Crystal Structure of the Active Site Mutant Form of Soluble Fumarate Reductase, Osm1
Abstract
1. Introduction
2. Materials and Methods
2.1. Site-Directed Mutagenesis
2.2. Protein Expression and Purification
2.3. Multi-Angle Light Scattering (MALS)
2.4. Structure Determination and Analysis
2.5. Protein Data Bank Accession Code
3. Results
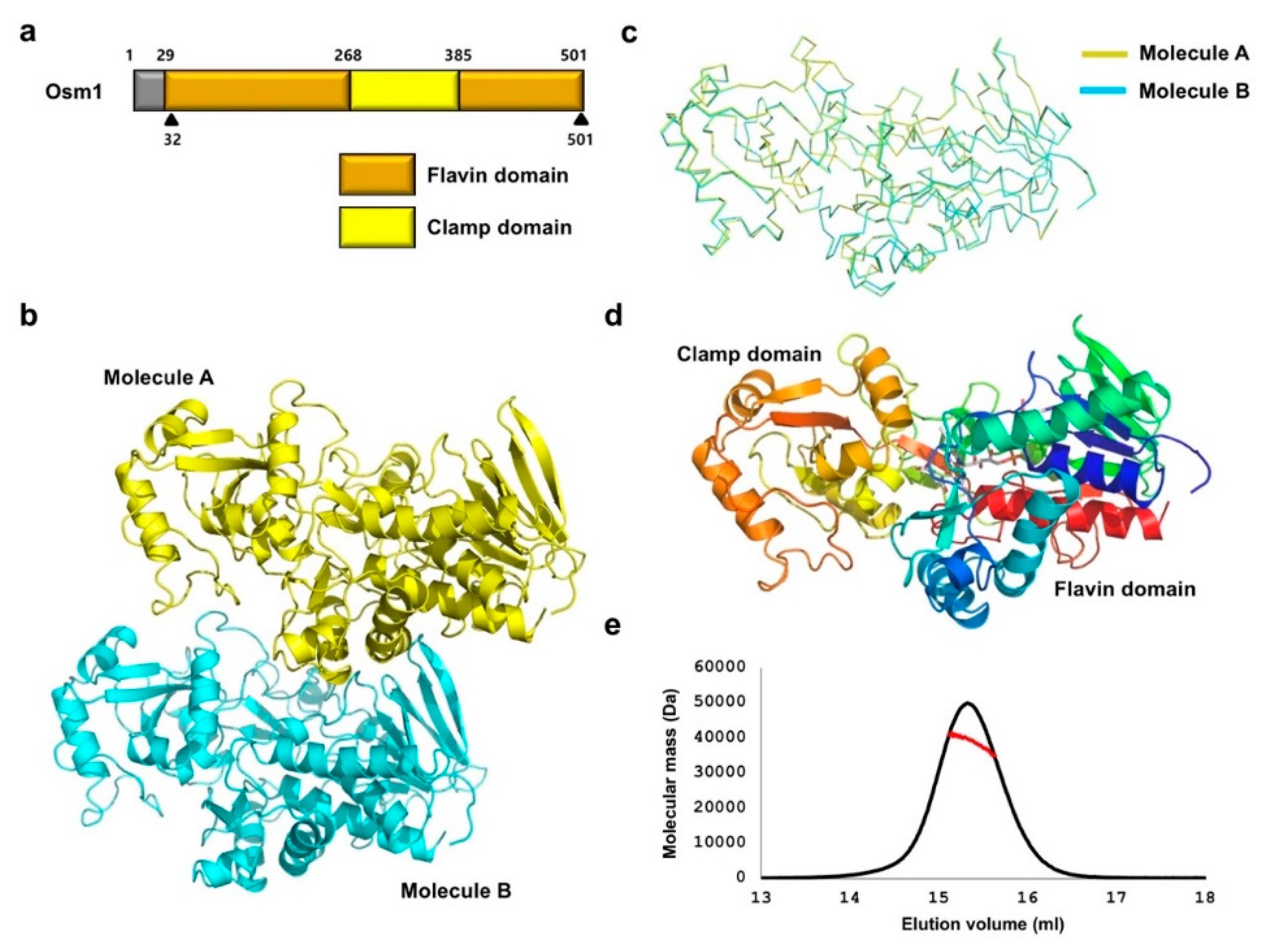
3.1. Overall Structure of the R326A Mutant Form of Osm1
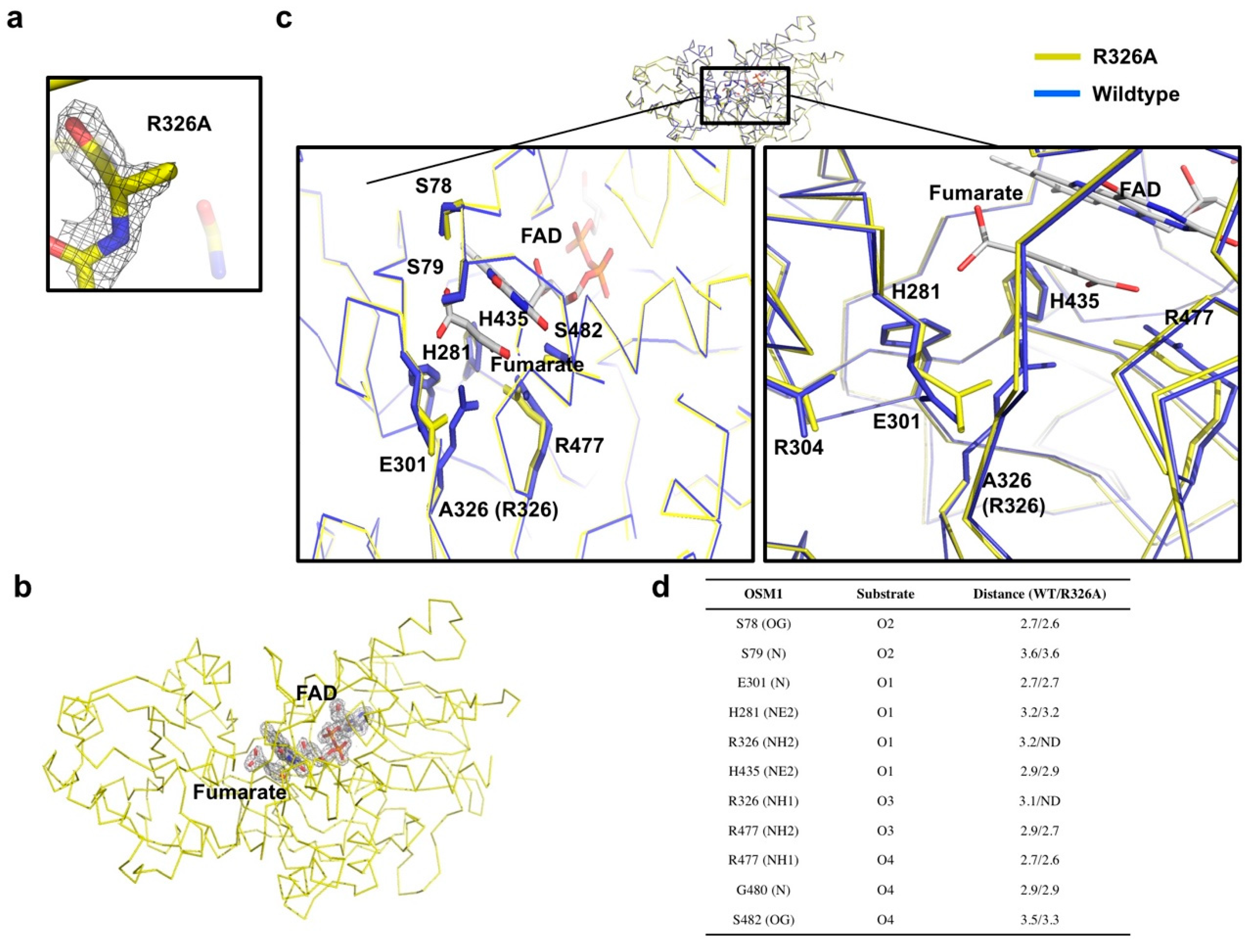
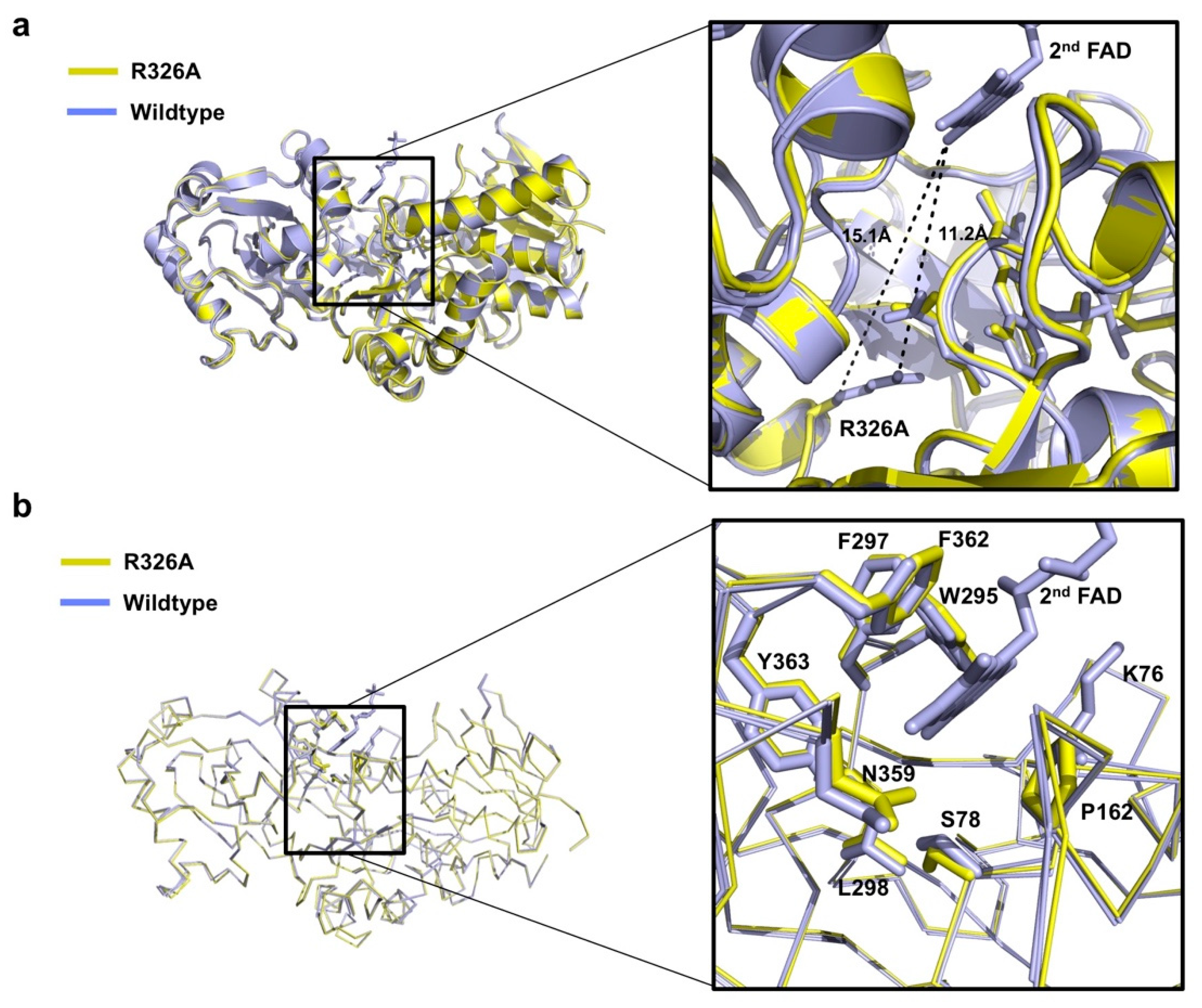
3.2. Comparison of the Structure of Osm1 R326A with the Structure of Wild-Type Osm1
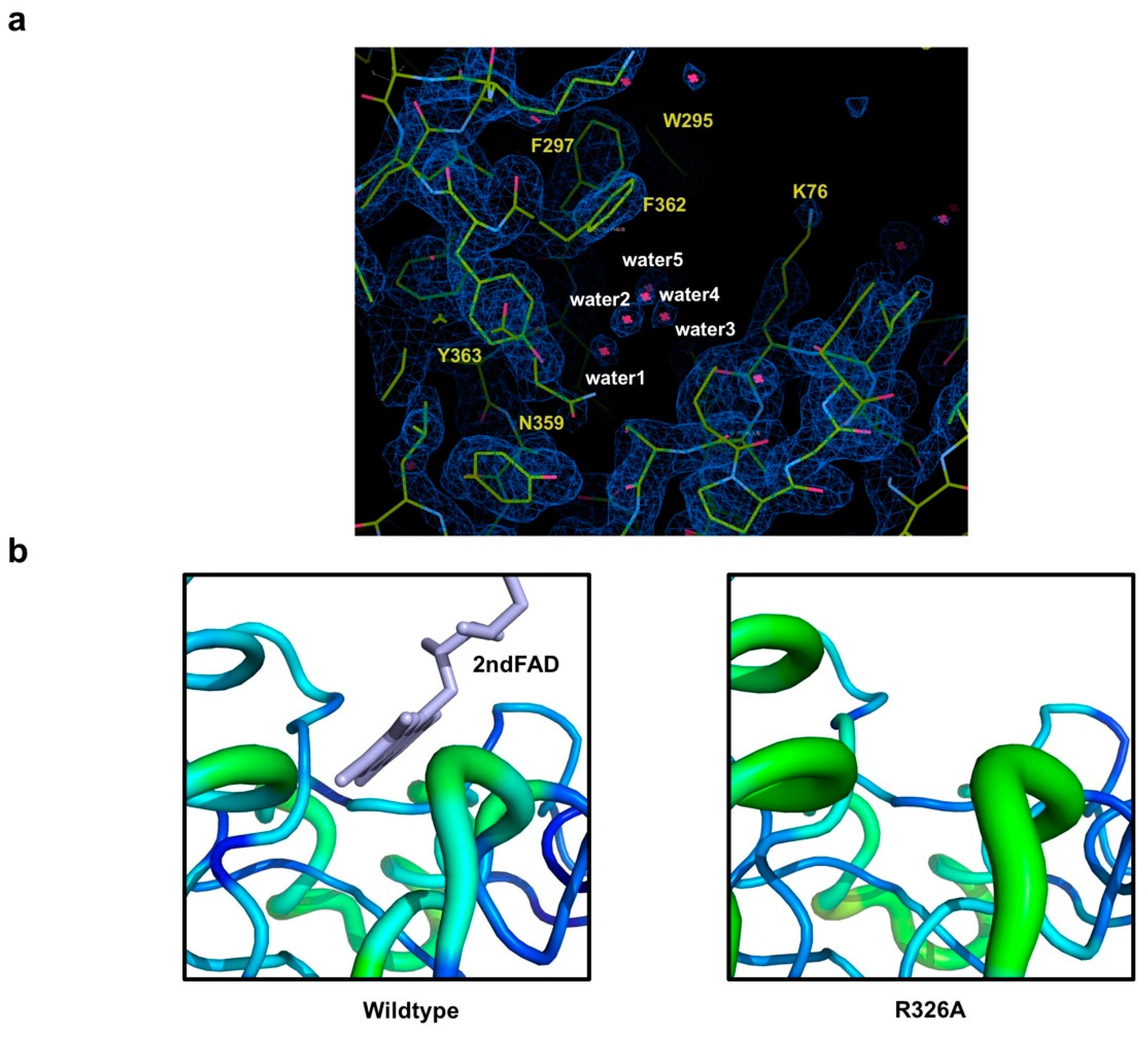
3.3. Undetectable 2nd FAD on Osm1 R326A at the Previously Defined 2nd FAD Binding Site
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Iverson, T.M.; Luna-Chavez, C.; Cecchini, G.; Rees, D.C. Structure of the escherichia coli fumarate reductase respiratory complex. Science 1999, 284, 1961–1966. [Google Scholar] [CrossRef] [PubMed]
