Crude Enzyme Concentrate of Filamentous Fungus Hydrolyzed Chitosan to Obtain Oligomers of Different Sizes
Abstract
1. Introduction
2. Materials and Methods
2.1. Chitosan
2.2. Commercial Enzyme
2.3. Microorganism
2.4. Identification of the Microorganism
2.5. Crude Enzyme Concentrate (CEC)
2.6. Protein Content
2.7. Endoglucanase Activity Assay (CMCase Activity)
2.8. Identification of Proteins in Crude Enzyme Concentrate
2.8.1. Protein Precipitation
2.8.2. Protein Digestion
2.8.3. Identification by LC-MS/MS
2.8.4. Data Processing and Search Parameters
2.8.5. Identification of Microorganisms Based on Peptides
2.9. Enzymatic Hydrolysis of Chitosan
2.9.1. Crude Enzyme Concentrate
2.9.2. Commercial Enzyme
2.10. Characterization of Chitosan and Oligomers
2.10.1. Molecular Mass Determination
2.10.2. FTIR Deacetylation Degree
3. Results and Discussion
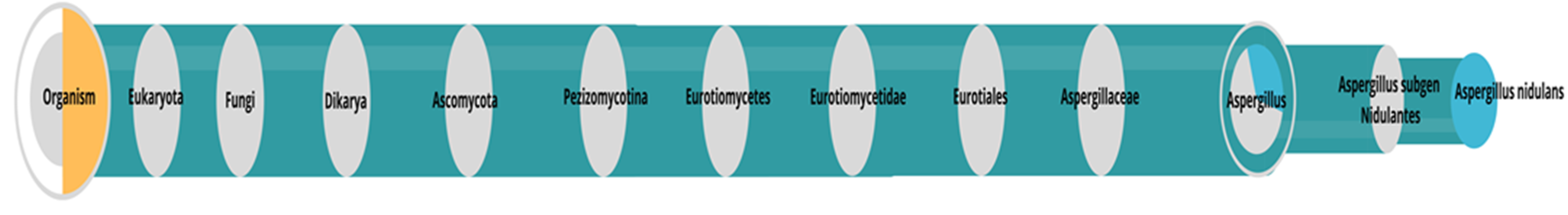
3.1. Identification of the Microorganism
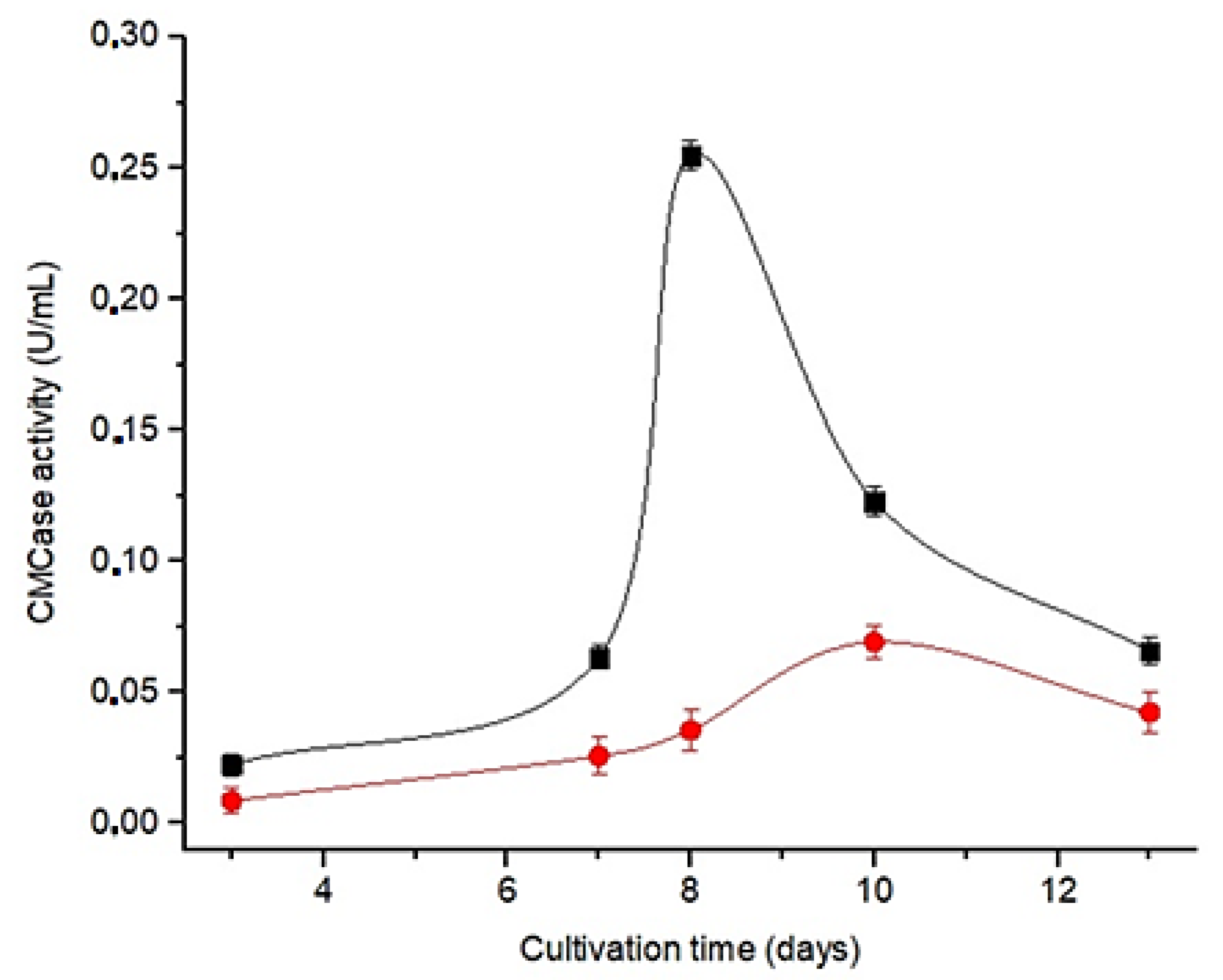
3.2. Selection of the Culture Medium
3.3. Crude Enzyme Concentrate (CEC)
3.4. Identification of Proteins in Crude Enzyme Concentrate
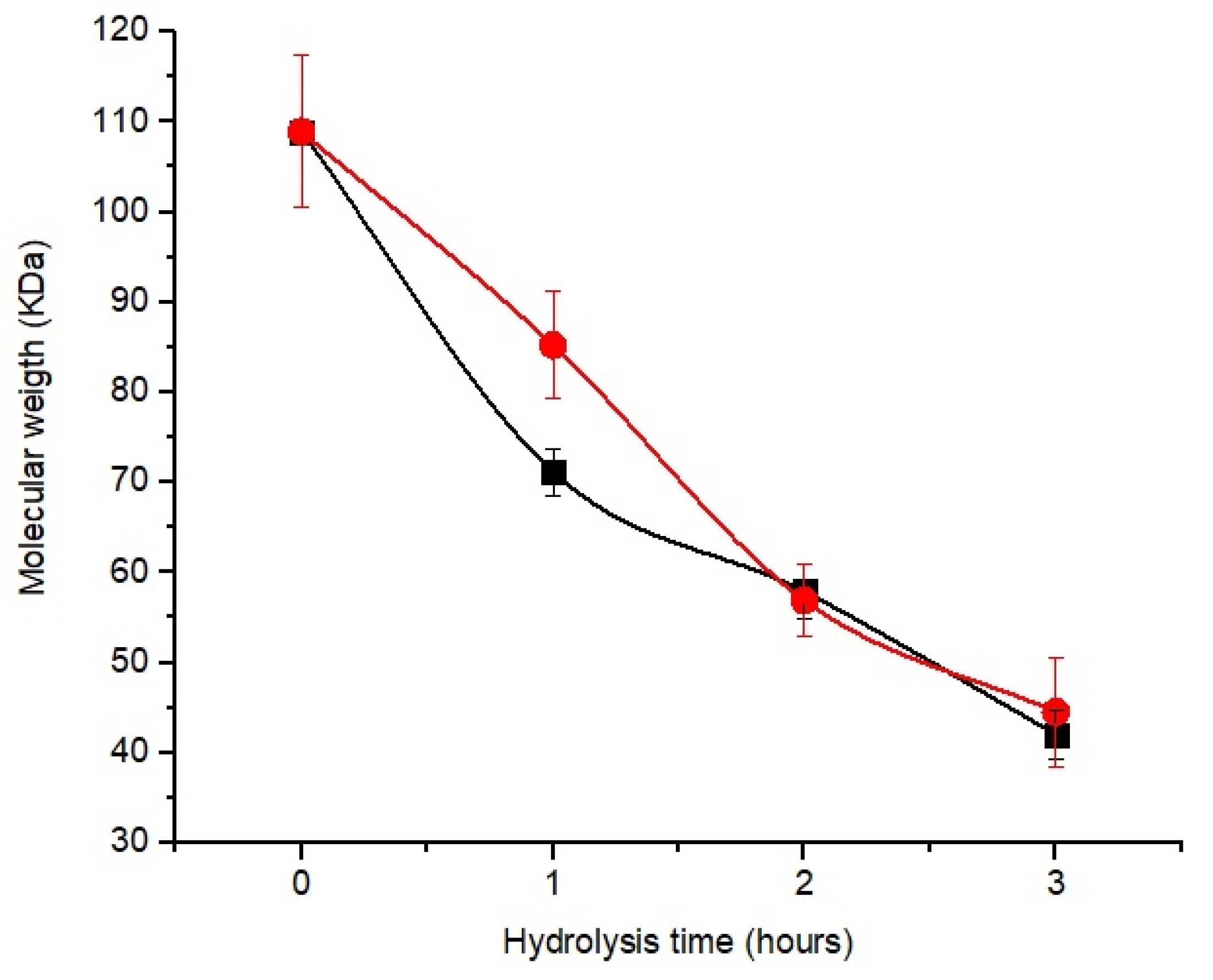
3.5. Chitosan Hydrolysis
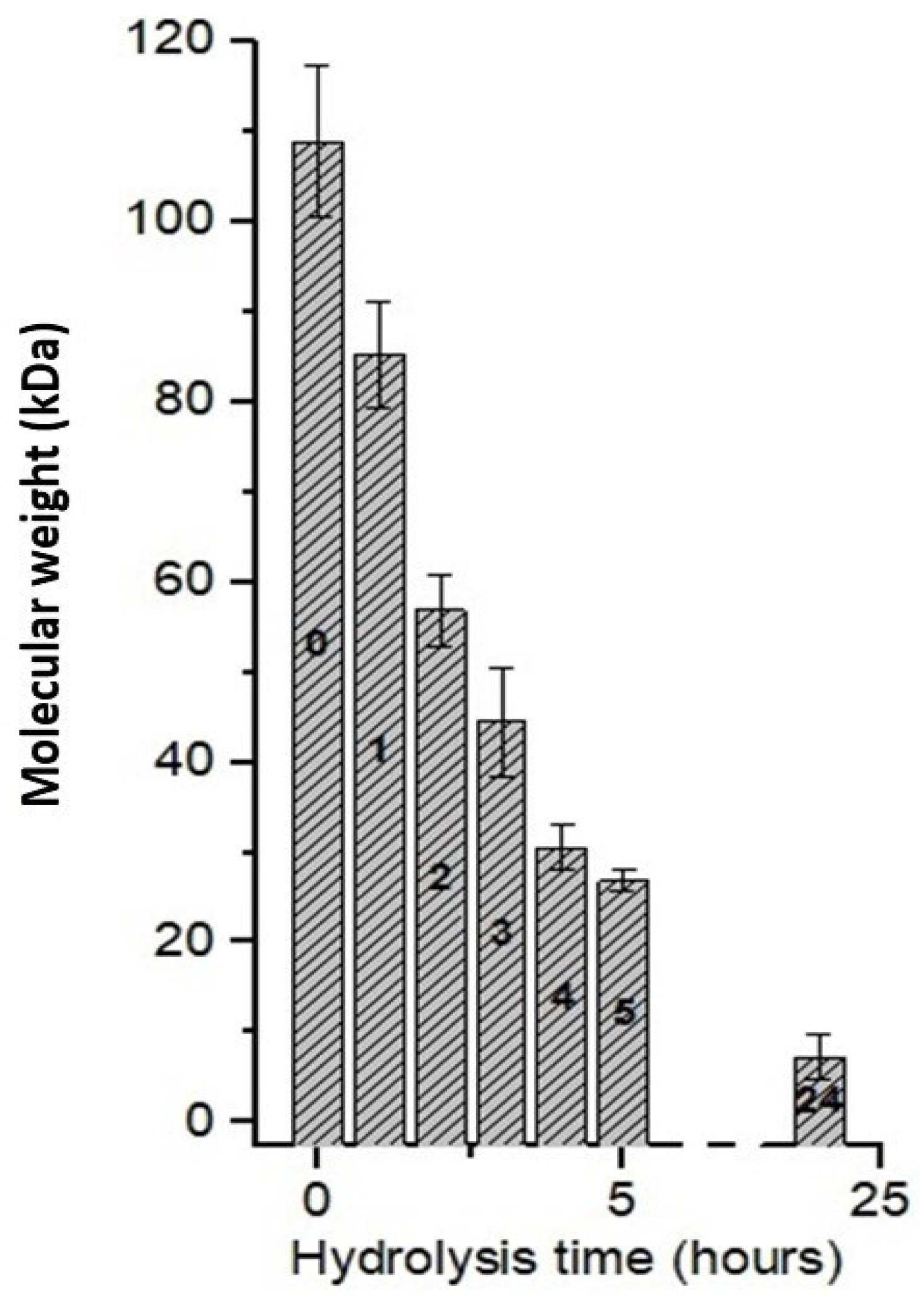
3.6. Effect of Hydrolysis Time Using CEC
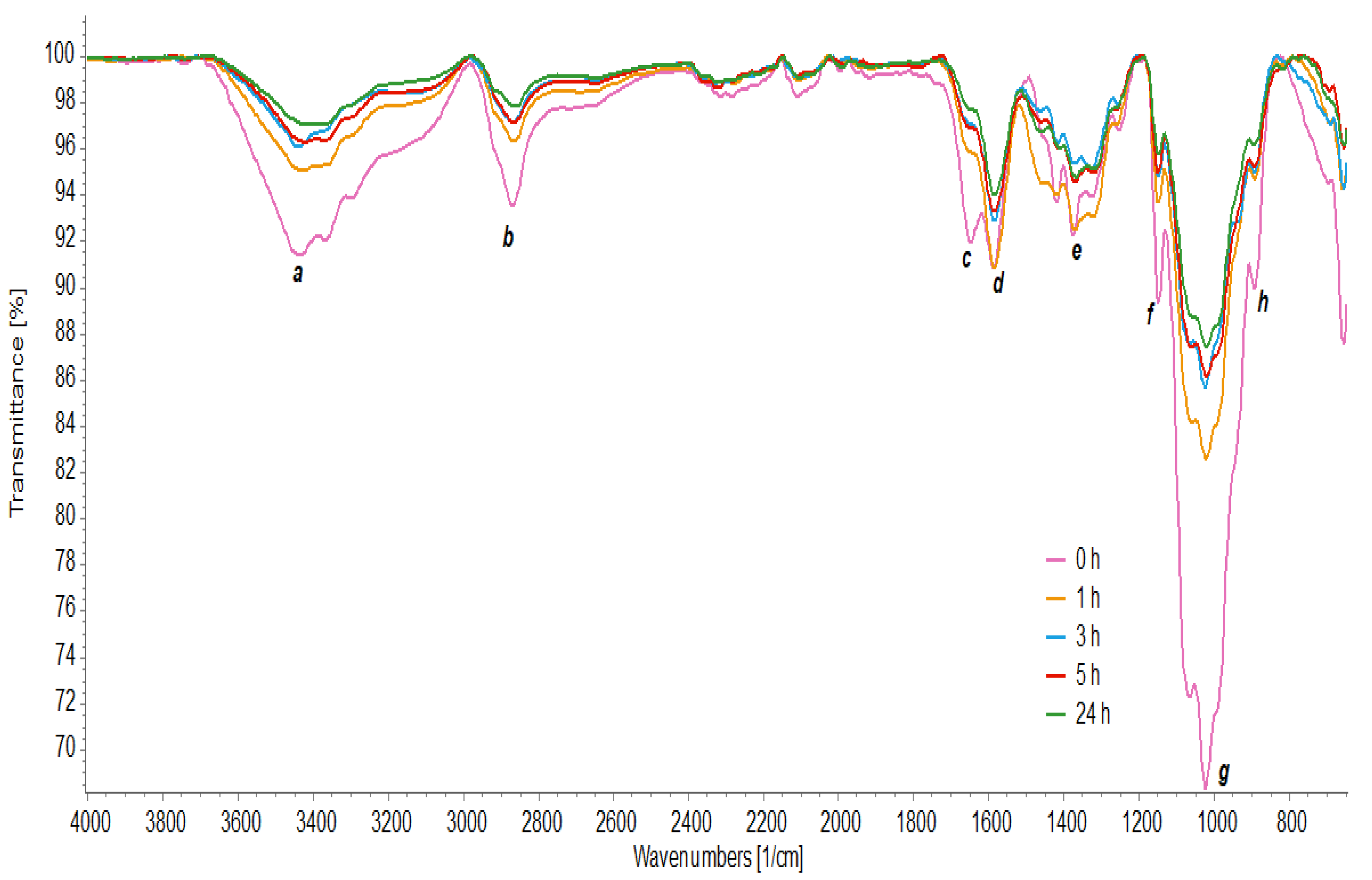
3.7. FTIR Analysis
3.8. Deacetylation Degree of Chitosan (DG)
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hamed, I.; Özogul, F.; Regenstein, J.M. Industrial applications of crustacean by-products (chitin, chitosan, and chitooligosaccharides): A review. Trends Food Sci. Technol. 2016, 48, 40–50. [Google Scholar] [CrossRef]
- Naveed, M.; Phil, L.; Sohail, M.; Hasnat, M.; Baig, M.M.F.A.; Ihsan, A.U.; Shumzaid, M.; Kakar, M.U.; Khan, T.M.; Akabar, M.D.; et al. Chitosan oligosaccharide (COS): An overview. Int. J. Biol. Macromol. 2019, 129, 827–843. [Google Scholar] [CrossRef] [PubMed]
- Santoso, J.; Adiputra, K.C.; Soerdirga, L.C.; Tarman, K. Effect of acetic acid hydrolysis on the characteristics of water soluble chitosan. IOP Conf. Ser. Earth Environ. Sci. 2020, 414, 012021. [Google Scholar] [CrossRef]
- Águila-Almanza, E.; Salgado-Delgado, R.; Vargas-Galarza, Z.; García-Hernández, E.; Hernández-Cocoletzi, H. Enzymatic depolimerization of chitosan for the preparation of functional membranes. J. Chem. 2019, 2019, 5416297. [Google Scholar] [CrossRef]
- Laokuldilok, T.; Potivas, T.; Kanha, N.; Surawang, S.; Seesuriyachan, P.; Wangtueai, S.; Phimolsiripol, Y.; Regenstein, J.M. Physicochemical, antioxidant, and antimicrobial properties of chitooligosaccharides produced using three different enzyme treatments. Food Biosci. 2017, 18, 28–33. [Google Scholar] [CrossRef]
