Plant Functional Diversity, Climate and Grazer Type Regulate Soil Activity in Natural Grasslands
Abstract
1. Introduction
2. Materials and Methods
2.1. Site Description
2.2. Study Design
2.3. Vegetation and Soil Analyses
2.4. Data Analysis
2.4.1. Gradient Analysis and Variation Partitioning
2.4.2. Generalized Diversity-Interaction (GDI) Models
3. Results
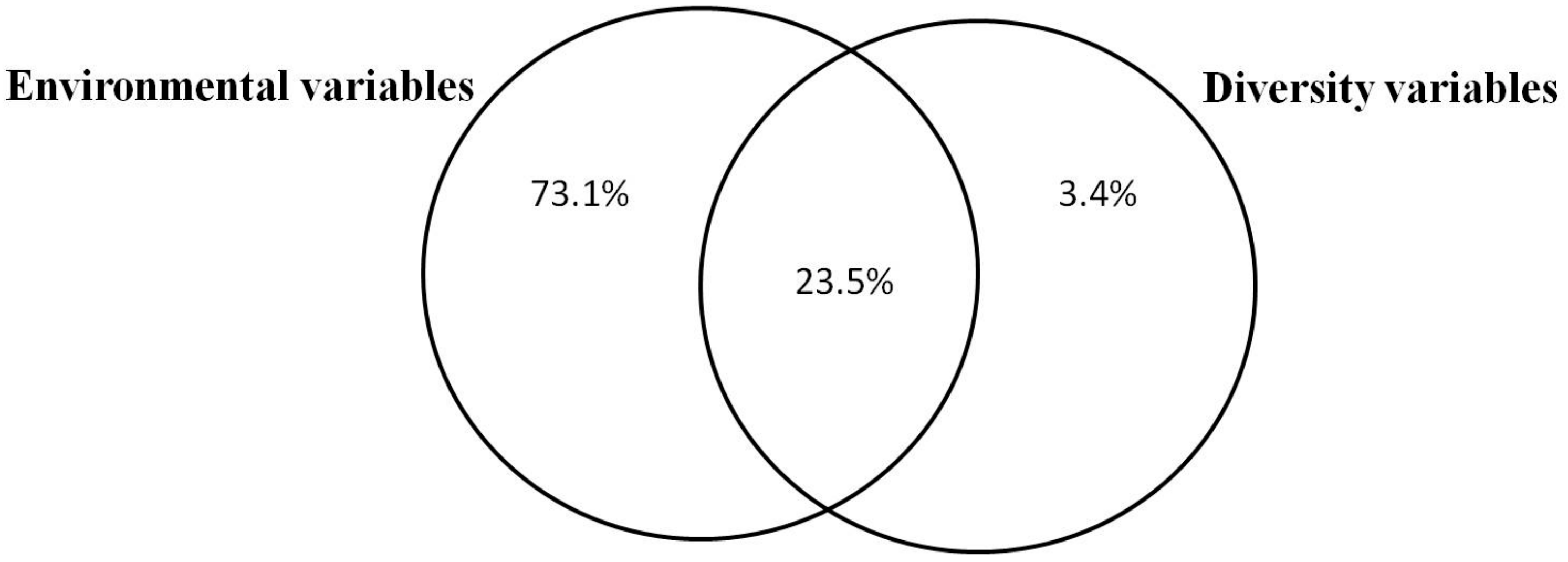
3.1. Relative Importance of Environmental Variables and Plant Functional Diversity on Soil Activity and Fertility
3.2. Plant Functional Diversity Effects on Soil Activity and Fertility
4. Discussion
4.1. Environmental Drivers of Soil Activity and Fertility
4.2. Plant Functional Diversity Effects on Soil Activity and Fertility
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Chapin, F.S.; Sala, O.E.; Huber-Sannwald, E. Global Biodiversity in a Changing Environment: Scenarios for the 21st Century; Springer: New York, NY, USA, 2001. [Google Scholar]
- FAO. FAOSTAT Statistical Databases; Food and Agricultural Organization of the United Nations: Rome, Italy, 2006. [Google Scholar]
- Bengtsson, J.; Bullock, J.M.; Egoh, B.; Everson, C.; Everson, T.; O’Connor, T.; O’Farrell, P.J.; Smith, H.G.; Lindborg, R. Grasslands—More important for ecosystem services than you might think. Ecosphere 2019, 10. [Google Scholar] [CrossRef]
- Lemaire, G.; Hodgson, J.; Chabbi, A. Grassland Productivity and Ecosystem Services; CAB International: Wallingford, UK; Cambridge, MA, USA, 2011; ISBN 978-1-84593-809-3. [Google Scholar]
- Noss, R.F. Biological hotspots and endangered ecosystems. In Forgotton Grasslands of the South-Natural History and Conservation; Island Press: Washington, DC, USA, 2013; pp. 73–115. ISBN 978-1-59726-362-7. [Google Scholar]
- Aguiar, M.R. Biodiversity in grasslands: Current changes and scenarios for the future. In Grasslands: Developments Opportunities Perspectives; Reynolds, S.G., Frame, J., Eds.; Science Publishers, Inc.: Einfield, NH, USA, 2005; pp. 261–280. [Google Scholar]
- Fry, E.L.; Manning, P.; Allen, D.G.P.; Hurst, A.; Everwand, G.; Rimmler, M.; Power, S.A. Plant functional group composition modifies the effects of precipitation change on grassland ecosystem function. PLoS ONE 2013, 8, e57027. [Google Scholar] [CrossRef] [PubMed]
- Sebastià, M.-T.; Kirwan, L.; Connolly, J. Strong shifts in plant diversity and vegetation composition in grassland shortly after climatic change. J. Veg. Sci. 2008, 19, 299–306. [Google Scholar] [CrossRef]
- Sebastià, M.T. Plant guilds drive biomass response to global warming and water availability in subalpine grassland. J. Appl. Ecol. 2007, 44, 158–167. [Google Scholar] [CrossRef]
- Sanz-Elorza, M.; Dana, E.D.; González, A.; Sobrino, E. Changes in the high-mountain vegetation of the central Iberian Peninsula as a probable sign of global warming. Ann. Bot. 2003, 92, 273–280. [Google Scholar] [CrossRef]
- Casasús, I.; Bernués, A.; Sanz, A.; Villalba, D.; Riedel, J.L.; Revilla, R. Vegetation dynamics in Mediterranean forest pastures as affected by beef cattle grazing. Agric. Ecosyst. Environ. 2007, 121, 365–370. [Google Scholar] [CrossRef]
- Mills, R.T.E.; Gavazov, K.S.; Spiegelberger, T.; Johnson, D.; Buttler, A. Diminished soil functions occur under simulated climate change in a sup-alpine pasture, but heterotrophic temperature sensitivity indicates microbial resilience. Sci. Total Environ. 2014, 473–474, 465–472. [Google Scholar] [CrossRef]
- Craine, J.M.; Nippert, J.B.; Elmore, A.J.; Skibbe, A.M.; Hutchinson, S.L.; Brunsell, N.A. Timing of climate variability and grassland productivity. Proc. Natl. Acad. Sci. USA 2012, 109, 3401–3405. [Google Scholar] [CrossRef]
