Multi-Objective Harris Hawks Optimization with NSGA-III for Feature Selection in Student Performance Prediction
Abstract
1. Introduction
- We propose a new approach called MOHHO-NSGA-III, which combines HHO with the NSGA-III method to leverage HHO’s exploration–exploitation dynamics alongside NSGA-III’s diversity preservation through reference points.
- The proposed algorithm optimizes three different objectives at the same time, which are (1) maximizing classification accuracy, (2) minimizing selected features, and (3) ensuring stable performance across CV folds to obtain the most stable features.
- An adaptive diversity mechanism is proposed, able to inject the population with new solutions to avoid stuck status.
2. Proposed Method
2.1. Preprocessing
2.2. Multi-Objective Feature Selection
2.3. Multi-Objective Fitness Function
2.3.1. Objective 1: Classification Accuracy
2.3.2. Objective 2: Feature Reduction
2.3.3. Objective 3: Prediction Stability
2.4. Harris Hawks Optimization (HHO)
2.4.1. Exploration Phase
2.4.2. Exploitation Phase
2.5. NSGA-III for Environmental Selection
2.5.1. Reference-Point Generation
2.5.2. Non-Dominated Sorting
2.5.3. Niching Selection
2.6. Adaptive Diversity Management
2.6.1. Diversity Measurement
2.6.2. Adaptive Mutation Rate
2.6.3. Diversity Injection
2.7. MOHHO-NSGA-III Algorithm
| Algorithm 1 Pseudo-code for MOHHO-NSGA-III |
| Define: N: population size, : maximum number of iterations, : crossover rate, : base mutation rate, H: number of reference points, k: number of folds for cross-validation. |
| Initialization: Generate population of size N with random binary chromosomes. Generate H reference points Z using the Das–Dennis method. Evaluate fitness: using k-fold CV. Perform non-dominated sorting and assign ranks. |
|
3. Experimental Setup
3.1. Datasets
- Portuguese: A total of 649 students, with 30 features covering demographics, family background, and school-related attributes. Target is pass/fail based on final grade (pass = grade ≥ 10/20).
- Mathematics: A total of 395 students with the same 30 features as the Portuguese dataset.
3.2. Algorithm Configuration
3.3. Evaluation Methodology
