J.; Liu, Q.C. Molecular diversity and genetic structure of 380 sweetpotato accessions as revealed by SSR markers. J. Integr. Agric. 2015, 14, 633–641. [Google Scholar] [CrossRef]
- Huang, K.; Dunn, D.W.; Ritland, K.; Li, B.G. polygene: Population genetics analyses for autopolyploids based on allelic phenotypes. Methods Ecol. Evol. 2020, 11, 448–456. [Google Scholar] [CrossRef]
- Paradis, E.; Schliep, K. ape 5.0: An environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 2018, 35, 526–528. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. Interactive Tree of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021, 49, W293–W296. [Google Scholar] [CrossRef] [PubMed]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software structure: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef] [PubMed]
- Earl, D.A.; vonHoldt, B.M. STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 2012, 4, 359–361. [Google Scholar] [CrossRef]
- Jakobsson, M.; Rosenberg, N.A. CLUMPP: A cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 2007, 23, 1801–1806. [Google Scholar] [CrossRef]
- Rosenberg, N.A. Distruct: A program for the graphical display of population structure. Mol. Ecol. Notes 2004, 4, 137–138. [Google Scholar] [CrossRef]
- Peakall, R.; Smouse, P.E. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research—An update. Bioinformatics 2012, 28, 2537–2539. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.W.; Chung, H.K.; Cho, G.T.; Ma, K.H.; Chandrabalan, D.; Gwag, J.G.; Kim, T.S.; Cho, E.G.; Park, Y.J. PowerCore: A program applying the advanced M strategy with a heuristic search for establishing core sets. Bioinformatics 2007, 23, 2155–2162. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.Q.; Gao, J.; Du, Z.F.; Li, D.K.; Wang, Z.; Li, Y.Y.; Pang, X.M. Identifying the genetic diversity, genetic structure and a core collection of Ziziphus jujuba Mill. var. jujuba accessions using microsatellite markers. Sci. Rep. 2016, 6, 31503. [Google Scholar] [CrossRef] [PubMed]
- Chen, R.K.; Hara, T.; Ohsawa, R.; Yoshioka, Y. Analysis of genetic diversity of rapeseed genetic resources in Japan and core collection construction. Breed. Sci. 2017, 67, 239–247. [Google Scholar] [CrossRef] [PubMed]
- Sun, M.H.; Xing, B.; Cui, H.L.; Zhou, B.W.; Zhang, Q.P.; Ren, G.X.; Qin, P.Y. Genetic diversity analysis of quinoa by SSR markers. J. Plant Genet. Resour. 2021, 22, 625–637. (In Chinese) [Google Scholar] [CrossRef]
- Sun, W.H.; Wu, F.F.; Cong, L.; Jin, M.Y.; Wang, X.G. Assessment of genetic diversity and population structure of the genus Vicia (Vicia L.) using simple sequence repeat markers. Grassl. Sci. 2022, 68, 205–213. [Google Scholar] [CrossRef]
- Liao, G.L.; Li, Z.Y.; Huang, C.H.; Zhong, M.; Tao, J.J.; Qu, X.Y.; Chen, L.; Xu, X.B. Genetic diversity of inner quality and SSR association analysis of wild kiwifruit (Actinidia eriantha). Sci. Hortic. 2019, 248, 241–247. [Google Scholar] [CrossRef]
- Lai, J.J.; Li, Z.Z.; Man, Y.P.; Lei, R.; Wang, Y.C. Genetic diversity of five wild Actinidia arguta populations native to China as revealed by SSR markers. Sci. Hortic. 2015, 191, 101–107. [Google Scholar] [CrossRef]
