Analysis on Microstructure–Property Linkages of Filled Rubber Using Machine Learning and Molecular Dynamics Simulations
Abstract
:1. Introduction
2. Problem Setting
3. Previous Work
3.1. CNN-Based CGMD Surrogate Model
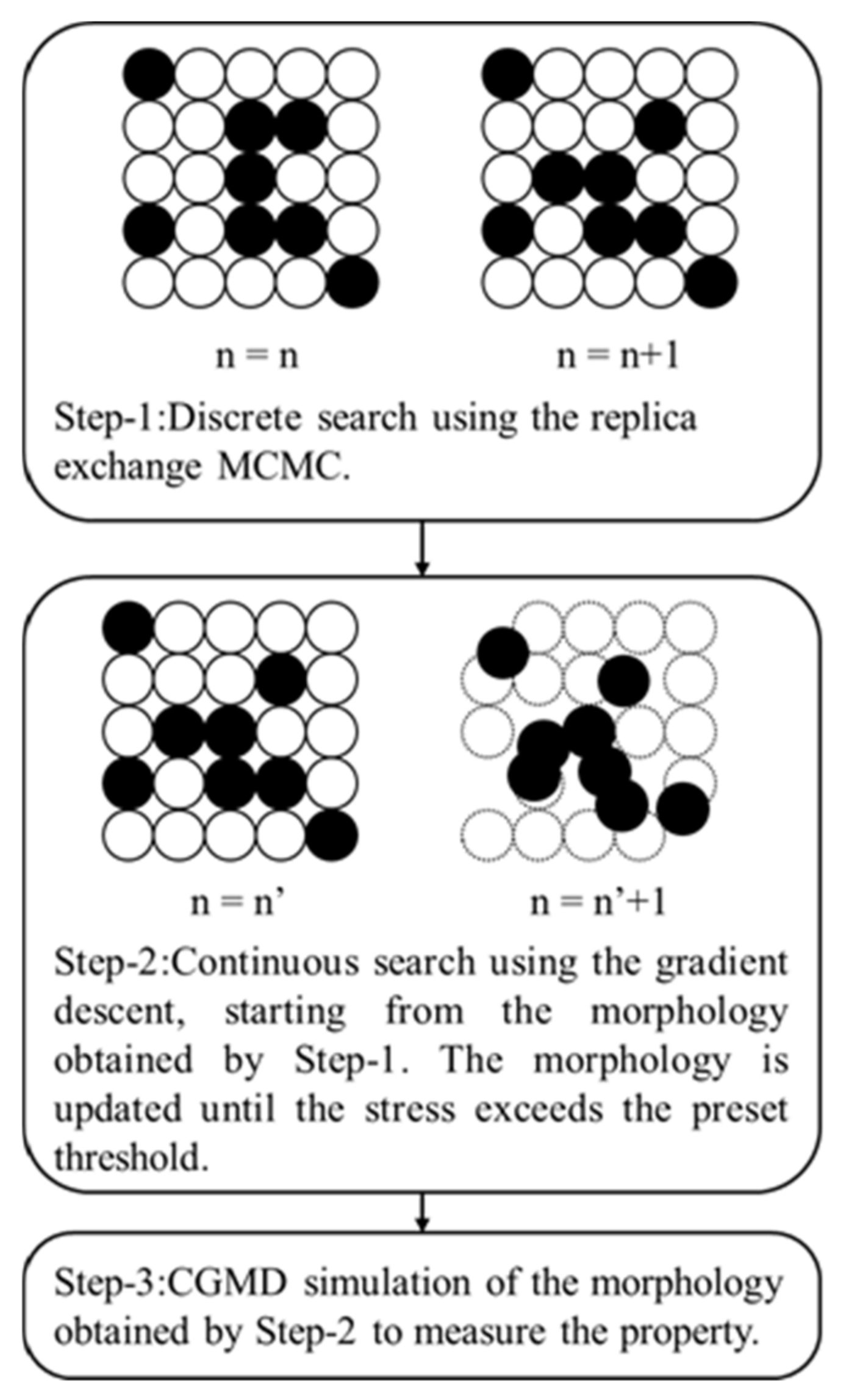
