Metabolomic, Ionomic and Microbial Characterization of Olive Xylem Sap Reveals Differences According to Plant Age and Genotype
Abstract
1. Introduction
2. Materials and Methods
2.1. Olive Plant Material and Sampling
2.2. Xylem Sap Extraction
2.3. Metabolomic and Ionomic Analysis
2.4. Microbiome Analysis
2.5. Statistical and Bioinformatics Analyses
3. Results
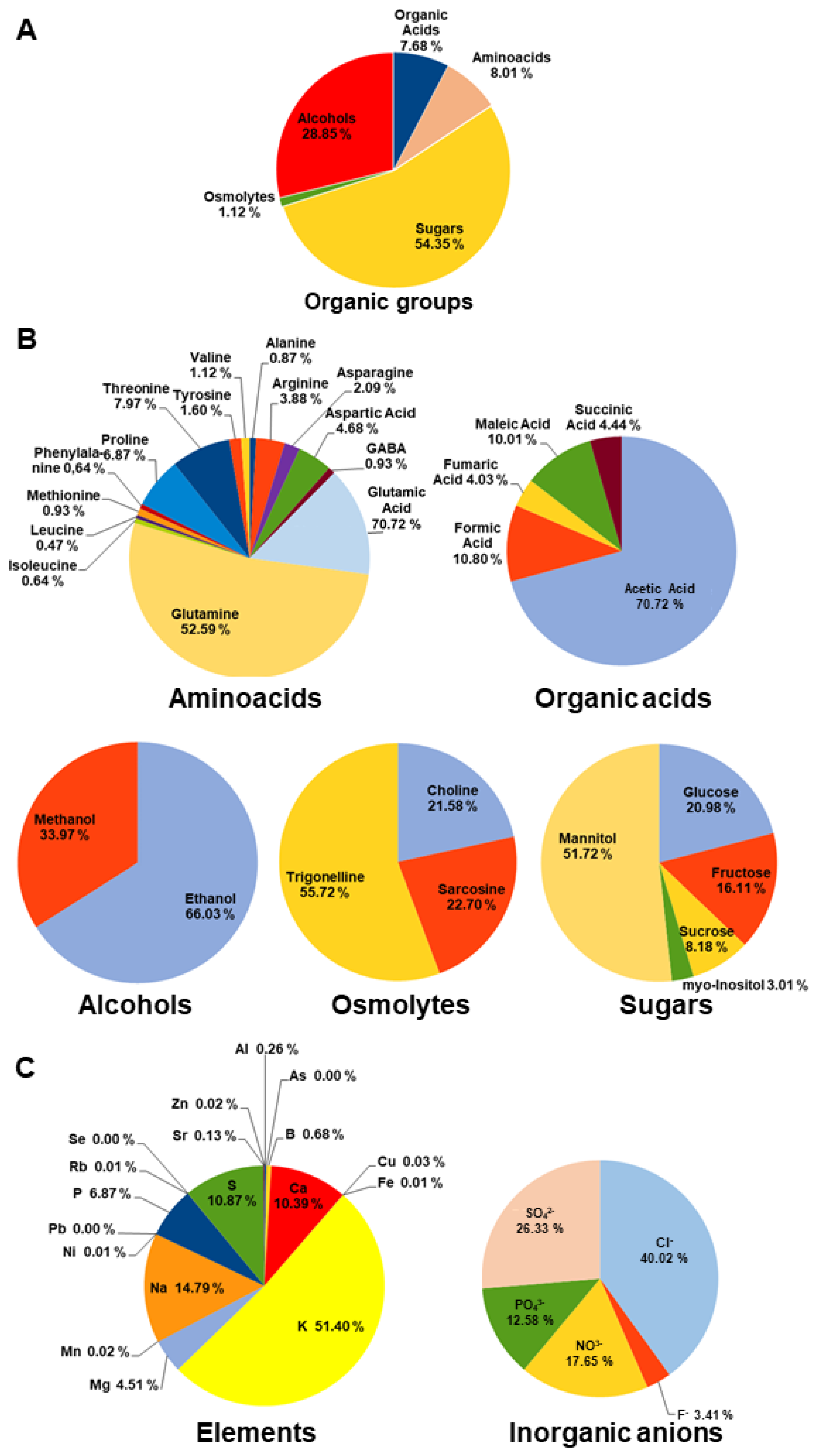
3.1. Metabolite and Ion Profiles in Olive Xylem Sap
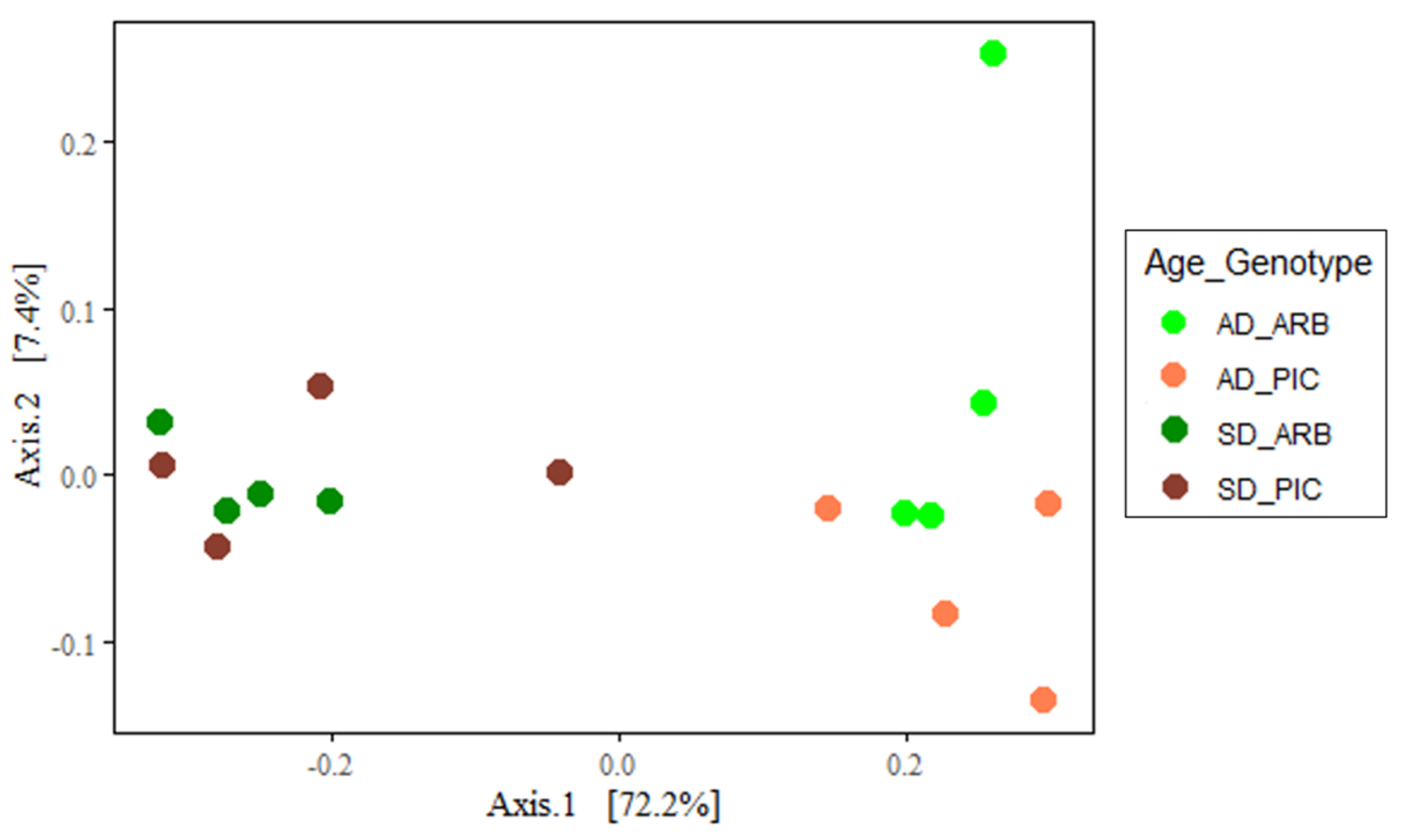
3.2. Alpha and Beta Microbial Diversity
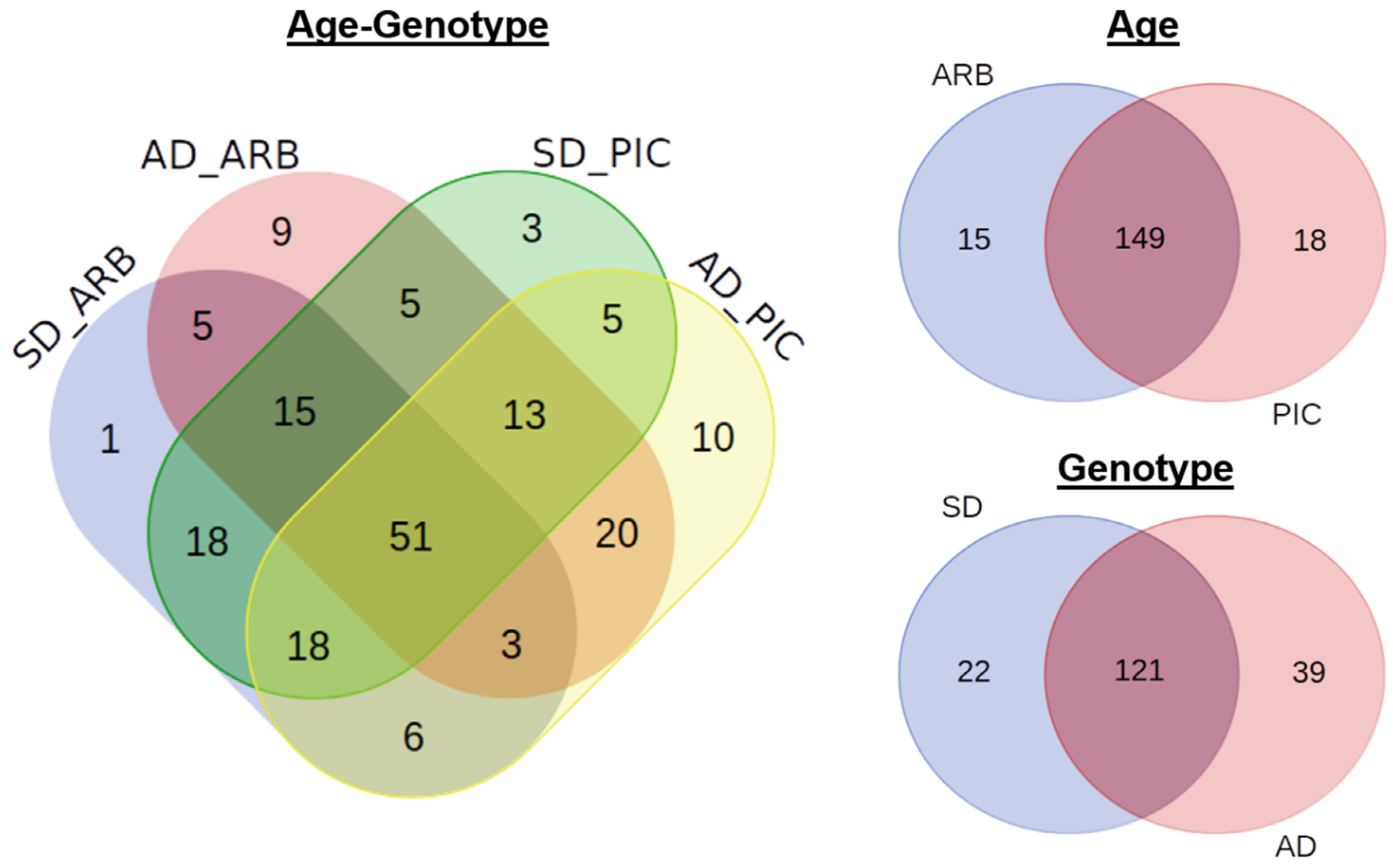
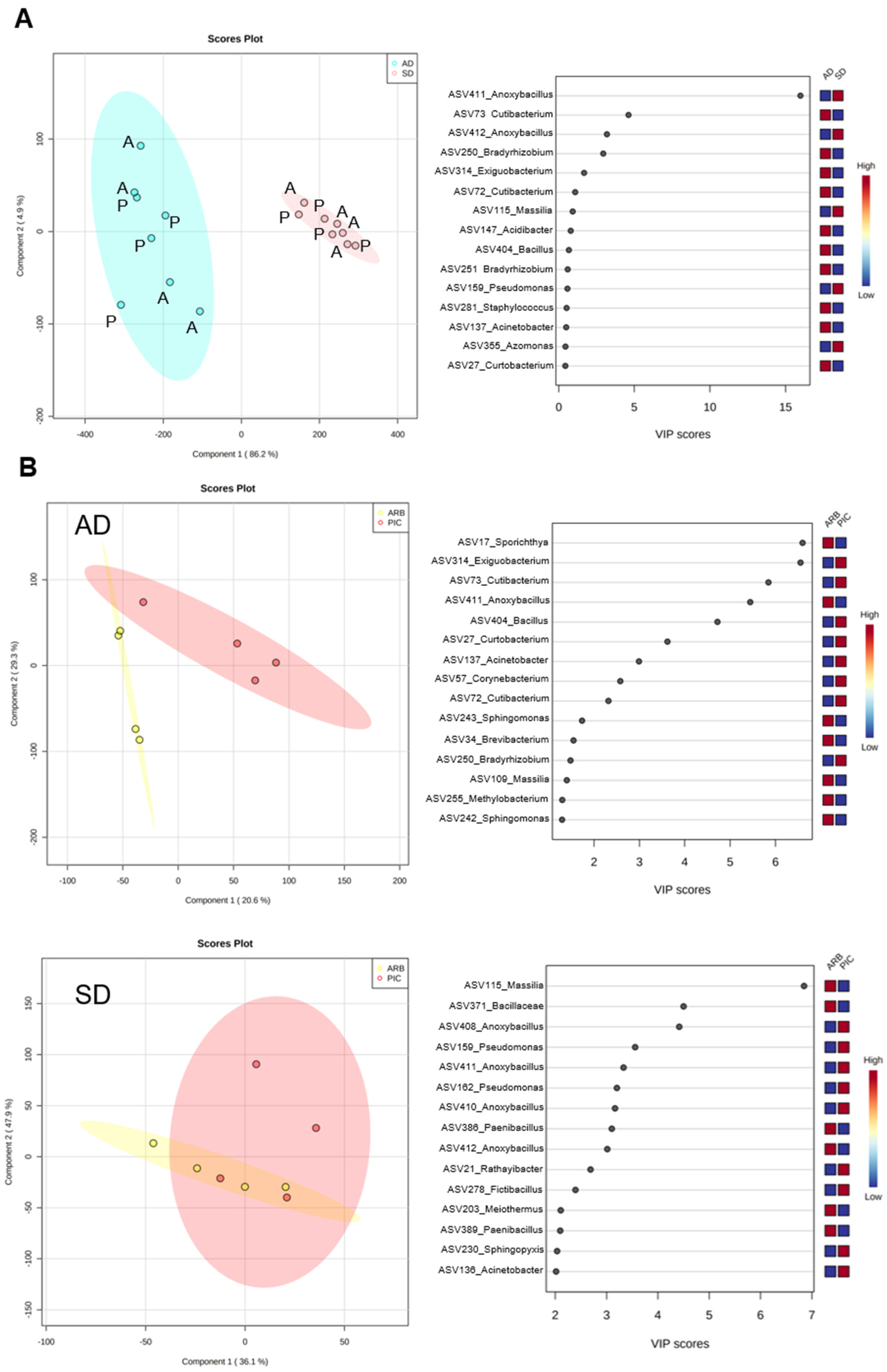
3.3. Composition and Abundance of Bacterial Communities in Olive Xylem Sap
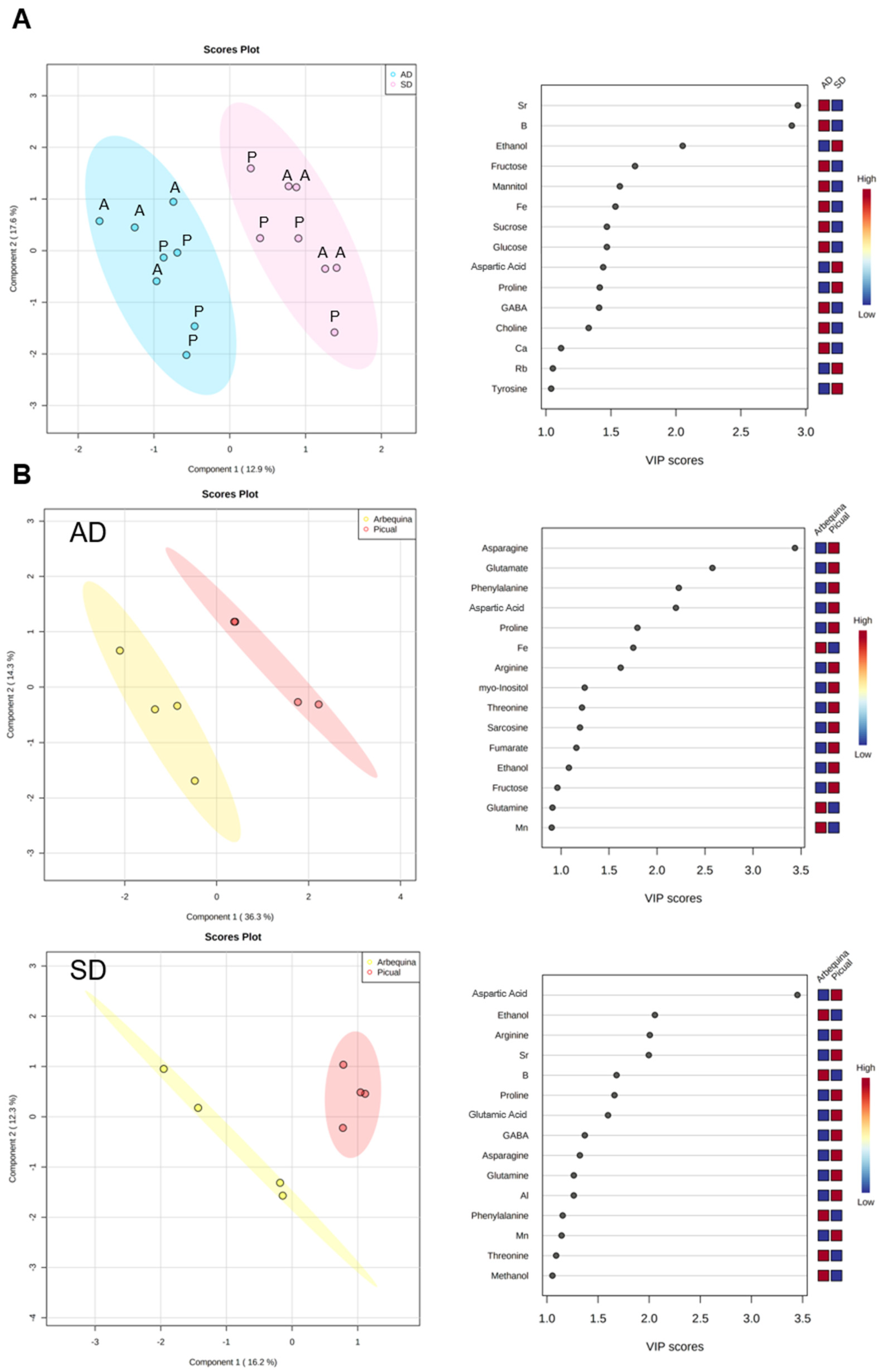
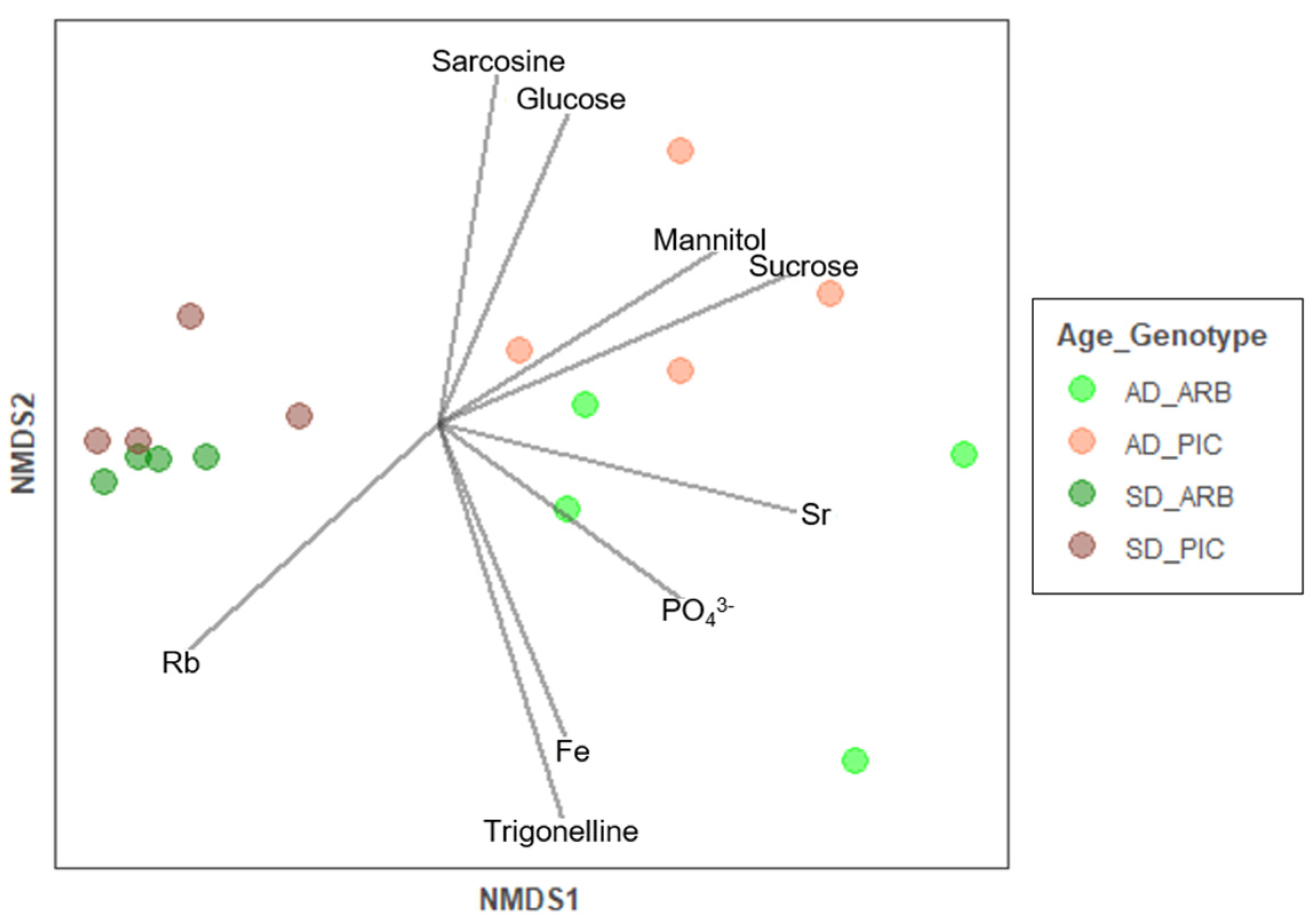
3.4. Effect of Olive Plant Age and Genotype in Xylem Sap Chemical and Microbial Composition
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Landa, B.B.; Pérez, A.G.; Luaces, P.; Montes-Borrego, M.; Navas-Cortés, J.A.; Sanz, C. Insights into the Effect of Verticillium dahliae Defoliating-Pathotype Infection on the Content of Phenolic and Volatile Compounds Related to the Sensory Properties of Virgin Olive Oil. Front. Plant Sci. 2019, 10, 232. [Google Scholar] [CrossRef]
- Allen, H.D.; Randall, R.E.; Amable, G.S.; Devereux, B.J. The impact of changing olive cultivation practices on the ground flora of olive groves in the Messara and Psiloritis regions, Crete, Greece. Land Degrad. Dev. 2006, 17, 249–273. [Google Scholar] [CrossRef]
- Gómez, A.J.; Infante-Amate, J.; De Molina, G.M.; Vanwalleghem, T.; Taguas, V.E.; Lorite, I. Olive Cultivation, its Impact on Soil Erosion and its Progression into Yield Impacts in Southern Spain in the Past as a Key to a Future of Increasing Climate Uncertainty. Agriculture 2014, 4, 170–198. [Google Scholar] [CrossRef]
- Tscheulin, T.; Neokosmidis, L.; Petanidou, T.; Settele, J. Influence of landscape context on the abundance and diversity of bees in Mediterranean olive groves. Bull. Èntomol. Res. 2011, 101, 557–564. [Google Scholar] [CrossRef] [PubMed]
- Jiménez-Díaz, R.M.; Cirulli, M.; Bubici, G.; Jiménez-Gasco, M.D.M.; Antoniou, P.P.; Tjamos, E.C. Verticillium Wilt, A Major Threat to Olive Production: Current Status and Future Prospects for its Management. Plant Dis. 2012, 96, 304–329. [Google Scholar] [CrossRef] [PubMed]
- Saponari, M.; Giampetruzzi, A.; Loconsole, G.; Boscia, D.; Saldarelli, P. Xylella fastidiosa in Olive in Apulia: Where We Stand. Phytopathology 2019, 109, 175–186. [Google Scholar] [CrossRef] [PubMed]
- White, P.J. Chapter 3—Long-distance Transport in the Xylem and Phloem. In Marschner’s Mineral Nutrition of Higher Plants; Third, E., Ed.; Academic Press: San Diego, CA, USA, 2012; pp. 49–70. ISBN 978-0-12-384905-2. [Google Scholar]
