The Development of a European and Mediterranean Chickpea Association Panel (EMCAP)
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material and Field Evaluation
2.2. Association Panel Development
2.3. UNIVPM_Ca_core2 Phenotypic Evaluation
2.4. Data Analysis
3. Results
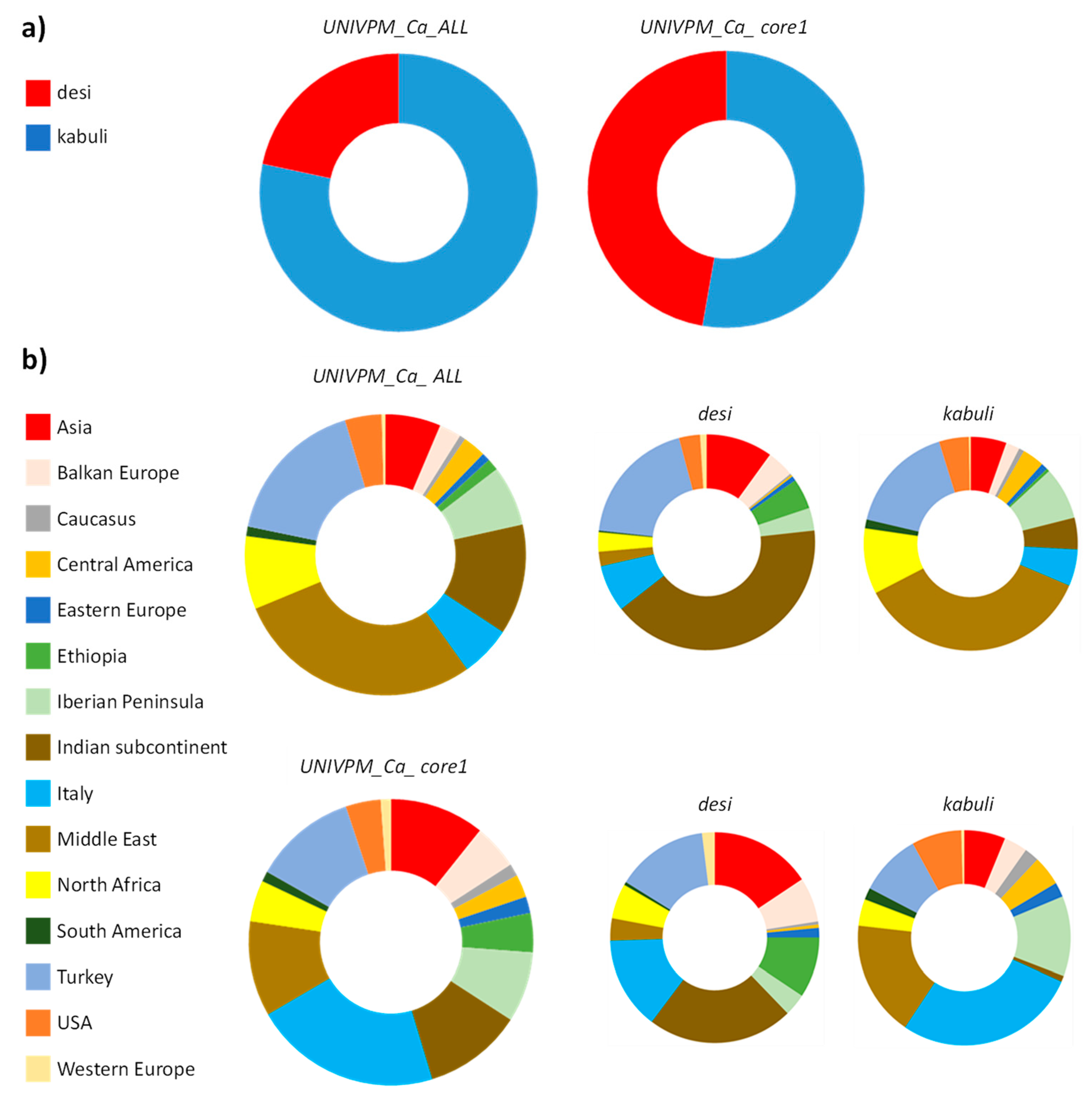
3.1. UNIVPM_Ca_ALL versus UNIVPM_Ca_core1
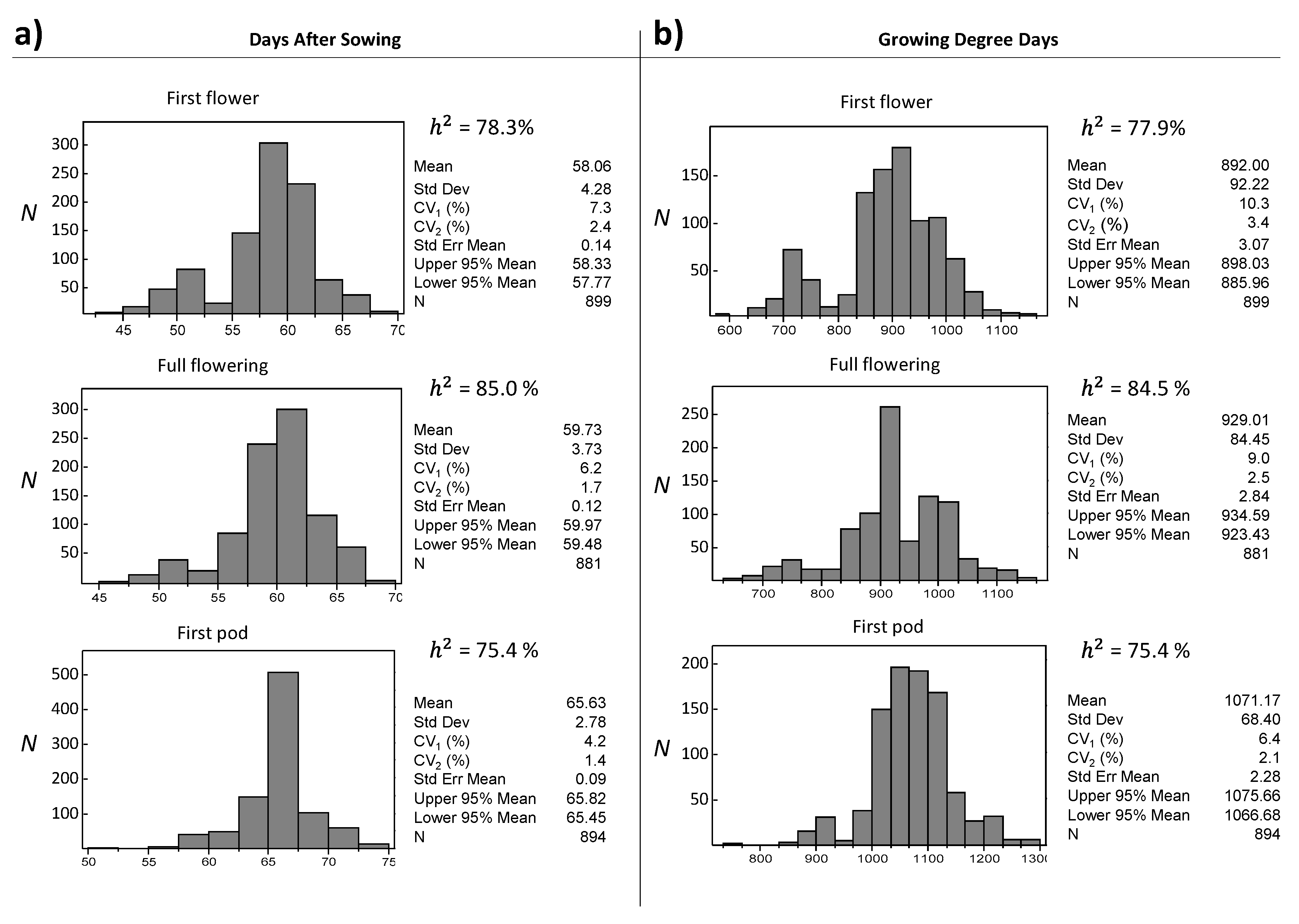
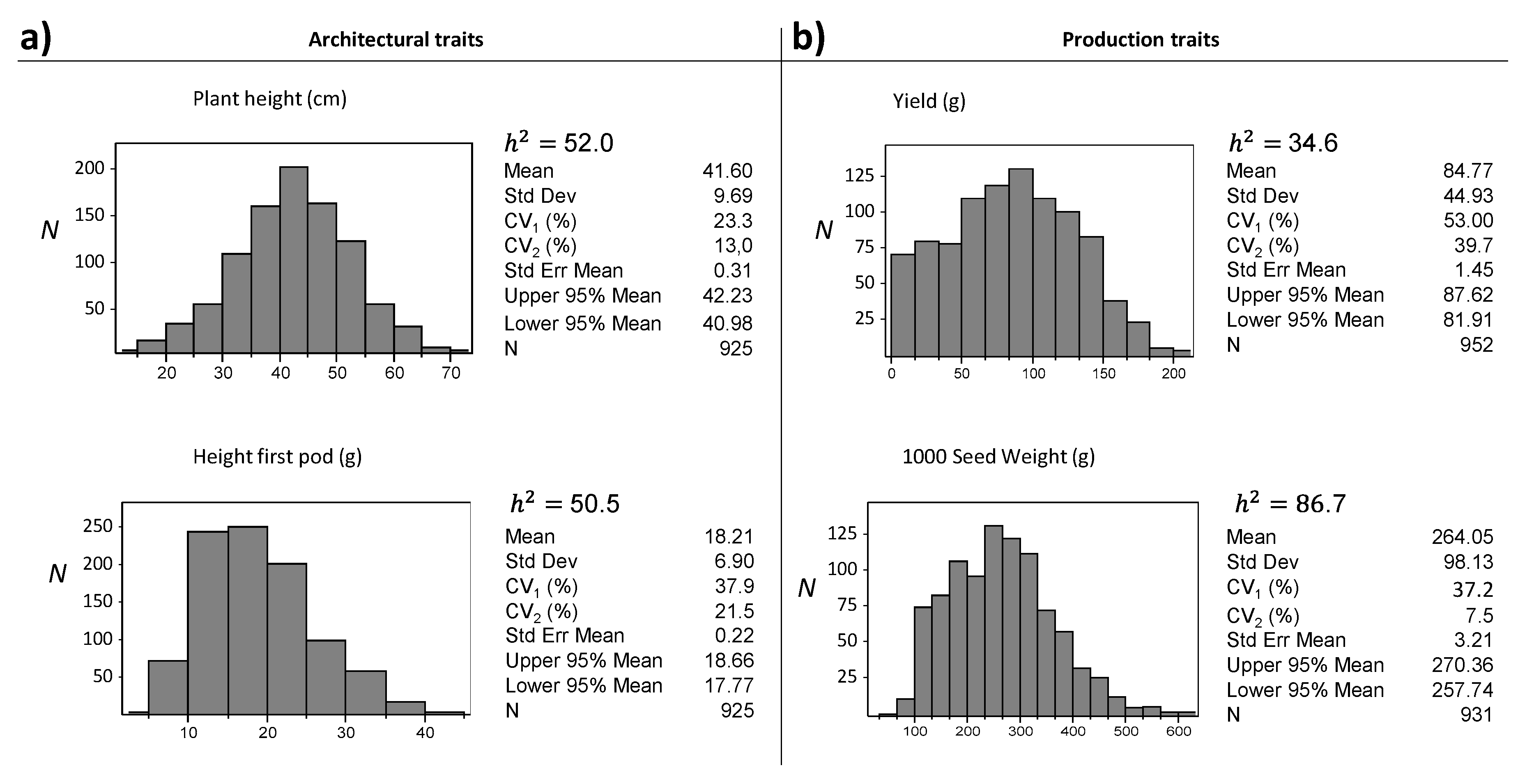
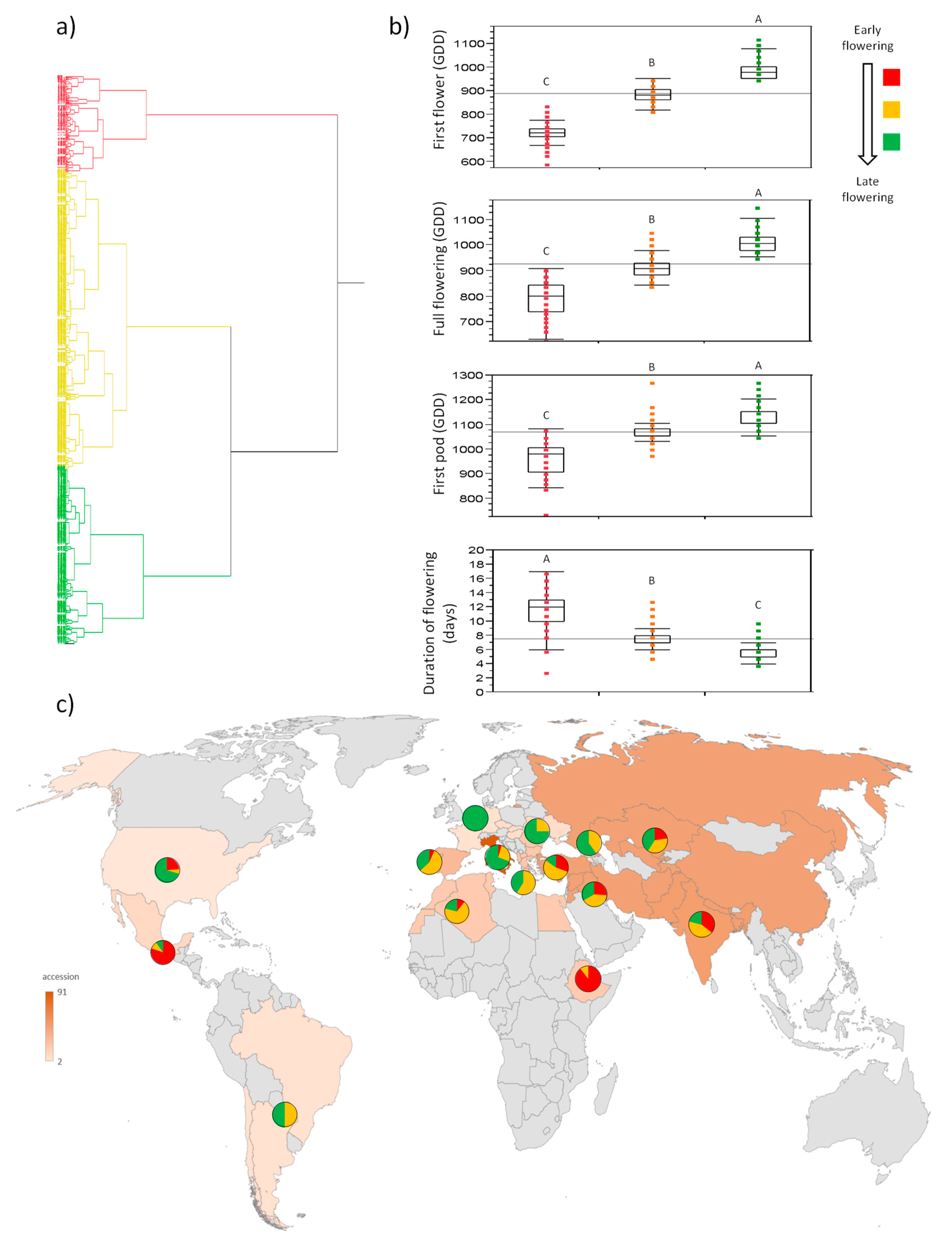
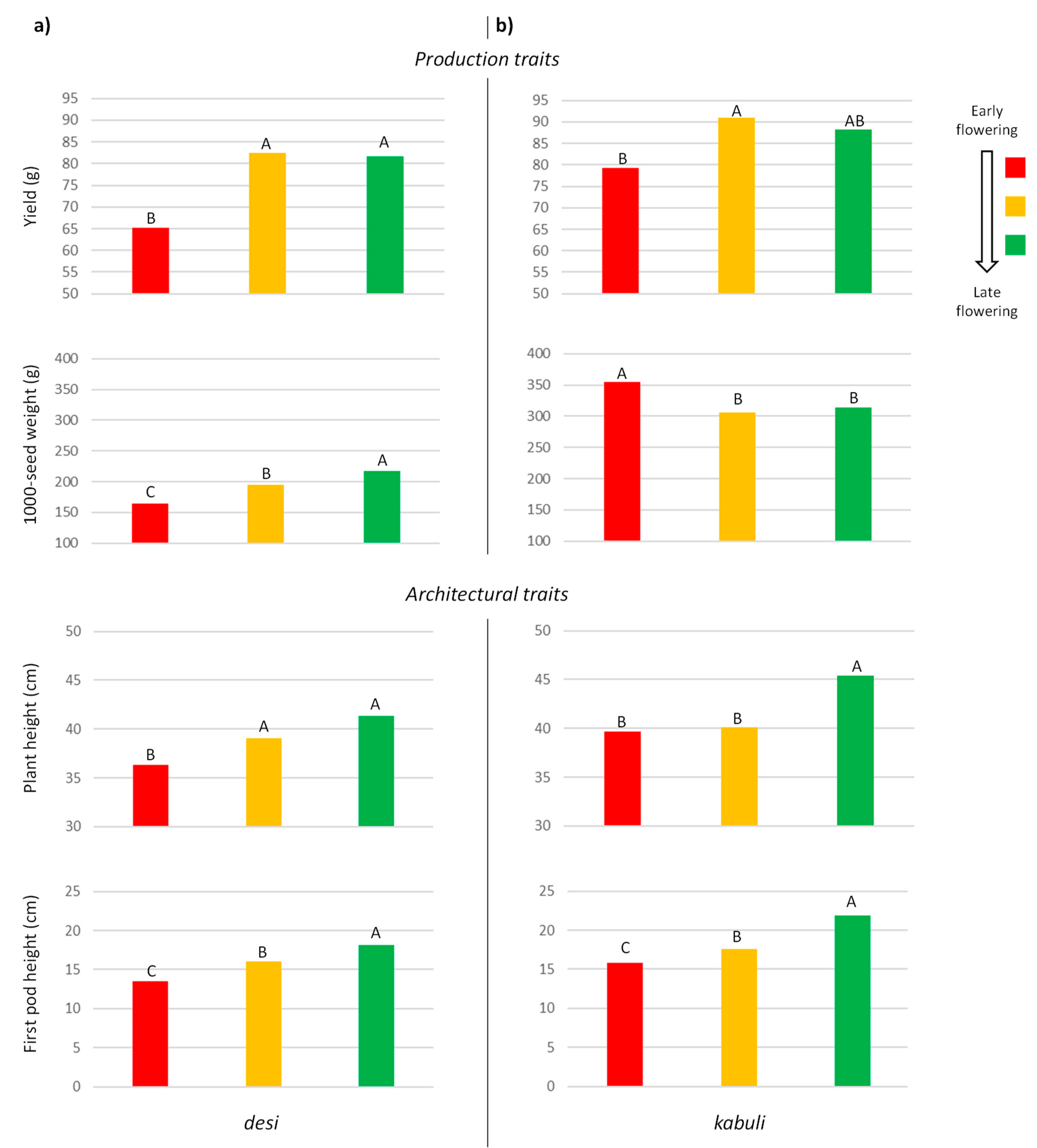
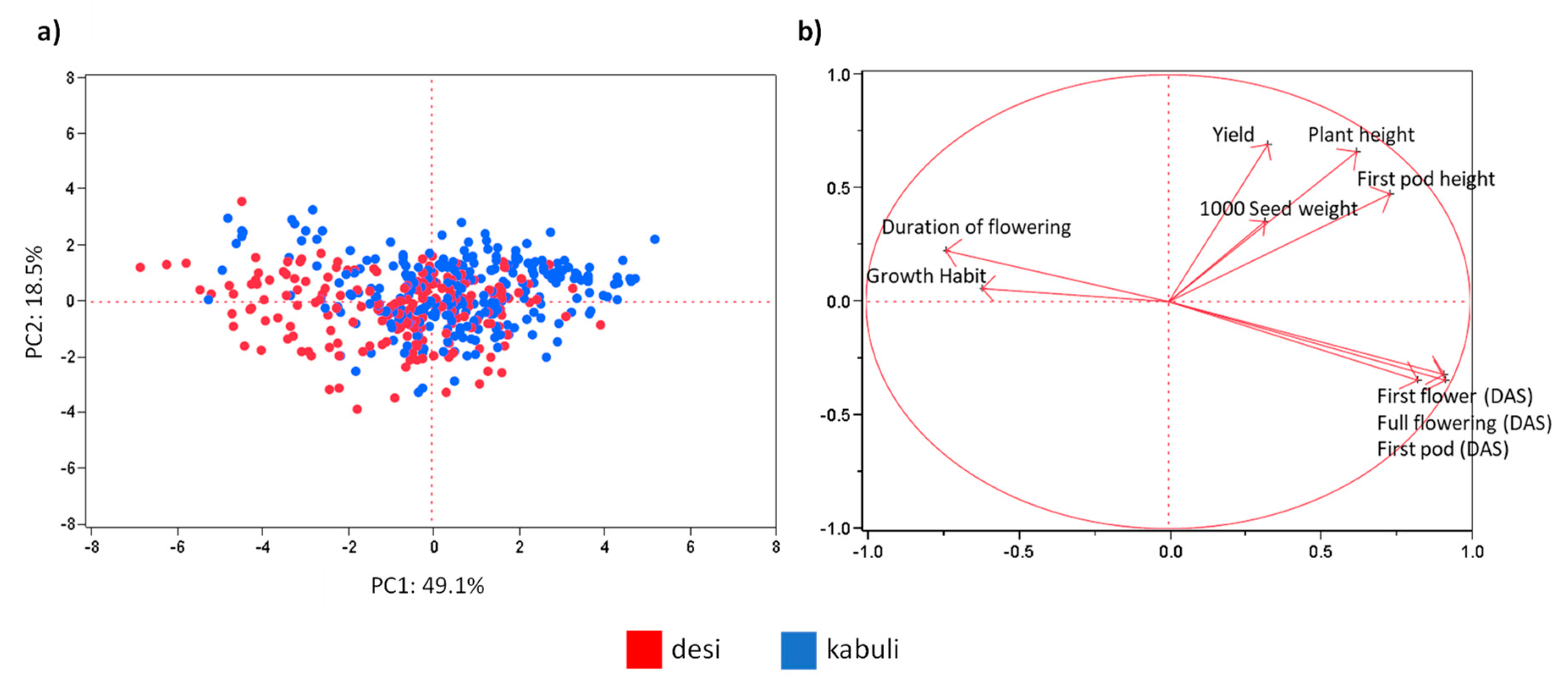
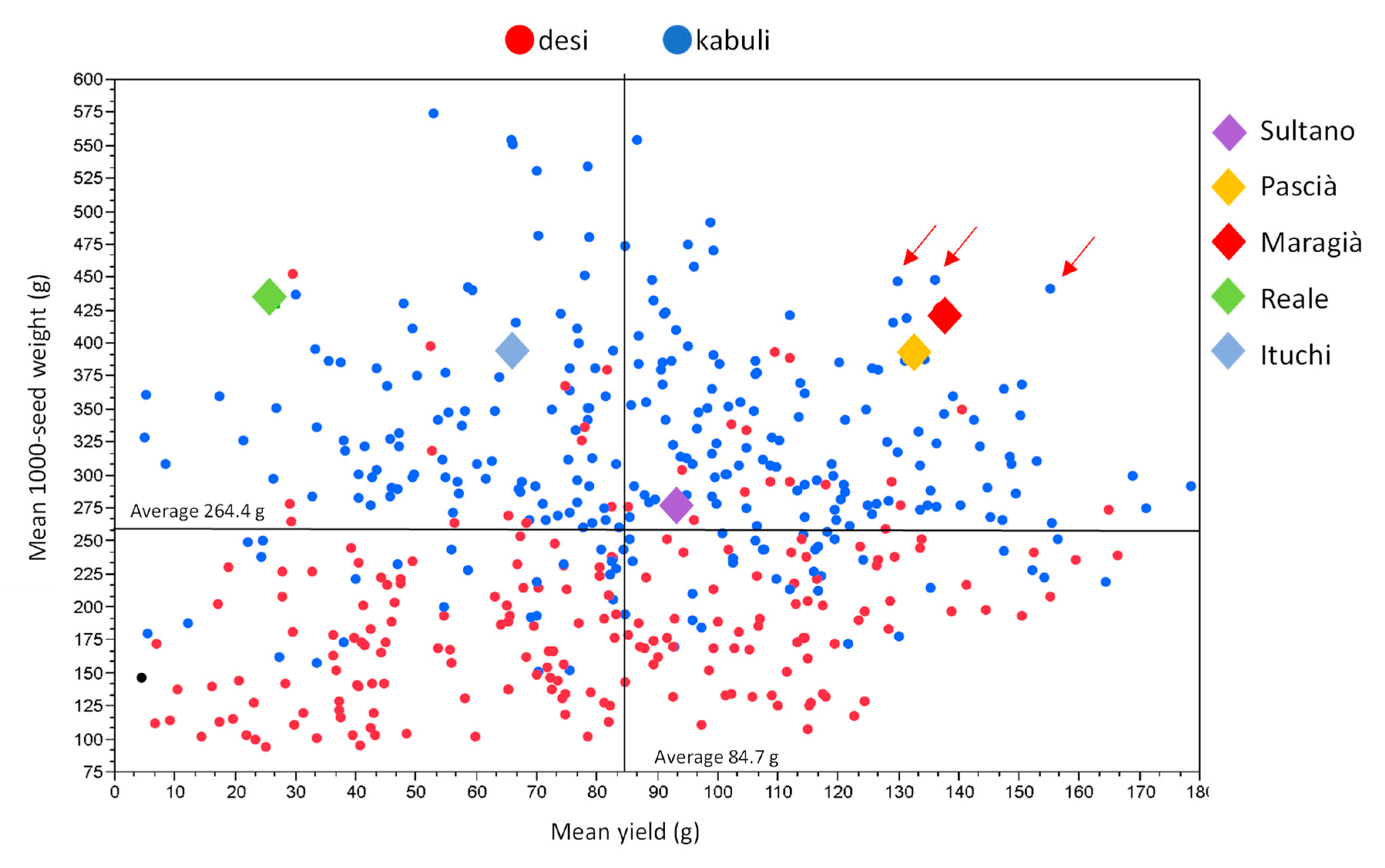
3.2. UNIVPM_Ca_core2 Phenotypic Evaluation
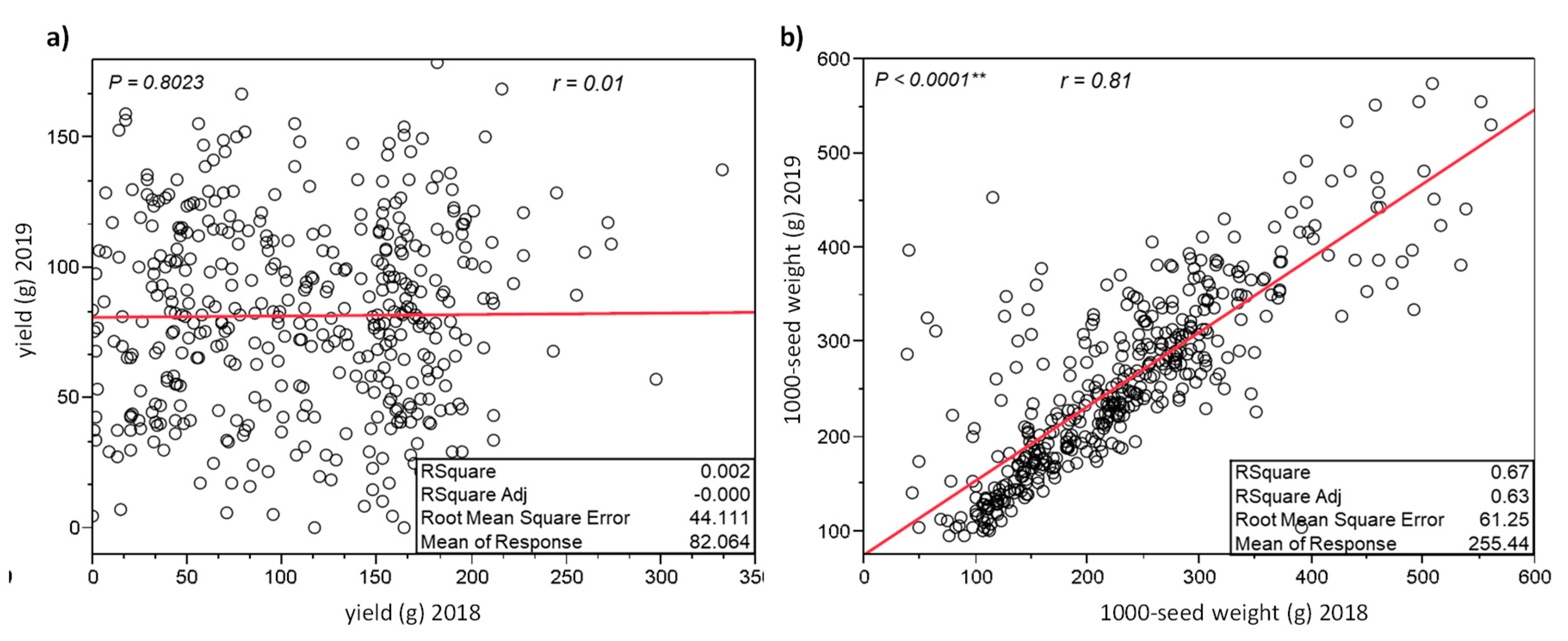
3.3. UNIVPM_Ca_core1: Comparison of the 2018 and 2019 Phenotypic Data
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Rawal, V.; Navarro, D.K. The Global Economy of Pulses; FAO: Rome, Italy, 2019. [Google Scholar]
- Abbo, S.; Berger, J.; Turner, N.C. Viewpoint: Evolution of cultivated chickpea: Four bottlenecks limit diversity andconstrain adaptation. Funct. Plant Boil. 2003, 30, 1081–1087. [Google Scholar] [CrossRef]
- Ladizinsky, G.; Adler, A. The origin of chickpea Cicer arietinum L. Euphytica 1976, 25, 211–217. [Google Scholar] [CrossRef]
- Varshney, R.; Thudi, M.; Roorkiwal, M.; He, W.; Upadhyaya, H.D.; Yang, W.; Bajaj, P.; Cubry, P.; Rathore, A.; Jian, J.; et al. Resequencing of 429 chickpea accessions from 45 countries provides insights into genome diversity, domestication and agronomic traits. Nat. Genet. 2019, 51, 857–864. [Google Scholar] [CrossRef]
- Fierro, M.; Palmieri, D.; Curtis, F.; De Vitullo, D.; Rubio, J.; Gil, J.; Lima, G.; Millan, T. Genetic and agronomic characterization of chickpea landraces for resistance to Fusarium oxysporum f. sp. ciceris. Phytopathol. Mediterr. 2019, 58, 239–248. [Google Scholar]
- Iliadis, C. Evaluation of six chickpea varieties for seed yield under autumn and spring sowing. J. Agric. Sci. 2001, 137, 439–444. [Google Scholar] [CrossRef]
- Singh, K.B.; Malhotra, R.S.; Saxena, M.C.; Bejiga, G. Superiority of Winter Sowing over Traditional Spring Sowing of Chickpea in the Mediterranean Region. Agron. J. 1907, 89, 112–118. [Google Scholar] [CrossRef]