- Reid, G.A.; Miles, C.S.; Moysey, R.K.; Pankhurst, K.L.; Chapman, S.K. Catalysis in fumarate reductase. Biochim. Biophys. Acta 2000, 1459, 310–315. [Google Scholar] [CrossRef]
- Tielens, A.G.; Van Hellemond, J.J. The electron transport chain in anaerobically functioning eukaryotes. Biochim. Biophys. Acta 1998, 1365, 71–78. [Google Scholar] [CrossRef]
- Lancaster, C.R.; Kroger, A.; Auer, M.; Michel, H. Structure of fumarate reductase from Wolinella succinogenes at 2.2 Å resolution. Nature 1999, 402, 377–385. [Google Scholar] [CrossRef] [PubMed]
- Leys, D.; Tsapin, A.S.; Nealson, K.H.; Meyer, T.E.; Cusanovich, M.A.; Van Beeumen, J.J. Structure and mechanism of the flavocytochrome c fumarate reductase of Shewanella putrefaciens MR-1. Nat. Struct. Biol. 1999, 6, 1113–1117. [Google Scholar]
- Arikawa, Y.; Enomoto, K.; Muratsubaki, H.; Okazaki, M. Soluble fumarate reductase isoenzymes from Saccharomyces cerevisiae are required for anaerobic growth. FEMS Microbiol. Lett. 1998, 165, 111–116. [Google Scholar] [CrossRef][Green Version]
- Enomoto, K.; Ohki, R.; Muratsubaki, H. Cloning and sequencing of the gene encoding the soluble fumarate reductase from Saccharomyces cerevisiae. DNA Res. 1996, 3, 263–267. [Google Scholar] [CrossRef]
- Muratsubaki, H.; Enomoto, K. One of the fumarate reductase isoenzymes from Saccharomyces cerevisiae is encoded by the OSM1 gene. Arch. Biochem. Biophys. 1998, 352, 175–181. [Google Scholar] [CrossRef]
- Camarasa, C.; Faucet, V.; Dequin, S. Role in anaerobiosis of the isoenzymes for saccharomyces cerevisiae fumarate reductase encoded by OSM1 and FRDS1. Yeast 2007, 24, 391–401. [Google Scholar] [CrossRef] [PubMed]
- Doherty, M.K.; Pealing, S.L.; Miles, C.S.; Moysey, R.; Taylor, P.; Walkinshaw, M.D.; Reid, G.A.; Chapman, S.K. Identification of the active site acid/base catalyst in a bacterial fumarate reductase: A kinetic and crystallographic study. Biochemistry 2000, 39, 10695–10701. [Google Scholar] [CrossRef]