- Lu, X.M.; Man, Y.P.; Lei, R.; Liu, Y.B.; Wu, J.H.; Wang, Y.C. Structural analysis of Actinidia arguta natural populations and preliminary application in association mapping of fruit traits. Sci. Hortic. 2022, 304, 304:111306. [Google Scholar] [CrossRef]
- Guo, R.; Landis, J.B.; Moore, M.J.; Meng, A.P.; Jian, S.G.; Yao, X.H.; Wang, H.C. Development and application of transcriptome-derived microsatellites in Actinidia eriantha (Actinidiaceae). Front. Plant Sci. 2017, 8, 1383. [Google Scholar] [CrossRef]
- Ouni, R.; Zborowska, A.; Sehic, J.; Choulak, S.; Hormaza, J.I.; Garkava-Gustavsson, L.; Mars, M. Genetic diversity and structure of Tunisian local pear germplasm as revealed by SSR markers. Hortic. Plant J. 2020, 6, 61–70. [Google Scholar] [CrossRef]
- Guan, C.F.; Zhang, P.X.; Hu, C.Q.; Chachar, S.; Riaz, A.; Wang, R.Z.; Yang, Y. Genetic diversity, germplasm identification and population structure of Diospyros kaki Thunb. from different geographic regions in China using SSR markers. Sci. Hortic. 2019, 251, 233–240. [Google Scholar] [CrossRef]
- Liu, C.Y.; Li, J.Y.; Qin, G.H. Genome-wide distribution of simple sequence repeats in pomegranate and their application to the analysis of genetic diversity. Tree Genet. Genomes 2020, 16, 36. [Google Scholar] [CrossRef]
- Pérez, V.; Larrañaga, N.; Abdallah, D.; Wünsch, A.; Hormaza, J.I. Genetic diversity of local peach (Prunus persica) accessions from La Palma Island (Canary Islands, Spain). Agronomy 2020, 10, 457. [Google Scholar] [CrossRef]
- Zhang, J.; Chen, T.; Wang, J.; Chen, Q.; Luo, Y.; Zhang, Y.; Tang, H.R.; Wang, X.R. Genetic diversity and population structure in cherry (Cerasus pseudocerasus (Lindl). G. Don) along Longmenshan Fault Zones in China with newly developed SSR markers. Sci. Hortic. 2016, 212, 11–19. [Google Scholar] [CrossRef]
- Urrestarazu, J.; Denancé, C.; Ravon, E.; Guyader, A.; Guisnel, R.; Feugey, L.; Poncet, C.; Lateur, M.; Houben, P.; Ordidge, M.; et al. Analysis of the genetic diversity and structure across a wide range of germplasm reveals prominent gene flow in apple at the European level. BMC Plant Biol. 2016, 16, 130. [Google Scholar] [CrossRef]
- Yu, W.H.; Wu, B.F.; Wang, X.Y.; Yao, Z.; Li, Y.H.; Liu, Y.B. Scale-dependent effects of habitat fragmentation on the genetic diversity of Actinidia chinensis populations in China. Hortic. Res. 2020, 7, 172. [Google Scholar] [CrossRef]
- Zhong, M.; Tao, J.J.; Huang, C.H.; Huang, Q.; Zou, L.F.; Liao, G.L.; Chen, L.; Xu, X.B. Analysis of genetic diversity of populations in Actinidia eriantha Benth. based on simple sequence repeat (SSR) markers. J. Nucl. Agric. Sci. 2019, 33, 863–869. (In Chinese) [Google Scholar] [CrossRef]
- Li, Y.; Li, Y.H.; Yang, Q.W.; Zhang, J.P.; Zhang, J.M.; Qiu, L.J.; Wang, T.Y. Genomics-based crop germplasm research: Advances and perspectives. Sci. Agric. Sin. 2015, 48, 3333–3353. (In Chinese) [Google Scholar] [CrossRef]