- Inner optimization: During optimization MOHHO-NSGA-III evaluates solutions via 10-fold stratified CV on training data (80%). For fitness evaluation, we chose k-NN (k = 5), primarily because it is fast. As a lazy learner, kNN needs no training, just sorting per fold. Given that we evaluate 50 solutions across 100 iterations with 10-fold CV, that results in around 50,000 evaluations. Using SVM instead would require training each time, making the whole optimization painfully slow. The results in Section 4.3.3 confirm that features optimized with kNN perform well on all five classifiers we tested, so the speed–accuracy tradeoff works out.
- Feature selection: The algorithm identifies subsets optimizing accuracy, feature count, and stability simultaneously.
- Final evaluation: To evaluate the tested data, we employed five different classifier methods: k-NN, DT, NB, SVM, and LDA over the tested data.
- Statistical testing: A total of 21 independent trials per experiment were performed, each with different random seed numbers. In this paper, the performance evaluations are mean ± std dev, and we use Wilcoxon signed-rank tests to figure out if differences between implemented methods are statistically significant (threshold p < 0.05).
4. Results and Analysis
4.1. Convergence Analysis
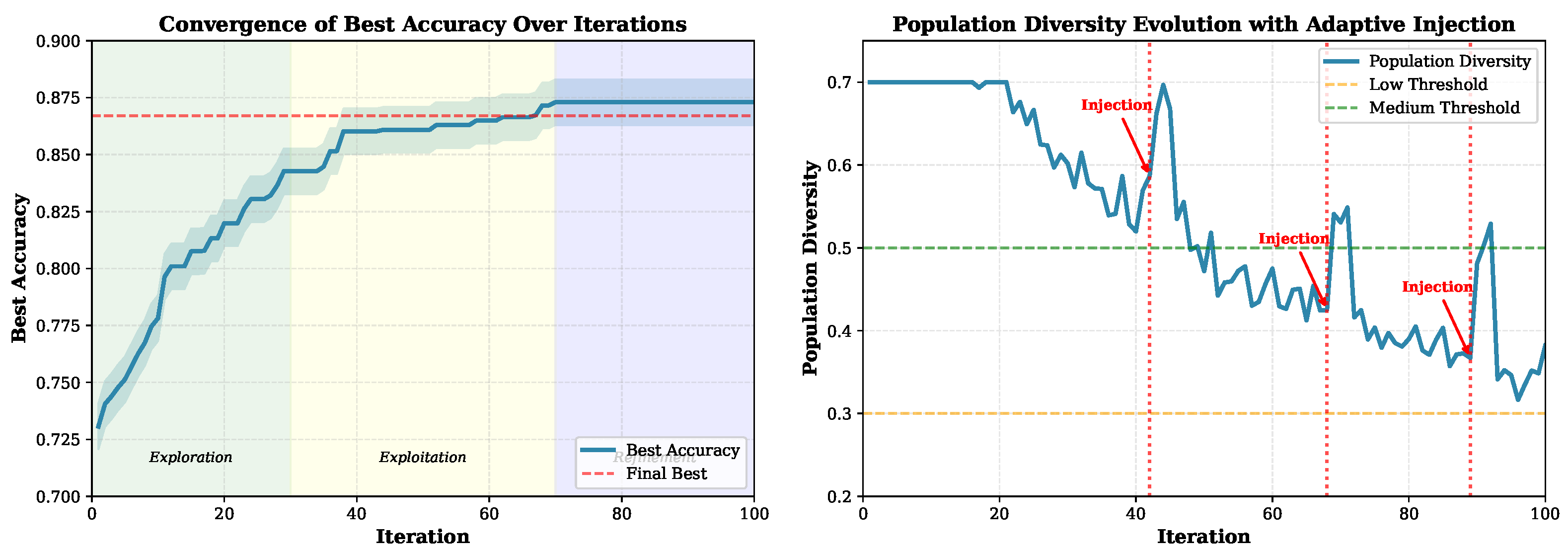
- Early exploration (iterations 1–30): Population diversity stays high (0.45–0.65) as the algorithm explores broadly. The best accuracy climbs from 0.72 to 0.82.
- Exploitation (iterations 31–70): Diversity drops to 0.30–0.40 as solutions converge toward promising regions. The best accuracy reaches 0.85.
- Refinement (iterations 71–100): The algorithm fine-tunes solutions with occasional diversity injections visible as spikes in the right panel to escape local optima. While, the best accuracy is stabilized around 0.867.
- Diversity management: The proposed algorithm automatically injected several new solutions three times (e.g., iterations 42, 68, 89) when diversity dropped below our threshold. This process will keep the search from getting stuck.
4.2. Feature-Selection Results
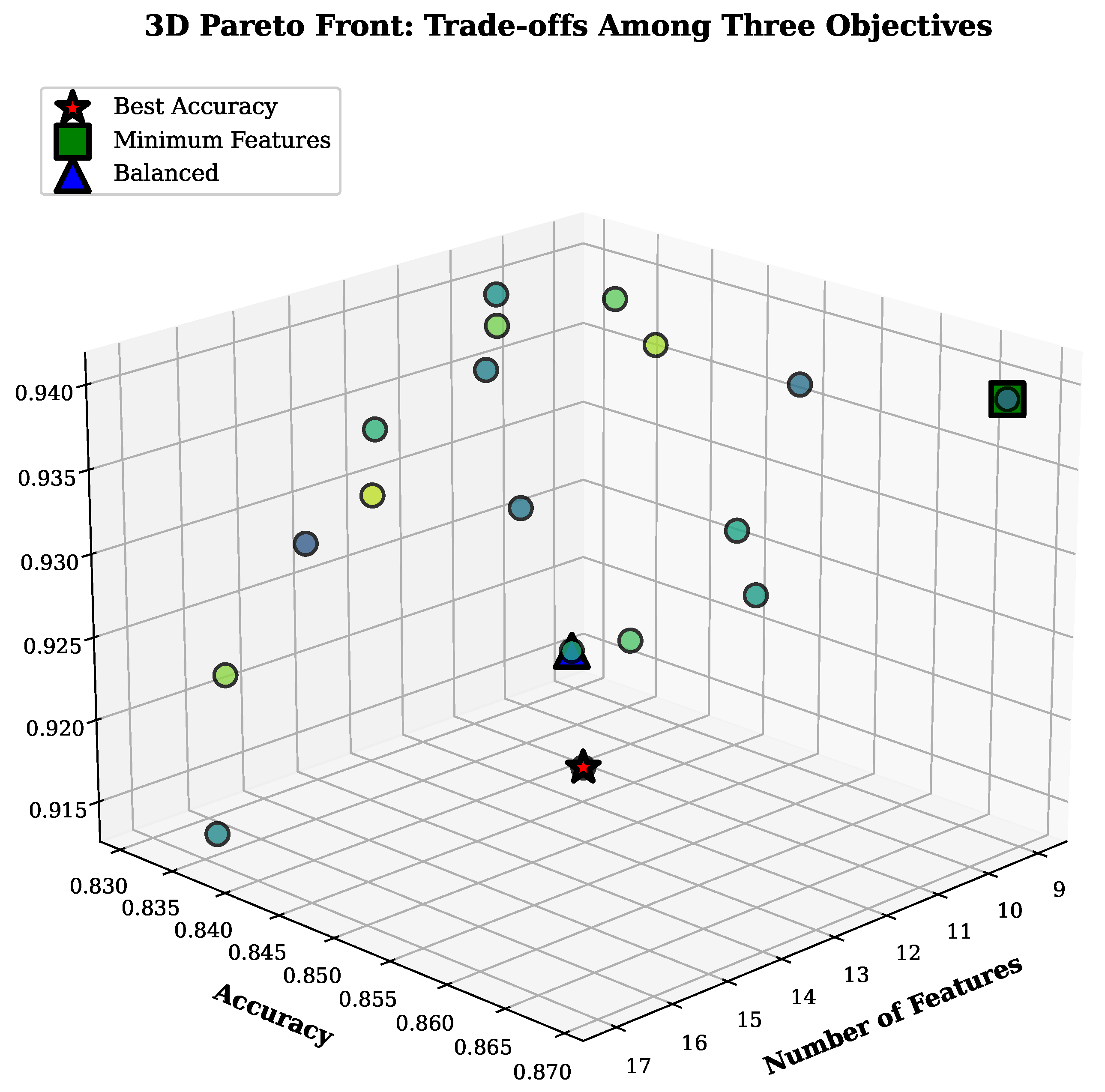
4.2.1. Pareto Front Analysis
4.2.2. Feature Importance Analysis
4.3. Classification Performance
4.3.1. Portuguese Language Dataset