3.2. Filler Morphology Search Method
4. Method
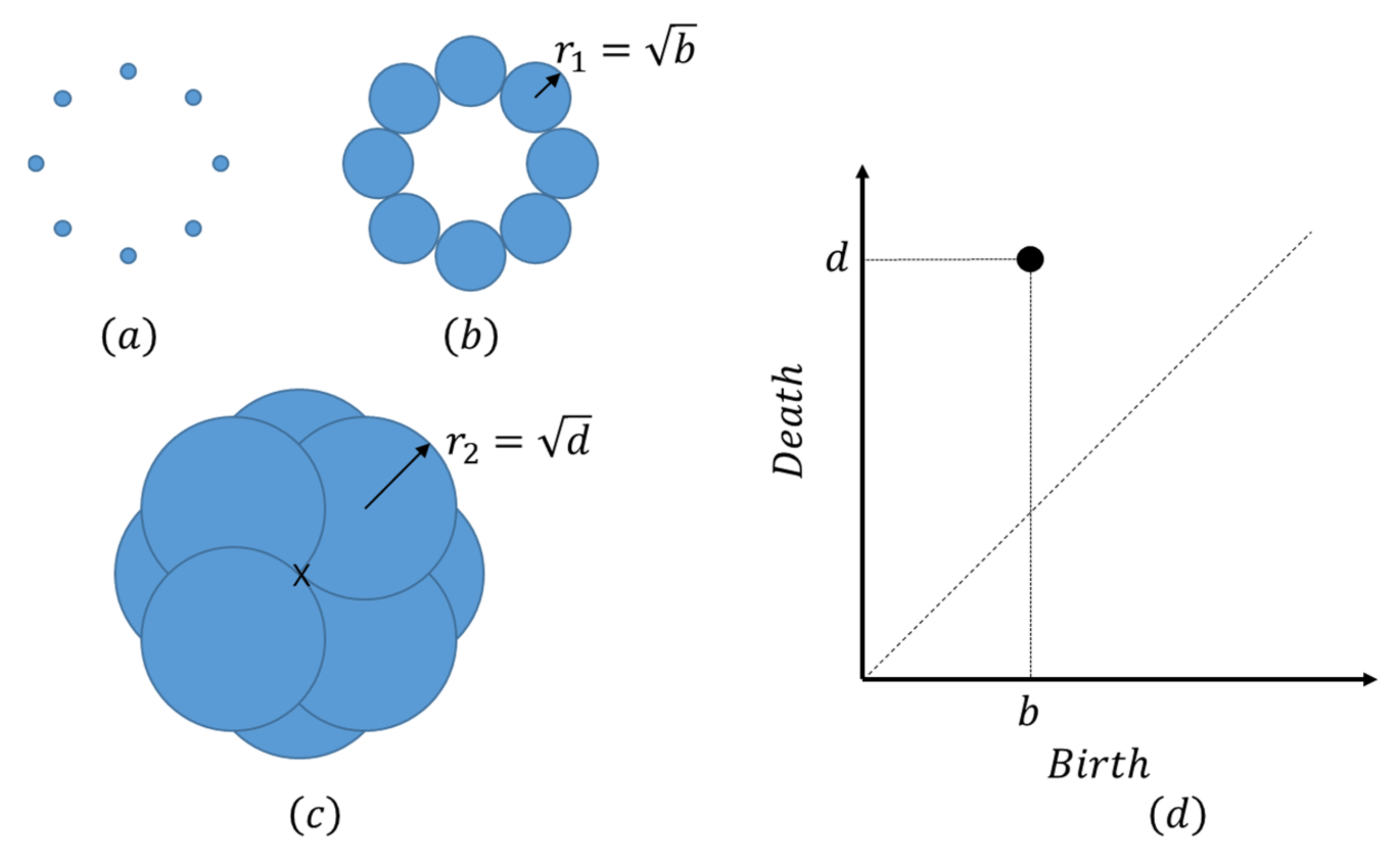
4.1. LR and PH Analyses
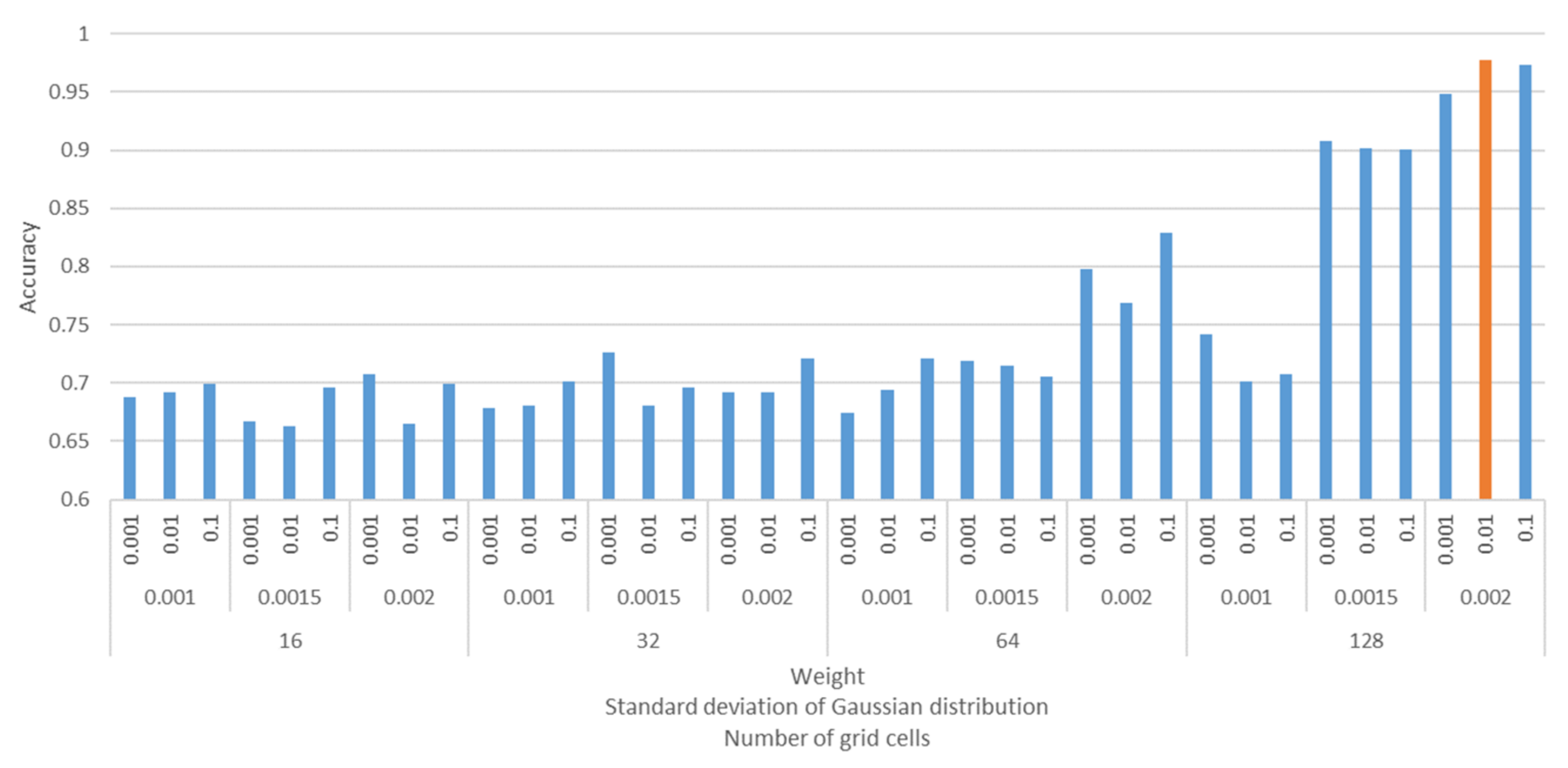
4.2. CNN-Based Analysis
5. Results and Discussion
5.1. Filler Aggregates Extracted by the PH and CNN Methods
