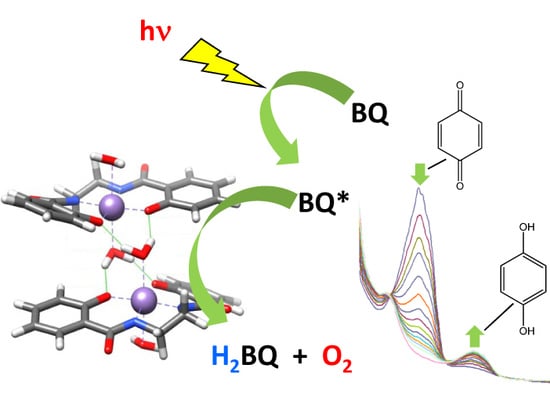
Modeling the OEC with Two New Biomimetic Models: Preparations, Structural Characterization, and Water Photolysis Studies of a Ba–Mn Box Type Complex and a Mn4N6 Planar-Diamond Cluster
Abstract
1. Introduction
2. Results
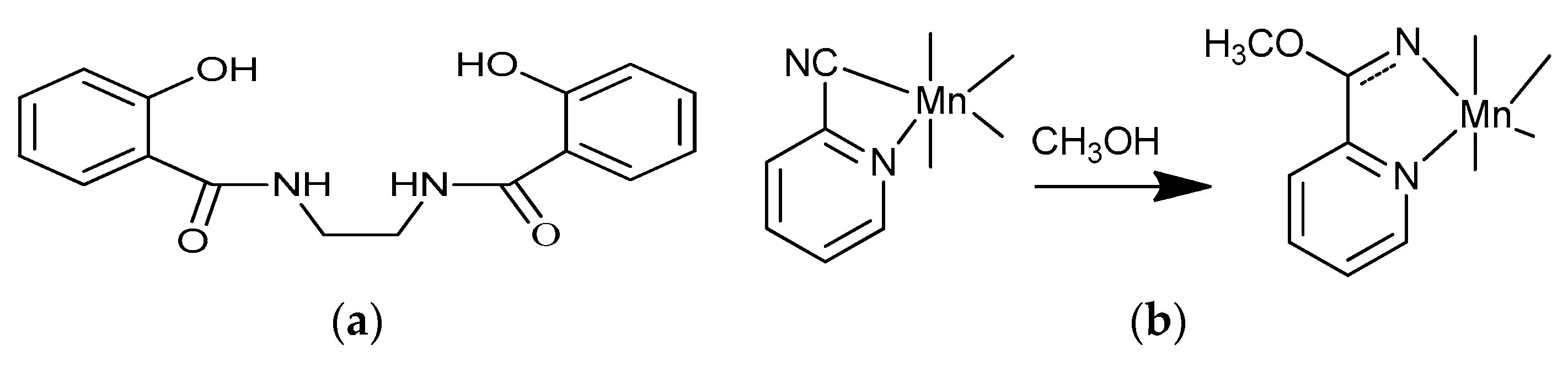
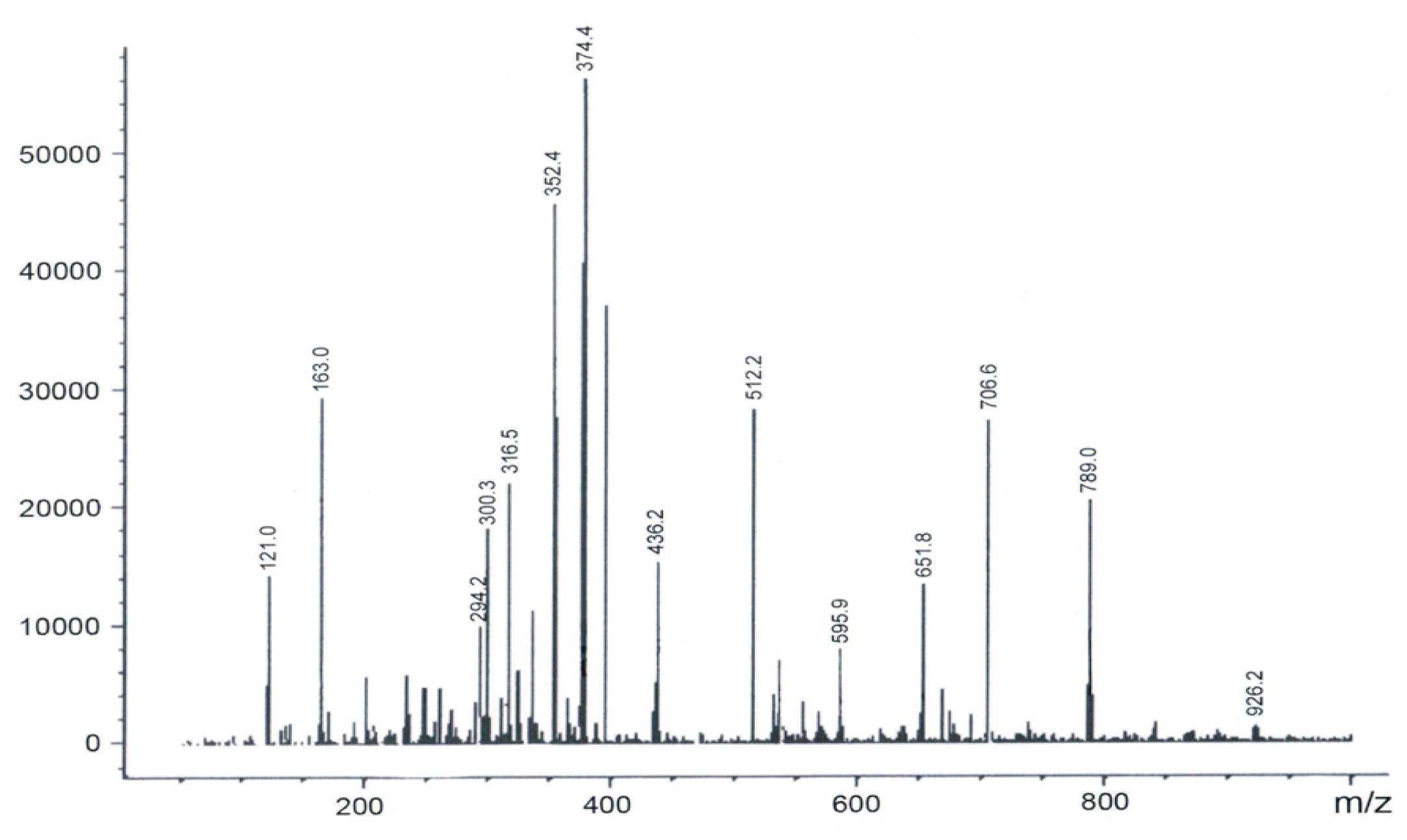
2.1. Preparation and Characterization of Biomimetic Model 1
2.2. Preparation and Characterization of Biomimetic Model 2
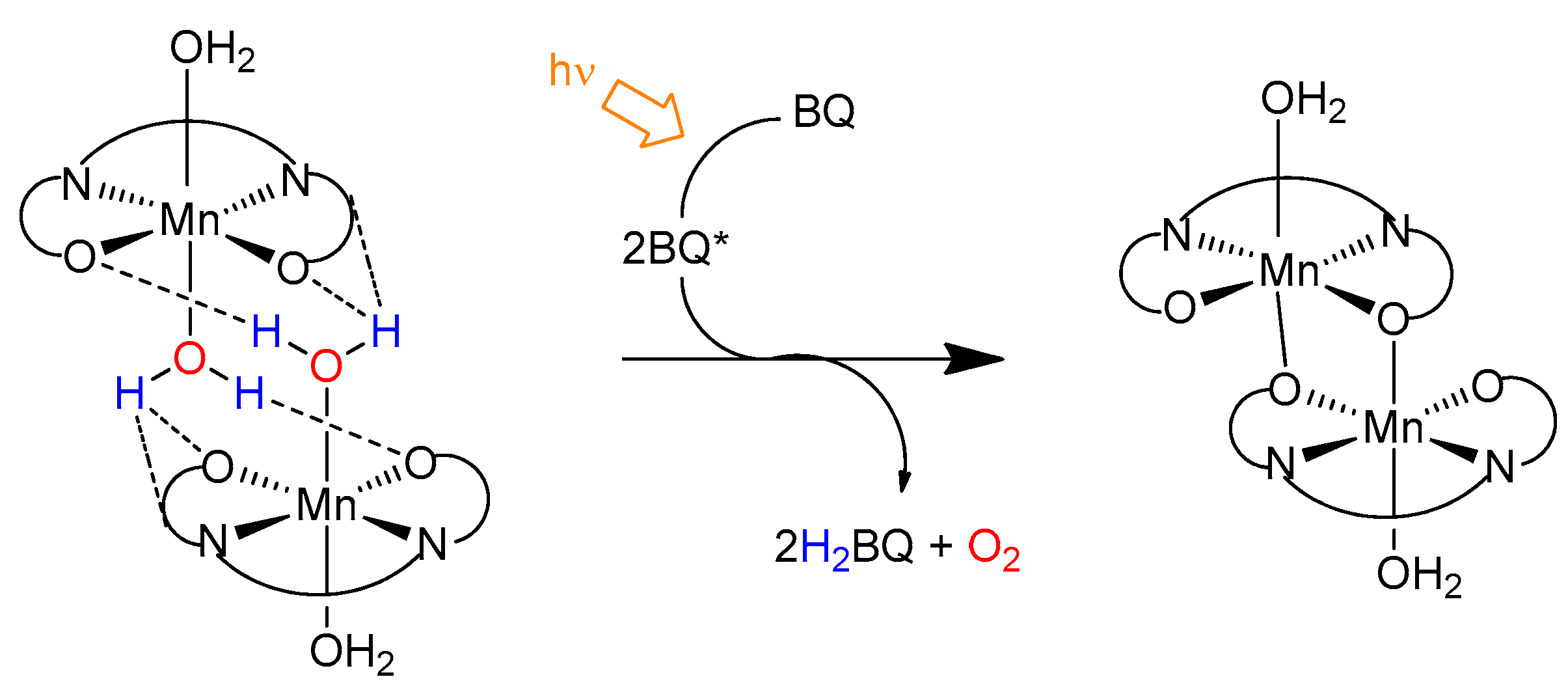
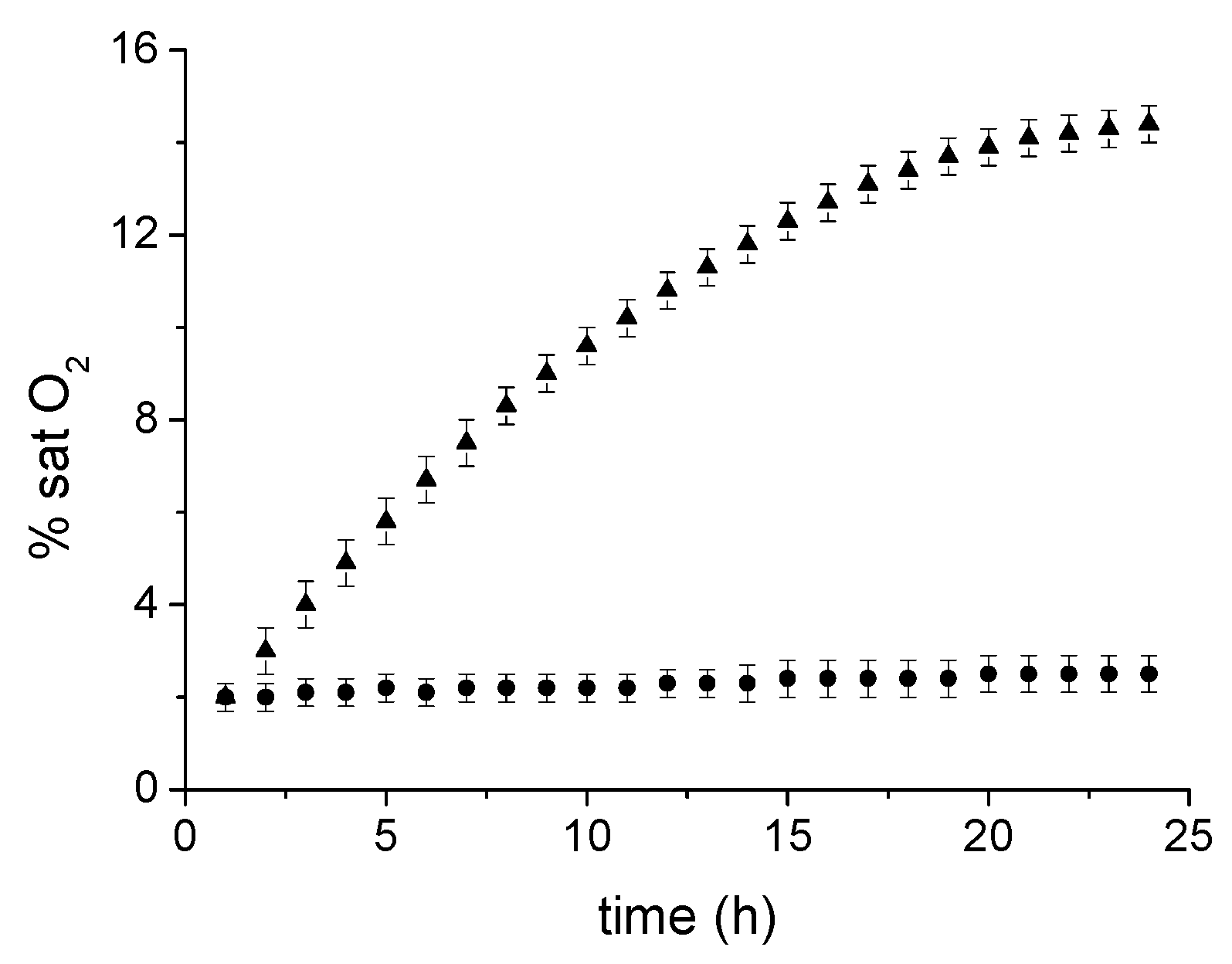
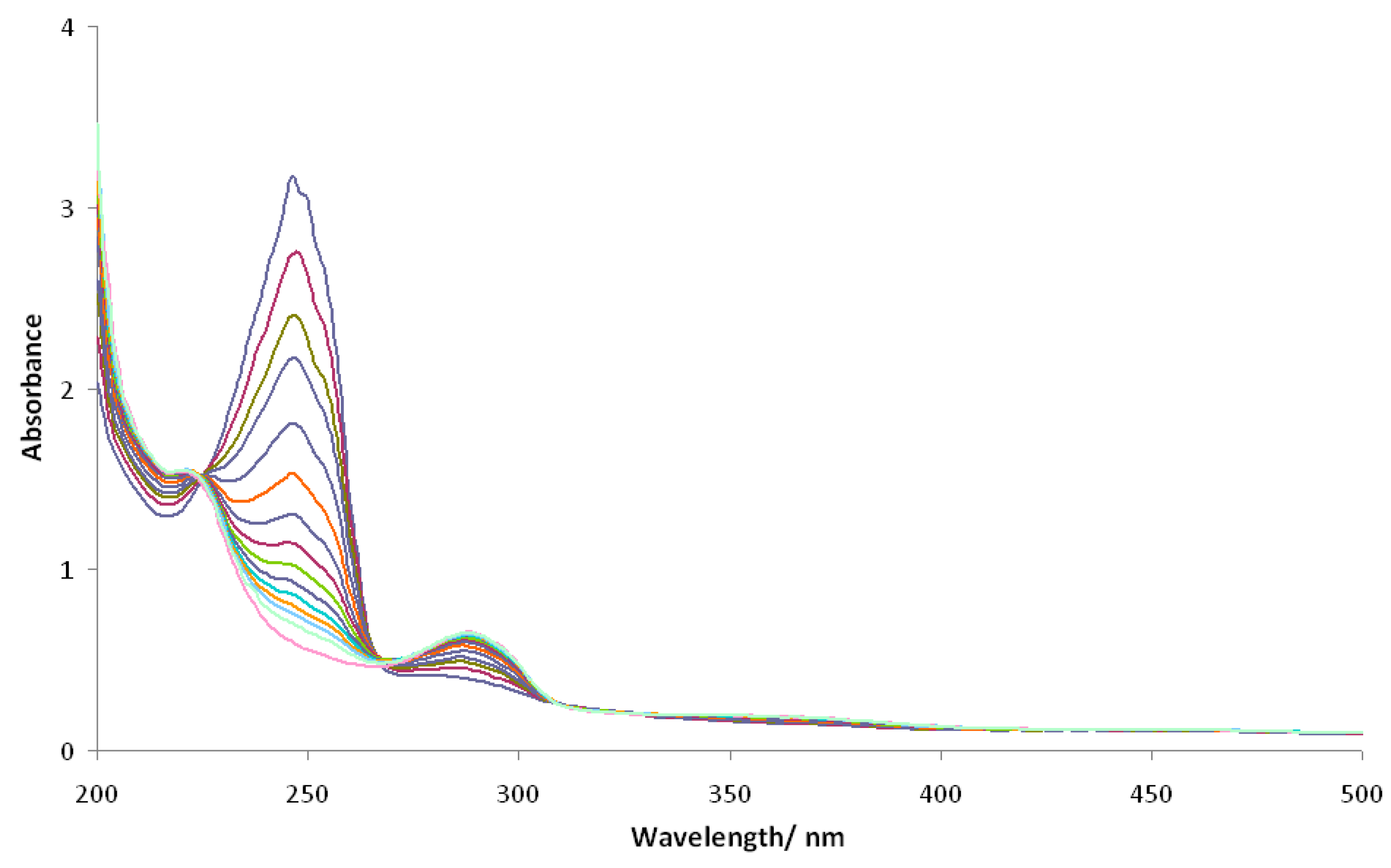
2.3. Photolytic Studies
3. Discussion
4. Materials and Methods
4.1. Chemical and Reagents
4.2. Physical Measurements
4.3. Synthesis of the Complexes
4.4. X-ray Crystallographic Studies
4.5. Photolysis Experiments
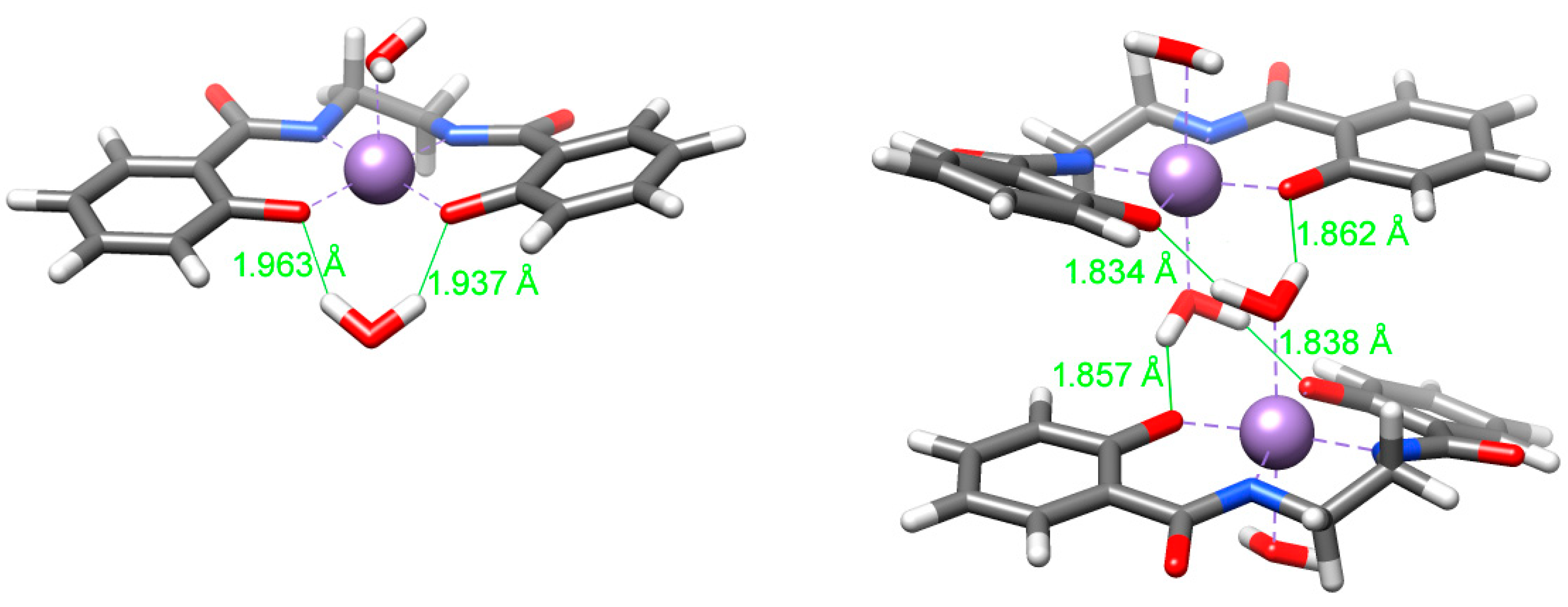
4.6. DFT Calculations
5. Conclusions
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
Nomenclature
| Acronyms | |
| BQ | benzoquinone |
| DFT | density functional theory |
| DMF | dimethylformamide |
| DMSO | dimethyl sulfoxide |
| ESI-MS | electrospray ionization mass spectroscopy |
| NAPDH | nicotinamide adenine dinucleotide phosphate |
| NMR | nuclear magnetic resonance |
| ONNO | oxygen-nitrogen-nitrogen-oxygen |
| ORTEP | Oak Ridge thermal ellipsoid plot |
| SCF | self consistent field |
| TPSSh/TZVP | hybrid functional using the Tao-Perdew-Staroverov-Scuseria functional–valence triple zeta polarization basis set |
References
- Umena, Y.; Kawakami, K.; Shen, J.R.; Kamiya, N. Crystal structure of oxygen-evolving photosystem II at a resolution of 1.9 Å. Nature 2011, 473, 55–60. [Google Scholar] [CrossRef] [PubMed]
- Yano, J.; Yachandra, V. Mn4Ca cluster in photosynthesis: Where and how water is oxidized to dioxygen. Chem. Rev. 2014, 114, 4175–4205. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Li, Y.; Zhao, G.; Yao, R.; Zhang, C. Natural and artificial Mn4Ca cluster for the water splitting reaction. ChemSusChem 2017, 10, 4403–4408. [Google Scholar] [CrossRef] [PubMed]
- McEvoy, J.P.; Brudvig, G.W. Water-splitting chemistry of photosystem II. Chem. Rev. 2006, 106, 4455–4483. [Google Scholar] [CrossRef] [PubMed]
- Cox, N.; Pantazis, D.A.; Neese, F.; Lubitz, W. Biological water oxidation. Acc. Chem. Res. 2013, 46, 1588–1596. [Google Scholar] [CrossRef] [PubMed]
- Suga, M.; Akita, F.; Hirata, K.; Ueno, G.; Murakami, H.; Nakajima, Y.; Shimizu, T.; Yamashita, K.; Yamamoto, M.; Ago, H.; et al. Native structure of photosystem II at 1.95 Å resolution viewed by femtosecond X-ray pulses. Nature 2015, 517, 99–103. [Google Scholar] [CrossRef] [PubMed]
- Blakemore, J.D.; Crabtree, R.H.; Brudvig, G.W. Molecular catalysts for water oxidation. Chem. Rev. 2015, 115, 12974–13005. [Google Scholar] [CrossRef] [PubMed]