- Brinkman, R.; Sombroek, W.G. The effects of global change on soil conditions in relation to plant growth and food produciton. In Global Climate Change and Agricultural Production. Direct and Indirect Effects of Changing Hydrological, Pedological and Plant Physiological Processes; Bazzaz, F., Ed.; Food and Agriculture Organization of the United Nations (FAO): Rome, Italy; John Wiley & Sons: Chichester, UK, 1996. [Google Scholar]
- Bot, A.; Benites, J. The Importance of Soil Organic Matter; Food and Agriculture Organization of the United Nations (FAO): Rome, Italy, 2005; ISBN 9251053669. [Google Scholar]
- Brookes, P. The soil microbial biomass: Concept, measurement and applications in soil ecosystem research. Microbes Environ. 2001, 16, 131–140. [Google Scholar] [CrossRef]
- Castro, H.F.; Classen, A.T.; Austin, E.E.; Norby, R.J.; Schadt, C.W. Soil microbial community responses to multiple experimental climate change drivers. Appl. Environ. Microbiol. 2010, 76, 999–1007. [Google Scholar] [CrossRef]
- Garten, C.T.; Classen, A.T.; Norby, R.J. Soil moisture surpasses elevated CO2 and temperature as a control on soil carbon dynamics in a multi-factor climate change experiment. Plant Soil 2009, 319, 85–94. [Google Scholar] [CrossRef]
- Le Roux, X.; Bardy, M.; Loiseau, P.; Louault, F. Stimulation of soil nitrification and denitrification by grazing in grasslands: Do changes in plant species composition matter? Oecologia 2003, 137, 417–425. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.; Yan, Y.; Fan, J.; Wang, X. Gross nitrification and denitrification in alpine grassland ecosystems on the Tibetan Plateau. Arct. Antarct. Alp. Res. 2012, 44, 188–196. [Google Scholar] [CrossRef]
- Emmett, B.A.; Beier, C.; Estiarte, M.; Tietema, A.; Kristensen, H.L.; Williams, D.; Peñuelas, J.; Schmidt, I.; Sowerby, A. The response of soil processes to climate change: Results from manipulation studies of shrublands across an environmental gradient. Ecosystems 2004, 7, 625–637. [Google Scholar] [CrossRef]
- Niklaus, P.A.; Wardle, D.A.; Tate, K.R. Effects of plant species diversity and composition on nitrogen cycling and the trace gas balance of soils. Plant Soil 2006, 282, 83–98. [Google Scholar] [CrossRef]
- Burns, R.G.; Dick, R.P. Enzymes in the Environment: Activity, Ecology, and Applications; Marcel Dekker, INC.: New York, NY, USA, 2002. [Google Scholar]
- Chróst, R. Microbial Enzymes in Aquatic Environments; Springer: New York, NY, USA, 1991; ISBN 978-1-4612-3090-8. [Google Scholar]
- Gianfreda, L. Enzymes of importance to rhizosphere processes. J. Soil Sci. Plant Nutr. 2015, 15, 283–306. [Google Scholar] [CrossRef]
- Sardans, J.; Peñuelas, J.; Estiarte, M. Changes in soil enzymes related to C and N cycle and in soil C and N content under prolonged warming and drought in a Mediterranean shrubland. Appl. Soil Ecol. 2008, 39, 223–235. [Google Scholar] [CrossRef]
- Steinweg, M.J.; Dukes, J.S.; Paul, E.A.; Wallenstein, M.D. Microbial responses to multi-factor climate change: Effects on soil enzymes. Front. Microbiol. 2013, 4, 1–11. [Google Scholar] [CrossRef]
- Wardle, D.A.; Bardgett, R.D.; Klironomos, J.N.; Setälä, H.; van der Putten, W.H.; Wall, D.H. Ecological Linkages between aboveground and belowground biota. Science 2004, 304, 1629–1633. [Google Scholar] [CrossRef]
- Pillar, V.D.; Blanco, C.C.; Müller, S.C.; Sosinski, E.E.; Joner, F.; Duarte, L.D.S. Functional redundancy and stability in plant communities. J. Veg. Sci. 2013, 24, 963–974. [Google Scholar] [CrossRef]
- Kang, S.; Ma, W.; Li, F.Y.; Zhang, Q.; Niu, J.; Ding, Y.; Han, F.; Sun, X. Functional redundancy instead of species redundancy determines community stability in a typical steppe of inner Mongolia. PLoS ONE 2015, 10, e0145605. [Google Scholar] [CrossRef] [PubMed]
- Kohler, M.; Devaux, C.; Grigulis, K.; Leitinger, G.; Lavorel, S.; Tappeiner, U. Plant functional assemblages as indicators of the resilience of grassland ecosystem service provision. Ecol. Indic. 2017, 73, 118–127. [Google Scholar] [CrossRef] [PubMed]