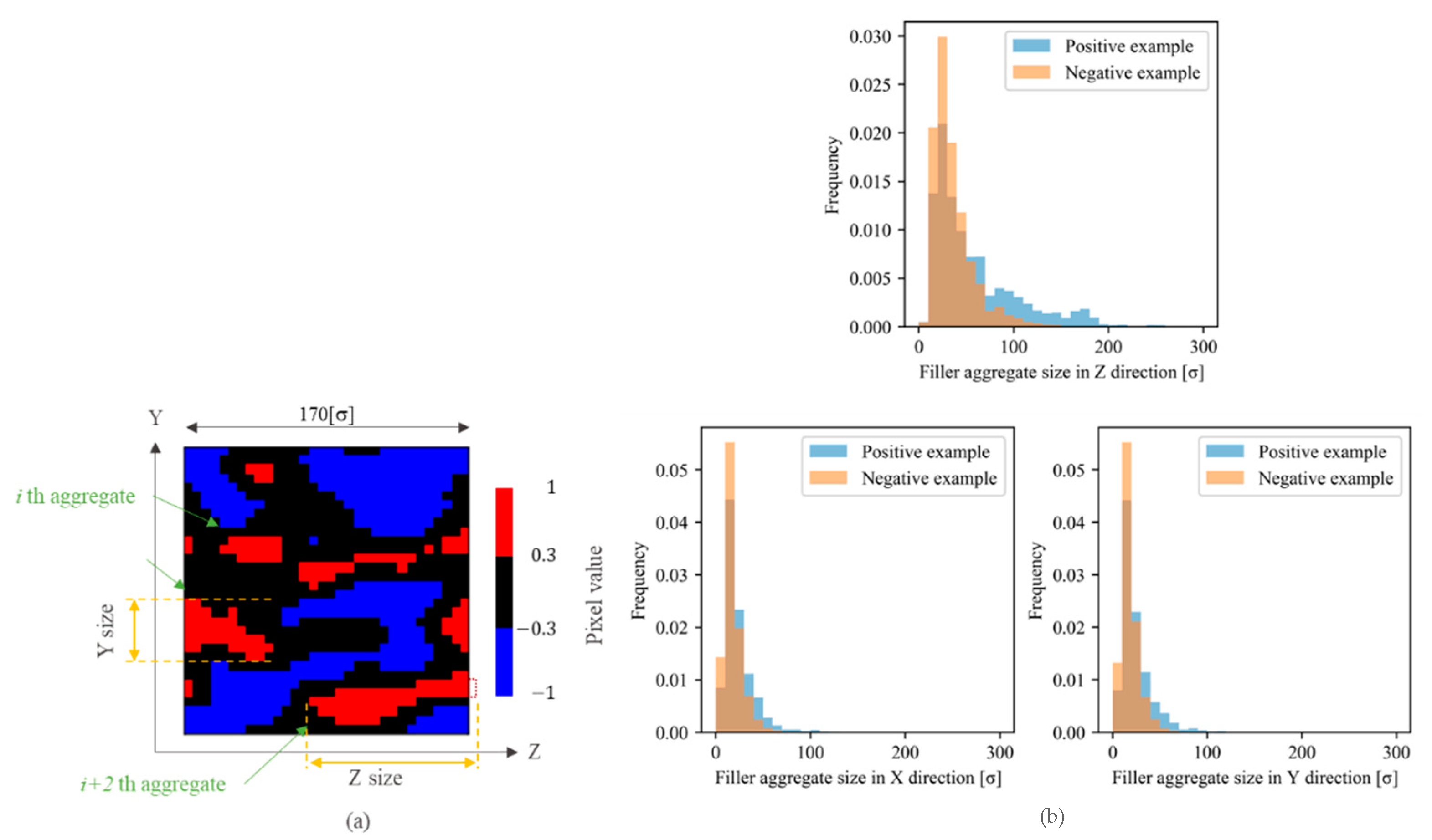
- The dense distribution of filler aggregates reflected the short distances between their surfaces. In addition, the sizes of agglomerates (quadratic aggregates) are small as shown in Figure 19.
