Genome-Wide Identification and Expression Analysis of MIKC-Type MADS-Box Gene Family in Punica granatum L.
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Collection for Expression Analysis
2.2. Identification and Sequence Analysis of Pomegranate MIKC-Type MADS-Box Transcription Factors
2.3. Phylogenetic Analysis of Pomegranate MIKC-Type MADS-Box Family
2.4. Gene Structure Analysis and Motif Identification of Pomegranate MIKC-Type MADS-Box Family
2.5. Prediction Promoter Elements and Protein Interaction Network of Pomegranate MIKC-Type MADS-Box Family
2.6. Expression Analysis for Pomegranate MIKC-Type MADS-Box Family
3. Results
3.1. Identification and Sequence Analysis of Pomegranate MIKC-Type MADS-Box Family
3.2. Phylogenetic Analysis of Pomegranate MIKC-Type MADS-Box Family
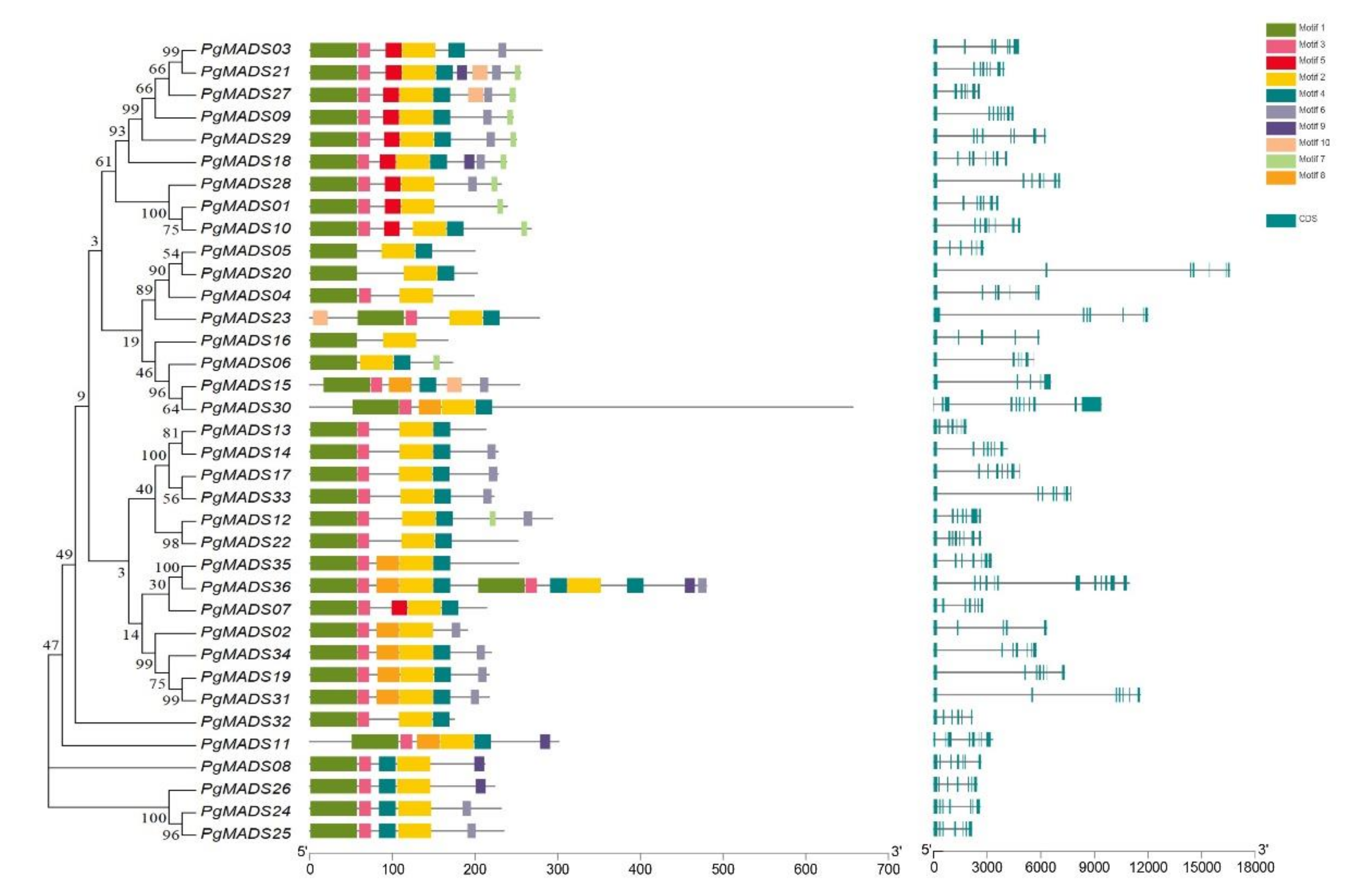
3.3. Gene Structure Analysis and Motif Identification of Pomegranate MIKC-Type MADS-Box Family
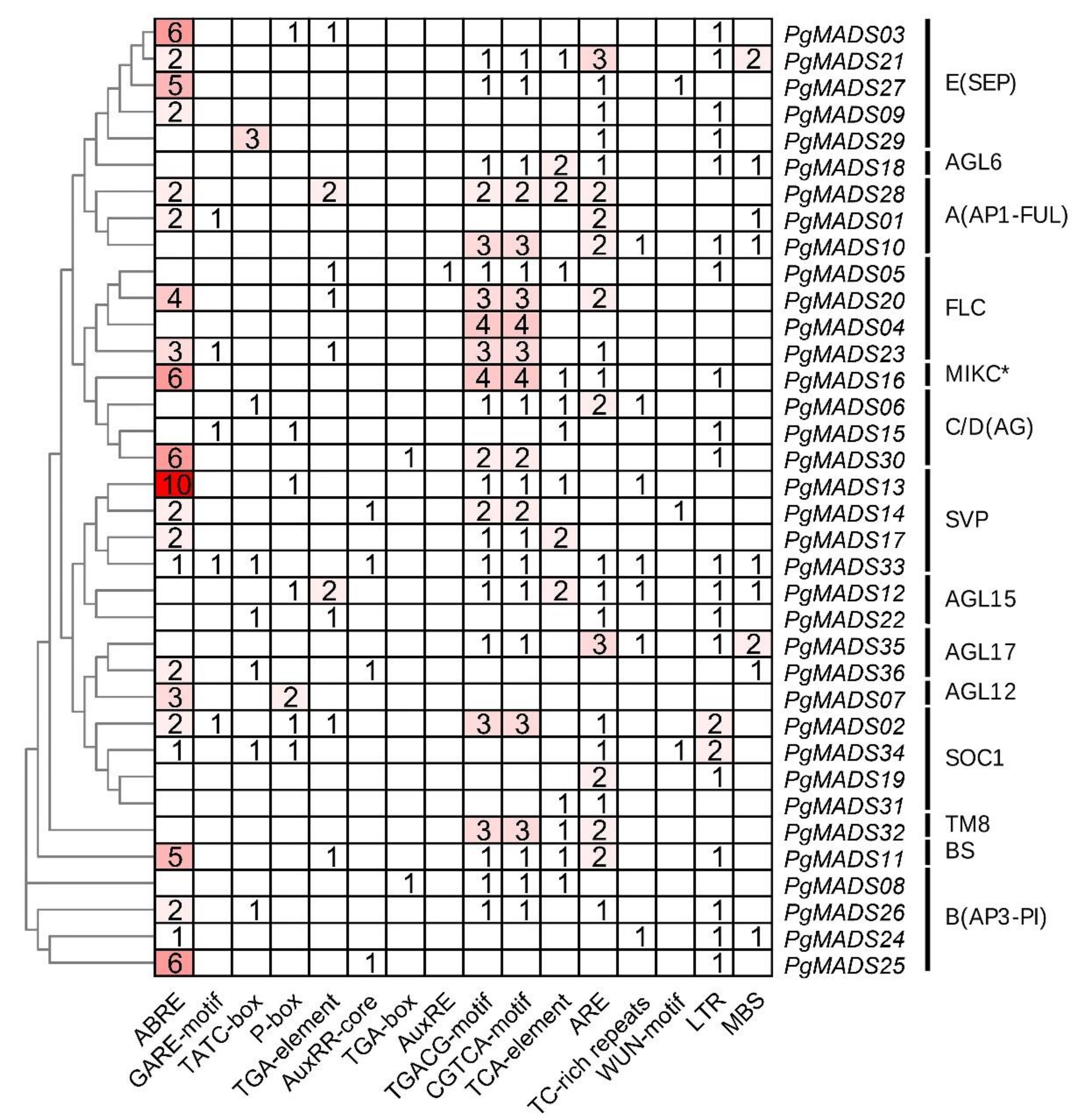
3.4. Prediction Promoter Elements of the Pomegranate MIKC-Type MADS-Box Family
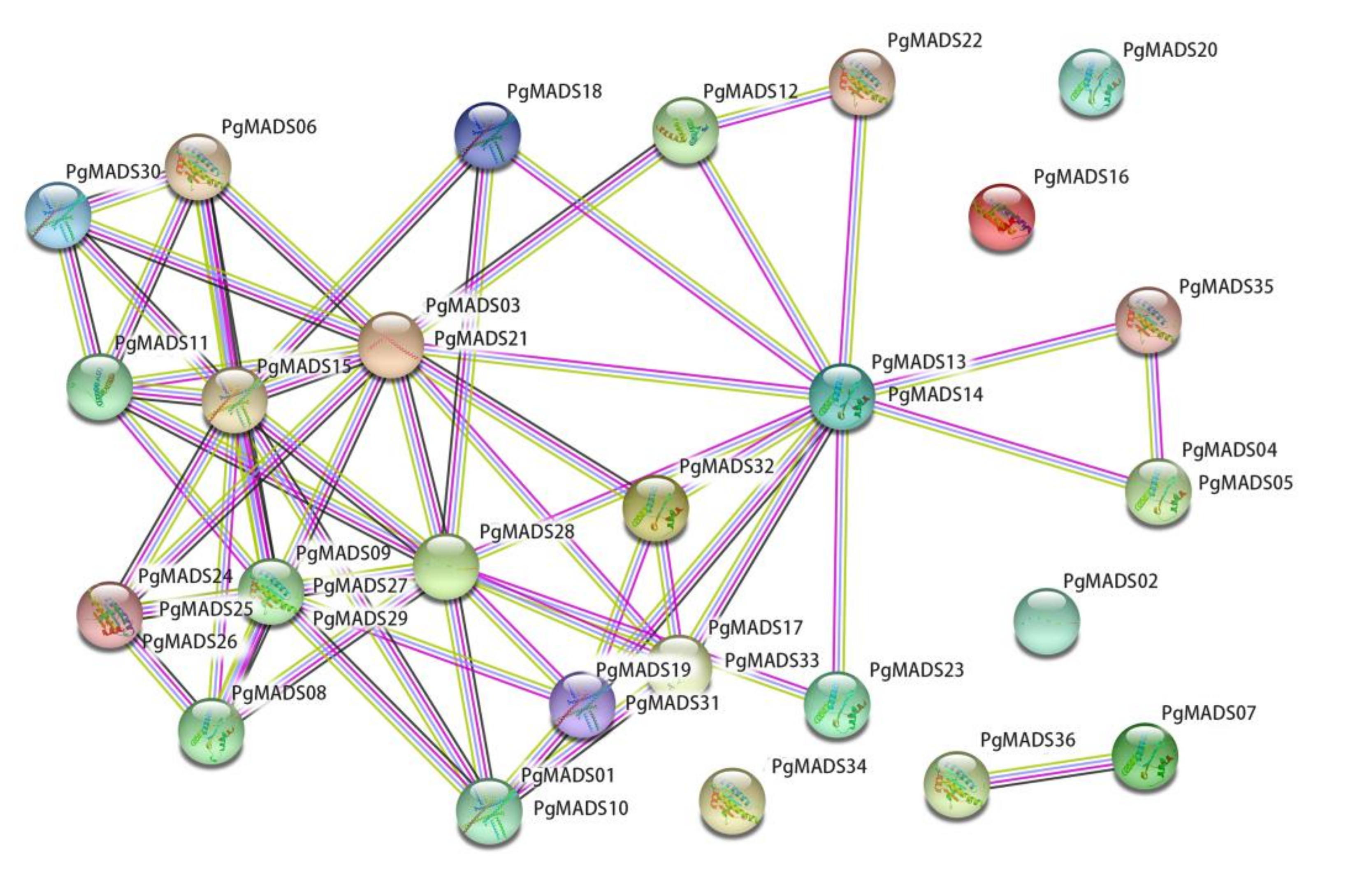
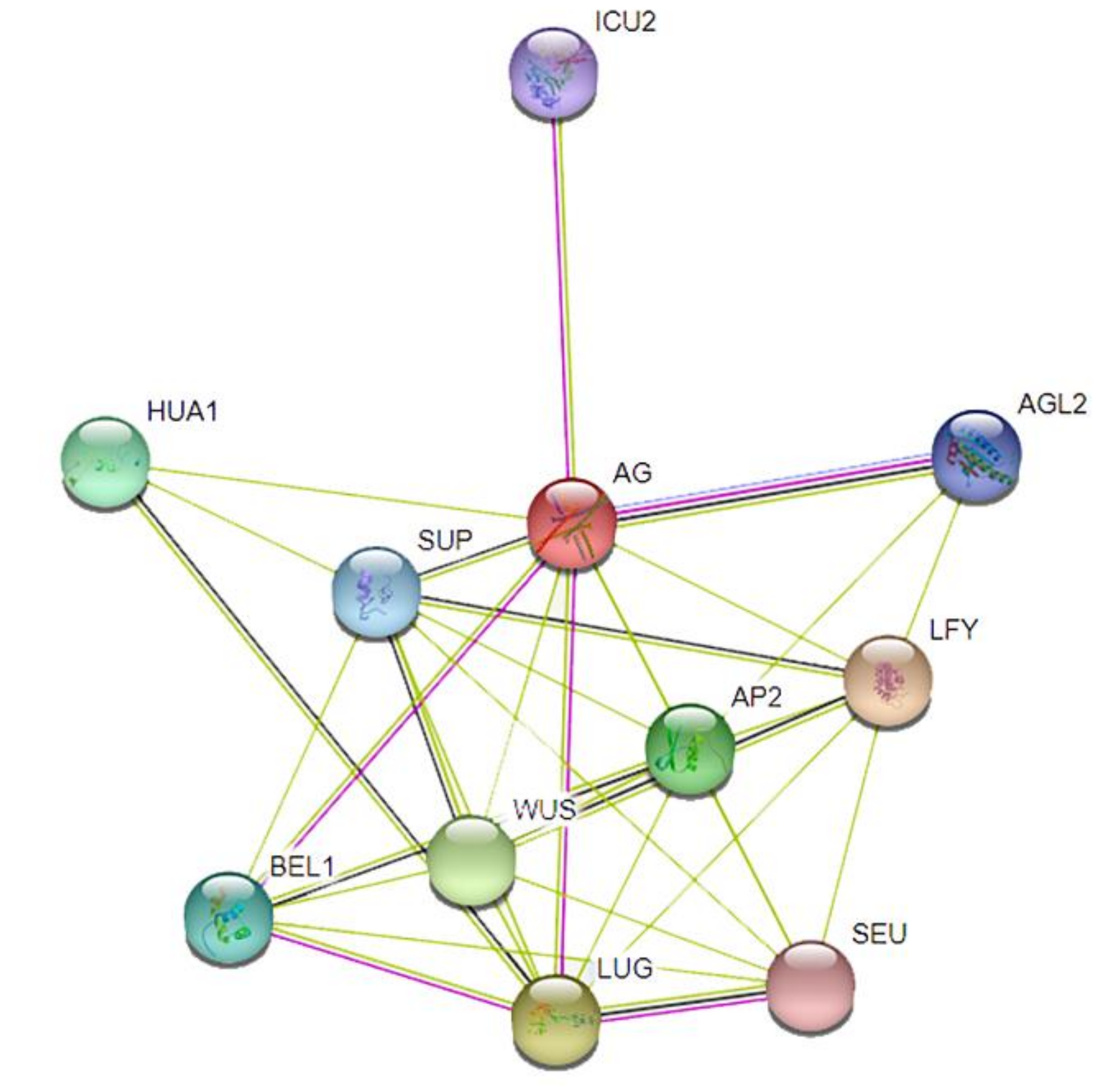
3.5. Protein Interaction Networks of the Pomegranate MIKC-Type MADS-Box Family
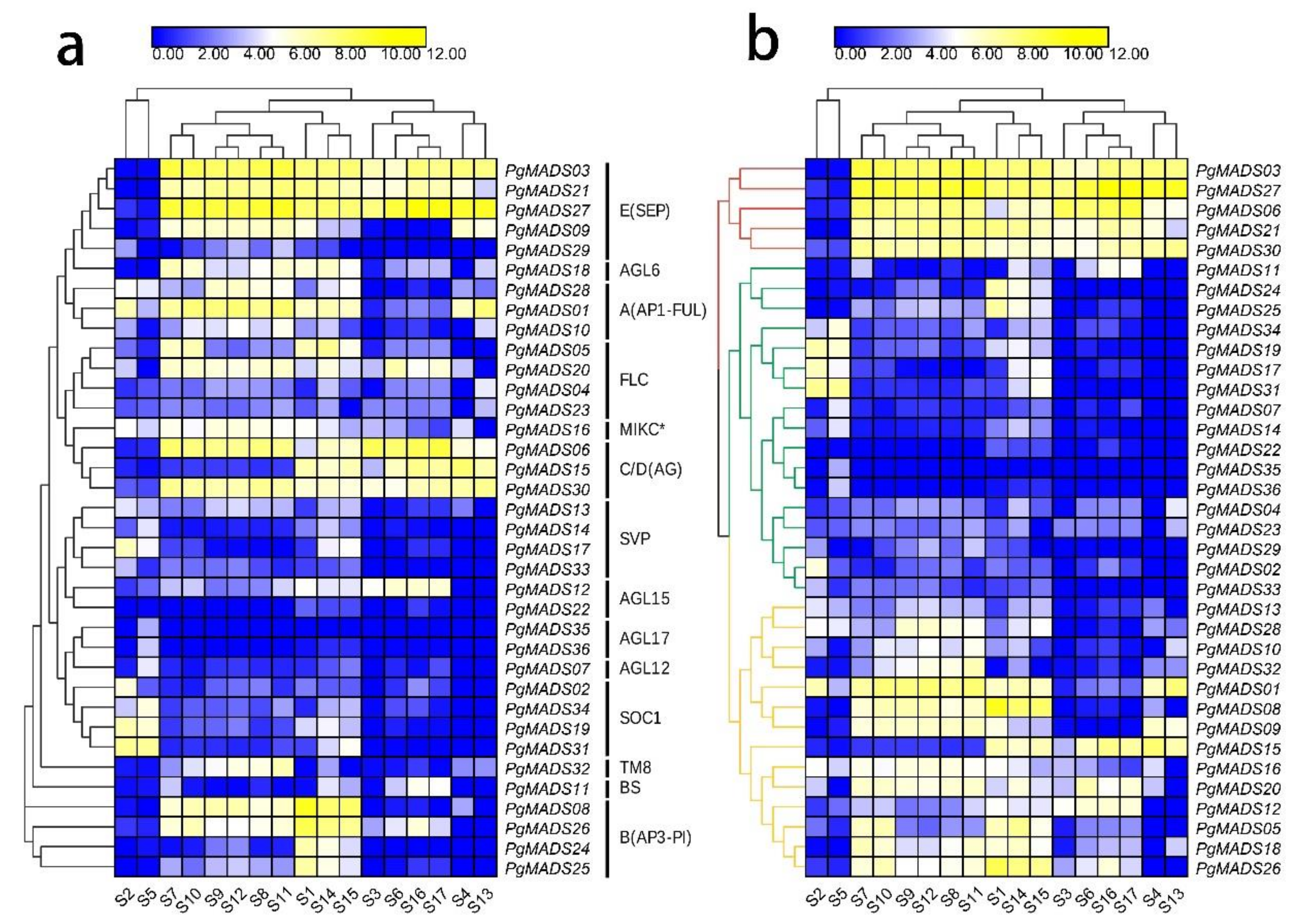
3.6. Expression Analysis for Pomegranate MIKC-Type MADS-Box Family
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Becker, A.; Theissen, G. The major clades of MADS-box genes and their role in the development and evolution of flowering plants. Mol. Phylogenet. Evol. 2003, 29, 464–489. [Google Scholar] [CrossRef]
- Messenguy, F.; Dubois, E. Role of MADS-box proteins and their cofactors in combinatorial control of gene expression and cell development. Gene 2003, 316, 1–21. [Google Scholar] [CrossRef]
- Alvarezbuylla, E.R.; Pelaz, S.; Liljegren, S.J.; Gold, S.E.; Burgeff, C.; Ditta, G.S.; De Pouplana, L.R.; Martinezcastilla, L.P.; Yanofsky, M.F. An ancestral MADS-box gene duplication occurred before the divergence of plants and animals. Proc. Natl. Acad. Sci. USA 2000, 97, 5328–5333. [Google Scholar] [CrossRef]
- Smaczniak, C.; Immink, R.G.H.; Angenent, G.C.; Kaufmann, K. Developmental and evolutionary diversity of plant MADS-domain factors: insights from recent studies. Development 2012, 139, 3081–3098. [Google Scholar] [CrossRef] [PubMed]
- Sommer, H.; Beltran, J.P.; Huijser, P.; Pape, H.; Lonnig, W.; Saedler, H.; Schwarzsommer, Z.D. A homeotic gene involved in the control of flower morphogenesis in Antirrhinum majus: the protein shows homology to transcription factors. EMBO J. 1990, 9, 605–613. [Google Scholar] [CrossRef] [PubMed]
- Yanofsky, M.F.; Ma, H.; Bowman, J.L.; Drews, G.N.; Feldmann, K.A.; Meyerowitz, E.M. The protein encoded by the Arabidopsis homeotic gene agamous resembles transcription factors. Nature 1990, 346, 35–39. [Google Scholar] [CrossRef] [PubMed]