- Foyer, C.H.; Lam, H.; Nguyen, H.T.; Siddique, K.H.M.; Varshney, R.; Colmer, T.D.; Cowling, W.A.; Bramley, H.; Mori, T.A.; Hodgson, J.M.; et al. Neglecting legumes has compromised human health and sustainable food production. Nat. Plants 2016, 2, 16112. [Google Scholar] [CrossRef]
- Papa, R. Increase Consortium: Increase-Intelligent Collections of Food Legumes Genetic Resources for European Agrofood Systems. Plant J. 2020, in press. [Google Scholar]
- Gupta, S.; Nawaz, K.; Parween, S.; Roy, R.; Sahu, K.; Pole, A.K.; Khandal, H.; Srivastava, R.; Parida, S.K.; Chattopadhyay, D. Draft genome sequence of Cicer reticulatum L., the wild progenitor of chickpea provides a resource for agronomic trait improvement. DNA Res. 2016, 24, 1–10. [Google Scholar] [CrossRef]
- Parween, S.; Nawaz, K.; Roy, R.; Pole, A.K.; Suresh, B.V.; Misra, G.; Jain, M.; Yadav, G.; Parida, S.K.; Tyagi, A.K.; et al. An advanced draft genome assembly of a desi type chickpea (Cicer arietinum L.). Sci. Rep. 2015, 5, 12806. [Google Scholar] [CrossRef]
- Varshney, R.; Song, C.; Saxena, R.; Azam, S.; Yu, S.; Sharpe, A.G.; Cannon, S.; Baek, J.; Rosen, B.D.; Tar’An, B.; et al. Draft genome sequence of chickpea (Cicer arietinum) provides a resource for trait improvement. Nat. Biotechnol. 2013, 31, 240–246. [Google Scholar] [CrossRef]
- Brown, A.H.D. Core collections: A practical approach to genetic resources management. Genome 1989, 31, 818–824. [Google Scholar] [CrossRef]
- El Bakkali, A.; Haouane, H.; Moukhli, A.; Costes, E.; Van Damme, P.; Khadari, B. Construction of Core Collections Suitable for Association Mapping to Optimize Use of Mediterranean Olive (Olea europaea L.) Genetic Resources. PLoS ONE 2013, 8, e61265. [Google Scholar] [CrossRef]
- Perseguini, J.M.K.C.; Silva, G.M.B.; Rosa, J.R.B.F.; Gazaffi, R.; Marçal, J.F.; Carbonell, S.A.M.; Chiorato, A.F.; Zucchi, M.I.; Garcia, A.A.F.; Benchimol, L.L. Developing a common bean core collection suitable for association mapping studies. Genet. Mol. Biol. 2015, 38, 67–78. [Google Scholar] [CrossRef]