- De Boer, A.H.; Volkov, V. Logistics of water and salt transport through the plant: Structure and functioning of the xylem. Plant Cell Environ. 2003, 26, 87–101. [Google Scholar] [CrossRef]
- Schurr, U. Xylem sap sampling—New approaches to an old topic. Trends Plant Sci. 1998, 3, 293–298. [Google Scholar] [CrossRef]
- Horsfield, D. Relationships between feeding of Philaenus spumarius (L.) and the amino acid concentration in the xylem sap. Ecol. Èntomol. 1977, 2, 259–266. [Google Scholar] [CrossRef]
- Sauter, J.J. Seasonal Variation of Amino Acids and Amides in the Xylem Sap of Salix. Z. Pflanzenphysiol. 1981, 101, 399–411. [Google Scholar] [CrossRef]
- Schurr, U.; Schulze, E.-D. The concentration of xylem sap constituents in root exudate, and in sap from intact, transpiring castor bean plants (Ricinus communis L.). Plant Cell Environ. 1995, 18, 409–420. [Google Scholar] [CrossRef]
- Tatar, E.; Mihucz, V.; Varga, A.; Zaray, G.; Fodor, F. Determination of Organic Acids in Xylem Sap of Cucumber: Effect of Lead Contamination. Microchem. J. 1998, 58, 306–314. [Google Scholar] [CrossRef]
- Garcia, N.F.; Hernandez, M.; Casado-Vela, J.; Bru, R.; Elortza, F.; Hedden, P.; Olmos, E. Changes to the proteome and targeted metabolites of xylem sap in Brassica oleracea in response to salt stress. Plant Cell Environ. 2011, 34, 821–836. [Google Scholar] [CrossRef]
- Krishnan, H.B.; Natarajan, S.S.; Bennett, J.O.; Sicher, R.C. Protein and metabolite composition of xylem sap from field-grown soybeans (Glycine max). Planta 2011, 233, 921–931. [Google Scholar] [CrossRef]
- Alvarez, S.; Marsh, E.L.; Schroeder, S.G.; Schachtman, D.P. Metabolomic and proteomic changes in the xylem sap of maize under drought. Plant Cell Environ. 2008, 31, 325–340. [Google Scholar] [CrossRef] [PubMed]
- Gollan, T.; Schurr, U.; Schulze, E.-D. Stomatal response to drying soil in relation to changes in the xylem sap composition of Helianthus annuus. I. The concentration of cations, anions, amino acids in, and pH of, the xylem sap. Plant Cell Environ. 1992, 15, 551–559. [Google Scholar] [CrossRef]
- Ferguson, A.R. Xylem Sap from Actinidia chinensis: Apparent Differences in Sap Composition Arising from the Method of Collection. Ann. Bot. 1980, 46, 791–801. [Google Scholar] [CrossRef]
- Dambrine, E.; Martin, F.; Carisey, N.; Granier, A.; Hällgren, J.-E.; Bishop, K. Xylem sap composition: A tool for investigating mineral uptake and cycling in adult spruce. Plant Soil 1995, 168, 233–241. [Google Scholar] [CrossRef]
- Jones, O.P. Effects of Rootstocks and Interstocks on the Xylem Sap Composition in Apple Trees: Effects on Nitrogen, Phosphorus, and Potassium Content. Ann. Bot. 1971, 35, 825–836. [Google Scholar] [CrossRef]
