Optimizing Molecular Minimal Residual Disease Analysis in Adult Acute Lymphoblastic Leukemia
Abstract
Simple Summary
Abstract
1. Introduction
2. IG/TR Gene Rearrangements and Fusion Gene Transcripts
2.1. IG/TR Gene Rearrangements
2.2. Fusion Gene Transcripts
3. Novel Molecular Techniques for MRD Monitoring
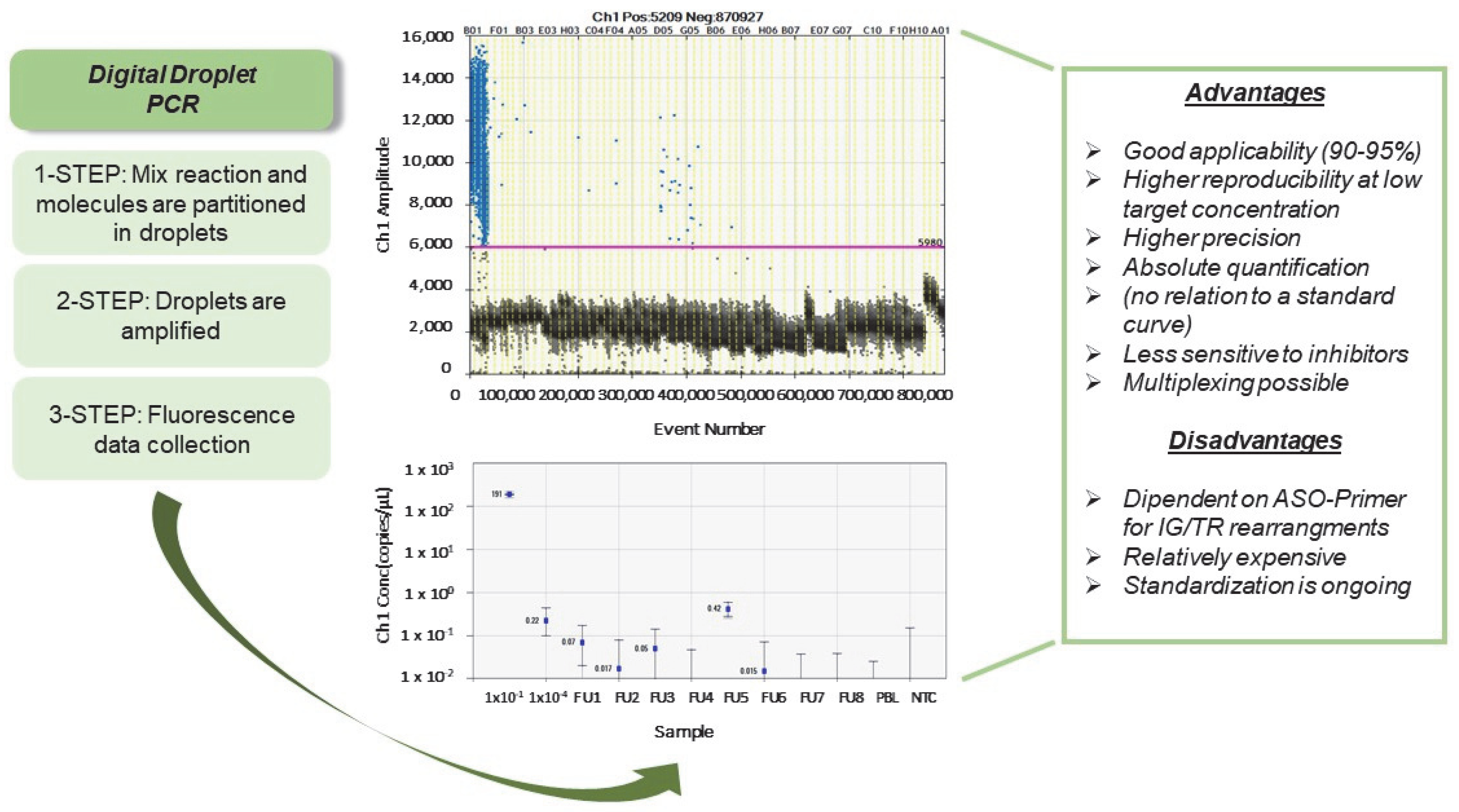
3.1. Digital Droplet PCR (ddPCR)
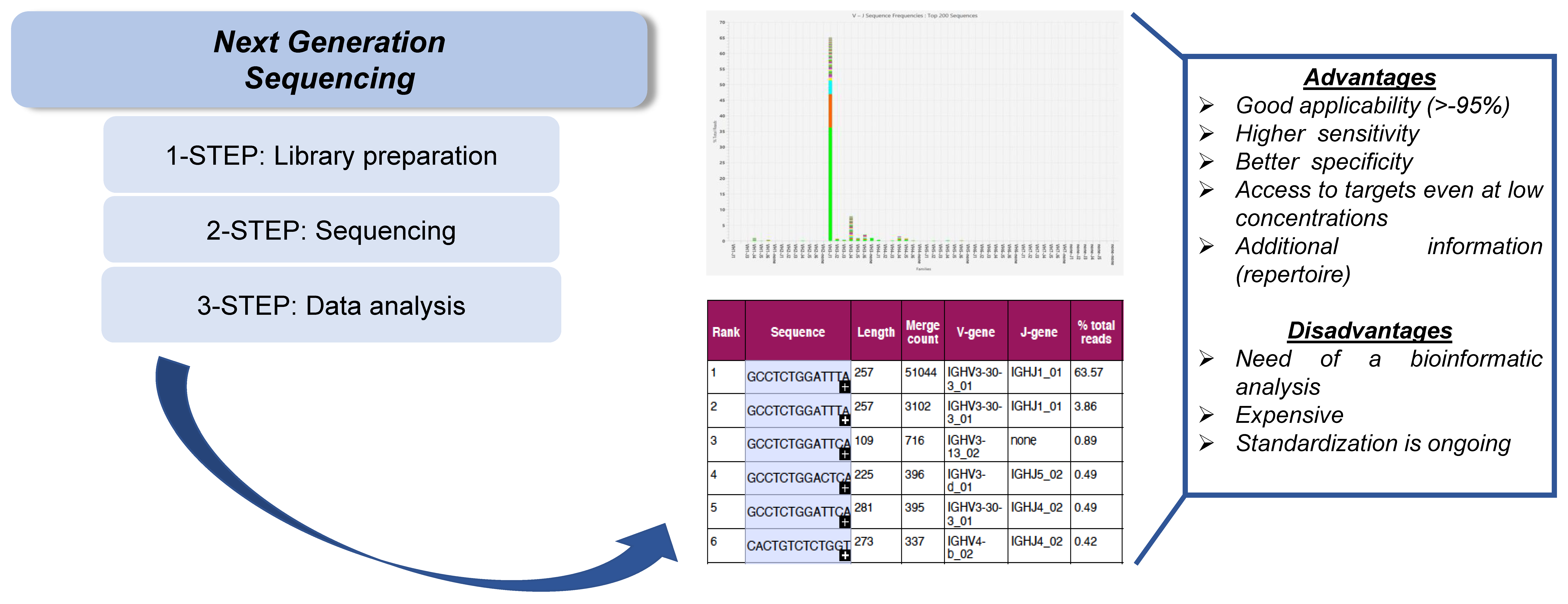
3.2. Next-Generation Sequencing (NGS)
4. Novel Compartments for MRD Monitoring
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Inaba, H.; Greaves, M.; Mullighan, C.G. Acute lymphoblastic leukaemia. Lancet 2013, 381, 1943–1955. [Google Scholar] [CrossRef] [PubMed]
- Faderl, S.; O’Brien, S.; Pui, C.H.; Stock, W.; Wetzler, M.; Hoelzer, D.; Kantarjian, H.M. Adult acute lymphoblastic leukemia: Concepts and strategies. Cancer 2010, 116, 1165–1176. [Google Scholar] [CrossRef] [PubMed]
- Redaelli, A.; Laskin, B.L.; Stephens, J.M.; Botteman, M.F.; Pashos, C.L. A systematic literature review of the clinical and epidemiological burden of acute lymphoblastic leukaemia (ALL). Eur. J. Cancer Care 2005, 14, 53–62. [Google Scholar] [CrossRef]
- Pui, C.H.; Evans, W.E. Treatment of acute lymphoblastic leukemia. N. Engl. J. Med. 2006, 354, 166–178. [Google Scholar] [CrossRef]
- Siegel, R.; Miller, K.; Jemal, A. Cancer statistics 2015. CA Cancer J. Clin. 2015, 65, 5–29. [Google Scholar] [CrossRef] [PubMed]
- Gökbuget, N.; Raff, R.; Brüggemann, M.; Flohr, T.; Scheuring, U.; Pfeifer, H.; Bartram, C.R.; Kneba, M.; Hoelzer, D. Risk/MRD adapted GMALL trials in adult ALL. Ann. Hematol. 2004, 83 (Suppl. S1), S129–S131. [Google Scholar] [PubMed]
- Conter, V.; Bartram, C.R.; Valsecchi, M.G.; Schrauder, A.; Panzer-Grumayer, R.; Moricke, A.; Aricò, M.; Zimmermann, M.; Mann, G.; De Rossi, G.; et al. Molecular response to treatment redefines all prognostic factors in children and adolescents with B-cell precursor acute lymphoblastic leukemia: Results in 3184 patients of the AIEOP-BFM ALL 2000 study. Blood 2010, 115, 3206–3214. [Google Scholar] [CrossRef] [PubMed]
- Stary, J.; Zimmermann, M.; Campbell, M.; Castillo, L.; Dibar, E.; Donska, S.; Gonzalez, A.; Izraeli, S.; Janic, D.; Jazbec, J.; et al. Intensive chemotherapy for childhood acute lymphoblastic leukemia: Results of the randomized intercontinental trial ALL IC-BFM 2002. J. Clin. Oncol. 2014, 32, 174–184. [Google Scholar] [CrossRef] [PubMed]
- Vora, A.; Goulden, N.; Wade, R.; Mitchell, C.; Hancock, J.; Hough, R.; Rowntree, C.; Richards, S. Treatment reduction for children and young adults with low-risk acute lymphoblastic leukaemia defined by minimal residual disease (UKALL 2003): A randomised controlled trial. Lancet Oncol. 2013, 14, 199–209. [Google Scholar] [CrossRef] [PubMed]
- Pieters, R.; de Groot-Kruseman, H.; Van der Velden, V.; Fiocco, M.; van den Berg, H.; de Bont, E.; Maarten Egeler, R.; Hoogerbrugge, P.; Kaspers, G.; Van der Schoot, E.; et al. Successful therapy reduction and intensification for childhood acute lymphoblastic leukemia based on minimal residual disease monitoring: Study ALL10 From the Dutch Childhood Oncology Group. J. Clin. Oncol. 2016, 34, 2591–2601. [Google Scholar] [CrossRef]
- Tzoneva, G.; Perez-Garcia, A.; Carpenter, Z.; Khiabanian, H.; Tosello, V.; Allegretta, M.; Paietta, E.; Racevskis, J.; Rowe, J.M.; Tallman, M.S.; et al. Activating mutations in the NT5C2 nucleotidase gene drive chemotherapy resistance in relapsed ALL. Nat. Med. 2013, 19, 368–371. [Google Scholar] [CrossRef] [PubMed]
- Einsiedel, H.G.; Von Stackelberg, A.; Hartmann, R.; Fengler, R.; Schrappe, M.; Janka-Schaub, G.; Mann, G.; Hählen, K.; Göbel, U.; Klingebiel, T.; et al. Long-term outcome in NGS for MRD Assessment in ALL 489 children with relapsed ALL by risk-stratified salvage therapy: Results of trial Acute Lymphoblastic Leukemia-Relapse Study of the Berlin-Frankfurt-Mu¨nster Group 87. J. Clin. Oncol. 2005, 23, 7942–7950. [Google Scholar] [CrossRef] [PubMed]
- Fielding, A.K.; Richards, S.M.; Chopra, R.; Lazarus, H.M.; Litzow, M.R.; Buck, G.; Durrant, I.J.; Luger, S.M.; Marks, D.I.; Franklin, I.M.; et al. Medical Research Council of the United Kingdom Adult ALL Working Party; Eastern Cooperative Oncology Group. Outcome of 609 adults after relapse of acute lymphoblastic leukemia (ALL); an MRC UKALL12/ECOG 2993 study. Blood 2007, 109, 944–950. [Google Scholar] [CrossRef] [PubMed]
- Gökbuget, N.; Stanze, D.; Beck, J.; Diedrich, H.; Horst, H.A.; Hüttmann, A.; Kobbe, G.; Kreuzer, K.A.; Leimer, L.; Reichle, A.; et al. German Multicenter Study Group for Adult Acute Lymphoblastic Leukemia. Outcome of relapsed adult lymphoblastic leukemia depends on response to salvage chemotherapy, prognostic factors, and performance of stem cell transplantation. Blood 2012, 120, 2032–2041. [Google Scholar] [CrossRef]
- Bassan, R.; Pavoni, C.; Intermesoli, T.; Spinelli, O.; Tosi, M.; Audisio, E.; Marmont, F.; Cattaneo, C.; Borlenghi, E.; Cortelazzo, S.; et al. Updated risk-oriented strategy for acute lymphoblastic leukemia in adult patients 18–65 years: NILG ALL 10/07. Blood Cancer J. 2020, 10, 119. [Google Scholar] [CrossRef]
