The Challenge of Environmental Samples for PCR Detection of Phytopathogenic Bacteria: A Case Study of Citrus Huanglongbing Disease
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material
2.2. Insect Material
2.3. DNA Extraction from Plant Material and Insects
2.4. Detection of ‘Ca. Liberibacter’ spp.
2.5. Verification of Positive Samples
2.6. Bioinformatic Analysis of the Sequences
2.7. Phylogenetic Analysis of the Undesirable Amplifications
3. Results
3.1. Analysis of Plant Samples
3.2. Analysis of Insect Samples
3.3. Verification of Positive Samples
3.3.1. Sequence Analyses of the Amplified Fragments
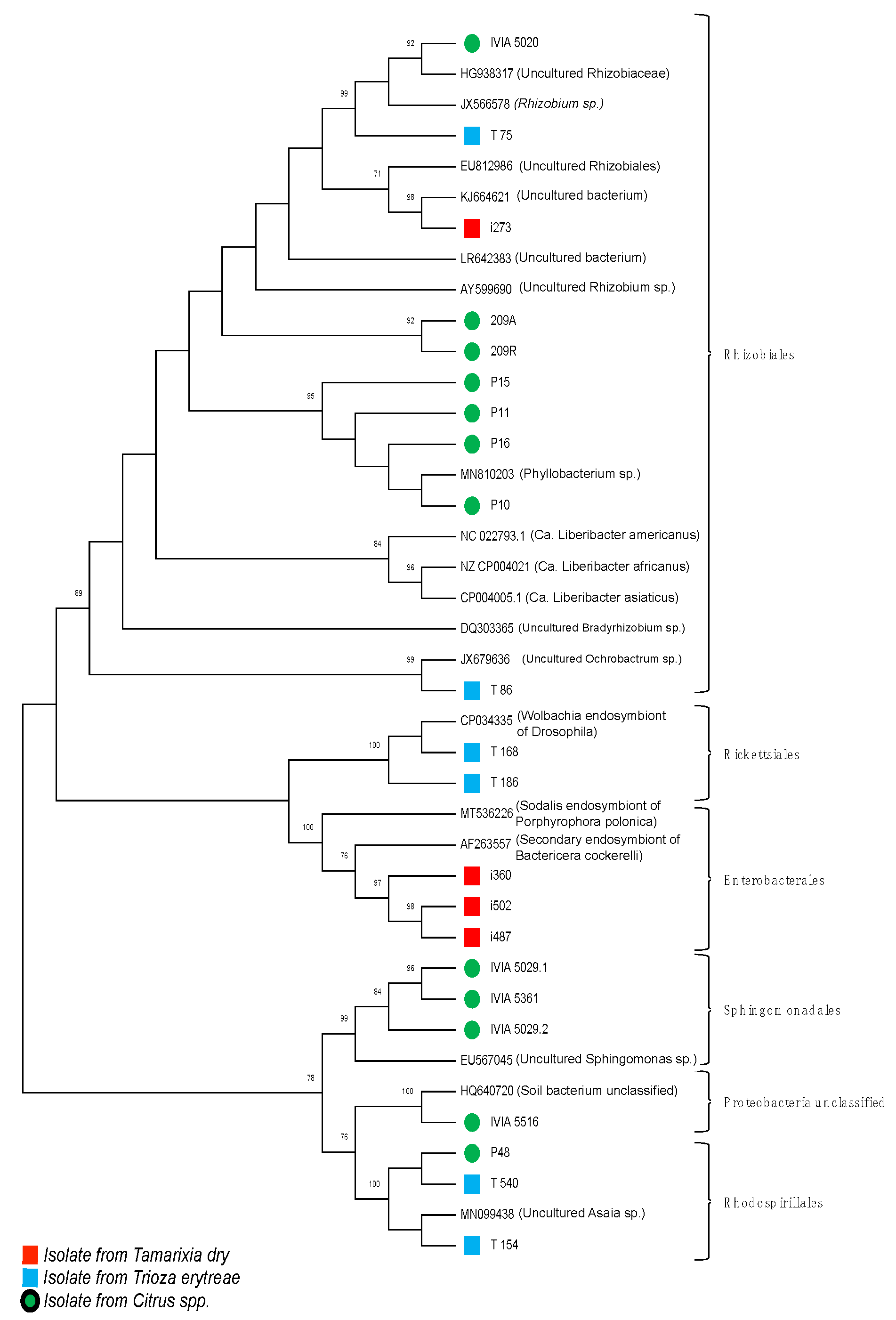
3.3.2. Phylogenetic Analysis of the Amplifications
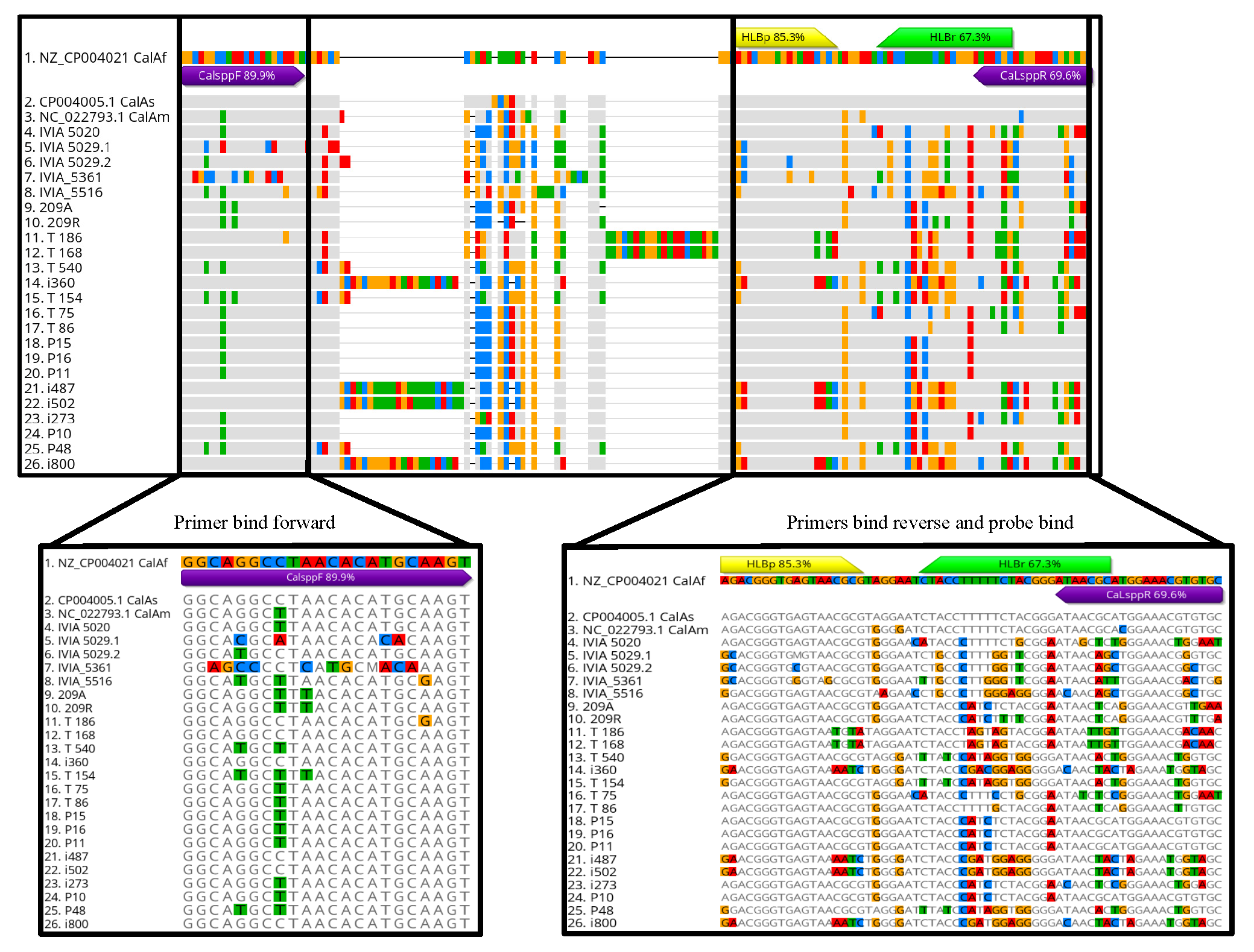
3.3.3. Bioinformatic Analyses of Primers, Probes and Undesired Amplicon Sequences
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Saingam, P.; Li, B.; Yan, T. Use of amplicon sequencing to improve sensitivity in PCR-based detection of microbial pathogen in environmental samples. J. Microbiol. Methods 2018, 149, 73–79. [Google Scholar] [CrossRef] [PubMed]
- Chabbert, E.; Lachaud, L.; Crobu, L.; Bastien, P. Comparison of Two Widely Used PCR Primer systems for detection of toxoplasma in amniotic fluid, blood, and tissues. J. Clin. Microbiol. 2004, 42, 1719–1722. [Google Scholar] [CrossRef] [PubMed]
- Bové, J.M. Huanglongbing: A destructive, newly-emerging, century-old disease of citrus. J. Plant Pathol. 2006, 88, 7–37. [Google Scholar]
- Ferrarezi, R.S.; Vincent, C.I.; Urbaneja, A.; Machado, M.A. Editorial: Unravelling citrus Huanglongbing disease. Front. Plant Sci. 2020, 11, 1635. [Google Scholar] [CrossRef] [PubMed]