- Herrero, C.; Nguyen-Thi, N.; Hammerer, F.; Banse, F.; Gagné, D.; Doucet, N.; Mahy, J.P.; Ricoux, R. Photoassisted oxidation of sulfides catalyzed by artificial metalloenzymes using water as an oxygen source. Catalysts 2016, 6, 202. [Google Scholar] [CrossRef]
- Garrido-Barros, P.; Gimbert-Suriñach, C.; Matheu, R.; Sala, X.; Llobet, A. How to make an efficient and robust molecular catalyst for water oxidation. Chem. Soc. Rev. 2017, 46, 6088–6098. [Google Scholar] [CrossRef] [PubMed]
- Paul, S.; Neese, F.; Pantazis, D.A. Structural models of the biological oxygen-evolving complex: Achievements, insights, and challenges for biomimicry. Green Chem. 2017, 19, 2309–2325. [Google Scholar] [CrossRef]
- Dismukes, G.C.; Siderer, Y. Intermediates of a polynuclear manganese center involved in photosynthetic oxidation of water. Proc. Natl. Acad. Sci. USA 1981, 78, 274–278. [Google Scholar] [CrossRef] [PubMed]
- Campbell, K.A.; Force, D.A.; Nixon, P.J.; Dole, F.; Diner, B.A.; Britt, R.D. Does aspartate 170 of the D1 polypeptide ligate the manganese cluster in photosystem II? An EPR and ESEEM study. J. Am. Chem. Soc. 2003, 42, 10600–10608. [Google Scholar] [CrossRef]
- Satadal, P.; Cox, N.; Pantazis, D.A. What can we learn from a biomimetic model of nature’s oxygen-evolving complex? Inorg. Chem. 2017, 56, 3875–3888. [Google Scholar] [CrossRef]
- Rompel, A.; Andrews, J.C.; Cinco, R.M.; Wemple, M.W.; Christou, G.; Law, N.A.; Pecoraro, V.L.; Sauer, K.; Yachandra, V.K.; Klein, M.P. Chlorine K-edge X-ray absorption spectroscopy as a probe of chlorine-manganese bonding: Model systems with relevance to the oxygen evolving complex in photosystem II. J. Am. Chem. Soc. 1997, 119, 4465–4470. [Google Scholar] [CrossRef]
- Grundmeier, A.; Dau, H. Structural models of the manganese complex of photosystem II and mechanistic implications. Biochim. Biophys. Acta-Bioenerg. 2012, 1817, 88–105. [Google Scholar] [CrossRef] [PubMed]
- Askerka, M.; Brudvig, G.W.; Batista, V.S. The O2-evolving complex of photosystem II: Recent insights from Quantum Mechanics/Molecular Mechanics (QM/MM), Extended X-ray Absorption Fine Structure (EXAFS), and Femtosecond X-ray Crystallography data. Acc. Chem. Res. 2017, 50, 41–48. [Google Scholar] [CrossRef] [PubMed]
- Mullins, C.S.; Pecoraro, V.L. Reflections on small molecule manganese models that seek to mimic photosynthetic water oxidation chemistry. Coord. Chem. Rev. 2008, 252, 416–443. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, K.; Shoji, M.; Isobe, H.; Yamanaka, S.; Umena, Y.; Kawakami, K.; Kamiya, N. On the guiding principles for understanding of geometrical structures of the CaMn4O5 cluster in oxygen-evolving complex of photosystem II. Proposal of estimation formula of structural deformations via the Jahn-Teller effects. Mol. Phys. 2017, 115, 636–666. [Google Scholar] [CrossRef]