- Foà, R.; Chiaretti, S. Philadelphia Chromosome-Positive Acute Lymphoblastic Leukemia. N. Engl. J. Med. 2022, 386, 2399–2411. [Google Scholar] [CrossRef]
- Cavé, H.; van der Werff ten Bosch, J.; Suciu, S.; Guidal, C.; Waterkeyn, C.; Otten, J.; Bakkus, M.; Thielemans, K.; Grandchamp, B.; Vilmer, E. European Organization for Research and Treatment of Cancer–Childhood Leukemia Cooperative Group. Clinical significance of minimal residual disease in childhood acute lymphoblastic leukemia. European Organization for Research and Treatment of Cancer—Childhood Leukemia Cooperative Group. N. Engl. J. Med. 1998, 339, 591–598. [Google Scholar]
- Borowitz, M.J.; Devidas, M.; Hunger, S.P.; Bowman, W.P.; Carroll, A.J.; Carroll, W.L.; Linda, S.; Martin, P.L.; Pullen, D.J.; Viswanatha, D.; et al. Children’s Oncology Group. Clinical significance of minimal residual disease in childhood acute lymphoblastic leukemia and its relationship to other prognostic factors: A Children’s Oncology Group study. Blood 2008, 111, 5477–5485. [Google Scholar] [CrossRef]
- Raff, T.; Gökbuget, N.; Lüschen, S.; Reutzel, R.; Ritgen, M.; Irmer, S.; Böttcher, S.; Horst, H.A.; Kneba, M.; Hoelzer, D.; et al. GMALL Study Group. Molecular relapse in adult standard-risk ALL patients detected by prospective MRD monitoring during and after maintenance treatment: Data from the GMALL 06/99 and 07/03 trials. Blood 2007, 109, 910–915. [Google Scholar] [CrossRef]
- Gökbuget, N.; Kneba, M.; Raff, T.; Trautmann, H.; Bartram, C.R.; Arnold, R.; Fietkau, R.; Freund, M.; Ganser, A.; Ludwig, W.D.; et al. German Multicenter Study Group for Adult Acute Lymphoblastic Leukemia. Adult patients with acute lymphoblastic leukemia and molecular failure display a poor prognosis and are candidates for stem cell transplantation and targeted therapies. Blood 2012, 120, 1868–1876. [Google Scholar] [CrossRef]
- van Dongen, J.J.M.; van der Velden, V.H.J.; Brüggemann, M.; Orfao, A. Minimal residual disease diagnostics in acute lymphoblastic leukemia: Need for sensitive, fast, and standardized technologies. Blood 2015, 125, 3996–4009. [Google Scholar] [CrossRef] [PubMed]
- Nyvold, C.G. Critical methodological factors in diagnosing minimal residual disease in hematological malignancies using quantitative PCR. Expert Rev. Mol. Diagn. 2015, 15, 581–584. [Google Scholar] [CrossRef] [PubMed]
- van der Velden, V.H.; Hochhaus, A.; Cazzaniga, G.; Szczepanski, T.; Gabert, J.; van Dongen, J.J. Detection of minimal residual disease in hematologic malignancies by real-time quantitative PCR: Principles, approaches, and laboratory aspects. Leukemia 2003, 17, 1013–1034. [Google Scholar] [CrossRef] [PubMed]
- van der Velden, V.H.; Cazzaniga, G.; Schrauder, A.; Hancock, J.; Bader, P.; Panzer-Grumayer, E.R.; Flohr, T.; Sutton, R.; Cave, H.; Madsen, H.O.; et al. European Study Group on MRD detection in ALL (ESG-MRD-ALL). Analysis of minimal residual disease by Ig/TCR gene rearrangements: Guidelines for interpretation of real-time quantitative PCR data. Leukemia 2007, 21, 604–611. [Google Scholar] [CrossRef] [PubMed]
- Campana, D. Status of minimal residual disease testing in childhood haematological malignancies. Br. J. Haematol. 2008, 143, 481–489. [Google Scholar] [CrossRef]
- Flohr, T.; Schrauder, A.; Cazzaniga, G.; Panzer-Grümayer, R.; van der Velden, V.; Fischer, S.; Stanulla, M.; Basso, G.; Niggli, F.K.; Schäfer, B.W.; et al. International BFM Study Group (I-BFM-SG). Minimal residual disease directed risk stratification using real-time quantitative PCR analysis of immunoglobulin and T-cell receptor gene rearrangements in the international multicenter trial AIEOP-BFM ALL 2000 for childhood acute lymphoblastic leukemia. Leukemia 2008, 22, 771–782. [Google Scholar]
- Dworzak, M.N.; Gaipa, G.; Ratei, R.; Veltroni, M.; Schumich, A.; Maglia, O.; Karawajew, L.; Benetello, A.; Pötschger, U.; Husak, Z.; et al. Standardization of flow cytometric minimal residual disease evaluation in acute lymphoblastic leukemia: Multicentric assessment is feasible. Cytom. B Clin. Cytom. 2008, 74, 331–340. [Google Scholar] [CrossRef]
- Cazzaniga, G.; Valsecchi, M.G.; Gaipa, G.; Conter, V.; Biondi, A. Defining the correct role of minimal residual disease tests in the management of acute lymphoblastic leukaemia. Br. J. Haematol. 2011, 155, 45–52. [Google Scholar] [CrossRef]
- Pfeifer, H.; Cazzaniga, G.; van der Velden, V.H.J.; Cayuela, J.M.; Schäfer, B.; Spinelli, O.; Akiki, S.; Avigad, S.; Bendit, I.; Borg, K.; et al. Standardisation and consensus guidelines for minimal residual disease assessment in Philadelphia-positive acute lymphoblastic leukemia (Ph + ALL) by real-time quantitative reverse transcriptase PCR of e1a2 BCR-ABL1. Leukemia 2019, 33, 1910–1922. [Google Scholar] [CrossRef]
- van der Velden, V.H.J.; van Dongen, J.J.M. MRD detection in acute lymphoblastic leukemia patients using Ig/TCR gene rearrangements as targets for real-time quantitative PCR. Methods Mol. Biol. 2009, 538, 115–150. [Google Scholar]