- Yin, N.; Du, R.; Zhao, F.; Han, Y.; Zhou, Z. Characterization of antibacterial bacterial cellulose composite membranes modified with chitosan or chitooligosaccharide. Carbohydr. Polym. 2020, 229, 4. [Google Scholar] [CrossRef]
- Xu, W.; Mohan, A.; Pitts, N.L.; Udenigwe, C.; Mason, B. Bile acid-binding capacity of lobster shell-derived chitin, chitosan and chitooligosaccharides. Food Biosci. 2020, 33, 100476. [Google Scholar] [CrossRef]
- Chang, S.-H.; Wu, C.-H.; Tsai, G.-J. Effects of chitosan molecular weight on its antioxidant and antimutagenic properties. Carbohydr. Polym. 2018, 18, 1026–1032. [Google Scholar] [CrossRef]
- Chang, Y.-W.; Zeng, X.Y.; Sung, W.-C. Effect of chitooligosaccharide and different low molecular weight chitosans on the formation of acrylamide and 5-hydroxymethylfurfural and Maillard reaction products in glucose/fructose-asparagine model systems. LWT 2020, 119, 108879. [Google Scholar] [CrossRef]
- Qin, C.; Zhou, B.; Zeng, L.; Zhang, Z.; Liu, Y.; Du, Y.; Xiao, L. The physicochemical properties and antitumor activity of cellulase-treated chitosan. Food Chem. 2004, 84, 107–115. [Google Scholar] [CrossRef]
- Xing, R.; Liu, Y.; Li, K.; Yu, H.; Liu, S.; Yang, Y.; Chen, X.; Li, P. Monomer composition of chitooligosaccharides obtained by different degradation methods and their effects on immunomodulatory activities. Carbohydr. Polym. 2017, 157, 1288–1297. [Google Scholar] [CrossRef]
- Kim, S.-K.; Rajapakse, N. Enzymatic production and biological activities of chitosan oligosaccharides (COS): A review. Carbohydr. Polym. 2005, 62, 357–368. [Google Scholar] [CrossRef]
- Sánchez, Á.; Mengíbar, M.; Rivera-Rodríguez, G.; Moerchbacher, B.; Acosta, N.; Heras, A. The effect of preparation processes on the physicochemical characteristics and antibacterial activity of chitooligosaccharides. Carbohydr. Polym. 2017, 157, 251–257. [Google Scholar] [CrossRef]
- Li, J.; Du, Y.; Yang, J.; Feng, T.; Li, A.; Chen, P. Preparation and characterisation of low molecular weight chitosan and chito-oligomers by a commercial enzyme. Polym. Degrad. Stab. 2005, 87, 441–448. [Google Scholar] [CrossRef]
- Aranaz, I.; Alcántara, A.R.; Civera, M.C.; Arias, C.; Elorza, B.; Caballero, A.H.; Acosta, N. Chitosan: An overview of its properties and applications. Polymers 2021, 13, 3256. [Google Scholar] [CrossRef]