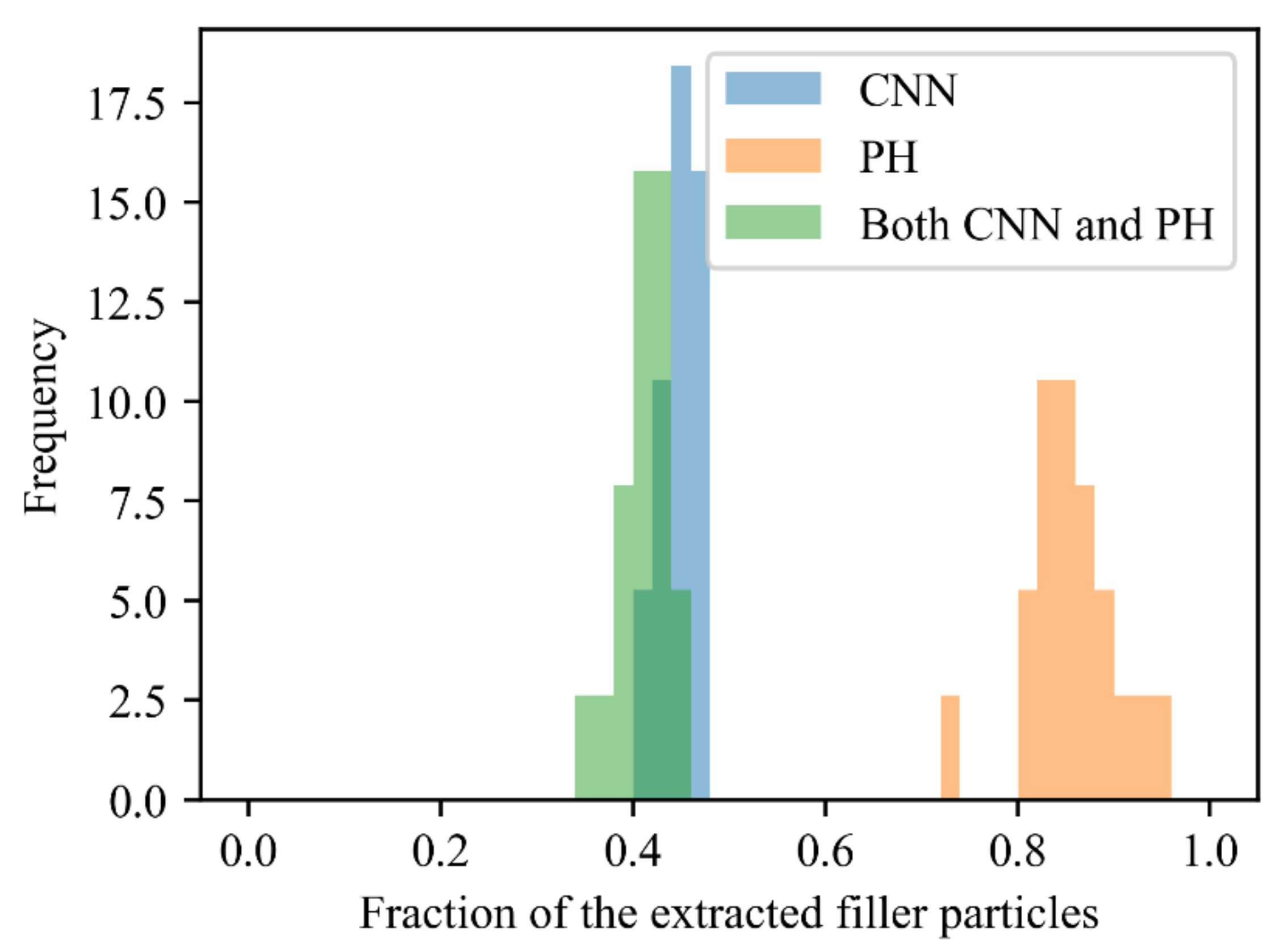
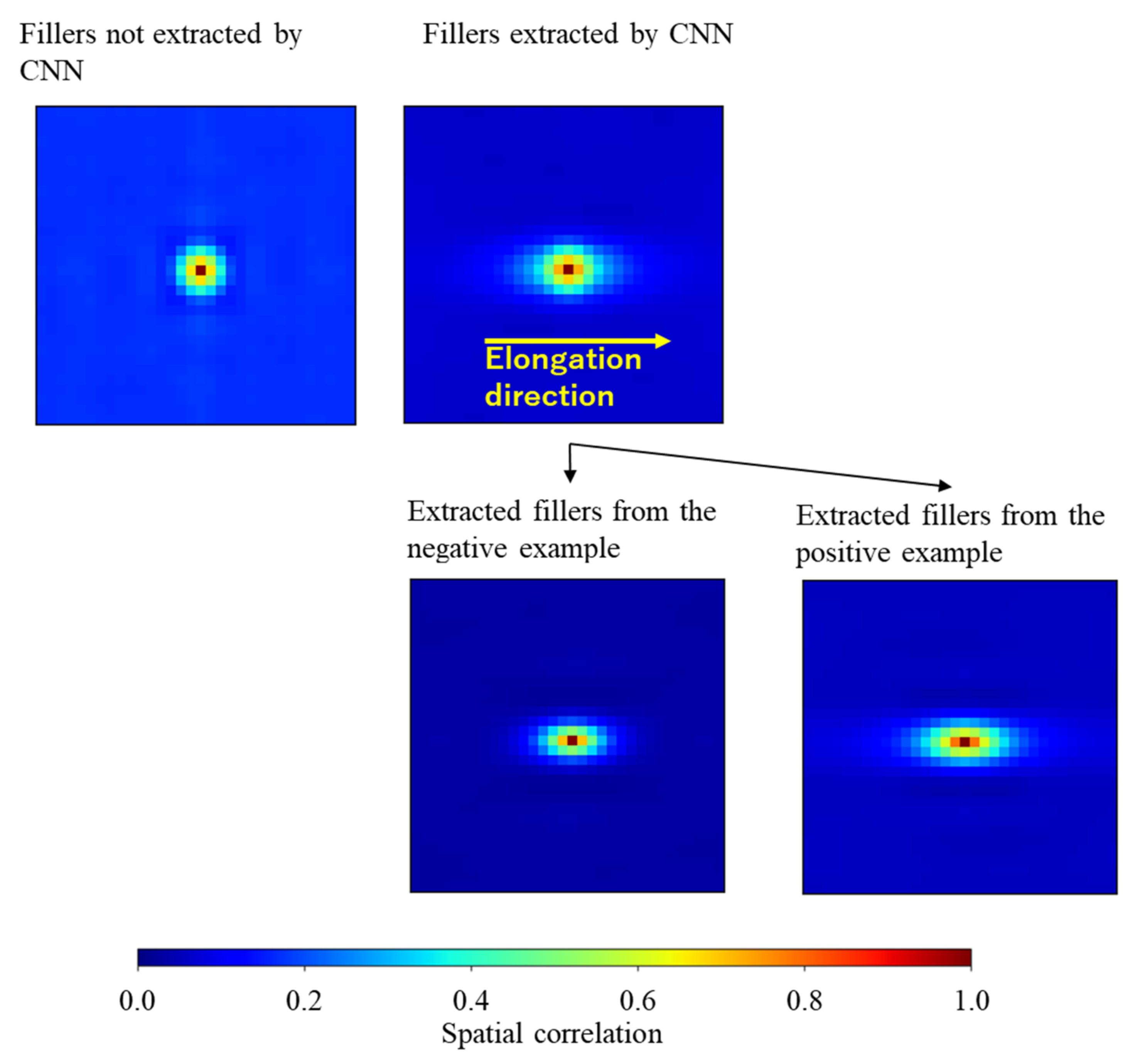
5.2. Comparison between the Extracted and Non-Extracted Filler Particles
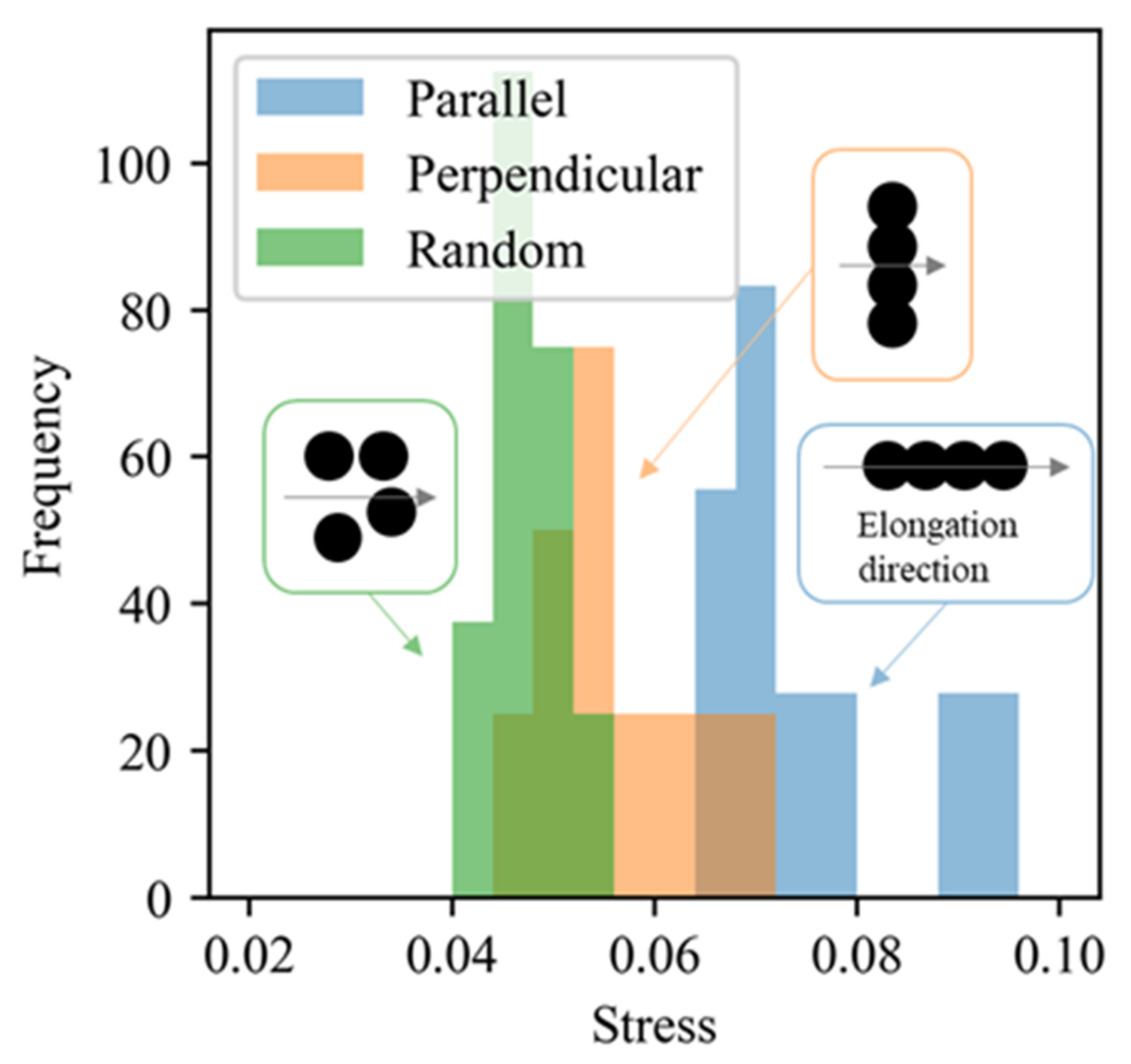
5.3. Validation Using CGMD Simulations
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Vilgis, T.A.; Heinrich, G.; Klüppel, M. Reinforcement of Polymer Nano-Composites-Theory, Experiments and Applications; Cambridge University Press: Cambridge, UK, 2009; ISBN 9780521874809. [Google Scholar]
- Tauban, M. Impact of Filler Morphology and Distribution on the Mechanical Properties of Filled Elastomers: Theory and Simulations; University of Lyon: Lyon, France, 2016; p. 199. [Google Scholar]
- Klüppel, M. The Role of Disorder in Filler Reinforcement of Elastomers on Various Length Scales. Adv. Polym. Sci. 2003, 164, 1–86. [Google Scholar] [CrossRef]
- Plagge, J.; Klüppel, M. Micromechanics of stress-softening and hysteresis of filler reinforced elastomers with applications to thermo-oxidative aging. Polymers 2020, 12, 1350. [Google Scholar] [CrossRef]
- Hashimoto, T.; Amino, N.; Nishitsuji, S.; Takenaka, M. Hierarchically self-organized filler particles in polymers: Cascade evolution of dissipative structures to ordered structures. Polym. J. 2019, 51, 109–130. [Google Scholar] [CrossRef]
- Nakajima, K.; Ito, M.; Nguyen, H.K.; Liang, X. Nanomechanics of the rubber-filler interface. Rubber Chem. Technol. 2017, 90, 272–284. [Google Scholar] [CrossRef]
- Baeza, G.P.; Genix, A.-C.; Degrandcourt, C.; Petitjean, L.; Gummel, J.; Couty, M.; Oberdisse, J. Multiscale Filler Structure in Simplified Industrial NanocompositeSilica/SBR Systems Studied by SAXS and TEM. Macromolecules 2013, 46, 317–329. [Google Scholar] [CrossRef]
- Litvinov, V.M.; Orza, R.A.; Klüppel, M.; Van Duin, M.; Magusin, P.C.M.M. Rubber-filler interactions and network structure in relation to stress-strain behavior of vulcanized, carbon black filled EPDM. Macromolecules 2011, 44, 4887–4900. [Google Scholar] [CrossRef]
- Lorenz, H.; Klüppel, M. Microstructure-based modelling of arbitrary deformation histories of filler-reinforced elastomers. J. Mech. Phys. Solids 2012, 60, 1842–1861. [Google Scholar] [CrossRef]
- Starr, F.W.; Douglas, J.F.; Glotzer, S.C. Origin of particle clustering in a simulated polymer nanocomposite and its impact on rheology. J. Chem. Phys. 2003, 119, 1777–1788. [Google Scholar] [CrossRef] [Green Version]
