Molecularly Imprinted Polymer-Based Electrochemical BioSensors for Haemophilus influenzae Rapid Detection
Abstract
1. Introduction
2. Materials and Methods
2.1. H. influenzae Preparation and Fixation Procedure
2.2. The Number of Imprinting H. influenzae on Electrode
2.3. Polymer Synthesis for H. influenzae Detection
2.4. Screen-Printed Electrode Preparation (SPE)
3. Results
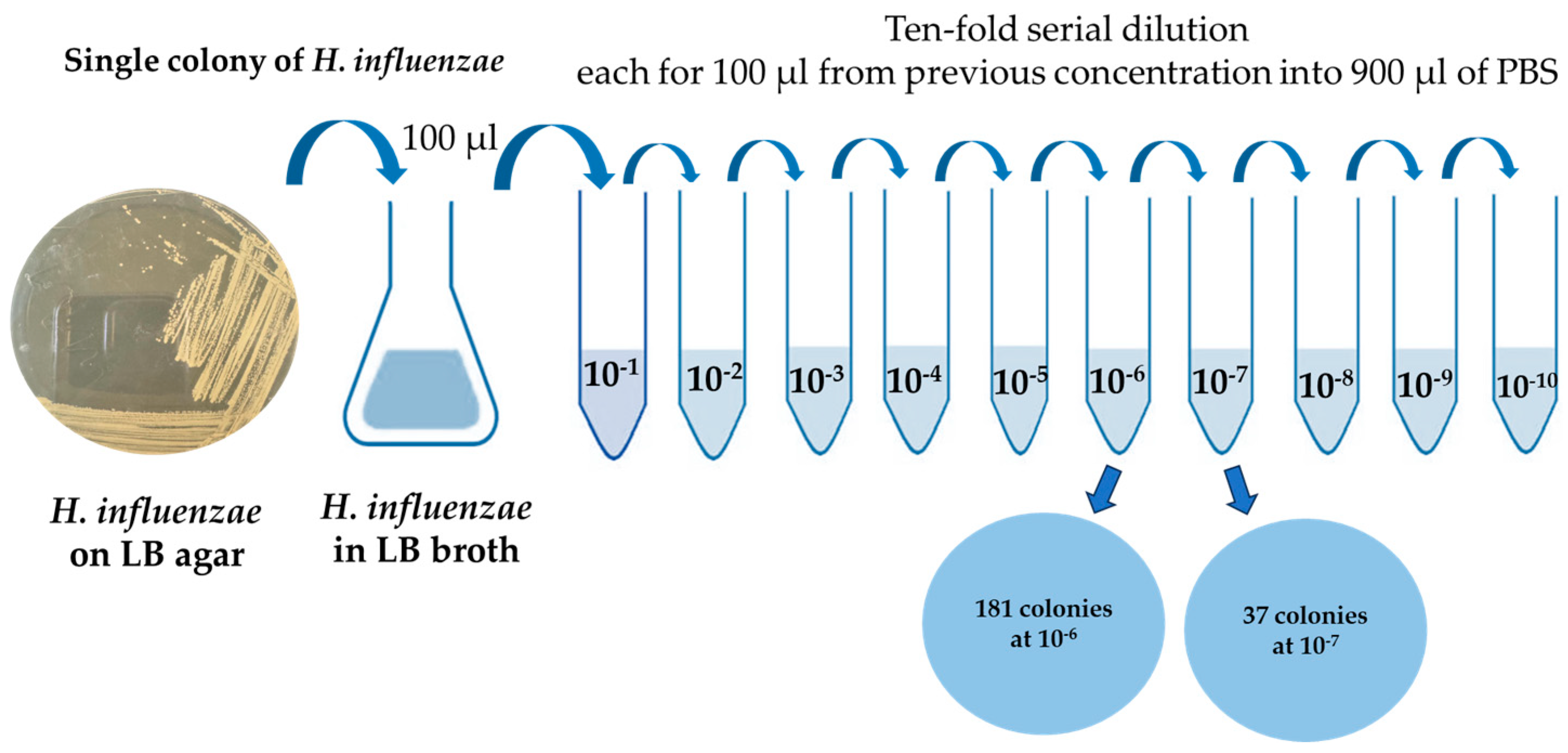
3.1. Initial H. influenzae Stock Concentration
3.2. Morphological Characterization of H. influenzae by SEM

3.3. Surface Characterization of Electrodes

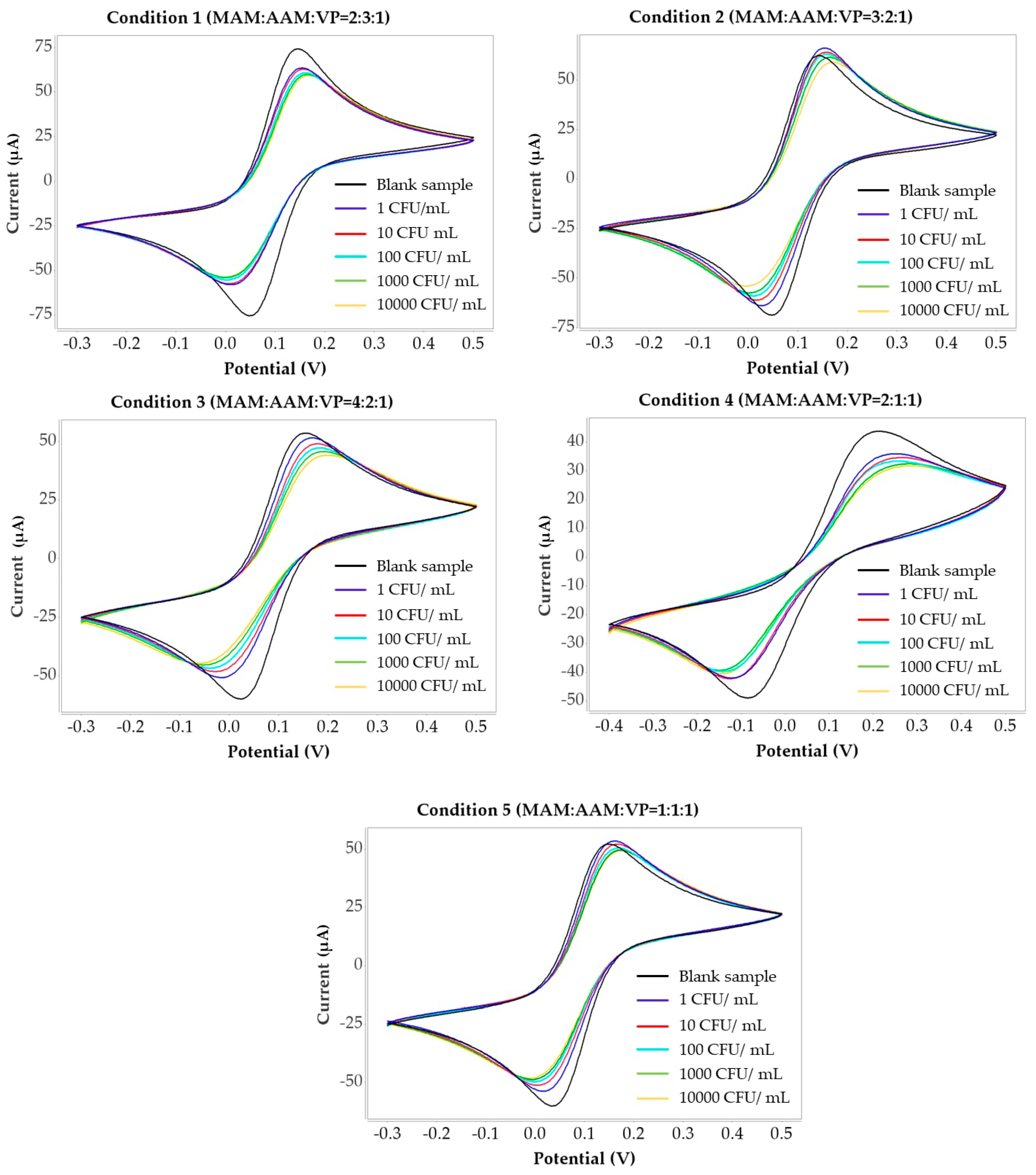
3.4. CV Characterization of H. influenzae Interaction
3.5. Repeatability Evaluation
3.6. Limit of Detection (LOD)
3.7. Comparison with Reported Method
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wen, S.; Feng, D.; Chen, D.; Yang, L.; Xu, Z. Molecular epidemiology and evolution of Haemophilus influenzae. Infect. Genet. Evol. 2020, 80, 104205. [Google Scholar] [CrossRef]
- Kalangi, H.; Ajit, A.A.; Camins, B.; Yancovitz, S.R. Haemophilus influenzae serotype a meningitis in an elderly patient: A case report and literature review. IDCases 2024, 38, e02094. [Google Scholar] [CrossRef]
- Patel, P.; Fogwe, D.; Patel, T.; Patil, S.; Bran-Acevedo, A.; Oba, Y.; Roland, W. A case of acute non-typeable Hemophilus influenza infective endocarditis in a patient with hypocomplementemia. IDCases 2023, 32, e01756. [Google Scholar] [CrossRef]
- Mangtani, P.; Mulholland, K.; Madhi, S.A.; Edmond, K.; O’Loughlin, R.; Hajjeh, R. Haemophilus influenzae type b disease in HIV-infected children: A review of the disease epidemiology and effectiveness of Hib conjugate vaccines. Vaccine 2010, 28, 1677–1683. [Google Scholar] [CrossRef] [PubMed]
- Abdeldaim, G.M.K.; Strålin, K.; Kirsebom, L.A.; Olcén, P.; Blomberg, J.; Herrmann, B. Detection of Haemophilus influenzae in respiratory secretions from pneumonia patients by quantitative real-time polymerase chain reaction. Diagn. Microbiol. Infect. Dis. 2009, 64, 366–373. [Google Scholar] [CrossRef] [PubMed]
- Templier, V.; Livache, T.; Boisset, S.; Maurin, M.; Slimani, S.; Mathey, R.; Roupioz, Y. Biochips for Direct Detection and Identification of Bacteria in Blood Culture-Like Conditions. Sci. Rep. 2017, 7, 9457. [Google Scholar] [CrossRef]