- Langerak, A.W.; Szczepański, T.; van der Burg, M.; Wolvers-Tettero, I.L.; van Dongen, J.J.M. Heteroduplex PCR analysis of rearranged T cell receptor genes for clonality assessment in suspect T cell proliferations. Leukemia 1997, 11, 2192–2199. [Google Scholar] [CrossRef] [PubMed]
- Germano, G.; del Giudice, L.; Palatron, S.; Giarin, E.; Cazzaniga, G.; Biondi, A.; Basso, G. Clonality profile in relapsed precursor-B-ALL children by GeneScan and sequencing analyses. Consequences on minimal residual disease monitoring. Leukemia 2003, 17, 1573–1582. [Google Scholar] [CrossRef] [PubMed]
- Verhagen, O.J.; Willemse, M.J.; Breunis, W.B.; Wijkhuijs, A.J.; Jacobs, D.C.; Joosten, S.A.; van Wering, E.R.; van Dongen, J.J.; van der Schoot, C.E. Application of germline IGH probes in real-time quantitative PCR for the detection of minimal residual disease in acute lymphoblastic leukemia. Leukemia 2000, 14, 1426–1435. [Google Scholar] [CrossRef] [PubMed]
- Szczepański, T.; van der Velden, V.H.; Raff, T.; Jacobs, D.C.; van Wering, E.R.; Brüggemann, M.; Kneba, M.; van Dongen, J.J.M. Comparative analysis of T-cell receptor gene rearrangements at diagnosis and relapse of T-cell acute lymphoblastic leukemia (T-ALL) shows high stability of clonal markers for monitoring of minimal residual disease and reveals the occurrence of second T-ALL. Leukemia 2003, 17, 2149–2156. [Google Scholar] [CrossRef]
- Eckert, C.; Flohr, T.; Koehler, R.; Hagedorn, N.; Moericke, A.; Stanulla, M.; Kirschner-Schwabe, R.; Cario, G.; Stackelberg, A.; Bartram, C.R.; et al. Very early/early relapses of acute lymphoblastic leukemia show unexpected changes of clonal markers and high heterogeneity in response to initial and relapse treatment. Leukemia 2011, 25, 1305–1313. [Google Scholar] [CrossRef]
- van Dongen, J.J.; Macintyre, E.A.; Gabert, J.A.; Delabesse, E.; Rossi, V.; Saglio, G.; Gottardi, E.; Rambaldi, A.; Dotti, G.; Griesinger, F.; et al. Standardized RT-PCR analysis of fusion gene transcripts from chromosome aberrations in acute leukemia for detection of minimal residual disease. Report of the BIOMED-1 Concerted Action: Investigation of minimal residual disease in acute leukemia. Leukemia 1999, 13, 1901–1928. [Google Scholar] [CrossRef]
- Gabert, J.; Beillard, E.; van der Velden, V.H.; Bi, W.; Grimwade, D.; Pallisgaard, N.; Barbany, G.; Cazzaniga, G.M.; Cayuela, J.M.; Cavé, H.; et al. Standardization and quality control studies of ‘real-time’ quantitative reverse transcriptase polymerase chain reaction of fusion gene transcripts for residual disease detection in leukemia—A Europe against cancer program. Leukemia 2003, 17, 2318–2357. [Google Scholar] [CrossRef]
- Chiaretti, S.; Vitale, A.; Cazzaniga, G.; Orlando, S.M.; Silvestri, D.; Fazi, P.; Valsecchi, M.G.; Elia, L.; Testi, A.M.; Mancini, F.; et al. Clinico-biological features of 5202 patients with acute lymphoblastic leukemia enrolled in the Italian AIEOP and GIMEMA protocols and stratified in age cohorts. Haematologica 2013, 98, 1702–1710. [Google Scholar] [CrossRef]
- D’Angiò, M.; Valsecchi, M.G.; Testi, A.M.; Conter, V.; Nunes, V.; Parasole, R.; Colombini, A.; Santoro, N.; Varotto, S.; Caniglia, M.; et al. Clinical features and outcome of SIL/TAL1-positive T-cell acute lymphoblastic leukemia in children and adolescents: A 10-year experience of the AIEOP group. Haematologica 2015, 100, e10–e13. [Google Scholar] [CrossRef]
- Elia, L.; Mancini, M.; Moleti, L.; Meloni, G.; Buffolino, S.; Krampera, M.; De Rossi, G.; Foà, R.; Cimino, G. A multiplex reverse transcriptase polymerase chain reaction strategy for the diagnostic molecular screening of chimeric genes: A clinical evaluation on 170 patients with acute lymphoblastic leukemia. Haematologica 2003, 88, 275–279. [Google Scholar]
- Campana, D. Determination of minimal residual disease in leukaemia patients. Br. J. Haematol. 2003, 121, 823–838. [Google Scholar] [CrossRef] [PubMed]
- Huggett, J.F.; Whale, A. Digital PCR as a novel technology and its potential implications for molecular diagnostics. Clin. Chem. 2013, 59, 1691–1693. [Google Scholar] [CrossRef] [PubMed]
- Dube, S.; Qin, J.; Ramakrishnan, R. Mathematical analysis of copy number variation in a DNA sample using digital PCR on a nanofluidic device. PLoS ONE 2008, 3, e2876. [Google Scholar] [CrossRef]
- Whale, A.S.; Cowen, S.; Foy, C.A.; Huggett, J.F. Methods for applying accurate digital PCR analysis on low copy DNA samples. PLoS ONE 2013, 8, e58177. [Google Scholar] [CrossRef] [PubMed]
- Williams, R.; Peisajovich, S.G.; Miller, O.J.; Magdassi, S.; Tawfik, D.S.; Griths, A.D. Amplification of complex gene libraries by emulsion PCR. Nat. Methods 2006, 3, 545–550. [Google Scholar] [CrossRef] [PubMed]
- Hindson, B.J.; Ness, K.D.; Masquelier, D.A.; Belgrader, P.; Heredia, N.J.; Makarewicz, A.J.; Bright, I.J.; Lucero, M.Y.; Hiddessen, A.L.; Legler, T.C.; et al. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal. Chem. 2011, 83, 8604–8610. [Google Scholar] [CrossRef] [PubMed]