- Li, M.; Han, J.; Xue, Y.; Dai, Y.; Liu, J.; Gan, L.; Xie, R.; Long, M. Hydrogen peroxide pretreatment efficiently assisting enzymatic hydrolysis of chitosan at high concentration for chitooligosaccharides. Polym. Degrad. Stab. 2019, 164, 177–186. [Google Scholar] [CrossRef]
- Pan, A.-D.; Zeng, H.-Y.; Foua, G.B.; Alain, C.; Li, Y.-Q. Enzymolysis of chitosan by papain and its kinetics. Carbohydr. Polym. 2016, 135, 199–206. [Google Scholar] [CrossRef]
- Roncal, T.; Oviedo, A.; Armentia, I.L.; Fernández, L.; Villarán, M.C. High yield production of monomer-free chitosan oligosaccharides by pepsin catalyzed hydrolysis of a high deacetylation degree chitosan. Carbohydr. Res. 2007, 342, 2750–2756. [Google Scholar] [CrossRef] [PubMed]
- Santos-Moriano, P.; Kidibule, P.E.; Alleyne, E.; Ballesteros, A.O.; Heras, A.; Fernandez-Lobato, M.; Plou, F.J. Efficient conversion of chitosan into chitooligosaccharides by a chitosanolytic activity from Bacillus thuringiensis. Process Biochem. 2018, 73, 102–108. [Google Scholar] [CrossRef]
- Wang, H.; Cui, H.; Wang, X.; Lin, C.; Xia, S.; Hayat, K.; Hussain, S.; Tahir, M.U.; Zhang, X. Metal complexed-enzymatic hydrolyzed chitosan improves moisture retention of fiber papers by migrating immobilized water to bound state. Carbohydr. Polym. 2020, 235, 115967. [Google Scholar] [CrossRef]
- Sirohi, R.; Singh, A.; Tarafdar, A.; Shahi, N.C.; Verma, A.K.; Kushwaha, A. Cellulase production from pre-treated pea hulls using Trichoderma reesei under submerged fermentation. Waste Biomass Valoriz. 2019, 10, 2651–2659. [Google Scholar] [CrossRef]
- Mrudula, S.; Murugammal, R. Production of cellulase by Aspergillus Niger under submerged and solid state fermentation using coir waste as a substrate. Braz. J. Microbiol. 2011, 42, 1119–1127. [Google Scholar] [CrossRef]
- Ferreira, N.R.; Sarquis, M.I.M.; Gobira, R.M.; Souza, M.G.; Santos, A.S. The multivariate statistical selection of fungal strains isolated from Neoteredo reynei, with the high hydrolytic potential to deconstruct celulose. Food Res. Int. 2019, 122, 402–410. [Google Scholar] [CrossRef]
- Chan, L.G.; Cohen, J.L.; Bell, J.M. Conversion of agricultural streams and food-processing by-products to value-added compounds using filamentous fungi. Annu. Rev. Food Sci. Technol. 2018, 9, 503–523. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.C.; Eisner, J.D.; Kattar, M.M.; Rassoulian-Barrett, S.L.; Lafe, K.; Bui, U.; Limaye, A.P.; Cookson, B.T. Polymorphic internal transcribed spacer region 1 DNA sequences identify medically important yeasts. J. Clin. Microbiol. 2001, 39, 4042–4051. [Google Scholar] [CrossRef]
