Convolutional Neural Networks for Crystal Material Property Prediction Using Hybrid Orbital-Field Matrix and Magpie Descriptors
Abstract
:1. Introduction
- (1)
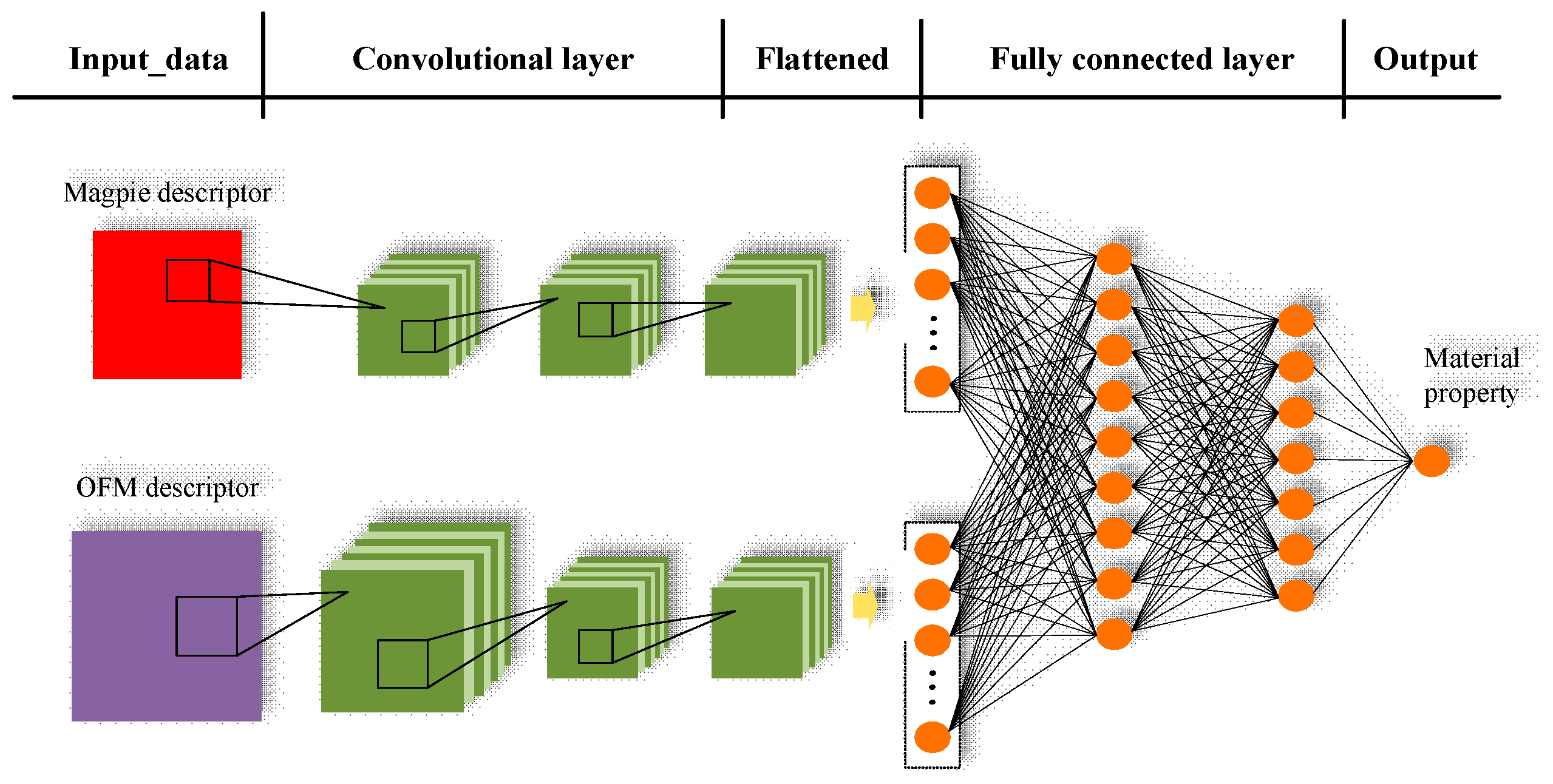
- We proposed CNN-OFM-Magpie, a convolution neural network model for materials formation energy prediction by exploiting its hierarchical feature extraction capabilities and fusion of two different types of features.