- Pinheiro, L.B.; Coleman, V.A.; Hindson, C.M.; Herrmann, J.; Hindson, B.J.; Bhat, S.; Emslie, K.R. Evaluation of a droplet digital polymerase chain reaction format for DNA copy number quantification. Anal. Chem. 2012, 84, 1003–1011. [Google Scholar] [CrossRef]
- Sanders, R.; Huggett, J.F.; Bushell, C.A.; Cowen, S.; Scott, D.J.; Foy, C.A. Evaluation of digital PCR for absolute DNA quantification. Anal. Chem. 2011, 83, 6474–6484. [Google Scholar] [CrossRef]
- Vincent, M.E.; Liu, W.; Haney, E.B.; Ismagilov, R.F. Microfluidic stochastic confinement enhances analysis of rare cells by isolating cells and creating high density environments for control of diffusible signals. Chem. Soc. Rev. 2010, 39, 974–984. [Google Scholar] [CrossRef]
- Goh, H.G.; Lin, M.; Fukushima, T.; Saglio, G.; Kim, D.; Choi, S.Y.; Kim, S.H.; Lee, J.; Lee, Y.S.; Oh, S.M.; et al. Sensitive quantitation of minimal residual disease in chronic myeloid leukemia using nanofluidic digital polymerase chain reaction assay. Leuk. Lymphoma 2011, 52, 896–904. [Google Scholar] [CrossRef]
- Alikian, M.; Whale, A.S.; Akiki, S.; Piechocki, K.; Torrado, C.; Myint, T.; Cowen, S.; Griths, M.; Reid, A.G.; Apperley, J.; et al. RT-qPCR and RT-digital PCR: A comparison of dierent platforms for the evaluation of residual disease in chronic myeloid leukemia. Clin. Chem. 2017, 63, 525–531. [Google Scholar] [CrossRef] [PubMed]
- Jennings, L.J.; George, D.; Czech, J.; Yu, M.; Joseph, L. Detection and quantification of BCR-ABL1 fusion transcripts by droplet digital PCR. J. Mol. Diagn. 2014, 16, 174–179. [Google Scholar] [CrossRef] [PubMed]
- Madic, J.; Zoetic, A.; Sengis, V.; Fradet, E.; Andre, B.; Muller, S.; Dangla, R.; Droniou, M.E. Three-color crystal digital PCR. Biomol. Detect. Quantif. 2016, 10, 34–46. [Google Scholar] [CrossRef] [PubMed]
- Low, H.; Chan, S.J.; Soo, G.H.; Ling, B.; Tan, E.L. Clarity TM digital PCR system: A novel platform for absolute quantification of nucleic acids. Anal. Bioanal. Chem. 2017, 409, 1869–1875. [Google Scholar] [CrossRef]
- Whale, A.S.; Huggett, J.F.; Cowen, S.; Speirs, V.; Shaw, J.; Ellison, S.; Foy, C.A.; Scott, D.J. Comparison of microfluidic digital PCR and conventional quantitative PCR for measuring copy number variation. Nucleic Acids Res. 2012, 40, e82. [Google Scholar] [CrossRef]
- Tsui, N.B.; Kadir, R.A.; Chan, K.C.; Chi, C.; Mellars, G.; Tuddenham, E.G.; Leung, T.Y.; Lau, T.K.; Chiu, R.W.K.; Lo, Y.M.D. Non-invasive prenatal diagnosis of hemophilia by microfluidics digital PCR analysis of maternal plasma DNA. Blood 2011, 117, 3684–3691. [Google Scholar] [CrossRef]
- Pornprasert, S.; Prasing, W. Detection of alpha (0)-thalassemia South-East Asian-type deletion by droplet digital PCR. Eur. J. Haematol. 2014, 92, 244–248. [Google Scholar] [CrossRef]
- Stahl, T.; Böhme, M.U.; Kröger, N.; Fehse, B. Digital PCR to assess hematopoietic chimerism after allogeneic stem cell transplantation. Exp. Hematol. 2015, 43, 462–468. [Google Scholar] [CrossRef]
- Zagaria, A.; Anelli, L.; Coccaro, N.; Tota, G.; Casieri, P.; Cellamare, A.; Impera, L.; Brunetti, C.; Minervini, A.; Minervini, C.F.; et al. BCR-ABL1 e6a2 transcript in chronic myeloid leukemia: Biological features and molecular monitoring by droplet digital PCR. Virchows Arch. 2015, 467, 357–363. [Google Scholar] [CrossRef]
- Albano, F.; Zagaria, A.; Anelli, L.; Coccaro, N.; Tota, G.; Brunetti, C.; Minervini, C.F.; Impera, L.; Minervini, A.; Cellamare, A.; et al. Absolute quantification of the pre-treatment PML-RARA transcript defines the relapse risk in acute promyelocytic leukemia. Oncotarget 2015, 6, 13269–13277. [Google Scholar] [CrossRef]
- Coccaro, N.; Anelli, L.; Zagaria, A.; Casieri, P.; Tota, G.; Orsini, P.; Impera, L.; Minervini, A.; Minervini, C.F.; Cumbo, C.; et al. Droplet digital PCR is a robust tool for monitoring minimal residual disease in adult Philadelphia-positive acute lymphoblastic leukemia. J. Mol. Diagn. 2018, 20, 474–482. [Google Scholar] [CrossRef] [PubMed]
- Drandi, D.; Kubiczkova-Besse, L.; Ferrero, S.; Dani, N.; Passera, R.; Mantoan, B.; Gambella, M.; Monitillo, L.; Saraci, F.; Ghione, P.; et al. Minimal residual disease detection by droplet digital PCR in multiple myeloma, mantle cell lymphoma, and follicular lymphoma: A comparison with real-time PCR. J. Mol. Diagn. 2015, 17, 652–660. [Google Scholar] [CrossRef] [PubMed]
- Della Starza, I.; Nunes, V.; Cavalli, M.; De Novi, L.A.; Ilari, C.; Apicella, V.; Vitale, A.; Testi, A.M.; Del Giudice, I.; Chiaretti, S.; et al. Comparative analysis between RQ-PCR and digital-droplet-PCR of immunoglobulin/Tcell receptor gene rearrangements to monitor minimal residual disease in acute lymphoblastic leukaemia. Br. J. Haematol. 2016, 174, 541–549. [Google Scholar] [CrossRef] [PubMed]
- Cavalli, M.; De Novi, L.A.; Della Starza, I.; Cappelli, L.V.; Nunes, V.; Pulsoni, A.; Del Giudice, I.; Guarini, A.; Foa, R. Comparative analysis between RQ-PCR and digital droplet PCR of BCL2/IGH gene rearrangement in the peripheral blood and bone marrow of early-stage follicular lymphoma. Br. J. Haematol. 2017, 177, 588–596. [Google Scholar] [CrossRef] [PubMed]