- Ma, H. Molecular genetic analyses of microsporogenesis and microgametogenesis in flowering plants. Annu. Rev. Plant. Biol. 2005, 56, 393–434. [Google Scholar] [CrossRef]
- Chi, Y.; Huang, F.; Liu, H.; Yang, S.; Yu, D. An APETALA1-like gene of soybean regulates flowering time and specifies floral organs. J. Plant. Physiol. 2011, 168, 2251–2259. [Google Scholar] [CrossRef]
- Galimba, K.D.; Stilio, V.S.D. Sub-functionalization to ovule development following duplication of a floral organ identity gene. Dev. Biol. 2015, 405, 158–172. [Google Scholar] [CrossRef]
- Gao, Y.; Sun, J.; Sun, Z.; Xing, Y.; Zhang, Q.; Fang, K.; Cao, Q.; Qin, L. The MADS-box transcription factor CmAGL11 modulates somatic embryogenesis in Chinese chestnut (Castanea mollissima Blume). J. Integr. Agric. 2020, 19, 1033–1043. [Google Scholar] [CrossRef]
- Parenicova, L.; De Folter, S.; Kieffer, M.; Horner, D.S.; Favalli, C.; Busscher, J.; Cook, H.; Ingram, R.; Kater, M.M.; Davies, B. Molecular and phylogenetic analyses of the complete MADS-box transcription factor family in Arabidopsis: New Openings to the MADS World. Plant Cell 2003, 15, 1538–1551. [Google Scholar] [CrossRef] [PubMed]
- Arora, R.; Agarwal, P.; Ray, S.; Singh, A.K.; Singh, V.P.; Tyagi, A.K.; Kapoor, S. MADS-box gene family in rice: genome-wide identification, organization and expression profiling during reproductive development and stress. BMC Genom. 2007, 8, 242. [Google Scholar] [CrossRef]
- Xu, Z.; Zhang, Q.; Sun, L.; Du, D.; Cheng, T.; Pan, H.; Yang, W.; Wang, J. Genome-wide identification, characterisation and expression analysis of the MADS-box gene family in Prunus mume. Mol. Genet. Genom. 2014, 289, 903–920. [Google Scholar] [CrossRef] [PubMed]
- Henschel, K.; Kofuji, R.; Hasebe, M.; Saedler, H.; Munster, T.; Theissen, G. Two ancient classes of MIKC-type MADS-box genes are present in the Moss Physcomitrella Patens. Mol. Biol. Evol. 2002, 19, 801–814. [Google Scholar] [CrossRef] [PubMed]
- Díaz-Riquelme, J.; Lijavetzky, D.; Martinezzapater, J.M.; Carmona, M.J. Genome-wide analysis of MIKCC-Type MADS-box genes in grapevine. Plant Physiol. 2009, 149, 354–369. [Google Scholar] [CrossRef]
- Heijmans, K.; Morel, P.; Vandenbussche, M. MADS-box genes and floral development: the Dark Side. J. Exp. Bot. 2012, 63, 5397–5404. [Google Scholar] [CrossRef]
- Theisen, G.; Melzer, R.; Rumpler, F. MADS-domain transcription factors and the floral quartet model of flower development: linking plant development and evolution. Development 2016, 143, 3259–3271. [Google Scholar] [CrossRef]
- Martín-Pizarro, C.; Triviño, J.C.; Posé, D. Functional analysis of TM6 MADS-box gene in the octoploid strawberry by CRISPR/Cas9 directed mutagenesis. J. Exp. Bot. 2019, 70, 885–895. [Google Scholar] [CrossRef]
- Winter, K.; Becker, A.; Munster, T.; Kim, J.T.; Saedler, H.; Theissen, G. MADS-box genes reveal that gnetophytes are more closely related to conifers than to flowering plants. Proc. Natl. Acad. Sci. USA 1999, 96, 7342–7347. [Google Scholar] [CrossRef]