- Subramanian, S.; Cho, U.-H.; Keyes, C.; Yu, O. Distinct changes in soybean xylem sap proteome in response to pathogenic and symbiotic microbe interactions. BMC Plant Biol. 2009, 9, 119. [Google Scholar] [CrossRef]
- Wallis, C.M.; Chen, J. Grapevine Phenolic Compounds in Xylem Sap and Tissues Are Significantly Altered During Infection by Xylella fastidiosa. Phytopathology 2012, 102, 816–826. [Google Scholar] [CrossRef] [PubMed]
- Thomas, R.J.; Feller, U.; Erismann, K.H. The Effect of Different Inorganic Nitrogen Sources and Plant Age on the Composition of Bleeding Sap of Phaseolus Vulgaris. New Phytol. 1979, 82, 657–669. [Google Scholar] [CrossRef]
- Tromp, J. Seasonal Variations in the Composition of Xylem Sap of Apple with Respect to K, Ca, Mg, and N. Z. Pflanzenphysiol. 1979, 94, 189–194. [Google Scholar] [CrossRef]
- Ferguson, A.R.; Eiseman, J.A.; Leonard, J.A. Xylem Sap from Actinidia chinensis: Seasonal Changes in Composition. Ann. Bot. 1983, 51, 823–833. [Google Scholar] [CrossRef]
- Rennenberg, H.; Schupp, R.; Glavac, V.; Jochheim, H. Xylem sap composition of beech (Fagus sylvatica L.) trees: Seasonal changes in the axial distribution of sulfur compounds. Tree Physiol. 1994, 14, 541–548. [Google Scholar] [CrossRef]
- Rellán-Álvarez, R.; El-Jendoubi, H.; Wohlgemuth, G.; Abadia, A.; Fiehn, O.; Abadía, J.; Álvarez-Fernández, A. Metabolite profile changes in xylem sap and leaf extracts of strategy I plants in response to iron deficiency and resupply. Front. Plant Sci. 2011, 2, 66. [Google Scholar] [CrossRef]
- Lima, M.R.M.; Machado, A.F.; Gubler, W.D. Metabolomic Study of Chardonnay Grapevines Double Stressed with Esca-Associated Fungi and Drought. Phytopathology 2017, 107, 669–680. [Google Scholar] [CrossRef]
- Gallinger, J.; Gross, J. Unraveling the Host Plant Alternation of Cacopsylla pruni—Adults but Not Nymphs Can Survive on Conifers Due to Phloem/Xylem Composition. Front. Plant Sci. 2018, 9, 484. [Google Scholar] [CrossRef]
- Sofo, A.; Fausto, C.; Mininni, A.N.; Dichio, B.; Lucini, L. Soil management type differentially modulates the metabolomic profile of olive xylem sap. Plant Physiol. Biochem. 2019, 139, 707–714. [Google Scholar] [CrossRef]
- McCully, M.E. Niches for bacterial endophytes in crop plants: A plant biologist’s view. Funct. Plant Biol. 2001, 28, 983. [Google Scholar] [CrossRef]
- Anguita-Maeso, M.; Olivares-García, C.; Haro, C.; Imperial, J.; Navas-Cortés, J.A.; Landa, B.B. Culture-Dependent and Culture-Independent Characterization of the Olive Xylem Microbiota: Effect of Sap Extraction Methods. Front. Plant Sci. 2020, 10, 1708. [Google Scholar] [CrossRef]
- Anguita-Maeso, M.; Trapero-Casas, J.L.; Olivares-García, C.; Ruano-Rosa, D.; Palomo-Ríos, E.; Jiménez-Díaz, R.M.; Navas-Cortés, J.A.; Landa, B.B. Verticillium dahliae Inoculation and in vitro Propagation Modify the Xylem Microbiome and Disease Reaction to Verticillium Wilt in a Wild Olive Genotype. Front. Plant Sci. 2021, 12, 250. [Google Scholar] [CrossRef] [PubMed]