- (2)
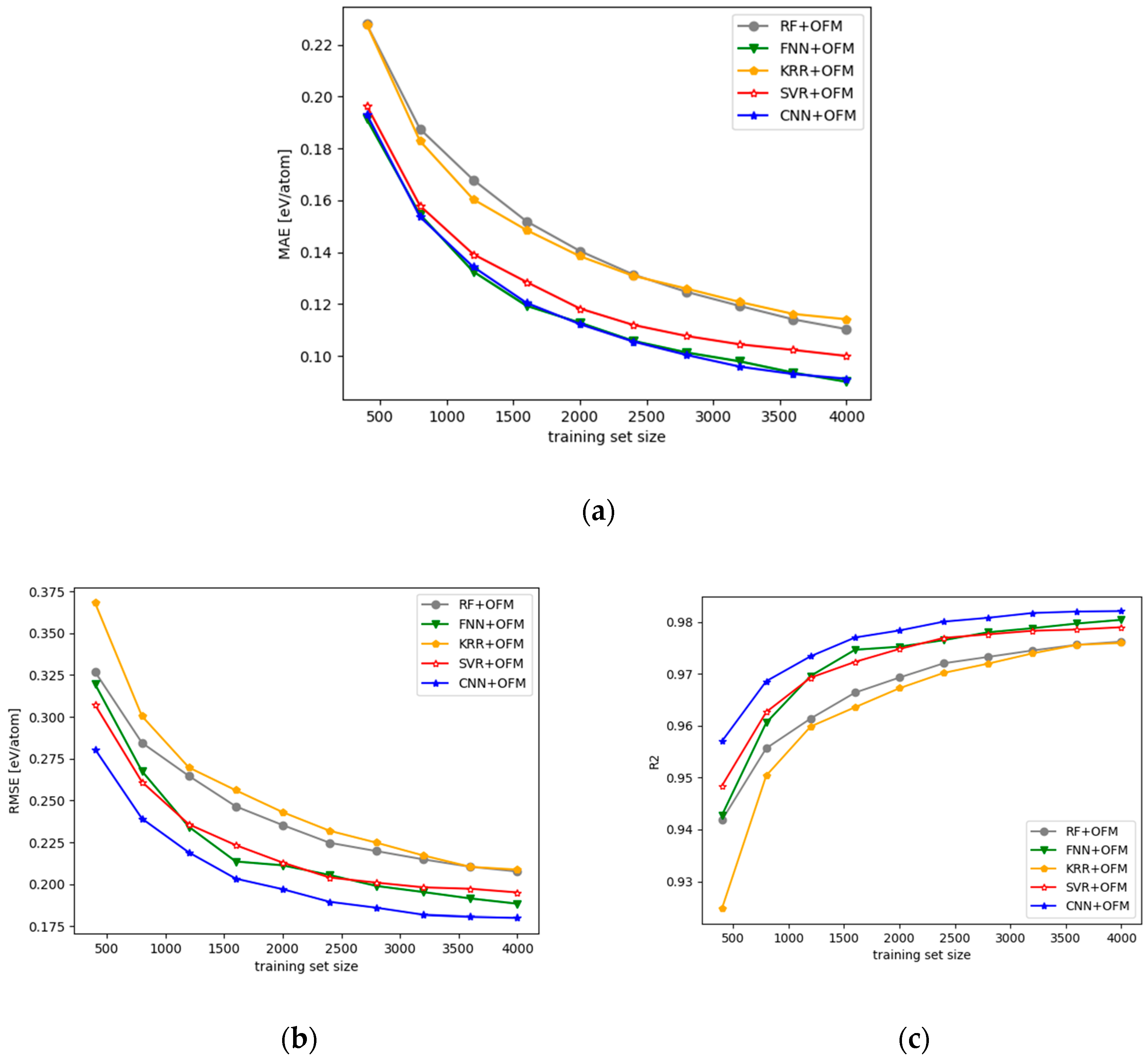
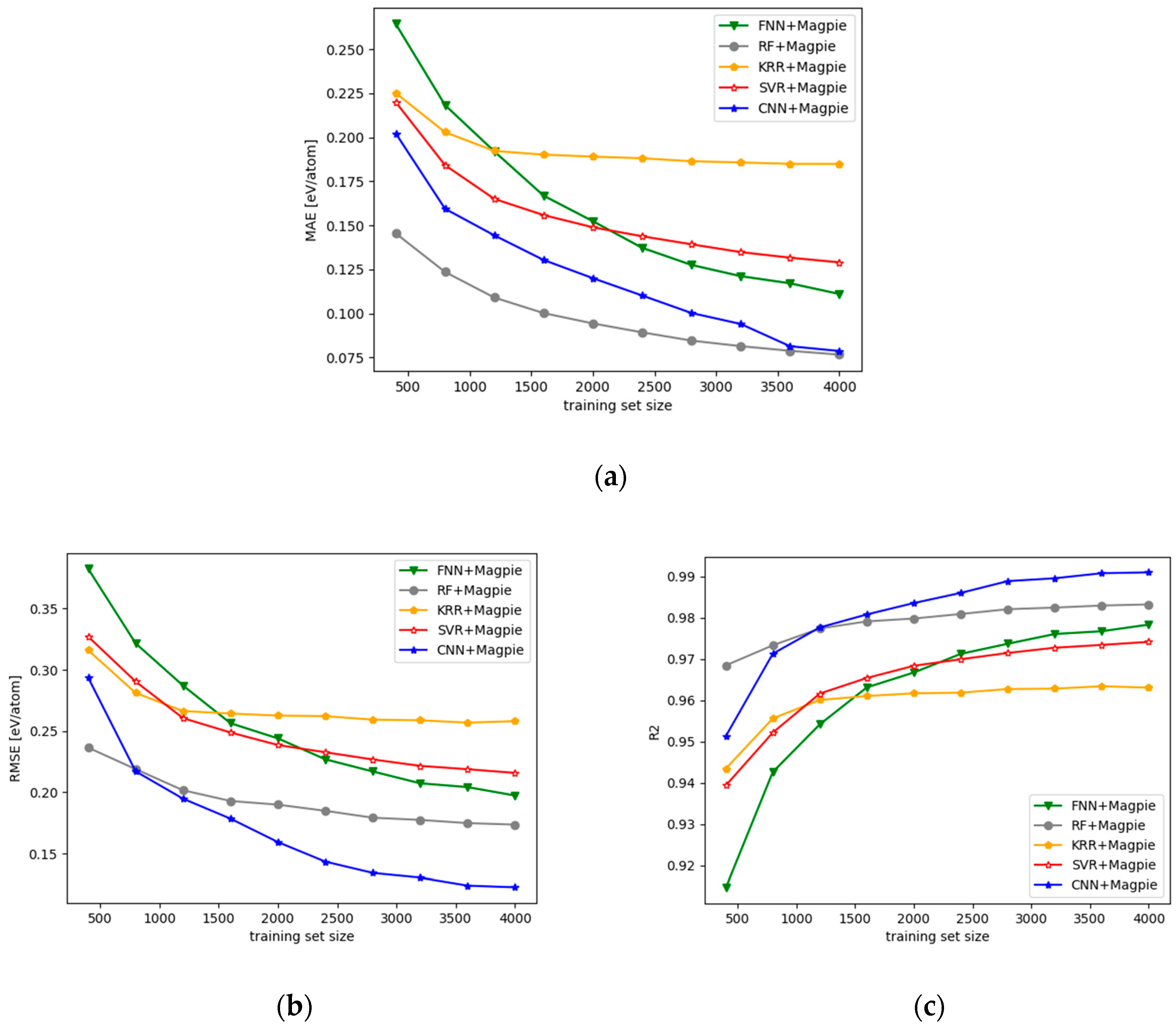
- We evaluated the performance of CNN-OFM and compared it with those of the regression prediction models based on conventional machine learning algorithms such as SVM, Random Forest, and KRR using OFM features and Magpie features, and showed the advantages of the CNN model.