- Ambreen, H.; Kumar, S.; Kumar, A.; Agarwal, M.; Jagannath, A.; Goel, S. Association Mapping for Important Agronomic Traits in Safflower (Carthamus tinctorius L.) Core Collection Using Microsatellite Markers. Front. Plant Sci. 2018, 9, 402. [Google Scholar] [CrossRef]
- Upadhyaya, H.D.; Ortiz, R. A mini core subset for capturing diversity and promoting utilization of chickpea genetic resources in crop improvement. Theor. Appl. Genet. 2001, 102, 1292–1298. [Google Scholar] [CrossRef]
- Upadhyaya, H.D.; Bramel, P.J.; Singh, S. Development of a Chickpea Core Subset Using Geographic Distribution and Quantitative Traits. Crop. Sci. 2001, 41, 206–210. [Google Scholar] [CrossRef]
- Upadhyaya, H.D.; Ortiz, R.; Bramel, P.J.; Singh, S. Phenotypic diversity for morphological and agronomic characteristics in chickpea core collection. Euphytica 2002, 123, 333–342. [Google Scholar] [CrossRef]
- Upadhyaya, H.D.; Furman, B.J.; Dwivedi, S.L.; Udupa, S.M.; Gowda, C.L.L.; Baum, M.; Crouch, J.H.; Buhariwalla, H.K.; Singh, S. Development of a composite collection for mining germplasm possessing allelic variation for beneficial traits in chickpea. Plant Genet. Resour. 2006, 4, 13–19. [Google Scholar] [CrossRef]
- Upadhyaya, H.D.; Dwivedi, S.L.; Baum, M.; Varshney, R.; Udupa, S.; Gowda, C.L.L.; Hoisington, D.; Singh, S. Genetic structure, diversity, and allelic richness in composite collection and reference set in chickpea (Cicer arietinum L.). BMC Plant Biol. 2008, 8, 106. [Google Scholar] [CrossRef]
- Dwivedi, S.L.; Upadhyaya, H.D.; Balaji, J.; Buhariwalla, H.K.; Blair, M.W.; Ortiz, R.; Crouch, J.H.; Serraj, R. Using Genomics to Exploit Grain Legume Biodiversity in Crop Improvement. Plant Breed. Rev. 2010, 26, 171–357. [Google Scholar] [CrossRef]
- Upadhyaya, H.D.; Bajaj, D.; Narnoliya, L.; Das, S.; Kumar, V.; Gowda, C.L.L.; Sharma, S.; Tyagi, A.K.; Parida, S.K. Genome-Wide Scans for Delineation of Candidate Genes Regulating Seed-Protein Content in Chickpea. Front. Plant Sci. 2016, 7, 302. [Google Scholar] [CrossRef]
- Basu, U.; Srivastava, R.; Bajaj, D.; Thakro, V.; Daware, A.; Malik, N.; Upadhyaya, H.D.; Parida, S.K. Genome-wide generation and genotyping of informative SNPs to scan molecular signatures for seed yield in chickpea. Sci. Rep. 2018, 8, 13240. [Google Scholar] [CrossRef]