- Tien, N.; You, B.-J.; Lin, H.-J.; Chang, C.-Y.; Chou, C.-Y.; Lin, H.-S.; Chang, C.-T.; Wang, C.C.N.; Chen, H.-C. Repeated centrifuging and washing concentrates bacterial samples in peritoneal dialysis for optimal culture: An original article. BMC Microbiol. 2020, 20, 365. [Google Scholar] [CrossRef]
- Akbari Nakhjavani, S.; Mirzajani, H.; Carrara, S.; Onbaşlı, M.C. Advances in biosensor technologies for infectious diseases detection. TrAC Trends Anal. Chem. 2024, 180, 117979. [Google Scholar] [CrossRef]
- Castillo-Henríquez, L.A.-O.; Brenes-Acuña, M.; Castro-Rojas, A.; Cordero-Salmerón, R.; Lopretti-Correa, M.; Vega-Baudrit, J.A.-O. Biosensors for the Detection of Bacterial and Viral Clinical Pathogens. Sensors 2020, 20, 6926. [Google Scholar] [CrossRef]
- Gaba, S.; Chauhan, N.; Chandra, R.; Jain, U. Future advances of artificial biosensor technology in biomedical applications. Talanta Open 2024, 9, 100301. [Google Scholar] [CrossRef]
- Erdem, A.; Senturk, H.; Karakus, M. Molecularly imprinted polymer-based sensors: Design and advances in the analysis of DNA and protein. Talanta Open 2025, 12, 100507. [Google Scholar] [CrossRef]
- Li, Y.; Luo, L.; Kong, Y.; Li, Y.; Wang, Q.; Wang, M.; Li, Y.; Davenport, A.; Li, B. Recent advances in molecularly imprinted polymer-based electrochemical sensors. Biosens. Bioelectron. 2024, 249, 116018. [Google Scholar] [CrossRef]
- Saylan, Y.; Kılıç, S.; Denizli, A. Biosensing Applications of Molecularly Imprinted-Polymer-Based Nanomaterials. Processes 2024, 12, 177. [Google Scholar] [CrossRef]
- Arreguin-Campos, R.; Jiménez-Monroy, K.L.; Diliën, H.A.-O.; Cleij, T.A.-O.; van Grinsven, B.A.-O.; Eersels, K.A.-O. Imprinted Polymers as Synthetic Receptors in Sensors for Food Safety. Biosensors 2021, 11, 46. [Google Scholar] [CrossRef]
- Asif, I.M.; Di Giulio, T.; Gagliani, F.; Malitesta, C.; Mazzotta, E. Advances in the Direct Nanoscale Integration of Molecularly Imprinted Polymers (MIPs) with Transducers for the Development of High-Performance Nanosensors. Biosensors 2025, 15, 509. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Dong, T.; Wei, X.; Yang, Z.; Matos Pires, N.M.; Ren, J.; Jiang, Z. Electrochemical methods for detection of biomarkers of Chronic Obstructive Pulmonary Disease in serum and saliva. Biosens. Bioelectron. 2019, 142, 111453. [Google Scholar] [CrossRef] [PubMed]