- Peláez, R.D.R.; Wischral, D.; Cunha, J.R.B.; Mendes, T.D.; Pacheco, T.F.; Siqueira, F.G.d.; Almeida, J.R.M.d. Production of Enzymatic Extract with High Cellulolytic and Oxidative Activities by Co-Culture of Trichoderma reesei and Panus lecomtei. Fermentation 2022, 8, 522. [Google Scholar] [CrossRef]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing theprincipleof protein-dyebinding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Miller, G.L. Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal. Chem. 1959, 31, 426–428. [Google Scholar] [CrossRef]
- Wessel, D.; Flugge, U.I. A method for the quantitative recovery of protein in dilute solution in the presence of detergents and lipids. Anal. Biochem. 1984, 138, 141–143. [Google Scholar] [CrossRef]
- Chapman, J.R. (Ed.) Mass Spectrometry of Proteins and Peptides; Humana Press: Totowa, NJ, USA, 2000. [Google Scholar]
- Garcia, L.G.; Guedes, G.M.; Silva, M.L.; Castelo-Branco, D.S.; Sidrim, J.J.; Cordeiro, R.; Rocha, M.F.; Vieira, R.S.; Brilhante, R.S. Effect of the molecular weight of chitosan on its antifungal activity against Candida spp. in planktonic cells and biofilm. Carbohydr. Polym. 2018, 195, 662–669. [Google Scholar] [CrossRef]
- Barragán, C.A.R.; Fornué, E.D.; Ortega, J.A.A. Determinación del grado de desacetilación de quitosana mediante titulación potenciométrica, FTIR y Raman. J. CIM 2016, 4, 769–777. [Google Scholar]
- Santos, L.N.; Santos, A.S.; Dantas, K.d.G.F.; Ferreira, N.R. Adsorption of Cu (II) Ions Present in the Distilled Beverage (Sugar Cane Spirit) Using Chitosan Derived from the Shrimp Shell. Polymers 2022, 14, 573. [Google Scholar] [CrossRef]
- Brugnerotto, J.; Lizardi, J.; Goycoolea, F.M.; Argüelles-Monal, W.; Desbrières, J.; Rinaudo, M. An infrared investigation in relation with chitin and chitosan characterization. Polymer 2001, 42, 3569–3580. [Google Scholar] [CrossRef]
- Xie, Y.; Wei, Y.; Hu, J.G. Depolymerization of Chitosan with a Crude Cellulase Preparation from Aspergillus niger. Appl. Biochem. Biotechnol. 2010, 160, 1074–1083. [Google Scholar] [CrossRef]
- Kittur, F.S.; Kumar, A.B.V.; Varadaraj, M.C.; Tharanathan, R.N. Chitooligosaccharides-preparation with the aid of pectinase isozyme from Aspergillus niger and their antibacterial activity. Carbohydr. Res. 2005, 340, 1239–1245. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Gay, L.M.; Tuveng, T.R.; Agger, J.W.; Westereng, B.; Mathiesen, G.; Horn, S.J.; Vaaje-Kolstad, G.; van Aalten, D.M.F.; Eijsink, V.G.H. Structure and function of a broad-specificity chitin deacetylase from Aspergillus nidulans FGSC A4. Sci. Rep. 2017, 7, 1746. [Google Scholar] [CrossRef]