- (3)
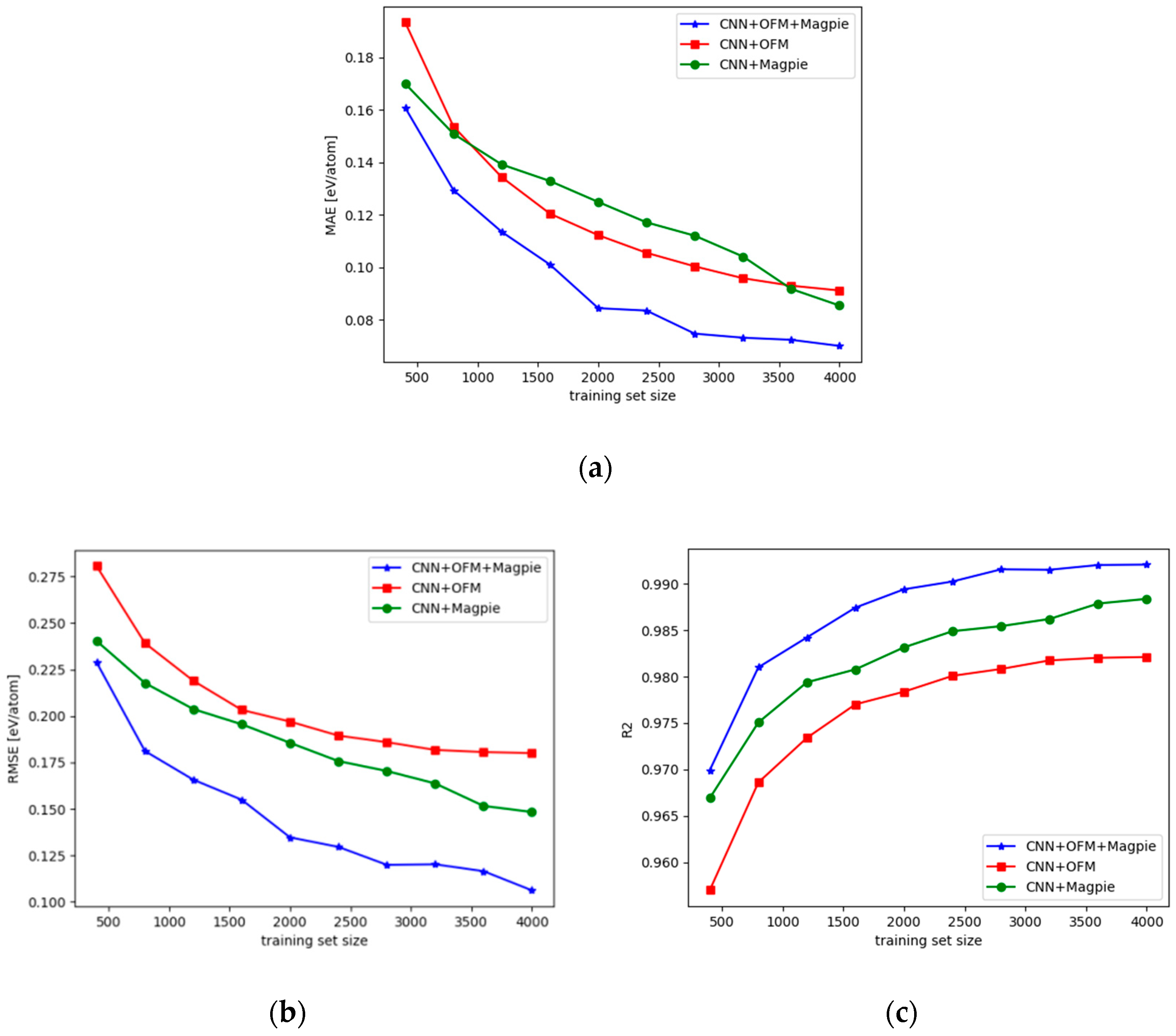
- We also compared the performance of the CNN models with hybrid descriptors with those with only one type of features. We found that feature fusion is important to achieve the highest formation energy prediction performance over the tested dataset.