- Ogulewe, F.E.; Oladipo, A.A.; Gazi, M. Molecularly imprinted polymers and metal-organic framework-based nanomaterial sensors for food and beverage analysis and safety—A review. Talanta Open 2025, 11, 100448. [Google Scholar] [CrossRef]
- Parisi, O.I.; Francomano, F.; Dattilo, M.; Patitucci, F.; Prete, S.; Amone, F.; Puoci, F. The Evolution of Molecular Recognition: From Antibodies to Molecularly Imprinted Polymers (MIPs) as Artificial Counterpart. J. Funct. Biomater. 2022, 13, 12. [Google Scholar] [CrossRef]
- Yang, Y.; He, X.; Xu, S.; Wang, D.; Liu, Z.; Xu, Z. Post-imprinting modification of molecularly imprinted polymer for proteins detection: A review. Int. J. Biol. Macromol. 2023, 253, 127104. [Google Scholar] [CrossRef]
- Agar, M.; Laabei, M.; Leese, H.S.; Estrela, P. Multi-Template Molecularly Imprinted Polymeric Electrochemical Biosensors. Chemosensors 2025, 13, 11. [Google Scholar] [CrossRef]
- Martín-Esteban, A. Molecularly-imprinted polymers as a versatile, highly selective tool in sample preparation. TrAC Trends Anal. Chem. 2013, 45, 169–181. [Google Scholar] [CrossRef]
- Karrat, A.; Amine, A. Innovative approaches to suppress non-specific adsorption in molecularly imprinted polymers for sensing applications. Biosens. Bioelectron. 2024, 250, 116053. [Google Scholar] [CrossRef]
- Pintavirooj, C.; Vongmanee, N.; Sukjee, W.; Sangma, C.; Visitsattapongse, S. Biosensors for Klebsiella pneumoniae with Molecularly Imprinted Polymer (MIP) Technique. Sensors 2022, 22, 4638. [Google Scholar] [CrossRef] [PubMed]
- Nampeng, J.; Vongmanee, N.; Pintavirooj, C.; Chiu, W.-T.; Visitsattapongse, S. Electrochemical Biosensors by Means of Molecularly Imprinted Polymers (MIPs) Cortisol Recognition. Polymers 2025, 17, 545. [Google Scholar] [CrossRef]
- Brazys, E.; Ratautaite, V.; Mohsenzadeh, E.; Boguzaite, R.; Ramanaviciute, A.; Ramanavicius, A. Formation of molecularly imprinted polymers: Strategies applied for the removal of protein template (review). Adv. Colloid Interface Sci. 2025, 337, 103386. [Google Scholar] [CrossRef] [PubMed]
- Abdella, A.A.; Ulber, R. Dummy-template molecularly imprinted polymers as an approach for improved efficiency for analytical applications: A mini review. Microchem. J. 2025, 209, 112768. [Google Scholar] [CrossRef]