- Cheng, C.-Y.; Chang, C.-H.; Wu, Y.-J.; Li, Y.-K. Exploration of glycosyl hydrolase family 75, a chitosanase from Aspergillus fumigatus. J. Biol. Chem. 2006, 28, 3137–3144. [Google Scholar] [CrossRef] [PubMed]
- Poshina, D.N.; Raik, S.V.; Poshin, A.N.; Skorik, Y.A. Accessibility of chitin and chitosan in enzymatic hydrolysis: A review. Polym. Degrad. Stab. 2018, 156, 269–278. [Google Scholar] [CrossRef]
- Su, P.C.; Hsueh, W.C.; Chang, W.S.; Chen, P.T. Enhancement of chitosanase secretion by Bacillus subtilis for production of chitosan oligosaccharides. J. Taiwan Inst. Chem. Eng. 2017, 79, 49–54. [Google Scholar] [CrossRef]
- Thadathil, N.; Velappan, S.P. Recent developments in chitosanase research and its biotechnological applications: A review. Food Chem. 2014, 150, 392–399. [Google Scholar] [CrossRef]
- Johnsen, M.G.; Hansen, O.C.; Stougaard, P. Isolation, characterization and heterologous expression of a novel chitosanase from Janthinobacterium sp. strain 4239. Microb. Cell Factories 2010, 9, 1–9. [Google Scholar] [CrossRef] [PubMed]
- da Silva, L.C.; Honorato, T.L.; Franco, T.T.; Rodrigues, S. Optimization of Chitosanase Production by Trichoderma koningii sp. Under Solid-State Fermentation. Food Bioprocess Technol. 2012, 5, 1564–1572. [Google Scholar] [CrossRef]
- Xia, W.; Yu, X. Purification and characterization of two types of chitosanase from Aspergillus sp. CJ22-326. Food Res. Int. 2005, 38, 315–322. [Google Scholar] [CrossRef]
- Embaby, A.M.; Melika, R.R.; Hussein, A.; El-Kamel, A.H.; Marey, H.S. Biosynthesis of chitosan-Oligosaccharides (COS) by nonaflatoxigenic Aspergillus sp. strain EGY1 DSM 101520: A robust biotechnological approach. Process Biochem. 2018, 64, 16–30. [Google Scholar] [CrossRef]
- Qu, T.; Zhang, C.; Qin, Z.; Fan, L.; Jiang, L.; Zhao, L. A novel GH family 20 β-N-acetylhexosaminidase with both chitosanase and chitinase activity from Aspergillus oryzae. Front. Mol. Biosci. 2021, 8, 684086. [Google Scholar] [CrossRef]
- Kidibule, P.E.; Santos-Moriano, P.; Plou, F.J.; Fernández-Lobato, M. Endo-chitinase Chit33 specificity on different chitinolytic materials allows the production of unexplored chitooligosaccharides with antioxidant activity. Biotechnol. Rep. 2020, 27, e00500. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Matsuo, I.; Ajisaka, K.; Nakajima, H.; Kitamoto, K. Cloning and characterization of the nagA gene that encodes beta-n-acetylglucosaminidase from Aspergillus nidulans and its expression in Aspergillus oryzae. Biosci. Biotechnol. Biochem. 2002, 66, 2168–2175. [Google Scholar] [CrossRef]