- Da Graça, J.V.; Korsten, L. Citrus Huanglongbing: Review, Present status and future strategies. In Diseases of Fruits and Vegetables; Springer: Dordrecht, The Netherlands, 2004; Volume I, pp. 229–245. [Google Scholar]
- Jagoueix, S.; Bové, J.M.; Garnier, M. PCR detection of the two ‘Candidatus Liberibacter’ species associated with greening disease of citrus. Mol. Cell. Probes 1996, 10, 43–50. [Google Scholar] [CrossRef]
- Del Guercio, G. Note ed osservazioni di entomologia agraria. Il cecidio delle foglie del limone ed il suo cecidozoo in Eritrea. L’Agricoltura Coloniale 1918, 12, 355–357. [Google Scholar]
- McClean, A.P.; Oberholzer, P.C. Citrus psylla, a vector of the greening disease, of sweet orange. S. Afr. J. Agric. Sci. 1965, 8, 297–298. [Google Scholar]
- Planet, P.; Jagoueix, S.; Bové, J.M.; Garnier, M. Detection and characterization of the African citrus greening Liberibacter by amplification, cloning, and sequencing of the rplKAJL-rpoBC operon. Curr. Microbiol. 1995, 30, 137–141. [Google Scholar] [CrossRef]
- Yamamoto, P.T.; Felippe, M.R.; Garbim, L.F.; Coelho, J.H.C.; Martins, E.C.; Leite, A.P.R.; Sousa, M.C.; Abrahão, D.P.; Braz, J. Diaphorina citri (Kuwayama) (Hemiptera: Psyllydae): Vector of the bacterium ‘Candidatus Liberibacter americanus’. In Proceedings of the Huanglongbing-Greening International Workshop Fundecitrus, Ribeirão Preto, São Paulo, Brazil, 16–20 July 2006. [Google Scholar]
- Do Carmo Teixeira, D.; Saillard, C.; Eveillard, S.; Danet, J.L.; da Costa, P.I.; Ayres, A.J.; Bové, J. ‘Candidatus Liberibacter americanus’, associated with citrus huanglongbing (greening disease) in São Paulo State, Brazil. Int. J. Syst. Evol. Microbiol. 2005, 55, 1857–1862. [Google Scholar] [CrossRef]