- Maneiro, M.; Ruettinger, W.F.; Bourles, E.; McLendon, G.L.; Dismukes, G.C. Kinetics of proton-coupled electron-transfer reactions to the manganese-oxo “cubane” complexes containing the Mn4O and Mn4O core types. Proc. Natl. Acad. Sci. USA 2003, 100, 3707–3712. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Concepcion, J.J.; Jurss, J.W.; Cardolaccia, T.; Templeton, J.L.; Meyer, T.J. Mechanisms of water oxidation from the blue dimer to photosystem II. Inorg. Chem. 2008, 47, 1727–1752. [Google Scholar] [CrossRef] [PubMed]
- Young, K.J.; Brennan, B.J.; Tagore, R.; Brudvig, G.W. Photosynthetic Water Oxidation: Insights from Manganese Model Chemistry. Acc. Chem. Res. 2015, 48, 567–574. [Google Scholar] [CrossRef] [PubMed]
- González-Riopedre, G.; Fernández-García, M.I.; González-Noya, A.M.; Vázquez-Fernández, M.A.; Bermejo, M.R.; Maneiro, M. Manganese-Schiff base complexes as catalysts for water photolysis. Phys. Chem. Chem. Phys. 2011, 13, 18069–18077. [Google Scholar] [CrossRef] [PubMed]
- González-Riopedre, G.; Bermejo, M.R.; Fernández-García, M.I.; González-Noya, A.M.; Pedrido, R.; Rodríguez-Doutón, M.J.; Maneiro, M. Alkali-metal-ion-directed self-assembly of redox-active manganese(III) supramolecular boxes. Inorg. Chem. 2015, 54, 2512–2521. [Google Scholar] [CrossRef] [PubMed]
- Bermejo, M.R.; Fernández, M.I.; González-Noya, A.M.; Maneiro, M.; Pedrido, R.; Rodríguez, M.J.; García-Monteagudo, J.C.; Donnadieu, B. Novel peroxidase mimics: μ-Aqua manganese–Schiff base dimers. J. Inorg. Biochem. 2006, 100, 1470–1478. [Google Scholar] [CrossRef] [PubMed]
- González-Riopedre, G.; Fernández-García, M.I.; Gómez-Fórneas, E.; Maneiro, M. Biomimetic catalysts for oxidation of veratryl alcohol, a lignin model compound. Catalysts 2013, 3, 232–246. [Google Scholar] [CrossRef]
- Geary, W.J.; William, J. The use of conductivity measurements in organic solvents for the characterisation of coordination compounds. Coord. Chem. Rev. 1971, 7, 81–122. [Google Scholar] [CrossRef]
- Daier, V.; Moreno, D.; Duhayon, C.; Tuchagues, J.; Signorella, S. Synthesis, characterization and combined superoxide dismutase and catalase activities of manganese complexes of 1,4-bis(salicylidenamino)butan-2-ol. Eur. J. Inorg. Chem. 2010, 965–974. [Google Scholar] [CrossRef]
- Collomb, M.N.; Mantel, C.; Romain, S.; Duboc, C.; Leprêtre, J.C.; Pécaut, J.; Deronzier, A. Redox-Induced μ-Acetato and μ-Oxo Core Interconversions in Dinuclear Manganese Tris (2-methylpyridyl) amine (tpa) Complexes: Isolation and Characterization of [Mn2III (μ-O)(μ-O2CCH3)(tpa) 2]3+. Eur. J. Inorg. Chem. 2007, 3179–3187. [Google Scholar] [CrossRef]