- Della Starza, I.; Cavalli, M.; De Novi, L.A.; Genuardi, E.; Mantoan, B.; Drandi, D.; Barbero, D.; Ciabatti, E.; Grassi, S.; Gazzola, A.; et al. Minimal residual disease (MRD) in non-Hodgkin lymphomas: Interlaboratory reproducibility on marrow samples with very low levels of disease within the FIL (Fondazione Italiana Linfomi) MRD Network. Hematol. Oncol. 2019, 37, 368–374. [Google Scholar] [CrossRef] [PubMed]
- Drandi, D.; Alcantara, M.; Benmaad, I.; Söhlbrandt, A.; Lhermitte, L.; Zaccaria, G.; Ferrante, M.; Genuardi, E.; Mantoan, B.; Villarese, P.; et al. Droplet digital PCR quantification of mantle cell lymphoma follow-up samples from four prospective trials of the European MCL network. HemaSphere 2020, 4, e347. [Google Scholar] [CrossRef] [PubMed]
- Della Starza, I.; De Novi, L.A.; Santoro, A.; Salemi, D.; Tam, W.; Cavalli, M.; Menale, L.; Soscia, R.; Apicella, V.; Ilari, C.; et al. Digital droplet PCR and next-generation sequencing refine minimal residual disease monitoring in acute lymphoblastic leukemia. Leuk. Lymphoma 2019, 60, 2838–2840. [Google Scholar] [CrossRef]
- Della Starza, I.; De Novi, L.A.; Santoro, A.; Salemi, D.; Spinelli, O.; Tosi, M.; Soscia, R.; Paoloni, F.; Cappelli, L.V.; Cavalli, M.; et al. Digital Droplet PCR Is a Reliable Tool to Improve Minimal Residual Disease Stratification in Adult Philadelphia-Negative Acute Lymphoblastic Leukemia. J. Mol. Diagn. 2022, 24, 893–900. [Google Scholar] [CrossRef]
- Schwinghammer, C.; Koopmann, J.; Chitadze, G.; Karawajew, L.; Brüggemann, M.; Eckert, C. A Droplet Digital PCR.: New View on Minimal Residual Disease Quantification in Acute Lymphoblastic Leukemia. J. Mol. Diagn. 2022, 24, 856–866. [Google Scholar] [CrossRef]
- Iacobucci, I.; Lonetti, A.; Venturi, C.; Ferrari, A.; Papayannidis, C.; Ottaviani, E.; Abbenante, M.C.; Paolini, S.; Bresciani, P.; Potenza, L.; et al. Use of a high sensitive nanofluidic array for the detection of rare copies of BCR-ABL1 transcript in patients with Philadelphia-positive acute lymphoblastic leukemia in complete response. Leuk. Res. 2014, 38, 581–585. [Google Scholar] [CrossRef]
- Foà, R.; Bassan, R.; Vitale, A.; Elia, L.; Piciocchi, A.; Puzzolo, M.C.; Canichella, M.; Viero, P.; Ferrara, F.; Lunghi, M.; et al. Dasatinib-Blinatumomab for Ph-Positive Acute Lymphoblastic Leukemia in Adults. N. Engl. J. Med. 2020, 383, 1613–1623. [Google Scholar] [CrossRef] [PubMed]
- Ansuinelli, M.; Della Starza, I.; Lauretti, A.; Elia, L.; Siravo, V.; Messina, M.; De Novi, L.A.; Taherinasab, A.; Canichella, M.; Guarini, A.; et al. Applicability of droplet digital polymerase chain reaction for minimal residual disease in Philadelphia-positive acute lymphoblastic leukaemia. Hematol. Oncol. 2021, 39, 680–686. [Google Scholar] [CrossRef] [PubMed]
- Hovorkova, L.; Zaliova, M.; Venn, N.C.; Bleckmann, K.; Trkova, M.; Potuckova, E.; Vaskova, M.; Linhartova, J.; Machova Polakova, K.; Fronkova, E.; et al. Monitoring of childhood ALL using BCR-ABL1 genomic breakpoints identifies a subgroup with CML-like biology. Blood 2017, 129, 2771–2781. [Google Scholar] [CrossRef] [PubMed]
- Cazzaniga, G.; De Lorenzo, P.; Alten, J.; Röttgers, S.; Hancock, J.; Saha, V.; Castor, A.; O Madsen, H.; Gandemer, V.; Cavé, H.; et al. Predictive value of minimal residual disease in Philadelphia-chromosome-positive acute lymphoblastic leukemia treated with imatinib in the European intergroup study of post-induction treatment of Philadelphia-chromosome-positive acute lymphoblastic leukemia, based on immunoglobulin/T-cell receptor and BCR/ABL1 methodologies. Haematologica 2018, 103, 107–115. [Google Scholar] [PubMed]
- Clappier, E.; Kim, R.; Cayuela, J.M.; Rousselot, P.; Chalandon, Y.; Passet, M.; Thomas, X.; Havelange, V.; Chevallier, P.; Huguet, F.; et al. Persistent BCR-ABL1 clonal hematopoiesis after blast clearance identifies a CML-like subgroup of patients with Philadelphia chromosome-positive (Ph+) ALL: Interim results from GRAAPH-2014 trial. In Proceedings of the 23rd EHA Annual Congress, Stockolm, Sweden, 14–17 June 2018; p. S1568. [Google Scholar]
- Della Starza, I.; Nunes, V.; Lovisa, V.; Silvestri, D.; Cavalli, M.; Garofalo, A.; Campeggio, M.; De Novi, L.A.; Soscia, R.; Oggioni, C.; et al. Droplet Digital PCR Improves IG-/TR-based MRD Risk Definition in Childhood B-cell Precursor Acute Lymphoblastic Leukemia. Hemasphere 2021, 5, e543. [Google Scholar] [CrossRef] [PubMed]
- Drandi, D.; Ferrero, S.; Ladetto, M. Droplet Digital PCR for Minimal Residual Disease Detection in Mature Lymphoproliferative Disorders. Methods Mol. Biol. 2018, 1768, 229–256. [Google Scholar] [PubMed]
- Whale, A.S.; De Spiegelaere, W.; Trypsteen, W.; Nour, A.A.; Bae, Y.K.; Benes, V.; Burke, D.; Cleveland, M.; Corbisier, P.; Devonshire, A.S.; et al. The Digital MIQE Guidelines Update: Minimum Information for Publication of Quantitative Digital PCR Experiments for 2020. Clin. Chem. 2020, 66, 1012–1029. [Google Scholar]