- Haro, C.; Anguita-Maeso, M.; Metsis, M.; Navas-Cortés, J.A.; Landa, B.B. Evaluation of Established Methods for DNA Extraction and Primer Pairs Targeting 16S rRNA Gene for Bacterial Microbiota Profiling of Olive Xylem Sap. Front. Plant Sci. 2021, 12, 296. [Google Scholar] [CrossRef]
- Deyett, E.; Roper, M.C.; Ruegger, P.; Yang, J.-I.; Borneman, J.; Rolshausen, P.E. Microbial Landscape of the Grapevine Endosphere in the Context of Pierce’s Disease. Phytobiomes J. 2017, 1, 138–149. [Google Scholar] [CrossRef]
- Fausto, C.; Mininni, A.N.; Sofo, A.; Crecchio, C.; Scagliola, M.; Dichio, B.; Xiloyannis, C. Olive orchard microbiome: Characterisation of bacterial communities in soil-plant compartments and their comparison between sustainable and conventional soil management systems. Plant Ecol. Divers. 2018, 11, 597–610. [Google Scholar] [CrossRef]
- Deyett, E.; Rolshausen, P.E. Temporal Dynamics of the Sap Microbiome of Grapevine Under High Pierce’s Disease Pressure. Front. Plant Sci. 2019, 10, 1246. [Google Scholar] [CrossRef] [PubMed]
- Zicca, S.; De Bellis, P.; Masiello, M.; Saponari, M.; Saldarelli, P.; Boscia, D.; Sisto, A. Antagonistic activity of olive endophytic bacteria and of Bacillus spp. strains against Xylella fastidiosa. Microbiol. Res. 2020, 236, 126467. [Google Scholar] [CrossRef] [PubMed]
- Hong, C.E.; Park, J.M. Endophytic bacteria as biocontrol agents against plant pathogens: Current state-of-the-art. Plant Biotechnol. Rep. 2016, 10, 353–357. [Google Scholar] [CrossRef]
- Giampetruzzi, A.; Baptista, P.; Morelli, M.; Cameirão, C.; Neto, T.L.; Costa, D.; D’Attoma, G.; Kubaa, R.A.; Altamura, G.; Saponari, M.; et al. Differences in the Endophytic Microbiome of Olive Cultivars Infected by Xylella Fastidiosa across Seasons. Pathogens 2020, 9, 723. [Google Scholar] [CrossRef]
- Barranco, D. Varieties and Rootstocks. In Olive Grow; Barranco, D., Fernández, E.-R., Escobar, R., Rallo, L., Eds.; Junta de Andalucía/Mundi Prensa/RIRDC/AOA: Pendle Hill, Australia, 2010. [Google Scholar]
- de la Rosa, G.; Peralta-Videa, J.R.; Montes, M.; Parsons, J.G.; Cano-Aguilera, I.; Gardea-Torresdey, J.L. Cadmium uptake and translocation in tumbleweed (Salsola kali), a potential Cd-hyperaccumulator desert plant species: ICP/OES and XAS studies. Chemosphere 2004, 55, 1159–1168. [Google Scholar] [CrossRef]
- Akhtar, S.S.; Andersen, M.N.; Liu, F. Biochar Mitigates Salinity Stress in Potato. J. Agron. Crop. Sci. 2015, 201, 368–378. [Google Scholar] [CrossRef]
- Kassambara, A. Rstatix: Pipe-Friendly Framework for Basic Statistical Tests. R Package Version 0.4.0. 2020. Available online: https://cran.r-project.org/web/packages/rstatix/index.html (accessed on 1 March 2021).