- Belasque, J.; Bassanezi, R.B.; Yamamoto, P.T.; Ayres, A.J.; Tachibana, A.; Violante, A.R.; Tank, A.; Di Giorgi, F.; Tersi, F.E.A.; Menezes, G.M.; et al. Lessons from huanglongbing management in São Paulo state, Brazil. J. Plant Pathol. 2010, 92, 285–302. [Google Scholar]
- Aubert, B. Trioza erytreae Del Guercio and Diaphorina citri Kuwayama (Homoptera: Psylloidea), the two vectors of citrus greening disease: Biological aspects andpossible control strategies. Fruits 1987, 42, 149–162. [Google Scholar]
- Qureshi, J.A.; Rogers, M.E.; Hall, D.G.; Stansly, P.A. Incidence of Invasive (Diaphorina citri) (Hemiptera: Psyllidae) and Its Introduced Parasitoid (Tamarixia radiata) (Hymenoptera: Eulophidae) in Florida Citrus. J. Econ. Entomol. 2009, 102, 247–256. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Rodríguez, J.; Krüger, K.; Pérez-Hedo, M.; Ruíz-Rivero, O.; Urbaneja, A.; Tena, A. Classical biological control of the African citrus psyllid Trioza erytreae, a major threat to the European citrus industry. Sci. Rep. 2019, 9, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Urbaneja-Bernat, P.; Pérez-Rodríguez, J.; Krüger, K.; Catalán, J.; Rizza, R.; Hernández-Suárez, E.; Urbaneja, A.; Tena, A. Host range testing of Tamarixia dryi (Hymenoptera: Eulophidae) sourced from South Africa for classical biological control of Trioza erytreae (Hemiptera: Psyllidae) in Europe. Biol. Control 2019, 135, 110–116. [Google Scholar] [CrossRef]
- Ministerio de Agricultura Alimentación y Medio Ambiente (MAPAMA). Real Decreto 23/2016, de 22 de Enero, por el que se Establece el Programa Nacional de Control y Erradicación de Trioza Erytreae, y el Programa Nacional de Prevención de Diaphorina Citri y ‘Candidatus Liberibacter spp’; BOE: Madrid, Spain, 2016; pp. 6432–6441.