- Della Starza, I.; Eckert, C.; Drandi, D.; Cazzaniga, G.; on behalf of the EuroMRD Consortium. Minimal residual Disease Analysis by Monitoring Immunoglobulin and T-Cell Receptor Gene rearrangements by Quantitative PCR and Droplet Digital PCR. Methods Mol. Biol. 2022, 2453, 79–89. [Google Scholar]
- van Bladel, D.A.G.; van der Last-Kempkes, J.L.M.; Scheijen, B.; Groenen, P.J.T.A.; on behalf of the EuroClonality Consortium. Next-Generation Sequencing-Based Clonality Detection of Immunoglobulin Gene Rearrangements in B-Cell Lymphoma. Methods Mol. Biol. 2022, 2453, 7–42. [Google Scholar]
- Scheijen, B.; Meijers, R.W.J.; Rijntjes, J.; van der Klift, M.Y.; Möbs, M.; Steinhilber, J.; Reigl, T.; van den Brand, M.; Kotrova, M.; Ritter, J.M.; et al. Next-generation sequencing of immunoglobulin gene rearrangements for clonality assessment: A technical feasibility study by EuroClonality-NGS. Leukemia 2019, 33, 2227–2240. [Google Scholar] [CrossRef]
- Merriman, B.; Rothberg, J.M. Progress in ion torrent semiconductor chip-based sequencing. Electrophoresis 2012, 33, 3397–3417. [Google Scholar] [CrossRef] [PubMed]
- Bentley, D.R.; Balasubramanian, S.; Swerdlow, H.P.; Smith, G.P.; Milton, J.; Brown, C.G.; Hall, K.P.; Evers, D.J.; Barnes, C.L.; Bignell, H.R.; et al. Accurate whole human genome sequencing using reversible terminator chemistry. Nature 2008, 456, 53–59. [Google Scholar] [CrossRef] [PubMed]
- Wu, D.; Sherwood, A.; Fromm, J.R.; Winter, S.S.; Dunsmore, K.P.; Loh, M.L.; Greisman, H.A.; Sabath, D.E.; Wood, B.L.; Robins, H. High-throughput sequencing detects minimal residual disease in acute T lymphoblastic leukemia. Sci. Transl. Med. 2012, 4, 134ra63. [Google Scholar] [CrossRef] [PubMed]
- Kotrova, M.; Muzikova, K.; Mejstrikova, E.; Novakova, M.; Bakardjieva-Mihaylova, V.; Fiser, K.; Stuchly, J.; Giraud, M.; Salson, M.I.; Pott, C.; et al. The predictive strength of next-generation sequencing MRD detection for relapse compared with current methods in childhood ALL. Blood 2015, 126, 1045–1047. [Google Scholar] [CrossRef] [PubMed]
- Ladetto, M.; Brïggemann, M.; Monitillo, L.; Ferrero, S.; Pepin, F.; Drandi, D.; Barbero, D.; Palumbo, A.; Passera, R.; Boccadoro, M.; et al. Next-generation sequencing and real-time quantitative PCR for minimal residual disease detection in B-cell disorders. Leukemia 2014, 28, 1299–1307. [Google Scholar] [CrossRef]
- Logan, A.C.; Zhang, B.; Narasimhan, B.; Carlton, V.; Zheng, J.; Moorhead, M.; Krampf, M.R.; Jones, C.D.; Waqar, A.N.; Faham, M.; et al. Minimal residual disease quantification using consensus primers and high-throughput IGH sequencing predicts post-transplant relapse in chronic lymphocytic leukemia. Leukemia 2013, 27, 1659–1665. [Google Scholar] [CrossRef] [PubMed]
- van Dongen, J.J.M.; Seriu, T.; Panzer-Grumayer, E.R.; Biondi, A.; Pongers-Willemse, M.J.; Corral, L.; Stolz, F.; Schrappe, M.; Masera, G.; Kamps, W.A.; et al. Prognostic value of minimal residual disease in acute lymphoblastic leukaemia in childhood. Lancet 1998, 352, 1731–1738. [Google Scholar] [CrossRef]
- Gawad, C.; Pepin, F.; Carlton, V.E.H.; Klinger, M.; Logan, A.C.; Miklos, D.B.; Faham, M.; Dahl, G.; Lacayo, N. Massive evolution of the immunoglobulin heavy chain locus in children with B precursor acute lymphoblastic leukemia. Blood 2012, 120, 4407–4417. [Google Scholar] [CrossRef]
- Kitchingman, G.R. Immunoglobulin heavy chain gene VH-D junctional diversity at diagnosis in patients with acute lymphoblastic leukemia. Blood 1993, 81, 775–782. [Google Scholar] [CrossRef]
- de Haas, V.; Verhagen, O.J.; von dem Borne, A.E.; Kroes, W.; van den Berg, H.; van der Schoot, C.E. Quantification of minimal residual disease in children with oligoclonal B-precursor acute lymphoblastic leukemia indicates that the clones that grow out during relapse already have the slowest rate of reduction during induction therapy. Leukemia 2001, 15, 134–140. [Google Scholar] [CrossRef]
- Theunissen, P.M.J.; van Zessen, D.; Stubbs, A.P.; Faham, M.; Zwaan, C.M.; van Dongen, J.J.M.; Van Der Velden, V.H.J. Antigen receptor sequencing of paired bone marrow samples shows homogeneous distribution of acute lymphoblastic leukemia subclones. Haematologica 2017, 102, 1869–1877. [Google Scholar] [CrossRef] [PubMed]
- Bashford-Rogers, R.J.M.; Nicolaou, K.A.; Bartram, J.; Goulden, N.J.; Loizou, L.; Koumas, L.; Chi, J.; Hubank, M.; Kellam, P.; Costeas, P.A.; et al. Eye on the B-ALL: B-cell receptor repertoires reveal persistence of numerous B-lymphoblastic leukemia subclones from diagnosis to relapse. Leukemia 2016, 30, 2312–2321. [Google Scholar] [CrossRef] [PubMed]
- Salson, M.; Giraud, M.; Caillault, A.; Grardel, N.; Duployez, N.; Ferret, Y.; Duez, M.; Herbert, R.; Rocher, T.; Sebda, S.; et al. High-throughput sequencing in acute lymphoblastic leukemia: Follow-up of minimal residual disease and emergence of new clones. Leuk. Res. 2017, 53, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Theunissen, P.M.J.; de Bie, M.; van Zessen, D.; de Haas, V.; Stubbs, A.P.; van der Velden, V.H.J. Next-generation antigen receptor sequencing of paired diagnosis and relapse samples of B-cell acute lymphoblastic leukemia: Clonal evolution and implications for minimal residual disease target selection. Leuk. Res. 2019, 76, 98–104. [Google Scholar] [CrossRef] [PubMed]