- Miller, P.; Lanier, W.; Brandt, S. Using growing degree days to predict plant stages. In Ag/Extension Communications Coordinator, Communications Services; Montana State University-Bozeman: Bozeman, MT, USA, 2001. [Google Scholar]
- IBPGR, ICRISAT and ICARDA. Descriptors for Chickpea (Cicer arietinum L.); International Board for Plant Genetic Resources (IBPGR): Rome, Italy; International Crops Research Institute for the Semi-Arid Tropics (ICRISAT): Patancheru, India; International Center for Agriculture Research in the Dry Areas (ICARDA): Aleppo, Syria, 1993. [Google Scholar]
- Ward, J.H. Hierarchical grouping to optimize an objective function. J. Am. Stat. Assoc. 1963, 58, 236–244. [Google Scholar]
- Visscher, P.M.; Hill, W.G.; Wray, N.R. Heritability in the genomics era—Concepts and misconceptions. Nat. Rev. Genet. 2008, 9, 255–266. [Google Scholar] [CrossRef]
- McCouch, S.; Navabi, K.; Abberton, M.; Anglin, N.L.; Barbieri, R.L.; Baum, M.; Bett, K.; Booker, H.; Brown, G.L.; Bryan, G.J.; et al. Mobilizing Crop Biodiversity. Mol. Plant 2020. [Google Scholar] [CrossRef]
- Cortinovis, G.; Di Vittori, V.; Bellucci, E.; Bitocchi, E.; Papa, R. Adaptation to novel environments during crop diversification. Curr. Opin. Plant Biol. 2020, 56, 203–217. [Google Scholar] [CrossRef]
- Korte, A.; Farlow, A. The advantages and limitations of trait analysis with GWAS: A review. Plant Methods 2013, 9, 29. [Google Scholar] [CrossRef]
- Canci, H.; Toker, C. Evaluation of Yield Criteria for Drought and Heat Resistance in Chickpea (Cicer arietinum L.). J. Agron. Crop. Sci. 2009, 195, 47–54. [Google Scholar] [CrossRef]
- Mallu, T.S.; Mwangi, S.G.; Nyende, A.B.; Ganga Rao, N.V.P.R.; Odeny, D.A.; Rathore, A.; Kumar, A. Assessment of genetic variation and heritability of agronomic traits in chickpea (Cicer arietinum L). Int. J. Agron. Agric. Res. 2015, 5, 76–88. [Google Scholar]
- Roberts, E.H.; Hadley, P.; Summerfield, R.J. Effects of Temperature and Photoperiod on Flowering in Chickpeas (Cicer arietinum L.). Ann. Bot. 1985, 55, 881–892. [Google Scholar] [CrossRef]