- Ostle, N.J.; Smith, P.; Fisher, R.; Woodward, F.I.; Fisher, J.B.; Smith, J.U.; Galbraith, D.; Levy, P.; Meir, P.; Mcnamara, N.P.; et al. Plant–soil interactions and the carbon cycle Integrating plant–soil interactions into global carbon cycle models. J. Ecol. 2009, 97, 851–863. [Google Scholar] [CrossRef]
- Wang, M.; Moore, T.R. Carbon, nitrogen, phosphorus, and potassium stoichiometry in an ombrotrophic peatland reflects plant functional type. Ecosystems 2014, 17, 673–684. [Google Scholar] [CrossRef]
- Wullschleger, S.D.; Epstein, H.E.; Box, E.O.; Euskirchen, E.S.; Goswami, S.; Iversen, C.M.; Kattge, J.; Norby, R.J.; Van Bodegom, P.M.; Xu, X. Plant functional types in Earth system models: Past experiences and future directions for application of dynamic vegetation models in high-latitude ecosystems. Ann. Bot. 2014, 114, 1–16. [Google Scholar] [CrossRef]
- Loreau, M.; Naeem, S.; Inchausti, P.; Bengtsson, J.; Grime, J.P.; Hector, A.; Hooper, D.U.; Huston, M.A.; Raffaelli, D.; Schmid, B.; et al. Biodiversity and ecosystem functioning: Current knowledge and future challenges. Science 2001, 294, 804–808. [Google Scholar] [CrossRef]
- Kirwan, L.; Lüscher, A.; Sebastià, M.T.; Finn, J.A.; Collins, R.P.; Porqueddu, C.; Helgadottir, A.; Baadshaug, O.H.; Brophy, C.; Coran, C.; et al. Evenness drives consistent diversity effects in intensive grassland systems across 28 European sites. J. Ecol. 2007, 95, 530–539. [Google Scholar] [CrossRef]
- Tilman, D.; Lehman, C.L.; Thomson, K.T. Plant diversity and ecosystem productivity: Theoretical considerations. Proc. Natl. Acad. Sci. USA 1997, 94, 1857–1861. [Google Scholar] [CrossRef]
- Kirwan, L.; Connolly, J.; Finn, J.; Brophy, C. Diversity-interaction modeling: Estimating contributions of species identities and interactions to ecosystem function. Ecology 2009, 90, 2032–2038. [Google Scholar] [CrossRef]
- Connolly, J.; Bell, T.; Bolger, T.; Brophy, C.; Carnus, T.; Finn, J.A.; Kirwan, L.; Isbell, F.; Levine, J.; Lüscher, A.; et al. An improved model to predict the effects of changing biodiversity levels on ecosystem function. J. Ecol. 2013, 101, 344–355. [Google Scholar] [CrossRef]
- Neilson, R.P.; Pitelka, L.F.; Solomon, A.M.; Nathan, R.; Midgley, G.F.; Fragoso, J.M.V.; Lischke, H.; Thompson, K. Forecasting regional to global plant migration in response to climate change. Bioscience 2005, 55, 749. [Google Scholar] [CrossRef]
- Pold, G.; DeAngelis, K. Up against the wall: The effects of climate warming on soil microbial diversity and the potential for feedbacks to the carbon cycle. Diversity 2013, 5, 409–425. [Google Scholar] [CrossRef]
- Classen, A.E.; Sundqvist, M.K.; Henning, J.A.; Newman, G.S.; M Moore, J.A.; Cregger, M.A.; Moorhead, L.C.; Patterson, C.M. Direct and indirect effects of climate change on soil microbial and soil microbial-plant interactions: What lies ahead? Ecosphere 2015, 6, 1–21. [Google Scholar] [CrossRef]
- Bru, D.; Ramette, A.; Saby, N.P.; Dequiedt, S.; Ranjard, L.; Jolivet, C.; Arrouays, D.; Philippot, L. Determinants of the distribution of nitrogen-cycling microbial communities at the landscape scale. ISME J. 2011, 5, 532–542. [Google Scholar] [CrossRef]
- Fierer, N.; Jackson, R.B. The diversity and biogeography of soil bacterial communities. Proc. Natl. Acad. Sci. USA 2006, 103, 626–631. [Google Scholar] [CrossRef] [PubMed]
- Jiao, F.; Shi, X.; Han, F.; Yuan, Z. Increasing aridity, temperature and soil pH induce soil C-N-P imbalance in grasslands. Sci. Rep. 2016, 6, 19601. [Google Scholar] [CrossRef]
- De Bolòs, O.; Vigo, J. Flora dels Països Catalans, Licopodiàcies-Capparàcies; Editorial Barcino: Barcelona, Spain, 1984; ISBN 8472265919. [Google Scholar]