- Dannenberg, E.M. Effects of Surface Chemical Interactions on the Properties of Filler-Reinforced Rubbers. Rubber Chem. Technol. 1975, 48, 410–444. [Google Scholar] [CrossRef]
- Miyata, T.; Nagao, T.; Watanabe, D.; Kumagai, A.; Akutagawa, K.; Morita, H.; Jinnai, H. Nanoscale Stress Distribution in Silica-Nanoparticle-Filled Rubber as Observed by Transmission Electron Microscopy: Implications for Tire Application. Appl. Nano Mater. 2021, 12, 4452–4461. [Google Scholar] [CrossRef]
- Figliuzzi, B.; Jeulin, D.; Faessel, M.; Willot, F.; Koishi, M.; Kowatari, N. Modelling the microstructure and the viscoelastic behaviour of carbon black filled rubber materials from 3D simulations. Tech. Mech. 2016, 36, 32–56. [Google Scholar]
- Silva Bellucci, F.; Salmazo, L.O.; Budemberg, E.R.; Guerrero, A.R.; Aroca, R.F.; de Lima Nobre, M.A.; Job, A.E. Morphological characterization by SEM, TEM and AFM of nanoparticles and functional nanocomposites based on natural rubber filled with oxide nanopowders. Mater. Sci. Forum 2014, 798–799, 426–431. [Google Scholar] [CrossRef] [Green Version]
- Koishi, M.; Miyajima, H.; Kowatari, N. Conceptual Design of Tires Using Multi-Objective Design Exploration. Available online: https://docplayer.net/134334670-Conceptual-design-of-tires-using-multi-objective-design-exploration.html (accessed on 1 June 2021).
- Kaga, H.; Okamoto, K.; Tozawa, I.; Stress, Y. Analysis of a Tire Under Vertical Load by a Finite Element Method "Stress Analysis of a Tire Under Vertical Load by a Finite Element Method. Tire Sci. Technol. TSTCA 1977, 5, 102–118. [Google Scholar] [CrossRef]
- Nakajima, Y.; Kadowaki, H.; Kamegawa, T.; Ueno, K. Application of a neural network for the optimization of tire design. Tire Sci. Technol. 1999, 27, 62–83. [Google Scholar] [CrossRef]
- Nakajima, Y. Application of computational mechanics to tire design-yesterday, today, and tomorrow. Tire Sci. Technol. 2011, 39, 223–244. [Google Scholar] [CrossRef]
- Hagita, K.; Morita, H.; Takano, H. Molecular dynamics simulation study of a fracture of filler-filled polymer nanocomposites. Polymer 2016, 99, 368–375. [Google Scholar] [CrossRef] [Green Version]
- Smith, J.S.; Bedrov, D.; Smith, G.D. A molecular dynamics simulation study of nanoparticle interactions in a model polymer-nanoparticle composite. Compos. Sci. Technol. 2003, 63, 1599–1605. [Google Scholar] [CrossRef]
