Long Non-Coding RNAs: Key Regulators of Epithelial-Mesenchymal Transition, Tumour Drug Resistance and Cancer Stem Cells
Abstract
1. Introduction
Epithelial-Mesenchymal Transition: Cancer’s Gateway to Metastasis
2. Non-Coding RNA
2.1. The Non-Coding RNA Revolution
2.2. Long Non-Coding RNAs
2.3. Competing Endogenous RNAs
2.4. miRNAs and EMT
2.5. LncRNAs as Mediators of Epigenetic Silencing Via PRC2 Recruitment
3. LncRNAs and EMT: The Main Players
3.1. Pro-EMT
3.1.1. LncRNAs with Pro-EMT Activity
3.1.2. Antisense lncRNAs
3.1.3. LincRNAs
3.2. Anti-EMT LncRNAs
3.2.1. lncRNAs
3.2.2. lincRNAs
3.3. LncRNAs with Varying Roles in EMT Regulation in Different Cancers
4. LncRNAs in Signalling Pathways Governing EMT
4.1. TGF-β Signalling
4.2. Wnt/β-Catenin Signalling
4.3. MAPK/ERK Signalling
4.4. PI3K/Akt Signalling
4.5. HIF-1α Signalling
4.6. p53
4.7. Other Pathways
5. LncRNAs, EMT and the Cancer Stem Cell Phenotype
6. LncRNAs, EMT and Drug Resistance
6.1. EMT LncRNAs and Platinum Based Resistance
6.2. EMT LncRNAs and Resistance to Other Chemotherapeutic Drugs
6.3. EMT lncRNAs and Resistance to Targeted Breast Cancer Therapies
6.4. EMT lncRNAs and Resistance to EGFR Tyrosine Kinase Inhibitors
7. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Kalluri, R.; Neilson, E.G. Epithelial-mesenchymal transition and its implications for fibrosis. J. Clin. Investig. 2003, 112, 1776–1784. [Google Scholar] [CrossRef] [PubMed]
- Kalluri, R.; Weinberg, R.A. The basics of epithelial-mesenchymal transition. J. Clin. Investig. 2009, 119, 1420–1428. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.M.; Dedhar, S.; Kalluri, R.; Thompson, E.W. The epithelial-mesenchymal transition: New insights in signaling, development, and disease. J. Cell Biol. 2006, 172, 973–981. [Google Scholar] [CrossRef] [PubMed]
- Nieto, M.A.; Huang, R.Y.; Jackson, R.A.; Thiery, J.P. Emt: 2016. Cell 2016, 166, 21–45. [Google Scholar] [CrossRef] [PubMed]
- Eger, A.; Aigner, K.; Sonderegger, S.; Dampier, B.; Oehler, S.; Schreiber, M.; Berx, G.; Cano, A.; Beug, H.; Foisner, R. Deltaef1 is a transcriptional repressor of e-cadherin and regulates epithelial plasticity in breast cancer cells. Oncogene 2005, 24, 2375–2385. [Google Scholar] [CrossRef] [PubMed]
- Batlle, E.; Sancho, E.; Franci, C.; Dominguez, D.; Monfar, M.; Baulida, J.; Garcia De Herreros, A. The transcription factor snail is a repressor of e-cadherin gene expression in epithelial tumour cells. Nat. Cell Biol. 2000, 2, 84–89. [Google Scholar] [CrossRef] [PubMed]
- Hajra, K.M.; Chen, D.Y.; Fearon, E.R. The SLUG zinc-finger protein represses E-cadherin in breast cancer. Cancer Res. 2002, 62, 1613–1618. [Google Scholar] [PubMed]
- Comijn, J.; Berx, G.; Vermassen, P.; Verschueren, K.; van Grunsven, L.; Bruyneel, E.; Mareel, M.; Huylebroeck, D.; van Roy, F. The two-handed E box binding zinc finger protein SIP1 downregulates E-cadherin and induces invasion. Mol. Cell 2001, 7, 1267–1278. [Google Scholar] [CrossRef]
- Kang, Y.; Massague, J. Epithelial-mesenchymal transitions: Twist in development and metastasis. Cell 2004, 118, 277–279. [Google Scholar] [CrossRef] [PubMed]
- Thiery, J.P. Epithelial-mesenchymal transitions in tumour progression. Nat. Rev. Cancer 2002, 2, 442–454. [Google Scholar] [CrossRef] [PubMed]
- Onder, T.T.; Gupta, P.B.; Mani, S.A.; Yang, J.; Lander, E.S.; Weinberg, R.A. Loss of E-cadherin promotes metastasis via multiple downstream transcriptional pathways. Cancer Res. 2008, 68, 3645–3654. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Weinberg, R.A. Epithelial-mesenchymal transition: At the crossroads of development and tumor metastasis. Dev. Cell 2008, 14, 818–829. [Google Scholar] [CrossRef] [PubMed]
- Cavallaro, U.; Christofori, G. Cell adhesion and signalling by cadherins and Ig-CAMs in cancer. Nat. Rev. Cancer 2004, 4, 118–132. [Google Scholar] [CrossRef] [PubMed]
- Tran, N.L.; Nagle, R.B.; Cress, A.E.; Heimark, R.L. N-cadherin expression in human prostate carcinoma cell lines. An epithelial-mesenchymal transformation mediating adhesion withstromal cells. Am. J. Pathol. 1999, 155, 787–798. [Google Scholar] [CrossRef]
- Zhou, B.; Liu, Y.; Kahn, M.; Ann, D.K.; Han, A.; Wang, H.; Nguyen, C.; Flodby, P.; Zhong, Q.; Krishnaveni, M.S.; et al. Interactions between beta-catenin and transforming growth factor-beta signaling pathways mediate epithelial-mesenchymal transition and are dependent on the transcriptional co-activator camp-response element-binding protein (Creb)-binding protein (Cbp). J. Biol. Chem. 2012, 287, 7026–7038. [Google Scholar] [CrossRef] [PubMed]
- Tse, J.C.; Kalluri, R. Mechanisms of metastasis: Epithelial-to-mesenchymal transition and contribution of tumor microenvironment. J. Cell Biochem. 2007, 101, 816–829. [Google Scholar] [CrossRef] [PubMed]
- Tam, W.L.; Weinberg, R.A. The epigenetics of epithelial-mesenchymal plasticity in cancer. Nat. Med. 2013, 19, 1438–1449. [Google Scholar] [CrossRef] [PubMed]
- Kung, J.T.; Colognori, D.; Lee, J.T. Long noncoding RNAs: Past, present, and future. Genetics 2013, 193, 651–669. [Google Scholar] [CrossRef] [PubMed]
- Kapranov, P.; Cheng, J.; Dike, S.; Nix, D.A.; Duttagupta, R.; Willingham, A.T.; Stadler, P.F.; Hertel, J.; Hackermuller, J.; Hofacker, I.L.; et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science 2007, 316, 1484–1488. [Google Scholar] [CrossRef] [PubMed]
- Eddy, S.R. Non-coding RNA genes and the modern RNA world. Nat. Rev. Genet. 2001, 2, 919–929. [Google Scholar] [CrossRef] [PubMed]
- Stefani, G.; Slack, F.J. Small non-coding RNAs in animal development. Nat. Rev. Mol. Cell Biol. 2008, 9, 219–230. [Google Scholar] [CrossRef] [PubMed]
- Ponting, C.P.; Oliver, P.L.; Reik, W. Evolution and functions of long noncoding RNAs. Cell 2009, 136, 629–641. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; Li, A.; Zou, D.; Xu, X.; Xia, L.; Yu, J.; Bajic, V.B.; Zhang, Z. Lncrnawiki: Harnessing community knowledge in collaborative curation of human long non-coding RNAs. Nucleic Acids Res. 2015, 43, D187–D192. [Google Scholar] [CrossRef] [PubMed]
- Fatica, A.; Bozzoni, I. Long non-coding RNAs: New players in cell differentiation and development. Nat. Rev. Genet. 2014, 15, 7–21. [Google Scholar] [CrossRef] [PubMed]
- Flynn, R.A.; Chang, H.Y. Long noncoding RNAs in cell-fate programming and reprogramming. Cell Stem Cell 2014, 14, 752–761. [Google Scholar] [CrossRef] [PubMed]
- Grote, P.; Wittler, L.; Hendrix, D.; Koch, F.; Wahrisch, S.; Beisaw, A.; Macura, K.; Blass, G.; Kellis, M.; Werber, M.; et al. The tissue-specific lncRNA Fendrr is an essential regulator of heart and body wall development in the mouse. Dev. Cell 2013, 24, 206–214. [Google Scholar] [CrossRef] [PubMed]
- Mercer, T.R.; Dinger, M.E.; Mattick, J.S. Long non-coding RNAs: Insights into functions. Nat. Rev. Genet. 2009, 10, 155–159. [Google Scholar] [CrossRef] [PubMed]
- Nagano, T.; Fraser, P. No-nonsense functions for long noncoding RNAs. Cell 2011, 145, 178–181. [Google Scholar] [CrossRef] [PubMed]
- Rinn, J.L.; Chang, H.Y. Genome regulation by long noncoding RNAs. Annu. Rev. Biochem. 2012, 81, 145–166. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.T. Epigenetic regulation by long noncoding RNAs. Science 2012, 338, 1435–1439. [Google Scholar] [CrossRef] [PubMed]
- Gibb, E.A.; Brown, C.J.; Lam, W.L. The functional role of long non-coding RNA in human carcinomas. Mol. Cancer 2011, 10, 38. [Google Scholar] [CrossRef] [PubMed]
- Prensner, J.R.; Chinnaiyan, A.M. The emergence of lncRNAs in cancer biology. Cancer Discov. 2011, 1, 391–407. [Google Scholar] [CrossRef] [PubMed]
- Ning, S.; Zhang, J.; Wang, P.; Zhi, H.; Wang, J.; Liu, Y.; Gao, Y.; Guo, M.; Yue, M.; Wang, L.; et al. Lnc2cancer: A manually curated database of experimentally supported lncRNAs associated with various human cancers. Nucleic Acids Res. 2016, 44, D980–D985. [Google Scholar] [CrossRef] [PubMed]
- Gutschner, T.; Diederichs, S. The hallmarks of cancer: A long non-coding RNA point of view. RNA Biol. 2012, 9, 703–719. [Google Scholar] [CrossRef] [PubMed]
- Salmena, L.; Poliseno, L.; Tay, Y.; Kats, L.; Pandolfi, P.P. A ceRNA hypothesis: The rosetta stone of a hidden RNA language? Cell 2011, 146, 353–358. [Google Scholar] [CrossRef] [PubMed]
- Cesana, M.; Cacchiarelli, D.; Legnini, I.; Santini, T.; Sthandier, O.; Chinappi, M.; Tramontano, A.; Bozzoni, I. A long noncoding RNA controls muscle differentiation by functioning as a competing endogenous RNA. Cell 2011, 147, 358–369. [Google Scholar] [CrossRef] [PubMed]
- Tay, Y.; Kats, L.; Salmena, L.; Weiss, D.; Tan, S.M.; Ala, U.; Karreth, F.; Poliseno, L.; Provero, P.; Di Cunto, F.; et al. Coding-independent regulation of the tumor suppressor PTEN by competing endogenous mRNAs. Cell 2011, 147, 344–357. [Google Scholar] [CrossRef] [PubMed]
- Friedman, R.C.; Farh, K.K.; Burge, C.B.; Bartel, D.P. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 2009, 19, 92–105. [Google Scholar] [CrossRef] [PubMed]
- Lewis, B.P.; Burge, C.B.; Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005, 120, 15–20. [Google Scholar] [CrossRef] [PubMed]
- Paraskevopoulou, M.D.; Georgakilas, G.; Kostoulas, N.; Reczko, M.; Maragkakis, M.; Dalamagas, T.M.; Hatzigeorgiou, A.G. Diana-lncbase: Experimentally verified and computationally predicted microRNA targets on long non-coding RNAs. Nucleic Acids Res. 2013, 41, D239–D245. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Target recognition and regulatory functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef] [PubMed]
- Calin, G.A.; Croce, C.M. MicroRNA signatures in human cancers. Nat. Rev. Cancer 2006, 6, 857–866. [Google Scholar] [CrossRef] [PubMed]
- Deng, S.; Calin, G.A.; Croce, C.M.; Coukos, G.; Zhang, L. Mechanisms of microRNA deregulation in human cancer. Cell Cycle 2008, 7, 2643–2646. [Google Scholar] [CrossRef] [PubMed]
- Visone, R.; Croce, C.M. miRNAs and cancer. Am. J. Pathol. 2009, 174, 1131–1138. [Google Scholar] [CrossRef] [PubMed]
- Bullock, M.D.; Sayan, A.E.; Packham, G.K.; Mirnezami, A.H. MicroRNAs: Critical regulators of epithelial to mesenchymal (EMT) and mesenchymal to epithelial transition (MET) in cancer progression. Biol. Cell 2012, 104, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Ding, X.M. MicroRNAs: Regulators of cancer metastasis and epithelial-mesenchymal transition (EMT). Chin. J. Cancer 2014, 33, 140–147. [Google Scholar] [CrossRef] [PubMed]
- Gregory, P.A.; Bert, A.G.; Paterson, E.L.; Barry, S.C.; Tsykin, A.; Farshid, G.; Vadas, M.A.; Khew-Goodall, Y.; Goodall, G.J. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat. Cell Biol. 2008, 10, 593–601. [Google Scholar] [CrossRef] [PubMed]
- Bracken, C.P.; Gregory, P.A.; Kolesnikoff, N.; Bert, A.G.; Wang, J.; Shannon, M.F.; Goodall, G.J. A double-negative feedback loop between ZEB1-SIP1 and the microRNA-200 family regulates epithelial-mesenchymal transition. Cancer Res. 2008, 68, 7846–7854. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.N.; Yin, J.J.; Abou-Kheir, W.; Hynes, P.G.; Casey, O.M.; Fang, L.; Yi, M.; Stephens, R.M.; Seng, V.; Sheppard-Tillman, H.; et al. MiR-1 and miR-200 inhibit EMT via slug-dependent and tumorigenesis via SLUG-independent mechanisms. Oncogene 2013, 32, 296–306. [Google Scholar] [CrossRef] [PubMed]
- Cicchini, C.; de Nonno, V.; Battistelli, C.; Cozzolino, A.M.; De Santis Puzzonia, M.; Ciafre, S.A.; Brocker, C.; Gonzalez, F.J.; Amicone, L.; Tripodi, M. Epigenetic control of EMT/MET dynamics: HNF4alpha impacts DNMT3S through miRs-29. Biochim. Biophys. Acta 2015, 1849, 919–929. [Google Scholar] [CrossRef] [PubMed]
- Kogure, T.; Kondo, Y.; Kakazu, E.; Ninomiya, M.; Kimura, O.; Shimosegawa, T. Involvement of miRNA-29a in epigenetic regulation of transforming growth factor-beta-induced epithelial-mesenchymal transition in hepatocellular carcinoma. Hepatol. Res. 2014, 44, 907–919. [Google Scholar] [CrossRef] [PubMed]
- Qiu, Y.H.; Wei, Y.P.; Shen, N.J.; Wang, Z.C.; Kan, T.; Yu, W.L.; Yi, B.; Zhang, Y.J. miR-204 inhibits epithelial to mesenchymal transition by targeting slug in intrahepatic cholangiocarcinoma cells. Cell Physiol. Biochem. 2013, 32, 1331–1341. [Google Scholar] [CrossRef] [PubMed]
- Ru, P.; Steele, R.; Newhall, P.; Phillips, N.J.; Toth, K.; Ray, R.B. miRNA-29b suppresses prostate cancer metastasis by regulating epithelial-mesenchymal transition signaling. Mol. Cancer Ther. 2012, 11, 1166–1173. [Google Scholar] [CrossRef] [PubMed]
- Kumarswamy, R.; Mudduluru, G.; Ceppi, P.; Muppala, S.; Kozlowski, M.; Niklinski, J.; Papotti, M.; Allgayer, H. MicroRNA-30a inhibits epithelial-to-mesenchymal transition by targeting snai1 and is downregulated in non-small cell lung cancer. Int. J. Cancer 2012, 130, 2044–2053. [Google Scholar] [CrossRef] [PubMed]
- Siemens, H.; Jackstadt, R.; Hunten, S.; Kaller, M.; Menssen, A.; Gotz, U.; Hermeking, H. miR-34 and SNAIL form a double-negative feedback loop to regulate epithelial-mesenchymal transitions. Cell Cycle 2011, 10, 4256–4271. [Google Scholar] [CrossRef] [PubMed]
- Moes, M.; Le Bechec, A.; Crespo, I.; Laurini, C.; Halavatyi, A.; Vetter, G.; Del Sol, A.; Friederich, E. A novel network integrating a miRNA-203/SNAI1 feedback loop which regulates epithelial to mesenchymal transition. PLoS ONE 2012, 7, e35440. [Google Scholar] [CrossRef] [PubMed]
- Ding, X.; Park, S.I.; McCauley, L.K.; Wang, C.Y. Signaling between transforming growth factor beta (TGF-beta) and transcription factor SNAI2 represses expression of microRNA miR-203 to promote epithelial-mesenchymal transition and tumor metastasis. J. Biol. Chem. 2013, 288, 10241–10253. [Google Scholar] [CrossRef] [PubMed]
- Yamasaki, T.; Seki, N.; Yamada, Y.; Yoshino, H.; Hidaka, H.; Chiyomaru, T.; Nohata, N.; Kinoshita, T.; Nakagawa, M.; Enokida, H. Tumor suppressive microRNA138 contributes to cell migration and invasion through its targeting of vimentin in renal cell carcinoma. Int. J. Oncol. 2012, 41, 805–817. [Google Scholar] [PubMed]
- Meng, Z.; Fu, X.; Chen, X.; Zeng, S.; Tian, Y.; Jove, R.; Xu, R.; Huang, W. Mir-194 is a marker of hepatic epithelial cells and suppresses metastasis of liver cancer cells in mice. Hepatology 2010, 52, 2148–2157. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, S.; Ueda, Y.; Akaboshi, S.; Hino, Y.; Sekita, Y.; Nakao, M. HMGA2 maintains oncogenic ras-induced epithelial-mesenchymal transition in human pancreatic cancer cells. Am. J. Pathol. 2009, 174, 854–868. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.S.; Dutta, A. The tumor suppressor microRNA let-7 represses the HMGA2 oncogene. Genes Dev. 2007, 21, 1025–1030. [Google Scholar] [CrossRef] [PubMed]
- Mayr, C.; Hemann, M.T.; Bartel, D.P. Disrupting the pairing between let-7 and HMGA2 enhances oncogenic transformation. Science 2007, 315, 1576–1579. [Google Scholar] [CrossRef] [PubMed]
- Thuault, S.; Valcourt, U.; Petersen, M.; Manfioletti, G.; Heldin, C.H.; Moustakas, A. Transforming growth factor-beta employs HMGA2 to elicit epithelial-mesenchymal transition. J. Cell Biol. 2006, 174, 175–183. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.; Veronese, A.; Pichiorri, F.; Lee, T.J.; Jeon, Y.J.; Volinia, S.; Pineau, P.; Marchio, A.; Palatini, J.; Suh, S.S.; et al. P53 regulates epithelial-mesenchymal transition through microRNAs targeting ZEB1 and ZEB2. J. Exp. Med. 2011, 208, 875–883. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.J.; Chao, C.H.; Xia, W.; Yang, J.Y.; Xiong, Y.; Li, C.W.; Yu, W.H.; Rehman, S.K.; Hsu, J.L.; Lee, H.H.; et al. p53 regulates epithelial-mesenchymal transition and stem cell properties through modulating miRNAs. Nat. Cell Biol. 2011, 13, 317–323. [Google Scholar] [CrossRef] [PubMed]
- Kim, N.H.; Kim, H.S.; Li, X.Y.; Lee, I.; Choi, H.S.; Kang, S.E.; Cha, S.Y.; Ryu, J.K.; Yoon, D.; Fearon, E.R.; et al. A p53/miRNA-34 axis regulates SNAIL1-dependent cancer cell epithelial-mesenchymal transition. J. Cell Biol. 2011, 195, 417–433. [Google Scholar] [CrossRef] [PubMed]
- Volinia, S.; Calin, G.A.; Liu, C.G.; Ambs, S.; Cimmino, A.; Petrocca, F.; Visone, R.; Iorio, M.; Roldo, C.; Ferracin, M.; et al. A microRNA expression signature of human solid tumors defines cancer gene targets. Proc. Natl. Acad. Sci. USA 2006, 103, 2257–2261. [Google Scholar] [CrossRef] [PubMed]