- Taylor, P.; Pealing, S.L.; Reid, G.A.; Chapman, S.K.; Walkinshaw, M.D. Structural and mechanistic mapping of a unique fumarate reductase. Nat. Struct. Biol. 1999, 6, 1108–1112. [Google Scholar] [PubMed]
- Kim, S.; Kim, C.M.; Son, Y.J.; Choi, J.Y.; Siegenthaler, R.K.; Lee, Y.; Jang, T.H.; Song, J.; Kang, H.; Kaiser, C.A.; et al. Molecular basis of maintaining an oxidizing environment under anaerobiosis by soluble fumarate reductase. Nat. Commun. 2018, 9, 4867. [Google Scholar] [CrossRef] [PubMed]
- Williams, C.C.; Jan, C.H.; Weissman, J.S. Targeting and plasticity of mitochondrial proteins revealed by proximity-specific ribosome profiling. Science 2014, 346, 748–751. [Google Scholar] [CrossRef] [PubMed]
- McCoy, A.J. Solving structures of protein complexes by molecular replacement with phaser. Acta Cryst. D Biol. Cryst. 2007, 63, 32–41. [Google Scholar] [CrossRef] [PubMed]
- Emsley, P.; Cowtan, K. Coot: Model-Building tools for molecular graphics. Acta Cryst. D Biol. Cryst. 2004, 60, 2126–2132. [Google Scholar] [CrossRef] [PubMed]
- Vagin, A.A.; Steiner, R.A.; Lebedev, A.A.; Potterton, L.; McNicholas, S.; Long, F.; Murshudov, G.N. REFMAC5 dictionary: Organization of prior chemical knowledge and guidelines for its use. Acta Cryst. D Biol. Cryst. 2004, 60, 2184–2195. [Google Scholar] [CrossRef] [PubMed]
- Chen, V.B.; Arendall, W.B., 3rd; Headd, J.J.; Keedy, D.A.; Immormino, R.M.; Kapral, G.J.; Murray, L.W.; Richardson, J.S.; Richardson, D.C. MolProbity: All-Atom structure validation for macromolecular crystallography. Acta Cryst. D Biol. Cryst. 2010, 66, 12–21. [Google Scholar] [CrossRef]
- Delano, W.L. The PyMol Molecular Graphics System. 2002. Available online: http://pymol.sourceforge.net/overview/index.htm (accessed on 27 September 2019).