- (4)
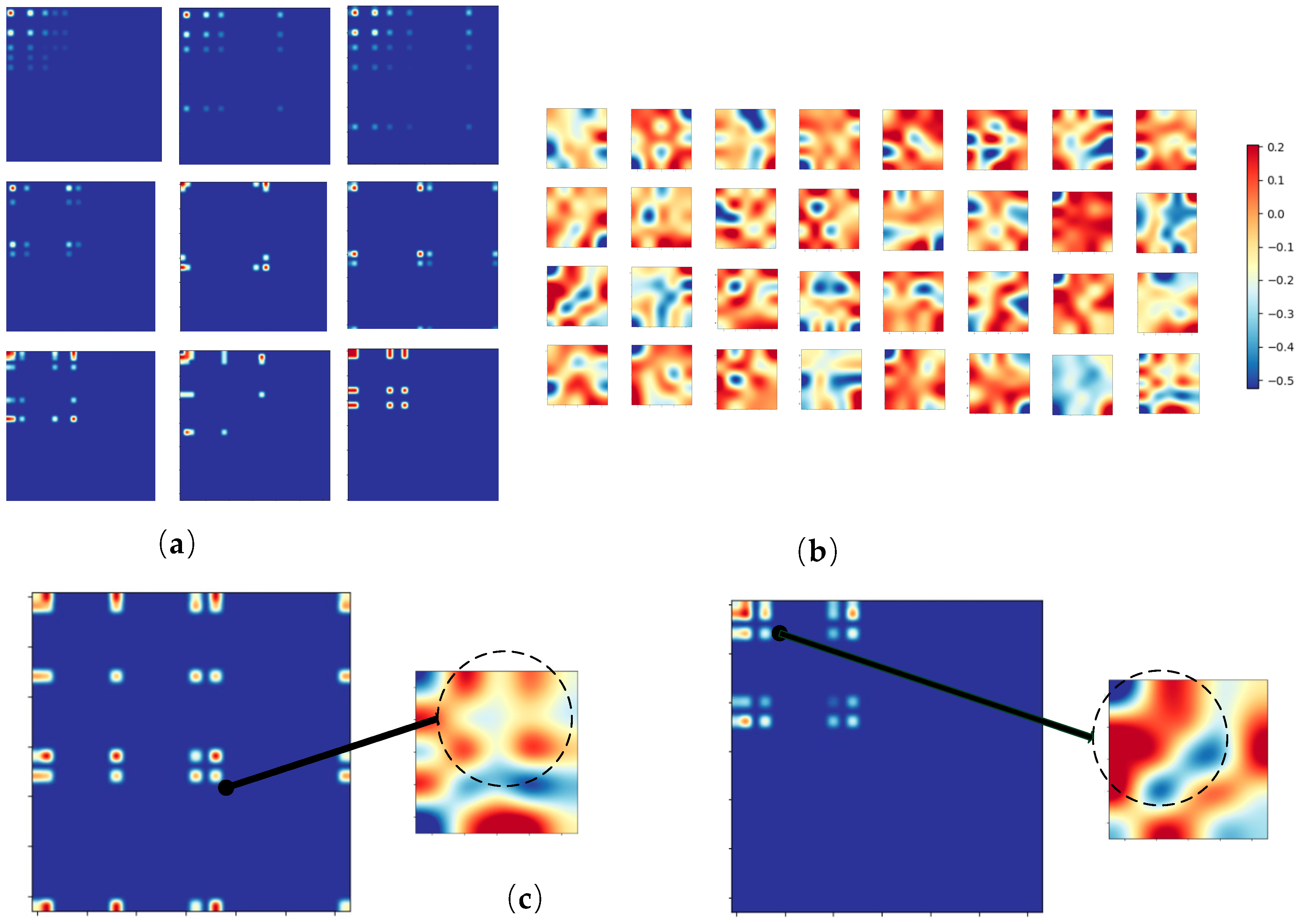
- Through visualization of the features extracted by the filters of the learned convolution neural network, interpretable analysis of CNN-OFM is provided.