- Chen, X. A MicroRNA as a Translational Repressor of APETALA2 in Arabidopsis Flower Development. Science 2004, 303, 2022–2025. [Google Scholar] [CrossRef]
- Xu, Z.; Zhong, K. Plant Cell Differentiation and Organogenesis; Science Press: Biejing, China, 2015. [Google Scholar]
- Ehlers, K.; Bhide, A.S.; Tekleyohans, D.G.; Wittkop, B.; Snowdon, R.J.; Becker, A. The MADS-box genes ABS, SHP1, and SHP2 are essential for the coordination of cell divisions in ovule and seed coat development and for endosperm formation in Arabidopsis thaliana. PLoS ONE 2016, 11, e0165075. [Google Scholar] [CrossRef] [PubMed]
- Pan, I.L.; McQuinn, R.; Giovannoni, J.J.; Irish, V.F. Functional diversification of AGAMOUS lineage genes in regulating tomato flower and fruit development. J. Exp. Bot. 2010, 61, 1795–1806. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Zhang, L.; Xie, H.; Zhang, Y.; Oliveira, M.M.; Ma, R. Expression analysis and genetic mapping of three SEPALLATA-like genes from peach (Prunus persica (L.) Batsch). Tree Genet. Genomes 2008, 4, 693–703. [Google Scholar] [CrossRef]
- Melzer, R.; Theissen, G. Reconstitution of ‘floral quartets’ in vitro involving class B and class E floral homeotic proteins. Nucleic Acids Res. 2009, 37, 2723–2736. [Google Scholar] [CrossRef] [PubMed]
- Pan, Z.J.; Chen, Y.Y.; Du, J.S.; Chen, Y.Y.; Chung, M.C.; Tsai, W.C.; Wang, C.; Chen, H. Flower development of phalaenopsis orchid involves functionally divergent SEPALLATA-like genes. New Phytol. 2014, 202, 1024–1042. [Google Scholar] [CrossRef] [PubMed]
- Honma, T.; Goto, K. Complexes of MADS-box proteins are sufficient to convert leaves into floral organs. Nature 2001, 409, 525–529. [Google Scholar] [CrossRef]
- Yuan, Z.H. Chinese Fruit Tree Science and Practice-Pomegranate; Shaanxi Science and Technology Press: Xi’an, China, 2015; pp. 1–10. [Google Scholar]
- Holland, D.; Hatib, K.; Bar-Yaakov, I. Pomegranate: botany, horticulture, breeding. In Horticultural Review; John Wiley & Sons Inc.: Hoboken, NJ, USA, 2009; Volume 35, pp. 127–191. [Google Scholar]
- Huang, T.H.; Peng, G.; Kota, B.P.; Li, G.Q.; Yamahara, J.; Roufogalis, B.D.; Li, Y. Anti-diabetic action of Punica granatum flower extract: activation of PPAR-gamma and identification of an active component. Toxicol. Appl. Pharm. 2005, 207, 160–169. [Google Scholar] [CrossRef]
- Lansky, E.P.; Newman, R.A. Punica granatum (pomegranate) and its potential for prevention and treatment of inflammation and cancer. J. Ethnopharmacol. 2007, 109, 177–206. [Google Scholar] [CrossRef]
- Basu, A.; Penugonda, K. Pomegranate juice: A heart healthy fruit juice. Nutr. Rev. 2009, 67, 49–56. [Google Scholar] [CrossRef]
- Qin, G.; Xu, C.; Ming, R.; Tang, H.; Guyot, R.; Kramer, E.M.; Hu, Y.; Yi, X.; Qi, Y.; Xu, X. The pomegranate (Punica granatum L.) genome and the genomics of punicalagin biosynthesis. Plant J. 2017, 91, 1108–1128. [Google Scholar] [CrossRef]