- Zavadil, J.; Narasimhan, M.; Blumenberg, M.; Schneider, R.J. Transforming growth factor-beta and microRNA:mRNA regulatory networks in epithelial plasticity. Cells Tissues Organs 2007, 185, 157–161. [Google Scholar] [CrossRef] [PubMed]
- Cottonham, C.L.; Kaneko, S.; Xu, L. MiR-21 and miR-31 converge on TIAM1 to regulate migration and invasion of colon carcinoma cells. J. Biol. Chem. 2010, 285, 35293–35302. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.H.; Huang, Q.; Jin, Z.Y.; Zhu, C.L.; Liu, Z.; Wang, C. Mir-21 and klf4 jointly augment epithelialmesenchymal transition via the AKT/ERK1/2 pathway. Int. J. Oncol. 2017, 50, 1109–1115. [Google Scholar] [PubMed]
- Han, M.; Wang, Y.; Liu, M.; Bi, X.; Bao, J.; Zeng, N.; Zhu, Z.; Mo, Z.; Wu, C.; Chen, X. Mir-21 regulates epithelial-mesenchymal transition phenotype and hypoxia-inducible factor-1alpha expression in third-sphere forming breast cancer stem cell-like cells. Cancer Sci. 2012, 103, 1058–1064. [Google Scholar] [CrossRef] [PubMed]
- Han, M.; Liu, M.; Wang, Y.; Chen, X.; Xu, J.; Sun, Y.; Zhao, L.; Qu, H.; Fan, Y.; Wu, C. Antagonism of miR-21 reverses epithelial-mesenchymal transition and cancer stem cell phenotype through AKT/ERK1/2 inactivation by targeting PTEN. PLoS ONE 2012, 7, e39520. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; Young, J.; Prabhala, H.; Pan, E.; Mestdagh, P.; Muth, D.; Teruya-Feldstein, J.; Reinhardt, F.; Onder, T.T.; Valastyan, S.; et al. miR-9, a MYC/MYCN-activated microRNA, regulates e-cadherin and cancer metastasis. Nat. Cell Biol. 2010, 12, 247–256. [Google Scholar] [CrossRef] [PubMed]
- Kong, W.; Yang, H.; He, L.; Zhao, J.J.; Coppola, D.; Dalton, W.S.; Cheng, J.Q. MicroRNA-155 is regulated by the transforming growth factor beta/SMAD pathway and contributes to epithelial cell plasticity by targeting RHOA. Mol. Cell Biol. 2008, 28, 6773–6784. [Google Scholar] [CrossRef] [PubMed]
- Margueron, R.; Reinberg, D. The polycomb complex PRC2 and its mark in life. Nature 2011, 469, 343–349. [Google Scholar] [CrossRef] [PubMed]
- Simon, J.A.; Kingston, R.E. Mechanisms of polycomb gene silencing: Knowns and unknowns. Nat. Rev. Mol. Cell Biol. 2009, 10, 697–708. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Ohsumi, T.K.; Kung, J.T.; Ogawa, Y.; Grau, D.J.; Sarma, K.; Song, J.J.; Kingston, R.E.; Borowsky, M.; Lee, J.T. Genome-wide identification of polycomb-associated RNAs by rip-seq. Mol. Cell 2010, 40, 939–953. [Google Scholar] [CrossRef] [PubMed]
- Kaneko, S.; Bonasio, R.; Saldana-Meyer, R.; Yoshida, T.; Son, J.; Nishino, K.; Umezawa, A.; Reinberg, D. Interactions between JARID2 and noncoding RNAs regulate PRC2 recruitment to chromatin. Mol. Cell 2014, 53, 290–300. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Sun, B.K.; Erwin, J.A.; Song, J.J.; Lee, J.T. Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science 2008, 322, 750–756. [Google Scholar] [CrossRef] [PubMed]
- Plath, K.; Fang, J.; Mlynarczyk-Evans, S.K.; Cao, R.; Worringer, K.A.; Wang, H.; de la Cruz, C.C.; Otte, A.P.; Panning, B.; Zhang, Y. Role of histone h3 lysine 27 methylation in x inactivation. Science 2003, 300, 131–135. [Google Scholar] [CrossRef] [PubMed]
- Gupta, R.A.; Shah, N.; Wang, K.C.; Kim, J.; Horlings, H.M.; Wong, D.J.; Tsai, M.C.; Hung, T.; Argani, P.; Rinn, J.L.; et al. Long non-coding RNA hotair reprograms chromatin state to promote cancer metastasis. Nature 2010, 464, 1071–1076. [Google Scholar] [CrossRef] [PubMed]
- Kotake, Y.; Nakagawa, T.; Kitagawa, K.; Suzuki, S.; Liu, N.; Kitagawa, M.; Xiong, Y. Long non-coding RNA anril is required for the PRC2 recruitment to and silencing of p15(INK4B) tumor suppressor gene. Oncogene 2011, 30, 1956–1962. [Google Scholar] [CrossRef] [PubMed]
- Beckedorff, F.C.; Ayupe, A.C.; Crocci-Souza, R.; Amaral, M.S.; Nakaya, H.I.; Soltys, D.T.; Menck, C.F.; Reis, E.M.; Verjovski-Almeida, S. The intronic long noncoding RNA ANRASSF1 recruits PRC2 to the RASSF1a promoter, reducing the expression of rassf1a and increasing cell proliferation. PLoS Genet. 2013, 9, e1003705. [Google Scholar] [CrossRef] [PubMed]
- Cao, Q.; Yu, J.; Dhanasekaran, S.M.; Kim, J.H.; Mani, R.S.; Tomlins, S.A.; Mehra, R.; Laxman, B.; Cao, X.; Kleer, C.G.; et al. Repression of E-cadherin by the polycomb group protein EZH2 in cancer. Oncogene 2008, 27, 7274–7284. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Liu, X.; Chen, Z.; Huang, H.; Jin, Y.; Kolokythas, A.; Wang, A.; Dai, Y.; Wong, D.T.; Zhou, X. Polycomb group protein EZH2-mediated E-cadherin repression promotes metastasis of oral tongue squamous cell carcinoma. Mol. Carcinog. 2013, 52, 229–236. [Google Scholar] [CrossRef] [PubMed]
- Luo, H.; Jiang, Y.; Ma, S.; Chang, H.; Yi, C.; Cao, H.; Gao, Y.; Guo, H.; Hou, J.; Yan, J.; et al. Ezh2 promotes invasion and metastasis of laryngeal squamous cells carcinoma via epithelial-mesenchymal transition through H3K27me3. Biochem. Biophys. Res. Commun. 2016, 479, 253–259. [Google Scholar] [CrossRef] [PubMed]
- Herranz, N.; Pasini, D.; Diaz, V.M.; Franci, C.; Gutierrez, A.; Dave, N.; Escriva, M.; Hernandez-Munoz, I.; Di Croce, L.; Helin, K.; et al. Polycomb complex 2 is required for E-cadherin repression by the SNAIL1 transcription factor. Mol. Cell Biol. 2008, 28, 4772–4781. [Google Scholar] [CrossRef] [PubMed]
- Chou, J.; Wang, B.; Zheng, T.; Li, X.; Zheng, L.; Hu, J.; Zhang, Y.; Xing, Y.; Xi, T. MALAT1 induced migration and invasion of human breast cancer cells by competitively binding miR-1 with CDC42. Biochem. Biophys. Res. Commun. 2016, 472, 262–269. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Wang, J.; Chen, Y.; Li, S.; Jin, M.; Wang, H.; Chen, Z.; Yu, W. LncRNA MALAT1 exerts oncogenic functions in lung adenocarcinoma by targeting miR-204. Am. J. Cancer Res. 2016, 6, 1099–1107. [Google Scholar] [PubMed]
- Hirata, H.; Hinoda, Y.; Shahryari, V.; Deng, G.; Nakajima, K.; Tabatabai, Z.L.; Ishii, N.; Dahiya, R. Long noncoding RNA MALAT1 promotes aggressive renal cell carcinoma through EZH2 and interacts with miR-205. Cancer Res. 2015, 75, 1322–1331. [Google Scholar] [CrossRef] [PubMed]
- Jiao, F.; Hu, H.; Han, T.; Yuan, C.; Wang, L.; Jin, Z.; Guo, Z. Long noncoding RNA MALAT-1 enhances stem cell-like phenotypes in pancreatic cancer cells. Int. J. Mol. Sci. 2015, 16, 6677–6693. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Shen, B.; Tan, M.; Mu, X.; Qin, Y.; Zhang, F.; Liu, Y. TGF-beta-induced upregulation of MALAT1 promotes bladder cancer metastasis by associating with SUZ12. Clin. Cancer Res. 2014, 20, 1531–1541. [Google Scholar] [CrossRef] [PubMed]
- Liang, J.; Liang, L.; Ouyang, K.; Li, Z.; Yi, X. MALAT1 induces tongue cancer cells’ EMT and inhibits apoptosis through Wnt/beta-catenin signaling pathway. J. Oral Pathol. Med. 2017, 46, 98–105. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.S.; Wang, X.A.; Wu, W.G.; Hu, Y.P.; Li, M.L.; Ding, Q.; Weng, H.; Shu, Y.J.; Liu, T.Y.; Jiang, L.; et al. Malat1 promotes the proliferation and metastasis of gallbladder cancer cells by activating the ERK/MAPK pathway. Cancer Biol. Ther. 2014, 15, 806–814. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Liang, G.; Yuan, B.; Yang, C.; Gao, R.; Zhou, X. MALAT1 promotes the proliferation and metastasis of osteosarcoma cells by activating the PI3K/Akt pathway. Tumour Biol. 2015, 36, 1477–1486. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.; Sui, S.; Zhang, J.; Bai, N.; Shi, Q.; Zhang, G.; Gao, S.; You, Z.; Zhan, C.; Liu, F.; et al. Downregulation of long noncoding RNA MALAT1 induces epithelial-to-mesenchymal transition via the PI3K-Akt pathway in breast cancer. Int. J. Clin. Exp. Pathol. 2015, 8, 4881–4891. [Google Scholar] [PubMed]
- Tripathi, V.; Shen, Z.; Chakraborty, A.; Giri, S.; Freier, S.M.; Wu, X.; Zhang, Y.; Gorospe, M.; Prasanth, S.G.; Lal, A.; et al. Long noncoding RNA MALAT1 controls cell cycle progression by regulating the expression of oncogenic transcription factor b-MYB. PLoS Genet. 2013, 9, e1003368. [Google Scholar] [CrossRef] [PubMed]
- Huang, N.S.; Chi, Y.Y.; Xue, J.Y.; Liu, M.Y.; Huang, S.; Mo, M.; Zhou, S.L.; Wu, J. Long non-coding RNA metastasis associated in lung adenocarcinoma transcript 1 (malat1) interacts with estrogen receptor and predicted poor survival in breast cancer. Oncotarget 2016, 7, 37957–37965. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Tang, X.; Shi, M.; Wen, C.; Shen, B. Mir-216a decreases malat1 expression, induces g2/m arrest and apoptosis in pancreatic cancer cells. Biochem. Biophys. Res. Commun. 2017, 483, 816–822. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.; Nong, K.; Zhu, H.; Wang, W.; Huang, X.; Yuan, Z.; Ai, K. H19 promotes pancreatic cancer metastasis by derepressing let-7’s suppression on its target HMGA2-mediated EMT. Tumour Biol. 2014, 35, 9163–9169. [Google Scholar] [CrossRef] [PubMed]
- Liang, W.C.; Fu, W.M.; Wong, C.W.; Wang, Y.; Wang, W.M.; Hu, G.X.; Zhang, L.; Xiao, L.J.; Wan, D.C.; Zhang, J.F.; et al. The lncRNA h19 promotes epithelial to mesenchymal transition by functioning as miRNA sponges in colorectal cancer. Oncotarget 2015, 6, 22513–22525. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Ye, F.; Yin, C.; Zhuang, Y.; Yue, G.; Zhang, G. The interaction between miR-141 and lncRNA-h19 in regulating cell proliferation and migration in gastric cancer. Cell Physiol. Biochem. 2015, 36, 1440–1452. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Lin, Y.; Yang, X.; Wu, X.; He, X. Long noncoding RNA h19 regulates EZH2 expression by interacting with miR-630 and promotes cell invasion in nasopharyngeal carcinoma. Biochem. Biophys. Res. Commun. 2016, 473, 913–919. [Google Scholar] [CrossRef] [PubMed]
- Luo, M.; Li, Z.; Wang, W.; Zeng, Y.; Liu, Z.; Qiu, J. Long non-coding RNA h19 increases bladder cancer metastasis by associating with EZH2 and inhibiting E-cadherin expression. Cancer Lett. 2013, 333, 213–221. [Google Scholar] [CrossRef] [PubMed]
- Matouk, I.J.; Halle, D.; Raveh, E.; Gilon, M.; Sorin, V.; Hochberg, A. The role of the oncofetal h19 lncRNA in tumor metastasis: Orchestrating the EMT-MET decision. Oncotarget 2016, 7, 3748–3765. [Google Scholar] [PubMed]
- Matouk, I.J.; Mezan, S.; Mizrahi, A.; Ohana, P.; Abu-Lail, R.; Fellig, Y.; Degroot, N.; Galun, E.; Hochberg, A. The oncofetal h19 RNA connection: Hypoxia, p53 and cancer. Biochim. Biophys. Acta 2010, 1803, 443–451. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Z.G.; Xu, H.; Suo, S.S.; Xu, X.L.; Ni, M.W.; Gu, L.H.; Chen, W.; Wang, L.Y.; Zhao, Y.; Tian, B.; et al. The essential role of h19 contributing to cisplatin resistance by regulating glutathione metabolism in high-grade serous ovarian cancer. Sci. Rep. 2016, 6, 26093. [Google Scholar] [CrossRef] [PubMed]
- Tsang, W.P.; Kwok, T.T. Riboregulator h19 induction of MDR1-associated drug resistance in human hepatocellular carcinoma cells. Oncogene 2007, 26, 4877–4881. [Google Scholar] [CrossRef] [PubMed]
- Jiang, P.; Wang, P.; Sun, X.; Yuan, Z.; Zhan, R.; Ma, X.; Li, W. Knockdown of long noncoding RNA h19 sensitizes human glioma cells to temozolomide therapy. Onco Targets Ther. 2016, 9, 3501–3509. [Google Scholar] [PubMed]
- Jiang, X.; Yan, Y.; Hu, M.; Chen, X.; Wang, Y.; Dai, Y.; Wu, D.; Zhuang, Z.; Xia, H. Increased level of h19 long noncoding RNA promotes invasion, angiogenesis, and stemness of glioblastoma cells. J. Neurosurg. 2016, 124, 129–136. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Jiang, P.; Sun, X.; Xu, S.; Ma, X.; Zhan, R. Suppressing h19 modulates tumorigenicity and stemness in u251 and u87mg glioma cells. Cell Mol. Neurobiol. 2016, 36, 1219–1227. [Google Scholar] [CrossRef] [PubMed]
- Zeira, E.; Abramovitch, R.; Meir, K.; Even Ram, S.; Gil, Y.; Bulvik, B.; Bromberg, Z.; Levkovitch, O.; Nahmansson, N.; Adar, R.; et al. The knockdown of h19lncRNA reveals its regulatory role in pluripotency and tumorigenesis of human embryonic carcinoma cells. Oncotarget 2015, 6, 34691–34703. [Google Scholar] [PubMed]
- Uehara, T.; Ainslie, G.R.; Kutanzi, K.; Pogribny, I.P.; Muskhelishvili, L.; Izawa, T.; Yamate, J.; Kosyk, O.; Shymonyak, S.; Bradford, B.U.; et al. Molecular mechanisms of fibrosis-associated promotion of liver carcinogenesis. Toxicol. Sci. 2013, 132, 53–63. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Zhang, L.; Wang, Y.; Li, H.; Ren, X.; Wei, F.; Yu, W.; Liu, T.; Wang, X.; Zhou, X.; et al. Long non-coding RNA hotair promotes tumor cell invasion and metastasis by recruiting EZH2 and repressing e-cadherin in oral squamous cell carcinoma. Int. J. Oncol. 2015, 46, 2586–2594. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.W.; Sun, M.; Xia, R.; Zhang, E.B.; Liu, X.H.; Zhang, Z.H.; Xu, T.P.; De, W.; Liu, B.R.; Wang, Z.X. Linchotair epigenetically silences miR34a by binding to PRC2 to promote the epithelial-to-mesenchymal transition in human gastric cancer. Cell Death Dis. 2015, 6, e1802. [Google Scholar] [CrossRef] [PubMed]
- Li, J.T.; Wang, L.F.; Zhao, Y.L.; Yang, T.; Li, W.; Zhao, J.; Yu, F.; Wang, L.; Meng, Y.L.; Liu, N.N.; et al. Nuclear factor of activated t cells 5 maintained by hotair suppression of mir-568 upregulates s100 calcium binding protein A4 to promote breast cancer metastasis. Breast Cancer Res. 2014, 16, 454. [Google Scholar] [CrossRef] [PubMed]
- Ge, X.S.; Ma, H.J.; Zheng, X.H.; Ruan, H.L.; Liao, X.Y.; Xue, W.Q.; Chen, Y.B.; Zhang, Y.; Jia, W.H. Hotair, a prognostic factor in esophageal squamous cell carcinoma, inhibits WIF-1 expression and activates wnt pathway. Cancer Sci. 2013, 104, 1675–1682. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Cai, K.; Wang, J.; Wang, X.; Cheng, K.; Shi, F.; Jiang, L.; Zhang, Y.; Dou, J. miR-7, inhibited indirectly by lincRNA hotair, directly inhibits SETDB1 and reverses the emt of breast cancer stem cells by downregulating the STAT3 pathway. Stem Cells 2014, 32, 2858–2868. [Google Scholar] [CrossRef] [PubMed]
- Battistelli, C.; Cicchini, C.; Santangelo, L.; Tramontano, A.; Grassi, L.; Gonzalez, F.J.; de Nonno, V.; Grassi, G.; Amicone, L.; Tripodi, M. The SNAIL repressor recruits EZH2 to specific genomic sites through the enrollment of the lncRNA hotair in epithelial-to-mesenchymal transition. Oncogene 2017, 36, 942–955. [Google Scholar] [CrossRef] [PubMed]
- Pádua Alves, C.; Fonseca, A.S.; Muys, B.R.; de Barros, E.L.B.R.; Burger, M.C.; de Souza, J.E.; Valente, V.; Zago, M.A.; Silva, W.A., Jr. Brief report: The lincRNA hotair is required for epithelial-to-mesenchymal transition and stemness maintenance of cancer cell lines. Stem Cells 2013, 31, 2827–2832. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Yang, S.; Su, N.; Wang, Y.; Yu, J.; Qiu, H.; He, X. Overexpression of long non-coding RNA hotair leads to chemoresistance by activating the Wnt/beta-catenin pathway in human ovarian cancer. Tumour Biol. 2016, 37, 2057–2065. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Sun, M.; Lu, K.; Liu, J.; Zhang, M.; Wu, W.; De, W.; Wang, Z.; Wang, R. The long noncoding RNA hotair contributes to cisplatin resistance of human lung adenocarcinoma cells via downregualtion of p21(WAF1/CIP1) expression. PLoS ONE 2013, 8, e77293. [Google Scholar]
- Wang, Y.; Wang, H.; Song, T.; Zou, Y.; Jiang, J.; Fang, L.; Li, P. Hotair is a potential target for the treatment of cisplatinresistant ovarian cancer. Mol. Med. Rep. 2015, 12, 2211–2216. [Google Scholar] [PubMed]
- Teschendorff, A.E.; Lee, S.H.; Jones, A.; Fiegl, H.; Kalwa, M.; Wagner, W.; Chindera, K.; Evans, I.; Dubeau, L.; Orjalo, A.; et al. Hotair and its surrogate DNA methylation signature indicate carboplatin resistance in ovarian cancer. Genome Med. 2015, 7, 108. [Google Scholar] [CrossRef] [PubMed]
- Deng, J.; Yang, M.; Jiang, R.; An, N.; Wang, X.; Liu, B. Long non-coding RNA hotair regulates the proliferation, self-renewal capacity, tumor formation and migration of the cancer stem-like cell (CSC) subpopulation enriched from breast cancer cells. PLoS ONE 2017, 12, e0170860. [Google Scholar] [CrossRef] [PubMed]
- Dou, J.; Ni, Y.; He, X.; Wu, D.; Li, M.; Wu, S.; Zhang, R.; Guo, M.; Zhao, F. Decreasing lncRNA hotair expression inhibits human colorectal cancer stem cells. Am. J. Transl. Res. 2016, 8, 98–108. [Google Scholar] [PubMed]