- Liu, Y.F.; Liu, Y.L.; Huang, H.W. Genetic variation and natural hybridization among sympatric Actinidia species and the implications for introgression breeding of kiwifruit. Tree Genet. Genomes 2010, 6, 801–813. [Google Scholar] [CrossRef]
- Wang, Z.; Zhong, C.H.; Li, D.W.; Yan, C.L.; Yao, X.H.; Li, Z.Z. Cytotype distribution and chloroplast phylogeography of the Actinidia chinensis complex. BMC Plant Biol. 2021, 21, 325. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Zhang, S.X.; Wu, S.H.; Tian, W.M.; Peng, X.L.; Liu, S.B. Genetic diversity of 33 kiwifruit germplasms based on AFLP markers. J. Biol. 2018, 35, 29–33. (In Chinese) [Google Scholar] [CrossRef]
- Jia, B.; Zhu, L.W.; Yu, X.; Lu, L.J.; Wang, Y.F. RAPD analysis on germplasm resources in genera Actinidia. J. Anhui Agric. Univ. 2005, 32, 381–384. (In Chinese) [Google Scholar] [CrossRef]
- Liu, Y.L.; Geng, Y.P.; Xie, X.D.; Zhang, P.F.; Hou, J.L.; Wang, W.Q. Core collection construction and evaluation of the genetic structure of Glycyrrhiza in China using markers for genomic simple sequence repeats. Genet. Resour. Crop Evol. 2020, 67, 1839–1852. [Google Scholar] [CrossRef]
- Li, S.K.; Wang, R.; Qi, X.J. Recent advances in research on the molecular markers, genetic map and QTL mapping in kiwifruit. J. Fruit Sci. 2022, 39, 662–673. (In Chinese) [Google Scholar] [CrossRef]
- Bourguiba, H.; Batnini, M.A.; Krichen, L.; Trifi-Farah, N.; Audergon, J.M. Population structure and core collection construction of apricot (Prunus armeniaca L.) in North Africa based on microsatellite markers. Plant Genet. Resour. 2017, 15, 21–28. [Google Scholar] [CrossRef]
- Li, H.G.; Xu, J.H.; Du, H.Y.; Wuyun, T.N.; Liu, P.F.; Du, Q.X. Preliminary construction of core collection of Eucommia ulmoides based on allele number maximization strategy. Sci. Silvae Sin. 2018, 54, 42–51. (In Chinese) [Google Scholar] [CrossRef]
- Li, D.W.; Liu, Y.F. Gene introgression from wild relatives. In The Kiwifruit Genome; Testolin, R., Huang, H.W., Ferguson, A.R., Eds.; Springer International Publishing: Cham, Switzerland, 2016; pp. 237–248. [Google Scholar] [CrossRef]
- Huang, H.W. Natural distribution of genus Actinidia. In Kiwifruit: The Genus ACTINIDIA; Huang, H.W., Ed.; Academic Press: San Diego, CA, USA, 2016; pp. 169–190. [Google Scholar] [CrossRef]

| Locus | Na | Ne | Ho | He | PIC | I | Fis |
|---|---|---|---|---|---|---|---|
| A-Geo101 | 25 | 12.317 | 0.712 | 0.919 | 0.913 | 2.677 | 0.225 |
| A-Geo114 | 28 | 7.392 | 0.665 | 0.865 | 0.855 | 2.485 | 0.23 |
| A-Geo131 | 22 | 4.554 | 0.499 | 0.78 | 0.764 | 2.078 | 0.36 |
| A-Geo014 | 28 | 12.012 | 0.7 | 0.917 | 0.911 | 2.71 | 0.236 |
| A-Geo142 | 21 | 10.792 | 0.732 | 0.907 | 0.9 | 2.6 | 0.194 |
| A-Geo156 | 38 | 15.964 | 0.599 | 0.937 | 0.934 | 3.106 | 0.361 |
| A-Geo158 | 22 | 13.153 | 0.789 | 0.924 | 0.919 | 2.717 | 0.146 |
| A-Geo167 | 16 | 2.448 | 0.612 | 0.591 | 0.571 | 1.461 | −0.034 |
| A-Geo018 | 18 | 8.358 | 0.606 | 0.88 | 0.87 | 2.365 | 0.312 |
| A-Geo188 | 26 | 12.697 | 0.687 | 0.921 | 0.916 | 2.782 | 0.254 |
| A-Geo198 | 23 | 7.788 | 0.645 | 0.872 | 0.86 | 2.38 | 0.259 |
| A-Geo218 | 19 | 5.226 | 0.628 | 0.809 | 0.793 | 2.098 | 0.223 |