- Carreras, J.; Carrillo, E.; Font, X.; Ninot, J.M.; Masalles, R.M.; Vilar, L. Vegetaciò arbustiva i herbàcia (Prats i Pastures). In Manual dels hàbitats de Catalunya-Volum 5; Vigo, J., Carreras, J., Ferrè, A., Eds.; Generalitat de Catalunya: Barcelona, Spain, 2006. [Google Scholar]
- San Emeterio, L.; Múgica, L.; Ugarte, M.D.; Goicoa, T.; Canals, R.M. Sustainability of traditional pastoral fires in highlands under global change: Effects on soil function and nutrient cycling. Agric. Ecosyst. Environ. 2016, 235, 155–163. [Google Scholar] [CrossRef]
- Black, C.A. Methods of Soil Analysis: Part I, Physical and Mineralogical Properties; American Society of Agronomy, Inc. & Soil Science Society of America, Inc.: Madison, WI, USA, 1965. [Google Scholar]
- Tian, L.; Dell, E.; Shi, W. Chemical composition of dissolved organic matter in agroecosystems: Correlations with soil enzyme activity and carbon and nitrogen mineralization. Appl. Soil Ecol. 2010, 46, 426–435. [Google Scholar] [CrossRef]
- Popova, I.E.; Deng, S. A high-throughput microplate assay for simultaneous colorimetric quantification of multiple enzyme activities in soil. Appl. Soil Ecol. 2010, 45, 315–318. [Google Scholar] [CrossRef]
- Kandeler, E.; Gerber, H. Short-term assay of soil urease activity using colorimetric determination of ammonium. Biol. Fertil. Soils 1988, 6, 68–72. [Google Scholar] [CrossRef]
- Rodríguez-Loinaz, G.; Onaindia, M.; Amezaga, I.; Mijangos, I.; Garbisu, C. Relationship between vegetation diversity and soil functional diversity in native mixed-oak forests. Soil Biol. Biochem. 2008, 40, 49–60. [Google Scholar] [CrossRef]
- Ter Braak, C.J.F.; Šmilauer, P. Canoco Reference Manual and User’s Guide: Software for Ordination, Version 5.0; Microcomputer Power: Ithaca, NY, USA, 2012. [Google Scholar]
- Rodríguez, A.; Canals, R.M.; Plaixats, J.; Albanell, E.; Debouk, H.; Garcia-Pausas, J.; San Emeterio, L.; Jimenez, J.J.; Sebstià, M.-T. Interactions between biogeochemical and management factors explain soil organic carbon in Pyrenean grasslands. Biogeosci. Discuss. 2020, 1–56. [Google Scholar] [CrossRef]
- Oksanen, J.; Blanchet, F.G.; Kindt, R.; Legendre, P.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; Stevens, M.H.H.; Wagner, H. Vegan: Community Ecology Package. R Package Version 2.0-7. 2013. Available online: http://CRAN.Rproject.org/package=vegan (accessed on 30 August 2020).
- Heiberger, R.M.; Holland, B. Statistical Analysis and Data Display: An Intermediate Course with Examples in S-Plus, R, and SAS. Springer Texts in Statistics; Springer Science + Business Media: New York, NY, USA, 2004; ISBN 0-387-40270-5. [Google Scholar]
- Symonds, M.R.E.; Moussalli, A. A brief guide to model selection, multimodel inference and model averaging in behavioural ecology using Akaike’s information criterion. Behav. Ecol. Sociobiol. 2011, 65, 13–21. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2013; Available online: http://www.R-project.org/ (accessed on 30 August 2020).
- Lenth, R.V. Response-surface methods in R, Using rsm. J. Stat. Softw. 2009, 32, 1–17. [Google Scholar] [CrossRef]
- Sarkar, D. Lattice: Multivariate Data Visualization with R; Springer: New York, NY, USA, 2008; ISBN 978-0-387-75968-5. [Google Scholar]
- Bates, D.; Maechler, M.; Bolker, B. lme4: Linear Mixed-Effects Models Using S4 Classes. J. Stat. Softw. 2012, 67. [Google Scholar]
- Gelman, A.; Su, Y. Package arm: Data Analysis Using Regression and Multilevel/Hierarchical Models; Cambridge University Press: New York, NY, USA, 2013; Available online: https://CRAN.R-project.org/package=arm (accessed on 30 August 2020).
- Neuwirth, E. RColorBrewer: ColorBrewer Palettes, Cynthia Brewer, Mark Harrower and The Pennsylvania State University. 2011. Available online: https://CRAN.R-project.org/package=RColorBrewer (accessed on 30 August 2020).