- Li, H.; An, J.; Wu, M.; Zheng, Q.; Gui, X.; Li, T.; Pu, H.; Lu, D. LncRNA hotair promotes human liver cancer stem cell malignant growth through downregulation of SETD2. Oncotarget 2015, 6, 27847–27864. [Google Scholar] [PubMed]
- Ye, P.; Wang, T.; Liu, W.H.; Li, X.C.; Tang, L.J.; Tian, F.Z. Enhancing hotair/miR-10b drives normal liver stem cells toward a tendency to malignant transformation through inducing epithelial- to-mesenchymal transition. Rejuv. Res. 2015, 18, 332–340. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Luo, F.; Xu, Y.; Wang, B.; Zhao, Y.; Xu, W.; Shi, L.; Lu, X.; Liu, Q. Epithelial-mesenchymal transition and cancer stem cells, mediated by a long non-coding RNA, hotair, are involved in cell malignant transformation induced by cigarette smoke extract. Toxicol. Appl. Pharmacol. 2015, 282, 9–19. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Chen, D.; He, X.; Zhang, Y.; Shi, F.; Wu, D.; Chen, J.; Zhao, F.; Dou, J. Downregulated lincRNA hotair expression in ovarian cancer stem cells decreases its tumorgeniesis and metastasis by inhibiting epithelial-mesenchymal transition. Cancer Cell Int. 2015, 15, 24. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Gao, Q.; Wang, J.; Zhang, X.; Liu, K.; Duan, Z. Linc-RNA-ROR acts as a “sponge” against mediation of the differentiation of endometrial cancer stem cells by microRNA-145. Gynecol. Oncol. 2014, 133, 333–339. [Google Scholar] [CrossRef] [PubMed]
- Hou, P.; Zhao, Y.; Li, Z.; Yao, R.; Ma, M.; Gao, Y.; Zhao, L.; Zhang, Y.; Huang, B.; Lu, J. LincRNA-ror induces epithelial-to-mesenchymal transition and contributes to breast cancer tumorigenesis and metastasis. Cell Death Dis. 2014, 5, e1287. [Google Scholar] [CrossRef] [PubMed]
- Zhang, A.; Zhou, N.; Huang, J.; Liu, Q.; Fukuda, K.; Ma, D.; Lu, Z.; Bai, C.; Watabe, K.; Mo, Y.Y. The human long non-coding RNA-ROR is a p53 repressor in response to DNA damage. Cell Res. 2013, 23, 340–350. [Google Scholar] [CrossRef] [PubMed]
- Zhan, H.X.; Wang, Y.; Li, C.; Xu, J.W.; Zhou, B.; Zhu, J.K.; Han, H.F.; Wang, L.; Wang, Y.S.; Hu, S.Y. LincRNA-ROR promotes invasion, metastasis and tumor growth in pancreatic cancer through activating ZEB1 pathway. Cancer Lett. 2016, 374, 261–271. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Yan, I.K.; Haga, H.; Patel, T. Modulation of hypoxia-signaling pathways by extracellular linc-ROR. J. Cell Sci. 2014, 127, 1585–1594. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Zhao, Z.; Zhou, Z.; Liu, R. Linc-ROR confers gemcitabine resistance to pancreatic cancer cells via inducing autophagy and modulating the miR-124/ptbp1/PKM2 axis. Cancer Chemother. Pharmacol. 2016, 78, 1199–1207. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.M.; Liu, Y.; Wei, H.Y.; Lv, K.Z.; Fu, P.F. Large intergenic non-coding RNA-ROR reverses gemcitabine-induced autophagy and apoptosis in breast cancer cells. Oncotarget 2016, 7, 59604–59617. [Google Scholar] [CrossRef] [PubMed]
- Rezaei, M.; Emadi-Baygi, M.; Hoffmann, M.J.; Schulz, W.A.; Nikpour, P. Altered expression of linc-ROR in cancer cell lines and tissues. Tumour Biol. 2016, 37, 1763–1769. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; He, L.; Du, Y.; Zhu, P.; Huang, G.; Luo, J.; Yan, X.; Ye, B.; Li, C.; Xia, P.; et al. The long noncoding RNA lncTCF7 promotes self-renewal of human liver cancer stem cells through activation of Wnt signaling. Cell Stem Cell 2015, 16, 413–425. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Zhang, J.; Shen, B.; Yin, K.; Xu, J.; Gao, W.; Zhang, L. Long noncoding RNA lncTCF7, induced by IL-6/STAT3 transactivation, promotes hepatocellular carcinoma aggressiveness through epithelial-mesenchymal transition. J. Exp. Clin. Cancer Res. 2015, 34, 116. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Wang, D. Long noncoding RNA TCF7 promotes invasiveness and self-renewal of human non-small cell lung cancer cells. Hum. Cell 2017, 30, 23–29. [Google Scholar] [CrossRef] [PubMed]
- Xia, T.; Liao, Q.; Jiang, X.; Shao, Y.; Xiao, B.; Xi, Y.; Guo, J. Long noncoding RNA associated-competing endogenous RNAs in gastric cancer. Sci. Rep. 2014, 4, 6088. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.N.; Chen, X.; Chen, Z.Y.; Nie, F.Q.; Wei, C.C.; Ma, H.W.; Wan, L.; Yan, S.; Ren, S.N.; Wang, Z.X. Long intergenic non-coding RNA 00152 promotes lung adenocarcinoma proliferation via interacting with EZH2 and repressing IL24 expression. Mol. Cancer 2017, 16, 17. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.M.; Huang, M.D.; Sun, D.P.; Kong, R.; Xu, T.P.; Xia, R.; Zhang, E.B.; Shu, Y.Q. Long intergenic non-coding RNA 00152 promotes tumor cell cycle progression by binding to EZH2 and repressing p15 and p21 in gastric cancer. Oncotarget 2016, 7, 9773–9787. [Google Scholar] [PubMed]
- Ji, J.; Tang, J.; Deng, L.; Xie, Y.; Jiang, R.; Li, G.; Sun, B. Linc00152 promotes proliferation in hepatocellular carcinoma by targeting epcam via the mtor signaling pathway. Oncotarget 2015, 6, 42813–42824. [Google Scholar] [PubMed]
- Zhou, J.; Zhi, X.; Wang, L.; Wang, W.; Li, Z.; Tang, J.; Wang, J.; Zhang, Q.; Xu, Z. Linc00152 promotes proliferation in gastric cancer through the EGFR-dependent pathway. J. Exp. Clin. Cancer Res. 2015, 34, 135. [Google Scholar] [CrossRef] [PubMed]
- Cai, Q.; Wang, Z.; Wang, S.; Weng, M.; Zhou, D.; Li, C.; Wang, J.; Chen, E.; Quan, Z. Long non-coding RNA linc00152 promotes gallbladder cancer metastasis and epithelial-mesenchymal transition by regulating hif-1alpha via miR-138. Open Biol. 2017, 7, 160247. [Google Scholar] [CrossRef] [PubMed]
- Cai, Q.; Wang, Z.Q.; Wang, S.H.; Li, C.; Zhu, Z.G.; Quan, Z.W.; Zhang, W.J. Upregulation of long non-coding RNA linc00152 by sp1 contributes to gallbladder cancer cell growth and tumor metastasis via PI3K/Akt pathway. Am. J. Transl. Res. 2016, 8, 4068–4081. [Google Scholar] [PubMed]
- Tani, H.; Torimura, M. Identification of short-lived long non-coding RNAs as surrogate indicators for chemical stress response. Biochem. Biophys. Res. Commun. 2013, 439, 547–551. [Google Scholar] [CrossRef] [PubMed]
- Yue, B.; Cai, D.; Liu, C.; Fang, C.; Yan, D. Linc00152 functions as a competing endogenous RNA to confer oxaliplatin resistance and holds Prognostic values in colon cancer. Mol. Ther. 2016, 24, 2064–2077. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Pan, C.; Cai, Y.; Wang, H. Interplay between long noncoding RNA ZEB1-AS1 and miR-200s regulates osteosarcoma cell proliferation and migration. J. Cell Biochem. 2017. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Lin, J. Long noncoding RNA ZEB1-AS1 acts as an oncogene in osteosarcoma by epigenetically activating ZEB1. Am. J. Transl. Res. 2016, 8, 4095–4105. [Google Scholar] [PubMed]
- Beltran, M.; Puig, I.; Pena, C.; Garcia, J.M.; Alvarez, A.B.; Pena, R.; Bonilla, F.; de Herreros, A.G. A natural antisense transcript regulates ZEB2/SIP1 gene expression during SNAIL1-induced epithelial-mesenchymal transition. Genes Dev. 2008, 22, 756–769. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, J.; Lu, Q.; Shen, B.; Huang, X.; Shen, L.; Zheng, X.; Huang, R.; Yan, J.; Guo, H. TGFβ1 secreted by cancer-associated fibroblasts induces epithelial-mesenchymal transition of bladder cancer cells through lncRNA-ZEB2NAT. Sci. Rep. 2015, 5, 11924. [Google Scholar] [CrossRef] [PubMed]
- Qi, H.L.; Li, C.S.; Qian, C.W.; Xiao, Y.S.; Yuan, Y.F.; Liu, Q.Y.; Liu, Z.S. The long noncoding RNA, EGFR-AS1, a target of GHR, increases the expression of EGFR in hepatocellular carcinoma. Tumour Biol. 2016, 37, 1079–1089. [Google Scholar] [CrossRef] [PubMed]
- Espada, J.; Peinado, H.; Lopez-Serra, L.; Setien, F.; Lopez-Serra, P.; Portela, A.; Renart, J.; Carrasco, E.; Calvo, M.; Juarranz, A.; et al. Regulation of SNAIL1 and E-cadherin function by DNMT1 in a DNA methylation-independent context. Nucleic Acids Res. 2011, 39, 9194–9205. [Google Scholar] [CrossRef] [PubMed]
- Fan, H.; Zhao, Z.; Quan, Y.; Xu, J.; Zhang, J.; Xie, W. DNA methyltransferase 1 knockdown induces silenced CDH1 gene reexpression by demethylation of methylated CPG in hepatocellular carcinoma cell line smmc-7721. Eur. J. Gastroenterol. Hepatol. 2007, 19, 952–961. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Liu, H.; Shi, X.; Yao, Y.; Yang, W.; Song, Y. The long non-coding RNA HNF1A-AS1 regulates proliferation and metastasis in lung adenocarcinoma. Oncotarget 2015, 6, 9160–9172. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Song, J.H.; Cheng, Y.; Wu, W.; Bhagat, T.; Yu, Y.; Abraham, J.M.; Ibrahim, S.; Ravich, W.; Roland, B.C.; et al. Long non-coding RNA HNF1A-AS1 regulates proliferation and migration in oesophageal adenocarcinoma cells. Gut 2014, 63, 881–890. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Wei, X.; Zhang, A.; Li, C.; Bai, J.; Dong, J. Long non-coding RNA HNF1A-AS1 functioned as an oncogene and autophagy promoter in hepatocellular carcinoma through sponging HSA-miR-30b-5p. Biochem. Biophys. Res. Commun. 2016, 473, 1268–1275. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Mou, L.; Chai, H.X.; Wang, F.; Yin, Y.Z.; Zhang, X.Y. Long non-coding RNA HNF1A-AS1 promotes hepatocellular carcinoma cell proliferation by repressing NKD1 and p21 expression. Biomed. Pharmacother. 2017, 89, 926–932. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Hou, W.; Tao, J.; Zhao, Y.; Wan, G.; Ma, C.; Xu, H. Upregulation of lncRNA HNF1A-AS1 promotes cell proliferation and metastasis in osteosarcoma through activation of the Wnt/β-catenin signaling pathway. Am. J. Transl. Res. 2016, 8, 3503–3512. [Google Scholar] [PubMed]
- Jin, Y.; Cui, Z.; Li, X.; Jin, X.; Peng, J. Upregulation of long non-coding RNA plncRNA-1 promotes proliferation and induces epithelial-mesenchymal transition in prostate cancer. Oncotarget 2017. [Google Scholar] [CrossRef] [PubMed]
- Fang, Z.; Xu, C.; Li, Y.; Cai, X.; Ren, S.; Liu, H.; Wang, Y.; Wang, F.; Chen, R.; Qu, M.; et al. A feed-forward regulatory loop between androgen receptor and plncRNA-1 promotes prostate cancer progression. Cancer Lett. 2016, 374, 62–74. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Cui, Z.L.; Wang, Q.; Jin, X.B.; Zhao, Y.; Wang, M.W.; Song, W.; Qu, H.W.; Kang, W.T. PlncRNA-1 induces apoptosis through the HER-2 pathway in prostate cancer cells. Asian J. Androl. 2016. [Google Scholar] [CrossRef]
- Ma, C.C.; Xiong, Z.; Zhu, G.N.; Wang, C.; Zong, G.; Wang, H.L.; Bian, E.B.; Zhao, B. Long non-coding RNA ATB promotes glioma malignancy by negatively regulating miR-200a. J. Exp. Clin. Cancer Res. 2016, 35, 90. [Google Scholar] [CrossRef] [PubMed]
- Shi, S.J.; Wang, L.J.; Yu, B.; Li, Y.H.; Jin, Y.; Bai, X.Z. LncRNA-ATB promotes trastuzumab resistance and invasion-metastasis cascade in breast cancer. Oncotarget 2015, 6, 11652–11663. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.H.; Yang, F.; Wang, F.; Ma, J.Z.; Guo, Y.J.; Tao, Q.F.; Liu, F.; Pan, W.; Wang, T.T.; Zhou, C.C.; et al. A long noncoding RNA activated by TGF-beta promotes the invasion-metastasis cascade in hepatocellular carcinoma. Cancer Cell 2014, 25, 666–681. [Google Scholar] [CrossRef] [PubMed]
- Saito, T.; Kurashige, J.; Nambara, S.; Komatsu, H.; Hirata, H.; Ueda, M.; Sakimura, S.; Uchi, R.; Takano, Y.; Shinden, Y.; et al. A long non-coding RNA activated by transforming growth factor-beta is an independent prognostic marker of gastric cancer. Ann. Surg. Oncol. 2015, 22 Suppl 3, S915–S922. [Google Scholar] [CrossRef] [PubMed]
- Lei, K.; Liang, X.; Gao, Y.; Xu, B.; Xu, Y.; Li, Y.; Tao, Y.; Shi, W.; Liu, J. Lnc-ATB contributes to gastric cancer growth through a miR-141–3p/TGFβ2 feedback loop. Biochem. Biophys. Res. Commun. 2017, 484, 514–521. [Google Scholar] [CrossRef] [PubMed]
- Yue, B.; Qiu, S.; Zhao, S.; Liu, C.; Zhang, D.; Yu, F.; Peng, Z.; Yan, D. LncRNA-ATB mediated e-cadherin repression promotes the progression of colon cancer and predicts poor prognosis. J. Gastroenterol. Hepatol. 2016, 31, 595–603. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.; Yi, X.M.; Tang, C.P.; Ge, J.P.; Zhang, Z.Y.; Zhou, W.Q. Long non-coding RNA ATB promotes growth and epithelial-mesenchymal transition and predicts poor prognosis in human prostate carcinoma. Oncol. Rep. 2016, 36, 10–22. [Google Scholar] [CrossRef] [PubMed]
- Bai, W.D.; Ye, X.M.; Zhang, M.Y.; Zhu, H.Y.; Xi, W.J.; Huang, X.; Zhao, J.; Gu, B.; Zheng, G.X.; Yang, A.G.; et al. miR-200c suppresses TGF-beta signaling and counteracts trastuzumab resistance and metastasis by targeting ZNF217 and ZEB1 in breast cancer. Int. J. Cancer 2014, 135, 1356–1368. [Google Scholar] [CrossRef] [PubMed]
- Li, S.P.; Xu, H.X.; Yu, Y.; He, J.D.; Wang, Z.; Xu, Y.J.; Wang, C.Y.; Zhang, H.M.; Zhang, R.X.; Zhang, J.J.; et al. LncRNA HULC enhances epithelial-mesenchymal transition to promote tumorigenesis and metastasis of hepatocellular carcinoma via the miR-200a-3p/ZEB1 signaling pathway. Oncotarget 2016, 7, 42431–42446. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Li, Y.; Chai, X.; Kang, Q.; Zhao, P.; Xiong, J.; Wang, J. Long noncoding RNA HULC promotes cell proliferation by regulating PI3K/Akt signaling pathway in chronic myeloid leukemia. Gene 2017, 607, 41–46. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Liu, X.; Wu, H.; Ni, P.; Gu, Z.; Qiao, Y.; Chen, N.; Sun, F.; Fan, Q. Creb up-regulates long non-coding RNA, HULC expression through interaction with microRNA-372 in liver cancer. Nucleic Acids Res. 2010, 38, 5366–5383. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Liu, X.; Zhou, J.; Hu, J.; Zhang, D.; Liu, J.; Qiao, Y.; Zhan, Q. LncRNA HULC modulates the phosphorylation of yb-1 through serving as a scaffold of ERK and yb-1 to enhance hepatocarcinogenesis. Hepatology 2016. [Google Scholar] [CrossRef]
- Yang, X.J.; Huang, C.Q.; Peng, C.W.; Hou, J.X.; Liu, J.Y. Long noncoding RNA HULC promotes colorectal carcinoma progression through epigenetically repressing NKD2 expression. Gene 2016, 592, 172–178. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Song, X.; Wang, X.; Hu, J.; Jiang, L. Silencing of lncrna hulc enhances chemotherapy induced apoptosis in human gastric cancer. J. Med. Biochem. 2015, 35, 137–143. [Google Scholar]
- Xiong, H.; Ni, Z.; He, J.; Jiang, S.; Li, X.; Gong, W.; Zheng, L.; Chen, S.; Li, B.; Zhang, N.; et al. LncRNA HULC triggers autophagy via stabilizing SIRT1 and attenuates the chemosensitivity of HCC cells. Oncogene 2017. [Google Scholar] [CrossRef] [PubMed]
- Gui, X.; Li, H.; Li, T.; Pu, H.; Lu, D. Long noncoding RNA CUDR regulates HULC and beta-catenin to govern human liver stem cell malignant differentiation. Mol. Ther. 2015, 23, 1843–1853. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.; Lin, Z.; Li, X.; Xin, X.; An, J.; Zheng, Q.; Yang, Y.; Lu, D. Hulc cooperates with MALAT1 to aggravate liver cancer stem cells growth through telomere repeat-binding factor 2. Sci. Rep. 2016, 6, 36045. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Jiao, C.; Lin, Y.; Chen, M.; Zhang, J.; Wang, J.; Zhang, Z. LncRNA UCA1 contributes to imatinib resistance by acting as a ceRNA against miR-16 in chronic myeloid leukemia cells. DNA Cell Biol. 2017, 36, 18–25. [Google Scholar] [CrossRef] [PubMed]
- Xue, M.; Pang, H.; Li, X.; Li, H.; Pan, J.; Chen, W. Long non-coding RNA urothelial cancer-associated 1 promotes bladder cancer cell migration and invasion by way of the HSA-miR-145-ZEB1/2-FSCN1 pathway. Cancer Sci. 2016, 107, 18–27. [Google Scholar] [CrossRef] [PubMed]