- Bonadies, J.A.; Maroney, M.L.; Pecoraro, V.L. Structurally diverse manganese (III) Schiff base complexes: Solution speciation via paramagnetic 1H NMR spectroscopy and electrochemistry. Inorg. Chem. 1989, 28, 2044–2051. [Google Scholar] [CrossRef]
- Zuniga, M.F.; Deacon, G.B.; Ruhlandt-Senge, K. Developments in heterobimetallic s-block systems: Synthesis and structural survey of molecular M/Ae (M = Li, Na, K, Cs; AE = Ca, Sr) aryloxo complexes. Inorg. Chem. 2008, 47, 4669–4681. [Google Scholar] [CrossRef] [PubMed]
- Costes, J.P.; Laurent, J.P.; Chabert, P.; Commenges, G.; Dahan, F. Solid state and solution study of trinuclear (Ni, Ba, Ni) complexes: (L12Ni)2Ba(ClO4)2·2H2O (1) and (L22Ni)2Ba(ClO4)2·2H2O (2) (L1 = 3-Methoxysalicylaldiminato and L2 = 3-(2-Methoxyethoxy)salicylaldiminato). Crystal and Molecular Structure of 2. Inorg. Chem. 1997, 36, 656–660. [Google Scholar] [CrossRef]
- Jamnicky, M.; Segla, P.; Koman, M. Methanolysis of pyridine-2-carbonitrile in the coordination sphere of copper(II), cobalt(II) and nickel(II). The structure of [Ni(O-methylpyridine-2-carboximidate)3]Br2·4H2O. Polyhedron 1995, 14, 1837–1847. [Google Scholar] [CrossRef]
- Garduno, J.A.; García, J.J. Synthesis of annidines and benzoxazoles from activated nitriles with Ni(0) catalysts. ACS Catal. 2015, 5, 3470–3477. [Google Scholar] [CrossRef]
- Amami, V.; Ahmadi, R.; Naseh, M.; Ebadi, A. Synthesis, spectroscopic characterization, crystal structure and thermal analyses of two zinc(II) complexes with methanolysis of 2-pyridinecarbonitrile as a chelating ligand. J. Iran. Chem. Soc. 2017, 14, 635–642. [Google Scholar] [CrossRef]
- Devi, S.P.; Devi, R.B.; Devi, N.S.; Singh, L.J.; Singh, R.H. Structural and spectroscopic investigations on bis(1-amidino-O-2-alkoxyethylurea) copper (II) perchlorate complexes (alkoxy = methoxy, ethoxy or butoxy). Polyhedron 2012, 47, 1–8. [Google Scholar] [CrossRef]
- Amani, V.; Safari, N.; Khavasi, H.R. Synthesis, characterization and crystal structure determination of iron (III) hetero-ligand complexes containing 2,2′-bipyridine, 5,5′-dimethyl-2,2′-bipyridine and chloride,[Fe(bipy)Cl4][bipy·H] and [Fe(dmbipy)2Cl2][FeCl4]. Polyhedron 2007, 26, 4257–4262. [Google Scholar] [CrossRef]
- Paulat, F.; Praneeth, V.K.K.; Näther, C.; Lehnert, N. Quantum chemistry-based analysis of the vibrational spectra of five-coordinate metalloporphyrins [M(TPP)Cl]. Inorg. Chem. 2006, 45, 2835–2856. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.I.; Lee, G.H.; Wur, C.S.; Lin, J.G.; Tsai, H.L. Syntheses, structures and single-molecule magnetic behaviors of two dicubane Mn4 complexes. Polyhedron 2005, 24, 2215–2221. [Google Scholar] [CrossRef]