- Kotrova, M.; van der Velden, V.H.J.; van Dongen, J.J.M.; Formankova, R.; Sedlacek, P.; Brüggemann, M.; Zuna, J.; Stary, J.; Trka, J.; Fronkova, E. Next-generation sequencing indicates false-positive MRD results and better predicts prognosis after SCT in patients with childhood ALL. Bone Marrow Transpl. 2017, 52, 962–968. [Google Scholar] [CrossRef]
- Bruggemann, M.; Kotrova, M.; Knecht, H.; Bartram, J.; Boudjogrha, M.; Bystry, V.; Fazio, G.; Froňková, E.; Giraud, M.; Grioni, A.; et al. Standardized next-generation sequencing of immunoglobulin and T-cell receptor gene recombinations for MRD marker identification in acute lymphoblastic leukaemia; a EuroClonality-NGS validation study. Leukemia 2019, 33, 2241–2253. [Google Scholar] [CrossRef]
- Knecht, H.; Reigl, T.; Kotrova, M.; Appelt, F.; Stewart, P.; Bystry, V.; Krejci, A.; Grioni, A.; Pal, K.; Stranska, K.; et al. Quality control and quantification in IG/TR next-generation sequencing marker identification: Protocols and bioinformatic functionalities by EuroClonality-NGS. Leukemia 2019, 33, 2254–2265. [Google Scholar] [CrossRef]
- Stewart, J.P.; Gazdova, J.; Darzentas, N.; Wren, D.; Proszek, P.; Fazio, G.; Songia, S.; Alcoceba, M.; Sarasquete, M.E.; Villarese, P.; et al. Validation of the EuroClonality-NGS DNA capture panel as an integrated genomic tool for lymphoproliferative disorders. Blood Adv. 2021, 5, 3188–3198. [Google Scholar]
- Bystry, V.; Reigl, T.; Krejci, A.; Demko, M.; Hanakova, B.; Grioni, A.; Knecht, H.; Schlitt, M.; Dreger, P.; Sellner, L.; et al. ARResT/Interrogate: An interactive immunoprofiler for IG/TR NGS data. Bioinformatics 2017, 33, 435–437. [Google Scholar] [CrossRef]
- ClonoSEQ Cleared for Residual Cancer Testing. Cancer Discov. 2018, 8, OF6. [CrossRef]
- Lay, L.; Stroup, B.; Payton, J.E. Validation and interpretation of IGH and TCR clonality testing by Ion Torrent S5 NGS for diagnosis and disease monitoring in B and T cell cancers. Pract. Lab. Med. 2020, 22, e00191. [Google Scholar] [CrossRef] [PubMed]
- Ho, C.C.; Tungy, J.K.; Zehnder, J.L.; Zhangy, B.M. Validation of a Next-Generation Sequencing Based T-Cell Receptor Gamma Gene Rearrangement Diagnostic Assay Transitioning from Capillary Electrophoresis to Next-Generation Sequencing. J. Mol. Diagn. 2021, 23, 805–815. [Google Scholar] [CrossRef] [PubMed]
- Ha, J.; Lee, H.; Shin, S.; Cho, H.; Chung, H.; Jang, J.E.; Kim, S.J.; Cheong, J.W.; Lee, S.T.; Kim, J.S.; et al. Ig Gene Clonality Analysis Using Next Generation Sequencing for Improved Minimal Residual Disease Detection with Significant Prognostic Value in Multiple Myeloma Patients. J. Mol. Diagn. 2022, 24, 48–56. [Google Scholar] [CrossRef] [PubMed]
- Arcila, M.E.; Yu, W.; Syed, M.; Kim, H.; Maciag, L.; Yao, J.J.; Ho, C.; Petrova, K.; Moung, C.; Salazar, P.; et al. Establishment of Immunoglobulin Heavy (IGH) Chain Clonality Testing by Next-Generation Sequencing for Routine Characterization of B-Cell and Plasma Cell Neoplasms. J. Mol. Diagn. 2019, 21, 330–342. [Google Scholar] [CrossRef]
- Takamatsu, H. Comparison of Minimal Residual Disease Detection by Multiparameter Flow Cytometry, ASO-qPCR, Droplet Digital PCR, and Deep Sequencing in Patients with Multiple Myeloma Who Underwent Autologous Stem Cell Transplantation. J. Clin. Med. 2017, 6, 91. [Google Scholar] [CrossRef]
- Pedini, P.; Cherouat, N.; Basire, A.; Simon, S.; Budon, L.; Pourtein, M.; Grondin, S.; Moskovtchenko, P.; Chiaroni, J.; Michel, G.; et al. Evaluation of Next-Generation Sequencing and Crystal Digital PCR for Chimerism Monitoring of Post-Allogeneic Hematopoietic Stem Cell Transplantation. Transplant. Cell. Ther. 2021, 27, 89.e1–89.e10. [Google Scholar] [CrossRef]
- Brüggemann, M.; Kotrova, M. Minimal residual disease in adult ALL: Technical aspects and implications for correct clinical interpretation. Blood Adv. 2017, 1, 2456–2466. [Google Scholar] [CrossRef]
- Coustan-Smith, E.; Sancho, J.; Hancock, M.L.; Razzouk, B.I.; Ribeiro, C.R.; Rivera, G.K.; Rubnitz, J.E.; Sandlund, J.T.; Pui, C.H.; Campana, D. Use of peripheral blood instead of bone marrow to monitor residual disease in children with acute lymphoblastic leukemia. Blood 2002, 100, 2399–2402. [Google Scholar] [CrossRef]
- Alix-Panabières, C.; Schwarzenbach, H.; Pantel, K. Circulating Tumor Cells and Circulating Tumor DNA. Annu. Rev. Med. 2012, 63, 199–215. [Google Scholar] [CrossRef]
- Siravegna, G.; Marsoni, S.; Siena, S.; Bardelli, A. Integrating Liquid Biopsies into the Management of Cancer. Nat. Rev. Clin. Oncol. 2017, 14, 531–548. [Google Scholar] [CrossRef]
- Gorgannezhad, L.; Umer, M.; Islam, M.N.; Nguyen, N.T.; Shiddiky, M.J.A. Circulating tumor DNA and liquid biopsy: Opportunities, challenges, and recent advances in detection technologies. Lab Chip 2018, 18, 1174–1196. [Google Scholar] [CrossRef] [PubMed]