- Fox, J.; Weisberg, S. An R Companion to Applied Regression, 3rd ed.; Sage: Thousand Oaks, CA, USA, 2019; Available online: https://socialsciences.mcmaster.ca/jfox/Books/Companion/ (accessed on 1 March 2021).
- Lee, L.C.; Liong, C.-Y.; Jemain, A.A. Partial least squares-discriminant analysis (PLS-DA) for classification of high-dimensional (HD) data: A review of contemporary practice strategies and knowledge gaps. Analyst 2018, 143, 3526–3539. [Google Scholar] [CrossRef]
- Chong, J.; Wishart, D.S.; Xia, J. Using MetaboAnalyst 4.0 for Comprehensive and Integrative Metabolomics Data Analysis. Curr. Protoc. Bioinform. 2019, 68, e86. [Google Scholar] [CrossRef]
- Callahan, B.J.; Mcmurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef] [PubMed]
- Mangiafico, S. Rcompanion: Functions to Support Extension Education Program Evaluation. R Package Version 2.3.25. Available online: https://CRAN.R-project.org/package=rcompanion (accessed on 3 May 2021).
- Garcia, E.J.; McDowell, T.; Ketola, C.; Jennings, M.; Miller, J.D.; Renaud, J.B. Metabolomics reveals chemical changes in Acer saccharum sap over a maple syrup production season. PLoS ONE 2020, 15, e0235787. [Google Scholar] [CrossRef] [PubMed]
- Sung, J.; Sonn, Y.; Lee, Y.; Kang, S.; Ha, S.; Krishnan, H.B.; Oh, T.-K. Compositional changes of selected amino acids, organic acids, and soluble sugars in the xylem sap of N, P, or K-deficient tomato plants. J. Plant Nutr. Soil Sci. 2015, 178, 792–797. [Google Scholar] [CrossRef]
- Hare, P.D.; Cress, W.A.; Van Staden, J. Dissecting the roles of osmolyte accumulation during stress. Plant Cell Environ. 1998, 21, 535–553. [Google Scholar] [CrossRef]
- Patel, T.K.; Williamson, J.D. Mannitol in Plants, Fungi, and Plant–Fungal Interactions. Trends Plant Sci. 2016, 21, 486–497. [Google Scholar] [CrossRef] [PubMed]
- Jlilat, A.; Ragone, R.; Gualano, S.; Santoro, F.; Gallo, V.; Varvaro, L.; Mastrorilli, P.; Saponari, M.; Nigro, F.; D’Onghia, A.M. A non-targeted metabolomics study on Xylella fastidiosa infected olive plants grown under controlled conditions. Sci. Rep. 2021, 11, 1–11. [Google Scholar] [CrossRef]
- Huang, H.; Thu, T.N.T.; He, X.; Gravot, A.; Bernillon, S.; Ballini, E.; Morel, J.-B. Increase of Fungal Pathogenicity and Role of Plant Glutamine in Nitrogen-Induced Susceptibility (NIS) To Rice Blast. Front. Plant Sci. 2017, 8, 265. [Google Scholar] [CrossRef]
- Seabra, A.R.; Carvalho, H. Inhibition of glutamine synthetase leads to a fast transcriptional activation of defense responses in root nodules. Model Legume Medicago Truncatula 2019, 665–673. [Google Scholar] [CrossRef]
- Wei, J.; Wang, S.; Pei, D.; Qu, L.; Liangjing, Q.; Chen, J.; Di, D.; Gao, K. Antibacterial Activity of Hydroxytyrosol Acetate from Olive Leaves (Olea Europaea L.). Nat. Prod. Res. 2018, 32, 1967–1970. [Google Scholar] [CrossRef]
- De Souza, L.P.; Garbowicz, K.; Brotman, Y.; Tohge, T.; Fernie, A.R. The Acetate Pathway Supports Flavonoid and Lipid Biosynthesis in Arabidopsis. Plant Physiol. 2020, 182, 857–869. [Google Scholar] [CrossRef]
- Putra, D.P.; Botton, B. Organic and inorganic compounds of xylem exudates from five woody plants at the stage of bud breaking. J. Plant Physiol. 1998, 153, 670–676. [Google Scholar] [CrossRef]
- Xu, X.; Du, X.; Wang, F.; Sha, J.; Chen, Q.; Tian, G.; Zhu, Z.; Ge, S.; Jiang, Y. Effects of Potassium Levels on Plant Growth, Accumulation and Distribution of Carbon, and Nitrate Metabolism in Apple Dwarf Rootstock Seedlings. Front. Plant Sci. 2020, 11, 904. [Google Scholar] [CrossRef] [PubMed]
- Puzanskiy, R.K.; Yemelyanov, V.V.; Shavarda, A.L.; Gavrilenko, T.A.; Shishova, M.F. Age- and Organ-Specific Differences of Potato (Solanum phureja) Plants Metabolome. Russ. J. Plant Physiol. 2018, 65, 813–823. [Google Scholar] [CrossRef]
- Yun, D.-Y.; Kang, Y.-G.; Kim, E.-H.; Kim, M.; Park, N.-H.; Choi, H.-T.; Go, G.H.; Lee, J.H.; Park, J.S.; Hong, Y.-S. Metabolomics approach for understanding geographical dependence of soybean leaf metabolome. Food Res. Int. 2018, 106, 842–852. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.-J.; Joo, S.C.; Shi, J.; Hu, C.; Quan, S.; Hu, J.; Sukweenadhi, J.; Mohanan, P.; Yang, D.-C.; Zhang, D. Metabolic dynamics and physiological adaptation of Panax ginseng during development. Plant Cell Rep. 2017, 37, 393–410. [Google Scholar] [CrossRef] [PubMed]
- Lawal, U.; Mediani, A.; Maulidiani, H.; Shaari, K.; Ismail, I.S.; Khatib, A.; Abas, F. Metabolite profiling of Ipomoea aquatica at different growth stages in correlation to the antioxidant and α-glucosidase inhibitory activities elucidated by 1H NMR-based metabolomics. Sci. Hortic. 2015, 192, 400–408. [Google Scholar] [CrossRef]
- Zheng, L.; Wang, M.; Ibarra-Estrada, E.; Wu, C.; Wilson, E.G.; Verpoorte, R.; Klinkhamer, P.G.L.; Choi, Y.H. Investigation of Chemomarkers of Astragali Radix of Different Ages and Geographical Origin by NMR Profiling. Molecules 2015, 20, 3389–3405. [Google Scholar] [CrossRef] [PubMed]
- Ossipov, V.; Ossipova, S.; Bykov, V.; Oksanen, E.; Koricheva, J.; Haukioja, E. Application of metabolomics to genotype and phenotype discrimination of birch trees grown in a long-term open-field experiment. Metabolomics 2007, 4, 39–51. [Google Scholar] [CrossRef]