- Yang, N.; Zou, D.; Yang, M.; Lin, Z. Variations in soil microbial biomass carbon and soil dissolved organic carbon in the re-vegetation of hilly slopes with purple soil. PLoS ONE 2016, e0166536. [Google Scholar] [CrossRef][Green Version]
- Paul, K.I.; Polglase, P.J.; O’Connell, A.M.; Carlyle, J.C.; Smethurst, P.J.; Khanna, P.K. Defining the relation between soil water content and net nitrogen mineralization. Eur. J. Soil Sci. 2003, 54, 39–47. [Google Scholar] [CrossRef]
- Mueller, K.E.; Hobbie, S.E.; Tilman, D.; Reich, P.B. Effects of plant diversity, N fertilization, and elevated carbon dioxide on grassland soil N cycling in a long-term experiment. Glob. Chang. Biol. 2013, 19, 1249–1261. [Google Scholar] [CrossRef]
- Zhalnina, K.; Dörr de Quadros, P.; Camargo, F.O.; Triplett, E.W. Drivers of archaeal ammonia-oxidizing communities in soil. Front. Microbiol. 2012, 3, 1–9. [Google Scholar] [CrossRef]
- Troy, C.; Tang, J. Effects of Temperature and Moisture Stress on Nitrous Oxide Production in Agricultural Soil; Villanova University: Vilanova, PA, USA, 2011. [Google Scholar]
- Sardans, J.; Peñuelas, J.; Estiarte, M. Seasonal patterns of root-surface phosphatase activities in a Mediterranean shrubland. Responses to experimental warming and drought. Biol. Fertil. Soils 2007, 43, 779–786. [Google Scholar] [CrossRef]
- Shen, J.; Yuan, L.; Zhang, J.; Li, H.; Bai, Z.; Chen, X.; Zhang, W.; Zhang, F. Phosphorus dynamics: From soil to plant. Plant Physiol. 2011, 156, 997–1005. [Google Scholar] [CrossRef] [PubMed]
- Leirós, M.C.; Trasar-Cepeda, C.; García-Fernández, F.; Gil-Sostres, F. Defining the validity of a biochemical index of soil quality. Biol. Fertil. Soils 1999, 30, 140–146. [Google Scholar] [CrossRef]
- Sardans, J.; Peñuelas, J. Drought decreases soil enzyme activity in a Mediterranean Quercus ilex L. forest. Soil Biol. Biochem. 2005, 37, 455–461. [Google Scholar] [CrossRef]
- Lauber, C.L.; Hamady, M.; Knight, R.; Fierer, N. Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl. Environ. Microbiol. 2009, 75, 5111–5120. [Google Scholar] [CrossRef] [PubMed]
- Andersson, S.; Nilsson, S.I.; Saetre, P. Leaching of dissolved organic carbon (DOC) and dissolved organic nitrogen (DON) in mor humus as affected by temperature and pH. Soil Biol. Biochem. 2000, 32, 1–10. [Google Scholar] [CrossRef]
- Fröberg, M.; Berggren, D.; Bergkvist, B.; Bryant, C.; Mulder, J. Concentration and fluxes of dissolved organic carbon (DOC) in three Norway spruce stands along a climatic gradient in Sweden. Biogeochemistry 2006, 77, 1–23. [Google Scholar] [CrossRef]
- Kalbitz, K.; Schmerwitz, J.; Schwesig, D.; Matzner, E. Biodegradation of soil-derived dissolved organic matter as related to its properties. Geoderma 2003, 113, 273–291. [Google Scholar] [CrossRef]
- Solinger, S.; Kalbitz, K.; Matzner, E. Controls on the dynamics of dissolved organic carbon and nitrogen in a Central European deciduous forest. Biogeochemistry 2001, 327–349. [Google Scholar] [CrossRef]
- Kaiser, K.; Guggenberger, G.; Haumaier, L.; Zech, W. The composition of dissolved organic matter in forest soil solutions: Changes induced by seasons and passage through the mineral soil. Org. Geochem. 2002, 33, 307–318. [Google Scholar] [CrossRef]
- Butterbach-Bahl, K.; Baggs, E.M.; Dannenmann, M.; Kiese, R.; Zechmeister-Boltenstern, S. Nitrous oxide emissions from soils: How well do we understand the processes and their controls? Philos. Trans. R. Soc. Lond. B Biol. Sci. 2013, 368, 20130122. [Google Scholar] [CrossRef]