- Patil, N.S.; Jadhav, J.P. Significance of Penicillium ochrochloron chitinase as a biocontrol agent against pest Helicoverpa armigera. Chemosphere 2015, 128, 231–235. [Google Scholar] [CrossRef] [PubMed]
- Han, P.; Yang, C.; Liang, X.; Li, L. Identification and characterization of a novel chitinase with antifungal activity from ‘Baozhu’ pear (Pyrus ussuriensis Maxim.). Food Chem. 2016, 196, 808–814. [Google Scholar] [CrossRef]
- Le, B.; Yang, S.H. Microbial chitinases: Properties, current state and biotechnological applications. World J. Microbiol. Biotechnol. 2019, 35, 144. [Google Scholar] [CrossRef]
- Purushotham, P.; Podile, A.R. Synthesis of Long-Chain Chitooligosaccharides by a Hypertransglycosylating Processive Endochitinase of Serratia proteamaculans 568. J. Bacteriol. 2016, 194, 4260–4271. [Google Scholar] [CrossRef]
- Al Abboud, M.A.; Al-Rajhi, A.M.H.; Shater, A.-R.M.; Alawlaqi, M.M.; Mashraqi, A.; Selim, S.; Al Jaouni, S.K.; Ghany, T.M.A. Halostability and Thermostability of Chitinase Produced by Fungi Isolated from Salt Marsh Soil in Subtropical Region of Saudi Arabia. BioResources 2022, 17, 4763–4780. [Google Scholar] [CrossRef]
- Abdel-Moneim, A.M.E.; El-Saadony, M.T.; Shehata, A.M.; Saad, A.M.; Aldhumri, S.A.; Ouda, S.M.; Mesalam, N.M. Antioxidant and antimicrobial activities of Spirulina platensis extracts and biogenic selenium nanoparticles against selected pathogenic bacteria and fungi. Saudi J. Biol. Sci. 2022, 29, 1197–1209. [Google Scholar] [CrossRef] [PubMed]
- Saykhedkar, S.; Ray, A.; Ayoubi-Canaan, P.; Hartson, S.D.; Prade, R.; Mort, A.J. A time course analysis of the extracellular proteome of Aspergillus nidulans growing on sorghum stover. Biotechnol. Biofuels 2012, 5, 52. [Google Scholar] [CrossRef] [PubMed]
- Tegl, G.; Öhlknecht, C.; Vielnascher, R.; Rollett, A.; Hofinger-Horvath, A.; Kosma, P.; Guebitz, G.M. Cellobiohydrolases produce different oligosaccharides from chitosan. Biomacromolecules 2016, 17, 2284–2292. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.-X.; Qian, L.; Zhou, Y. Stimulation effect of chitosan on the immunity of radiotherapy patients suffered from lung câncer. Int. J. Biol. Macromol. 2015, 72, 195–198. [Google Scholar] [CrossRef]
- Squillaci, G.; Esposito, A.; La Cara, F.; Morana, A. Free and immobilized thermophilic alpha-L-arabinofuranosidase for arabinose production. Arch. Ind. Biotechnol. 2017, 1, 27–34. [Google Scholar]