| A-Geo229 | 32 | 14.543 | 0.673 | 0.931 | 0.927 | 2.969 | 0.277 |
| A-Geo249 | 21 | 9.654 | 0.67 | 0.896 | 0.888 | 2.503 | 0.252 |
| A-Geo257 | 19 | 10.17 | 0.683 | 0.902 | 0.893 | 2.496 | 0.243 |
| A-Geo266 | 20 | 4.45 | 0.61 | 0.775 | 0.758 | 2.059 | 0.213 |
| A-Geo275 | 20 | 3.678 | 0.495 | 0.728 | 0.707 | 1.834 | 0.32 |
| A-Geo290 | 22 | 10.877 | 0.665 | 0.908 | 0.901 | 2.57 | 0.268 |
| A-Geo293 | 21 | 10.728 | 0.617 | 0.907 | 0.9 | 2.587 | 0.319 |
| A-Geo302 | 23 | 11.647 | 0.655 | 0.914 | 0.908 | 2.634 | 0.284 |
| A-Geo307 | 20 | 11.45 | 0.731 | 0.913 | 0.906 | 2.572 | 0.199 |
| A-Geo316 | 22 | 9.692 | 0.606 | 0.897 | 0.888 | 2.463 | 0.324 |
| A-Geo323 | 15 | 1.721 | 0.497 | 0.419 | 0.41 | 1.091 | −0.186 |
| A-Geo335 | 21 | 11.157 | 0.683 | 0.91 | 0.904 | 2.617 | 0.249 |
| A-Geo341 | 20 | 7.343 | 0.626 | 0.864 | 0.852 | 2.336 | 0.275 |
| A-Geo350 | 22 | 8.995 | 0.607 | 0.889 | 0.879 | 2.45 | 0.317 |
| A-Geo365 | 19 | 7.945 | 0.647 | 0.874 | 0.863 | 2.35 | 0.26 |
| A-Geo369 | 14 | 1.901 | 0.347 | 0.474 | 0.448 | 1.076 | 0.269 |
| A-Geo377 | 18 | 7.377 | 0.63 | 0.864 | 0.853 | 2.322 | 0.272 |
| A-Geo038 | 29 | 7.166 | 0.659 | 0.86 | 0.849 | 2.469 | 0.235 |
| A-Geo393 | 20 | 10.711 | 0.549 | 0.907 | 0.899 | 2.555 | 0.395 |
| A-Geo401 | 21 | 7.018 | 0.644 | 0.858 | 0.842 | 2.22 | 0.249 |
| A-Geo407 | 26 | 13.683 | 0.657 | 0.927 | 0.922 | 2.82 | 0.291 |
| A-Geo415 | 18 | 7.864 | 0.632 | 0.873 | 0.861 | 2.33 | 0.276 |
| A-Geo042 | 16 | 7.81 | 0.606 | 0.872 | 0.859 | 2.264 | 0.305 |
| A-Geo426 | 24 | 11.999 | 0.716 | 0.917 | 0.911 | 2.693 | 0.219 |
| A-Geo427 | 21 | 3.301 | 0.479 | 0.697 | 0.684 | 1.873 | 0.312 |
| A-Geo054 | 31 | 14.269 | 0.711 | 0.93 | 0.926 | 2.878 | 0.236 |
| A-Geo068 | 23 | 3.974 | 0.597 | 0.748 | 0.729 | 1.97 | 0.202 |
| A-Geo083 | 24 | 6.5 | 0.32 | 0.846 | 0.832 | 2.318 | 0.622 |
| Mean | 22.2 | 8.758 | 0.622 | 0.846 | 0.835 | 2.369 | 0.257 |
| Population | n | Na | Ne | Ho | He | PIC | I | Fis |
|---|---|---|---|---|---|---|---|---|
| A. chinensis | 240 | 21.725 | 8.689 | 0.625 | 0.849 | 0.838 | 2.367 | 0.257 |
| A. deliciosa | 54 | 17.150 | 6.442 | 0.611 | 0.786 | 0.771 | 2.086 | 0.225 |
| Mean | 19.438 | 7.566 | 0.618 | 0.818 | 0.805 | 2.227 | 0.241 |
| Source of Variation | Degrees of Freedom | Sum of Square | Mean of Square | Variance Component | Percentage of Variation (%) | p |
|---|---|---|---|---|---|---|
| Among variants | 1 | 259.56 | 259.56 | 2.13 | 2.90 | <0.001 |
| Within variants | 292 | 20,869 | 71.47 | 71.47 | 97.10 | <0.001 |
| Total | 293 | 21,128.56 | - | 73.60 | 100 | - |
| Number | Name | Code | Variant | Category | Selection Method |
|---|---|---|---|---|---|