- Pan, J.; Li, X.; Wu, W.; Xue, M.; Hou, H.; Zhai, W.; Chen, W. Long non-coding RNA UCA1 promotes cisplatin/gemcitabine resistance through Creb modulating miR-196a-5p in bladder cancer cells. Cancer Lett. 2016, 382, 64–76. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Ying, H.Q.; He, B.S.; Pan, Y.Q.; Deng, Q.W.; Sun, H.L.; Chen, J.; Liu, X.; Wang, S.K. Upregulated lncRNA-UCA1 contributes to progression of hepatocellular carcinoma through inhibition of miR-216b and activation of FGFR1/ERK signaling pathway. Oncotarget 2015, 6, 7899–7917. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Jiang, Y.; Wan, Y.; Zhang, L.; Qiu, J.; Zhou, S.; Cheng, W. Uca1 functions as a competing endogenous RNA to suppress epithelial ovarian cancer metastasis. Tumour Biol. 2016, 37, 10633–10641. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Shen, B.; Tan, M.; Mu, X.; Qin, Y.; Zhang, F.; Liu, Y. Long non-coding RNA UCA1 increases chemoresistance of bladder cancer cells by regulating Wnt signaling. FEBS J. 2014, 281, 1750–1758. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Wang, G.; Yang, L.; Qu, J.; Yang, Z.; Zhou, X. Knockdown of long non-coding RNA UCA1 increases the tamoxifen sensitivity of breast cancer cells through inhibition of Wnt/beta-catenin pathway. PLoS ONE 2016, 11, e0168406. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Gao, Z.; Liao, J.; Shang, M.; Li, X.; Yin, L.; Pu, Y.; Liu, R. LncRNA UCA1 inhibits esophageal squamous-cell carcinoma growth by regulating the Wnt signaling pathway. J. Toxicol. Environ. Health A 2016, 79, 407–418. [Google Scholar] [CrossRef] [PubMed]
- Xiao, C.; Wu, C.H.; Hu, H.Z. LncRNA UCA1 promotes epithelial-mesenchymal transition (EMT) of breast cancer cells via enhancing Wnt/beta-catenin signaling pathway. Eur. Rev. Med. Pharmacol. Sci. 2016, 20, 2819–2824. [Google Scholar] [PubMed]
- Yang, Y.T.; Wang, Y.F.; Lai, J.Y.; Shen, S.Y.; Wang, F.; Kong, J.; Zhang, W.; Yang, H.Y. Long non-coding RNA UCA1 contributes to the progression of oral squamous cell carcinoma by regulating the Wnt/beta-catenin signaling pathway. Cancer Sci. 2016, 107, 1581–1589. [Google Scholar] [CrossRef] [PubMed]
- Cheng, N.; Cai, W.; Ren, S.; Li, X.; Wang, Q.; Pan, H.; Zhao, M.; Li, J.; Zhang, Y.; Zhao, C.; et al. Long non-coding RNA UCA1 induces non-T790M acquired resistance to EGFR-TKIS by activating the Akt/mtor pathway in EGFR-mutant non-small cell lung cancer. Oncotarget 2015, 6, 23582–23593. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Wu, Y.; Liu, A.; Tang, X. Long non-coding RNA UCA1 enhances tamoxifen resistance in breast cancer cells through a miR-18a-HIF1α feedback regulatory loop. Tumour Biol. 2016, 37, 14733–14743. [Google Scholar] [CrossRef] [PubMed]
- Zuo, Z.K.; Gong, Y.; Chen, X.H.; Ye, F.; Yin, Z.M.; Gong, Q.N.; Huang, J.S. TGFβ1-induced lncRNA UCA1 upregulation promotes gastric cancer invasion and migration. DNA Cell Biol. 2017, 36, 159–167. [Google Scholar] [CrossRef] [PubMed]
- Fang, Q.; Chen, X.; Zhi, X. Long non-coding RNA (lncRNA) urothelial carcinoma associated 1 (UCA1) increases multi-drug resistance of gastric cancer via downregulating miR-27b. Med. Sci. Monit. 2016, 22, 3506–3513. [Google Scholar] [CrossRef] [PubMed]
- Shang, C.; Guo, Y.; Zhang, J.; Huang, B. Silence of long noncoding RNA UCA1 inhibits malignant proliferation and chemotherapy resistance to adriamycin in gastric cancer. Cancer Chemother. Pharmacol. 2016, 77, 1061–1067. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Zhou, J.; Xie, X.; Hu, J.; Chen, L.; Hu, Q.; Guo, H.; Yu, C. Involvement of SRPK1 in cisplatin resistance related to long non-coding RNA UCA1 in human ovarian cancer cells. Neoplasma 2015, 62, 432–438. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Yang, B.; Ma, B. The UCA1/miR-204/SIRT1 axis modulates docetaxel sensitivity of prostate cancer cells. Cancer Chemother. Pharmacol. 2016, 78, 1025–1031. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.G.; Yang, M.F.; Ren, Y.Q.; Wu, C.H.; Wang, L.Q. Exosomes mediated transfer of lncRNA UCA1 results in increased tamoxifen resistance in breast cancer cells. Eur. Rev. Med. Pharmacol. Sci. 2016, 20, 4362–4368. [Google Scholar] [PubMed]
- Li, T.; Zheng, Q.; An, J.; Wu, M.; Li, H.; Gui, X.; Pu, H.; Lu, D. SET1a cooperates with cudr to promote liver cancer growth and hepatocyte-like stem cell malignant transformation epigenetically. Mol. Ther. 2016, 24, 261–275. [Google Scholar] [CrossRef] [PubMed]
- Pu, H.; Zheng, Q.; Li, H.; Wu, M.; An, J.; Gui, X.; Li, T.; Lu, D. CUDR promotes liver cancer stem cell growth through upregulating TERT and c-MYC. Oncotarget 2015, 6, 40775–40798. [Google Scholar] [PubMed]
- Tan, J.; Qiu, K.; Li, M.; Liang, Y. Double-negative feedback loop between long non-coding RNA TUG1 and miR-145 promotes epithelial to mesenchymal transition and radioresistance in human bladder cancer cells. FEBS Lett. 2015, 589, 3175–3181. [Google Scholar] [CrossRef] [PubMed]
- Ma, F.; Wang, S.H.; Cai, Q.; Jin, L.Y.; Zhou, D.; Ding, J.; Quan, Z.W. Long non-coding RNA TUG1 promotes cell proliferation and metastasis by negatively regulating mir-300 in gallbladder carcinoma. Biomed. Pharmacother. 2017, 88, 863–869. [Google Scholar] [CrossRef] [PubMed]
- Jiang, L.; Wang, W.; Li, G.; Sun, C.; Ren, Z.; Sheng, H.; Gao, H.; Wang, C.; Yu, H. High tug1 expression is associated with chemotherapy resistance and poor prognosis in esophageal squamous cell carcinoma. Cancer Chemother. Pharmacol. 2016, 78, 333–339. [Google Scholar] [CrossRef] [PubMed]
- Katsushima, K.; Natsume, A.; Ohka, F.; Shinjo, K.; Hatanaka, A.; Ichimura, N.; Sato, S.; Takahashi, S.; Kimura, H.; Totoki, Y.; et al. Targeting the notch-regulated non-coding RNA TUG1 for glioma treatment. Nat. Commun. 2016, 7, 13616. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Li, T.; Wei, G.; Liu, L.; Chen, Q.; Xu, L.; Zhang, K.; Zeng, D.; Liao, R. The long non-coding RNA NEAT1 regulates epithelial to mesenchymal transition and radioresistance in through mir-204/zeb1 axis in nasopharyngeal carcinoma. Tumour Biol. 2016, 37, 11733–11741. [Google Scholar] [CrossRef] [PubMed]
- Qian, K.; Liu, G.; Tang, Z.; Hu, Y.; Fang, Y.; Chen, Z.; Xu, X. The long non-coding RNA NEAT1 interacted with miR-101 modulates breast cancer growth by targeting EZH2. Arch. Biochem. Biophys. 2017, 615, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Jiang, P.; Wu, X.; Wang, X.; Huang, W.; Feng, Q. NEAT1 upregulates EGCG-induced CTR1 to enhance cisplatin sensitivity in lung cancer cells. Oncotarget 2016, 7, 43337–43351. [Google Scholar] [CrossRef] [PubMed]
- Gao, C.; Zhang, J.; Wang, Q.; Ren, C. Overexpression of lncRNA NEAT1 mitigates multidrug resistance by inhibiting ABCG2 in leukemia. Oncol. Lett. 2016, 12, 1051–1057. [Google Scholar] [CrossRef] [PubMed]
- Gong, W.; Zheng, J.; Liu, X.; Ma, J.; Liu, Y.; Xue, Y. Knockdown of NEAT1 restrained the malignant progression of glioma stem cells by activating microRNA let-7e. Oncotarget 2016, 7, 62208–62223. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Xiao, Z.; Du, X.; Huang, L.; Du, G. Silencing of the long non-coding RNA NEAT1 suppresses glioma stem-like properties through modulation of the miR-107/CDK6 pathway. Oncol. Rep. 2017, 37, 555–562. [Google Scholar] [CrossRef] [PubMed]
- Lo, P.K.; Zhang, Y.; Wolfson, B.; Gernapudi, R.; Yao, Y.; Duru, N.; Zhou, Q. Dysregulation of the brca1/long non-coding RNA NEAT1 signaling axis contributes to breast tumorigenesis. Oncotarget 2016, 7, 65067–65089. [Google Scholar] [CrossRef] [PubMed]
- Sun, N.X.; Ye, C.; Zhao, Q.; Zhang, Q.; Xu, C.; Wang, S.B.; Jin, Z.J.; Sun, S.H.; Wang, F.; Li, W. Long noncoding RNA-ebic promotes tumor cell invasion by binding to EZH2 and repressing e-cadherin in cervical cancer. PLoS ONE 2014, 9, e100340. [Google Scholar] [CrossRef] [PubMed]
- Zhou, B.; Wang, Y.; Jiang, J.; Jiang, H.; Song, J.; Han, T.; Shi, J.; Qiao, H. The long noncoding RNA colon cancer-associated transcript-1/miR-490 axis regulates gastric cancer cell migration by targeting hnRNPA1. IUBMB Life 2016, 68, 201–210. [Google Scholar] [CrossRef] [PubMed]
- Deng, L.; Yang, S.B.; Xu, F.F.; Zhang, J.H. Long noncoding RNA CCAT1 promotes hepatocellular carcinoma progression by functioning as let-7 sponge. J. Exp. Clin. Cancer Res. 2015, 34, 18. [Google Scholar] [CrossRef] [PubMed]
- Ma, M.Z.; Chu, B.F.; Zhang, Y.; Weng, M.Z.; Qin, Y.Y.; Gong, W.; Quan, Z.W. Long non-coding RNA CCAT1 promotes gallbladder cancer development via negative modulation of miRNA-218–5p. Cell Death Dis. 2015, 6, e1583. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Wang, W.; Cao, L.; Li, Z.; Wang, X. Long non-coding RNA CCAT1 acts as a competing endogenous RNA to regulate cell growth and differentiation in acute myeloid leukemia. Mol. Cells 2016, 39, 330–336. [Google Scholar] [PubMed]
- Kim, T.; Cui, R.; Jeon, Y.J.; Lee, J.H.; Sim, H.; Park, J.K.; Fadda, P.; Tili, E.; Nakanishi, H.; Huh, M.I.; et al. Long-range interaction and correlation between myc enhancer and oncogenic long noncoding RNA carlo-5. Proc. Natl. Acad. Sci. USA 2014, 111, 4173–4178. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.Q.; Zhou, X.; Chang, H.; Li, H.G.; Liu, F.F.; Ma, C.Q.; Lu, J. Aberrant expression of CCAT1 regulated by c-MYC predicts the prognosis of hepatocellular carcinoma. Asian Pac. J. Cancer Prev. 2015, 16, 5181–5185. [Google Scholar] [CrossRef] [PubMed]
- Xiang, J.F.; Yin, Q.F.; Chen, T.; Zhang, Y.; Zhang, X.O.; Wu, Z.; Zhang, S.; Wang, H.B.; Ge, J.; Lu, X.; et al. Human colorectal cancer-specific CCAT1-l lncRNA regulates long-range chromatin interactions at the MYC locus. Cell Res. 2014, 24, 513–531. [Google Scholar] [CrossRef] [PubMed]
- McCleland, M.L.; Mesh, K.; Lorenzana, E.; Chopra, V.S.; Segal, E.; Watanabe, C.; Haley, B.; Mayba, O.; Yaylaoglu, M.; Gnad, F.; et al. CCAT1 is an enhancer-templated RNA that predicts bet sensitivity in colorectal cancer. J. Clin. Investig. 2016, 126, 639–652. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.J.; Liu, J.Z.; Lv, P.; Dang, Y.; Gao, J.Y.; Wang, Y. Long non-coding RNA CCAT2 promotes gastric cancer proliferation and invasion by regulating the e-cadherin and lats2. Am. J. Cancer Res. 2016, 6, 2651–2660. [Google Scholar] [PubMed]
- Redis, R.S.; Vela, L.E.; Lu, W.; Ferreira de Oliveira, J.; Ivan, C.; Rodriguez-Aguayo, C.; Adamoski, D.; Pasculli, B.; Taguchi, A.; Chen, Y.; et al. Allele-specific reprogramming of cancer metabolism by the long non-coding RNA CCAT2. Mol. Cell 2016, 61, 520–534. [Google Scholar] [CrossRef] [PubMed]
- Cai, Y.; He, J.; Zhang, D. Long noncoding RNA CCAT2 promotes breast tumor growth by regulating the Wnt signaling pathway. Onco Targets Ther. 2015, 8, 2657–2664. [Google Scholar] [PubMed]
- Wu, Z.J.; Li, Y.; Wu, Y.Z.; Wang, Y.; Nian, W.Q.; Wang, L.L.; Li, L.C.; Luo, H.L.; Wang, D.L. Long non-coding RNA CCAT2 promotes the breast cancer growth and metastasis by regulating TGF-beta signaling pathway. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 706–714. [Google Scholar] [PubMed]
- Gong, W.J.; Yin, J.Y.; Li, X.P.; Fang, C.; Xiao, D.; Zhang, W.; Zhou, H.H.; Li, X.; Liu, Z.Q. Association of well-characterized lung cancer lncRNA polymorphisms with lung cancer susceptibility and platinum-based chemotherapy response. Tumour Biol. 2016, 37, 8349–8358. [Google Scholar] [CrossRef] [PubMed]
- Jia, X.; Wang, Z.; Qiu, L.; Yang, Y.; Wang, Y.; Chen, Z.; Liu, Z.; Yu, L. Upregulation of lncRNA-hit promotes migration and invasion of non-small cell lung cancer cells by association with ZEB1. Cancer Med. 2016, 5, 3555–3563. [Google Scholar] [CrossRef] [PubMed]
- Richards, E.J.; Zhang, G.; Li, Z.P.; Permuth-Wey, J.; Challa, S.; Li, Y.; Kong, W.; Dan, S.; Bui, M.M.; Coppola, D.; et al. Long non-coding RNAs (lncRNA) regulated by transforming growth factor (TGF) beta: LncRNA-HIT-mediated TGFβ-induced epithelial to mesenchymal transition in mammary epithelia. J. Biol. Chem. 2015, 290, 6857–6867. [Google Scholar] [CrossRef] [PubMed]
- Zhou, M.; Hou, Y.; Yang, G.; Zhang, H.; Tu, G.; Du, Y.E.; Wen, S.; Xu, L.; Tang, X.; Tang, S.; et al. LncRNA-HH strengthen cancer stem cells generation in twist-positive breast cancer via activation of hedgehog signaling pathway. Stem Cells 2016, 34, 55–66. [Google Scholar] [CrossRef] [PubMed]
- Pan, H.; Jiang, T.; Cheng, N.; Wang, Q.; Ren, S.; Li, X.; Zhao, C.; Zhang, L.; Cai, W.; Zhou, C. Long non-coding RNA bc087858 induces non-t790m mutation acquired resistance to EGFR-TKIS by activating PI3K/Akt and MEK/ERK pathways and emt in non-small-cell lung cancer. Oncotarget 2016, 7, 49948–49960. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Zhu, L.; Xu, L.; Qin, K.; Liu, C.; Yu, Y.; Su, D.; Wu, K.; Sheng, Y. Long noncoding RNA linc00617 exhibits oncogenic activity in breast cancer. Mol. Carcinog. 2017, 56, 3–17. [Google Scholar] [CrossRef] [PubMed]
- Gao, K.; Ji, Z.; She, K.; Yang, Q.; Shao, L. Long non-coding RNA ZFAS1 is an unfavourable prognostic factor and promotes glioma cell progression by activation of the notch signaling pathway. Biomed. Pharmacother. 2017, 87, 555–560. [Google Scholar] [CrossRef] [PubMed]
- Shi, X.; Sun, M.; Liu, H.; Yao, Y.; Kong, R.; Chen, F.; Song, Y. A critical role for the long non-coding RNA GAS5 in proliferation and apoptosis in non-small-cell lung cancer. Mol. Carcinog. 2015, 54 (Suppl. 1), E1–E12. [Google Scholar] [CrossRef] [PubMed]
- Mazar, J.; Rosado, A.; Shelley, J.; Marchica, J.; Westmoreland, T.J. The long non-coding RNA GAS5 differentially regulates cell cycle arrest and apoptosis through activation of BRCA1 and p53 in human neuroblastoma. Oncotarget 2017, 8, 6589–6607. [Google Scholar] [CrossRef] [PubMed]
- Dong, S.; Qu, X.; Li, W.; Zhong, X.; Li, P.; Yang, S.; Chen, X.; Shao, M.; Zhang, L. The long non-coding RNA, GAS5, enhances gefitinib-induced cell death in innate EGFR tyrosine kinase inhibitor-resistant lung adenocarcinoma cells with wide-type EGFR via downregulation of the IGF-1R expression. J. Hematol. Oncol. 2015, 8, 43. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Wang, A.Y.; Wang, X.K.; Sun, X.M.; Xue, H.Z. GAS5 is downregulated in gastric cancer cells by promoter hypermethylation and regulates adriamycin sensitivity. Eur. Rev. Med. Pharmacol. Sci. 2016, 20, 3199–3205. [Google Scholar] [PubMed]
- Zhang, N.; Yang, G.Q.; Shao, X.M.; Wei, L. GAS5 modulated autophagy is a mechanism modulating cisplatin sensitivity in NSCLC cells. Eur. Rev. Med. Pharmacol. Sci. 2016, 20, 2271–2277. [Google Scholar] [PubMed]
- Pickard, M.R.; Williams, G.T. Regulation of apoptosis by long non-coding RNA GAS5 in breast cancer cells: Implications for chemotherapy. Breast Cancer Res. Treat. 2014, 145, 359–370. [Google Scholar] [CrossRef] [PubMed]
- Mourtada-Maarabouni, M.; Pickard, M.R.; Hedge, V.L.; Farzaneh, F.; Williams, G.T. GAS5, a non-protein-coding RNA, controls apoptosis and is downregulated in breast cancer. Oncogene 2009, 28, 195–208. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Guo, Y.; Song, Y.; Shang, C. Long noncoding RNA GAS5 inhibits malignant proliferation and chemotherapy resistance to doxorubicin in bladder transitional cell carcinoma. Cancer Chemother. Pharmacol. 2017, 79, 49–55. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.F.; Guo, Y.J.; Zhao, C.X.; Yuan, S.X.; Wang, Y.; Tang, G.N.; Zhou, W.P.; Sun, S.H. Hepatitis b virus X protein (HBX)-related long noncoding RNA (lncRNA) down-regulated expression by HBX (DREH) inhibits hepatocellular carcinoma metastasis by targeting the intermediate filament protein vimentin. Hepatology 2013, 57, 1882–1892. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Liu, Z.; Yao, B.; Dou, C.; Xu, M.; Xue, Y.; Ding, L.; Jia, Y.; Zhang, H.; Li, Q.; et al. Long non-coding RNA TUSC7 acts a molecular sponge for miR-10a and suppresses EMT in hepatocellular carcinoma. Tumour Biol. 2016, 37, 11429–11441. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Zhang, R.; Zhao, J. The novel long noncoding RNA TUSC7 inhibits proliferation by sponging miR-211 in colorectal cancer. Cell Physiol. Biochem. 2017, 41, 635–644. [Google Scholar] [CrossRef] [PubMed]
- Shang, C.; Guo, Y.; Hong, Y.; Xue, Y.X. Long non-coding RNA tusc7, a target of miR-23b, plays tumor-suppressing roles in human gliomas. Front. Cell Neurosci. 2016, 10, 235. [Google Scholar] [CrossRef] [PubMed]
- Qi, P.; Xu, M.D.; Shen, X.H.; Ni, S.J.; Huang, D.; Tan, C.; Weng, W.W.; Sheng, W.Q.; Zhou, X.Y.; Du, X. Reciprocal repression between TUSC7 and miR-23b in gastric cancer. Int. J. Cancer 2015, 137, 1269–1278. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Huang, J.; Zhou, N.; Zhang, Z.; Zhang, A.; Lu, Z.; Wu, F.; Mo, Y.Y. LncRNA loc285194 is a p53-regulated tumor suppressor. Nucleic Acids Res. 2013, 41, 4976–4987. [Google Scholar] [CrossRef] [PubMed]