- Bibi, F.; Irshad, M.; Anwar, Z.; Bhatti, K.H.; Raza, A. Improved catalytic functionalities of purified pristine and chitosan-immobilized polygalacturonase, and pectin lyase. Chem. Eng. Res. Des. 2019, 128, 146–154. [Google Scholar] [CrossRef]
- Kittur, F.S.; Kumar, A.B.V.; Tharanat Purushotham and Podile, R.N. Low molecular weight chitosans/preparation by depolymerization with Aspergillus niger pectinase, and characterization. Carbohydr. Res. 2003, 33, 1283–1290. [Google Scholar] [CrossRef]
- Tishchenko, G.; Šimůnek, J.; Brus, J.; Netopilík, M.; Pekárek, M.; Walterová, Z.; Koppová, I.; Lenfeld, J. Low-molecular-weight chitosans: Preparation and characterization. Carbohydr. Polym. 2011, 86, 1077–1081. [Google Scholar] [CrossRef]
- Lin, S.B.; Lin, Y.C.; Chen, H.H. Low molecular weight chitosan prepared with the aid of cellulase, lysozyme and chitinase: Characterisation and antibacterial activity. Food Chem. 2009, 116, 47–53. [Google Scholar] [CrossRef]
- Beer, B.; Bartolome, M.J.; Berndorfer, L.; Bochmann, G.; Guebitz, G.M.; Nyanhongo, G.S. Controlled enzymatic hydrolysis and synthesis of lignin cross-linked chitosan functional hydrogels. Int. J. Biol. Macromol. 2020, 161, 1440–1446. [Google Scholar] [CrossRef] [PubMed]
- Luo, W.; Han, Z.; Zeng, X.; Yu, S.; Kennedy, J.F. Study on the degradation of chitosan by pulsed electric fields treatment. Innov. Food Sci. Emerg. Technol. 2010, 11, 587–591. [Google Scholar] [CrossRef]
| Volume (mL) | Total Activity (U/mL) | Total Protein (mg/mL) | Specific Activity (U/mg) | |
|---|---|---|---|---|
| Crude enzyme | 500.0 | 0.22 ± 0.031 | 0.0165 ± 0.001 | 13.3 ± 0.97 |
| Crude enzyme concentrate | 25.0 | 2.23 ± 0.19 | 0.14 ± 0.013 | 15.9 ± 1.38 |
| Data on Proteins Identified by LC-MS/MS against the Aspergillus sp. Database | Data on Proteins Identified by LC-MS/MS against the Aspergillus nidulans Database | ||||||
|---|---|---|---|---|---|---|---|
| Access Number—Proteins Identified against the Aspergillus sp. Database | Peptide | Mass | Ppm | m/z | Accession Number | Coverage (%) | Catalytic Class (E.C) |
| A1DMA5—Probable 1,4-beta-D-glucan cellobiohydrolase A OS = Neosartorya fischeri | VIANSVSNVADVSGNSISSDFC(+57.02)TAQK | 26.692.603 | 16.7 | 8.907.712 | Q5B2Q4—Probable 1,4-beta-D-glucan cellobiohydrolase A; flags: Precursor [Aspergillus nidulans FGSC A4] | 99% | Glycosidase, Hydrolase (3.2.1.91) |
| Q8NK02—1 4-beta-D-glucan cellobiohydrolase B OS = Emericella nidulans | YGTGYC(+57.02)DSQC(+57.02)PR LNFVTQSQQK | 14.625.605 11.916.248 | 0.9 −1.9 | 7.322.845 5.968.185 | Q8NK02—1 4-beta-D-glucan cellobiohydrolase B OS = Emericella nidulans | 100% | Glycosidase, Hydrolase (3.2.1.91) |