- Vivaldi, F.; Torrini, F.; Spiaggia, F.; Di Francesco, F.; Minunni, M. Exploring affinity receptors in bioanalysis: From natural binders to biomimetics. Sens. Actuators Rep. 2025, 10, 100359. [Google Scholar] [CrossRef]
- Ma, X.; Allahou, L.W.; Yang, R.; Ma, Y.; Dimoula, M.; Chau, D.Y.S.; Williams, G.R.; Knowles, J.C.; Poma, A. Antiviral molecularly imprinted polymers: Engineered precision for multifunctional therapeutic strategies. Mater. Sci. Eng. R Rep. 2026, 167, 101099. [Google Scholar] [CrossRef]
- Guo, X.; Bai, X.; Zheng, Z.; Qiu, Z.; Qiao, X.; Qiao, Y. Optimization of Culture Conditions for Bacteriocin Production by Pediococcus Acidilactici CCFM18 and Characterization of Its Biological Properties. Fermentation 2025, 11, 470. [Google Scholar] [CrossRef]
- Nikara, S.; Ahmadi, E.; Nia, A.A. Effects of different preparation techniques on the microstructural features of biological materials for scanning electron microscopy. J. Agric. Food Res. 2020, 2, 100036. [Google Scholar] [CrossRef]
- Yadav, A.K.; Verma, D.; Dalal, N.; Kumar, A.; Solanki, P.R. Molecularly imprinted polymer-based nanodiagnostics for clinically pertinent bacteria and virus detection for future pandemics. Biosens. Bioelectron. X 2022, 12, 100257. [Google Scholar] [CrossRef]
- Vongmanee, N.; Nampeng, J.; Pintavirooj, C.; Visitsattapongse, S. Biosensor Based on Electrochemical Analysis for Staphylococcus aureus Detection with Molecular Imprinted Polymer Technique. Polymers 2025, 17, 2826. [Google Scholar] [CrossRef] [PubMed]
- Rabchinskii, M.K.; Shiyanova, K.A.; Brzhezinskaya, M.; Gudkov, M.V.; Saveliev, S.D.; Stolyarova, D.Y.; Torkunov, M.K.; Chumakov, R.G.; Vdovichenko, A.Y.; Cherviakova, P.D.; et al. Chemistry of Reduced Graphene Oxide: Implications for the Electrophysical Properties of Segregated Graphene–Polymer Composites. Nanomaterials 2024, 14, 1664. [Google Scholar] [CrossRef]
- Elfadil, D.; Lamaoui, A.; Della Pelle, F.; Amine, A.; Compagnone, D. Molecularly Imprinted Polymers Combined with Electrochemical Sensors for Food Contaminants Analysis. Molecules 2021, 26, 4607. [Google Scholar] [CrossRef]
- Resende, S.; Fernandes, J.; Sousa, P.C.; Calaza, C.; Frasco, M.F.; Freitas, P.P.; Goreti, F.; Sales, M. Fabrication and sensing properties of a molecularly imprinted polymer on a photonic PDMS substrate for the optical detection of C-reactive protein. Chem. Eng. J. 2024, 485, 149924. [Google Scholar] [CrossRef]