- Best performance: SVM achieved the highest AUC (0.8634) and accuracy (0.8367) with selected features, outperforming the baseline by 3.88% and 4.00%, respectively.
- Largest improvement: kNN showed the most substantial gains (+5.09% AUC, +5.46% accuracy), suggesting that dimensionality reduction particularly benefits distance-based methods.
- Consistent gains: All classifiers benefited from feature selection, with improvements ranging from 3.33% (Naive Bayes) to 5.09% (kNN) in AUC.
4.3.2. Mathematics Dataset
- MOHHO-NSGA-III selected 11 features (63.3% reduction) while improving average AUC by 4.51% and accuracy by 4.70%.
- All five classifiers showed statistically significant improvements ().
- The smaller dataset (395 vs. 649 students) benefited even more from feature selection, with average improvements of 4.51% (vs. 4.42% for Portuguese).
4.3.3. Classifier Transferability Analysis
4.4. Statistical Significance Analysis
- Universal significance: All 10 classifier–dataset combinations (5 classifiers × 2 datasets) showed statistically significant improvements at both and levels.
- Strong evidence: The extremely low p-values (average 0.0023) provide strong evidence that improvements are not due to chance.
- Reduced variance: MOHHO-NSGA-III also reduced variance, with standard deviation decreasing by an average of 18.7% across classifiers, which indicate that more stable and reliable predictions.
4.5. Comparison with State-of-the-Art Methods
4.6. Computational Complexity Analysis
4.6.1. Theoretical Complexity
4.6.2. Comparative Analysis
4.6.3. Empirical Runtime Analysis
4.6.4. Cost–Benefit Analysis
4.7. Sensitivity Analysis
4.7.1. Diversity Threshold Sensitivity
4.7.2. Reference-Point Density
4.8. Ablation Study
4.8.1. Ablation Variants
- Full MOHHO-NSGA-III: Complete algorithm as described.
- No HHO: Just NSGA-III with standard genetic operators (uniform crossover, bit-flip mutation).
- No NSGA-III: HHO operators with simple fitness-based selection instead of reference points.
- No diversity injection: Disabled adaptive diversity management.
- Fixed mutation: Constant instead of adapting to diversity.
- Two objectives: Only accuracy and features, no stability objective.