- Yuan, Z.; Fang, Y.; Zhang, T.; Fei, Z.; Han, F.; Liu, C.; Liu, M.; Xiao, W.; Zhang, W.; Wu, S. The pomegranate (Punica granatum L.) genome provides insights into fruit quality and ovule developmental biology. Plant Biotechnol. J. 2018, 16, 1363–1374. [Google Scholar] [CrossRef] [PubMed]
- Luo, X.; Li, H.; Wu, Z.; Yao, W.; Zhao, P.; Cao, D.; Yu, H.; Li, K.; Poudel, K.L.; Zhao, D. The pomegranate (Punica granatum L.) draft genome dissects genetic divergence between soft- and hard-seeded cultivars. Plant Biotechnol. J. 2020, 18, 955–968. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.N.; Zhang, J.; Li, H.X.; Niu, J.; Xue, H.; Liu, B.B.; Wang, Q.; Luo, X.; Zhang, F.H.; Zhao, D.G.; et al. Transcriptomic analysis reveals candidate genes for female sterility in pomegranate flowers. Front Plant Sci. 2017, 8, 1430. [Google Scholar] [CrossRef] [PubMed]
- Ono, N.N.; Britton, M.T.; Fass, J.N.; Nicolet, C.M.; Lin, D.; Tian, L. Exploring the transcriptome landscape of pomegranate fruit peel for natural product biosynthetic gene and SSR marker discovery. J. Integr. Plant Biol. 2011, 53, 800–813. [Google Scholar] [CrossRef] [PubMed]
- Ophir, R.; Sherman, A.; Rubinstein, M.; Eshed, R.; Schwager, M.S.; Harel-Beja, R.; Bar-Ya’akov, I.; Holland, D. Single-nucleotide polymorphism markers from de-novo assembly of the pomegranate transcriptome reveal germplasm genetic diversity. PLoS ONE 2014, 9, e88998. [Google Scholar] [CrossRef]
- Marchler-Bauer, A.; Bo, Y.; Han, L.; He, J.; Lanczycki, C.J.; Lu, S.; Chitsaz, F.; Derbyshire, M.K.; Geer, R.C.; Gonzales, N.R.; et al. CDD/SPARCLE: functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. 2017, 45, D200–D203. [Google Scholar] [CrossRef]
- Jin, J.P.; Zhang, H.; Kong, L.; Gao, G.; Luo, J.C. PlantTFDB 3.0: a portal for the functional and evolutionary study of plant transcription factors. Nucleic Acids Res. 2014, 42, D1182-7. [Google Scholar] [CrossRef]
- Gao, H.H.; Zhang, Y.X.; Hu, S.W.; Guo, Y. Genome-wide survey and phylogenetic analysis of MADS-box gene family in Brassica napus. Chin. Bull. Botany 2017, 52, 699–712. [Google Scholar]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Chen, C.J.; Xia, R.; Chen, H.; He, Y.H. TBtools, a Toolkit for biologists integrating various HTS-data handling tools with a user-friendly interface. BioRxiv 2018. [Google Scholar] [CrossRef]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef]
- Nicolas, L.B.; Harold, P.; Páll, M.; Lior, P. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 2016, 34, 525–527. [Google Scholar]
- Leseberg, C.H.; Li, A.L.; Kang, H.; Duvall, M.; Mao, L. Genome-wide analysis of the MADS-box gene family in Populus trichocarpa. Gene 2006, 378, 84–94. [Google Scholar] [CrossRef] [PubMed]
- Hobo, T.; Asada, M.; Kowyama, Y.; Hattori, T. ACGT-containing abscisic acid response element (ABRE) and coupling element 3 (CE3) are functionally equivalent. Plant J. 1999, 19, 679–689. [Google Scholar] [CrossRef] [PubMed]