- Tan, L.; Jin, Z.; Ge, Y.; Nadeem, H.; Cheng, Z.; Azeem, F.; Zhan, R. Comprehensive ESI-Q TRAP-MS/MS based characterization of metabolome of two mango (Mangifera indica L) cultivars from China. Sci. Rep. 2020, 10, 20017. [Google Scholar] [CrossRef] [PubMed]
- Vasmatkar, P.; Kaur, K.; Pannu, P.; Kaur, G.; Kaur, H. Unraveling the metabolite signatures of maize genotypes showing differential response towards southern corn leaf blight by 1H-NMR and FTIR spectroscopy. Physiol. Mol. Plant Pathol. 2019, 108, 101441. [Google Scholar] [CrossRef]
- Billet, K.; Houillé, B.; De Bernonville, T.D.; Besseau, S.; Oudin, A.; Courdavault, V.; Delanoue, G.; Guérin, L.; Clastre, M.; Giglioli-Guivarc’H, N.; et al. Field-Based Metabolomics of Vitis vinifera L. Stems Provides New Insights for Genotype Discrimination and Polyphenol Metabolism Structuring. Front. Plant Sci. 2018, 9, 798. [Google Scholar] [CrossRef] [PubMed]
- Bundy, J.; Davey, M.P.; Viant, M.R. Environmental metabolomics: A critical review and future perspectives. Metabolomics 2009, 5, 3–21. [Google Scholar] [CrossRef]
- Rosati, A.; Cafiero, C.; Paoletti, A.; Alfei, B.; Caporali, S.; Casciani, L.; Valentini, M. Effect of agronomical practices on carpology, fruit and oil composition, and oil sensory properties, in olive (Olea europaea L.). Food Chem. 2014, 159, 236–243. [Google Scholar] [CrossRef] [PubMed]
- Guodong, R.; Xiaoxia, L.; Weiwei, Z.; Wenjun, W.; Jianguo, Z. Metabolomics reveals variation and correlation among different tissues of olive (Olea europaea L.). Biol. Open 2017, 6, 1317–1323. [Google Scholar] [CrossRef]
- Merchak, N.; El Bacha, E.; Khouzam, R.B.; Rizk, T.; Akoka, S.; Bejjani, J. Geoclimatic, morphological, and temporal effects on Lebanese olive oils composition and classification: A 1H NMR metabolomic study. Food Chem. 2017, 217, 379–388. [Google Scholar] [CrossRef]
- Rongai, D.; Sabatini, N.; Del Coco, L.; Perri, E.; Del Re, P.; Simone, N.; Marchegiani, D.; Fanizzi, F.P. 1H NMR and Multivariate Analysis for Geographic Characterization of Commercial Extra Virgin Olive Oil: A Possible Correlation with Climate Data. Foods 2017, 6, 96. [Google Scholar] [CrossRef]
- Culeddu, N.; Chessa, M.; Bandino, G.; Sedda, P.; Zurru, R.; Anedda, R.; Motroni, A.; Molinu, M.G.; Dettori, S.; Santona, M. Classification of Monovarietal Sardinian Extra Virgin Olive Oils by 1 H NMR Metabolomic. Eur. J. Lipid Sci. Technol. 2017, 119, 1700035. [Google Scholar] [CrossRef]
- Fausto, C.; Araniti, F.; Mininni, A.N.; Crecchio, C.; Scagliola, M.; Bleve, G.; Dichio, B.; Sofo, A. Differential olive grove management regulates the levels of primary metabolites in xylem sap. Plant Soil 2021, 460, 281–296. [Google Scholar] [CrossRef]
- Kim, H.K.; Choi, Y.H.; Verpoorte, R. NMR-based metabolomic analysis of plants. Nat. Protoc. 2010, 5, 536–549. [Google Scholar] [CrossRef]
- Mahrous, E.A.; Farag, M.A. Two dimensional NMR spectroscopic approaches for exploring plant metabolome: A review. J. Adv. Res. 2015, 6, 3–15. [Google Scholar] [CrossRef]
- Müller, H.; Berg, C.; Landa, B.B.; Auerbach, A.; Moissl-Eichinger, C.; Berg, G. Plant genotype-specific archaeal and bacterial endophytes but similar Bacillus antagonists colonize Mediterranean olive trees. Front. Microbiol. 2015, 6, 138. [Google Scholar] [CrossRef]
- Mercado-Blanco, J.; RodrÍguez-Jurado, D.; Hervás, A.; Jiménez-DÍaz, R.M. Suppression of Verticillium wilt in olive planting stocks by root-associated fluorescent Pseudomonas spp. Biol. Control 2004, 30, 474–486. [Google Scholar] [CrossRef]
- Weller, D.M. Pseudomonas Biocontrol Agents of Soilborne Pathogens: Looking Back Over 30 Years. Phytopathology 2007, 97, 250–256. [Google Scholar] [CrossRef] [PubMed]
- Loper, J.E.; Hassan, K.A.; Mavrodi, D.; Ii, E.W.D.; Lim, C.K.; Shaffer, B.T.; Elbourne, L.D.H.; Stockwell, V.O.; Hartney, S.L.; Breakwell, K.; et al. Comparative Genomics of Plant-Associated Pseudomonas spp.: Insights into Diversity and Inheritance of Traits Involved in Multitrophic Interactions. PLoS Genet. 2012, 8, e1002784. [Google Scholar] [CrossRef] [PubMed]
- Ofek, M.; Hadar, Y.; Minz, D. Ecology of Root Colonizing Massilia (Oxalobacteraceae). PLoS ONE 2012, 7, e40117. [Google Scholar] [CrossRef] [PubMed]
- López-Escudero, F.; Del Río, C.; Caballero, J.; Blanco-López, M. Evaluation of Olive Cultivars for Resistance to Verticillium dahliae. Eur. J. Plant Pathol. 2004, 110, 79–85. [Google Scholar] [CrossRef]
- Martos-Moreno, C.; López-Escudero, F.J.; Blanco-López, M.A. Resistance of Olive Cultivars to the Defoliating Pathotype of Verticillium dahliae. HortScience 2006, 41, 1313–1316. [Google Scholar] [CrossRef]
- D’Attoma, G.; Morelli, M.; Saldarelli, P.; Saponari, M.; Giampetruzzi, A.; Boscia, D.; Savino, V.N.; De La Fuente, L.; Cobine, P.A. Ionomic Differences between Susceptible and Resistant Olive Cultivars Infected by Xylella fastidiosa in the Outbreak Area of Salento, Italy. Pathogens 2019, 8, 272. [Google Scholar] [CrossRef]
- Vergine, M.; Meyer, J.B.; Cardinale, M.; Sabella, E.; Hartmann, M.; Cherubini, P.; De Bellis, L.; Luvisi, A. The Xylella fastidiosa-Resistant Olive Cultivar “Leccino” Has Stable Endophytic Microbiota during the Olive Quick Decline Syndrome (OQDS). Pathogens 2019, 9, 35. [Google Scholar] [CrossRef] [PubMed]