2. Materials and Methods
2.1. Materials Dataset Preparation
2.2. Orbital Field Matrix Representation of Materials
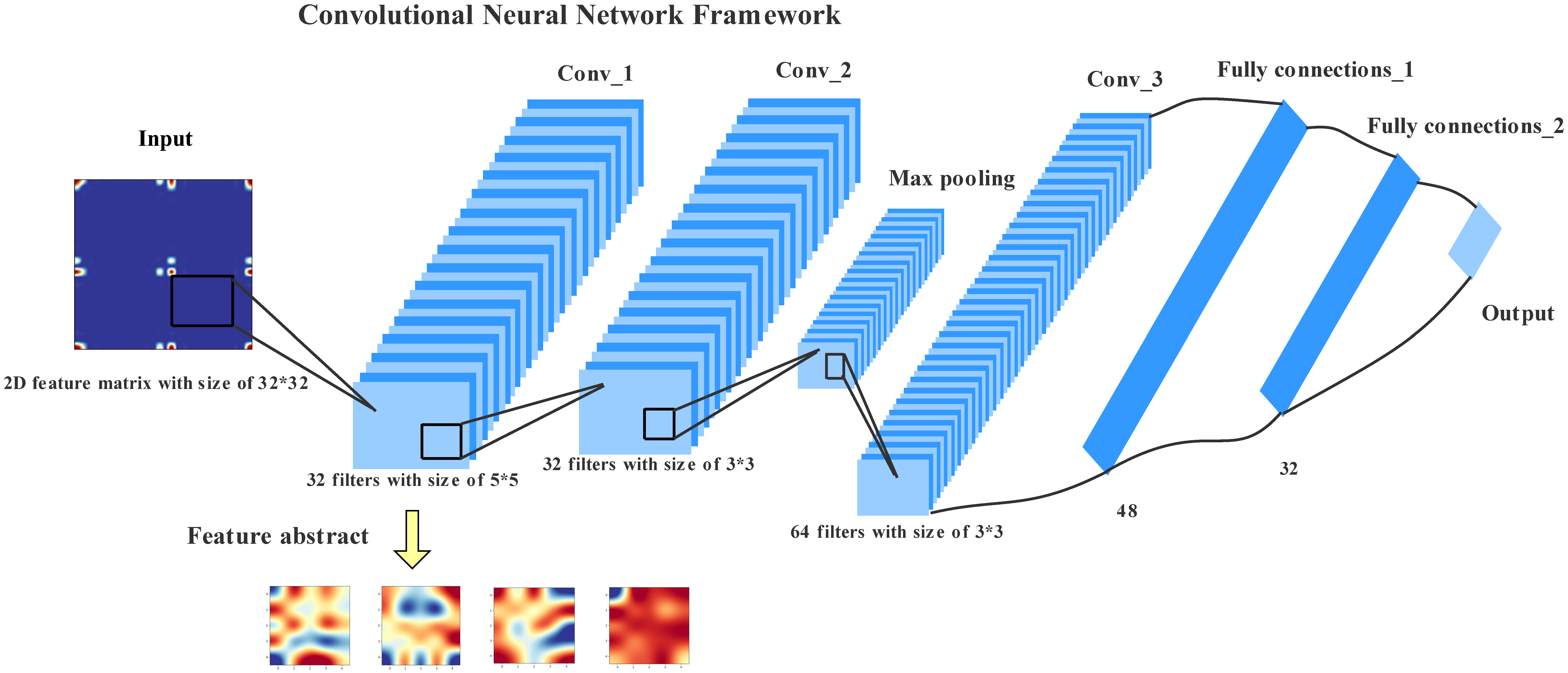
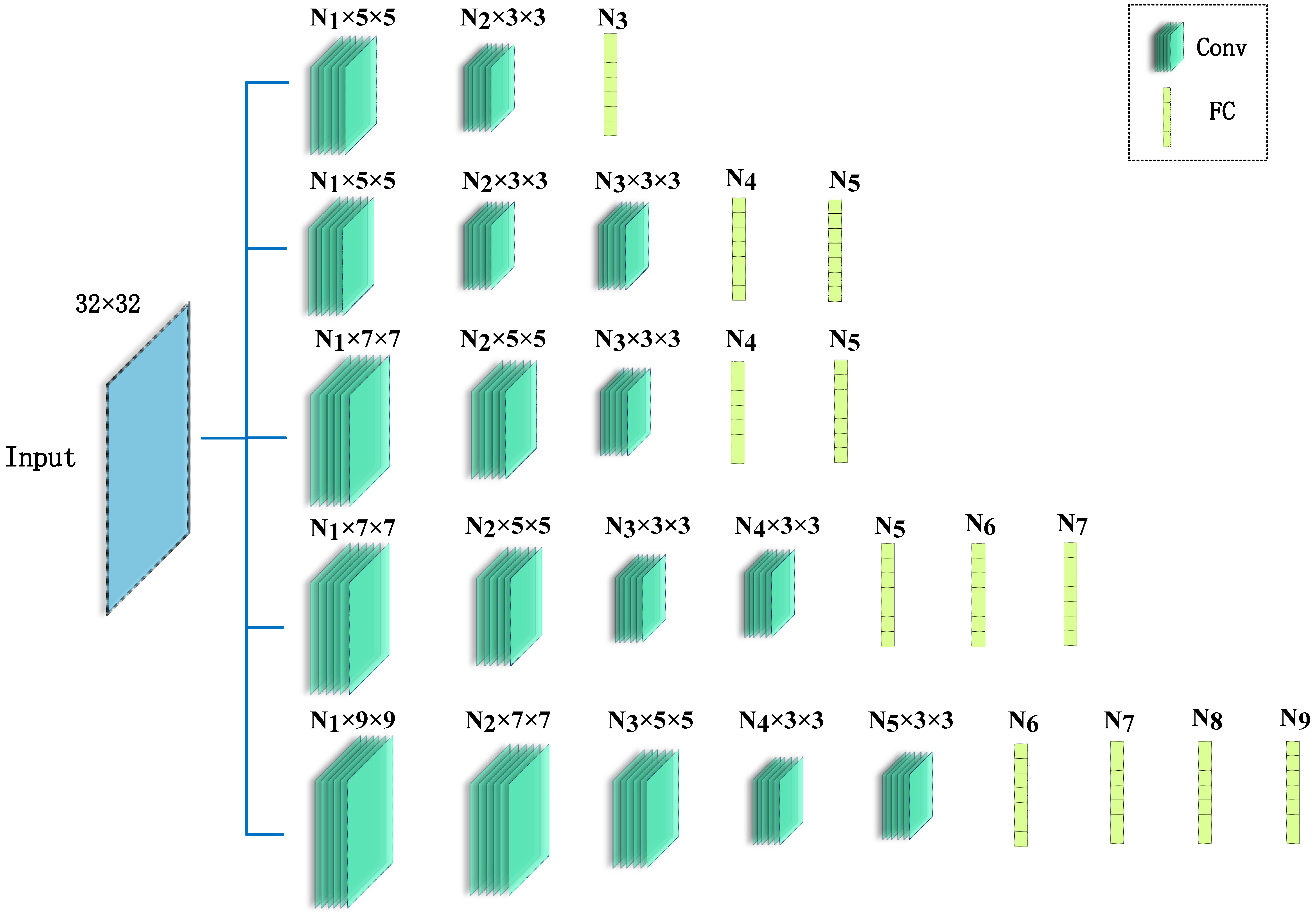
2.3. Convolutional Neural Networks Model
2.4. Regression Algorithms with One-Dimensional Input
2.5. Hyperparameters Tuning Strategies
3. Results and Discussions
3.1. Performance of the CNN Models with 2D OFM Features
3.2. Analysis over the Features Extracted by the CNN Model
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Takahashi, K.; Tanaka, Y. Materials informatics: A journey towards material design and synthesis. Dalton Trans. 2016, 45, 1497–1499. [Google Scholar] [CrossRef]
- Butler, K.T.; Davies, D.W.; Cartwright, H.; Isayev, O.; Walsh, A. Machine learning for molecular and materials science. Nature 2018, 559, 547–555. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Liu, L.; Chen, W.; An, L. Materials genome: Research progress, challenges and outlook. Sci. Sin. Chim. 2018, 48, 243–255. [Google Scholar] [CrossRef]
- Ramprasad, R.; Batra, R.; Pilania, G.; Mannodi-Kanakkithodi, A.; Kim, C. Machine learning in materials informatics: Recent applications and prospects. NPJ Comput. Mater. 2017, 3, 54. [Google Scholar] [CrossRef]
- Ward, L.; Wolverton, C. Atomistic calculations and materials informatics: A review. Curr. Opin. Solid State Mat. Sci. 2017, 21, 167–176. [Google Scholar] [CrossRef]
- Meredig, B.; Agrawal, A.; Kirklin, S.; Saal, J.E.; Doak, J.W.; Thompson, A.; Zhang, K.; Choudhary, A.; Wolverton, C. Combinatorial screening for new materials in unconstrained composition space with machine learning. Phys. Rev. B 2014, 89, 0941049. [Google Scholar] [CrossRef]
- Seko, A.; Togo, A.; Tanaka, I. Descriptors for Machine Learning of Materials Data. In Nanoinformatics; Tanaka, I., Ed.; Springer: Singapore, 2018. [Google Scholar]