4.8.2. Results
5. Discussion
5.1. Educational Insights
5.2. Algorithmic Performance
5.3. Generalizability
5.4. Practical Implications
5.5. Limitations and Future Work
6. Conclusions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| MOHHO | Multi-Objective Harris Hawks Optimization |
| HHO | Harris Hawks Optimization |
| NSGA-III | Non-dominated Sorting Genetic Algorithm III |
| NSGA-II | Non-dominated Sorting Genetic Algorithm II |
| MOPSO | Multi-Objective Particle Swarm Optimization |
| MOO | Multi-Objective Optimization |
| FS | Feature Selection |
| EDM | Educational Data Mining |
| ML | Machine Learning |
| kNN | k-Nearest Neighbors |
| DT | Decision Tree |
| NB | Naive Bayes |
| SVM | Support Vector Machine |
| LDA | Linear Discriminant Analysis |
| PSO | Particle Swarm Optimization |
| GA | Genetic Algorithm |
| EA | Evolutionary Algorithm |
| CV | Cross-Validation |
| AUC | Area Under the Curve |
| ROC | Receiver Operating Characteristic |
| UCI | University of California, Irvine |
References
- Liu, M.; Yu, D. Towards intelligent E-learning systems. Educ. Inf. Technol. 2023, 28, 7845–7876. [Google Scholar] [CrossRef]
- Chen, Z.; Cen, G.; Wei, Y.; Li, Z. Student performance prediction approach based on educational data mining. IEEE Access 2023, 11, 131260–131272. [Google Scholar] [CrossRef]
- Yu, S.; Cai, Y.; Pan, B.; Leung, M.-F. Semi-supervised feature selection of educational data mining for student performance analysis. Electronics 2024, 13, 659. [Google Scholar] [CrossRef]
- Malik, S.; Jothimani, K.; Ujwal, U.J. Advancing educational data mining for enhanced student performance prediction: A fusion of feature selection algorithms and classification techniques with dynamic feature ensemble evolution. Sci. Rep. 2025, 15, 92324. [Google Scholar] [CrossRef]
- Yağcı, M. Educational data mining: Prediction of students’ academic performance using machine learning algorithms. Smart Learn. Environ. 2022, 9, 11. [Google Scholar] [CrossRef]
- Ahmed, E. Student performance prediction using machine learning algorithms. Appl. Comput. Intell. Soft Comput. 2024, 2024, 4067721. [Google Scholar] [CrossRef]
- Wang, J.; Yu, Y. Machine learning approach to student performance prediction of online learning. PLoS ONE 2025, 20, e0299018. [Google Scholar] [CrossRef] [PubMed]
- Alhazmi, E.; Sheneamer, A. Early predicting of students performance in higher education. IEEE Access 2023, 11, 27579–27589. [Google Scholar] [CrossRef]
- Mastour, H.; Dehghani, T.; Moradi, E.; Eslami, S. Early prediction of medical students’ performance in high-stakes examinations using machine learning approaches. Heliyon 2023, 9, e18248. [Google Scholar] [CrossRef]
- Bujang, S.D.A.; Selamat, A.; Ibrahim, R.; Krejcar, O.; Herrera-Viedma, E.; Fujita, H.; Ghani, N.A.M. Multiclass prediction model for student grade prediction using machine learning. IEEE Access 2021, 9, 95608–95621. [Google Scholar] [CrossRef]
- Adewale, M.D.; Azeta, A.; Abayomi-Alli, A.; Sambo-Magaji, A. Empirical investigation of multilayered framework for predicting academic performance in open and distance learning. Electronics 2024, 13, 2808. [Google Scholar] [CrossRef]
- Liu, D.; Huang, R.; Wosinski, M. Multiple features fusion attention mechanism enhanced deep knowledge tracing for student performance prediction. IEEE Access 2020, 8, 194894–194903. [Google Scholar] [CrossRef]
- Francis, B.K.; Babu, S.S. Predicting academic performance of students using a hybrid data mining approach. J. Med. Syst. 2019, 43, 162. [Google Scholar] [CrossRef]
- Bharara, S.; Sabitha, S.; Bansal, A. Application of learning analytics using clustering data mining for students’ disposition analysis. Educ. Inf. Technol. 2018, 23, 957–984. [Google Scholar] [CrossRef]