- Ellis, R.H.; Lawn, R.J.; Summerfield, R.J.; Qi, A.; Roberts, E.H.; Chay, P.M.; Brouwer, J.B.; Rose, J.L.; Yeates, S.J.; Sandover, S. Towards the Reliable Prediction of Time to Flowering in Six Annual Crops. V. Chickpea (Cicer arietinum). Exp. Agric. 1994, 30, 271–282. [Google Scholar] [CrossRef]
- Berger, J.D.; Ali, M.; Basu, P.; Chaudhary, B.; Chaturvedi, S.; Deshmukh, P.; Dharmaraj, P.; Dwivedi, S.; Gangadhar, G.; Gaur, P.; et al. Genotype by environment studies demonstrate the critical role of phenology in adaptation of chickpea (Cicer arietinum L.) to high and low yielding environments of India. Field Crop. Res. 2006, 98, 230–244. [Google Scholar] [CrossRef]
- Berger, J.D.; Milroy, S.; Turner, N.C.; Siddique, K.H.M.; Imtiaz, M.; Malhotra, R. Chickpea evolution has selected for contrasting phenological mechanisms among different habitats. Euphytica 2011, 180, 1–15. [Google Scholar] [CrossRef]
- Bajaj, D.; Upadhyaya, H.D.; Khan, Y.; Das, S.; Badoni, S.; Shree, T.; Kumar, V.; Tripathi, S.; Gowda, C.L.L.; Singh, S.; et al. A combinatorial approach of comprehensive QTL-based comparative genome mapping and transcript profiling identified a seed weight-regulating candidate gene in chickpea. Sci. Rep. 2015, 5, 9264. [Google Scholar] [CrossRef]
- Sharma, K.D.; Chen, W.; Muehlbauer, F.J. Genetics of Chickpea Resistance to Five Races of Fusarium Wilt and a Concise Set of Race Differentials for Fusarium oxysporum f. sp. ciceris. Plant Dis. 2005, 89, 385–390. [Google Scholar] [CrossRef]
- Penmetsa, R.V.; Carrasquilla-Garcia, N.; Bergmann, E.M.; Vance, L.; Castro, B.; Kassa, M.T.; Sarma, B.K.; Datta, S.; Farmer, A.D.; Baek, J.-M.; et al. Multiple post-domestication origins of kabuli chickpea through allelic variation in a diversification-associated transcription factor. New Phytol. 2016, 211, 1440–1451. [Google Scholar] [CrossRef]
- De Giovanni, C.; Pavan, S.; Taranto, F.; Di Rienzo, V.; Miazzi, M.M.; Marcotrigiano, A.R.; Mangini, G.; Montemurro, C.; Ricciardi, L.; Lotti, C. Genetic variation of a global germplasm collection of chickpea (Cicer arietinum L.) including Italian accessions at risk of genetic erosion. Physiol. Mol. Biol. Plants 2016, 23, 197–205. [Google Scholar] [CrossRef]