| 1 | CSXZSIII | 3-0-2-4 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 2 | CXHY4 | 3-2-24 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 3 | Taibao No. 2 | 4-1-1-2 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 4 | Jinyang | 4-3-1-1 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 5 | Jinyi | 4-3-2-1 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 6 | Kuimi | 4-3-3-1 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 7 | Huayou | 4-4-2-2 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 8 | DH-1 | 4-6-5-2N | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 9 | DD-111 | 4-7-5-2 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 10 | Hongyang | 4-8-4-2 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 11 | Fengyue | 4-9-2-2 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 12 | Wanjin | 5-10-1-3 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 13 | Chuhong | 5-11-3-4 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 14 | Jinxia | 5-2-2-1 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 15 | Hort16A | 5-4-2-4 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 16 | Cuiyu | 5-7-2-1 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 17 | Taishanghuang | 5-8-1-1 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 18 | ZH8792 ♂ | 8-9-7-2 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 19 | FC-1-1-4 | A-13-2-3 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 20 | XB-48-2-10 | A-1-6-3 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 21 | XB-49-5-9 | A-2-5-1 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 22 | XB-51-1-3 | A-3-6-1 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 23 | CD-19-1-28 | B-20-1-1 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 24 | Wanmi | 5-8-2-2 | A. chinensis var. chinensis | Cultivars (lines) | PowerCore |
| 25 | P610726070 | 5-8-4-1 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 26 | 17-SCLS-41 | A-10-2-1 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 27 | 17-SCLS-50 | A-11-5-2 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 28 | 18-ANXN-31 | A-5-2-3 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 29 | 18-ANXN-33 | A-5-3-4 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 30 | 17-SCLS-2 | A-5-5-3 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 31 | 17-SCLS-33 | A-9-2-1 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 32 | 18-HN-1 | B-22-1-2 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 33 | 18-HN-13 | B-23-1-2 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 34 | 18-ZHLS-1 | B-23-6-5 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 35 | 17-ZET-2 | B-25-7-3 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 36 | JXRJ | B-27-5-2 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 37 | YSZY44 ♂ | B-29-2-2 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 38 | 18-AHJX-02 | B-30-3-2 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 39 | 18-AHJX-04 | B-30-4-5 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 40 | 18-AHQM-12 | B-31-7-3 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 41 | 18-AHQM-21 | B-32-2-2 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 42 | 18-AHQM-25 | B-32-3-5 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 43 | 18-AHQM-28 | B-32-5-2 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 44 | 18-AHQS-07 | B-32-7-5 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 45 | 18-AHQS-08 | B-32-8-3 