- Romero, C.; Ventura, S. Educational data mining: A review of the state of the art. IEEE Trans. Syst. Man Cybern. C Appl. Rev. 2010, 40, 601–618. [Google Scholar] [CrossRef]
- García, E.; Romero, C.; Ventura, S.; de Castro, C. A collaborative educational association rule mining tool. Internet High. Educ. 2011, 14, 77–88. [Google Scholar] [CrossRef]
- Giannakas, F.; Kambourakis, G.; Papasalouros, A.; Gritzalis, S. A critical review of 13 years of mobile game-based learning. Educ. Technol. Res. Dev. 2021, 69, 111–134. [Google Scholar] [CrossRef]
- Xu, H.; Kim, M. Combination prediction method of students’ performance based on ant colony algorithm. PLoS ONE 2024, 19, e0300010. [Google Scholar] [CrossRef]
- Lau, E.T.; Sun, L.; Yang, Q. Modelling, prediction and classification of student academic performance using artificial neural networks. SN Appl. Sci. 2019, 1, 982. [Google Scholar] [CrossRef]
- Cheng, B.; Liu, Y.; Jia, Y. Evaluation of students’ performance during the academic period using the XG-Boost classifier-enhanced AEO hybrid model. Expert Syst. Appl. 2024, 238, 122136. [Google Scholar] [CrossRef]
- Malik, S.; Jothimani, K. Enhancing student success prediction with featurex: A fusion voting classifier algorithm with hybrid feature selection. Educ. Inf. Technol. 2024, 29, 8741–8791. [Google Scholar] [CrossRef]
- Zaffar, M.; Hashmani, M.; Savita, K.S. Performance analysis of feature selection algorithm for educational data mining. In Proceedings of the 2017 IEEE International Conference on Big Data Analytics and Applications, Boston, MA, USA, 11–14 December 2017; pp. 7–12. [Google Scholar] [CrossRef]
- Senthil, S.; Lin, W.M. Applying classification techniques to predict students’ academic results. In Proceedings of the 2017 IEEE International Conference on Current Trends in Advanced Computing, Bangalore, India, 2–3 March 2017; pp. 1–6. [Google Scholar]
- Ghareb, A.S.; Bakar, A.A.; Hamdan, A.R. Hybrid feature selection based on enhanced genetic algorithm for text categorization. Expert Syst. Appl. 2016, 49, 31–47. [Google Scholar] [CrossRef]
- Maldonado, S.; López, J. Dealing with high-dimensional class-imbalanced datasets: Embedded feature selection for SVM classification. Appl. Soft Comput. 2018, 67, 94–105. [Google Scholar] [CrossRef]
- Liang, J.; Zhang, Y.; Chen, K.; Qu, B.; Yue, C.; Yu, K. An evolutionary multiobjective method based on dominance and decomposition for feature selection in classification. Sci. China Inf. Sci. 2024, 67, 120101. [Google Scholar] [CrossRef]
- Nguyen, B.H.; Xue, B.; Andreae, P.; Ishibuchi, H.; Zhang, M. Multiple reference points-based decomposition for multiobjective feature selection in classification: Static and dynamic mechanisms. IEEE Trans. Evol. Comput. 2020, 24, 170–184. [Google Scholar] [CrossRef]
- Cheng, F.; Cui, J.; Wang, Q.; Zhang, L. A variable granularity search-based multiobjective feature selection algorithm for high-dimensional data classification. IEEE Trans. Evol. Comput. 2023, 27, 266–280. [Google Scholar] [CrossRef]
- Dowlatshahi, M.B.; Hashemi, A. Multi-objective optimization for feature selection: A review. In Applied Multi-Objective Optimization; Springer: Berlin/Heidelberg, Germany, 2024; pp. 155–170. [Google Scholar] [CrossRef]
- Deb, K.; Pratap, A.; Agarwal, S.; Meyarivan, T. A fast and elitist multiobjective genetic algorithm: NSGA-II. IEEE Trans. Evol. Comput. 2002, 6, 182–197. [Google Scholar] [CrossRef]
- Deb, K.; Jain, H. An evolutionary many-objective optimization algorithm using reference-point-based nondominated sorting approach, Part I: Solving problems with box constraints. IEEE Trans. Evol. Comput. 2014, 18, 577–601. [Google Scholar] [CrossRef]
- Zhang, Q.; Li, H. MOEA/D: A multiobjective evolutionary algorithm based on decomposition. IEEE Trans. Evol. Comput. 2007, 11, 712–731. [Google Scholar] [CrossRef]