- European and Mediterranean Plant Protection Organization. PM 7/121 (1) ‘Candidatus Liberibacter africanus’, ‘Candidatus Liberibacter americanus’ and ‘Candidatus Liberibacter asiaticus’. EPPO Bull. 2014, 44, 376–389. [Google Scholar] [CrossRef][Green Version]
- Bertolini, E.; Felipe, R.T.A.; Sauer, A.V.; Lopes, S.A.; Arilla, A.; Vidal, E.; Mourão Filho, F.A.A.; Nunes, W.M.C.; Bové, J.M.; López, M.M.; et al. Tissue-print and squash real-time PCR for direct detection of ‘Candidatus Liberibacter’ species in citrus plants and psyllid vectors. Plant Pathol. 2014, 63, 1149–1158. [Google Scholar] [CrossRef]
- Teixeira, D.D.C.; Danet, J.L.; Eveillard, S.; Martins, E.C.; De Jesus, W.C.; Yamamoto, P.T.; Lopes, S.A.; Bassanezi, R.B.; Ayres, A.J.; Saillard, C.; et al. Citrus huanglongbing in São Paulo State, Brazil: PCR detection of the ‘Candidatus Liberibacter’ species associated with the disease. Mol. Cell. Probes 2005, 19, 173–179. [Google Scholar] [CrossRef]
- Li, W.; Hartung, J.S.; Levy, L. Quantitative real-time PCR for detection and identification of ‘Candidatus Liberibacter’ species associated with citrus huanglongbing. J. Microbiol. Methods 2006, 66, 104–115. [Google Scholar] [CrossRef]
- Morgan, J.K.; Zhou, L.; Li, W.; Shatters, R.G.; Keremane, M.; Duan, Y.P. Improved real-time PCR detection of ‘Candidatus Liberibacter asiaticus’ from citrus and psyllid hosts by targeting the intragenic tandem-repeats of its prophage genes. Mol. Cell. Probes 2012, 26, 90–98. [Google Scholar] [CrossRef]
- Bao, M.; Zheng, Z.Z.; Sun, X.; Chen, J.; Deng, X. Enhancing PCR capacity to detect ‘Candidatus Liberibacter asiaticus’ utilizing whole genome sequence information. Plant Dis. 2020, 104, 527–532. [Google Scholar] [CrossRef]
- Ananthakrishnan, G.; Choudhary, N.; Roy, A.; Sengoda, V.G.; Postnikova, E.; Hartung, J.S.; Stone, A.L.; Damsteegt, V.D.; Schneider, W.L.; Munyaneza, J.E.; et al. Development of primers and probes for genus and species specific detection of ‘Candidatus Liberibacter´ species by real-time PCR. Plant Dis. 2013, 97, 1235–1243. [Google Scholar] [CrossRef] [PubMed]
- Okuda, M.; Matsumoto, M.; Tanaka, Y.; Subandiyah, S.; Iwanami, T. Characterization of the tufB-secE-nusG-rplKAJL-rpoB gene cluster of the citrus greening organism and detection by loop-mediated isothermal amplification. Plant Dis. 2005, 89, 705–711. [Google Scholar] [CrossRef] [PubMed]
- Kogenaru, S.; Yan, Q.; Riera, N.; Roper, M.C.; Deng, X.; Ebert, T.A.; Rogers, M.; Irey, M.E.; Pietersen, G.; Rush, C.M.; et al. Repertoire of novel sequence signatures for the detection of ‘Candidatus Liberibacter asiaticus’ by quantitative real-time PCR. BMC Microbiol. 2014, 14, 39. [Google Scholar] [CrossRef] [PubMed]