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 46 | 18-AHQS-06 | B-33-1-3 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 47 | 18-AHQS-10 | B-33-3-3 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 48 | 18-AHQS-22 | B-33-7-5 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 49 | 18-AHQY-19 | B-34-5-2 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 50 | 18-AHXN-11 | B-35-2-4 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 51 | 18-AHXN-12 | B-35-3-3 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 52 | 18-AHXN-13 | B-35-3-4 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 53 | 18-AHXN-16 | B-35-4-1 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 54 | 18-AHXN-20 | B-35-4-4 | A. chinensis var. chinensis | Wild resources | PowerCore |
| 55 | Donghong | 3-10-9 | A. chinensis var. chinensis | Cultivars (lines) | Supplementary cultivar |
| 56 | Moshan No. 2 ♂ | 3-7-4-2 | A. chinensis var. chinensis | Cultivars (lines) | Supplementary cultivar |
| 57 | Lushanxiang | 4-1-4-2 | A. chinensis var. chinensis | Cultivars (lines) | Supplementary cultivar |
| 58 | Tongshan 5 | 4-2-1-1S | A. chinensis var. chinensis | Cultivars (lines) | Supplementary cultivar |
| 59 | Moshan No. 4 | 4-2-2-2N | A. chinensis var. chinensis | Cultivars (lines) | Supplementary cultivar |
| 60 | Jinnong | 4-2-4-1 | A. chinensis var. chinensis | Cultivars (lines) | Supplementary cultivar |
| 61 | Moshan No. 1 ♂ | 4-3-3-2 | A. chinensis var. chinensis | Cultivars (lines) | Supplementary cultivar |
| 62 | Zaoxian | 4-4-1-1N | A. chinensis var. chinensis | Cultivars (lines) | Supplementary cultivar |
| 63 | Huabao No. 1 | 4-6-1-1 | A. chinensis var. chinensis | Cultivars (lines) | Supplementary cultivar |
| 64 | Golden peach | 4-6-11-1 | A. chinensis var. chinensis | Cultivars (lines) | Supplementary cultivar |
| 65 | Xiaya No. 1 | 4-6-4-1 | A. chinensis var. chinensis | Cultivars (lines) | Supplementary cultivar |
| 66 | Guihai No. 4 | 4-8-1-1 | A. chinensis var. chinensis | Cultivars (lines) | Supplementary cultivar |
| 67 | Moshan No. 7 ♂ | 4-8-1-2 | A. chinensis var. chinensis | Cultivars (lines) | Supplementary cultivar |
| 68 | Jinyuan | 2-2-3-3 | A. chinensis var. chinensis | Cultivars (lines) | Supplementary cultivar |
| 69 | Jinyan | 4-8-10-2 | A. chinensis var. chinensis | Cultivars (lines) | Supplementary cultivar |
| 70 | Jinmei | 7-2-3-1 | A. chinensis var. chinensis | Cultivars (lines) | Supplementary cultivar |
| 71 | Jinmei | 2-4-3-2 | A. chinensis var. deliciosa | Cultivars (lines) | PowerCore |
| 72 | Longshanhong | 4-12-5-1 | A. chinensis var. deliciosa | Cultivars (lines) | PowerCore |
| 73 | Xinguan No. 2 | 4-3-5-2E | A. chinensis var. deliciosa | Cultivars (lines) | PowerCore |
| 74 | Shixuana No. 2 | 4-4-5-3 | A. chinensis var. deliciosa | Cultivars (lines) | PowerCore |
| 75 | Dongmei | 4-5-1-3 | A. chinensis var. deliciosa | Cultivars (lines) | PowerCore |
| 76 | Qinxiang | 4-5-2-1N | A. chinensis var. deliciosa | Cultivars (lines) | PowerCore |
| 77 | Chang’an No. 1 | 4-6-3-1 | A. chinensis var. deliciosa | Cultivars (lines) | PowerCore |