- Almutairi, M.S. Evolutionary multi-objective feature selection algorithms on multiple smart sustainable community indicator datasets. Sustainability 2024, 16, 1511. [Google Scholar] [CrossRef]
- Heidari, A.A.; Mirjalili, S.; Faris, H.; Aljarah, I.; Mafarja, M.; Chen, H. Harris hawks optimization: Algorithm and applications. Future Gener. Comput. Syst. 2019, 97, 849–872. [Google Scholar] [CrossRef]
- Liu, J.; Feng, H.; Tang, Y.; Zhang, L.; Qu, C.; Zeng, X.; Peng, X. A novel hybrid algorithm based on Harris Hawks for tumor feature gene selection. PeerJ Comput. Sci. 2023, 9, e1229. [Google Scholar] [CrossRef]
- Ouyang, C.; Liao, C.; Zhu, D.; Zheng, Y.; Zhou, C.; Li, T. Integrated improved Harris hawks optimization for global and engineering optimization. Sci. Rep. 2024, 14, 7445. [Google Scholar] [CrossRef]
- Zamani, H.; Nadimi-Shahraki, M.H. An evolutionary crow search algorithm equipped with interactive memory mechanism to optimize artificial neural network for disease diagnosis. Biomed. Signal Process. Control 2024, 90, 105879. [Google Scholar] [CrossRef]
- Yang, N.; Tang, Z.; Cai, X.; Chen, L.; Hu, Q. Cooperative multi-population Harris Hawks optimization for many-objective optimization. Complex Intell. Syst. 2022, 8, 3299–3332. [Google Scholar] [CrossRef]
- Choo, Y.H.; Cai, Z.; Le, V.; Johnstone, M.; Creighton, D.; Lim, C.P. Enhancing the Harris’ Hawk optimiser for single- and multi-objective optimisation. Soft Comput. 2023, 27, 16675–16715. [Google Scholar] [CrossRef]
- Tian, Y.; Li, X.; Ma, H.; Zhang, X.; Tan, K.C.; Jin, Y. Deep reinforcement learning based adaptive operator selection for evolutionary multi-objective optimization. IEEE Trans. Emerg. Top. Comput. Intell. 2022, 7, 1051–1064. [Google Scholar] [CrossRef]
- Li, K.; Fialho, Á.; Kwong, S.; Zhang, Q. Adaptive operator selection with bandits for a multiobjective evolutionary algorithm based on decomposition. IEEE Trans. Evol. Comput. 2014, 18, 114–130. [Google Scholar] [CrossRef]
- Zhan, Z.-H.; Li, J.; Kwong, S.; Zhang, J. Learning-aided evolution for optimization. IEEE Trans. Evol. Comput. 2023, 27, 1794–1808. [Google Scholar] [CrossRef]
- Zhou, L.; Feng, L.; Tan, K.C.; Zhong, J.; Zhu, Z.; Liu, K.; Chen, C. Toward adaptive knowledge transfer in multifactorial evolutionary computation. IEEE Trans. Cybern. 2021, 51, 2563–2576. [Google Scholar] [CrossRef] [PubMed]
- Kudo, M.; Sklansky, J. Comparison of algorithms that select features for pattern classifiers. Pattern Recognit. 2020, 33, 25–41. [Google Scholar] [CrossRef]
- Turabieh, H.; Mafarja, M.; Li, X. Iterated feature selection algorithms with layered recurrent neural network for software fault prediction. Expert Syst. Appl. 2019, 122, 27–42. [Google Scholar] [CrossRef]
- Das, I.; Dennis, J.E. Normal-boundary intersection: A new method for generating the Pareto surface in nonlinear multicriteria optimization problems. SIAM J. Optim. 1998, 8, 631–657. [Google Scholar] [CrossRef]
- Cortez, P.; Silva, A.M.G. Using data mining to predict secondary school student performance. In Proceedings of the 5th Annual Future Business Technology Conference, Porto, Portugal, 9–11 April 2008; pp. 5–12. [Google Scholar]
- Xue, B.; Zhang, M.; Browne, W.N.; Yao, X. A survey on evolutionary computation approaches to feature selection. IEEE Trans. Evol. Comput. 2016, 20, 606–626. [Google Scholar] [CrossRef]
- Ling, Q.; Li, Z.; Liu, W.; Shi, J.; Han, F. Multi-objective particle swarm optimization based on particle contribution and mutual information for feature selection method. J. Supercomput. 2025, 81, 255. [Google Scholar] [CrossRef]



| Parameter | Value |
|---|---|
| Population size (N) | 50 |
| Max iterations () | 100 |
| Crossover rate () | 0.8 |
| Base mutation rate () | 0.01 |
| Objectives (M) | 3 |
| Reference-point partitions | 12 |