| Q5AUX2—Alpha-L-arabinofuranosidase axhA-2 OS = Emericella nidulans | ANSGATWTDDISHGDLVR | 19.138.867 | 2.8 | 6.389.697 | Q5AUX2—Alpha-L-arabinofuranosidase axhA-2 OS = Emericella nidulans | 100% | Glicosidase, Hidrolase (3.2.1.55) |
| Q5BA61 Pectin lyase B OS = Emericella nidulans | SLVGEGSSGVIK | 11.316.135 | 1.5 | 5.668.149 | Q5BA61 Pectin lyase B OS = Emericella nidulans | 100% | Liase (4.2.2.10) |
| E9QRF2 Endochitinase B1 OS = Neosartorya fumigata | IVLGMPLYGR | 11.176.317 | −0.6 | 5.598.228 | G5EAZ3—Endochitinase B; AltName: Full = Chitinase B [Aspergillus nidulans FGSC A4] | 90% | Glycosidase, Hydrolase (3.2.1.14) |
| Q9HGI3—Beta-hexosaminidase OS = Emericella nidulans OX = 162425 GN = nagA PE = 1 SV = 1 | HISWGHSGPKPLSDVSLRTERDTDDSILTNAWNR | 38.598.989 | −5.7 | 12.876.329 | Q9HGI3—Beta-hexosaminidase OS = Emericella nidulans OX = 162425 GN = nagA PE = 1 SV = 1 | 100% | Glycosidase, Hydrolase (3.2.1.52) |
| Hydrolysis Time | Integrated Area (A1345) | Integrated Area (A1460) | DG (%) | CV (%) |
|---|---|---|---|---|
| Without hydrolysis | 1.051 | 1.062 | 80 | 0.49 |
| 1 h | 1.030 | 1.154 | 83 | 3.23 |
| 3 h | 1.175 | 1.150 | 79 | 1.74 |
| 5 h | 0.991 | 1.010 | 80 | 0.49 |
| 24 h | 1.220 | 1.991 | 80 | 0.49 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gonçalves, C.G.e.; Lourenço, L.d.F.H.; Philippsen, H.K.; Santos, A.S.; Santos, L.N.d.; Ferreira, N.R. Crude Enzyme Concentrate of Filamentous Fungus Hydrolyzed Chitosan to Obtain Oligomers of Different Sizes. Polymers 2023, 15, 2079. https://doi.org/10.3390/polym15092079
Gonçalves CGe, Lourenço LdFH, Philippsen HK, Santos AS, Santos LNd, Ferreira NR. Crude Enzyme Concentrate of Filamentous Fungus Hydrolyzed Chitosan to Obtain Oligomers of Different Sizes. Polymers. 2023; 15(9):2079. https://doi.org/10.3390/polym15092079
Chicago/Turabian StyleGonçalves, Cleidiane Gonçalves e, Lúcia de Fátima Henriques Lourenço, Hellen Kempfer Philippsen, Alberdan Silva Santos, Lucely Nogueira dos Santos, and Nelson Rosa Ferreira. 2023. "Crude Enzyme Concentrate of Filamentous Fungus Hydrolyzed Chitosan to Obtain Oligomers of Different Sizes" Polymers 15, no. 9: 2079. https://doi.org/10.3390/polym15092079
APA StyleGonçalves, C. G. e., Lourenço, L. d. F. H., Philippsen, H. K., Santos, A. S., Santos, L. N. d., & Ferreira, N. R. (2023). Crude Enzyme Concentrate of Filamentous Fungus Hydrolyzed Chitosan to Obtain Oligomers of Different Sizes. Polymers, 15(9), 2079. https://doi.org/10.3390/polym15092079