- Bamford, V.; Dobbin, P.S.; Richardson, D.J.; Hemmings, A.M. Open conformation of a flavocytochrome c3 fumarate reductase. Nat. Struct. Biol. 1999, 6, 1104–1107. [Google Scholar]
- Edwards, C.B.; Copes, N.; Brito, A.G.; Canfield, J.; Bradshaw, P.C. Malate and fumarate extend lifespan in caenorhabditis elegans. PLoS ONE 2013, 8, e58345. [Google Scholar] [CrossRef]
- Butler, J.A.; Ventura, N.; Johnson, T.E.; Rea, S.L. Long-Lived mitochondrial (Mit) mutants of caenorhabditis elegans utilize a novel metabolism. FASEB J. 2010, 24, 4977–4988. [Google Scholar] [CrossRef]
- Rea, S.; Johnson, T.E. A metabolic model for life span determination in Caenorhabditis elegans. Dev. Cell 2003, 5, 197–203. [Google Scholar] [CrossRef]
- Enomoto, K.; Arikawa, Y.; Muratsubaki, H. Physiological role of soluble fumarate reductase in redox balancing during anaerobiosis in Saccharomyces cerevisiae. FEMS Microbiol. Lett. 2002, 215, 103–108. [Google Scholar] [CrossRef] [PubMed][Green Version]
| Data Collection | Native |
|---|---|
| X-ray source | Synchrotron (PAL 5C) |
| Detector | Eiger 9M |
| Wavelength | 0.97950 |
| Space group | P2 |
| Cell dimensions | |
| a, b, c | 51.9 Å, 109.9 Å, 77.1 Å |
| β | 98.5° |
| Resolution | 50–2.0 Å |
| Wilson B-factor | 20.8 Å2 |
| † No. of unique reflections overall | 5,6568 (2,799) |
| † Rsym | 15.6% (49.1%) |
| † I/I | 15.1 (2.9) |
| † Completeness | 99.9% (100 %) |
| † Redundancy | 6.5 (6.0) |
| Refinement | |
| Resolution | 38–2.0 Å |
| No. of reflections used (completeness) | 5,6437 (98.8%) |
| No. of non-H protein atoms | 7943 |
| No. of water molecules | 599 |
| No. of ions | 0 |
| † Rwork | 20.0% (20.0%) |
| † Rfree | 25.1% (25.0%) |
| Average B-factors | |
| Protein | 22.9 Å2 |
| Water and other small molecules | 34.4 Å2 |
| r.m.s. deviations | |
| B-factor for bonded atoms | 3.17 |
| Bond lengths | 0.007 Å |
| Bond angles | 0.973° |
| MolProbity analysis | |
| Ramachandran outliers | 0.00% |
| Ramachandran favored | 97.12% |
| Ramachandran allowed | 2.88% |
| Rotamer outliers | 1.00% |
| Clash score | 5.00 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kim, C.M.; Kwon, S.; Jung, K.H.; Chun, H.L.; Ha, H.J.; Park, H.H. Crystal Structure of the Active Site Mutant Form of Soluble Fumarate Reductase, Osm1. Crystals 2019, 9, 504. https://doi.org/10.3390/cryst9100504
Kim CM, Kwon S, Jung KH, Chun HL, Ha HJ, Park HH. Crystal Structure of the Active Site Mutant Form of Soluble Fumarate Reductase, Osm1. Crystals. 2019; 9(10):504. https://doi.org/10.3390/cryst9100504
Chicago/Turabian StyleKim, Chang Min, Sunghark Kwon, Kyung Ho Jung, Hye Lin Chun, Hyun Ji Ha, and Hyun Ho Park. 2019. "Crystal Structure of the Active Site Mutant Form of Soluble Fumarate Reductase, Osm1" Crystals 9, no. 10: 504. https://doi.org/10.3390/cryst9100504
APA StyleKim, C. M., Kwon, S., Jung, K. H., Chun, H. L., Ha, H. J., & Park, H. H. (2019). Crystal Structure of the Active Site Mutant Form of Soluble Fumarate Reductase, Osm1. Crystals, 9(10), 504. https://doi.org/10.3390/cryst9100504