- Tong, Y.S.; Zhou, X.L.; Wang, X.W.; Wu, Q.Q.; Yang, T.X.; Lv, J.; Yang, J.S.; Zhu, B.; Cao, X.F. Association of decreased expression of long non-coding RNA loc285194 with chemoradiotherapy resistance and poor prognosis in esophageal squamous cell carcinoma. J. Transl. Med. 2014, 12, 233. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.H.; Lin, Y.S.; Chen, Y.; Yeh, C.T.; Huang, Y.L.; Hsieh, T.H.; Shieh, T.M.; Hsueh, C.; Chen, T.C. Long non-coding RNA AOC4P suppresses hepatocellular carcinoma metastasis by enhancing vimentin degradation and inhibiting epithelial-mesenchymal transition. Oncotarget 2015, 6, 23342–23357. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Ye, J.; Wu, D.; Wu, P.; Chen, Z.; Chen, J.; Gao, S.; Huang, J. Leigc long non-coding RNA acts as a tumor suppressor in gastric carcinoma by inhibiting the epithelial-to-mesenchymal transition. BMC Cancer 2014, 14, 932. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Huang, S.; Zhang, W.; He, K.; Zhao, M.; Lin, H.; Li, D.; Zhang, H.; Zheng, Z.; Huang, C. Decreased expression of lncRNA slc25a25-as1 promotes proliferation, chemoresistance, and emt in colorectal cancer cells. Tumour Biol. 2016, 37, 14205–14215. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.; Yu, H.; Shen, Y.; Liu, Y.; Yang, Z.; Sun, T. miR-146b-5p overexpression attenuates stemness and radioresistance of glioma stem cells by targeting HUR/lincRNA-p21/β-catenin pathway. Oncotarget 2016, 7, 41505–41526. [Google Scholar] [CrossRef] [PubMed]
- Yoon, J.H.; Abdelmohsen, K.; Srikantan, S.; Yang, X.; Martindale, J.L.; De, S.; Huarte, M.; Zhan, M.; Becker, K.G.; Gorospe, M. LincRNA-p21 suppresses target mRNA translation. Mol. Cell 2012, 47, 648–655. [Google Scholar] [CrossRef] [PubMed]
- Jia, M.; Jiang, L.; Wang, Y.D.; Huang, J.Z.; Yu, M.; Xue, H.Z. LincRNA-p21 inhibits invasion and metastasis of hepatocellular carcinoma through notch signaling-induced epithelial-mesenchymal transition. Hepatol. Res. 2016, 46, 1137–1144. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Zhang, H.; Mei, Y.; Wu, M. Reciprocal regulation of HIF-1alpha and lincRNA-p21 modulates the warburg effect. Mol. Cell 2014, 53, 88–100. [Google Scholar] [CrossRef] [PubMed]
- Huarte, M.; Guttman, M.; Feldser, D.; Garber, M.; Koziol, M.J.; Kenzelmann-Broz, D.; Khalil, A.M.; Zuk, O.; Amit, I.; Rabani, M.; et al. A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell 2010, 142, 409–419. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Ruan, Y.; Zhao, W.; Jiang, Q.; Jiang, C.; Zhao, Y.; Xu, Y.; Sun, F.; Zhu, Y.; Xia, S.; et al. Long intragenic non-coding RNA lincRNA-p21 suppresses development of human prostate cancer. Cell Prolif. 2017, 50, e12318. [Google Scholar] [CrossRef] [PubMed]
- Spurlock, C.F., 3rd; Tossberg, J.T.; Matlock, B.K.; Olsen, N.J.; Aune, T.M. Methotrexate inhibits NF-kappab activity via long intergenic (noncoding) RNA-p21 induction. Arthritis Rheumatol. 2014, 66, 2947–2957. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Lei, Z.J.; Guo, Y.; Wang, T.; Qin, Z.Y.; Xiao, H.L.; Fan, L.L.; Chen, D.F.; Bian, X.W.; Liu, J.; et al. miRNA-regulated delivery of lincRNA-p21 suppresses beta-catenin signaling and tumorigenicity of colorectal cancer stem cells. Oncotarget 2015, 6, 37852–37870. [Google Scholar] [PubMed]
- Bao, X.; Wu, H.; Zhu, X.; Guo, X.; Hutchins, A.P.; Luo, Z.; Song, H.; Chen, Y.; Lai, K.; Yin, M.; et al. The p53-induced lincRNA-p21 derails somatic cell reprogramming by sustaining H3K9ME3 and CPG methylation at pluripotency gene promoters. Cell Res. 2015, 25, 80–92. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.H.; Yu, C.C.; Lin, Y.S.; Chen, T.C.; Yeh, C.T.; Liang, K.H.; Shieh, T.M.; Chen, C.Y.; Hsueh, C. Long noncoding RNA CPS1-IT1 suppresses the metastasis of hepatocellular carcinoma by regulating HIF-1α activity and inhibiting epithelial-mesenchymal transition. Oncotarget 2016, 7, 43588–43603. [Google Scholar] [PubMed]
- Yu, Y.; Li, L.; Zheng, Z.; Chen, S.; Chen, E.; Hu, Y. Long non-coding RNA linc00261 suppresses gastric cancer progression via promoting SLUG degradation. J. Cell Mol. Med. 2017, 21, 955–967. [Google Scholar] [CrossRef] [PubMed]
- Ru, N.; Liang, J.; Zhang, F.; Wu, W.; Wang, F.; Liu, X.; Du, Y. SPRY4 intronic transcript 1 promotes epithelial-mesenchymal transition through association with SNAIL1 in osteosarcoma. DNA Cell Biol. 2016, 35, 290–295. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Li, Y.; Luo, G.; Xiao, X.; Tao, D.; Wu, X.; Wang, M.; Huang, C.; Wang, L.; Zeng, F.; et al. LncRNA SPRY4-IT1 sponges miR-101–3p to promote proliferation and metastasis of bladder cancer cells through up-regulating EZH2. Cancer Lett. 2017, 388, 281–291. [Google Scholar] [CrossRef] [PubMed]
- Zhou, M.; Zhang, X.Y.; Yu, X. Overexpression of the long non-coding RNA SPRY4-IT1 promotes tumor cell proliferation and invasion by activating EZH2 in hepatocellular carcinoma. Biomed. Pharmacother. 2017, 85, 348–354. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.Y.; Li, R.K.; Qi, Y.; Li, X.N.; Yang, Y.; Liu, D.L.; Zhao, J.; Zhu, D.Y.; Wu, K.; Zhou, X.D.; et al. Upregulation of long noncoding RNA SPRY4-IT1 promotes metastasis of esophageal squamous cell carcinoma via induction of epithelial-mesenchymal transition. Cell Biol. Toxicol. 2016, 32, 391–401. [Google Scholar] [CrossRef] [PubMed]
- Kong, J.; Sun, W.; Li, C.; Wan, L.; Wang, S.; Wu, Y.; Xu, E.; Zhang, H.; Lai, M. Long non-coding RNA linc01133 inhibits epithelial-mesenchymal transition and metastasis in colorectal cancer by interacting with SRSF6. Cancer Lett. 2016, 380, 476–484. [Google Scholar] [CrossRef] [PubMed]
- Zang, C.; Nie, F.Q.; Wang, Q.; Sun, M.; Li, W.; He, J.; Zhang, M.; Lu, K.H. Long non-coding RNA linc01133 represses KLF2, p21 and E-cadherin transcription through binding with EZH2, LSD1 in non small cell lung cancer. Oncotarget 2016, 7, 11696–11707. [Google Scholar] [PubMed]
- Guo, Q.; Zhao, Y.; Chen, J.; Hu, J.; Wang, S.; Zhang, D.; Sun, Y. Braf-activated long non-coding RNA contributes to colorectal cancer migration by inducing epithelial-mesenchymal transition. Oncol. Lett. 2014, 8, 869–875. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Wang, N.; Long, Z.; Ren, X. Long non-coding RNA BANCR promotes endometrial cancer cell proliferation and invasion by regulating MMP2 and MMP1 via ERK/MAPK signaling pathway. Cell Physiol. Biochem. 2016, 40, 644–656. [Google Scholar] [CrossRef] [PubMed]
- Liao, T.; Qu, N.; Shi, R.L.; Guo, K.; Ma, B.; Cao, Y.M.; Xiang, J.; Lu, Z.W.; Zhu, Y.X.; Li, D.S.; et al. BRAF-activated lncRNA functions as a tumor suppressor in papillary thyroid cancer. Oncotarget 2017, 8, 238–247. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.X.; Liu, Z.Q.; Jiang, B.; Lu, X.Y.; Ning, X.F.; Yuan, C.T.; Wang, A.L. braf activated non-coding RNA (BANCR) promoting gastric cancer cells proliferation via regulation of Nf-KAPPAB1. Biochem. Biophys. Res. Commun. 2015, 465, 225–231. [Google Scholar] [CrossRef] [PubMed]
- Ji, P.; Diederichs, S.; Wang, W.; Boing, S.; Metzger, R.; Schneider, P.M.; Tidow, N.; Brandt, B.; Buerger, H.; Bulk, E.; et al. MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene 2003, 22, 8031–8041. [Google Scholar] [CrossRef] [PubMed]
- Jiao, F.; Hu, H.; Yuan, C.; Wang, L.; Jiang, W.; Jin, Z.; Guo, Z. Elevated expression level of long noncoding RNA MALAT-1 facilitates cell growth, migration and invasion in pancreatic cancer. Oncol. Rep. 2014, 32, 2485–2492. [Google Scholar] [CrossRef] [PubMed]
- Ying, L.; Chen, Q.; Wang, Y.; Zhou, Z.; Huang, Y.; Qiu, F. Upregulated MALAT-1 contributes to bladder cancer cell migration by inducing epithelial-to-mesenchymal transition. Mol. Biosyst. 2012, 8, 2289–2294. [Google Scholar] [CrossRef] [PubMed]
- Shen, L.; Chen, L.; Wang, Y.; Jiang, X.; Xia, H.; Zhuang, Z. Long noncoding RNA MALAT1 promotes brain metastasis by inducing epithelial-mesenchymal transition in lung cancer. J. Neurooncol. 2015, 121, 101–108. [Google Scholar] [CrossRef] [PubMed]
- Sun, R.; Qin, C.; Jiang, B.; Fang, S.; Pan, X.; Peng, L.; Liu, Z.; Li, W.; Li, Y.; Li, G. Down-regulation of MALAT1 inhibits cervical cancer cell invasion and metastasis by inhibition of epithelial-mesenchymal transition. Mol. Biosyst. 2016, 12, 952–962. [Google Scholar] [CrossRef] [PubMed]
- Ning, L.; Li, Z.; Wei, D.; Chen, H.; Yang, C. LncRNA, NEAT1 is a prognosis biomarker and regulates cancer progression via epithelial-mesenchymal transition in clear cell renal cell carcinoma. Cancer Biomark 2017. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Wu, W.B.; Wang, Z.W.; Wang, X.H. LncRNA NEAT1 is closely related with progression of breast cancer via promoting proliferation and emt. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 1020–1026. [Google Scholar] [PubMed]
- Jin, C.; Yan, B.; Lu, Q.; Lin, Y.; Ma, L. Reciprocal regulation of HSA-miR-1 and long noncoding RNA MALAT1 promotes triple-negative breast cancer development. Tumour Biol. 2016, 37, 7383–7394. [Google Scholar] [CrossRef] [PubMed]
- Phillips, S.; Kuperwasser, C. SLUG: Critical regulator of epithelial cell identity in breast development and cancer. Cell Adh. Migr. 2014, 12, 578–587. [Google Scholar]
- Raveh, E.; Matouk, I.J.; Gilon, M.; Hochberg, A. The h19 long non-coding RNA in cancer initiation, progression and metastasis—A proposed unifying theory. Mol. Cancer 2015, 14, 184. [Google Scholar] [CrossRef] [PubMed]
- Matouk, I.J.; Raveh, E.; Abu-lail, R.; Mezan, S.; Gilon, M.; Gershtain, E.; Birman, T.; Gallula, J.; Schneider, T.; Barkali, M.; et al. Oncofetal h19 RNA promotes tumor metastasis. Biochim. Biophys. Acta 2014, 1843, 1414–1426. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.H.; Wu, X.C.; Zhang, M.D.; Weng, M.Z.; Zhou, D.; Quan, Z.W. Upregulation of h19 indicates a poor prognosis in gallbladder carcinoma and promotes epithelial-mesenchymal transition. Am. J. Cancer Res. 2016, 6, 15–26. [Google Scholar] [PubMed]
- Huang, C.; Cao, L.; Qiu, L.; Dai, X.; Ma, L.; Zhou, Y.; Li, H.; Gao, M.; Li, W.; Zhang, Q.; et al. Upregulation of h19 promotes invasion and induces epithelial-to-mesenchymal transition in esophageal cancer. Oncol. Lett. 2015, 10, 291–296. [Google Scholar] [CrossRef] [PubMed]
- Cai, X.; Cullen, B.R. The imprinted h19 noncoding RNA is a primary microRNA precursor. RNA 2007, 13, 313–316. [Google Scholar] [CrossRef] [PubMed]
- Rokavec, M.; Hermeking, H. Cellular model of colon cancer progression reveals signatures of mRNAs, miRNA, lncRNAs and epigenetic modifications associated with metastasis. Cancer Res. 2017, 77, 1854–1867. [Google Scholar] [CrossRef] [PubMed]
- Kim, N.H.; Choi, S.H.; Lee, T.R.; Lee, C.H.; Lee, A.Y. Cadherin 11, a mir-675 target, induces N-cadherin expression and epithelial-mesenchymal transition in melasma. J. Investig. Dermatol. 2014, 134, 2967–2976. [Google Scholar] [CrossRef] [PubMed]
- Hernandez, J.M.; Elahi, A.; Clark, C.W.; Wang, J.; Humphries, L.A.; Centeno, B.; Bloom, G.; Fuchs, B.C.; Yeatman, T.; Shibata, D. miR-675 mediates downregulation of TWIST1 and RB in AFP-secreting hepatocellular carcinoma. Ann. Surg. Oncol. 2013, 20 (Suppl. 3), S625–S635. [Google Scholar] [CrossRef] [PubMed]
- Ke, L.; Xu, S.B.; Wang, J.; Jiang, X.L.; Xu, M.Q. High expression of long non-coding RNA ATB indicates a poor prognosis and regulates cell proliferation and metastasis in non-small cell lung cancer. Clin. Transl. Oncol. 2017, 19, 599–605. [Google Scholar] [CrossRef] [PubMed]
- Xiong, J.; Liu, Y.; Jiang, L.; Zeng, Y.; Tang, W. High expression of long non-coding RNA lncRNA-ATB is correlated with metastases and promotes cell migration and invasion in renal cell carcinoma. Jpn. J. Clin. Oncol. 2016, 46, 378–384. [Google Scholar] [CrossRef] [PubMed]
- Qu, S.; Yang, X.; Song, W.; Sun, W.; Li, X.; Wang, J.; Zhong, Y.; Shang, R.; Ruan, B.; Zhang, Z.; et al. Downregulation of lncRNA-ATB correlates with clinical progression and unfavorable prognosis in pancreatic cancer. Tumour Biol. 2016, 37, 3933–3938. [Google Scholar] [CrossRef] [PubMed]
- Li, A.; Omura, N.; Hong, S.M.; Vincent, A.; Walter, K.; Griffith, M.; Borges, M.; Goggins, M. Pancreatic cancers epigenetically silence SIP1 and hypomethylate and overexpress miR-200a/200b in association with elevated circulating miR-200a and miR-200b levels. Cancer Res. 2010, 70, 5226–5237. [Google Scholar] [CrossRef] [PubMed]
- Panzitt, K.; Tschernatsch, M.M.; Guelly, C.; Moustafa, T.; Stradner, M.; Strohmaier, H.M.; Buck, C.R.; Denk, H.; Schroeder, R.; Trauner, M.; et al. Characterization of HULC, a novel gene with striking up-regulation in hepatocellular carcinoma, as noncoding RNA. Gastroenterology 2007, 132, 330–342. [Google Scholar] [CrossRef] [PubMed]
- Du, Y.; Kong, G.; You, X.; Zhang, S.; Zhang, T.; Gao, Y.; Ye, L.; Zhang, X. Elevation of highly up-regulated in liver cancer (HULC) by hepatitis b virus x protein promotes hepatoma cell proliferation via down-regulating p18. J. Biol. Chem. 2012, 287, 26302–26311. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Li, Z.; Zhong, Q.; Chen, Q.; Zhang, L. Molecular mechanism of heih and HULC in the proliferation and invasion of hepatoma cells. Int. J. Clin. Exp. Med. 2015, 8, 12956–12962. [Google Scholar] [PubMed]
- Zhao, Y.; Guo, Q.; Chen, J.; Hu, J.; Wang, S.; Sun, Y. Role of long non-coding RNA HULC in cell proliferation, apoptosis and tumor metastasis of gastric cancer: A clinical and in vitro investigation. Oncol. Rep. 2014, 31, 358–364. [Google Scholar] [PubMed]
- Subramanyam, D.; Lamouille, S.; Judson, R.L.; Liu, J.Y.; Bucay, N.; Derynck, R.; Blelloch, R. Multiple targets of miR-302 and miR-372 promote reprogramming of human fibroblasts to induced pluripotent stem cells. Nat. Biotechnol. 2011, 29, 443–448. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Li, Q.; Wang, S.; Zhang, J. miR4855p inhibits bladder cancer metastasis by targeting HMGA2. Int. J. Mol. Med. 2015, 36, 1136–1142. [Google Scholar] [PubMed]
- Hu, Y.; Sun, X.; Mao, C.; Guo, G.; Ye, S.; Xu, J.; Zou, R.; Chen, J.; Wang, L.; Duan, P.; et al. Upregulation of long noncoding RNA TUG1 promotes cervical cancer cell proliferation and migration. Cancer Med. 2017, 6, 471–482. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Ding, C.; Yang, Z.; Liu, T.; Zhang, X.; Zhao, C.; Wang, J. The long non-coding RNA TUG1 indicates a poor prognosis for colorectal cancer and promotes metastasis by affecting epithelial-mesenchymal transition. J. Transl. Med. 2016, 14, 42. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Zhao, Z.; Feng, W.; Ye, Z.; Dai, W.; Zhang, C.; Peng, J.; Wu, K. Long non-coding RNA TUG1 promotes colorectal cancer metastasis via EMT pathway. Oncotarget 2016, 7, 51713–51719. [Google Scholar] [CrossRef] [PubMed]
- Fu, J.W.; Kong, Y.; Sun, X. Long noncoding RNA NEAT1 is an unfavorable prognostic factor and regulates migration and invasion in gastric cancer. J. Cancer Res. Clin. Oncol. 2016, 142, 1571–1579. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Zhang, X.; Lin, L.; Chen, G.; Chen, J. Silencing of the long non-coding RNA NEAT1 suppresses ovarian cancer cell proliferation, migration and invasion. Int. J. Clin. Exp. Pathol. 2016, 9, 5138–5147. [Google Scholar]
- Nissan, A.; Stojadinovic, A.; Mitrani-Rosenbaum, S.; Halle, D.; Grinbaum, R.; Roistacher, M.; Bochem, A.; Dayanc, B.E.; Ritter, G.; Gomceli, I.; et al. Colon cancer associated transcript-1: A novel RNA expressed in malignant and pre-malignant human tissues. Int. J. Cancer 2012, 130, 1598–1606. [Google Scholar] [CrossRef] [PubMed]
- Luo, J.; Tang, L.; Zhang, J.; Ni, J.; Zhang, H.P.; Zhang, L.; Xu, J.F.; Zheng, D. Long non-coding RNA Carlo-5 is a negative prognostic factor and exhibits tumor pro-oncogenic activity in non-small cell lung cancer. Tumour Biol. 2014, 35, 11541–11549. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Xie, C.; Zhao, W.; Deng, Z.; Yang, H.; Fang, Q. Long non-coding RNA Carlo-5 expression is associated with disease progression and predicts outcome in hepatocellular carcinoma patients. Clin. Exp. Med. 2017, 17, 33–43. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Ma, M.; Liu, W.; Ding, W.; Yu, H. Enhanced expression of long noncoding RNA Carlo-5 is associated with the development of gastric cancer. Int. J. Clin. Exp. Pathol. 2014, 7, 8471–8479. [Google Scholar] [PubMed]