- Trapero, C.; Díez, C.M.; Rallo, L.; Barranco, D.; López-Escudero, F.J. Effective inoculation methods to screen for resistance to Verticillium wilt in olive. Sci. Hortic. 2013, 162, 252–259. [Google Scholar] [CrossRef]
- Luvisi, A.; Aprile, A.; Sabella, E.; Vergine, M.; Nicolì, F.; Nutricati, E.; Miceli, A.; Negro, C.; Bellis, L. De Xylella fastidiosa subsp. pauca (CoDiRO strain) infection in four olive (Olea europaea L.) cultivars: Profile of phenolic compounds in leaves and progression of leaf scorch symptoms. Phytopathol. Mediterr. 2017, 56, 259–273. [Google Scholar] [CrossRef]
- Girelli, C.R.; Del Coco, L.; Scortichini, M.; Petriccione, M.; Zampella, L.; Mastrobuoni, F.; Cesari, G.; Bertaccini, A.; D’Amico, G.; Contaldo, N.; et al. Xylella fastidiosa and olive quick decline syndrome (CoDiRO) in Salento (southern Italy): A chemometric 1H NMR-based preliminary study on Ogliarola salentina and Cellina di Nardò cultivars. Chem. Biol. Technol. Agric. 2017, 4, 1–9. [Google Scholar] [CrossRef]
- Girelli, C.R.; Angilè, F.; Del Coco, L.; Migoni, D.; Zampella, L.; Marcelletti, S.; Cristella, N.; Marangi, P.; Scortichini, M.; Fanizzi, F.P. 1H-NMR Metabolite Fingerprinting Analysis Reveals a Disease Biomarker and a Field Treatment Response in Xylella fastidiosa subsp. pauca-Infected Olive Trees. Plants 2019, 8, 115. [Google Scholar] [CrossRef] [PubMed]
- Markakis, E.A.; Tjamos, S.E.; Antoniou, P.P.; Roussos, P.A.; Paplomatas, E.J.; Tjamos, E.C. Phenolic Responses of Resistant and Susceptible Olive Cultivars Induced by Defoliating and Nondefoliating Verticillium dahliae Pathotypes. Plant Dis. 2010, 94, 1156–1162. [Google Scholar] [CrossRef] [PubMed]
| Metabolite | Min–Max | Adult Trees | Plantlets | ANOVA | ||||
|---|---|---|---|---|---|---|---|---|
| Arbequina | Picual | Arbequina | Picual | Age (A) | Genotype (G) | A × G | ||
| Organic Acids | ||||||||
| Acetate | 97.9–264.6 | 134.93 ± 14.05 (4/4) | 127.28 ± 14.29 (4/4) | 137.20 ± 30.15 (4/4) | 142.73 ± 81.42 (4/4) | 0.925 | 0.715 | 0.951 |
| Formic Acid | 16.0–36.0 | 22.43 ± 5.39 (4/4) | 16.98 ± 1.62 (4/4) | 24.70 ± 9.49 (4/4) | 18.73 ± 1.54 (4/4) | 0.509 | 0.065 | 0.883 |
| Fumaric Acid | 1.6–17.4 | 5.70 ± 4.30 (4/4) | 11.25 ± 4.95 (4/4) | 7.68 ± 1.64 (4/4) | 6.25 ± 2.23 (4/4) | 0.991 | 0.270 | 0.065 |
| Maleic Acid | 9.8–29.4 | 26.30 ± 3.80 (4/4) | 15.40 ± 2.77 (4/4) | 16.93 ± 8.62 (4/4) | 18.20 ± 5.41 (4/4) | 0.229 | 0.195 | 0.042 |
| Succinic Acid | 1.6–29.5 | 6.98 ± 5.00 (4/4) | 10.83 ± 6.14 (4/4) | 6.18 ± 3.47 (4/4) | 10.08 ± 13.17 (4/4) | 0.446 | 0.576 | 0.551 |
| Aminoacids | ||||||||
| Alanine | 2.7–11.5 | 4.93 ± 1.12 (4/4) | 4.35 ± 0.94 (4/4) | 7.53 ± 3.43 (3/4) | 5.75 ± 2.60 (4/4) | 0.873 | 0.918 | 0.607 |
| Arginine | 16.1–115.7 | 106.40 ± 0.00 (1/4) | 30.67 ± 26.71 (3/4) | 25.00 ± 0.00 (1/4) | 69.80 ± 64.91 (2/4) | 0.712 | 0.262 | 0.982 |
| Asparagine | 23.6–36.9 | nd * (0/4) | 32.17 ± 5.39 (3/4) | 36.20 ± 0.00 (1/4) | 28.85 ± 7.42 (2/4) | 0.987 | 0.054 | 0.272 |
| Aspartate | 14.7–137.9 | 34.90 ± 0.00 (1/4) | 48.07 ± 54.61 (3/4) | 23.50 ± 7.35 (2/4) | 53.05 ± 57.33 (4/4) | 0.345 | 0.050 | 0.833 |
| GABA | 4.4–24.9 | 8.00 ± 5.09 (2/4) | 18.70 ± 7.49 (2/4) | nd (0/4) | 24.90 ± 0.00 (1/4) | 0.239 | 0.354 | 0.798 |
| Glutamate | 4.5–856.1 | 21.10 ± 10.15 (3/4) | 84.75 ± 71.41 (4/4) | 36.83 ± 26.74 (3/4) | 224.63 ± 421.06 (4/4) | 0.858 | 0.115 | 0.563 |
| Glutamine | 12.7–2482.1 | 142.55 ± 90.44 (4/4) | 63.33 ± 45.89 (4/4) | 510.57 ± 474.87 (3/4) | 672.90 ± 1208.3 (4/4) | 0.990 | 0.981 | 0.513 |
| Isoleucine | 2.4–9.2 | 3.43 ± 1.03 (4/4) | 5.03 ± 2.88 (4/4) | 4.07 ± 1.81 (3/4) | 4.47 ± 2.54 (3/4) | 0.326 | 0.585 | 0.683 |
| Leucine | 1.4–6.3 | 3.15 ± 2.26 (4/4) | 3.20 ± 1.06 (4/4) | 2.37 ± 0.85 (3/4) | 3.63 ± 2.19 (3/4) | 0.339 | 0.483 | 0.865 |
| Methionine | 3.7–17.8 | 7.05 ± 4.73 (2/4) | 12.85 ± 1.06 (2/4) | 13.70 ± 5.79 (2/4) | 7.15 ± 0.21 (2/4) | 0.953 | 0.964 | 0.625 |
| Phenylalanine | 3.9–7.8 | nd (0/4) | 5.55 ± 1.65 (4/4) | 7.37 + 0.23 (3/4) | 5.30 ± 1.69 (2/4) | 0.388 | 0.182 | 0.006 |
| Proline | 6.3–402.1 | 12.00 ± 0.00 (1/4) | 53.30 ± 15.27 (2/4) | 26.25 ± 70.63 (2/4) | 140.10 ± 226.90 (3/4) | 0.456 | 0.292 | 0.859 |
| Threonine | 5.3–186.1 | 15.55 ± 3.15 (4/4) | 43.68 ± 22.58 (4/4) | 90.43 ± 78.78 (4/4) | 41.58 ± 27.60 (4/4) | 0.304 | 0.771 | 0.125 |
| Tyrosine | 4.5–17.7 | 8.97 ± 0.40 (3/4) | 7.65 ± 2.81 (4/4) | 12.83 ± 3.40 (4/4) | 10.95 ± 5.20 (4/4) | 0.090 | 0.868 | 0.403 |
| Valine | 2.5–16.4 | 8.73 ± 6.53 (4/4) | 5.98 ± 1.54 (4/4) | 6.10 ± 1.94 (4/4) | 6.08 ± 3.57 (4/4) | 0.675 | 0.653 | 0.952 |
| Sugars | ||||||||
| Glucose | 45.2–752.9 | 333.43 ± 162.12 (4/4) | 443.83 ± 216.31 (4/4) | 145.13 ± 77.58 (4/4) | 216.13 ± 132.94 (4/4) | 0.029 | 0.378 | 0.918 |
| Fructose | 39.2–634.6 | 222.75 ± 130.41 (4/4) | 387.38 ± 187.14 (4/4) | 110.78 ± 129.78 (4/4) | 153.50 ± 141.01 (4/4) | 0.037 | 0.170 | 0.773 |
| Sucrose | 25.4–220.1 | 145.35 ± 45.20 (4/4) | 157.15 ± 46.49 (4/4) | 89.18 ± 32.03 (4/4) | 52.45 ± 28.95 (4/4) | 0.001 | 0.219 | 0.113 |
| myo-Inositol | 14.1–90.2 | 42.53 ± 2.68 (3/4) | 53.60 ± 35.38 (4/4) | 32.88 ± 11.15 (4/4) | 44.53 ± 20.85 (4/4) | 0.576 | 0.278 | 0.537 |
| Mannitol | 72.6–1334.0 | 839.73 ± 76.19 (4/4) | 1032.9 ± 202.75 (4/4) | 370.70 ± 256.06 (4/4) | 564.03 ± 251.7 (4/4) | 0.010 | 0.204 | 0.513 |