- Duan, Y.; Zhou, L.; Hall, D.G.; Li, W.; Doddapaneni, H.; Lin, H.; Liu, L.; Vahling, C.M.; Gabriel, D.W.; Williams, K.P.; et al. Complete genome sequence of citrus huanglongbing bacterium, ‘Candidatus Liberibacter asiaticus’ obtained through metagenomics. Mol. Plant-Microbe Interact. 2009, 22, 1011–1020. [Google Scholar] [CrossRef]
- Kim, J.S.; Wang, N. Characterization of copy numbers of 16S rDNA and 16S rRNA of ‘Candidatus Liberibacter asiaticus’ and the implication in detection in planta using quantitative PCR. BMC Res. Notes 2009, 2, 37. [Google Scholar] [CrossRef]
- Cellier, G.; Redondo, C.; Cubero, J.; Roselló, M.; de Andrade, E.; Cruz, L.; Ince, E.; Yildiz, H.N.; Güler, P.G.; D’Onghia, A.M.; et al. Comparison of the performance of the main real-time and conventional PCR detection tests for ‘Candidatus Liberibacter’ spp., plant pathogenic bacteria causing the huanglongbing disease in Citrus spp. Eur. J. Plant Pathol. 2020. [Google Scholar] [CrossRef]
- Kunta, M.; Viloria, Z.; del Rio, H.S.; Louzada, E.S. Diverse DNA extraction methods and PCR primers for detection of huanglongbing-associated bacteria from roots of ‘Valencia’ sweet orange on sour orange rootstock. Sci. Hortic. (Amsterdam) 2014, 178, 23–30. [Google Scholar] [CrossRef]
- Louzada, E.S.; Vazquez, O.E.; Braswell, W.E.; Yanev, G.; Devanaboina, M.; Kunta, M. Distribution of ‘Candidatus Liberibacter asiaticus’ Above and below ground in Texas citrus. Phytopathology 2016, 106, 702–709. [Google Scholar] [CrossRef]
- Zheng, Z.; Xu, M.; Bao, M.; Wu, F.; Chen, J.; Deng, X. Unusual five copies and dual forms of nrdB in ‘Candidatus Liberibacter asiaticus’: Biological Implications and PCR Detection Application. Sci. Rep. 2016, 6, 1–9. [Google Scholar] [CrossRef]
- Folmer, O.; Black, M.; Hoeh, W.; Lutz, R.; Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome C oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 1994, 3, 294–299. [Google Scholar]
- National Center for Biotechnology Information. Available online: https://www.ncbi.nlm.nih.gov/ (accessed on 6 October 2020).
- Hill, J.T.; Demarest, B.L.; Bisgrove, B.W.; Su, Y.C.; Smith, M.; Yost, H.J. Poly peak parser: Method and software for identification of unknown indels using sanger sequencing of polymerase chain reaction products. Dev. Dyn. 2014, 243, 1632–1636. [Google Scholar] [CrossRef]
- Bunn, A.; Korpela, M. A dendrochronology program library in R (dplR9). Dendro 2018, 26, 115–124. [Google Scholar] [CrossRef]
- BLAST: Basic Local Alignment Search Tool. Available online: https://blast.ncbi.nlm.nih.gov/Blast.cgi (accessed on 5 October 2020).