- Langley, S.K.; Chilton, N.F.; Massi, M.; Moubaraki, B.; Berry, K.J.; Murray, K.S. Synthesis and characterization of homo- and heterovalent tetra- hexa- hepta- and decanuclear manganese clusters using pyridyl functionalized β-diketone, carboxylate and triethanolamine ligands. Dalton Trans. 2010, 39, 7236–7249. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Ma, C.B.; Yuan, D.Q.; Wang, H.S.; Chen, C.N.; Liu, Q.T. Synthesis and characterization of a family of penta- and tetra-manganese(III) complexes derived from an assembly system containing tert-butylphosphonic acid. Inorg. Chem. 2008, 47, 5580–5590. [Google Scholar] [CrossRef] [PubMed]
- Wemple, M.W.; Tsai, H.-L.; Wang, S.; Claude, J.P.; Streib, W.E.; Huffman, J.C.; Hendrickson, D.N.; Christou, G. Tetranuclear and octanuclear manganese carboxylate clusters: Preparation and reactivity of (NBun4)[Mn4O2(O2CPh)9(H2O)] and synthesis of (NBun4)2[Mn8O4(O2CPh)12(Et2mal)2(H2O)2] with a “linked-butterfly” structure. Inorg. Chem. 1996, 35, 6437–6449. [Google Scholar] [CrossRef] [PubMed]
- Ruettinger, W.; Yagi, M.; Wolf, K.; Bernasek, S.; Dismukes, G.C. O2 Evolution from the manganese–oxo cubane core Mn4O46+: A molecular mimic of the photosynthetic water oxidation enzyme? J. Am. Chem. Soc. 2000, 122, 10353–10357. [Google Scholar] [CrossRef]
- Hendrickson, D.N.; Christou, G.; Schmitt, E.A.; Libby, E.; Bashkin, J.S.; Wang, S.; Tsai, H.L.; Vincent, J.B.; Boyd, P.D.W. Photosynthetic water oxidation center: Spin frustration in distorted cubane MnIVMnIII3 model complexes. J. Am. Chem. Soc. 1992, 114, 2455–2471. [Google Scholar] [CrossRef]
- Pedrido, R.; Bermejo, M.R.; García-Deibe, A.M.; González-Noya, A.M.; Maneiro, M.; Vázquez, M. Metal complexes of a novel achiral symmetric pentadentate ligand–Crystal structures of monohelical zinc(II) and cadmium(II) complexes. Eur. J. Inorg. Chem. 2003, 3193–3200. [Google Scholar] [CrossRef]
- Addison, A.; Nageswara, R.T.; Reedijk, J.; Van Rijn, J.; Verschoor, G.C. Synthesis, structure, and spectroscopic properties of copper(II) compounds containing nitrogen–sulphur donor ligands; the crystal and molecular structure of aqua[1,7-bis(N-methylbenzimidazol-2′-yl)-2,6-dithiaheptane]copper(II) perchlorate. J. Chem. Soc. Dalton Trans. 1984, 1349–1356. [Google Scholar] [CrossRef]
- Awad, M.K.; Anderson, A.B. Photoactivation of H2O by p-benzoquinone and the role of MnIII complexes in O2 evolution: Molecular orbital theory. J. Am. Chem. Soc. 1989, 111, 802–806. [Google Scholar] [CrossRef]
- Ashmawy, F.M.; McAuliffe, C.A.; Parish, R.V.; Tames, J. Water photolysis. Part 1. The photolysis of coordinated water in [{MnL(H2O)}2][ClO4]2 (L = dianion of tetradentate O2N2-donor Schiff bases). A model for the manganese site in photosystem II of green plant photosynthesis. J. Chem. Soc. Dalton Trans. 1985, 1391–1397. [Google Scholar] [CrossRef]