| 78 | Xiangma No. 6 | 4-6-4-3 | A. chinensis var. deliciosa | Cultivars (lines) | PowerCore |
| 79 | Xuxiang | 4-7-3-1 | A. chinensis var. deliciosa | Cultivars (lines) | PowerCore |
| 80 | Chuanmi No. 1 | 4-7-4-2 | A. chinensis var. deliciosa | Cultivars (lines) | PowerCore |
| 81 | Xuxiang ♂ | 4-8-4-3 | A. chinensis var. deliciosa | Cultivars (lines) | PowerCore |
| 82 | Qinmei | 5-10-4-3 | A. chinensis var. deliciosa | Cultivars (lines) | PowerCore |
| 83 | Guichang | 5-11-6-3 | A. chinensis var. deliciosa | Cultivars (lines) | PowerCore |
| 84 | Hayward | 5-9-1-4 | A. chinensis var. deliciosa | Cultivars (lines) | PowerCore |
| 85 | Huamei No. 1 | 5-9-2-1 | A. chinensis var. deliciosa | Cultivars (lines) | PowerCore |
| 86 | 18-20161395 | B-26-5-1 | A. chinensis var. deliciosa | Cultivars (lines) | PowerCore |
| 87 | Shixuana No. 1 | B-28-2-2 | A. chinensis var. deliciosa | Cultivars (lines) | PowerCore |
| 88 | Miliang No. 1 B2831 | B-14-1-1 | A. chinensis var. deliciosa | Cultivars (lines) | PowerCore |
| 89 | 18-AHHS-01 | B-29-5-1 | A. chinensis var. deliciosa | Wild resources | PowerCore |
| 90 | Moshan No. 3 ♂ | 3-19-2 | A. chinensis var. deliciosa | Cultivars (lines) | Supplementary cultivar |
| 91 | Sanxia No. 1 | 4-2-5-2 | A. chinensis var. deliciosa | Cultivars (lines) | Supplementary cultivar |
| 92 | Xianglv | 4-5-4-2E | A. chinensis var. deliciosa | Cultivars (lines) | Supplementary cultivar |
| 93 | Bruno | 4-7-5-3 | A. chinensis var. deliciosa | Cultivars (lines) | Supplementary cultivar |
| Population | n | Na | Ne | Ho | He | PIC | I |
|---|---|---|---|---|---|---|---|
| All collection | 294 (100%) | 22.2 | 8.758 | 0.622 | 0.846 | 0.835 | 2.369 |
| Core collection 1 | 73 (24.83%) | 21.1 | 9.081 | 0.635 | 0.849 | 0.84 | 2.407 |
| Core collection 2 | 93 (31.63%) | 21.975 | 9.058 | 0.627 | 0.848 | 0.838 | 2.401 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hu, G.; Jiang, Q.; Wang, Z.; Li, Z.; Liao, W.; Shen, D.; Zhong, C. Genetic Diversity Analysis and Core Collection Construction of the Actinidia chinensis Complex (Kiwifruit) Based on SSR Markers. Agronomy 2022, 12, 3078. https://doi.org/10.3390/agronomy12123078
Hu G, Jiang Q, Wang Z, Li Z, Liao W, Shen D, Zhong C. Genetic Diversity Analysis and Core Collection Construction of the Actinidia chinensis Complex (Kiwifruit) Based on SSR Markers. Agronomy. 2022; 12(12):3078. https://doi.org/10.3390/agronomy12123078
Chicago/Turabian StyleHu, Guangming, Quan Jiang, Zhi Wang, Zuozhou Li, Wenyue Liao, Dandan Shen, and Caihong Zhong. 2022. "Genetic Diversity Analysis and Core Collection Construction of the Actinidia chinensis Complex (Kiwifruit) Based on SSR Markers" Agronomy 12, no. 12: 3078. https://doi.org/10.3390/agronomy12123078
APA StyleHu, G., Jiang, Q., Wang, Z., Li, Z., Liao, W., Shen, D., & Zhong, C. (2022). Genetic Diversity Analysis and Core Collection Construction of the Actinidia chinensis Complex (Kiwifruit) Based on SSR Markers. Agronomy, 12(12), 3078. https://doi.org/10.3390/agronomy12123078