- Salehi, M.; Sharifi, M. Exosomal MiRNAs as Novel Cancer Biomarkers: Challenges and Opportunities. J. Cell. Physiol. 2018, 233, 6370–6380. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Bode, A.M.; Dong, Z. Circulating Tumor Cells: Moving Biological Insights into Detection. Theranostics 2017, 7, 2606–2619. [Google Scholar] [CrossRef] [PubMed]
- Becker, A.; Thakur, B.K.; Weiss, J.M.; Kim, H.S.; Peinado, H.; Lyden, D. Extracellular Vesicles in Cancer: Cell-to-Cell Mediators of Metastasis. Cancer Cell 2016, 30, 836–848. [Google Scholar] [CrossRef] [PubMed]
- Buedts, L.; Vandenberghe, P. Circulating cell-free DNA in hematological malignancies. Haematologica 2016, 101, 997–999. [Google Scholar] [CrossRef] [PubMed]
- Rossi, D.; Diop, F.; Spaccarotella, E.; Monti, S.; Zanni, M.; Rasi, S.; Deambrogi, C.; Spina, V.; Bruscaggin, A.; Favini, C.; et al. Diffuse large B-cell lymphoma genotyping on the liquid biopsy. Blood 2017, 129, 1947–1957. [Google Scholar] [CrossRef] [PubMed]
- Lim, J.K.; Kuss, B.; Talaulikar, D. Role of cell-free DNA in haematological malignancies. Pathology 2021, 53, 416–426. [Google Scholar] [CrossRef]
- Bingham, N.; Spencer, A. The role of cell free DNA and liquid biopsies in haematological conditions. Cancer Drug Resist. 2020, 3, 521–531. [Google Scholar] [CrossRef]
- Colmenares, R.; Álvarez, N.; Barrio, S.; Martinez-López, J.; Ayala, R. The Minimal Residual Disease Using Liquid Biopsies in Hematological Malignancies. Cancers 2022, 14, 1310. [Google Scholar] [CrossRef]
- Schwarz, A.K.; Stanulla, M.; Cario, G.; Flohr, T.; Sutton, R.; Möricke, A.; Anker, P.; Stroun, M.; Welte, K.; Bartram, C.R.; et al. Quantification of Free Total Plasma DNA and Minimal Residual Disease Detection in the Plasma of Children with Acute Lymphoblastic Leukemia. Ann. Hematol. 2009, 88, 897–905. [Google Scholar] [CrossRef]
- Cheng, S.H.; Lau, K.M.; Li, C.K.; Chan, N.P.H.; Ip, R.K.L.; Cheng, C.K.; Lee, V.; Shing, M.M.K.; Leung, A.W.K.; Ha, S.Y.; et al. Minimal Residual Disease-Based Risk Stratification in Chinese Childhood Acute Lymphoblastic Leukemia by Flow Cytometry and Plasma DNA Quantitative Polymerase Chain Reaction. PLoS ONE 2013, 8, e69467. [Google Scholar] [CrossRef] [PubMed]
- Ikeuchi, K.; Doki, N.; Takao, A.; Hishima, T.; Yamada, Y.; Konishi, T.; Nagata, A.; Takezaki, T.; Kaito, S.; Kurosawa, S.; et al. Extramedullary Gastric Relapse at the Time of Bone Marrow Relapse of Acute Lymphoblastic Leukemia after Allogeneic Bone Marrow Transplantation. Intern. Med. 2017, 56, 3215–3217. [Google Scholar] [CrossRef] [PubMed]
- Aldoss, I.; Song, J.Y. Extramedullary relapse of KMT2A(MLL)-rearranged acute lymphoblastic leukemia with lineage switch following blinatumomab. Blood 2018, 131, 2507. [Google Scholar] [CrossRef] [PubMed]
- Demosthenous, C.; Lalayanni, C.; Iskas, M.; Douka, V.; Pastelli, N.; Anagnostopoulos, A. Extramedullary relapse and discordant CD19 expression between bone marrow and extramedullary sites in relapsed acute lymphoblastic leukemia after blinatumomab treatment. Curr. Probl. Cancer 2019, 43, 222–227. [Google Scholar] [CrossRef] [PubMed]
- Bronkhorst, A.J.; Aucamp, J.; Pretorius, P.J. Cell-free DNA: Preanalytical variables. Clin. Chim. Acta 2015, 450, 243–253. [Google Scholar] [CrossRef] [PubMed]
- Cavallone, L.; Aldamry, M.; Lafleur, J.; Lan, C.; Gonzalez Ginestet, P.; Alirezaie, N.; Ferrario, C.; Aguilar-Mahecha, A.; Basik, M. A Study of Pre-Analytical Variables and Optimization of Extraction Method for Circulating Tumor DNA Measurements by Digital Droplet PCR. Cancer Epidemiol. Biomark. Prev. 2019, 28, 909–916. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Della Starza, I.; De Novi, L.A.; Elia, L.; Bellomarino, V.; Beldinanzi, M.; Soscia, R.; Cardinali, D.; Chiaretti, S.; Guarini, A.; Foà, R. Optimizing Molecular Minimal Residual Disease Analysis in Adult Acute Lymphoblastic Leukemia. Cancers 2023, 15, 374. https://doi.org/10.3390/cancers15020374
Della Starza I, De Novi LA, Elia L, Bellomarino V, Beldinanzi M, Soscia R, Cardinali D, Chiaretti S, Guarini A, Foà R. Optimizing Molecular Minimal Residual Disease Analysis in Adult Acute Lymphoblastic Leukemia. Cancers. 2023; 15(2):374. https://doi.org/10.3390/cancers15020374
Chicago/Turabian StyleDella Starza, Irene, Lucia Anna De Novi, Loredana Elia, Vittorio Bellomarino, Marco Beldinanzi, Roberta Soscia, Deborah Cardinali, Sabina Chiaretti, Anna Guarini, and Robin Foà. 2023. "Optimizing Molecular Minimal Residual Disease Analysis in Adult Acute Lymphoblastic Leukemia" Cancers 15, no. 2: 374. https://doi.org/10.3390/cancers15020374
APA StyleDella Starza, I., De Novi, L. A., Elia, L., Bellomarino, V., Beldinanzi, M., Soscia, R., Cardinali, D., Chiaretti, S., Guarini, A., & Foà, R. (2023). Optimizing Molecular Minimal Residual Disease Analysis in Adult Acute Lymphoblastic Leukemia. Cancers, 15(2), 374. https://doi.org/10.3390/cancers15020374