- Edgar, R.C. MUSCLE: A multiple sequence alignment method with reduced time and space complexity. BMC Bioinform. 2004, 5, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Nei, M.; Kumar, S. Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc. Natl. Acad. Sci. USA 2004, 101, 11030–11035. [Google Scholar] [CrossRef] [PubMed]
- López, M.M.; Llop, P.; Olmos, A.; Marco-Noales, E.; Cambra, M.; Bertolini, E. Are molecular tools solving the challenges posed by detection of plant pathogenic bacteria and viruses? Curr. Issues Mol. Biol. 2009, 11, 13–46. [Google Scholar] [CrossRef]
- Louca, S.; Mazel, F.; Doebeli, M.; Parfrey, L.W. A census-based estimate of earth’s bacterial and archaeal diversity. PLoS Biol. 2019, 17, e3000106. [Google Scholar] [CrossRef]
- Siverio, F.; Marco-Noales, E.; Bertolini, E.; Teresani, G.R.; Peñalver, J.; Mansilla, P.; Aguín, O.; Pérez-Otero, R.; Abelleira, A.; Guerra-García, J.A.; et al. Survey of huanglongbing associated with ‘Candidatus Liberibacter’ species in Spain: Analyses of citrus plants and Trioza erytreae. Phytopathol. Mediterr. 2017, 56, 98–110. [Google Scholar] [CrossRef]
- Tyler, H.L.; Roesch, L.F.W.; Gowda, S.; Dawson, W.O.; Triplett, E.W. Confirmation of the sequence of ‘Candidatus Liberibacter asiaticus’ and assessment of microbial diversity in huanglongbing-infected citrus phloem using a metagenomic approach. Mol. Plant-Microbe Interact. 2009, 22, 1624–1634. [Google Scholar] [CrossRef][Green Version]
- Damsteegt, V.D.; Postnikova, E.N.; Stone, A.L.; Kuhlmann, M.; Wilson, C.; Sechler, A.; Schaad, N.W.; Brlansky, R.H.; Schneider, W.L. Murraya paniculata and related species as potential hosts and inoculum reservoirs of ‘Candidatus Liberibacter asiaticus’, causal agent of huanglongbing. Plant Dis. 2010, 94, 528–533. [Google Scholar] [CrossRef][Green Version]
- Lin, H.; Doddapaneni, H.; Bai, X.; Yao, J.; Zhao, X.; Civerolo, E.L. Acquisition of uncharacterized sequences from ‘Candidatus Liberibacter’, an unculturable bacterium, using an improved genomic walking method. Mol. Cell. Probes 2008, 22, 30–37. [Google Scholar] [CrossRef] [PubMed]
- Oyaizu-Masuchi, Y.; Komagata, K. Isolation of free-living nitrogen-fixing bacteria from the rhizosphere of rice. J. Gen. Appl. Microbiol. 1988, 34, 127–164. [Google Scholar] [CrossRef]
- Wang, J.; Haapalainen, M.; Schott, T.; Thompson, S.M.; Smith, G.R.; Nissinen, A.I.; Pirhonen, M. Genomic sequence of ‘Candidatus Liberibacter solanacearum’ haplotype C and its comparison with haplotype A and B genomes. PLoS ONE 2017, 12, e0171531. [Google Scholar] [CrossRef] [PubMed]
- Mantelin, S.; Fischer-Le Saux, M.; Zakhia, F.; Béna, G.; Bonneau, S.; Jeder, H.; de Lajudie, P.; Cleyet-Marel, J.C. Emended description of the genus Phyllobacterium and description of four novel species associated with plant roots: Phyllobacterium bourgognense sp. nov., Phyllobacterium ifriqiyense sp. nov., Phyllobacterium leguminum sp. nov. and Phyllobacterium brassicacearum sp. nov. Int. J. Syst. Evol. Microbiol. 2006, 56, 827–839. [Google Scholar] [CrossRef] [PubMed]
- Hartung, J.S.; Shao, J.; Kuykendall, L.D. Comparison of the ‘Candidatus Liberibacter asiaticus’ genome adapted for an intracellular lifestyle with other members of the Rhizobiales. PLoS ONE 2011, 6, e23289. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Trivedi, P. Citrus huanglongbing: A newly relevant disease presents unprecedented challenges. Phytopathology 2013, 103, 652–665. [Google Scholar] [CrossRef] [PubMed]
- Toft, C.; Andersson, S.G.E. Evolutionary microbial genomics: Insights into bacterial host adaptation. Nat. Rev. Genet. 2010, 11, 465–475. [Google Scholar] [CrossRef]