- Haynes, R.J. Nitrification. In Mineral Nitrogen in the Plant-Soil System; Haynes, R.J., Ed.; Academic Press: New York, NY, USA, 1986; pp. 127–165. [Google Scholar]
- De Boer, W.; Kowalchuk, G.A. Nitrification in acid soils: Micro-organisms and mechanisms. Soil Biol. Biochem. 2001, 33, 853–866. [Google Scholar] [CrossRef]
- Burns, R.G.; DeForest, J.L.; Marxsen, J.; Sinsabaugh, R.L.; Stromberger, M.E.; Wallenstein, M.D.; Weintraub, M.N.; Zoppini, A. Soil enzymes in a changing environment: Current knowledge and future directions. Soil Biol. Biochem. 2013, 58, 216–234. [Google Scholar] [CrossRef]
- Caldwell, B.A. Enzyme activities as a component of soil biodiversity: A review. Pedobiologia 2005, 49, 637–644. [Google Scholar] [CrossRef]
- Olivera, N.L.; Prieto, L.; Carrera, A.L.; Cisneros, H.S.; Bertiller, M.B. Do soil enzymes respond to long-term grazing in an arid ecosystem? Plant Soil 2014, 1–14, 35–48. [Google Scholar] [CrossRef]
- Prieto, L.H.; Bertiller, M.B.; Carrera, A.L.; Olivera, N.L. Soil enzyme and microbial activities in a grazing ecosystem of Patagonian Monte, Argentina. Geoderma 2011, 162, 281–287. [Google Scholar] [CrossRef]
- Sebastià, M.T.; de Bello, F.; Puig, L.; Taull, M. Grazing as a factor structuring grasslands in the Pyrenees. Appl. Veg. Sci. 2008, 11, 215–222. [Google Scholar] [CrossRef]
- Tabatabai, M.A. Soil enzymes. In Methods of Soil Analysis: Microbiological and biochemical Properties Part 2; Weaver, R.W., Angle, J.R., Bottomley, P.S., Eds.; Soil Science Society America: Madison, WI, USA, 1994; pp. 775–833. [Google Scholar]
- Hao, Y.; He, Z. Effects of grazing patterns on grassland biomass and soil environments in China: A meta-analysis. PLoS ONE 2019, 14, e0215223. [Google Scholar] [CrossRef]
- Ding, K.; Zhong, L.; Xin, X.P.; Xu, Z.H.; Kang, X.M.; Liu, W.J.; Rui, Y.C.; Jiang, L.L.; Tang, L.; Wang, Y.F. Effect of grazing on the abundance of functional genes associated with N cycling in three types of grassland in Inner Mongolia. J. Soils Sediments 2015, 15, 683–693. [Google Scholar] [CrossRef]
- Klumpp, K.; Fontaine, S.; Attard, E.; Le Roux, X.; Gleixner, G.; Soussana, J.F. Grazing triggers soil carbon loss by altering plant roots and their control on soil microbial community. J. Ecol. 2009, 97, 876–885. [Google Scholar] [CrossRef]
- Wardle, D.A.; Yeates, G.W.; Williamson, W.; Bonner, K.I. The response of a three trophic level soil food web to the identity and diversity of plant species and functional groups. Oikos 2003, 102, 45–56. [Google Scholar] [CrossRef]
- Cui, H.-J.; Wang, G.-X.; Yang, Y.; Yang, Y.; Chang, R.-Y.; Ran, F. Soil microbial community composition and its driving factors in alpine grasslands along a mountain elevational gradient. J. Mt. Sci. 2016, 13, 1013–1023. [Google Scholar] [CrossRef]
- Canals, R.M.; Eviner, V.T.; Herman, D.J.; Chapin, F.S. Plant colonizers shape early N-dynamics in gopher-mounds. Plant Soil 2005, 276, 327–334. [Google Scholar] [CrossRef]
- Strecker, T.; Barnard, R.L.; Niklaus, P.A.; Scherer-Lorenzen, M.; Weigelt, A.; Scheu, S.; Eisenhauer, N. Effects of plant diversity, functional group composition, and fertilization on soil microbial properties in experimental grassland. PLoS ONE 2015, 10, e0125678. [Google Scholar] [CrossRef] [PubMed]
- Fang, S.; Liu, D.; Tian, Y.; Deng, S.; Shang, X. Tree species composition influences enzyme activities and microbial biomass in the Rhizosphere: A rhizobox approach. PLoS ONE 2013, 8, e61461. [Google Scholar] [CrossRef]