- Mascher, M.; Schreiber, M.; Scholz, U.; Graner, A.; Reif, J.C.; Stein, N. Genebank genomics bridges the gap between the conservation of crop diversity and plant breeding. Nat. Genet. 2019, 51, 1076–1081. [Google Scholar] [CrossRef]


| Geographic Area | Country | N. of Accessions | |
|---|---|---|---|
| UNIVPM_Ca_ALL | UNIVPM_Ca_core1 | ||
| Asia | Total | 121 | 46 |
| Tajikistan | 54 | 30 | |
| Uzbekistan | 31 | 5 | |
| Afghanistan | 22 | 6 | |
| China | 4 | 2 | |
| Kazakhstan | 1 | 1 | |
| Russia | 9 | 2 | |
| Balkan Europe | Total | 48 | 22 |
| North Macedonia | 19 | 4 | |
| Greece | 18 | 10 | |
| Bulgaria | 4 | 3 | |
| Albania | 3 | 2 | |
| Moldova | 2 | 2 | |
| Romania | 2 | 1 | |
| Caucasus | Total | 13 | 6 |
| Armenia | 6 | 2 | |
| Azerbaijan | 4 | 1 | |
| Georgia | 3 | 3 | |
| Central America | Mexico (total) | 50 | 11 |
| Eastern Europe | Total | 18 | 8 |
| Ukraine | 8 | 3 | |
| Hungary | 7 | 2 | |
| Slovakia | 2 | 2 | |
| Czech Republic | 1 | 1 | |
| Ethiopia | Ethiopia (total) | 26 | 19 |
| Iberian Peninsula | Total | 133 | 30 |
| Spain | 102 | 20 | |
| Portugal | 31 | 10 | |
| Indian subcontinent | Total | 241 | 48 |
| India | 198 | 31 | |
| Pakistan | 37 | 12 | |
| Bangladesh | 4 | 3 | |
| Nepal | 2 | 2 | |
| Italy | Italy (total) | 111 | 91 |
| Middle East | Total | 542 | 46 |
| Iran | 117 | 4 | |
| Syria | 361 | 32 | |
| Jordan | 43 | 1 | |
| Iraq | 14 | 2 | |
| Cyprus | 4 | 4 | |
| Lebanon | 3 | 3 | |
| North Africa | Total | 159 | 20 |
| Morocco | 89 | 6 | |
| Tunisia | 50 | 11 | |
| Algeria | 15 | 2 | |
| Egypt | 5 | 1 | |
| Others | Total | 6 | 2 |
| Australia | 5 | 1 | |
| Malawi | 1 | 1 | |
| South America | Total | 21 | 5 |
| Chile | 19 | 3 | |
| Argentina | 1 | 1 | |
| Brazil | 1 | 1 | |
| Turkey | Turkey (total) | 324 | 46 |
| USA | USA (total) | 80 | 17 |
| Western Europe | Total | 8 | 5 |
| France | 5 | 2 | |
| Germany | 3 | 3 | |
| Overall total | 1901 1 | 422 1 | |
| Geographic Area | Collection | Deviation 1 (%) | |||||||
|---|---|---|---|---|---|---|---|---|---|
| UNIVPM_Ca_ALL | UNIVPM_Ca_core1 | ||||||||
| (n) | (%) | Desi (%) | Kabuli (%) | (n) | (%) | Desi (%) | Kabuli (%) | ||
| Asia | 121 | 6.4 | 33.9 | 66.1 | 46 | 10.9 | 69.6 | 30.4 | 4.5 |
| Balkan Europe | 48 | 2.5 | 35.4 | 64.6 | 22 | 5.2 | 63.6 | 36.4 | 2.7 |