- Zhao, X.; Wei, X.; Zhao, L.; Shi, L.; Cheng, J.; Kang, S.; Zhang, H.; Zhang, J.; Li, L.; Zhao, W. The rs6983267 SNP and long non-coding RNA Carlo-5 are associated with endometrial carcinoma. Environ. Mol. Mutagen. 2016, 57, 508–515. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Zhou, X.; Chang, H.; Li, H.; Liu, F.; Ma, C.; Lu, J. Ccat1 promotes hepatocellular carcinoma cell proliferation and invasion. Int. J. Clin. Exp. Pathol. 2015, 8, 5427–5434. [Google Scholar] [PubMed]
- Ling, H.; Spizzo, R.; Atlasi, Y.; Nicoloso, M.; Shimizu, M.; Redis, R.S.; Nishida, N.; Gafa, R.; Song, J.; Guo, Z.; et al. Ccat2, a novel noncoding RNA mapping to 8q24, underlies metastatic progression and chromosomal instability in colon cancer. Genome Res. 2013, 23, 1446–1461. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.Y.; Hua, L.; Yao, K.H.; Chen, J.T.; Zhang, J.J.; Hu, J.H. Long non-coding RNA CCAT2 is up-regulated in gastric cancer and associated with poor prognosis. Int. J. Clin. Exp. Pathol. 2015, 8, 779–785. [Google Scholar] [PubMed]
- Chen, S.; Wu, H.; Lv, N.; Wang, H.; Wang, Y.; Tang, Q.; Shao, H.; Sun, C. LncRNA CCAT2 predicts poor prognosis and regulates growth and metastasis in small cell lung cancer. Biomed. Pharmacother. 2016, 82, 583–588. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Liu, L.; Zhu, W. Up-regulation of long non-coding RNA CCAT2 correlates with tumor metastasis and poor prognosis in cervical squamous cell cancer patients. Int. J. Clin. Exp. Pathol. 2015, 8, 13261–13266. [Google Scholar] [PubMed]
- Huang, S.; Qing, C.; Huang, Z.; Zhu, Y. The long non-coding RNA Ccat2 is up-regulated in ovarian cancer and associated with poor prognosis. Diagn. Pathol. 2016, 11, 49. [Google Scholar] [CrossRef] [PubMed]
- Kasagi, Y.; Oki, E.; Ando, K.; Ito, S.; Iguchi, T.; Sugiyama, M.; Nakashima, Y.; Ohgaki, K.; Saeki, H.; Mimori, K.; et al. The expression of Ccat2, a novel long noncoding RNA transcript, and rs6983267 single-nucleotide polymorphism genotypes in colorectal cancers. Oncology 2017, 92, 48–54. [Google Scholar] [CrossRef] [PubMed]
- Qiu, M.; Xu, Y.; Yang, X.; Wang, J.; Hu, J.; Xu, L.; Yin, R. Ccat2 is a lung adenocarcinoma-specific long non-coding RNA and promotes invasion of non-small cell lung cancer. Tumour Biol. 2014, 35, 5375–5380. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.; Jin, L.; Zhang, W.; Zhang, L. Roles of long non-coding RNA CCAT2 in cervical cancer cell growth and apoptosis. Med. Sci. Monit. 2016, 22, 875–879. [Google Scholar] [CrossRef] [PubMed]
- Zeng, J.; Du, T.; Song, Y.; Gao, Y.; Li, F.; Wu, R.; Chen, Y.; Li, W.; Zhou, H.; Yang, Y.; et al. Knockdown of long noncoding RNA CCAT2 inhibits cellular proliferation, invasion, and EMT in glioma cells. Oncol. Res. 2016. [Google Scholar] [CrossRef]
- Zhang, X.; Xu, Y.; He, C.; Guo, X.; Zhang, J.; Zhang, L.; Kong, M.; Chen, B.; Zhu, C. Elevated expression of Ccat2 is associated with poor prognosis in esophageal squamous cell carcinoma. J. Surg. Oncol. 2015, 111, 834–839. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Wang, J.; Wang, S.; Chang, H.; Zhang, T.; Qu, J. LncRNA CCAT2 promotes tumorigenesis by over-expressed pokemon in non-small cell lung cancer. Biomed. Pharmacother. 2017, 87, 692–697. [Google Scholar] [CrossRef] [PubMed]
- Zheng, J.; Zhao, S.; He, X.; Zheng, Z.; Bai, W.; Duan, Y.; Cheng, S.; Wang, J.; Liu, X.; Zhang, G. The up-regulation of long non-coding RNA CCAT2 indicates a poor prognosis for prostate cancer and promotes metastasis by affecting epithelial-mesenchymal transition. Biochem. Biophys. Res. Commun. 2016, 480, 508–514. [Google Scholar] [CrossRef] [PubMed]
- Zhou, N.; Si, Z.; Li, T.; Chen, G.; Zhang, Z.; Qi, H. Long non-coding RNA CCAT2 functions as an oncogene in hepatocellular carcinoma, regulating cellular proliferation, migration and apoptosis. Oncol. Lett. 2016, 12, 132–138. [Google Scholar] [CrossRef] [PubMed]
- Tseng, Y.Y.; Moriarity, B.S.; Gong, W.; Akiyama, R.; Tiwari, A.; Kawakami, H.; Ronning, P.; Reuland, B.; Guenther, K.; Beadnell, T.C.; et al. PVT1 dependence in cancer with MYC copy-number increase. Nature 2014, 512, 82–86. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.; Hu, H.; Li, S. High expression of lncRNA PVT1 promotes invasion by inducing epithelial-to-mesenchymal transition in esophageal cancer. Oncol. Lett. 2016, 12, 2357–2362. [Google Scholar] [CrossRef] [PubMed]
- Wu, B.Q.; Jiang, Y.; Zhu, F.; Sun, D.L.; He, X.Z. Long noncoding RNA PVT1 promotes emt and cell proliferation and migration through downregulating p21 in pancreatic cancer cells. Technol Cancer Res. Treat. 2017. [Google Scholar] [CrossRef] [PubMed]
- Zheng, J.; Yu, F.; Dong, P.; Wu, L.; Zhang, Y.; Hu, Y.; Zheng, L. Long non-coding RNA PVT1 activates hepatic stellate cells through competitively binding microRNA-152. Oncotarget 2016, 7, 62886–62897. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Xie, J.; Shen, C.; Cheng, D.; Shi, Y.; Wu, Z.; Deng, X.; Chen, H.; Shen, B.; Peng, C.; et al. Upregulation of long noncoding RNA Zeb1-AS1 promotes tumor metastasis and predicts poor prognosis in hepatocellular carcinoma. Oncogene 2016, 35, 1575–1584. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.L.; Bai, Y.; Yao, W.J.; Guo, L.; Wang, Z.M. Expression of long non-coding RNA ZEB1-AS1 in esophageal squamous cell carcinoma and its correlation with tumor progression and patient survival. Int. J. Clin. Exp. Pathol. 2015, 8, 11871–11876. [Google Scholar] [PubMed]
- Lv, Q.L.; Hu, L.; Chen, S.H.; Sun, B.; Fu, M.L.; Qin, C.Z.; Qu, Q.; Wang, G.H.; He, C.J.; Zhou, H.H. A long noncoding RNA ZEB1-AS1 promotes tumorigenesis and predicts poor prognosis in glioma. Int. J. Mol. Sci. 2016, 17, 1431. [Google Scholar] [CrossRef] [PubMed]
- Lan, T.; Chang, L.; Wu, L.; Yuan, Y. Downregulation of ZEB2-AS1 decreased tumor growth and metastasis in hepatocellular carcinoma. Mol. Med. Rep. 2016, 14, 4606–4612. [Google Scholar] [CrossRef] [PubMed]
- Dong, L.; Ni, J.; Hu, W.; Yu, C.; Li, H. Upregulation of long non-coding RNA plncRNA-1 promotes metastasis and induces epithelial-mesenchymal transition in hepatocellular carcinoma. Cell Physiol. Biochem. 2016, 38, 836–846. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Cao, R.; Mu, H. Long non-coding RNA UCA1 may be a novel diagnostic and predictive biomarker in plasma for early gastric cancer. Int. J. Clin. Exp. Pathol. 2015, 8, 12936–12942. [Google Scholar] [PubMed]
- Wang, C.M.; Wu, Q.Q.; Li, S.Q.; Chen, F.J.; Tuo, L.; Xie, H.W.; Tong, Y.S.; Ji, L.; Zhou, G.Z.; Cao, G.; et al. Upregulation of the long non-coding RNA plncRNA-1 promotes esophageal squamous carcinoma cell proliferation and correlates with advanced clinical stage. Dig. Dis. Sci. 2014, 59, 591–597. [Google Scholar] [CrossRef] [PubMed]
- Cui, Z.; Ren, S.; Lu, J.; Wang, F.; Xu, W.; Sun, Y.; Wei, M.; Chen, J.; Gao, X.; Xu, C.; et al. The prostate cancer-up-regulated long noncoding RNA plncRNA-1 modulates apoptosis and proliferation through reciprocal regulation of androgen receptor. Urol. Oncol. 2013, 31, 1117–1123. [Google Scholar] [CrossRef] [PubMed]
- Askarian-Amiri, M.E.; Crawford, J.; French, J.D.; Smart, C.E.; Smith, M.A.; Clark, M.B.; Ru, K.; Mercer, T.R.; Thompson, E.R.; Lakhani, S.R.; et al. Snord-host RNA ZFAS1 is a regulator of mammary development and a potential marker for breast cancer. RNA 2011, 17, 878–891. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Wang, F.; Chen, H.; Tan, Q.; Qiu, S.; Chen, S.; Jing, W.; Yu, M.; Liang, C.; Ye, S.; et al. Increased expression of long-noncoding RNA ZFAS1 is associated with epithelial-mesenchymal transition of gastric cancer. Aging 2016, 8, 2023–2038. [Google Scholar] [CrossRef] [PubMed]
- Fang, C.; Zan, J.; Yue, B.; Liu, C.; He, C.; Yan, D. Long noncoding RNA ZFAS1 promotes the progression of colonic cancer by modulating ZEB1 expression. J. Gastroenterol. Hepatol. 2016. [Google Scholar] [CrossRef] [PubMed]
- Wei, W.; Pelechano, V.; Jarvelin, A.I.; Steinmetz, L.M. Functional consequences of bidirectional promoters. Trends Genet. 2011, 27, 267–276. [Google Scholar] [CrossRef] [PubMed]
- Kanduri, C. Long noncoding RNAs: Lessons from genomic imprinting. Biochim. Biophys. Acta 2016, 1859, 102–111. [Google Scholar] [CrossRef] [PubMed]
- Tsai, M.C.; Manor, O.; Wan, Y.; Mosammaparast, N.; Wang, J.K.; Lan, F.; Shi, Y.; Segal, E.; Chang, H.Y. Long noncoding RNA as modular scaffold of histone modification complexes. Science 2010, 329, 689–693. [Google Scholar] [CrossRef] [PubMed]
- Rinn, J.L.; Kertesz, M.; Wang, J.K.; Squazzo, S.L.; Xu, X.; Brugmann, S.A.; Goodnough, L.H.; Helms, J.A.; Farnham, P.J.; Segal, E.; et al. Functional demarcation of active and silent chromatin domains in human hox loci by noncoding RNAs. Cell 2007, 129, 1311–1323. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Zhang, L.; Wang, Y.; Li, H.; Ren, X.; Wei, F.; Yu, W.; Wang, X.; Yu, J.; Hao, X. Long noncoding RNA hotair involvement in cancer. Tumour Biol. 2014, 35, 9531–9538. [Google Scholar] [CrossRef] [PubMed]
- Yao, Y.; Li, J.; Wang, L. Large intervening non-coding RNA hotair is an indicator of poor prognosis and a therapeutic target in human cancers. Int. J. Mol. Sci. 2014, 15, 18985–18999. [Google Scholar] [CrossRef] [PubMed]
- Berrondo, C.; Flax, J.; Kucherov, V.; Siebert, A.; Osinski, T.; Rosenberg, A.; Fucile, C.; Richheimer, S.; Beckham, C.J. Expression of the long non-coding RNA hotair correlates with disease progression in bladder cancer and is contained in bladder cancer patient urinary exosomes. PLoS ONE 2016, 11, e0147236. [Google Scholar] [CrossRef] [PubMed]
- Peng, L.; Yuan, X.; Jiang, B.; Tang, Z.; Li, G.C. LncRNAs: Key players and novel insights into cervical cancer. Tumour Biol. 2016, 37, 2779–2788. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.J.; Lee, D.W.; Yim, G.W.; Nam, E.J.; Kim, S.; Kim, S.W.; Kim, Y.T. Long non-coding RNA hotair is associated with human cervical cancer progression. Int. J. Oncol. 2015, 46, 521–530. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.H.; Wang, X.L.; Tang, H.M.; Jiang, T.; Chen, J.; Lu, S.; Qiu, G.Q.; Peng, Z.H.; Yan, D.W. Long non-coding RNA hotair is a powerful predictor of metastasis and poor prognosis and is associated with epithelial-mesenchymal transition in colon cancer. Oncol. Rep. 2014, 32, 395–402. [Google Scholar] [CrossRef] [PubMed]
- Qiu, J.J.; Lin, Y.Y.; Ye, L.C.; Ding, J.X.; Feng, W.W.; Jin, H.Y.; Zhang, Y.; Li, Q.; Hua, K.Q. Overexpression of long non-coding RNA hotair predicts poor patient prognosis and promotes tumor metastasis in epithelial ovarian cancer. Gynecol. Oncol. 2014, 134, 121–128. [Google Scholar] [CrossRef] [PubMed]
- Ren, D.; Wang, M.; Guo, W.; Huang, S.; Wang, Z.; Zhao, X.; Du, H.; Song, L.; Peng, X. Double-negative feedback loop between ZEB2 and miR-145 regulates epithelial-mesenchymal transition and stem cell properties in prostate cancer cells. Cell Tissue Res. 2014, 358, 763–778. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.; Zeng, H.; Chen, W.; Zheng, R.; Zhang, Y.; Li, Z.; Qi, J.; Wang, M.; Chen, T.; Lou, J.; et al. Helicobacter pylori infection, h19 and linc00152 expression in serum and risk of gastric cancer in a chinese population. Cancer Epidemiol. 2016, 44, 147–153. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Liu, Y.; Zhang, W.; Zhou, Z.; Wu, J.; Cui, P.; Zhang, Y.; Huang, G. Long non-coding RNA linc00152 is involved in cell cycle arrest, apoptosis, epithelial to mesenchymal transition, cell migration and invasion in gastric cancer. Cell Cycle 2015, 14, 3112–3123. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Tan, C.; Weng, W.W.; Deng, Y.; Zhang, Q.Y.; Yang, X.Q.; Gan, H.L.; Wang, T.; Zhang, P.P.; Xu, M.D.; et al. Long non-coding RNA linc00152 is a positive prognostic factor for and demonstrates malignant biological behavior in clear cell renal cell carcinoma. Am. J. Cancer Res. 2016, 6, 285–299. [Google Scholar] [PubMed]
- Muller, S.; Raulefs, S.; Bruns, P.; Afonso-Grunz, F.; Plotner, A.; Thermann, R.; Jager, C.; Schlitter, A.M.; Kong, B.; Regel, I.; et al. Next-generation sequencing reveals novel differentially regulated mRNAs, lncRNAs, mirnas, sdRNAs and a piRNA in pancreatic cancer. Mol. Cancer 2015, 14, 94. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.H.; Fu, J.; Zhang, Z.J.; Ge, C.C.; Yi, Y. LncRNA-linc00152 down-regulated by miR-376c-3p restricts viability and promotes apoptosis of colorectal cancer cells. Am. J. Transl. Res. 2016, 8, 5286–5297. [Google Scholar] [PubMed]
- Li, Y.; Huang, S.; He, K.; Zhao, M.; Lin, H.; Li, D.; Qian, J.; Zhou, C.; Chen, Y.; Huang, C. Long non-coding RNA growth arrest specific transcript 5 acts as a tumour suppressor in colorectal cancer by inhibiting interleukin-10 and vascular endothelial growth factor expression. Oncotarget 2017, 8, 13690–13702. [Google Scholar] [CrossRef] [PubMed]
- Renganathan, A.; Kresoja-Rakic, J.; Echeverry, N.; Ziltener, G.; Vrugt, B.; Opitz, I.; Stahel, R.A.; Felley-Bosco, E. GAS5 long non-coding RNA in malignant pleural mesothelioma. Mol. Cancer 2014, 13, 119. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Li, Z. Long non-coding RNA growth arrest-specific transcript 5 in tumor biology. Oncol. Lett. 2015, 10, 1953–1958. [Google Scholar] [PubMed]
- Chang, L.; Li, C.; Lan, T.; Wu, L.; Yuan, Y.; Liu, Q.; Liu, Z. Decreased expression of long non-coding RNA GAS5 indicates a poor prognosis and promotes cell proliferation and invasion in hepatocellular carcinoma by regulating vimentin. Mol. Med. Rep. 2016, 13, 1541–1550. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Lyu, H.; Liu, H.; Shi, X.; Song, Y.; Liu, B. Downregulation of the long noncoding RNA GAS5-AS1 contributes to tumor metastasis in non-small cell lung cancer. Sci. Rep. 2016, 6, 31093. [Google Scholar] [CrossRef] [PubMed]
- Gong, X.; Wei, W.; Chen, L.; Xia, Z.; Yu, C. Comprehensive analysis of long non-coding RNA expression profiles in hepatitis b virus-related hepatocellular carcinoma. Oncotarget 2016, 7, 42422–42430. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.L.; Ren, T.Y.; Cao, S.W.; Zheng, S.H.; Hu, X.M.; Hu, Y.W.; Lin, L.; Chen, J.; Zheng, L.; Wang, Q. Hbx-related long non-coding RNA DBH-AS1 promotes cell proliferation and survival by activating mapk signaling in hepatocellular carcinoma. Oncotarget 2015, 6, 33791–33804. [Google Scholar] [PubMed]
- Zhang, Q.; Matsuura, K.; Kleiner, D.E.; Zamboni, F.; Alter, H.J.; Farci, P. Analysis of long noncoding RNA expression in hepatocellular carcinoma of different viral etiology. J. Transl. Med. 2016, 14, 328. [Google Scholar] [CrossRef] [PubMed]
- Cong, M.; Li, J.; Jing, R.; Li, Z. Long non-coding RNA tumor suppressor candidate 7 functions as a tumor suppressor and inhibits proliferation in osteosarcoma. Tumour Biol. 2016, 37, 9441–9450. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Jin, Y.; Ren, H.; Ma, X.; Wang, B.; Wang, Y. Downregulation of the long non-coding RNA TUSC7 promotes NSCLC cell proliferation and correlates with poor prognosis. Am. J. Transl. Res. 2016, 8, 680–687. [Google Scholar] [PubMed]
- He, A.; Liu, Y.; Chen, Z.; Li, J.; Chen, M.; Liu, L.; Liao, X.; Lv, Z.; Zhan, Y.; Zhuang, C.; et al. Over-expression of long noncoding RNA BANCR inhibits malignant phenotypes of human bladder cancer. J. Exp. Clin. Cancer Res. 2016, 35, 125. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Liu, Y.; Wang, J.; Jie, D.; Yun, T.; Li, W.; Yan, L.; Wang, K.; Feng, J. Downregulated long noncoding RNA BANCR promotes the proliferation of colorectal cancer cells via downregualtion of p21 expression. PLoS ONE 2015, 10, e0122679. [Google Scholar] [CrossRef] [PubMed]
- Sun, M.; Liu, X.H.; Wang, K.M.; Nie, F.Q.; Kong, R.; Yang, J.S.; Xia, R.; Xu, T.P.; Jin, F.Y.; Liu, Z.J.; et al. Downregulation of braf activated non-coding RNA is associated with poor prognosis for non-small cell lung cancer and promotes metastasis by affecting epithelial-mesenchymal transition. Mol. Cancer 2014, 13, 68. [Google Scholar] [CrossRef] [PubMed]
- Flockhart, R.J.; Webster, D.E.; Qu, K.; Mascarenhas, N.; Kovalski, J.; Kretz, M.; Khavari, P.A. Brafv600e remodels the melanocyte transcriptome and induces BANCR to regulate melanoma cell migration. Genome Res. 2012, 22, 1006–1014. [Google Scholar] [CrossRef] [PubMed]