- Favia, G.; Ricci, I.; Marzorati, M.; Negri, I.; Alma, A.; Sacchi, L.; Bandi, C.; Daffonchio, D. Bacteria of the genus Asaia: A potential paratransgenic weapon against malaria. Adv. Exp. Med. Biol. 2008, 627, 49–59. [Google Scholar]
- Yamada, Y.; Katsura, K.; Kawasaki, H.; Widyastuti, Y.; Saono, S.; Seki, T.; Uchimura, T.; Komagata, K. Asaia bogorensis gen. nov., sp. nov., an unusual acetic acid bacterium in the α-Proteobacteria. Int. J. Syst. Evol. Microbiol. 2000, 50, 823–829. [Google Scholar] [CrossRef]
- Schäfer, A. Hemicellulose-degrading bacteria and yeasts from the termite gut. J. Appl. Bacteriol. 1996, 80, 471–478. [Google Scholar] [CrossRef]
- Eleftherianos, L.; Atri, J.; Accetta, J.; Castillo, J.C. Endosymbiotic bacteria in insects: Guardians of the immune system? Front. Physiol. 2013, 46. [Google Scholar] [CrossRef] [PubMed]
- Rasowo, B.A.; Copeland, R.S.; Khamis, F.M.; Aidoo, O.F.; Ajene, I.J.; Mohamed, S.A.; Sétamou, M.; Ekesi, S.; Borgemeister, C. Diversity and phylogenetic analysis of endosymbionts from Trioza erytreae (Del Guercio) and its parasitoids in Kenya. J. Appl. Entomol. 2020, jen.12807. [Google Scholar] [CrossRef]
- Werren, J.H.; Baldo, L.; Clark, M.E. Wolbachia: Master manipulators of invertebrate biology. Nat. Rev. Microbiol. 2008, 6, 741–751. [Google Scholar] [CrossRef] [PubMed]
- Hancock, P.A.; Sinkins, S.P.; Godfray, H.C.J. Strategies for introducing Wolbachia to reduce transmission of mosquito-borne diseases. PLoS Negl. Trop. Dis. 2011, 5, e1024. [Google Scholar] [CrossRef] [PubMed]
- Tamborindeguy, C.; Huot, O.B.; Ibanez, F.; Levy, J. The influence of bacteria on multitrophic interactions among plants, psyllids, and pathogen. Insect Sci. 2017, 24, 961–974. [Google Scholar] [CrossRef] [PubMed]
- Arahal, D.R.; Llop, P.; Pérez Alonso, M.; López, M.M. In silico evaluation of molecular probes for detection and identification of Ralstonia solanacearum and Clavibacter michiganensis subsp. sepedonicus. Syst. Appl. Microbiol. 2004, 27, 581–591. [Google Scholar] [CrossRef]
- Pritchard, L.; Holden, N.J.; Bielaszewska, M.; Karch, H.; Toth, I.K. Alignment-free design of highly discriminatory diagnostic primer Sets for Escherichia coli O104:H4 Outbreak Strains. PLoS ONE 2012, 7, e34498. [Google Scholar] [CrossRef]
- Coy, M.R.; Hoffmann, M.; Kingdom Gibbard, H.N.; Kuhns, E.H.; Pelz-Stelinski, K.S.; Stelinski, L.L. Nested-quantitative PCR approach with improved sensitivity for the detection of low titer levels of ‘Candidatus Liberibacter asiaticus’ in the Asian citrus psyllid, Diaphorina citri Kuwayama. J. Microbiol. Methods 2014, 102, 15–22. [Google Scholar] [CrossRef]
- Wilson, I.G. Inhibition and facilitation of nucleic acid amplification. Appl. Environ. Microbiol. 1997, 63, 3741–3751. [Google Scholar] [CrossRef]
| Source | Origin | Number Total Samples | Number Positive Samples by Bertolini et al. [19] | Number of Positive Samples by Li et al. [21] | |||
|---|---|---|---|---|---|---|---|
| CaLspp | CaLam | CaLas | CaLaf | ||||
| Plant | Citrus sp. | Spain | 2780 | 210 | 0 | 0 | 3 |
| Thailand | 1 | 1 | 1 | 1 | 0 | ||
| Murraya koenegii | Netherlands | 1 | 1 | 0 | 0 | 0 | |
| Uganda | 1 | 1 | 1 | 0 | 0 | ||
| Psyllid | Trioza erytreae | Spain a | 1051 | 9 | 0 | 0 | 0 |
| Parasitoid | Tamarixia dryi | South Africa | 82 | 25 | 0 | 0 | 0 |
| Spain a (Isoline F1) | 317 | 57 | 0 | 0 | 0 | ||
| Spain a (Isoline F2–4) | 596 | 142 | 0 | 0 | 2 | ||
| Spain a (Isoline F5–9) | 86 | 13 | 0 | 0 | 5 | ||
| Total | 4915 | 459 | 2 | 2 | 10 | ||