- Wu, T.; Yagati, A.K.; Min, J. Electrochemical Detection of Different Foodborne Bacteria for Point-of-Care Applications. Biosensors 2023, 13, 641. [Google Scholar] [CrossRef] [PubMed]
- Oğuzhan Kaya, H.; Tekintaş, Y.; Kurul, F.; Cetin, A.E.; Nur Topkaya, S. Targeted microorganism detection with molecularly imprinted polymer biosensors. J. Electroanal. Chem. 2024, 971, 118575. [Google Scholar] [CrossRef]
- Liu, R.; Cai, T.; Huang, Z.; Zhu, Q.; Wang, X. Novel ultrasensitive impedimetric biosensor for rapid detection of Pseudomonas aeruginosa via recombinant lectin-functionalized nanoporous gold biointerface. Biosens. Bioelectron. 2025, 288, 117788. [Google Scholar] [CrossRef]
- Qi, W.; Wang, S.; Wang, L.; Xi, X.; Wu, S.; Li, Y.; Liao, M.; Lin, J. A portable viable Salmonella detection device based on microfluidic chip and recombinase aided amplification. Chin. Chem. Lett. 2023, 34, 107360. [Google Scholar] [CrossRef]
- Qazi, R.A.; Aman, N.; Ullah, N.; Jamila, N.; Bibi, N. Recent advancement for enhanced e. Coli detection in electrochemical biosensors. Microchem. J. 2024, 196, 109673. [Google Scholar] [CrossRef]
- Bayliss, N.; Schmidt, B.V.K.J. Hydrophilic polymers: Current trends and visions for the future. Prog. Polym. Sci. 2023, 147, 101753. [Google Scholar] [CrossRef]
- Alizadeh, N.; Georgas, A.; Argirusis, C.; Sourkouni, G.; Argirusis, N. Polymer Coatings for Electrochemical Biosensors. Coatings 2026, 16, 164. [Google Scholar] [CrossRef]




| Condition | Ratio (n:n) | MAM (mg) | AAM (mg) | VP (µL) |
|---|---|---|---|---|
| 1 | 2:3:1 | 17.0 | 21.8 | 10.7 |
| 2 | 3:2:1 | 25.5 | 14.2 | 10.7 |
| 3 | 4:2:1 | 34.0 | 14.2 | 10.7 |
| 4 | 2:1:1 | 17.0 | 7.1 | 10.7 |
| 5 | 1:1:1 | 8.5 | 7.1 | 10.7 |
| Repetition 1 | Current (µA) | ∆I (µA) | Relative Current Changing (%) |
| Blank | 50.47 | ||
| 1 CFU/mL | 44.35 | 11.09 | 20.00 |
| 10 CFU/mL | 39.49 | 15.95 | 28.77 |
| 100 CFU/mL | 33.24 | 22.20 | 40.04 |
| 1000 CFU/mL | 27.87 | 27.57 | 49.73 |
| 10,000 CFU/mL | 22.23 | 33.21 | 59.91 |
| Repetition 2 | Current (µA) | ∆I (µA) | Relative Current Changing (%) |
| Blank | 48.56 | ||
| 1 CFU/mL | 42.45 | 12.99 | 23.43 |