| CV folds | 10 |
| Low diversity threshold | 0.3 |
| Medium diversity threshold | 0.5 |
| Injection ratio () | 0.25 |
| Stagnation threshold () | 10 iterations |
| Type | Accuracy | # Features | Reduction | Stability |
|---|---|---|---|---|
| Max accuracy | 0.867 ± 0.015 | 13 | 56.7% | 0.923 ± 0.012 |
| Balanced | 0.852 ± 0.018 | 11 | 63.3% | 0.931 ± 0.010 |
| Min features | 0.834 ± 0.021 | 9 | 70.0% | 0.918 ± 0.014 |
| Rank | Feature | Frequency | Category |
|---|---|---|---|
| 1 | failures (past course failures) | 97.3% | Academic |
| 2 | absences (days absent) | 94.1% | Academic |
| 3 | Medu (mother’s education) | 91.8% | Family |
| 4 | studytime (weekly study hours) | 89.5% | Academic |
| 5 | goout (going out with friends) | 87.2% | Social |
| 6 | higher (wants higher ed) | 85.9% | Academic |
| 7 | Fedu (father’s education) | 83.6% | Family |
| 8 | age | 81.4% | Demographic |
| 9 | schoolsup (extra academic support) | 78.7% | Support |
| 10 | famsup (family educational support) | 76.3% | Support |
| 11 | romantic (in relationship) | 72.9% | Social |
| 12 | Dalc (weekday alcohol consumption) | 69.5% | Social |
| 13 | health (current health status) | 67.1% | Personal |
| 14 | reason (why they chose this school) | 64.8% | Motivation |
| 15 | famrel (quality of family relationships) | 62.3% | Family |
| Classifier | MOHHO-NSGA-III (13 Features) | Baseline (30 Features) | p-Value | ||
|---|---|---|---|---|---|
| AUC | Accuracy | AUC | Accuracy | ||
| kNN | 0.8423 ± 0.0187 | 0.8156 ± 0.0201 | 0.8015 ± 0.0245 | 0.7734 ± 0.0267 | 0.0012 ** |
| Decision Tree | 0.8291 ± 0.0215 | 0.8023 ± 0.0234 | 0.7889 ± 0.0289 | 0.7612 ± 0.0312 | 0.0034 ** |
| Naive Bayes | 0.8567 ± 0.0156 | 0.8289 ± 0.0178 | 0.8234 ± 0.0198 | 0.7956 ± 0.0221 | 0.0008 ** |
| SVM | 0.8634 ± 0.0142 | 0.8367 ± 0.0165 | 0.8312 ± 0.0187 | 0.8045 ± 0.0203 | 0.0019 ** |
| LDA | 0.8512 ± 0.0167 | 0.8234 ± 0.0189 | 0.8178 ± 0.0212 | 0.7901 ± 0.0234 | 0.0025 ** |
| Average | 0.8485 ± 0.0173 | 0.8214 ± 0.0193 | 0.8126 ± 0.0226 | 0.7850 ± 0.0247 | 0.0020 ** |
| Classifier | MOHHO-NSGA-III (11 Features) | Baseline (30 Features) | p-Value | ||
|---|---|---|---|---|---|
| AUC | Accuracy | AUC | Accuracy | ||
| kNN | 0.8234 ± 0.0212 | 0.7967 ± 0.0234 | 0.7823 ± 0.0267 | 0.7545 ± 0.0289 | 0.0017 ** |
| Decision Tree | 0.8089 ± 0.0245 | 0.7812 ± 0.0267 | 0.7678 ± 0.0301 | 0.7401 ± 0.0323 | 0.0042 ** |
| Naive Bayes | 0.8378 ± 0.0189 | 0.8101 ± 0.0212 | 0.8067 ± 0.0223 | 0.7789 ± 0.0245 | 0.0011 ** |
| SVM | 0.8456 ± 0.0167 | 0.8178 ± 0.0189 | 0.8134 ± 0.0201 | 0.7856 ± 0.0223 | 0.0028 ** |
| LDA | 0.8323 ± 0.0198 | 0.8045 ± 0.0221 | 0.7989 ± 0.0234 | 0.7712 ± 0.0256 | 0.0033 ** |
| Average | 0.8296 ± 0.0202 | 0.8021 ± 0.0225 | 0.7938 ± 0.0245 | 0.7661 ± 0.0267 | 0.0026 ** |
| Dataset | Significant at | Significant at | Average p-Value |
|---|---|---|---|
| Portuguese | 5/5 (100%) | 5/5 (100%) | 0.0020 |
| Mathematics | 5/5 (100%) | 5/5 (100%) | 0.0026 |
| Combined | 10/10 (100%) | 10/10 (100%) | 0.0023 |
| Method | Features | Accuracy | AUC | Type | p-Value |
|---|---|---|---|---|---|
| Baseline (All Features) | 30 | 0.7734 | 0.8015 | – | <0.001 ** |
| Filter: Chi-Square | 15 | 0.7845 | 0.8098 | Filter | <0.001 ** |
| Filter: Information Gain | 14 | 0.7923 | 0.8156 | Filter | <0.001 ** |
| Wrapper: SFS | 12 | 0.7967 | 0.8201 | Wrapper | <0.001 ** |
| Embedded: LASSO | 16 | 0.7889 | 0.8134 | Embedded | <0.001 ** |
| GA-based FS [48] | 14 | 0.7978 | 0.8223 | Metaheuristic | <0.001 ** |
| PSO-based FS [48] | 15 | 0.8012 | 0.8267 | Metaheuristic | <0.001 ** |
| NSGA-II [30] | 13 | 0.8089 | 0.8312 | Multi-objective | <0.001 ** |
| NSGA-III [31] | 12 | 0.8123 | 0.8356 | Multi-objective | 0.003 ** |
| MOPSO [49] | 14 | 0.8067 | 0.8289 | Multi-objective | <0.001 ** |