- Hagita, K.; Tominaga, T.; Hatazoe, T.; Sone, T.; Takano, H. Filler network model of filled rubber materials to estimate system size dependence of two-dimensional small-angle scattering patterns. J. Phys. Soc. Jpn. 2018, 87, 1–10. [Google Scholar] [CrossRef]
- Hagita, K. Nanovoids in uniaxially elongated polymer network filled with polydisperse nanoparticles via coarse-grained molecular dynamics simulation and two-dimensional scattering patterns. Polymer 2019, 174, 218–233. [Google Scholar] [CrossRef]
- Raos, G.; Moreno, M.; Elli, S. Computational experiments on filled rubber viscoelasticity: What is the role of particle-Particle interactions? Macromolecules 2006, 39, 6744–6751. [Google Scholar] [CrossRef]
- Kojima, T.; Koishi, M. Mechanisms of Mechanical Behavior of Filled Rubber by Coarse-Grained Molecular Dynamics Simulations. Tire Sci. Technol. 2020, 48, 1–30. [Google Scholar] [CrossRef]
- Kojima, T.; Koishi, M. Influence of filler dispersion on mechanical behavior with large-scale coarse-grained molecular dynamics simulation. Tech. Mech. 2018, 38, 41–54. [Google Scholar] [CrossRef]
- Kojima, T.; Washio, T.; Hara, S.; Koishi, M. Synthesis of computer simulation and machine learning for achieving the best material properties of filled rubber. Sci. Rep. 2020, 10, 1–11. [Google Scholar] [CrossRef]
- Nishi, K.; Shibayama, M. 2D pair distribution function analysis of anisotropic small-angle scattering patterns from elongated nano-composite hydrogels. Soft Matter 2017, 13, 3076–3083. [Google Scholar] [CrossRef]
- Baeza, G.P.; Genix, A.C.; Degrandcourt, C.; Petitjean, L.; Gummel, J.; Schweins, R.; Couty, M.; Oberdisse, J. Effect of grafting on rheology and structure of a simplified industrial nanocomposite silica/sbr. Macromolecules 2013, 46, 6621–6633. [Google Scholar] [CrossRef]
- Koishi, M.; Kowatari, N.; Figliuzzi, B.; Faessel, M.; Willot, F.; Jeulin, D. Computational material design of filled rubbers using multi-objective design exploration. In Constitutive Models for Rubber X; CRC Press: Boca Raton, FL, USA, 2017; pp. 467–473. [Google Scholar] [CrossRef]
- Kojima, T.; Washio, T.; Hara, S.; Koishi, M. Search Strategy for Rare Microstructure to Optimize Material Properties of Filled Rubber using Machine Learning Based Simulation. (submitted).