- Su, S.; Gao, J.; Wang, T.; Wang, J.; Li, H.; Wang, Z. Long non-coding RNA BANCR regulates growth and metastasis and is associated with poor prognosis in retinoblastoma. Tumour Biol. 2015, 36, 7205–7211. [Google Scholar] [CrossRef] [PubMed]
- Zhou, T.; Gao, Y. Increased expression of lncRNA BANCR and its prognostic significance in human hepatocellular carcinoma. World J. Surg. Oncol. 2016, 14, 8. [Google Scholar] [CrossRef] [PubMed]
- Tang, S.S.; Zheng, B.Y.; Xiong, X.D. LincRNA-p21: Implications in human diseases. Int. J. Mol. Sci. 2015, 16, 18732–18740. [Google Scholar] [CrossRef] [PubMed]
- Castellano, J.J.; Navarro, A.; Vinolas, N.; Marrades, R.M.; Moises, J.; Cordeiro, A.; Saco, A.; Munoz, C.; Fuster, D.; Molins, L.; et al. LincRNA-p21 impacts prognosis in resected non-small cell lung cancer patients through angiogenesis regulation. J. Thorac. Oncol. 2016, 11, 2173–2182. [Google Scholar] [CrossRef] [PubMed]
- Dimitrova, N.; Zamudio, J.R.; Jong, R.M.; Soukup, D.; Resnick, R.; Sarma, K.; Ward, A.J.; Raj, A.; Lee, J.T.; Sharp, P.A.; et al. LincRNA-p21 activates p21 in cis to promote polycomb target gene expression and to enforce the G1/S checkpoint. Mol. Cell 2014, 54, 777–790. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Liu, M.; Zhang, J.; Zhang, R.; Zhang, Q.; Tan, X. Long noncoding RNA CPS1-IT1 suppresses cell proliferation and metastasis in human lung cancer. Oncol. Res. 2017, 25, 373–380. [Google Scholar]
- Wang, Y.; Xue, K.; Guan, Y.; Jin, Y.; Liu, S.; Wang, L.; Han, L. Long non-coding RNA linc00261 suppresses cell proliferation and invasion and promotes cell apoptosis in human choriocarcinoma. Oncol. Res. 2016. [Google Scholar] [CrossRef] [PubMed]
- Cao, W.J.; Wu, H.L.; He, B.S.; Zhang, Y.S.; Zhang, Z.Y. Analysis of long non-coding RNA expression profiles in gastric cancer. World J. Gastroenterol. 2013, 19, 3658–3664. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Wang, Y.F.; Su, H.F.; Fang, N.; Zou, C.; Li, W.F.; Fei, Z.H. Decreased expression of the long noncoding RNA linc00261 indicate poor prognosis in gastric cancer and suppress gastric cancer metastasis by affecting the epithelial-mesenchymal transition. J. Hematol. Oncol. 2016, 9, 57. [Google Scholar] [CrossRef] [PubMed]
- Cao, D.; Ding, Q.; Yu, W.; Gao, M.; Wang, Y. Long noncoding RNA SPRY4-IT1 promotes malignant development of colorectal cancer by targeting epithelial-mesenchymal transition. Onco Targets Ther. 2016, 9, 5417–5425. [Google Scholar] [PubMed]
- Cui, F.; Wu, D.; He, X.; Wang, W.; Xi, J.; Wang, M. Long noncoding RNA SPRY4-IT1 promotes esophageal squamous cell carcinoma cell proliferation, invasion, and epithelial-mesenchymal transition. Tumour Biol. 2016, 37, 10871–10876. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Lv, Z.; Guo, E. Knockdown of long noncoding RNA SPRY4-IT1 suppresses glioma cell proliferation, metastasis and epithelial-mesenchymal transition. Int. J. Clin. Exp. Pathol. 2015, 8, 9140–9146. [Google Scholar] [PubMed]
- Shen, F.; Cai, W.S.; Feng, Z.; Chen, J.W.; Feng, J.H.; Liu, Q.C.; Fang, Y.P.; Li, K.P.; Xiao, H.Q.; Cao, J.; et al. Long non-coding RNA SPRY4-IT1 pormotes colorectal cancer metastasis by regulate epithelial-mesenchymal transition. Oncotarget 2017, 8, 14479–14486. [Google Scholar] [CrossRef] [PubMed]
- Xie, H.W.; Wu, Q.Q.; Zhu, B.; Chen, F.J.; Ji, L.; Li, S.Q.; Wang, C.M.; Tong, Y.S.; Tuo, L.; Wu, M.; et al. Long noncoding RNA SPRY4-IT1 is upregulated in esophageal squamous cell carcinoma and associated with poor prognosis. Tumour Biol. 2014, 35, 7743–7754. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.L.; Zhao, Z.H.; Xu, W.C.; Hou, J.Q.; Du, X.Y. Increased expression of SPRY4-IT1 predicts poor prognosis and promotes tumor growth and metastasis in bladder cancer. Int. J. Clin. Exp. Pathol. 2015, 8, 1954–1960. [Google Scholar] [PubMed]
- Sun, M.; Liu, X.H.; Lu, K.H.; Nie, F.Q.; Xia, R.; Kong, R.; Yang, J.S.; Xu, T.P.; Liu, Y.W.; Zou, Y.F.; et al. EZH2-mediated epigenetic suppression of long noncoding RNA SPRY4-IT1 promotes NSCLC cell proliferation and metastasis by affecting the epithelial-mesenchymal transition. Cell Death Dis. 2014, 5, e1298. [Google Scholar] [CrossRef] [PubMed]
- Tennis, M.A.; Van Scoyk, M.M.; Freeman, S.V.; Vandervest, K.M.; Nemenoff, R.A.; Winn, R.A. Sprouty-4 inhibits transformed cell growth, migration and invasion, and epithelial-mesenchymal transition, and is regulated by Wnt7a through ppargamma in non-small cell lung cancer. Mol. Cancer Res. 2010, 8, 833–843. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Zhu, N.; Chen, X. A novel long noncoding RNA linc01133 is upregulated in lung squamous cell cancer and predicts survival. Tumour Biol. 2015, 36, 7465–7471. [Google Scholar] [CrossRef] [PubMed]
- Peng, W.; Wu, G.; Fan, H.; Wu, J.; Feng, J. Long noncoding RNA SPRY4-IT1 predicts poor patient prognosis and promotes tumorigenesis in gastric cancer. Tumour Biol. 2015, 36, 6751–6758. [Google Scholar] [CrossRef] [PubMed]
- Xie, M.; Nie, F.Q.; Sun, M.; Xia, R.; Liu, Y.W.; Zhou, P.; De, W.; Liu, X.H. Decreased long noncoding RNA SPRY4-IT1 contributing to gastric cancer cell metastasis partly via affecting epithelial-mesenchymal transition. J. Transl. Med. 2015, 13, 250. [Google Scholar] [CrossRef] [PubMed]
- Moustakas, A.; Heldin, C.H. Signaling networks guiding epithelial-mesenchymal transitions during embryogenesis and cancer progression. Cancer Sci. 2007, 98, 1512–1520. [Google Scholar] [CrossRef] [PubMed]
- Thiery, J.P.; Sleeman, J.P. Complex networks orchestrate epithelial-mesenchymal transitions. Nat. Rev. Mol. Cell Biol. 2006, 7, 131–142. [Google Scholar] [CrossRef] [PubMed]
- Moustakas, A.; Heldin, C.H. Mechanisms of TGFβ-induced epithelial-mesenchymal transition. J. Clin. Med. 2016. [Google Scholar] [CrossRef] [PubMed]
- Ikushima, H.; Miyazono, K. TGFβ signalling: A complex web in cancer progression. Nat. Rev. Cancer 2010, 10, 415–424. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Shao, N.; Ding, X.; Tan, B.; Song, Q.; Wang, N.; Jia, Y.; Ling, H.; Cheng, Y. Crosstalk between transforming growth factor-beta signaling pathway and long non-coding RNAs in cancer. Cancer Lett. 2016, 370, 296–301. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.Y.; Bai, W.D.; Li, C.; Zheng, Z.; Guan, H.; Liu, J.Q.; Yang, X.K.; Han, S.C.; Gao, J.X.; Wang, H.T.; et al. Knockdown of lncRNA-ATB suppresses autocrine secretion of tgf-beta2 by targeting znf217 via mir-200c in keloid fibroblasts. Sci. Rep. 2016, 6, 24728. [Google Scholar] [CrossRef] [PubMed]
- Iempridee, T. Long non-coding RNA h19 enhances cell proliferation and anchorage-independent growth of cervical cancer cell lines. Exp. Biol. Med. 2017, 242, 184–193. [Google Scholar] [CrossRef] [PubMed]
- Terashima, M.; Tange, S.; Ishimura, A.; Suzuki, T. MEG3 long noncoding RNA contributes to the epigenetic regulation of epithelial-mesenchymal transition in lung cancer cell lines. J. Biol. Chem. 2017, 292, 82–99. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Qian, J.; Li, X.; Chen, W.; Xu, A.; Zhao, K.; Hua, Y.; Huang, Z.; Zhang, J.; Liang, C.; et al. Long noncoding RNA bx357664 regulates cell proliferation and epithelial-to-mesenchymal transition via inhibition of TGF-β1/p38/HSP27 signaling in renal cell carcinoma. Oncotarget 2016, 7, 81410–81422. [Google Scholar] [PubMed]
- Fodde, R.; Brabletz, T. Wnt/β-catenin signaling in cancer stemness and malignant behavior. Curr. Opin. Cell Biol. 2007, 19, 150–158. [Google Scholar] [CrossRef] [PubMed]
- Schmalhofer, O.; Brabletz, S.; Brabletz, T. E-cadherin, β-catenin, and ZEB1 in malignant progression of cancer. Cancer Metastasis Rev. 2009, 28, 151–166. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Tian, X.J.; Xing, J. Signal transduction pathways of emt induced by TGF-β, shh, and wnt and their crosstalks. J. Clin. Med. 2016, 5, 41. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.; Lu, Z.; Hay, E.D. Direct evidence for a role of β-catenin/LEF-1 signaling pathway in induction of emt. Cell Biol. Int. 2002, 26, 463–476. [Google Scholar] [CrossRef] [PubMed]
- Vassallo, I.; Zinn, P.; Lai, M.; Rajakannu, P.; Hamou, M.F.; Hegi, M.E. Wif1 re-expression in glioblastoma inhibits migration through attenuation of non-canonical Wnt signaling by downregulating the lncRNA MALAT1. Oncogene 2016, 35, 12–21. [Google Scholar] [CrossRef] [PubMed]
- Jiao, J.; Zhao, X.; Liang, Y.; Tang, D.; Pan, C. FGF1-FGFR1 axis promotes tongue squamous cell carcinoma (TSCC) metastasis through epithelial-mesenchymal transition (EMT). Biochem. Biophys. Res. Commun. 2015, 466, 327–332. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, P.T.; Tsunematsu, T.; Yanagisawa, S.; Kudo, Y.; Miyauchi, M.; Kamata, N.; Takata, T. The FGFR1 inhibitor pd173074 induces mesenchymal-epithelial transition through the transcription factor AP-1. Br. J. Cancer 2013, 109, 2248–2258. [Google Scholar] [CrossRef] [PubMed]
- Tomlinson, D.C.; Baxter, E.W.; Loadman, P.M.; Hull, M.A.; Knowles, M.A. FGFR1-induced epithelial to mesenchymal transition through MAPK/PLCGAMMA/COX-2-mediated mechanisms. PLoS ONE 2012, 7, e38972. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Yang, L.; Hu, X.; Jiang, Y.; Hu, Y.; Liu, Z.; Liu, J.; Wen, T.; Ma, Y.; An, G.; et al. Upregulated NNT-AS1, a long noncoding RNA, contributes to proliferation and migration of colorectal cancer cells in vitro and in vivo. Oncotarget 2017, 8, 3441–3453. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Wu, Z.; Wu, T.; Huang, Y.; Cheng, Z.; Li, X.; Sun, T.; Xie, X.; Zhou, Y.; Du, Z. Tumor-suppressive function of long noncoding RNA MALAT1 in glioma cells by downregulation of MMP2 and inactivation of ERK/MAPK signaling. Cell Death Dis. 2016, 7, e2123. [Google Scholar] [CrossRef] [PubMed]
- Peng, W.; Fan, H. Long noncoding RNA CCHE1 indicates a poor prognosis of hepatocellular carcinoma and promotes carcinogenesis via activation of the ERK/MAPK pathway. Biomed. Pharmacother. 2016, 83, 450–455. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [PubMed]
- Liberti, M.V.; Locasale, J.W. The warburg effect: How does it benefit cancer cells? Trends Biochem. Sci. 2016, 41, 211–218. [Google Scholar] [CrossRef] [PubMed]
- Choudhry, H.; Harris, A.L.; McIntyre, A. The tumour hypoxia induced non-coding transcriptome. Mol. Aspects Med. 2016, 47, 35–53. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.N.; Zhang, K.; Hu, Z.M.; Qi, H.X.; Shi, Z.M.; Han, X.H.; Han, Y.W.; Hong, W. Hypoxia-regulated lncRNAs in cancer. Gene 2016, 575, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Higgins, D.F.; Kimura, K.; Bernhardt, W.M.; Shrimanker, N.; Akai, Y.; Hohenstein, B.; Saito, Y.; Johnson, R.S.; Kretzler, M.; Cohen, C.D.; et al. Hypoxia promotes fibrogenesis in vivo via HIF-1 stimulation of epithelial-to-mesenchymal transition. J. Clin. Investig. 2007, 117, 3810–3820. [Google Scholar] [CrossRef] [PubMed]
- Jiang, J.; Tang, Y.L.; Liang, X.H. Emt: A new vision of hypoxia promoting cancer progression. Cancer Biol. Ther. 2011, 11, 714–723. [Google Scholar] [CrossRef] [PubMed]
- Xue, M.; Li, X.; Li, Z.; Chen, W. Urothelial carcinoma associated 1 is a hypoxia-inducible factor-1alpha-targeted long noncoding RNA that enhances hypoxic bladder cancer cell proliferation, migration, and invasion. Tumour Biol. 2014, 35, 6901–6912. [Google Scholar] [CrossRef] [PubMed]
- Puisieux, A.; Brabletz, T.; Caramel, J. Oncogenic roles of EMT-inducing transcription factors. Nat. Cell Biol. 2014, 16, 488–494. [Google Scholar] [CrossRef] [PubMed]
- Engelmann, D.; Putzer, B.M. Emerging from the shade of p53 mutants: N-terminally truncated variants of the p53 family in emt signaling and cancer progression. Sci. Signal. 2014, 7. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.P.; Wang, W.L.; Chang, Y.L.; Wu, C.T.; Chao, Y.C.; Kao, S.H.; Yuan, A.; Lin, C.W.; Yang, S.C.; Chan, W.K.; et al. P53 controls cancer cell invasion by inducing the MDM2-mediated degradation of SLUG. Nat. Cell Biol. 2009, 11, 694–704. [Google Scholar] [CrossRef] [PubMed]
- Shiota, M.; Izumi, H.; Onitsuka, T.; Miyamoto, N.; Kashiwagi, E.; Kidani, A.; Hirano, G.; Takahashi, M.; Naito, S.; Kohno, K. Twist and p53 reciprocally regulate target genes via direct interaction. Oncogene 2008, 27, 5543–5553. [Google Scholar] [CrossRef] [PubMed]
- Barsotti, A.M.; Beckerman, R.; Laptenko, O.; Huppi, K.; Caplen, N.J.; Prives, C. P53-dependent induction of PVT1 and miR-1204. J. Biol. Chem. 2012, 287, 2509–2519. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.; Kim, H.J.; Kim, S.W.; Park, S.A.; Chun, K.H.; Cho, N.H.; Song, Y.S.; Kim, Y.T. The long non-coding RNA hotair increases tumour growth and invasion in cervical cancer by targeting the notch pathway. Oncotarget 2016, 7, 44558–44571. [Google Scholar] [CrossRef] [PubMed]
- Pan, L.; Liang, W.; Fu, M.; Huang, Z.H.; Li, X.; Zhang, W.; Zhang, P.; Qian, H.; Jiang, P.C.; Xu, W.R.; et al. Exosomes-mediated transfer of long noncoding RNA ZFAS1 promotes gastric cancer progression. J. Cancer Res. Clin. Oncol. 2017. [Google Scholar] [CrossRef] [PubMed]
- Visvader, J.E.; Lindeman, G.J. Cancer stem cells in solid tumours: Accumulating evidence and unresolved questions. Nat. Rev. Cancer 2008, 8, 755–768. [Google Scholar] [CrossRef] [PubMed]
- MacDonagh, L.; Gray, S.G.; Breen, E.; Cuffe, S.; Finn, S.P.; O’Byrne, K.J.; Barr, M.P. Lung cancer stem cells: The root of resistance. Cancer Lett. 2016, 372, 147–156. [Google Scholar] [CrossRef] [PubMed]
- Dean, M.; Fojo, T.; Bates, S. Tumour stem cells and drug resistance. Nat. Rev. Cancer 2005, 5, 275–284. [Google Scholar] [CrossRef] [PubMed]
- Mladinich, M.; Ruan, D.; Chan, C.H. Tackling cancer stem cells via inhibition of emt transcription factors. Stem Cells Int. 2016, 2016, 5285892. [Google Scholar] [CrossRef] [PubMed]
- Mani, S.A.; Guo, W.; Liao, M.J.; Eaton, E.N.; Ayyanan, A.; Zhou, A.Y.; Brooks, M.; Reinhard, F.; Zhang, C.C.; Shipitsin, M.; et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 2008, 133, 704–715. [Google Scholar] [CrossRef] [PubMed]
- Boyer, L.A.; Lee, T.I.; Cole, M.F.; Johnstone, S.E.; Levine, S.S.; Zucker, J.P.; Guenther, M.G.; Kumar, R.M.; Murray, H.L.; Jenner, R.G.; et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 2005, 122, 947–956. [Google Scholar] [CrossRef] [PubMed]
- Loh, Y.H.; Wu, Q.; Chew, J.L.; Vega, V.B.; Zhang, W.; Chen, X.; Bourque, G.; George, J.; Leong, B.; Liu, J.; et al. The Oct4 and nanog transcription network regulates pluripotency in mouse embryonic stem cells. Nat. Genet. 2006, 38, 431–440. [Google Scholar] [CrossRef] [PubMed]
- Leis, O.; Eguiara, A.; Lopez-Arribillaga, E.; Alberdi, M.J.; Hernandez-Garcia, S.; Elorriaga, K.; Pandiella, A.; Rezola, R.; Martin, A.G. SOX2 expression in breast tumours and activation in breast cancer stem cells. Oncogene 2012, 31, 1354–1365. [Google Scholar] [CrossRef] [PubMed]
- Herreros-Villanueva, M.; Zhang, J.S.; Koenig, A.; Abel, E.V.; Smyrk, T.C.; Bamlet, W.R.; de Narvajas, A.A.; Gomez, T.S.; Simeone, D.M.; Bujanda, L.; et al. SOX2 promotes dedifferentiation and imparts stem cell-like features to pancreatic cancer cells. Oncogenesis 2013, 2, e61. [Google Scholar] [CrossRef] [PubMed]
- Chiou, S.H.; Wang, M.L.; Chou, Y.T.; Chen, C.J.; Hong, C.F.; Hsieh, W.J.; Chang, H.T.; Chen, Y.S.; Lin, T.W.; Hsu, H.S.; et al. Coexpression of Oct4 and nanog enhances malignancy in lung adenocarcinoma by inducing cancer stem cell-like properties and epithelial-mesenchymal transdifferentiation. Cancer Res. 2010, 70, 10433–10444. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Zhou, L.; Wu, T.; Huang, Y.; Cheng, Z.; Li, X.; Sun, T.; Zhou, Y.; Du, Z. Downregulation of lncRNA-MALAT1 affects proliferation and the expression of stemness markers in glioma stem cell line SHG139S. Cell Mol. Neurobiol. 2016, 36, 1097–1107. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Lu, R.; Gu, J.; Chen, Y.; Zhang, X.; Zhang, L.; Wu, H.; Hua, W.; Zeng, J. Aldehyde dehydrogenase 1A1 up-regulates stem cell markers in benzo[a]pyrene-induced malignant transformation of BEAS-2b cells. Environ. Toxicol. Pharmacol. 2016, 45, 241–250. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Liu, F.; Deng, J.; Cai, X.; Han, J.; Liu, Q. Long noncoding RNA ROR regulates proliferation, invasion, and stemness of gastric cancer stem cell. Cell Reprogr. 2016, 18, 319–326. [Google Scholar] [CrossRef] [PubMed]