- Dijkstra, F.A.; West, J.B.; Hobbie, S.E.; Reich, P.B.; Trost, J. Plant diversity, CO2, and N influence inorganic and organic N leaching in grasslands. Ecology 2007, 88, 490–500. [Google Scholar] [CrossRef]
- Eisenhauer, A.N.; Beßler, H.; Engels, C.; Gleixner, G.; Habekost, M.; Milcu, A.; Partsch, S.; Sabais, A.C.W.; Scherber, C.; Steinbeiss, S.; et al. Plant diversity effects on soil microorganisms support the singular hypothesis. Ecology 2010, 91, 485–496. [Google Scholar] [CrossRef]
- Roscher, C.; Schumacher, J.; Gubsch, M.; Lipowsky, A.; Weigelt, A.; Buchmann, N.; Schmid, B.; Schulze, E.D. Using plant functional traits to explain diversity-productivity relationships. PLoS ONE 2012, 7, e36760. [Google Scholar] [CrossRef]
- Moran-Zuloaga, D.; Dippold, M.; Glaser, B.; Kuzyakov, Y. Organic nitrogen uptake by plants: Reevaluation by position-specific labeling of amino acids: Reevaluation of organic N uptake by plants by position-specific labeling. Biogeochemistry 2015, 125, 359–374. [Google Scholar] [CrossRef]
- Nyfeler, D.; Huguenin-Elie, O.; Suter, M.; Frossard, E.; Lüscher, A. Grass-legume mixtures can yield more nitrogen than legume pure stands due to mutual stimulation of nitrogen uptake from symbiotic and non-symbiotic sources. Agric. Ecosyst. Environ. 2011, 140, 155–163. [Google Scholar] [CrossRef]
- Suter, M.; Connolly, J.; Finn, J.A.; Loges, R.; Kirwan, L.; Sebastià, M.T.; Lüscher, A. Nitrogen yield advantage from grass-legume mixtures is robust over a wide range of legume proportions and environmental conditions. Glob. Chang. Biol. 2015, 21, 2424–2438. [Google Scholar] [CrossRef]
- Oelmann, Y.; Buchmann, N.; Gleixner, G.; Habekost, M.; Roscher, C.; Rosenkranz, S.; Schulze, E.D.; Steinbeiss, S.; Temperton, V.M.; Weigelt, A.; et al. Plant diversity effects on aboveground and belowground N pools in temperate grassland ecosystems: Development in the first 5 years after establishment. Glob. Biogeochem. Cycles 2011, 25, 1–11. [Google Scholar] [CrossRef]
- Liang, Y.; Yang, Y.; Yang, C.; Shen, Q.; Zhou, J.; Yang, L. Soil enzymatic activity and growth of rice and barley as influenced by organic manure in an anthropogenic soil. Geoderma 2003, 115, 149–160. [Google Scholar] [CrossRef]
- Zhang, T.; Wan, S.; Kang, Y.; Feng, H. Urease activity and its relationships to soil physiochemical properties in a highly saline-sodic soil. J. Soil Sci. Plant Nutr. 2014, 14, 304–315. [Google Scholar] [CrossRef]
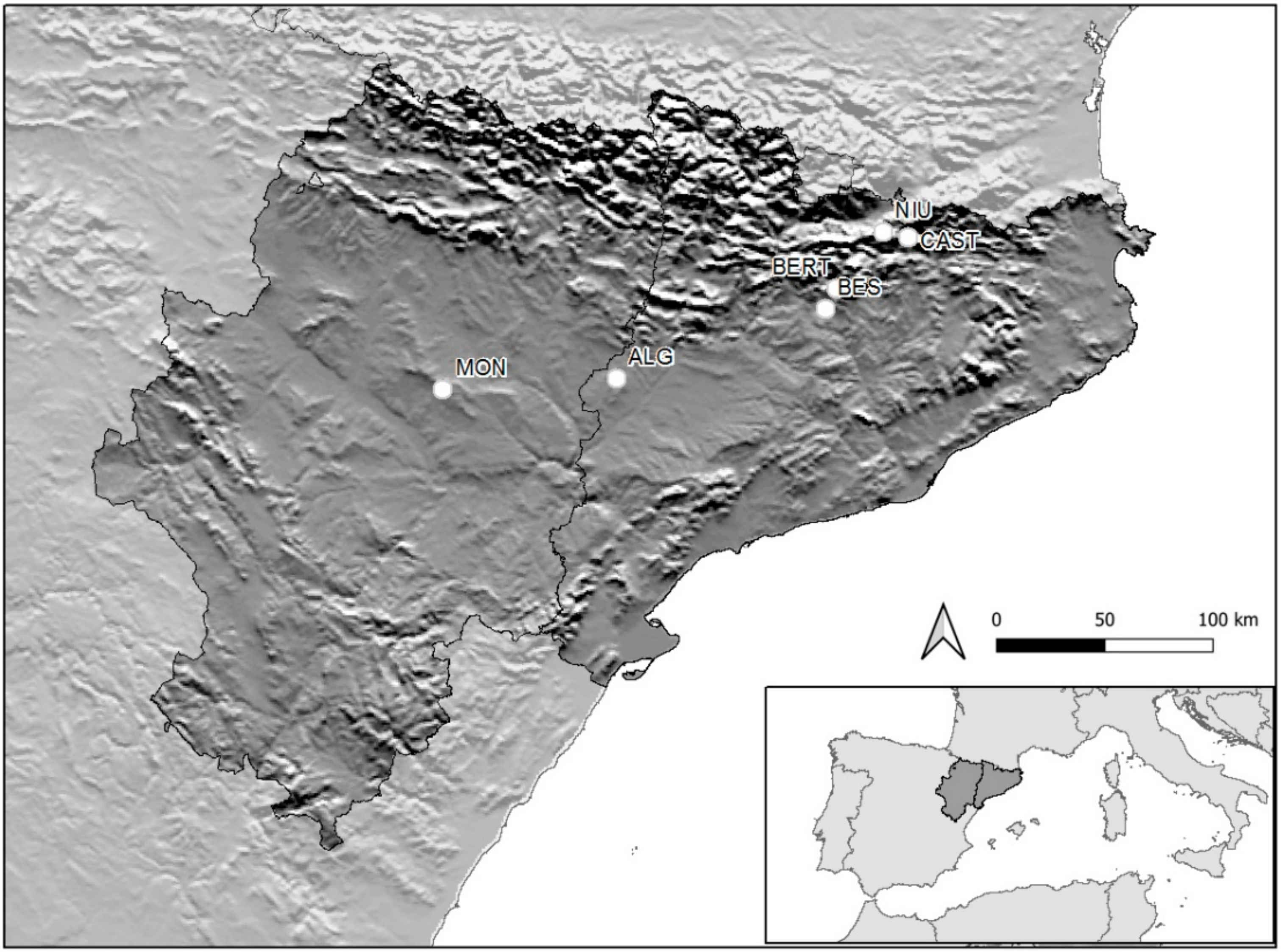


| Variables | ALG | MON | BES | BERT | CAST | NIU |
|---|---|---|---|---|---|---|
| Latitude | 41°43′46,60″ | 41°39′45,62″ | 42°1′13,07″ | 42°5′57,33″ | 42°18′18,84″ | 42°19′18,91″ |
| Longitude | 0°31′30″ | −1°37′22,15″ | 1°36′23,48″ | 1°39′48,63″ | 2°1′57,58″ | 1°54′4,47″ |
| Altitude (m) | 334 | 657 | 712 | 1276 | 1850 | 2479 |
| MAP (mm) | 421.4 | 225.2 | 720.5 | 954.8 | 1199 | 1302 |
| MAT (°C) | 13.9 | 12.6 | 11.7 | 8.7 | 5.4 | 2.4 |
| Lang’s index | 30.32 | 17.87 | 61.58 | 109.75 | 222.04 | 542.5 |
| MST (°C) | 22.8 | 23 | 20 | 18.2 | 15.3 | 13.8 |
| MSP (mm) | 95.7 | 93.8 | 237.5 | 465.2 | 319.1 | 278.8 |
| MTmin (°C) | 0 | 1 | 0 | −1 | −5 | −7 |
| MTmax (°C) | 28 | 28 | 25 | 24 | 23 | 22 |
| TSIS | 8.7 | 8.6 | 8.2 | 7.8 | 7.3 | 7.4 |
| pH | 8.57 ± 0.1 | 8.46 ± 0.01 | 6.77 ± 0.19 | 6.87 ± 0.06 | 6.02 ± 0.32 | 6.1 ± 0.15 |
| Soil moisture (%) | 5 ± 1 | 18 ± 2 | 17 ± 2 | 11 ± 1 | 42 ± 2 | 34 ± 2 |
| Grazer type | Sheep | Sheep | Cattle | Cattle | Cattle, sheep | Cattle, horses |