- Aurangzeb, N.; Hulme, C.E.; McAuliffe, C.A.; Pritchard, R.G.; Watkinson, M.; Bermejo, M.R.; Garcia-Deibe, A.; Sanmartin, J.; Sousa, A. Crystallographic characterization of a possible model for photosystem II. J. Chem. Soc. Chem. Commun. 1994, 1153–1155. [Google Scholar] [CrossRef]
- Ononye, A.I.; McIntosh, A.R.; Bolton, J.R. Mechanism of the photochemistry of p-benzoquinone in aqueous solutions. 1. Spin trapping and flash photolysis electron paramagnetic resonance studies. J. Phys. Chem. 1986, 90, 6266–6270. [Google Scholar] [CrossRef]
- Taguchi, T.; Stone, K.L.; Gupta, R.; Kaiser-Lassalle, B.; Yano, J.; Hendrich, M.P.; Borovik, A.S. Preparation and properties of an MnIV-hydroxide complex: Proton and electron transfer at a mononuclear manganese site and its relationship to the Oxygen Evolving Complex within Photosystem II. Chem. Sci. 2014, 5, 3064–3071. [Google Scholar] [CrossRef] [PubMed]
- Gupta, R.; Taguchi, T.; Lassalle-Kaiser, B.; Bominaar, E.L.; Yano, J.; Hendrich, M.P.; Borovik, A.S. High-spin Mn-oxo complexes and their relevance to the oxygen-evolving complex within photosystem II. Proc. Natl. Acad. Sci. USA 2015, 112, 5319–5324. [Google Scholar] [CrossRef] [PubMed]
- Bruker. SAINT, Siemens Area Detector Integration Software; Bruker AXS Inc.: Madison, WI, USA, 2003. [Google Scholar]
- Sheldrick, G.M. SADABS, Program for Scaling and Correction of Area Detector Data; University of Göttingen: Göttingen, Germany, 1996. [Google Scholar]
- Altomare, A.; Burla, M.C.; Camalli, M.; Cascarano, G.L.; Giacovazzo, C.; Guagliardi, A.; Moliterni, A.G.G.; Polidori, G.; Spagna, R. SIR97: A new tool for crystal structure determination and refinement. J. Appl. Crystallogr. 1999, 32, 115–119. [Google Scholar] [CrossRef]
- Sheldrick, G.M. A short history of SHELX. Acta Cryst. 2008, A64, 112–122. [Google Scholar] [CrossRef] [PubMed]
- Farrugia, L.J. ORTEP-3 for Windows—A version of ORTEP-III with a Graphical User Interface (GUI). J. Appl. Cryst. 1997, 30, 565. [Google Scholar] [CrossRef]
- Macrae, C.F.; Bruno, I.J.; Chisholm, J.A.; Edgington, P.R.; McCabe, P.; Pidcock, E.; Rodriguez-Monge, L.; Taylor, R.; van de Streek, J.; Wood, P.A. MERCURY CSD 2.0—New features for the visualization and investigation of crystal structures. J. Appl. Crystallogr. 2008, 41, 466–470. [Google Scholar] [CrossRef]
- Tao, J.M.; Perdew, J.P.; Staroverov, V.N.; Scuseria, G.E. Climbing the density functional ladder: Nonempirical meta-generalizated gradient approximation designed for molecules and solids. Phys. Rev. Lett. 2003, 91, 146401–146405. [Google Scholar] [CrossRef] [PubMed]
- Schaefer, A.; Huber, C.; Ahlrichs, R. Fully optimized contracted Gaussian basis sets of triple zeta valence quality for atoms Li to Kr. J. Chem. Phys. 1994, 100, 5829–5835. [Google Scholar] [CrossRef]