| 10 CFU/mL | 36.98 | 18.46 | 33.30 |
| 100 CFU/mL | 34.12 | 21.32 | 38.45 |
| 1000 CFU/mL | 27.56 | 27.88 | 50.29 |
| 10,000 CFU/mL | 22.78 | 32.66 | 58.91 |
| Repetition 3 | Current (µA) | ∆I (µA) | Relative Current Changing (%) |
| Blank | 49.22 | ||
| 1 CFU/mL | 42.37 | 13.07 | 23.57 |
| 10 CFU/mL | 36.52 | 18.92 | 34.13 |
| 100 CFU/mL | 34.65 | 20.79 | 37.50 |
| 1000 CFU/mL | 25.43 | 30.01 | 54.13 |
| 10,000 CFU/mL | 21.39 | 34.05 | 61.42 |
| Concentration | Average (µA) | SD | % RSD |
|---|---|---|---|
| 1 CFU/mL | 22.34 | 2.02 | 9.05 |
| 10 CFU/mL | 32.06 | 2.88 | 8.99 |
| 100 CFU/mL | 38.67 | 1.28 | 3.32 |
| 1000 CFU/mL | 51.38 | 2.40 | 4.66 |
| 10,000 CFU/mL | 60.08 | 1.26 | 2.10 |
| Sensor Platform | Target | Recognition Element | Electrode | Linear Range | LOD | Reference |
|---|---|---|---|---|---|---|
| This work | H. influenzae | MIP | Gold SPE | 1–104 CFU/mL | 1.03 CFU/mL | This study |
| Electrochemical aptasensor | Salmonella spp. | DNA aptamer | Carbon SPE | 10–104 CFU/mL | 50 CFU/mL | [36] |
| MIP electrochemical sensor | E. coli | MIP | Glassy carbon | 10–105 CFU/mL | 9.40 CFU/mL | [37] |
| Electrochemical immunosensor | P. aeruginosa | Antibody | Au electrode | 10–106 CFU/mL | 8 CFU/mL | [38] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Vongmanee, N.; Nampeng, J.; Pintavirooj, C.; Visitsattapongse, S. Molecularly Imprinted Polymer-Based Electrochemical BioSensors for Haemophilus influenzae Rapid Detection. Polymers 2026, 18, 726. https://doi.org/10.3390/polym18060726
Vongmanee N, Nampeng J, Pintavirooj C, Visitsattapongse S. Molecularly Imprinted Polymer-Based Electrochemical BioSensors for Haemophilus influenzae Rapid Detection. Polymers. 2026; 18(6):726. https://doi.org/10.3390/polym18060726
Chicago/Turabian StyleVongmanee, Naphatsawan, Jindapa Nampeng, Chuchart Pintavirooj, and Sarinporn Visitsattapongse. 2026. "Molecularly Imprinted Polymer-Based Electrochemical BioSensors for Haemophilus influenzae Rapid Detection" Polymers 18, no. 6: 726. https://doi.org/10.3390/polym18060726
APA StyleVongmanee, N., Nampeng, J., Pintavirooj, C., & Visitsattapongse, S. (2026). Molecularly Imprinted Polymer-Based Electrochemical BioSensors for Haemophilus influenzae Rapid Detection. Polymers, 18(6), 726. https://doi.org/10.3390/polym18060726