- Gubler, F.; Jacobsen, J.V. Gibberellin-responsive elements in the promoter of a barley high-pI alpha-amylase gene. Plant Cell 1992, 4, 1435–1441. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.R.; Hu, J.Y.; Ali, G.M. Reciprocal loss of CArG-boxes and auxin response elements drives expression divergence of MPF2-Like MADS-box genes controlling calyx inflation. PloS ONE 2012, 7, e42781. [Google Scholar] [CrossRef]
- Rouster, J.; Leah, R.; Mundy, J.; Cameron-Mills, V. Identification of a methyl jasmonate-responsive region in the promoter of a lipoxygenase 1 gene expressed in barley grain. Plant J. 1997, 11, 513–523. [Google Scholar] [CrossRef]
- Shah, J.; Klessig, D.F. Identification of a salicylic acid-responsive element in the promoter of the tobacco pathogenesis-related beta-1,3-glucanase gene, PR-2d. Plant J. 1996, 10, 1089–1101. [Google Scholar] [CrossRef]
- Yoshida, H.; Haze, K.; Yanagi, H.; Yura, T.; Mori, K. Identification of the cis-acting endoplasmic reticulum stress response element responsible for transcriptional induction of mammalian glucose-regulated proteins. Involvement of basic leucine zipper transcription factors. J. Biol. Chem. 1998, 273, 33741–33749. [Google Scholar] [CrossRef]
- Winter, K.; Weiser, C.; Kaufmann, K.; Bohne, A.; Kirchner, C.; Kanno, A.; Saedler, H.; Theissen, G. Evolution of class B floral homeotic proteins: Obligate heterodimerization originated from homodimerization. Mol. Biol. Evol. 2002, 19, 587–596. [Google Scholar] [CrossRef]
- Folter, S.D.; Shchennikova, A.V.; Franken, J.; Busscher, M.; Baskar, R.; Grossniklaus, U.; Angenent, G.C.; Immink, R.G.H. A Bsister MADS-box gene involved in ovule and seed development in petunia and Arabidopsis. Plant J. 2006, 47, 934–946. [Google Scholar] [CrossRef] [PubMed]
- Mondragonpalomino, M.; Theisen, G. Conserved differential expression of paralogous DEFICIENS- and GLOBOSA-like MADS-box genes in the flowers of Orchidaceae: refining the ‘orchid code’. Plant J. 2011, 66, 1008–1019. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Li, G.; Zhao, X.; Kang, Z.; Guo, A. Genome-Wide sequence identification of MADS-box transcription factor gene family from Fragaria vesca and cloning of FaMADS1 gene from F. ananassa Fruit. Plant Phys. J. 2014, 50, 1630–1638. [Google Scholar]
- Wells, C.E.; Vendramin, E.; Tarodo, S.J.; Verde, I.; Bielenberg, D.G. A genome-wide analysis of MADS-box genes in peach [Prunus persica (L.) Batsch]. BMC Plant Biol. 2015, 15, 41. [Google Scholar] [CrossRef]
- Shu, Y.; Yu, D.; Wang, D.; Guo, D.; Guo, C. Genome-wide survey and expression analysis of the MADS-box gene family in soybean. Mol. Biol. Rep. 2013, 40, 3901–3911. [Google Scholar] [CrossRef]
- Tian, Y.; Dong, Q.; Ji, Z.; Chi, F.; Zhou, Z. Genome-wide identification and analysis of the MADS-box gene family in apple. Gene 2014, 555, 277–290. [Google Scholar] [CrossRef]