- Frisch, M.J.; Trucks, G.W.; Schlegel, H.B.; Scuseria, G.E.; Robb, M.A.; Cheeseman, J.R.; Scalmani, G.; Barone, V.; Mennucci, B.; Petersson, G.A.; et al. Gaussian 09, Revision, A.01; Gaussian, Inc.: Wallingford, CT, USA, 2009. [Google Scholar]
- Tomasi, J.; Mennucci, B.; Cammi, R. Quantum mechanical continuum solvation models. Chem. Rev. 2005, 105, 2999–3094. [Google Scholar] [CrossRef] [PubMed]
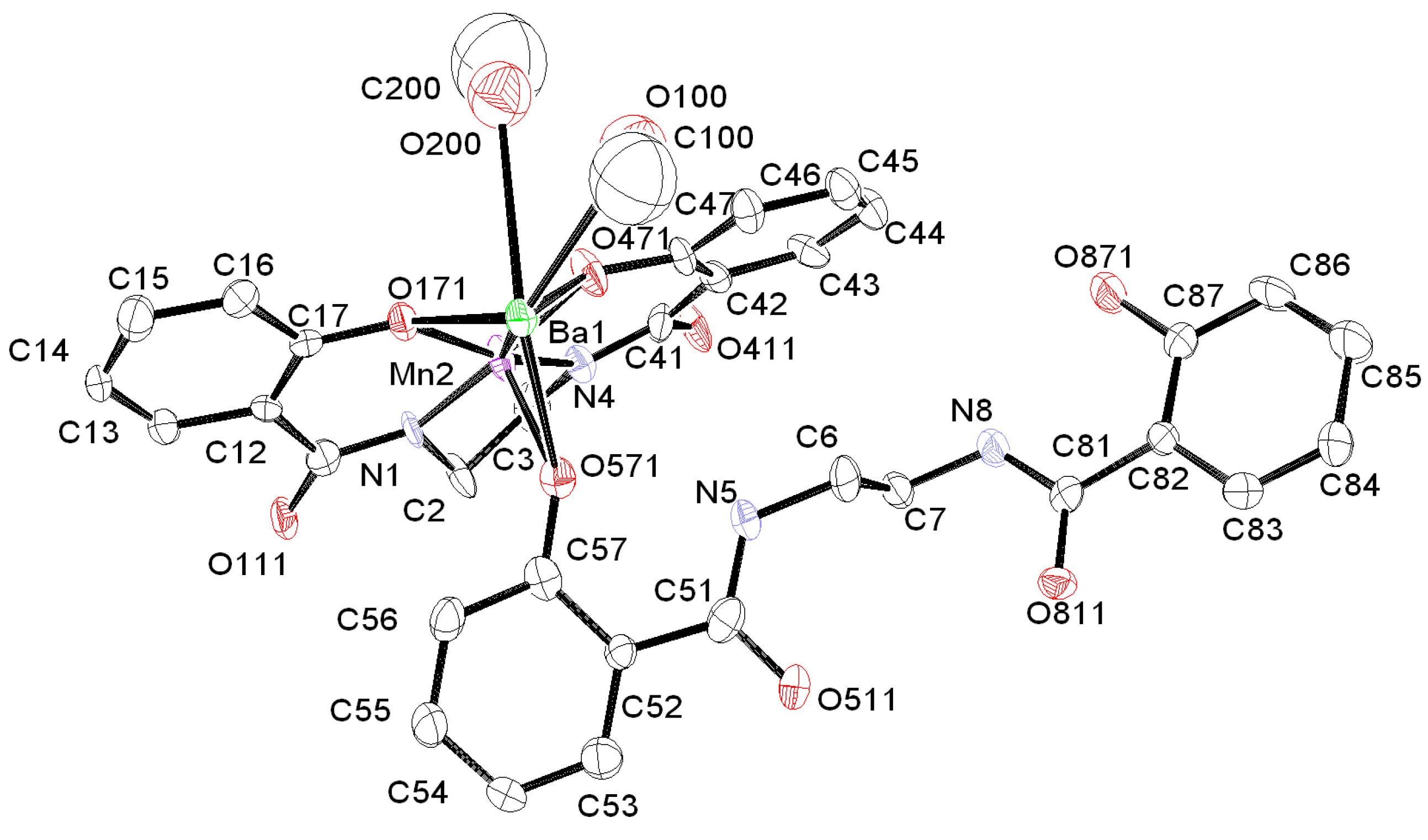
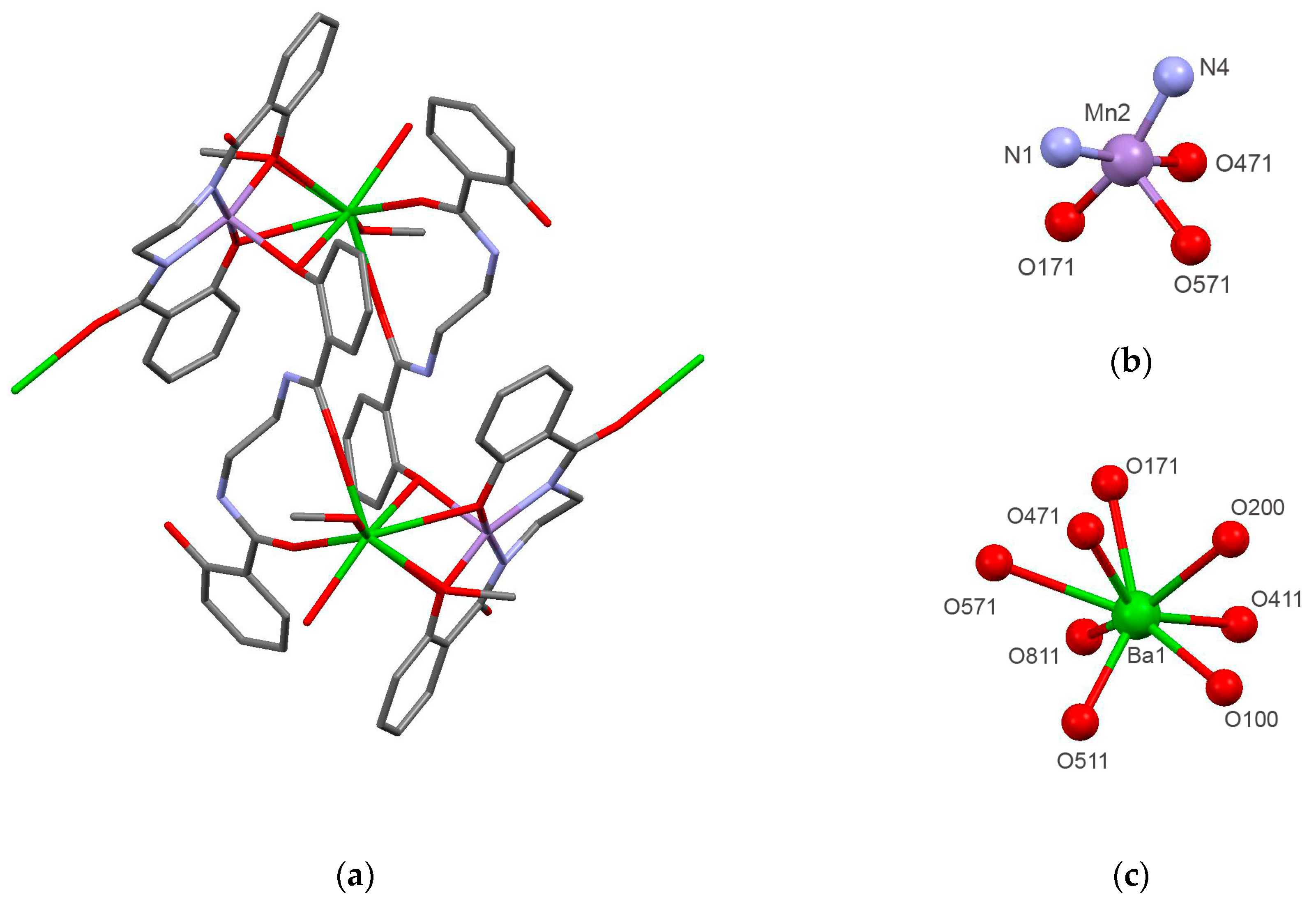
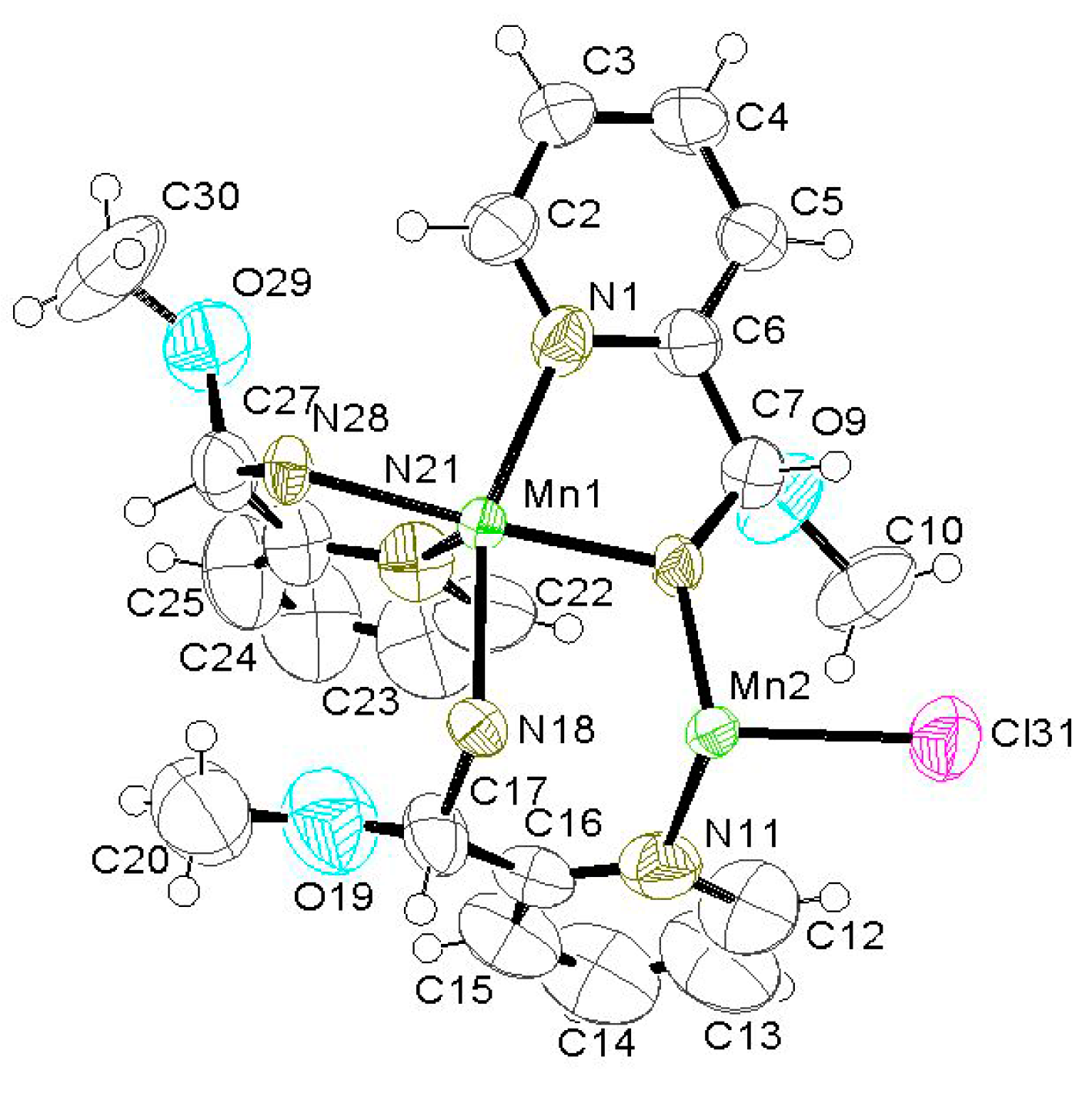
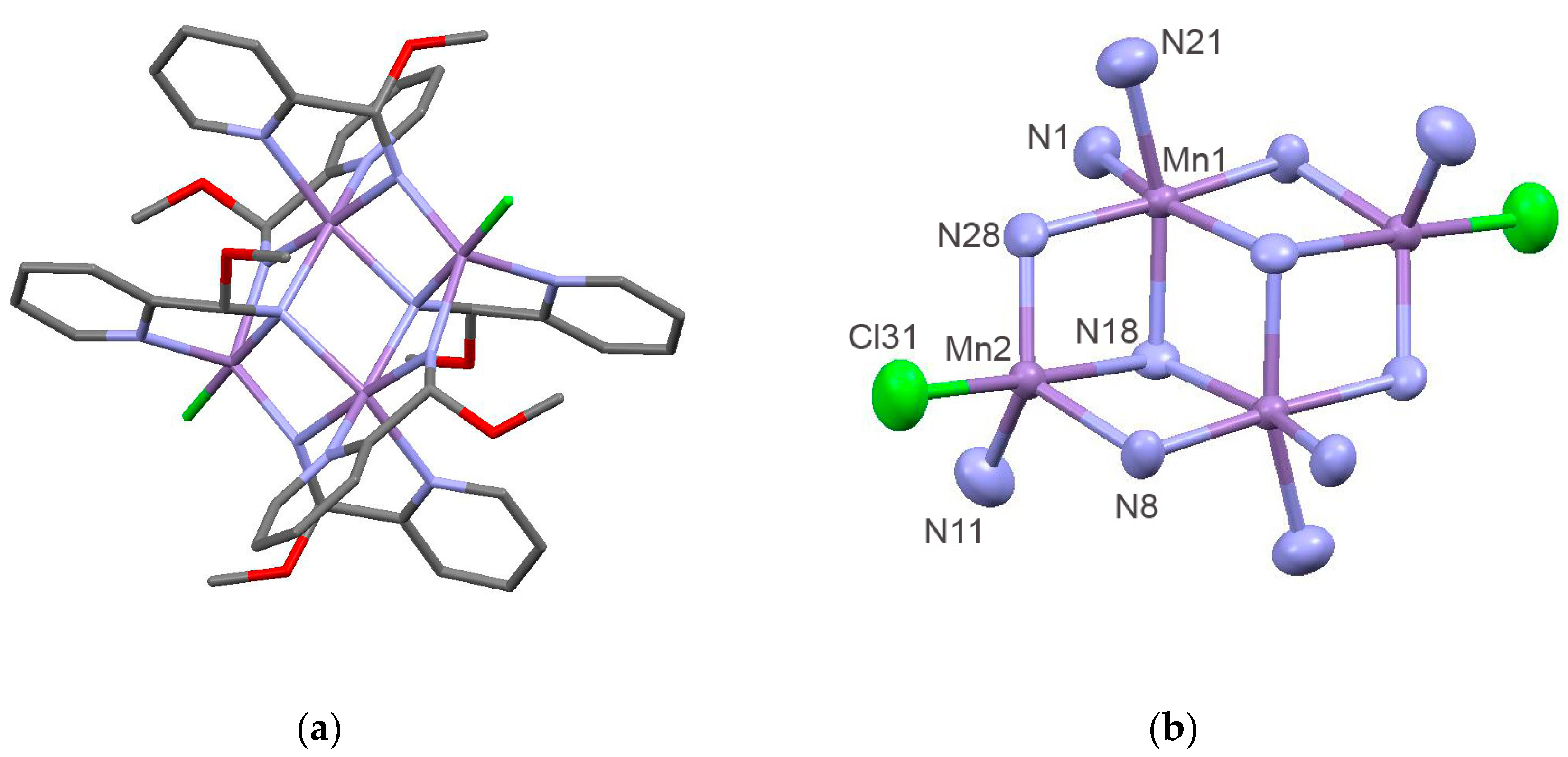
| Compound | 1 | 2 |
|---|---|---|
| Empirical formula | C34H35BaMnN4O10 | C21H24ClMn2N6O3 |
| Formula weight | 851.93 | 553.79 |
| Temperature (K) | 100(2) | 293(2) |
| Wavelength (Å) | 0.71073 | 0.71069 |
| Crystal system | Monoclinic | Monoclinic |
| Space group | P21/c | P21/n |
| a (Å) | 12.245(2) | 11.953(5) |
| b (Å) | 17.345(3) | 11.256(5) |
| c (Å) | 18.041(4) | 17.889(5) |
| α (°) | 90 | 90 |
| β (°) | 106.38(3) | 99.051(5) |
| γ (°) | 90 | 90 |
| Volume (Å3) | 3676.2(13) | 2376.9(16) |
| Z | 4 | 4 |
| Dcalcd. (g cm−3) | 1.525 | 1.548 |
| μ (mm−1) | 1.467 | 1.21 |
| F (000) | 1712 | 1132 |
| θmin/max (°) | 2.62/21.14 | 1.92/24.73 |
| Goodness-of-fit on F2 | 1.005 | 1.067 |
| Total data | 27,410 | 4038 |
| Unique data | 6303 | 4038 |
| Data/restrains/parameters | 6303/3/440 | 4038/0/299 |
| Final R indices (I > 2σ(I)) | R1 = 0.0521; wR2 = 0.1220 | R1 = 0.0894; wR2 = 0.2658 |
| R indices (all data) | R1 = 0.0941; wR2 =0.1356 | R1 = 0.1223; wR2 =0.2812 |
| Mn(2)–N(1) | 1.938(5) | Ba(1)–O(411) | 2.636(5) |
| Mn(2)–N(4) | 1.944(6) | Ba(1)–O(511) | 2.648(5) |
| Mn(2)–O(471) | 1.877(5) | Ba(1)–O(811) | 2.658(5) |
| Mn(2)–O(171) | 1.883(5) | Ba(1)–O(171) | 2.754(5) |
| Mn(2)–O(571) | 2.125(5) | Ba(1)–O(471) | 2.816(5) |
| Ba(1)–Mn(2) | 3.5510(12) | Ba(1)–O(571) | 3.074(5) |
| Ba(1)–O(200) | |||
| Ba(1)–O(100) | |||
| O(471)–Mn(2)–O(171) | 88.6(2) | O(411)–Ba(1)–O(511) | 120.81(16) |
| O(471)–Mn(2)–N(1) | 173.4(2) | O(411)–Ba(1)–O(811) | 75.91(14) |
| O(171)–-Mn(2)–N(1) | 92.4(2) | O(511)–Ba(1)–O(811) | 91.95(14) |
| O(471)–-Mn(2)–N(4) | 92.7(2) | O(411)–Ba(1)–O(171) | 97.69(15) |
| O(171)–-Mn(2)–N(4) | 164.2(2) | O(511)–Ba(1)–O(171) | 140.07(14) |
| N(1)–Mn(2)–N(4) | 84.6(2) | O(811)–Ba(1)–O(171) | 87.29(14) |
| O(471)–Mn(2)–O(571) | 84.4(2) | O(411)–Ba(1)–O(471) | 139.51(16) |
| O(171)–Mn(2)–O(571) | 92.40(19) | O(511)–Ba(1)–O(471) | 94.13(15) |
| N(1)–Mn(2)–O(571) | 102.1(2) | O(811)–Ba(1)–O(471) | 126.37(14) |
| N(4)–Mn(2)–O(571) | 103.4(2) | O(171)–Ba(1)–O(471) | 56.24(14) |
| O(411)–Ba(1)–O(571) | 142.68(14) | ||
| O(511)–Ba(1)–O(571) | 82.08(13) | ||
| O(811)–Ba(1)–O(571) | 74.13(13) | ||
| O(171)–Ba(1)–O(571) | 59.34(13) | ||