- Swann, E.; Sun, B.; Cleland, D.M.; Barnard, A.S. Representing molecular and materials data for unsupervised machine learning. Mol. Simul. 2018, 44, 905–920. [Google Scholar] [CrossRef]
- Ghiringhelli, L.M.; Vybiral, J.; Levchenko, S.V.; Draxl, C.; Scheffler, M. Big Data of Materials Science: Critical Role of the Descriptor. Phys. Rev. Lett. 2015, 114, 105503. [Google Scholar] [CrossRef]
- Ziletti, A.; Kumar, D.; Scheffler, M.; Ghiringhelli, L.M. Insightful classification of crystal structures using deep learning. Nat. Commun. 2018, 9, 2775. [Google Scholar] [CrossRef]
- Calfa, B.A.; Kitchin, J.R. Property Prediction of Crystalline Solids from Composition and Crystal Structure. Aiche J. 2016, 62, 2605–2613. [Google Scholar] [CrossRef]
- Ward, L.; Agrawal, A.; Choudhary, A.; Wolverton, C. A general-purpose machine learning framework for predicting properties of inorganic materials. NPJ Comput. Mater. 2016, 2, 16028. [Google Scholar] [CrossRef]
- Faber, F.; Lindmaa, A.; Von Lilienfeld, O.A.; Armiento, R. Crystal structure representations for machine learning models of formation energies. Int. J. Quantum Chem. 2015, 115, 1094–1101. [Google Scholar] [CrossRef]
- Tien, L.P.; Kino, H.; Terakura, K.; Miyake, T.; Tsuda, K.; Takigawa, I.; Hieu, C.D. Machine learning reveals orbital interaction in materials. Sci. Technol. Adv. Mater. 2017, 18, 756–765. [Google Scholar]
- Zhou, Q.; Tang, P.; Liu, S.; Pan, J.; Yan, Q.; Zhang, S. Learning atoms for materials discovery. PNAS 2018, 115, E6411–E6417. [Google Scholar] [CrossRef] [PubMed]
- Hansen, K.; Montavon, G.; Biegler, F.; Fazli, S.; Rupp, M.; Scheffler, M.; von Lilienfeld, O.A.; Tkatchenko, A.; Müller, K. Assessment and Validation of Machine Learning Methods for Predicting Molecular Atomization Energies. J. Chem. Theory Comput. 2013, 9, 3404–3419. [Google Scholar] [CrossRef]
- Kajita, S.; Ohba, N.; Jinnouchi, R.; Asahi, R. A Universal 3D Voxel Descriptor for Solid-State Material Informatics with Deep Convolutional Neural Networks. Sci. Rep. 2017, 7, 16991. [Google Scholar] [CrossRef]