- Bai, G.; Yang, D.; Cao, P.; Yao, H.; Zhang, Y.; Chen, X.; Xiao, B.; Li, F.; Wang, Z.; Yang, J. Genome-wide identification, gene structure and expression analysis of the MADS-Box gene family indicate their function in the development of tobacco (Nicotiana tabacum L.). Int. J. Mol. Sci. 2019, 20, 5043. [Google Scholar] [CrossRef]
- Schilling, S.; Kennedy, A.; Pan, S.; Jermiin, L.S.; Melzer, R. Genome-wide analysis of MIKC-type MADS-box genes in wheat: pervasive duplications, functional conservation and putative neofunctionalization. New Phytol. 2020, 225, 511–529. [Google Scholar] [CrossRef]
- Xu, Y.; Ding, Z.; Yao, Y.; Gong, R.; Wen, Y. Karyotype analysis of four pomegranate cultivars. Nonwood For. Res. 2008, 26, 47–52. [Google Scholar]
- De Bodt, S.; Raes, J.; De Peer, Y.V.; Theisen, G. And then there were many: MADS goes genomic. Trends Plant Sci. 2003, 8, 475–483. [Google Scholar] [CrossRef]
- Tang, W.; Tu, Y.; Cheng, X.; Zhang, L.; Meng, H.; Zhao, X.; Zhang, W.; He, B. Genome-wide identification and expression profile of the MADS-box gene family in Erigeron breviscapus. PLoS ONE 2019, 14, e0226599. [Google Scholar] [CrossRef]
- Riechmann, J.L.; Heard, J.E.; Martin, G.M.; Reuber, L.; Jiang, C.; Keddie, J.; Adam, L.; Pineda, O.; Ratcliffe, O.; Samaha, R. Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science 2000, 290, 2105–2110. [Google Scholar] [CrossRef]
- Zhang, Y.; Tang, D.; Lin, X.; Ding, M.; Tong, Z. Genome-wide identification of MADS-box family genes in moso bamboo (Phyllostachys edulis) and a functional analysis of PeMADS5 in flowering. BMC Plant Biol. 2018, 18, 176. [Google Scholar] [CrossRef]
- Cui, M.; Wang, C.; Wu, W.; Wang, J.; Tang, W.; Zhang, W.; Zhu, X.; Jia, H.; Shen, W.; Fang, J. Expression characteristic analysis of MADS-box family members of grape transcription factors in berry development and ripening process. J. Fruit Sci. 2017, 34, 1497–1508. [Google Scholar]
- Hundertmark, M.; Hincha, D.K. LEA (Late Embryogenesis Abundant) proteins and their encoding genes in Arabidopsis thaliana. BMC Genom. 2008, 9, 118. [Google Scholar] [CrossRef] [PubMed]
- Li, P.C.; Yu, S.W.; Shen, J.; Li, Q.Q.; Li, D.P.; Li, D.Q.; Zheng, C.C.; Shu, H.R. The transcriptional response of apple alcohol acyltransferase (MdAAT2) to salicylic acid and ethylene is mediated through two apple MYB TFs in transgenic tobacco. Plant Mol. Biol. 2014, 85, 627–638. [Google Scholar] [CrossRef] [PubMed]
- Kaufmann, K.; Melzer, R.; Theissen, G. MIKC-type MADS-domain proteins: Structural modularity, protein interactions and network evolution in land plants. Gene 2005, 347, 183–198. [Google Scholar] [CrossRef] [PubMed]
- Ma, H.; Depamphilis, C.W. The ABCs of floral evolution. Cell 2000, 101, 5–8. [Google Scholar] [CrossRef]