| Grasses (%) | 12–98 | 7–99 | 18–75 | 6–95 | 37–97 | 16.3–86 |
| Forbs (%) | 0.2–88 | 0–93 | 19–79 | 0–77 | 2.6–62 | 6.5–81 |
| Legumes (%) | 0–0.4 | 0–0.2 | 0.1–48 | 1.2–74 | 0–45 | 0–44 |
| Variable | Conditional Term Effects | Simple Term Effects | ||||
|---|---|---|---|---|---|---|
| % Explained | F | Padj | % Explained | F | Padj | |
| Large-scale climatic and management factors | ||||||
| MTmin | 0.3 | 0.7 | n.s. | 57.9 | 63.2 | <0.001 |
| TSIS | 0.4 | 1 | n.s. | 69.3 | 104 | <0.001 |
| Sheep grazing | 2 | 4.9 | 0.06 | 0.5 | 0.2 | n.s. |
| Cattle grazing | 0.1 | 0.3 | n.s. | 34.6 | 24.3 | <0.001 |
| Site-specific soil factors | ||||||
| Moisture | 73 | 124 | 0.001 | 73 | 124 | <0.001 |
| pH | 2.8 | 6.4 | <0.05 | 58.5 | 65 | <0.001 |
| PFT main effects | ||||||
| Grass | 0.8 | 2.1 | n.s. | 0.4 | 0.2 | n.s. |
| Forb | 0.7 | 1.7 | n.s. | 1.4 | 0.6 | n.s. |
| Legume | 0.2 | 0.4 | n.s. | 0.8 | 0.4 | n.s. |
| PFT interactions | ||||||
| FG | 4.9 | 10 | <0.01 | 8.9 | 4.5 | 0.08 |
| FL | 0.6 | 1.5 | n.s. | 0.6 | 0.3 | n.s. |
| GL | 0.2 | 0.6 | n.s. | 5.2 | 2.5 | n.s. |
| Soil Parameter | Model | Adjusted r2 | p-Value |
|---|---|---|---|
| Model 1. Only environmental variables included | |||
| Nitrate | (+) MTmax *** (+) Cattle ** | 0.38 | <0.001 |
| Ammonium | (−) MTmax *** (−) MAP+ (+) Sheep *** (+) Cattle | 0.84 | <0.001 |
| DOC | (+) MTmax ** (−) MTmin *** (+) MAP ** + (1) Sheep *** (+) Cattle *** | 0.79 | <0.001 |
| MBC | (−) MTmax *** (+) MTmin * (−) MAP * (+) Sheep *** | 0.75 | <0.001 |
| Glucosidase | (+) MTmax * (−) MTmin (+) MAP+ (+) Cattle+ | 0.17 | <0.05 |
| Model 3. Environmental variables + PFT identity effects + PFT pairwise interactions | |||
| MBN | (+) MTmin *** (−) MTmax *** (−) MAP ** (+) Sheep *** (−) G (−) F (−) L (+) FG *(−) GL (+) FL (−) MTmin:G * (−) MTmin:F+ (−) MTmin:L | 0.88 | <0.001 |
| DON | (+) MTmin *** (−) MST ** (+) Sheep *** (−) G (−) F+ (−) L (+) FG (+) GL ** (+) FL | 0.60 | <0.001 |
| Phosphatase | (−) MTmin *** (+) Sheep ** (−) G (−) F+ (−) L (+) FG *** (−) GL (+) FL | 0.63 | <0.001 |
| Model 4. Environmental variables + PFT identity effects + evenness | |||
| Total N | (−) MTmin *** (−) MST ** (+) Sheep *** (−) G (−) F (−) L (+) Ek * | 0.84 | <0.001 |
| Urease | (+) MTmin ** (+) MTmax *** (−) MST+ (−) Sheep *** (+) Cattle *** (−) G (−) F ** (+) L (+) Ek ** | 0.79 | <0.001 |
| Nitrification Potential | (+) MTmin *** (+) MTmax *** (−) Sheep *** (−) G+ (−) F (−) L (+) Ek+ | 0.78 | <0.001 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Debouk, H.; San Emeterio, L.; Marí, T.; Canals, R.M.; Sebastià, M.-T. Plant Functional Diversity, Climate and Grazer Type Regulate Soil Activity in Natural Grasslands. Agronomy 2020, 10, 1291. https://doi.org/10.3390/agronomy10091291
Debouk H, San Emeterio L, Marí T, Canals RM, Sebastià M-T. Plant Functional Diversity, Climate and Grazer Type Regulate Soil Activity in Natural Grasslands. Agronomy. 2020; 10(9):1291. https://doi.org/10.3390/agronomy10091291
Chicago/Turabian StyleDebouk, Haifa, Leticia San Emeterio, Teresa Marí, Rosa M. Canals, and Maria-Teresa Sebastià. 2020. "Plant Functional Diversity, Climate and Grazer Type Regulate Soil Activity in Natural Grasslands" Agronomy 10, no. 9: 1291. https://doi.org/10.3390/agronomy10091291
APA StyleDebouk, H., San Emeterio, L., Marí, T., Canals, R. M., & Sebastià, M.-T. (2020). Plant Functional Diversity, Climate and Grazer Type Regulate Soil Activity in Natural Grasslands. Agronomy, 10(9), 1291. https://doi.org/10.3390/agronomy10091291