| O(471)–Ba(1)–O(571) | 54.23(13) |
| Mn(1)–N(28) | 2.048(8) | Mn(1)–Mn(2)#1 | 3.206(2) |
| Mn(1)–N(8) | 2.051(8) | Mn(2)–N(28)#1 | 1.939(8) |
| Mn(1)–N(18)#1 | 2.120(7) | Mn(2)–N(8) | 1.971(8) |
| Mn(1)–N(1) | 2.123(9) | Mn(2)–N(11) | 2.081(11) |
| Mn(1)–N(18) | 2.141(8) | Mn(2)–Cl(31) | 2.305(4) |
| Mn(1)–N(21) | 2.143(10) | Mn(2)–N(18) | 2.353(8) |
| Mn(1)–Mn(2) | 3.203(2) | Mn(2)–Mn(1)#1 | 3.206(2) |
| N(28)–Mn(1)–N(8) | 175.2(3) | N(28)#1–Mn(2)–N(8) | 124.9(4) |
| N(28)–Mn(1)–N(18)#1 | 80.3(3) | N(28)#1–Mn(2)–N(11) | 118.5(4) |
| N(8)–Mn(1)–N(18)#1 | 102.2(3) | N(8)–Mn(2)–N(11) | 101.3(4) |
| N(28)–Mn(1)–N(1) | 98.5(3) | N(28)#1–Mn(2)–Cl(31) | 101.6(2) |
| N(8)–Mn(1)–N(1) | 77.1(3) | N(8)–Mn(2)–Cl(31) | 102.9(2) |
| N(18)#1–Mn(1)–N(1) | 98.9(3) | N(11)–Mn(2)–Cl(31) | 105.2(3) |
| N(28)–Mn(1)–N(18) | 102.7(3) | N(28)#1–Mn(2)–N(18) | 76.8(3) |
| N(8)–Mn(1)–N(18) | 81.9(3) | N(8)–Mn(2)–N(18) | 78.4(3) |
| N(18)#1–Mn(1)–N(18) | 78.2(3) | N(11)–Mn(2)–N(18) | 75.4(4) |
| N(1)–Mn(1)–N(18) | 157.7(3) | Cl(31)–Mn(2)–N(18) | 178.4(2) |
| N(28)–Mn(1)–N(21) | 77.9(4) | ||
| N(8)–Mn(1)–N(21) | 100.4(4) | ||
| N(18)#1–Mn(1)–N(21) | 155.7(4) | ||
| N(1)–Mn(1)–N(21) | 94.8(4) | ||
| N(18)–Mn(1)–N(21) | 96.2(4) |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rouco, L.; Fernández-García, M.I.; Pedrido, R.; Botana, L.M.; Esteban-Gómez, D.; Platas-Iglesias, C.; Maneiro, M. Modeling the OEC with Two New Biomimetic Models: Preparations, Structural Characterization, and Water Photolysis Studies of a Ba–Mn Box Type Complex and a Mn4N6 Planar-Diamond Cluster. Catalysts 2018, 8, 382. https://doi.org/10.3390/catal8090382
Rouco L, Fernández-García MI, Pedrido R, Botana LM, Esteban-Gómez D, Platas-Iglesias C, Maneiro M. Modeling the OEC with Two New Biomimetic Models: Preparations, Structural Characterization, and Water Photolysis Studies of a Ba–Mn Box Type Complex and a Mn4N6 Planar-Diamond Cluster. Catalysts. 2018; 8(9):382. https://doi.org/10.3390/catal8090382
Chicago/Turabian StyleRouco, Lara, M. Isabel Fernández-García, Rosa Pedrido, Luis M. Botana, David Esteban-Gómez, Carlos Platas-Iglesias, and Marcelino Maneiro. 2018. "Modeling the OEC with Two New Biomimetic Models: Preparations, Structural Characterization, and Water Photolysis Studies of a Ba–Mn Box Type Complex and a Mn4N6 Planar-Diamond Cluster" Catalysts 8, no. 9: 382. https://doi.org/10.3390/catal8090382
APA StyleRouco, L., Fernández-García, M. I., Pedrido, R., Botana, L. M., Esteban-Gómez, D., Platas-Iglesias, C., & Maneiro, M. (2018). Modeling the OEC with Two New Biomimetic Models: Preparations, Structural Characterization, and Water Photolysis Studies of a Ba–Mn Box Type Complex and a Mn4N6 Planar-Diamond Cluster. Catalysts, 8(9), 382. https://doi.org/10.3390/catal8090382