- Yoo, S.K.; Wu, X.; Lee, J.S.; Ji, H.A. AGAMOUS-LIKE 6 is a floral promoter that negatively regulates the FLC/MAF clade genes and positively regulates FT in Arabidopsis. Plant J. 2011, 65, 62–76. [Google Scholar] [CrossRef]
- Ferrandiz, C.; Liljegren, S.J.; Yanofsky, M.F. Negative Regulation of the SHATTERPROOF Genes by FRUITFULL during Arabidopsis fruit development. Science 2000, 289, 436–438. [Google Scholar] [CrossRef]
- Pinyopich, A.; Ditta, G.S.; Savidge, B.; Liljegren, S.J.; Baumann, E.; Wisman, E.; Yanofsky, M.F. Assessing the redundancy of MADS-box genes during carpel and ovule development. Nature 2003, 424, 85–88. [Google Scholar] [CrossRef] [PubMed]
- Seymour, G.B.; Ryder, C.D.; Cevik, V.; Hammond, J.P.; Popovich, A.; King, G.J.W.; Vrebalov, J.; Giovannoni, J.J.; Manning, K. A SEPALLATA gene is involved in the development and ripening of strawberry (Fragaria x ananassa Duch.) fruit, a non-climacteric tissue. J. Ex. Bot. 2011, 62, 1179–1188. [Google Scholar] [CrossRef] [PubMed]
- Theißen, G. Development of floral organ identity: Stories from the MADS house. Curr. Opin. Plant Biol. 2001, 4, 75–85. [Google Scholar] [CrossRef]

| Gene Name | Gene ID | Location | CDS (bp) | Exon No. | Strand | Amino Acid Residues | MW(Da) | PI | Subfamily |
|---|---|---|---|---|---|---|---|---|---|
| PgMADS01 | Pg000170.1 | scaffold1:4730801:4734403 | 720 | 7 | − | 239 | 27591.1 | 9.76 | A(AP1-FUL) |
| PgMADS02 | Pg001270.1 | scaffold10:781136:787463 | 576 | 5 | − | 191 | 22251.1 | 8.76 | SOC1 |
| PgMADS03 | Pg002695.1 | scaffold11:837228:841996 | 846 | 7 | − | 281 | 31958.1 | 8.19 | E(SEP) |
| PgMADS04 | Pg002696.1 | scaffold11:827349:833260 | 600 | 7 | − | 199 | 22764.8 | 9.58 | FLC |
| PgMADS05 | Pg002697.1 | scaffold11:821950:824737 | 603 | 6 | − | 200 | 22793.1 | 10.17 | FLC |
| PgMADS06 | Pg003192.1 | scaffold111:252873:258483 | 522 | 6 | − | 173 | 20047.1 | 9.59 | C/D(AG) |
| PgMADS07 | Pg003228.1 | scaffold111:423542:426294 | 645 | 7 | + | 214 | 24079.5 | 8.95 | AGL12 |
| PgMADS08 | Pg003757.1 | scaffold119:139771:142418 | 639 | 7 | − | 212 | 24900.2 | 9.28 | B(AP3-PI) |
| PgMADS09 | Pg005184.1 | scaffold13:3171597:3176055 | 741 | 8 | − | 246 | 28261 | 8.58 | E(SEP) |
| PgMADS10 | Pg005185.1 | scaffold13:3153310:3158145 | 807 | 8 | − | 268 | 30475.7 | 8.8 | A(AP1-FUL) |
| PgMADS11 | Pg006006.1 | scaffold136:390163:393471 | 906 | 9 | − | 301 | 34973.4 | 9.9 | BS |
| PgMADS12 | Pg008155.1 | scaffold16:570614:573248 | 885 | 7 | − | 294 | 33598.8 | 6.39 | AGL15 |
| PgMADS13 | Pg008465.1 | scaffold16:2554235:2556059 | 642 | 7 | + | 213 | 23721.7 | 8.48 | SVP |
| PgMADS14 | Pg008776.1 | scaffold17:2387895:2392008 | 687 | 8 | − | 228 | 25779.9 | 5.52 | SVP |
| PgMADS15 | Pg009904.1 | scaffold184:267398:273943 | 765 | 5 | − | 254 | 28775.2 | 8.88 | C/D(AG) |
| PgMADS16 | Pg014226.1 | scaffold24:1271362:1277266 | 504 | 5 | + | 167 | 19349.0 | 5.97 | MIKC* |
| PgMADS17 | Pg014283.1 | scaffold24:2253081:2257895 | 687 | 8 | + | 228 | 25738.3 | 6.53 | SVP |