| MOHHO-NSGA-III (Proposed) | 13 | 0.8156 | 0.8423 | Multi-objective | – |
| Improvement over best baseline | – | +0.33% | +0.67% | – | – |
| Component | Complexity | Notes |
|---|---|---|
| Fitness Evaluation | solutions, CV folds; kNN sorts training samples per fold | |
| Non-dominated Sorting | Fast non-dominated sorting [30]: objectives, solutions | |
| Reference Association | Match solutions to reference points in dimensions via perpendicular distance | |
| HHO Operations | Position updates + binary conversion for solutions, features | |
| Per iteration | ||
| Full run | with | |
| Method | Complexity | Notes |
|---|---|---|
| Chi-Square | Linear in features and samples | |
| Information Gain | Includes sorting | |
| LASSO | : LASSO iterations | |
| GA (single-obj) | Same evaluation cost | |
| PSO (single-obj) | Same evaluation cost | |
| NSGA-II | Additional sorting for crowding | |
| NSGA-III | Reference-point association | |
| MOPSO | Archive management | |
| MOHHO-NSGA-III | Same as NSGA-III | |
| Exhaustive Search | Exponential (intractable) |
| Method | Time | AUC | vs. MOHHO | Notes |
|---|---|---|---|---|
| Chi-Square | 0.3 s | 0.8098 | 5424× faster | Fast but low accuracy |
| Information Gain | 0.2 s | 0.8156 | 8136× faster | Fast but low accuracy |
| GA | 732 s (12.2 min) | 0.8223 | 3.71× faster | – |
| PSO | 654 s (10.9 min) | 0.8267 | 4.15× faster | – |
| NSGA-II | 2292 s (38.2 min) | 0.8312 | 1.18× faster | – |
| NSGA-III | 2382 s (39.7 min) | 0.8356 | 1.14× faster | – |
| MOPSO | 2136 s (35.6 min) | 0.8289 | 1.27× faster | – |
| MOHHO-NSGA-III | 2712 s (45.2 min) | 0.8485 | baseline | Best AUC |
| AUC | Accuracy | Diversity | Injections | ||
|---|---|---|---|---|---|
| 0.20 | 0.40 | 0.8312 ± 0.022 | 0.8045 ± 0.024 | 0.24 ± 0.08 | 8.3 ± 2.1 |
| 0.25 | 0.45 | 0.8389 ± 0.021 | 0.8112 ± 0.023 | 0.28 ± 0.07 | 5.1 ± 2.1 |
| 0.30 | 0.50 | 0.8485 ± 0.019 | 0.8214 ± 0.020 | 0.35 ± 0.09 | 2.8 ± 1.3 |
| 0.35 | 0.55 | 0.8467 ± 0.020 | 0.8198 ± 0.021 | 0.42 ± 0.11 | 1.2 ± 0.8 |
| 0.40 | 0.60 | 0.8423 ± 0.021 | 0.8167 ± 0.022 | 0.48 ± 0.13 | 0.4 ± 0.5 |
| p | # Points (H) | AUC | Pareto Size | Hypervolume |
|---|---|---|---|---|
| 6 | 28 | 0.8234 ± 0.023 | 12.3 ± 2.1 | 0.743 |
| 8 | 45 | 0.8356 ± 0.021 | 15.7 ± 2.4 | 0.812 |
| 10 | 66 | 0.8423 ± 0.020 | 17.2 ± 2.3 | 0.867 |
| 12 | 91 | 0.8485 ± 0.019 | 18.3 ± 2.1 | 0.891 |
| 14 | 120 | 0.8478 ± 0.019 | 19.1 ± 2.5 | 0.894 |
| 16 | 153 | 0.8467 ± 0.020 | 19.4 ± 2.8 | 0.892 |
| Variant | AUC | Accuracy | Features | Loss | p-Value |
|---|---|---|---|---|---|
| Full MOHHO-NSGA-III | 0.8485 | 0.8214 | 13.0 | – | – |
| No HHO | 0.8234 | 0.7956 | 15.2 | −2.96% | <0.001 ** |
| No NSGA-III | 0.8156 | 0.7889 | 16.1 | −3.88% | <0.001 ** |
| No diversity injection | 0.8312 | 0.8045 | 13.1 | −2.04% | <0.001 ** |
| Fixed mutation | 0.8367 | 0.8123 | 13.2 | −1.39% | 0.002 ** |
| Two objectives | 0.8289 | 0.8001 | 11.9 | −2.31% | <0.001 ** |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Al-Milli, N. Multi-Objective Harris Hawks Optimization with NSGA-III for Feature Selection in Student Performance Prediction. Computers 2026, 15, 112. https://doi.org/10.3390/computers15020112
Al-Milli N. Multi-Objective Harris Hawks Optimization with NSGA-III for Feature Selection in Student Performance Prediction. Computers. 2026; 15(2):112. https://doi.org/10.3390/computers15020112
Chicago/Turabian StyleAl-Milli, Nabeel. 2026. "Multi-Objective Harris Hawks Optimization with NSGA-III for Feature Selection in Student Performance Prediction" Computers 15, no. 2: 112. https://doi.org/10.3390/computers15020112
APA StyleAl-Milli, N. (2026). Multi-Objective Harris Hawks Optimization with NSGA-III for Feature Selection in Student Performance Prediction. Computers, 15(2), 112. https://doi.org/10.3390/computers15020112