- De, S.; Bartok, A.P.; Csanyi, G.; Ceriotti, M. Comparing molecules and solids across structural and alchemical space. Phys. Chem. Chem. Phys. 2016, 18, 13754–13769. [Google Scholar] [CrossRef]
- Cecen, A.; Dai, H.; Yabansu, Y.C.; Kalidindi, S.R.; Song, L. Material structure-property linkages using three-dimensional convolutional neural networks. Acta Mater. 2018, 146, 76–84. [Google Scholar] [CrossRef]
- Xie, T.; Grossman, J.C. Crystal Graph Convolutional Neural Networks for an Accurate and Interpretable Prediction of Material Properties. Phys. Rev. Lett. 2018, 120, 145301. [Google Scholar] [CrossRef] [PubMed]
- Jain, A.; Shyue, P.O.; Hautier, G.; Chen, W.; Richards, W.D.; Dacek, S.; Cholia, S.; Gunter, D.; Skinner, D.; Ceder, G.; et al. Commentary: The Materials Project: A materials genome approach to accelerating materials innovation. APL Mater. 2013, 1, 0110021. [Google Scholar] [CrossRef]
- Ward, L.; Dunn, A.; Faghaninia, A.; Zimmermann, N.E.R.; Bajaj, S.; Wang, Q.; Montoya, J.; Chen, J.; Bystrom, K.; Dylla, M.; et al. Matminer: An open source toolkit for materials data mining. Comput. Mater. Sci. 2018, 152, 60–69. [Google Scholar] [CrossRef]
- Ong, S.P.; Richards, W.D.; Jain, A.; Hautier, G.; Kocher, M.; Cholia, S.; Gunter, D.; Chevrier, V.L.; Persson, K.A.; Ceder, G. Python Materials Genomics (pymatgen): A robust, open-source python library for materials analysis. Comput. Mater. Sci. 2013, 68, 314–319. [Google Scholar] [CrossRef]
- Azimi, S.M.; Britz, D.; Engstler, M.; Fritz, M.; Mücklich, F. Advanced Steel Microstructural Classification by Deep Learning Methods. Sci. Rep. 2018, 8, 2128. [Google Scholar] [CrossRef]
- Srivastava, R.K.; Greff, K.; Schmidhuber, J. Training Very Deep Networks. 2015. Eprint arXiv:1507.06228. Available online: https://arxiv.org/pdf/1507.06228.pdf (accessed on 9 March 2019).