| PgMADS18 | Pg014451.1 | scaffold25:814656:818749 | 717 | 8 | − | 238 | 27124.6 | 9.52 | AGL6 |
| PgMADS19 | Pg014630.1 | scaffold25:796521:803849 | 654 | 7 | + | 217 | 24695 | 7.93 | SOC1 |
| PgMADS20 | Pg016044.1 | scaffold3:3612017:3628588 | 612 | 7 | − | 203 | 23056.1 | 4.99 | FLC |
| PgMADS21 | Pg016673.1 | scaffold3:3637896:3641838 | 768 | 8 | − | 255 | 29008.8 | 8.85 | E(SEP) |
| PgMADS22 | Pg017747.1 | scaffold33:1901905:1904548 | 759 | 8 | + | 252 | 28454 | 7.62 | AGL15 |
| PgMADS23 | Pg018533.1 | scaffold37:1603815:1615806 | 837 | 7 | − | 278 | 31322 | 6.89 | FLC |
| PgMADS24 | Pg020204.1 | scaffold40:966778:969371 | 699 | 7 | + | 232 | 26761 | 4.9 | B(AP3-PI) |
| PgMADS25 | Pg020205.1 | scaffold40:971279:973422 | 708 | 7 | + | 235 | 27089.6 | 6.31 | B(AP3-PI) |
| PgMADS26 | Pg022433.1 | scaffold5:1599117:1601542 | 675 | 7 | − | 224 | 25754 | 9.07 | B(AP3-PI) |
| PgMADS27 | Pg023007.1 | scaffold50:465601:468168 | 750 | 7 | + | 249 | 28634.4 | 8.57 | E(SEP) |
| PgMADS28 | Pg023008.1 | scaffold50:480602:487666 | 699 | 7 | + | 232 | 26732.3 | 9.43 | A(AP1-FUL) |
| PgMADS29 | Pg024984.1 | scaffold60:1113804:1120065 | 753 | 8 | − | 250 | 28461.2 | 9.5 | E(SEP) |
| PgMADS30 | Pg026995.1 | scaffold7:90410:99792 | 1974 | 11 | + | 657 | 74414.5 | 9.56 | C/D(AG) |
| PgMADS31 | Pg027190.1 | scaffold71:566889:578438 | 654 | 7 | + | 217 | 24935.5 | 9.15 | SOC1 |
| PgMADS32 | Pg027284.1 | scaffold72:255144:257303 | 528 | 6 | − | 175 | 20534.3 | 10.89 | TM8 |
| PgMADS33 | Pg028066.1 | scaffold79:321847:329529 | 672 | 8 | − | 223 | 25020.5 | 10.39 | SVP |
| PgMADS34 | Pg028180.1 | scaffold8:2827697:2833438 | 663 | 7 | − | 220 | 25172.5 | 10.04 | SOC1 |
| PgMADS35 | Pg029246.1 | scaffold86:465224:468460 | 762 | 7 | − | 253 | 28671.7 | 8.89 | AGL17 |
| PgMADS36 | Pg029338.1 | scaffold86:426938:437880 | 1443 | 13 | − | 480 | 55016.3 | 6.61 | AGL17 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, Y.; Zhao, H.; Wang, Y.; Zhang, X.; Zhao, X.; Yuan, Z. Genome-Wide Identification and Expression Analysis of MIKC-Type MADS-Box Gene Family in Punica granatum L. Agronomy 2020, 10, 1197. https://doi.org/10.3390/agronomy10081197
Zhao Y, Zhao H, Wang Y, Zhang X, Zhao X, Yuan Z. Genome-Wide Identification and Expression Analysis of MIKC-Type MADS-Box Gene Family in Punica granatum L. Agronomy. 2020; 10(8):1197. https://doi.org/10.3390/agronomy10081197
Chicago/Turabian StyleZhao, Yujie, Honglian Zhao, Yuying Wang, Xinhui Zhang, Xueqing Zhao, and Zhaohe Yuan. 2020. "Genome-Wide Identification and Expression Analysis of MIKC-Type MADS-Box Gene Family in Punica granatum L." Agronomy 10, no. 8: 1197. https://doi.org/10.3390/agronomy10081197
APA StyleZhao, Y., Zhao, H., Wang, Y., Zhang, X., Zhao, X., & Yuan, Z. (2020). Genome-Wide Identification and Expression Analysis of MIKC-Type MADS-Box Gene Family in Punica granatum L. Agronomy, 10(8), 1197. https://doi.org/10.3390/agronomy10081197