| Osmolytes | ||||||||
| Choline | 2.1–7.2 | 4.15 ± 0.37 (4/4) | 4.90 ± 2.16 (4/4) | 3.95 ± 3.60 (2/4) | 3.53 ± 2.28 (3/4) | 0.038 | 0.599 | 0.760 |
| Sarcosine | 2.2–13.1 | 3.20 ± 1.11 (3/4) | 7.03 ± 4.85 (4/4) | 3.20 ± 2.26 (2/4) | 4.20 ± 2.25 (3/4) | 0.167 | 0.095 | 0.566 |
| Trigonelline | 5.4–22.8 | 11.80 ± 7.68 (4/4) | 9.73 ± 3.15 (4/4) | 7.23 ± 1.68 (4/4) | 8.50 ± 0.82 (4/4) | 0.231 | 0.817 | 0.474 |
| Alcohols | ||||||||
| Ethanol | 29.0–606.7 | 43.43 ± 6.42 (4/4) | 144.50 ± 161.52 (4/4) | 409.45 ± 162.01 (4/4) | 163.55 ± 142.90 (4/4) | 0.008 | 0.609 | 0.018 |
| Methanol | 35–276.4 | 54.88 ± 13.32 (4/4) | 81.10 ± 64.90 (4/4) | 147.53 ± 34.86 (4/4) | 107.93 ± 112.69 (4/4) | 0.071 | 0.492 | 0.180 |
| Compound | Min–Max | Adult Trees | Plantlets | ANOVA | ||||
|---|---|---|---|---|---|---|---|---|
| Arbequina | Picual | Arbequina | Picual | Age (A) | Genotype (G) | A × G | ||
| Element | ||||||||
| Al | 5.4–41.4 | 9.63 ± 5.94 (2/4) | 33.22 ± 11.61 (2/4) | 13.05 ± 3.82 (2/4) | 12.38 ± 8.77 (3/4) | 0.825 | 0.517 | 0.968 |
| As | 0.1–0.5 | 0.21 ± 0.08 (2/4) | nd (0/4) | 0.41 ± 0.00 (1/4) | 0.05 ± 0.00 (1/4) | 0.373 | 0.538 | 0.536 |
| B | 1.4–67.9 | 49.41 ± 15.68 (4/4) | 28.19 ± 10.03 (4/4) | 12.78 ± 7.37 (4/4) | 10.32 ± 7.78 (3/4) | 0.002 | 0.089 | 0.630 |
| Ca | 104.0–1208.0 | 465.31 ± 42.70 (4/4) | 372.00 ± 55.43 (4/4) | 222.61 ± 216.07 (4/4) | 452.08 ± 517.03 (4/4) | 0.086 | 0.694 | 0.306 |
| Cu | 0.3–1.6 | 1.20 ± 0.35 (4/4) | 0.91 ± 0.09 (4/4) | 1.26 ± 0.63 (3/4) | 0.70 ± 0.28 (4/4) | 0.195 | 0.544 | 0.807 |
| Fe | 0.3–1.2 | 1.03 ± 0.21 (4/4) | 0.37 ± 0.02 (3/4) | 0.47 ± 0.00 (1/4) | 0.58 ± 0.00 (1/4) | 0.007 | 0.022 | 0.044 |
| K | 1246.6–3571.2 | 1743.8 ± 160.4 (4/4) | 1808.8 ± 320.31 (4/4) | 1556.3 ± 277.21 (4/4) | 2371.4 ± 1056.8 (4/4) | 0.765 | 0.177 | 0.244 |
| Mg | 50.4–879.4 | 90.00 ± 9.33 (4/4) | 92.84 ± 16.94 (4/4) | 156.56 ± 173.11 (4/4) | 316.95 ± 388.10 (4/4) | 0.347 | 0.570 | 0.607 |
| Mn | 0.4–4.3 | 0.89 ± 0.05 (4/4) | 0.46 ± 0.12 (4/4) | 1.53 ± 0.00 (1/4) | 4.33 ± 0.00 (1/4) | 0.435 | 0.991 | 0.175 |
| Na | 156.5–1281.1 | 410.28 ± 43.45 (4/4) | 709.86 ± 22.76 (4/4) | 365.80 ± 178.67 (4/4) | 666.55 ± 433.10 (4/4) | 0.332 | 0.024 | 0.997 |
| Ni | 0.2–0.7 | nd * (0/4) | 0.33 ± 0.00 (1/4) | 0.32 ± 0.00 (1/4) | 0.50 ± 0.40 (2/4) | 0.538 | 0.504 | 0.991 |
| P | 34.7–556.0 | 360.25 ± 162.99 (4/4) | 198.94 ± 25.91 (4/4) | 204.80 ± 155.74 (4/4) | 235.52 ± 66.57 (4/4) | 0.652 | 0.805 | 0.153 |
| Pb | 0.07–0.1 | 0.07 ± 0.00 (1/4) | nd (0/4) | nd (0/4) | 0.10 ± 0.00 (1/4) | 0.298 | 0.889 | 0.144 |
| Rb | 0.2–0.8 | 0.39 ± 0.01 (4/4) | 0.35 ± 0.08 (3/4) | 0.62 ± 0.20 (4/4) | 0.59 ± 0.14 (4/4) | 0.006 | 0.266 | 0.357 |
| S | 48.6–2330.9 | 249.43 ± 45.61 (4/4) | 194.87 ± 18.93 (4/4) | 341.73 ± 367.40 (4/4) | 796.26 ± 1075.4 (4/4) | 0.746 | 0.971 | 0.815 |
| Se | 0.1–0.3 | 0.13 ± 0.00 (1/4) | 0.30 ± 0.00 (1/4) | nd (0/4) | nd (0/4) | 0.848 | 0.913 | 0.751 |
| Sr | 0.3–10.3 | 9.02 ± 1.04 (4/4) | 6.24 ± 0.99 (4/4) | 0.85 ± 0.35 (4/4) | 2.75 ± 1.55 (4/4) | <0.001 | 0.077 | 0.003 |
| Zn | 0.3–1.3 | 0.77 ± 0.10 (4/4) | 0.58 ± 0.07 (4/4) | 0.44 ± 0.08 (4/4) | 0.91 ± 0.46 (3/4) | 0.233 | 0.872 | 0.377 |
| Inorganic Anions | ||||||||
| Cl− | 2098.9–3925.2 | 2422.6 ± 67.18 (4/4) | 2545.2 ± 219.52 (4/4) | 2539.1 ± 668.37 (4/4) | 3048.1 ± 593.46 (4/4) | 0.236 | 0.151 | 0.918 |
| F− | 121.2–295.4 | 245.14 ± 28.32 (4/4) | 245.81 ± 35.64 (4/4) | 195.70 ± 58.43 (4/4) | 213.64 ± 2.98 (4/4) | 0.058 | 0.514 | 0.773 |
| NO3− | 762.8–2581.9 | 941.12 ± 77.57 (4/4) | 787.04 ± 30.56 (4/4) | 1488.7 ± 376.20 (4/4) | 1438.2 ± 822.63 (4/4) | 0.009 | 0.327 | 0.113 |
| PO43− | 741.8–1038.7 | 927.47 ± 104.85 (4/4) | 813.42 ± 17.03 (4/4) | 789.21 ± 60.03 (4/4) | 788.06 ± 30.53 (4/4) | 0.021 | 0.104 | 0.537 |
| SO42− | 1470.2–3005.4 | 1657.8 ± 28.86 (4/4) | 1620.5 ± 14.80 (4/4) | 1694.7 ± 218.43 (4/4) | 1970.4 ± 722.29 (4/4) | 0.370 | 0.629 | 0.513 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Anguita-Maeso, M.; Haro, C.; Montes-Borrego, M.; De La Fuente, L.; Navas-Cortés, J.A.; Landa, B.B. Metabolomic, Ionomic and Microbial Characterization of Olive Xylem Sap Reveals Differences According to Plant Age and Genotype. Agronomy 2021, 11, 1179. https://doi.org/10.3390/agronomy11061179
Anguita-Maeso M, Haro C, Montes-Borrego M, De La Fuente L, Navas-Cortés JA, Landa BB. Metabolomic, Ionomic and Microbial Characterization of Olive Xylem Sap Reveals Differences According to Plant Age and Genotype. Agronomy. 2021; 11(6):1179. https://doi.org/10.3390/agronomy11061179
Chicago/Turabian StyleAnguita-Maeso, Manuel, Carmen Haro, Miguel Montes-Borrego, Leonardo De La Fuente, Juan A. Navas-Cortés, and Blanca B. Landa. 2021. "Metabolomic, Ionomic and Microbial Characterization of Olive Xylem Sap Reveals Differences According to Plant Age and Genotype" Agronomy 11, no. 6: 1179. https://doi.org/10.3390/agronomy11061179
APA StyleAnguita-Maeso, M., Haro, C., Montes-Borrego, M., De La Fuente, L., Navas-Cortés, J. A., & Landa, B. B. (2021). Metabolomic, Ionomic and Microbial Characterization of Olive Xylem Sap Reveals Differences According to Plant Age and Genotype. Agronomy, 11(6), 1179. https://doi.org/10.3390/agronomy11061179