- Rickman, J.M.; Lookman, T.; Kalinin, S.V. Materials informatics: From the atomic-level to the continuum. Acta Mater. 2019, 168, 473–510. [Google Scholar] [CrossRef]
- Agrawal, A.; Choudhary, A. Deep materials informatics: Applications of deep learning in materials science. MRS Commun. 2019, 9, 779–792. [Google Scholar] [CrossRef] [Green Version]
- Yang, Z.; Yabansu, Y.C.; Jha, D.; Liao, W.; Choudhary, A.N.; Kalidindi, S.R.; Agrawal, A. Acta Materialia Establishing structure-property localization linkages for elastic deformation of three-dimensional high contrast composites using deep learning approaches. Acta Mater. 2019, 166, 335–345. [Google Scholar] [CrossRef]
- Yang, Z.; Yabansu, Y.C.; Al-Bahrani, R.; Liao, W.K.; Choudhary, A.N.; Kalidindi, S.R.; Agrawal, A. Deep learning approaches for mining structure-property linkages in high contrast composites from simulation datasets. Comput. Mater. Sci. 2018, 151, 278–287. [Google Scholar] [CrossRef]
- Mulholland, G.J.; Paradiso, S.P. Perspective: Materials informatics across the product lifecycle: Selection, manufacturing, and certification. APL Mater. 2016, 4, 053207. [Google Scholar] [CrossRef]
- Liu, Z.; Wu, C.T.; Koishi, M. Transfer learning of deep material network for seamless structure–property predictions. Comput. Mech. 2019, 64, 451–465. [Google Scholar] [CrossRef]
- Han, J.; Zhang, L.; Car, R. Deep Potential: A General Representation of a Many-Body Potential Energy Surface. Commun. Comput. Phys. 2018, 23, 629–639. [Google Scholar] [CrossRef] [Green Version]
- Iwasaki, Y.; Kusne, A.G.; Takeuchi, I. Comparison of dissimilarity measures for cluster analysis of X-ray diffraction data from combinatorial libraries. NPJ Comput. Mater. 2017, 3, 1–8. [Google Scholar] [CrossRef]
- Iwasaki, Y.; Sawada, R.; Stanev, V.; Ishida, M.; Kirihara, A.; Omori, Y.; Someya, H.; Takeuchi, I.; Saitoh, E.; Yorozu, S. Identification of advanced spin-driven thermoelectric materials via interpretable machine learning. NPJ Comput. Mater. 2019, 5, 6–11. [Google Scholar] [CrossRef] [Green Version]
- Yang, Z.; Yu, C.H.; Buehler, M.J. Deep learning model to predict complex stress and strain fields in hierarchical composites. Sci. Adv. 2021, 7, eabd7416. [Google Scholar] [CrossRef] [PubMed]
- Kopal, I.; Labaj, I.; Harničárová, M.; Valíček, J.; Hrubý, D. Prediction of the tensile response of carbon black filled rubber blends by artificial neural network. Polymers 2018, 10, 644. [Google Scholar] [CrossRef] [Green Version]
- Tolles, J.; Meurer, W.J. Logistic Regression Relating Patient Characteristics to Outcomes. JAMA 2016, 316, 533–534. [Google Scholar] [CrossRef]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. Available online: https://dl.acm.org/doi/10.5555/2999134.2999257 (accessed on 1 June 2021).
- LeCun, Y.; Bottou, L.; Bengio, Y.; Haffner, P. Gradient-based learning applied to document recognition. Proc. IEEE 1998, 86, 2278–2323. [Google Scholar] [CrossRef] [Green Version]
- JSOL Corporation, Japan. Available online: https://www.j-octa.com/ (accessed on 1 June 2021).
- Plimpton, S. Short-Range Molecular Dynamics. J. Comput. Phys. 1997, 117, 1–42. [Google Scholar] [CrossRef] [Green Version]
- Guth, E. Theory of Filler Reinforcement. Rubber Chem. Technol. 1945, 18, 596–604. [Google Scholar] [CrossRef]
- Karasek, L.; Meissner, B.; Asai, S.; Sumita, M. Percolation Concept: Polymer-Filler Gel Formation, Electrical Conductivity and Dynamic Electrical Properties of Carbon-Black-Filled Rubbers. Polym. J. 1996, 28, 121–126. [Google Scholar] [CrossRef] [Green Version]
- Chong, S.; Zhang, P.; Wrana, C.; Schuster, R.; Zhao, S. Combined Dielectric and Mechanical Investigation of Filler Network Percolation Behavior, Filler—Filler Contact, and Filler—Polymer Interaction on Carbon Black—Filled Hydrogenated Acrylonitrile—Butadiene Rubber. RUBBER Chem. Technol. 2014, 87, 647–663. [Google Scholar] [CrossRef]
- Hastings, W.K. Monte carlo sampling methods using Markov chains and their applications. Biometrika 1970, 57, 97–109. [Google Scholar] [CrossRef]
- PyTorch. Available online: https://pytorch.org/ (accessed on 1 June 2021).