| Host | Origin | Sample | Real-Time PCR According to Bertolini et al. [19] | Real-Time PCR According to Li et al. [21] | Sequence Analysis by n from Checking Sample Results (Identity%—GenBank Accession Number) | ||
|---|---|---|---|---|---|---|---|
| CaLam | CaLas | CaLaf | |||||
| Murraya koenegii | Uganda | IVIA 5020 | 34.78 ± 0.5 | - | - | - | Rhizobium sp. (98%—HG938317) |
| IVIA 5029.2 | 35.79± 0.7 | 37.52± 0.1 | - | - | Sphingomonas sp. (98%—JX174255.1) | ||
| Citrus unshiu | Spain (mainland Spain) | IVIA 5361 | 35.64± 0.2 | - | - | 36.83± 0.8 | Sphingomonas sp. (94%—JX660147) |
| IVIA 5516 | 36.22 ± 0.1 | - | - | - | Soil bacterium from enrichment culture (100%—HQ640720) * | ||
| Citrus spp. | Thailand | IVIA 5029.1 | 34.48± 0.5 | 37.89± 0.2 | 38.47± 0.4 | - | Sphingomonas sp. (90%—EU567045.2) |
| Spain (Canary Islands) | 209R | 32.82± 0.3 | - | - | - | Bradyrhizobium sp. (93%—AY599690). | |
| 209A | 35.96± 0.5 | - | - | - | Rhizobiales bacterium (96%—LR642383.1) | ||
| P10; P11 | 29.48± 0.7 | - | - | 36.7± 0.3 | Phyllobacterium sp. (100%—MN810203) | ||
| P15; P16 | 28.66± 0.7 | - | - | - | Phyllobacterium sp. (100%—MN810203) | ||
| P 48 | 37.34± 0.1 | - | - | 36.50± 0.1 | Asaia sp. (99%—MN099438.1) | ||
| Tamarixia dryi | South Africa | T: 7; 31; 33; 38; 46; 66; 67; 147; 151; 154; 170; 204; 212; 219; 227; 244; 247; 257; 260; 267; 278 | 34.84± 0.1 | - | - | - | Asaia sp. (99%—MN099438.1) |
| T75 | 32.76± 0.8 | - | - | - | Rhizobium sp. (99%—JX566578.1) | ||
| T86 | 33.47± 0.1 | - | - | - | Ochrobactrum sp. (100%—CP015776) | ||
| T: 168; 186 | 35.34± 0.1 | - | - | - | Wolbachia endosymbiont of Drosophila sp. (99%—CP034335.1) | ||
| T: 540; 555; 567; 575; 576 | 26.29± 1.7 | ND | - | 35.62± 0.7 | Asaia sp. (99%—MN099438.1) | ||
| Tryoza erytreae | Spain (Canary Islands) | i273 | 34.51 ± 0.5 | - | - | - | Uncultured bacterium. (99%—KJ664621) * |
| i800; i360 | 35.31 ± 0.2 | - | - | - | Sodalis sp. (91%—MT536226.1) | ||
| i487; i502 | 32.41 ± 0.6 | - | - | - | Secondary endosymbiont of Bactericera cockerelli (91%—AF263557.1) | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Morán, F.; Barbé, S.; Bastin, S.; Navarro, I.; Bertolini, E.; López, M.M.; Hernández-Suárez, E.; Urbaneja, A.; Tena, A.; Siverio, F.; et al. The Challenge of Environmental Samples for PCR Detection of Phytopathogenic Bacteria: A Case Study of Citrus Huanglongbing Disease. Agronomy 2021, 11, 10. https://doi.org/10.3390/agronomy11010010
Morán F, Barbé S, Bastin S, Navarro I, Bertolini E, López MM, Hernández-Suárez E, Urbaneja A, Tena A, Siverio F, et al. The Challenge of Environmental Samples for PCR Detection of Phytopathogenic Bacteria: A Case Study of Citrus Huanglongbing Disease. Agronomy. 2021; 11(1):10. https://doi.org/10.3390/agronomy11010010
Chicago/Turabian StyleMorán, Félix, Silvia Barbé, Saskia Bastin, Inmaculada Navarro, Edson Bertolini, María M. López, Estrella Hernández-Suárez, Alberto Urbaneja, Alejandro Tena, Felipe Siverio, and et al. 2021. "The Challenge of Environmental Samples for PCR Detection of Phytopathogenic Bacteria: A Case Study of Citrus Huanglongbing Disease" Agronomy 11, no. 1: 10. https://doi.org/10.3390/agronomy11010010
APA StyleMorán, F., Barbé, S., Bastin, S., Navarro, I., Bertolini, E., López, M. M., Hernández-Suárez, E., Urbaneja, A., Tena, A., Siverio, F., & Marco-Noales, E. (2021). The Challenge of Environmental Samples for PCR Detection of Phytopathogenic Bacteria: A Case Study of Citrus Huanglongbing Disease. Agronomy, 11(1), 10. https://doi.org/10.3390/agronomy11010010