| Caucasus | 13 | 0.7 | 7.7 | 92.3 | 6 | 1.4 | 16.7 | 83.3 | 0.7 |
| Central America | 50 | 2.6 | 2.0 | 98.0 | 11 | 2.6 | 9.1 | 90.9 | 0.0 |
| Eastern Europe | 18 | 0.9 | 16.7 | 83.3 | 8 | 1.9 | 37.5 | 62.5 | 1.0 |
| Ethiopia | 26 | 1.4 | 73.1 | 26.9 | 19 | 4.5 | 100.0 | 0.0 | 3.1 |
| Iberian Peninsula | 133 | 7.0 | 10.5 | 88.7 | 30 | 7.1 | 16.7 | 83.3 | 0.1 |
| Indian subcontinent | 241 | 12.7 | 70.5 | 29.5 | 48 | 11.4 | 95.8 | 4.2 | −1.2 |
| Italy | 111 | 5.8 | 26.1 | 73.9 | 91 | 21.4 | 31.9 | 68.1 | 15.6 |
| Middle East | 542 | 28.5 | 1.7 | 98.3 | 46 | 10.8 | 15.2 | 84.8 | −17.3 |
| North Africa | 159 | 8.4 | 7.6 | 92.5 | 20 | 4.7 | 55.0 | 45.0 | −3.5 |
| other | 6 | 0.3 | 33.3 | 66.7 | 2 | 0.5 | 0.5 | 0.5 | 0.2 |
| South America | 21 | 1.1 | 4.8 | 95.2 | 5 | 1.2 | 20.0 | 80.0 | 0.1 |
| Turkey | 324 | 17.0 | 24.4 | 75.6 | 46 | 10.9 | 56.5 | 43.5 | −6.0 |
| USA | 80 | 4.2 | 16.3 | 83.8 | 17 | 4.0 | 0.0 | 100.0 | −0.1 |
| Western Europe | 8 | 0.4 | 50.0 | 50.0 | 5 | 1.2 | 80.0 | 20.0 | 0.8 |
| Over total | 1901 | 422 | |||||||
| Yield Class (g) | Collection | Deviation 1 (%) | |||
|---|---|---|---|---|---|
| UNIVPM_Ca_ALL | UNIVPM_Ca_core1 | ||||
| (n) | (%) | (n) | (%) | ||
| <50 | 694 | 35.9 | 103 | 24.2 | −11.7 |
| 50–100 | 670 | 34.7 | 88 | 20.7 | −14.0 |
| 101–150 | 404 | 20.9 | 72 | 16.9 | −4.0 |
| 151–200 | 135 | 7.0 | 135 | 31.7 | 24.7 |
| 201–250 | 22 | 1.1 | 22 | 5.2 | 4.1 |
| >250 | 6 | 0.3 | 6 | 1.4 | 1.1 |
| 1000-Seed Weight Class (g) | Collection | Deviation 1 (%) | |||
|---|---|---|---|---|---|
| UNIVPM_Ca_ALL | UNIVPM_Ca_core1 | ||||
| (n) | (%) | (n) | (%) | ||
| <100 | 158 | 8.1 | 21 | 4.9 | −3.2 |
| 100–200 | 571 | 29.6 | 154 | 36.2 | 6.6 |
| 201–300 | 769 | 39.8 | 158 | 37.1 | −2.7 |
| 301–400 | 378 | 19.6 | 65 | 15.3 | −4.3 |
| 401–500 | 47 | 2.4 | 20 | 4.7 | 2.3 |
| >500 | 8 | 0.4 | 8 | 1.9 | 1.5 |
| Flowering Trait | Class | Collection | Deviation 1 (%) | |||
|---|---|---|---|---|---|---|
| UNIVPM_Ca_ALL | UNIVPM_Ca_core1 | |||||
| (n) | (%) | (n) | (%) | |||
| Days to first flower | <72 | 14 | 1.1 | 14 | 5.1 | 4.0 |
| 72–85 | 680 | 53.9 | 157 | 57.5 | 3.6 | |
| 86–92 | 554 | 43.9 | 100 | 36.6 | −7.3 | |
| 93–99 | 14 | 1.1 | 2 | 0.7 | −0.4 | |