- Earl, D.J.; Deem, M.W. Parallel tempering: Theory, applications, and new perspectives. Phys. Chem. Chem. Phys. 2005, 7, 3910–3916. [Google Scholar] [CrossRef] [Green Version]
- Zomorodian, A. Computing Persistent Homology. Discret. Comput. Geom. 2005, 33, 249–274. [Google Scholar] [CrossRef] [Green Version]
- Hiraoka, Y.; Nakamura, T.; Hirata, A.; Escolar, E.G.; Matsue, K. Hierarchical structures of amorphous solids characterized by persistent homology. Proc. Natl. Acad. Sci. USA 2016, 113, 7035–7040. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ichinomiya, T.; Obayashi, I.; Hiraoka, Y. Persistent homology analysis of craze formation. Phys. Rev. E 2017, 95, 012504. [Google Scholar] [CrossRef] [Green Version]
- Adams, H.; Emerson, T.; Kirby, M.; Neville, R.; Peterson, C.; Shipman, P.; Chepushtanova, S.; Hanson, E.; Motta, F.; Ziegelmeier, L. Persistence images: A stable vector representation of persistent homology. J. Mach. Learn. Res. 2017, 18, 1–35. [Google Scholar]
- Obayashi, I.; Hiraoka, Y.; Kimura, M. Persistence diagrams with linear machine learning models. J. Appl. Comput. Topol. 2018, 1, 421–449. [Google Scholar] [CrossRef] [Green Version]
- Maas, A.L.; Hannun, A.Y.; Ng, A.Y. Rectifier nonlinearities improve neural network acoustic models. ICML Work. Deep Learn. Audio Speech Lang. Process. 2013, 30, 1. [Google Scholar]
- Lin, M.; Chen, Q.; Yan, S. Network in network. arXiv 2014, arXiv:1312.4400. [Google Scholar]
- Ioffe, S.; Szegedy, C. Batch normalization: Accelerating deep network training by reducing internal covariate shift. In Proceedings of the 32nd International Conference on Machine Learning ICML 2015, Lile, France, 6–11 July 2015; Volume 1, pp. 448–456. Available online: http://proceedings.mlr.press/v37/ioffe15.html (accessed on 1 June 2021).
- Peng, X.; Li, L.; Wang, F.-Y. Accelerating Minibatch Stochastic Gradient Descent Using Typicality Sampling. IEEE Trans. Neural Netw. Learn. Syst. 2019, 31, 1–11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kingma, D.P.; Ba, J.L. Adam: A method for stochastic optimization. arXiv 2017, arXiv:1412.6980. [Google Scholar]
- Akutagawa, K.; Yamaouchi, K.; Yamamoto, A.; Heouri, H.; Jinnai, H.; Shinbori, Y. Mesoscopic mechanical analysis of filled elastomer with 3D-finite element analysis and transmission electron microtomography. Rubber Chem. Technol. 2008, 81, 182–189. [Google Scholar] [CrossRef]
- Torquato, S.; Beasley, J.D.; Chiew, Y.C. Two-point cluster function for continuum percolation. J. Chem. Phys. 1988, 88, 6540–6547. [Google Scholar] [CrossRef] [Green Version]
- Cecen, A.; Dai, H.; Yabansu, Y.C.; Kalidindi, S.R.; Song, L. Material structure-property linkages using three-dimensional convolutional neural networks. Acta Mater. 2018, 146, 76–84. [Google Scholar] [CrossRef]
- Latypov, M.I.; Kalidindi, S.R. Data-driven reduced order models for effective yield strength and partitioning of strain in multiphase materials. J. Comput. Phys. 2017, 346, 242–261. [Google Scholar] [CrossRef]
- Cang, R.; Li, H.; Yao, H.; Jiao, Y.; Ren, Y. Improving direct physical properties prediction of heterogeneous materials from imaging data via convolutional neural network and a morphology-aware generative model. Comput. Mater. Sci. 2018, 150, 212–221. [Google Scholar] [CrossRef] [Green Version]









Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kojima, T.; Washio, T.; Hara, S.; Koishi, M.; Amino, N. Analysis on Microstructure–Property Linkages of Filled Rubber Using Machine Learning and Molecular Dynamics Simulations. Polymers 2021, 13, 2683. https://doi.org/10.3390/polym13162683
Kojima T, Washio T, Hara S, Koishi M, Amino N. Analysis on Microstructure–Property Linkages of Filled Rubber Using Machine Learning and Molecular Dynamics Simulations. Polymers. 2021; 13(16):2683. https://doi.org/10.3390/polym13162683
Chicago/Turabian StyleKojima, Takashi, Takashi Washio, Satoshi Hara, Masataka Koishi, and Naoya Amino. 2021. "Analysis on Microstructure–Property Linkages of Filled Rubber Using Machine Learning and Molecular Dynamics Simulations" Polymers 13, no. 16: 2683. https://doi.org/10.3390/polym13162683
APA StyleKojima, T., Washio, T., Hara, S., Koishi, M., & Amino, N. (2021). Analysis on Microstructure–Property Linkages of Filled Rubber Using Machine Learning and Molecular Dynamics Simulations. Polymers, 13(16), 2683. https://doi.org/10.3390/polym13162683