- Voulgari, A.; Pintzas, A. Epithelial-mesenchymal transition in cancer metastasis: Mechanisms, markers and strategies to overcome drug resistance in the clinic. Biochim. Biophys. Acta 2009, 1796, 75–90. [Google Scholar] [CrossRef] [PubMed]
- Arumugam, T.; Ramachandran, V.; Fournier, K.F.; Wang, H.; Marquis, L.; Abbruzzese, J.L.; Gallick, G.E.; Logsdon, C.D.; McConkey, D.J.; Choi, W. Epithelial to mesenchymal transition contributes to drug resistance in pancreatic cancer. Cancer Res. 2009, 69, 5820–5828. [Google Scholar] [CrossRef] [PubMed]
- Byers, L.A.; Diao, L.; Wang, J.; Saintigny, P.; Girard, L.; Peyton, M.; Shen, L.; Fan, Y.; Giri, U.; Tumula, P.K.; et al. An epithelial-mesenchymal transition gene signature predicts resistance to EGFR and PI3K inhibitors and identifies axl as a therapeutic target for overcoming egfr inhibitor resistance. Clin. Cancer Res. 2013, 19, 279–290. [Google Scholar] [CrossRef] [PubMed]
- Hollier, B.G.; Evans, K.; Mani, S.A. The epithelial-to-mesenchymal transition and cancer stem cells: A coalition against cancer therapies. J. Mammary Gland Biol. Neoplasia 2009, 14, 29–43. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.; Settleman, J. EMT, cancer stem cells and drug resistance: An emerging axis of evil in the WAR on cancer. Oncogene 2010, 29, 4741–4751. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.Y.; Li, X.Q.; Gao, T.H.; Cui, Y.; Ma, N.; Zhou, Y.; Zhang, G.J. Elevated hotair expression associated with cisplatin resistance in non-small cell lung cancer patients. J. Thorac. Dis. 2016, 8, 3314–3322. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Zhou, L.; Wu, L.M.; Lai, M.C.; Xie, H.Y.; Zhang, F.; Zheng, S.S. Overexpression of long non-coding RNA hotair predicts tumor recurrence in hepatocellular carcinoma patients following liver transplantation. Ann. Surg. Oncol. 2011, 18, 1243–1250. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Li, S.; Chen, Z.; Wang, J.; Chen, Y.; Xu, Z.; Jin, M.; Yu, W. miR-326 reverses chemoresistance in human lung adenocarcinoma cells by targeting specificity protein 1. Tumour Biol. 2016, 37, 13287–13294. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.; Dang, Y.; Liu, S.; Zhang, Y.; Zhang, G. LncRNA hotair promotes cisplatin resistance in gastric cancer by targeting miR-126 to activate the PI3K/AKT/MRP1 genes. Tumour Biol. 2016, 37, 16345–16355. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.L.; Wang, W.W.; Zhu, W.G.; Yu, C.H.; Tao, G.Z.; Wu, Q.Q.; Song, Y.Q.; Pan, P.; Tong, Y.S. High expression of long non-coding RNA AFAP1-AS1 predicts chemoradioresistance and poor prognosis in patients with esophageal squamous cell carcinoma treated with definitive chemoradiotherapy. Mol. Carcinog. 2016, 55, 2095–2105. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Cheng, N.; Li, X.; Pan, H.; Li, C.; Ren, S.; Su, C.; Cai, W.; Zhao, C.; Zhang, L.; et al. Correlation of long non-coding RNA h19 expression with cisplatin-resistance and clinical outcome in lung adenocarcinoma. Oncotarget 2017, 8, 2558–2567. [Google Scholar] [CrossRef] [PubMed]
- Yin, X.; Zheng, S.S.; Zhang, L.; Xie, X.Y.; Wang, Y.; Zhang, B.H.; Wu, W.; Qiu, S.; Ren, Z.G. Identification of long noncoding RNA expression profile in oxaliplatin-resistant hepatocellular carcinoma cells. Gene 2017, 596, 53–88. [Google Scholar] [CrossRef] [PubMed]
- Damas, N.D.; Marcatti, M.; Come, C.; Christensen, L.L.; Nielsen, M.M.; Baumgartner, R.; Gylling, H.M.; Maglieri, G.; Rundsten, C.F.; Seemann, S.E.; et al. Snhg5 promotes colorectal cancer cell survival by counteracting STAU1-mediated mRNA destabilization. Nat. Commun. 2016, 7, 13875. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Zhang, X.; Wang, H.; Wang, L.; Liu, T.; Du, L.; Yang, Y.; Wang, C. MALAT1 is associated with poor response to oxaliplatin-based chemotherapy in colorectal cancer patients and promotes chemoresistance through ezh2. Mol. Cancer Ther. 2017. [Google Scholar] [CrossRef] [PubMed]
- Gao, H.; Song, X.; Kang, T.; Yan, B.; Feng, L.; Gao, L.; Ai, L.; Liu, X.; Yu, J.; Li, H. Long noncoding RNA crnde functions as a competing endogenous RNA to promote metastasis and oxaliplatin resistance by sponging miR-136 in colorectal cancer. Onco Targets Ther. 2017, 10, 205–216. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.M.; Liu, Y.; Wei, H.Y.; Lv, K.Z.; Fu, P. Linc-ROR induces epithelial-mesenchymal transition and contributes to drug resistance and invasion of breast cancer cells. Tumour Biol. 2016, 37, 10861–10870. [Google Scholar] [CrossRef] [PubMed]
- Krell, J.; Frampton, A.E.; Mirnezami, R.; Harding, V.; De Giorgio, A.; Roca Alonso, L.; Cohen, P.; Ottaviani, S.; Colombo, T.; Jacob, J.; et al. Growth arrest-specific transcript 5 associated snoRNA levels are related to p53 expression and DNA damage in colorectal cancer. PLoS ONE 2014, 9, e98561. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Sun, S.; Yu, W.; Jiang, J.; Zhuo, F.; Qiu, G.; Xu, S.; Jiang, X. Altered expression of long non-coding RNAs during genotoxic stress-induced cell death in human glioma cells. J. Neurooncol. 2015, 122, 283–292. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Yan, I.K.; Kogure, T.; Haga, H.; Patel, T. Extracellular vesicle-mediated transfer of long non-coding RNA ror modulates chemosensitivity in human hepatocellular cancer. FEBS Open Biol. 2014, 4, 458–467. [Google Scholar] [CrossRef] [PubMed]
- Lukong, K.E. Understanding breast cancer—The long and winding road. BBA Clin. 2017, 7, 64–77. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.; Luo, J. Long non-coding RNA (lncRNA) urothelial carcinoma-associated 1 (UCA1) enhances tamoxifen resistance in breast cancer cells via inhibiting mtor signaling pathway. Med. Sci. Monit. 2016, 22, 3860–3867. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.Y.; Liang, F.; Zhang, J.W.; Wang, F.; Wang, L.; Kang, X.G. Effects of long noncoding RNA-ROR on tamoxifen resistance of breast cancer cells by regulating microRNA-205. Cancer Chemother. Pharmacol. 2017, 79, 327–337. [Google Scholar] [CrossRef] [PubMed]
- Xue, X.; Yang, Y.A.; Zhang, A.; Fong, K.W.; Kim, J.; Song, B.; Li, S.; Zhao, J.C.; Yu, J. LncRNA hotair enhances ER signaling and confers tamoxifen resistance in breast cancer. Oncogene 2016, 35, 2746–2755. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Zhai, L.; Wang, H.; Liu, C.; Zhang, J.; Chen, W.; Wei, Q. Downregulation of lncRNA GAS5 causes trastuzumab resistance in breast cancer. Oncotarget 2016, 7, 27778–27786. [Google Scholar] [CrossRef] [PubMed]
- Nagini, S. Breast cancer: Current molecular therapeutic targets and new players. Anticancer Agents Med. Chem. 2017, 17, 152–163. [Google Scholar] [CrossRef] [PubMed]
- Barr, S.; Thomson, S.; Buck, E.; Russo, S.; Petti, F.; Sujka-Kwok, I.; Eyzaguirre, A.; Rosenfeld-Franklin, M.; Gibson, N.W.; Miglarese, M.; et al. Bypassing cellular EGF receptor dependence through epithelial-to-mesenchymal-like transitions. Clin. Exp. Metastasis 2008, 25, 685–693. [Google Scholar] [CrossRef] [PubMed]
- Yauch, R.L.; Januario, T.; Eberhard, D.A.; Cavet, G.; Zhu, W.; Fu, L.; Pham, T.Q.; Soriano, R.; Stinson, J.; Seshagiri, S.; et al. Epithelial versus mesenchymal phenotype determines in vitro sensitivity and predicts clinical activity of erlotinib in lung cancer patients. Clin. Cancer Res. 2005, 11, 8686–8698. [Google Scholar] [CrossRef] [PubMed]
- Xue, D.; Zhou, C.; Lu, H.; Xu, R.; Xu, X.; He, X. LncRNA GAS5 inhibits proliferation and progression of prostate cancer by targeting miR-103 through AKT/MTOR signaling pathway. Tumour Biol. 2016, 37, 16187–16197. [Google Scholar] [CrossRef] [PubMed]
- Reya, T.; Morrison, S.J.; Clarke, M.F.; Weissman, I.L. Stem cells, cancer, and cancer stem cells. Nature 2001, 414, 105–111. [Google Scholar] [CrossRef] [PubMed]
- Gezer, U.; Ozgur, E.; Cetinkaya, M.; Isin, M.; Dalay, N. Long non-coding RNAs with low expression levels in cells are enriched in secreted exosomes. Cell Biol. Int. 2014, 38, 1076–1079. [Google Scholar] [CrossRef] [PubMed]
- Conigliaro, A.; Costa, V.; Lo Dico, A.; Saieva, L.; Buccheri, S.; Dieli, F.; Manno, M.; Raccosta, S.; Mancone, C.; Tripodi, M.; et al. Cd90+ liver cancer cells modulate endothelial cell phenotype through the release of exosomes containing h19 lncRNA. Mol. Cancer 2015, 14, 155. [Google Scholar] [CrossRef] [PubMed]
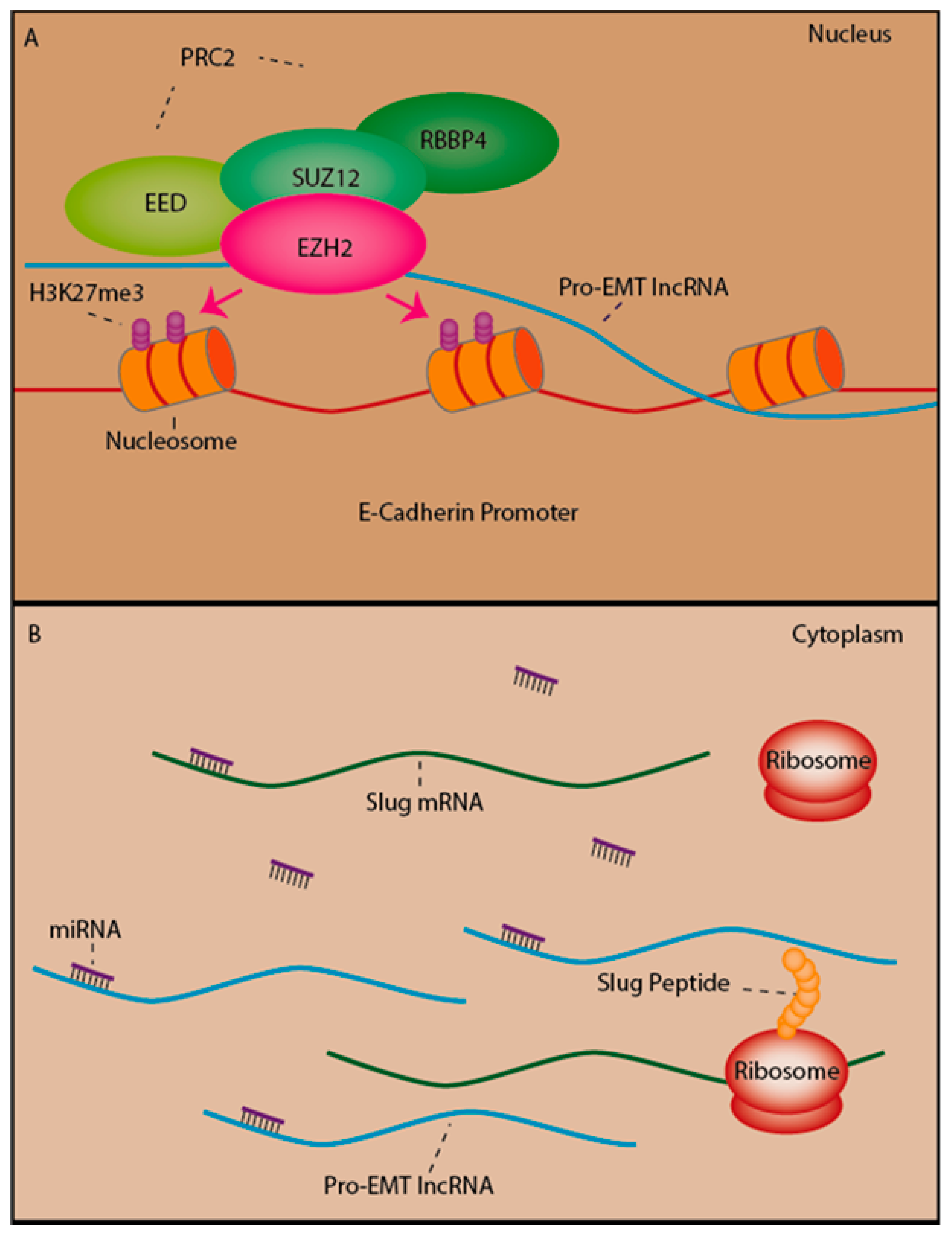
| Pro-EMT LncRNAs | Molecular Mechanisms of Action in EMT | Signalling Pathways Involved | Drug Sensitivity/Resistance | Cancer Stem Cells |
|---|---|---|---|---|
| MALAT-1 (NEAT2) | Acts as ceRNA for miR-1 [89] and miR-204 [90] to derepress Slug; also acts as a ceRNA for miR-205 [91] and predicted to bind miR-200c and miR-145 (in silico analysis) [92]; recruits EZH2 and Suz12 to CDH1 [91,93] | TGF-β [93], Wnt [91,94], MAPK [95], PI3K/Akt [96,97], p53 [98] | Tamoxifen [99], Gemcitabine [100] | Pancreatic Cancer [92] |
| H19 | Acts as a ceRNA for let-7 [101], miR-138 [102], miR-200 family members miR-141 [103] and miR-200a [102] and miR-630 [104] and recruits EZH2 to CDH1 [105] and NKD1 [105,106] | TGF-β [106], Wnt [105], p53 [107] | Cisplatin [108], Doxorubicin [109], Temozolomide [110] | Glioblastoma [111,112], Embryonal Carcinoma [113] Liver Cancer [114] |
| HOTAIR | Recruits PRC2 to CDH1 [115], miR-34 [116], miR-568 [117] and WIF1 [118]; indirectly inhibits miR-7 through suppression of HoxD10 [119], and as part of a SNAIL/EZH2 tripartite complex to EMT genes [120] | TGF-β [121], Wnt [118,122] | Cisplatin [122,123,124] Carboplatin [125] | Breast Cancer [119,121,126], Colon Cancer [121] Colorectal Cancer [127] Liver Cancer [128,129] Lung Cancer [130] Ovarian Cancer [131] |
| Linc-ROR | Acts as a ceRNA for miR-145 [132] and miR-205 [133]; Possibly through interaction with hnRNP 1 to prevent p53 translation [134] Upregulates ZEB1 [135], and prevents degradation of ZEB2 [133] | HIF [136], p53 [134] | 5-FU [137], Paclitaxel [137] Gemcitabine [137,138] | Breast Cancer [133] various cancers [139] |
| lncTCF7 | Possibly through recruitment of Swi/SNF to the TCF7 promoter, triggering TCF7 expression and activating Wnt signalling [140] | Wnt [140], IL-6/STAT3 [141] | - | HCC [140] NSCLC [142] |
| Linc00152 | Acts as a ceRNA for miR-139-5p [143] Binds to EZH2 [144,145] | mTOR [146] EGFR [147] HIF-1α [148] PI3K/Akt [149] | Cisplatin [150] Oxaliplatin [151] | human-induced pluripotent stem cells (hiPSCs) [150] |
| ZEB1-AS1 | Upregulates ZEB1 by acting as ceRNA for miR-200 species [152], or by recruiting histone acetyltransferases to the ZEB1 promoter [153] | - | - | - |
| ZEB2-AS1 | Binds to IRES in ZEB2 mRNA to increase ZEB2 translation [154] | TGF-β [155] | - | - |
| EGFR-AS1 | Unknown | GHR modulates EGFR by regulating EGFRAS1 expression [156] | - | - |
| HNF1A-AS1 | Interacts with DNMT1 to possibly silence CDH1 [157,158,159] Upstream regulator of H19 [160] Acts as a ceRNA for hsa-miR-30b-5p [161] Interacts with EZH2 [162] | Wnt/β-catenin [163] | - | - |
| CBR3-AS1 (PlncRNA-1) | Unknown | TGF-β [164] AR [165] HER2 [166] | - | - |
| LncRNA-ATB | Acts as ceRNA for miR-200 family [167,168,169,170] Acts as a ceRNA for mir-141-3p [171] Suppresses E-Cadherin expression [172] | TGF-β [169], PI3K/Akt [173] | Trastuzumab [168,174] | - |
| HULC | Acts as ceRNA for miR-200a [175,176] and miR-372 [177] Binds to YB-1 promoting its release cyclin D1, cyclin E1, and matrix metalloproteinase 3 [178] Binds to and stabilizes Sirt1 Interacts with EZH2 [179] | PI3K/Akt [176] ERK [178] | Cisplatin [180] Oxaliplatin, 5-FU and THP1 [181] | Liver Cancer [182,183] |
| UCA1 (aka CUDR) | Acts as ceRNA for miR-16 [184], miR-145 [185], miR-196a-5p [186], miR-216b [187], and miR-485-5p [188] | Wnt [189,190,191,192,193] MAPK [187], Akt/mTOR [194], HIF-1α [185,195] TGF-β [196] | Adriamycin [197,198] Cisplatin [186,189,197,199] Docetaxol [200] Gefitinib [194] Imatinib [184] Multi-drug resistance [197] Tamoxifen [195,201] | [182,202,203] |
| TUG1 | Acts as ceRNA for miR-145 [204], and mir-300 [205] | TGF-β [205] | platinum-based chemotherapy combined with 5-fluorouracil (FU) or paclitaxel [206] | Glioma [207] |
| NEAT1 | Acts as ceRNA for miR-204 and miR-101 [208,209] | - | Cisplatin [210] multi-drug resistance [211] | Glioma [212,213] Breast Cancer [214] |
| lncRNA-EBIC | Recruits PRC2 to CDH1 [215] | - | - | - |
| CCAT1 (aka CARLo-5) | Interacts with miR-490 [216] Acts as a ceRNA for let-7 [217] Acts as a ceRNA for miR-218-5p [218] Acts as a ceRNA for miR-155 [219] | c-MYC [220,221,222] | Predicts sensitivity to BET inhibitors in colorectal cancer [223] | - |
| CCAT2 | Interacts with EZH2 [224] | CCAT2 has been shown to regulate cancer cell metabolism [225] Wnt [226] TGFβ [227] | Genetic polymorphisms in CCAT2 have been linked to cisplatin resistance [228] | - |
| lncRNA-HIT | Stabilization of ZEB1 protein [229] | TGF-β [230] | - | - |
| lncRNA-HH | Directly targets GAS1 [231] | Hedgehog [231] | - | Breast Cancer [231] |
| BC087858 | Unknown | MAPK [232], PI3K/Akt [232] | Gefitinib [232] | - |
| Linc00617 | Possibly through recruiting hnRNP-K to SOX2 promoter [233] | - | - | Breast Cancer [233] |
| ZFAS1 | - | Notch [234] | - | - |
| Anti-EMT LncRNAs | Molecular Mechanisms of Action | Signalling Pathways Regulated | Drug Resistance | Cancer Stem Cells |
| GAS5 | Unknown | P53 [235,236] Inhibits IGF1R signalling [237] BRCA1 [236] | Adriamycin [238] Cisplatin [239], Docetaxel [240], Doxorubicin [241,242], 5-FU [240], Gefitinib [237] | - |
| GAS5-AS1 | Unknown | - | - | - |
| LncRNA-Dreh | Interacts with vimentin protein and represses vimentin expression [243] | - | - | - |
| TUSC7 | Acts as ceRNA for miR-10a [244] Acts as a ceRNA for miR-211-3p [245] Binds to miR-23b [246] Directly regulates miR-23b [247] | integrin β1 pathway signalling [244] p53 [247,248] | 5-FU [249], Cisplatin [249] | - |
| AOC4P | Promotes ubiquitination and degradation of vimentin protein [250] | - | - | - |
| LEIGC | Unknown | - | 5-FU [251] | - |
| SLC25A25-AS1 | Unknown | MAPK [252] | 5-FU [252], Doxorubicin [252] | - |
| LincRNA-p21 | Unknown | β-catenin [253,254], Notch [255], HIF [256], p53 [257,258] | Methotrexate [259] | CRC [260], Glioma [253] preiPSC to iPSC conversion [261] |
| CPS1-IT1 | Possibly through interaction with Hsp90 and inhibition of its activation of HIF-1α HIF-1α [262] | HIF-1α [262] | - | - |
| Linc00261 | Binds to Slug protein and promotes its degradation [263] | - | - | - |
| LncRNAs with Variable Roles in EMT | Molecular Mechanisms of Action | Signalling Pathways Regulated | Drug Resistance | Cancer Stem Cells |
| SPRY4-IT1 | Promotes EMT by interacting with Snail and regulating its stability [264] Acts as a ceRNA for miR-101-3p, resulting in up-regulation of EZH2 [265] Interacts with EZH2 to epigenetically repress CDH1 expression [266] | TGF-β [267] | - | - |
| Linc01133 | Inhibits EMT by interacting with and inhibiting SRSF6 [268]. Promotes EMT by recruiting EZH2 to CDH1 [269] | - | - | - |
| BANCR | Unknown | MAPK [270,271] ERK [272] NFκB [273] | - | - |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Heery, R.; Finn, S.P.; Cuffe, S.; Gray, S.G. Long Non-Coding RNAs: Key Regulators of Epithelial-Mesenchymal Transition, Tumour Drug Resistance and Cancer Stem Cells. Cancers 2017, 9, 38. https://doi.org/10.3390/cancers9040038
Heery R, Finn SP, Cuffe S, Gray SG. Long Non-Coding RNAs: Key Regulators of Epithelial-Mesenchymal Transition, Tumour Drug Resistance and Cancer Stem Cells. Cancers. 2017; 9(4):38. https://doi.org/10.3390/cancers9040038
Chicago/Turabian StyleHeery, Richard, Stephen P. Finn, Sinead Cuffe, and Steven G. Gray. 2017. "Long Non-Coding RNAs: Key Regulators of Epithelial-Mesenchymal Transition, Tumour Drug Resistance and Cancer Stem Cells" Cancers 9, no. 4: 38. https://doi.org/10.3390/cancers9040038
APA StyleHeery, R., Finn, S. P., Cuffe, S., & Gray, S. G. (2017). Long Non-Coding RNAs: Key Regulators of Epithelial-Mesenchymal Transition, Tumour Drug Resistance and Cancer Stem Cells. Cancers, 9(4), 38. https://doi.org/10.3390/cancers9040038