| Days to full flowering | <92 | 1026 | 65.9 | 232 | 67.2 | 1.3 |
| 92–99 | 524 | 33.7 | 112 | 32.5 | −1.2 | |
| 106 | 6 | 0.4 | 1 | 0.3 | −0.1 | |
| Days to first pod | <92 | 17 | 0.9 | 6 | 1.4 | 0.5 |
| 92–99 | 1295 | 67.2 | 289 | 67.8 | 0.6 | |
| 100–106 | 604 | 31.3 | 129 | 30.3 | −1.1 | |
| 107–111 | 11 | 0.6 | 2 | 0.5 | −0.1 | |
| Trait | Units | Genotype | Prob > F | |
|---|---|---|---|---|
| Desi | Kabuli | |||
| Days to first flower | DAS | 56.8 | 58.9 | <0.0001 |
| Days to full flowering | DAS | 58.7 | 60.5 | <0.0001 |
| Days to first pod | DAS | 64.8 | 66.2 | <0.0001 |
| Duration of flowering | days | 8 | 7.2 | <0.0001 |
| Plant height | cm | 38.9 | 43.4 | <0.0001 |
| First pod height | cm | 15.9 | 19.9 | <0.0001 |
| Yield | g | 78.3 | 90.1 | <0.0001 |
| 1000-seed weight | g | 192.9 | 316.2 | <0.0001 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rocchetti, L.; Bellucci, E.; Cortinovis, G.; Di Vittori, V.; Lanzavecchia, G.; Frascarelli, G.; Nanni, L.; Del Gatto, A.; Pieri, S.; Mangoni, L.; et al. The Development of a European and Mediterranean Chickpea Association Panel (EMCAP). Agronomy 2020, 10, 1417. https://doi.org/10.3390/agronomy10091417
Rocchetti L, Bellucci E, Cortinovis G, Di Vittori V, Lanzavecchia G, Frascarelli G, Nanni L, Del Gatto A, Pieri S, Mangoni L, et al. The Development of a European and Mediterranean Chickpea Association Panel (EMCAP). Agronomy. 2020; 10(9):1417. https://doi.org/10.3390/agronomy10091417
Chicago/Turabian StyleRocchetti, Lorenzo, Elisa Bellucci, Gaia Cortinovis, Valerio Di Vittori, Giovanna Lanzavecchia, Giulia Frascarelli, Laura Nanni, Andrea Del Gatto, Sandro Pieri, Lorella Mangoni, and et al. 2020. "The Development of a European and Mediterranean Chickpea Association Panel (EMCAP)" Agronomy 10, no. 9: 1417. https://doi.org/10.3390/agronomy10091417
APA StyleRocchetti, L., Bellucci, E., Cortinovis, G., Di Vittori, V., Lanzavecchia, G., Frascarelli, G., Nanni, L., Del Gatto, A., Pieri, S., Mangoni, L., Bitocchi, E., & Papa, R. (2020). The Development of a European and Mediterranean Chickpea Association Panel (EMCAP). Agronomy, 10(9), 1417. https://doi.org/10.3390/agronomy10091417