- Lecun, Y.; Bottou, L.; Bengio, Y.; Haffner, P. Gradient-based learning applied to document recognition. Proc. IEEE 1998, 86, 2278–2324. [Google Scholar] [CrossRef]
- Glorot, X.; Bordes, A.; Bengio, Y. Deep sparse rectifier neural networks. In Proceedings of the 14th International Conference on Artificial Intelligence and Statistics, Ft. Lauderdale, FL, USA, 11–13 April 2011; pp. 315–323. [Google Scholar]
- Kingma, D.; Ba, J. Adam: A Method for Stochastic Optimization. In Proceedings of the 3rd International Conference for Learning Representations, San Diego, CA, USA, 7–9 May 2015. [Google Scholar]
- Chollet, F. Keras. 2015. Available online: https://keras.io (accessed on 9 March 2019).
- Abadi, M.; Agarwal, A.; Barham, P.; Brevdo, E.; Chen, Z.; Citro, C.; Corrado, G.S.; Davis, A.; Dean, J.; Devin, M.; et al. TensorFlow: Large-Scale Machine Learning on Heterogeneous Distributed Systems. 2016. Available online: https://www.tensorflow.org (accessed on 9 March 2019).
- Pedregosa, F.; Varoquaux, G.E.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Shahriari, B.; Swersky, K.; Wang, Z.; Adams, R.P.; de Freitas, N. Taking the Human Out of the Loop: A Review of Bayesian Optimization. Proc. IEEE 2016, 104, 148–175. [Google Scholar] [CrossRef]
- Hertel, L.; Collado, J.; Sadowski, P.; Baldi, P. Sherpa: Hyperparameter Optimization for Machine Learning Models. In Proceedings of the 32nd Conference on Neural Information Processing Systems, Montreal, QC, Canada, 3–8 December 2018. [Google Scholar]
- Ahneman, D.T.; Estrada, J.G.; Lin, S.; Dreher, S.D.; Doyle, A.G. Predicting reaction performance in C–N cross-coupling using machine learning. Science 2018, 360, 186–190. [Google Scholar] [CrossRef]
- Schütt, K.T.; Sauceda, H.E.; Kindermans, P.J.; Tkatchenko, A.; Müller, K.R. SchNet—A deep learning architecture for molecules and materials. J. Chem. Phys. 2017, 148, 241722. [Google Scholar] [CrossRef] [PubMed]
| Regression Model | RMSE | MAE | R2 |
|---|---|---|---|
| SVR | 0.1950 | 0.1000 | 0.9790 |
| KRR | 0.2054 | 0.1174 | 0.9767 |
| RF | 0.2075 | 0.1103 | 0.9762 |
| FNN | 0.1941 | 0.1037 | 0.9791 |
| CNN | 0.1800 | 0.0911 | 0.9821 |
| Regression Model | RMSE | MAE | R2 |
|---|---|---|---|
| SVR | 0.2158 | 0.1290 | 0.9741 |
| KRR | 0.2580 | 0.1849 | 0.9630 |
| RF | 0.1736 | 0.0778 | 0.9832 |
| FNN | 0.1973 | 0.1110 | 0.9783 |
| CNN | 0.1227 | 0.0786 | 0.9910 |
| Descriptor | RMSE | MAE | R2 |
|---|---|---|---|
| OFM | 0.1800 | 0.0911 | 0.9821 |
| Magpie | 0.1227 | 0.0786 | 0.9910 |
| OFM + Magpie | 0.1062 | 0.0700 | 0.9920 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cao, Z.; Dan, Y.; Xiong, Z.; Niu, C.; Li, X.; Qian, S.; Hu, J. Convolutional Neural Networks for Crystal Material Property Prediction Using Hybrid Orbital-Field Matrix and Magpie Descriptors. Crystals 2019, 9, 191. https://doi.org/10.3390/cryst9040191
Cao Z, Dan Y, Xiong Z, Niu C, Li X, Qian S, Hu J. Convolutional Neural Networks for Crystal Material Property Prediction Using Hybrid Orbital-Field Matrix and Magpie Descriptors. Crystals. 2019; 9(4):191. https://doi.org/10.3390/cryst9040191
Chicago/Turabian StyleCao, Zhuo, Yabo Dan, Zheng Xiong, Chengcheng Niu, Xiang Li, Songrong Qian, and Jianjun Hu. 2019. "Convolutional Neural Networks for Crystal Material Property Prediction Using Hybrid Orbital-Field Matrix and Magpie Descriptors" Crystals 9, no. 4: 191. https://doi.org/10.3390/cryst9040191
APA StyleCao, Z., Dan, Y., Xiong, Z., Niu, C., Li, X., Qian, S., & Hu, J. (2019). Convolutional Neural Networks for Crystal Material Property Prediction Using Hybrid Orbital-Field Matrix and Magpie Descriptors. Crystals, 9(4), 191. https://doi.org/10.3390/cryst9040191





